Abstract
Background:
Recent efforts to improve outcomes for high-grade serous ovarian cancer, a leading cause of cancer death in women, have focused on identifying molecular subtypes and prognostic gene signatures, but existing subtypes have poor cross-study robustness. We tested the contribution of cell admixture in published ovarian cancer molecular subtypes and prognostic gene signatures.
Methods:
Gene signatures of tumor and stroma were developed using paired microdissected tissue from two independent studies. Stromal genes were investigated in two molecular subtype classifications and 61 published gene signatures. Prognostic performance of gene signatures of stromal admixture was evaluated in 2,527 ovarian tumors (16 studies). Computational simulations of increasing stromal cell proportion were performed by mixing gene expression profiles of paired microdissected ovarian tumor and stroma.
Results:
Recently described ovarian cancer molecular subtypes are strongly associated with the cell admixture. Tumors were classified as different molecular subtypes in simulations where the percentage of stromal cells increased. Stromal gene expression in bulk tumors was associated with overall survival (hazard ratio 1.17, 95% confidence interval 1.11–1.23), and in one dataset, increased stroma was associated with anatomic sampling location. Five published prognostic gene signatures were no longer prognostic in a multivariate model that adjusted for stromal content.
Conclusions:
Cell admixture impacts the interpretation and reproduction of ovarian cancer molecular subtypes and gene signatures derived from bulk tissue. Elucidating the role of stroma in the tumor microenvironment and in prognosis is important.
Impact:
Single-cell analyses may be required to refine the molecular subtypes of high-grade serous ovarian cancer.
Keywords: Ovarian cancer, molecular diagnosis and prognosis, inflammation and tumor development, functional genomics, microarrays, gene expression profiling
Introduction
There have been several attempts to identify molecular subtypes and prognostic genomic signatures for advanced-stage serous ovarian cancers (1–4). The first large study to report molecular subtypes studied mostly serous tumors and identified a subtype that was associated with poor overall survival, C1 (1). Subsequently, The Cancer Genome Atlas (TCGA) study reported four similar subtypes in high-grade serous ovarian cancer (HGSOC) but found no difference in clinical outcome between the subtypes (2), which led researchers to question the clinical utility of these classifications (5). Subtype predictions using the TCGA classification are not consistently robust, sometimes producing overlapping clusters, with most tumors assigned to more than one subtype (6,7). A recent comprehensive meta-analysis could not robustly classify subtypes across datasets (8), leading to alternative subtype classifications that were associated with survival, tumor purity, age, and lymphocyte infiltration (8).
Molecular subtype and biomarker discovery in HGSOC has mostly been performed on gene expression profiles of bulk tumors that may include fibroblasts, immune cells, and endothelial cells (9). The gene expression profile of a bulk tumor is the mean of the admixture of malignant epithelial, stromal, and other cells, but gene expression studies of bulk tumors rarely record or adjust for tumor purity, which can be highly variable within and between studies. Stromal content is variable in two of the largest gene expression profiling studies (1,2). In the Australian Ovarian Cancer Study (AOCS), 40% of tumors in the molecular subtype C1 had low tumor percentage (≤ 50%) compared with 9% in the other molecular subtypes combined (1). In the TCGA study, the proportion of stromal cells exceeded 50% in about one percent of tumors, and the median proportion of stromal cells was 10% (2).
The relative proportions of tumor and stromal cells may also be an independent prognostic indicator in several epithelial cancers, including breast (10), colon (11), and possibly ovarian cancer (12). Cells in the tumor microenvironment are important for chemo-sensitivity (13), and genes expressed by cells in the tumor microenvironment are prognostic in ovarian cancer (4,14–16). For example, activation of Smad signaling in cancer-associated fibroblasts in the tumor microenvironment is associated poor patient survival (16).
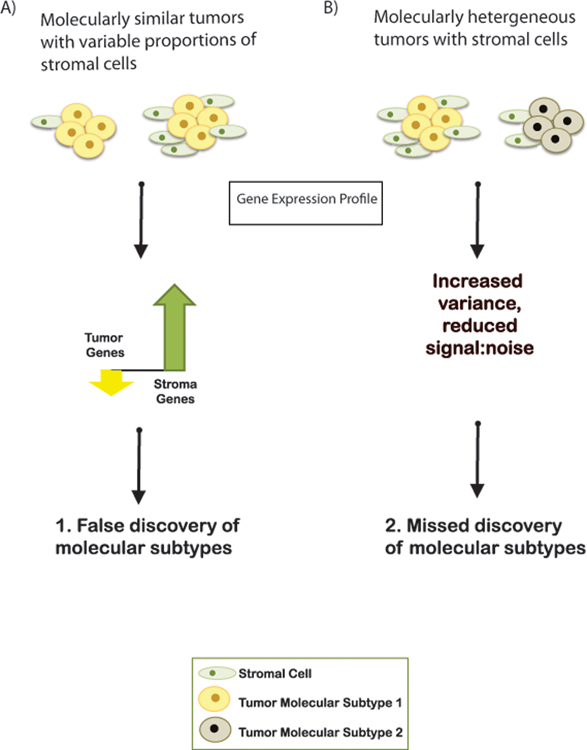
Additionally, cancer type- or subtype-specific biomarkers obtained in bulk gene expression profiling studies may include genes expressed by stromal or immune cells. Recent studies report discovery of six immune subtypes in 33 cancer types (17) and two stromal subtypes in ovarian cancer (16). The potentially complex relationships between non-malignant cells in HGSOC tumor microenvironment and malignant epithelial cell gene expression are unknown, but sampling variability in the cell admixture may impact subtype discovery (Figure 1). Single-cell gene expression profiling may resolve many of these issues, but the extent to which non-malignant cell admixture has influenced the field to date is unknown.
Figure 1: Variability in stromal proportions may present challenges in gene expression analysis of bulk tissue.
A) If two tumors of the same molecular subtype have different proportions of stromal cells, the observed average gene expression in the tumor with more stromal cells will show an increase in stroma-associated gene expression with a concurrent decrease in tumor-associated gene expression. This apparent decrease in tumor gene expression may result in false discovery of molecular subtypes. B) Different proportions of stromal cells will increase noise in analysis, which may cause researchers not to discover underlying molecular subtypes.
We systematically explored the extent to which stromal cell admixture in tumor samples influences molecular subtype and prognostic gene signature discovery (Figure 1). We hypothesized that, if unadjusted in statistical analyses, variance in the percent of stromal cells may lead to false discovery of tumor molecular subtypes, low precision of tumor classification, and reduced power to detect true ovarian epithelial carcinoma molecular subtypes and prognostic signatures. We tested this by identifying genes that were differentially expressed between microdissected tumor and stromal cells and examining whether these genes were enriched in published serous ovarian cancer subtypes and prognostic gene expression signatures. We show that stromal gene expression and gene signatures of stromal cell admixture are associated with overall survival in a meta-analysis of HGSOC gene expression datasets. We also investigate the number of published prognostic ovarian cancer gene signatures that add prognostic information to tumor-stroma proportion.
Materials and Methods
Data
Details and sources of all datasets are listed in Table S1. Raw gene expression data for two Australian Ovarian Cancer Study (AOCS) datasets (2) were downloaded from Gene Expression Omnibus (GEO). These AOCS datasets were 1) high-grade (G2, G3) ovarian adenocarcinomas (n=215; 11 endometrioid, 203 serous, one undifferentiated) that were previously classified as high-grade molecular subtypes C1, C2, C4, or C5 (the “AOCS dataset,” GEO GSE9891) and 2) a subset of four C1 tumors that had matched microdissected tumor and stroma (the “AOCS microdissected dataset,” n=8, GEO GSE9890). We excluded low grade and low malignant potential AOCS subtypes (C3 and C6 subtypes, n=36) and unclassified tumors (n=34). The Massachusetts General Hospital (MGH) ovarian cancer dataset was received directly from the authors and included Affymetrix gene expression profiles of pairs (n=38) of microdissected tumor and stroma of HGSOC from women hospitalized at the MGH (GSE18520, GSE40595, the “MGH dataset”) (18,19). The MGH dataset also included unmatched normal ovary stroma (n=10). In the Cancer Genome Atlas (TCGA) ovarian cancer dataset (n=518, downloaded November 24, 2010), all but one tumor was high-grade serous (2). Public gene expression data from 16 studies were extracted from the curatedOvarianData (20) database. Non-ovarian validation datasets were downloaded from GEO (Table S1) and included microarray gene expression profiles of microdissected stroma and tumor from non-inflammatory breast cancer (n=34 pairs) (GSE5847, Boersma et al.) (21) and invasive breast cancer (n=28 pairs) (GSE10797, Casey et al.) (22). These also included two prostate cancer datasets with known percent stromal content (GSE17951, n=136; GSE8218, n=109) (9).
To compare bulk and microdissected AOCS molecular profiles, raw data (cel files) were normalized using frozen robust multi-array analysis (fRMA) (23), which uses an external reference normalization approach, allowing one to combine data from different batches (when these data were generated using the same platform). Other GEO raw datasets were normalized using RMA (24,25). Data in curatedOvarianData were used as provided because these have been extensively manually curated, uniformly pre-processed, and normalized.
Statistical analysis
Analyses were performed using R and Bioconductor, versions 2.15 and 2.10, respectively, or later.
Gene signatures of tumor and stroma
Genome-wide differential gene expression analyses were performed using a moderated t-test, and p-values were adjusted for multiple testing using the Benjamini-Hochberg method (26) from the Bioconductor package limma (27). For paired microdissected stroma and tumor samples, a paired moderated t-test with a p-value cutoff of 0.01 was used. A separate analysis of the AOCS microdissected dataset with a less stringent cutoff (p<0.05) was also performed to identify a broader set of tumor and stromal genes. Genes that were differentially expressed between each AOCS or TCGA molecular subtypes were detected using an unpaired test. In differential gene expression analysis of the MGH dataset, a more stringent p-value cut-off was applied to generate gene lists of similar length. Hierarchical clustering was performed using Pearson correlation with average linkage clustering (28).
Tissue-specific tumor or stroma marker genes were identified by transforming gene expression to binary on/off calls (29) and ranking genes that were expressed in one tissue and not expressed in the other microdissected tissues. Genes were considered tissue-specific if they had high sensitivity and specificity (area under a receiver operating curve ≥0.9).
To generate a single gene signature covariate in statistical models, a gene signature score was computed using a weighted sum approach as previously described (30). Positive weights (+1) were applied to genes up-regulated in poor-prognosis tumors, BRCA mutation-like tumors, metastases, an angiogenic subtype, or malignant tumors. Negative weights (−1) were applied to the other genes in the signature. Linear regression was used to test for association between continuous variables and percent stromal content. Fisher’s exact test was used to test for overlap between gene sets, with the background number of genes equal to the number of probes or genes on the platform (for example the AOCS n=18,769).
Molecular subtype prediction
The gene lists reported to predict AOCS and TCGA molecular subtypes were downloaded as online publication supplements (1,2), curated, and submitted to GeneSigDB (31). To reproduce the AOCS and TCGA molecular subtype classifiers, gene lists were used to build subtype classifiers using the single sample predictor (SSP) approach (32). The centroids of gene expression profiles of the four molecular subtypes in the AOCS study (C1, C2, C4, C5) (1) and TCGA study (mesenchymal, immunoreactive, differentiated, proliferative) (2) were calculated (Figure S1), and cases (tumor or stroma) were assigned to the molecular subtype with highest Pearson correlation coefficient. A case was not classified to any molecular subtype (“unclassified”) if its correlation to each one of the subtypes was less than 0.7. There was high concordance in classification between our implementation of the subtype classifiers and their original studies. The AOCS SSP molecular subtype classifier had a self-validation accuracy of 95%, sensitivity 0.93–0.97, and specificity 0.96–1.0. (Figure S1C). The TCGA molecular subtype SSP classifier had an overall self-validation accuracy of 89%, 0.84–0.94 sensitivity, and 0.95–0.98 specificity for each subtype (Figure S1D).
Computational mixing of tumor and stroma fractions
Paired microdissected tumor and stroma (n=38) were computationally mixed in increasing increments of 10%, as described previously (33). The gene expression values of each tumor (Gt) were mixed with gene expression values of its adjacent stroma (Gs) using the linear combinations mGt + (1-m)Gs. Mixing parameter m is the proportion of tumor and 1-m is the proportion of stroma, incremented by 0.1 from 0 to 1. This was performed for all genes in the molecular subtype classifier to produce a computationally simulated mixture of tumor and stroma. The assumption of linearity has been previously supported by comparing the computational values to those observed in gene expression of co-cultured cells for the breast molecular subtype PAM50 gene signature (33).
Survival analysis
Cox proportional hazards regression was used to test for association between continuous variables and survival variables (relapse-free survival (RFS), overall survival (OS)). In survival analyses of gene signatures using independent datasets, the p-values were not corrected for multiple testing. Meta-analysis of the tumor-stroma gene signatures was performed using the metafor R package and summary hazard ratios and confidence intervals of the 16 studies (n=2,527 ovarian tumors) were calculated with a fixed effects model.
Exploratory data analysis of stromal genes and TCGA Immune Landscape Signatures
Additional clinical data, stromal fraction, leukocyte fraction, five core immune signatures (wound healing, macrophage regulation, lymphocyte infiltration, interferon gamma response, and TGF-β response), CIBERSORT estimates of >25 immune cell types, and mutation load for each TCGA HGSOC tumor were downloaded from the TCGA immune landscape study (17) article supplement (https://isb-cgc.shinyapps.io/shiny-iatlas/). Spearman Rank correlation coefficient was used to summarize the relationship between stromal gene expression score and immune features in TCGA HGSOC tumors.
Results
We investigated the extent to which non-tumor cell gene expression distinguishes HGSOC molecular subtypes using three different approaches: 1) cluster analysis to explore correlations between gene expression profiles of microdissected tumor, microdissected stroma, and HGSOC tumors of different molecular subtypes, 2) differential gene expression analysis to determine if genes that are differentially expressed between molecular subtypes are also differentially expressed in microdissected tumor or stroma, and 3) in silico cell admixture analysis to discover the robustness of molecular subtype classifications to increasing proportion of stromal cells.
Defining HGSOC tumor and stroma gene signatures
We defined gene expression signatures that characterize HGSOC and its adjacent stroma using paired laser microdissected tissue from two independent studies. Using the AOCS microdissected dataset, we defined a 688-gene signature using C1/stromal tumors (n=8) (p<0.01, paired t-test with modified variance (27) and FDR correction (26); Table S2). This AOCS tumor-stroma gene signature contained 461 and 227 unique genes with increased expression in C1 stroma and tumor, respectively, which are referred to as the “stromal gene set” and “tumor gene set.” A list with a larger number of genes extracted using lower differential expression stringency (p<0.05) is available in Table S2. In a second, independent analysis of paired microdissected tumor and stroma from HGSOC tumors of 38 patients in a Massachusetts General Hospital (MGH) study (18), we identified a similar gene signature. This MGH tumor-stroma gene signature of 519 unique genes contained 429 and 90 genes that had higher expression in stroma and tumor, respectively (Table S2). Despite the small sample size from which the AOCS tumor-stroma signature was derived, the MGH signature had significant overlap with it: 24/90 genes over-expressed in tumor (Fisher’s exact test p<10−10) and 122/429 genes over-expressed in stroma (p<10−25). The concordance of these signatures was supported by the observation that the AOCS signature correctly discriminated all but one of the microdissected tumor and stroma MGH specimens using unsupervised hierarchical cluster analysis (Figure S2).
Cluster analysis of microdissected tumor, stroma, and their bulk gene expression profiles
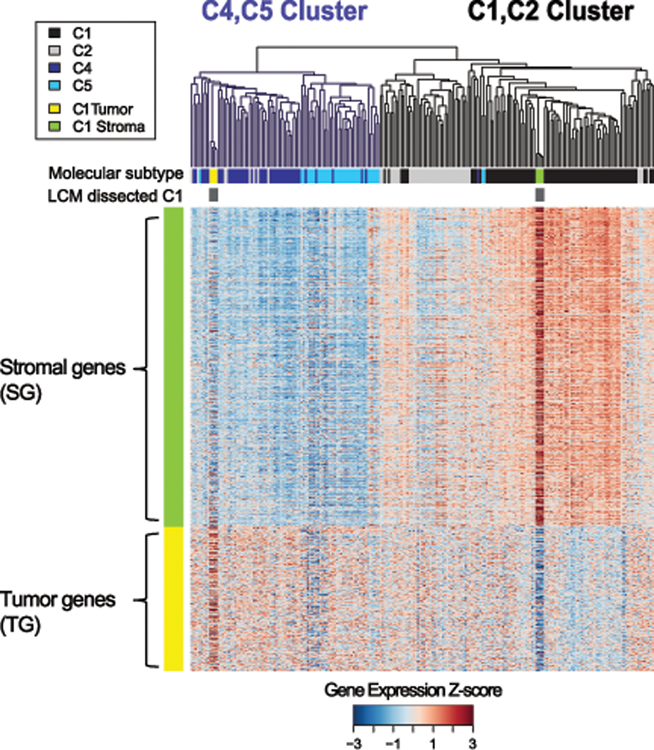
Using these tumor-stroma gene signatures, we investigated the contribution of stromal cell gene expression to the HGSOC molecular subtype using unsupervised cluster analysis. The 688-gene AOCS tumor-stroma gene signature partitioned the AOCS high-grade serous tumors (n=215) into two clusters (Figure 2): those with low vs. high stromal gene expression. These two clusters could be further subdivided to form four clusters, which largely reflected the four AOCS HGSOC molecular subtypes: C4/CA125, C5/MYCN deregulation (34), C2/immunoreactive, and C1/stromal. We observed that matched microdissected tumor samples did not cluster with the corresponding bulk tumor. Instead, microdissected C1 tumors clustered with C4/CA125 molecular subtype tumors while microdissected C1 stroma clustered with the C1 bulk tumors.
Figure 2: AOCS molecular subtypes can be distinguished by stromal and tumor gene expression.
Analysis of gene expression profiles of microdissected epithelial tumor cells and stroma from four C1 tumors identified a 688-gene ovarian tumor-stroma gene signature (FDR-adjusted p<0.01). The signature contained 461 genes that were over-expressed in microdissected stroma and 227 genes with increased expression in epithelial tumor cells, and these distinguished ovarian tumor and stroma in an independent MGH dataset (Figure S2). The heatmap shows the tumor and stromal gene expression profiles in the AOCS’s HGSOC tumors and microdissected C1 stroma and tumor samples. C1 microdissected stroma samples clustered with C1 subtype tumors, while the C1 microdissected tumor samples clustered with C4 subtype tumors. All of these analyses were done using unsupervised hierarchical cluster analysis. Colors of the heatmap were scaled (z-score values across rows).
The MGH tumor-stroma gene signature similarly partitioned the AOCS dataset into clusters (Figure S2C) which reflected AOCS HGSOC molecular subtypes, particularly the split of C1 compared to others. The observation that C1 tumors had higher stromal content is known, but the high concordance of tumor and stroma clusters with published HGSOC molecular subtypes suggested that genes which are differentially expressed between tumor and stroma might be sufficient to discriminate the HGSOC molecular subtypes.
Stromal genes are differentially expressed between ovarian cancer molecular subtypes
These molecular subtypes were then further examined using gene expression analysis, in which we quantified the relative contribution of stroma and tumor gene expression to each molecular subtype. We performed global differential gene expression analysis between each pair of AOCS tumor molecular subtypes (C1, C2, C4, and C5) (limma (27), 2-fold change and p<0.001 after FDR correction, Table S3A). The number of genes that discriminated between each pair of subtypes varied in size from 343 (C1 vs. C2) to 1,267 (C1 vs. C5) (Figure 3A). We tested the overlap between these gene lists and the AOCS stromal and tumor gene sets. The stromal and tumor gene sets derived from the AOCS microdissected dataset’s C1 tumors were again used so that the gene sets were most comparable. We observed that any contrast involving C1 had strong overlap with the stromal gene set (Fisher’s test p<10−61), as did the contrast of C2 vs. C4 (Fisher’s test p<10−73). Specifically, 54% and 41% of genes significantly different in contrasts of C1 vs. C2 or C1 vs. C4, respectively, were up-regulated in stromal cells (Figure 3A). This suggests that at least C1 is primarily driven by different amounts of stromal gene expression. In contrast, only 9% of genes differentially expressed between C4 and C5 were stromal genes (p = 1), consistent with the subtypes’ reported low amount of desmoplasia (1). The tumor gene set and “other” genes dominated C5 differential gene expression, consistent with C5’s distinct molecular properties (2,34).
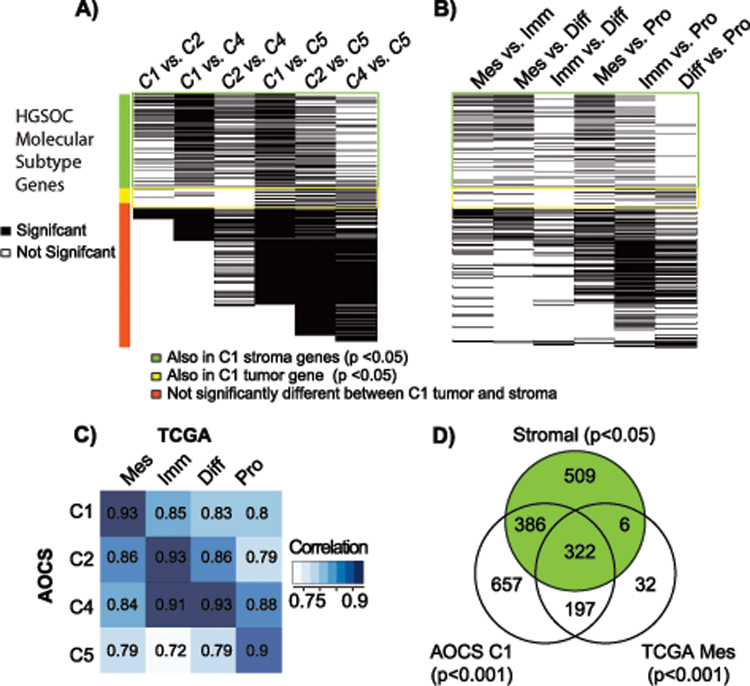
Figure 3: Overlap between genes that are differentially expressed between the AOCS and TCGA molecular subtypes and tumor-stroma genes sets.
Black and white lines indicate genes that were or were not strongly differentially expressed (limma, moderated t-statistics, p<0.001 after FDR correction), respectively, between each pair of A) AOCS and B) TCGA molecular subtypes. The color bar indicates whether these genes were also differentially expressed in laser capture microdissected C1 tumor (yellow) and stroma (green) at p<0.05, or not significantly differentially expressed between C1 tumor and stroma (orange). Stromal and tumor genes with cutoff p<0.05 rather than 0.01 were used in order to explore the extent of stromal gene enrichment in these subtypes. C) Correlation between average expression of AOCS and TCGA molecular subtypes, which was calculated using the intersection of all differentially expressed genes between any pair of AOCS or TCGA subtypes. Mes, Imm, Diff, and Pro are the TCGA molecular subtypes mesenchymal, immunoreactive, differentiated, and proliferative, respectively. D) There was a large overlap between genes expressed in C1 microdissected stroma and genes that were differentially expressed either between C1 vs. other AOCS subtypes, or between the TCGA mesenchymal subtype vs. other TCGA subtypes.
Based on these results, we also investigated the influence of stromal gene expression in TCGA HGSOC molecular subtypes (Figure 3B, Table S3B). The TCGA mesenchymal, immunoreactive, differentiated, and proliferative subtypes have highest global gene expression correlation with the AOCS subtypes C1, C2, C4, and C5, respectively (Figure 3C), and the mesenchymal subtype has significantly higher percent stromal content than other subtypes (t-test p=1.35 × 10−7). Results from analysis of the TCGA study confirmed findings in the AOCS subtype analysis. Genes differentially expressed (p<0.001, FDR correction) in the TCGA mesenchymal subtype, similar to C1, were mostly in the stromal gene set (Figure 3D, 59%), and overlapped strongly (35%) with genes differentially expressed in C1 (Figure 3D). Similar to the AOCS C4 and C5 subtypes, few genes in the stromal gene set were differentially expressed between TCGA’s differentiated and proliferative molecular subtypes (<5%, p=1). Lastly, genes that were differentially expressed in the proliferative subtype, similar to C5, were relatively enriched in tumor-specific and “other” genes.
Similar results were observed when these analyses were repeated with the MGH tumor and stroma signatures (Table S3C). Similar to the AOCS tumor signature, the MGH tumor signature did not significantly overlap with gene lists from pairwise comparisons of any two AOCS or TCGA molecular subtypes (p=1 for overlap with any comparison). Consistent with the AOCS stromal signature, the MGH stromal signature strongly overlapped with comparisons between the AOCS C1 subtype and either C2 or C4 and similar comparisons between the TCGA mesenchymal subtype and either the immune or differentiated subtypes. We observed that 24% and 19% of genes differentially expressed between C1 vs. C2 (p<10−13) and C1 vs. C4 (p<10−15), respectively, were MGH stromal genes. There was no significant overlap between MGH stromal genes and any pairwise comparison involving the AOCS C5 subtype (p=1). Therefore, in two independent studies, stroma-associated expression explains a large portion of the differential gene expression of select subtypes.
Molecular subtype classifications change with the proportion of stromal gene expression
We next investigated if the gene signature classifiers for these molecular subtypes were stable when the proportion of stromal cells changed. We trained a Single Sample Predictor (SSP) classifier using the gene lists reported to predict each of the AOCS C1, C2, C4, and C5 molecular subtypes (1) (Figure S1A,C). When applying the AOCS classifier to epithelial carcinoma cells microdissected from HGSOC tumors in an independent cohort (MGH dataset n=38), most microdissected tumors (20/38) were assigned to subtype C4 (Figure S1E,I). Matched microdissected stroma was all classified as C1 (n=23) or unclassified (n=15, Figure S1G,J). However, computational mixing of only 10% stromal gene expression with paired microdissected tumor caused 6/38 tumors to be reclassified (Figure S3A), including C4 tumors that were reclassified as C1 (n=2) or C2 (n=2). Molecular subtype C5 tumors remained stable, reflecting the lack of stromal gene expression determining this subtype. Five additional tumors were predicted to have a different molecular subtype when the stromal content was increased to 20%, and at 30% stroma, over one third (15/38) of tumors’ subtypes were reassigned.
We observed similar results regarding the robustness of TCGA molecular subtypes with increasing proportions of stroma. We built an SSP classifier of TCGA molecular subtypes using their 100-gene signature (7) (Figure S1B,D). When it was applied to classify microdissected tumors in the MGH dataset, most were assigned either to the differentiated subtype (23/38), which is similar to the AOCS C4 subtype, or the proliferative subtype (8/38), which is similar to C5 (Figure S1F,I). Microdissected stroma was classified as mesenchymal (20/38) or was unclassified (10/38, Figure S1H,J). With 10%, 20%, and 30% stroma, 0/38, 5/38, and 8/38 tumors, respectively, were classified to a different molecular subtype (Figure S3B). Thus, percent stromal content influences subtype classification.
The association between the proportion of stroma in tumors and overall survival in HGSOC
The AOCS C1/stromal subtype is reported to have a worse prognosis, and therefore we explored whether percent stroma itself was prognostic. We confirmed that the AOCS C1/stromal subtype, which has high stromal proportion, was associated with poorer overall survival. C1 had a hazard ratio (HR) of 1.72 (95% confidence interval 1.15–2.58, p=0.0083). The AOCS C1/stromal molecular subtype had a HR of 1.49 (95% CI 0.99–2.26, p=0.056) within high stage (III or IV) tumors.
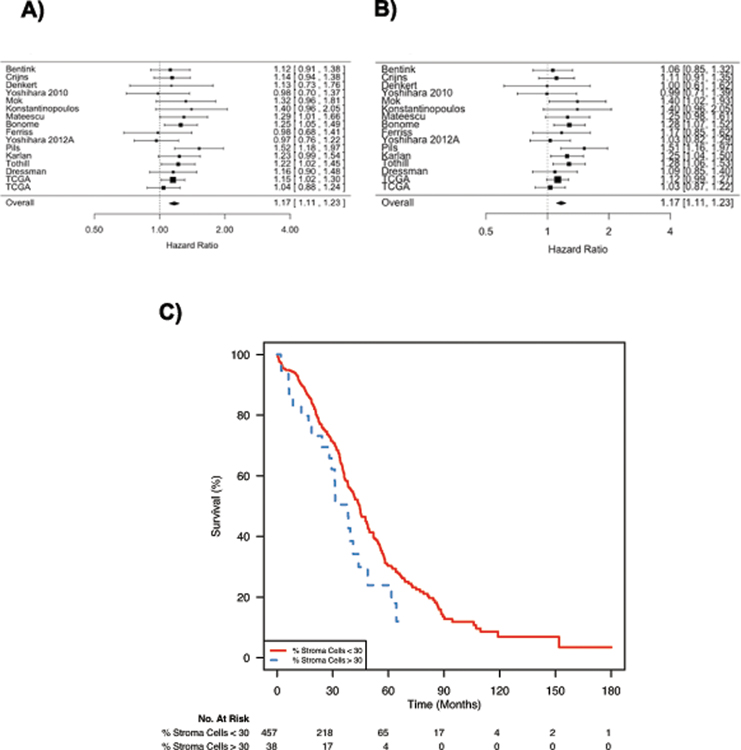
In the TCGA data, pathologist scores of percent stromal content had a weak but significant (HR 1.01 for each percent increase in stromal content, CI 1.0–1.02, p=0.024) association with overall survival in univariate Cox proportional hazards regression analyses of high stage tumors, but not across all tumors (HR 1.0, CI 0.99–1.02, p=0.088, Figure 4C for survival association with stromal content > 30%). In other studies, no estimate of percent stroma was available, but we assessed the prognostic power of a tumor-stroma gene signature in an aggregated cohort of 2,527 HGSOC patients from 16 studies, which included the TCGA and AOCS datasets (20). The 688-gene AOCS tumor-stroma signature had a HR of 1.17 (95% confidence interval 1.11–1.23, Figure 4A).
Figure 4: Stromal content’s association with overall survival.
These forest plots examine association with overall survival across ovarian cancer studies (n=16) using A) the AOCS tumor and stromal gene signatures or B) only the genes that were specific to tumor-associated stroma from the MGH study. The two TCGA gene expression datasets are n=510 Affymetrix HT_HG-U133A and n=256 Illumina HiSeq RNA sequencing. C) Kaplan-Meier curves of pathologist assessment of percent stroma (cutoff of 30%) in high stage TCGA tumors (HR 1.01 for each percent increase in stromal content, CI 1.0–1.02, p=0.024). A 30% stroma threshold is shown since it is between the original TCGA consortium tumor purity threshold of at least 80% and the later reduced threshold of 60% tumor cells.
The AOCS and MGH tumor-stroma signatures were generated using differential gene expression analysis and might include genes that are expressed in both tumor and stroma cells, albeit at different levels. Therefore, we derived a list of markers specific to tumor or stroma that better reflect cell admixture. Using expression profiles of paired microdissected epithelial tumor (MGH dataset n=38), adjacent tumor stroma (n=38), and unrelated normal ovarian stroma (n=10), we applied a barcode approach (29) to convert gene expression to binary expressed or not expressed calls. We then extracted a list of 28, 11, and 38 tissue markers with high specificity (Figure S4) for tumor, stroma adjacent to tumor, or normal ovary stroma (Table S2). The 11 cancer-associated stromal tissue markers had similar prognostic power (HR 1.17, CI 1.11–1.23, Figure 4B), but the tumor markers were not prognostic (HR 0.96, CI 0.96–1.01) and performed comparably to a randomly selected 20-gene signature (HR 0.96, CI 0.96–1.02). These results support that a higher stroma proportion in HGSOC tumors is associated with worse overall survival.
Some prognostic ovarian cancer signatures predict stroma proportion in HGSOC
Given that stromal gene expression was associated with survival in HGSOC tumors, we hypothesized that some published prognostic ovarian cancer gene signatures might incidentally be proxies for the presence of stromal tissue. Published gene signatures of ovarian cancer (n=61) were downloaded from the database GeneSigDB (31) or curated from the literature (Table S4). Gene signatures varied in size from four to 1,903 genes, with a median of 47 genes. Most gene signatures, including the TCGA study’s prognostic signature and Konstantinopoulos et al. BRCAness signature (35), were not significantly enriched in stromal genes (Table S4) and not associated with percent stromal content in TCGA samples (Table 1).
Table 1:
Associations between ovarian cancer gene signatures, stromal genes (SG), tumor genes (TG), and percent stromal content.
| Published gene signature | # Number of genesa | A) Overlap with SGs and TGs | B) % stroma contentb | C) Breast stroma vs. epitheliumb |
D) Prostate % stromal contentb |
||||
|---|---|---|---|---|---|---|---|---|---|
| Study | Signature components | # TGs (%) | # SGs (%) | TCGA (n = 518) |
Boersma (n = 34) |
Casey (n = 28) |
Wang (n = 109) |
Wang (n = 136) |
|
| AOCS C1 subtype stromal and tumor genes | Stromal genes (SG) Tumor genes (TG) |
461 227 |
0 227 |
461 0 |
5.6×10−13 | 4.3×10−8 | 7.1×10−8 | 1.5×10−9 | 4.9×10−17 |
| AOCS signatures (Tothill et al.) | Up in C1 Down in C1 |
287 147 |
0 2 (1) |
176 (61)* 2 (1) |
3.8×10−11 | 2.8×10−8 | 5.1×10−7 | 4.9×10−8 | 1.1×10−14 |
| Poor PFS Good PFS |
135 143 |
0 16 (11)* |
103 (76)* 0 |
8.6×10−13 | 4.0×10−9 | 4.1×10−8 | 1.2×10−9 | 2.0×10−17 | |
| Poor OS Good OS |
147 146 |
0 23 (16)* |
101 (69)* 0 |
5.3×10−11 | 4.6×10−9 | 9.7×10−8 | 1.7×10−5 | 6.5×10−13 | |
| MGH stroma vs. tumor signature | Tumor-associated stroma | 431 81 |
0 23 (28%)* |
125 (29%)* 0 |
6.5×10−15 | 1.6×10−5 | 2.2×10−8 | 1.6×10−11 | 3.6×10−17 |
| Tumor | |||||||||
| MGH tissue-specific genes | Tumor-associated stroma | 11 28 |
0 17 (57%)* |
5 (42%)* 0 |
2.1×10−14 0.136 |
8.1×10−5 0.001 (−) |
0.04 1.8×10−8 (−) |
1.6×10−6 3.7×10−9 (−) |
1.0×10−7 1.2×10−19 (−) |
| Tumor | |||||||||
| Bentink et al. | Angiogenic Non-angiogenic |
74 19 |
0 0 |
35 (47)* 1 (5) |
1.6×10−9 | 1.5×10−7 | 0.93 | 1.1×10−6 | 6.8×10−14 |
| Bignotti et al. | Up in metastasis Up in primary tumor |
89 36 |
0 1 (3) |
54 (61)* 0 |
1.2×10−9 | 4.5×10−7 | 5.2×10−5 | 7.2×10−4 | 7.2×10−8 |
| Spentzos et al. | Unfavorable prognosis Favorable prognosis |
73 43 |
0 4 (9)* |
19 (26)* 0 |
1.7×10−10 | 1.8×10−5 | 0.0074 (−) | 4.1×10−14 | 1.0×10−7 |
| Bonome et al. | Poor prognosis Good prognosis |
288 272 |
0 5 (2) |
37 (13)* 1 |
9.4×10−11 | 3.5×10−6 | 1.5×10−6 | 7.7×10−5 | 3.0×10−11 |
| Biade et al. | Malignant cluster Benign cluster |
21 15 |
4 (27)* 1 |
0 6 (29)* |
1.1×10−12 (−) | 9.8×10−8 (−) | 5.2×10−6 (−) | 6.5×10−10 (−) | 7.1×10−13 (−) |
| Konstantinopoulos et al. | BRCA-like Non-BRCA-like |
32 27 |
0 0 |
2 (6) 2 (7) |
0.7279 (−) | 0.0043 (−) | 7.6×10−6 (−) | 0.34 | 0.19 |
Details of published signatures in Table S4A
p < 0.001
A) Gene overlap between microdissected ovarian cancer gene signatures and both SGs and TGs of the AOCS microdissected dataset. P-values calculated with Fisher’s exact test. Association between gene signatures and B) the % stromal proportion in TCGA ovarian cancer tumors or microdissected stroma in C) two breast cancer datasets, or D) two prostate cancer datasets. P-values were calculated using B, D) linear regression or C) t-tests. Associations with stroma were mostly positive. Negative associations with stroma are indicated in parentheses (−) next to p-values, and otherwise, associations were positive. No multiple testing correction was used.
Total number of unique gene symbols reported in the specified gene signature from each study.
P-values derived using paired t-test or linear regression.
Over a third of published prognostic gene signatures (n=21/61) were enriched in AOCS stromal genes, and 9/61 were enriched in MGH stroma genes (p<0.05 after FDR correction). Eight signatures were further analyzed (Table 1); seven had strong enrichment (p<0.001) in both the AOCS and MGH stromal genes (Table S4), and the Biade et al. benign tumor signature (36) nearly met these p-value cutoffs. All of these eight signatures (AOCS signatures; Biade et al. benign tumor signature (36); Bentink et al. angiogenic signatures (14); and prognostic signatures from Bignotti et al. (37), Bonome et al. (38), and Spentzos et al. (39)) were positively associated with pathological scores of percent stromal content in TCGA tumors (linear regression, p<10−8, Table 1B). Genes reported to be associated with poor prognosis overlapped with the stromal gene set in 7/8 gene signatures (p<0.001), whereas the good prognosis genes were often enriched in the tumor gene sets (Table 1A). The Biade et al. signature (36) was an exception, which had increased stromal cell proportions in benign tumor samples (36).
Gene signatures enriched in stromal genes may provide little additional prognostic information beyond tumor-stroma ratio
Most of the eight gene signatures that were enriched in stromal genes were no longer associated with poor overall survival (OS) when adjusted for pathologist’s estimates of percent stromal content in TCGA tumors in Cox proportional hazard regression analysis (Table 2A). The AOCS prognostic gene signatures remained associated with OS and progression free survival (PFS) when adjusted for stromal content (p<0.01 and p<0.05, respectively, Table 2A) but were no longer significant after adjusting for both stromal content and stage. Gene signatures not enriched in stromal genes, such as the BRCAness signature, remained associated with OS in multivariate models that adjusted for both stromal content and stage (p<0.05, Table 2A).
Table 2:
Univariate and multivariate analysis of association between ovarian cancer gene signatures and overall survival (OS) of patients in A) the TCGA dataset or B) the smaller MGH study.
| Published gene signature | A) HGSOC tumors (TCGA, n=518)b | B) MGH Data (n=38) | |||||
|---|---|---|---|---|---|---|---|
| Study | Signature or % stromal content | OS | b OS, adjust stage | b OS, adjust % stroma | b OS, adjust % stroma + stage | OS, gene expression in microdissected epithelium | OS, gene expression in microdissected stroma |
| AOCS C1 stroma vs. tumora | Stromal genes vs. tumor genes | 0.0870 | 0.137 | 0.2730 | 0.484 | 0.2440 | 0.1800 |
| MGH stroma vs. tumora | Tumor-associated stroma vs. tumor | 0.0336 | 0.0544 | 0.148 | 0.289 | 0.555 | 0.156 |
| MGH tissue- specific genesa | Tumor | 0.778 | 0.640 | 0.996 | 0.906 | 0.924 | 0.718 |
| Tumor-associated stroma | 0.0430 | 0.0732 | 0.177 | 0.337 | 0.540 | 0.164 | |
| Normal or tumor-associated stroma | 0.0395 | 0.0707 | 0.140 | 0.294 | 0.598 | 0.175 | |
| TCGA stromal content | Pathologist assessment of % stroma in TCGA samples | 0.0979 | 0.0499 | N/A | N/A | N/A | N/A |
| AOCS (Tothill et al.) | C1 | 0.0175 | 0.0658 | 0.0755 | 0.276 | 0.343 | 0.316 |
| PFS | 0.0045 | 0.0195 | 0.0346 | 0.148 | 0.242 | 0.377 | |
| OS | 0.0010 | 0.0068 | 0.0076 | 0.0541 | 0.356 | 0.105 | |
| Bentink et al. | Angiogenic vs. non-angiogenic | 0.0313 | 0.115 | 0.109 | 0.374 | 0.552 | 0.324 |
| Bignotti et al. | Metastasis vs. primary | 0.0165 | 0.0921 | 0.0613 | 0.311 | 0.383 | 0.229 |
| Spentzos et al. | Prognosis | 0.0118 | 0.0330 | 0.0507 | 0.163 | 0.608 | 0.145 |
| Bonome et al. | Prognosis | 0.0426 | 0.0571 | 0.128 | 0.208 | 0.366 | 0.0352 |
| Biade et al. | Malignant vs. benign | 0.310 | 0.266 | 0.587 | 0.584 | 0.642 | 0.101 |
| Konstantinopoulos et al. | BRCA-like | 0.0167 | 0.0281 | 0.0272 | 0.0386 | 0.696 | 0.387 |
Significance (p < 0.05) is highlighted in bold.
Multivariate survival analyses of TCGA data adjusted for percent stromal content and/or stage.
The stromal signature is not specific to HGSOC
Several types of carcinoma cells have complex interactions with cells in the tumor microenvironment. Therefore, we asked whether stromal genes derived from ovarian cancer are specific to HGSOC or might more broadly characterize epithelial cancers. Each of the eight ovarian cancer signatures enriched in stromal genes were strongly associated with stroma vs. epithelial components of breast cancer (Boersma et al. (21), Casey et al. (22)) or percent stromal content in prostate cancer (Wang et al. (9)) (Table 1, linear regression or paired t-test p<10−4 in at least 3 of 4 datasets).
Clinical variables that explain stromal genes’ association with survival
We observed that the strength of association between greater stromal proportion and survival was variable across datasets. We hypothesized that this could be related to sampling heterogeneity. In the AOCS C1 subtype, 63% of tumor samples were from extra-ovarian sites (most commonly peritoneum, as defined by the AOCS study), which often reflects a higher stage and thus worse prognosis, compared to 19% extra-ovarian sampling in other AOCS subtypes (1). In the TCGA dataset, every tumor except for four were reported to be sampled at the ovary/pelvis (2). After adjusting for whether the sampling location was extra-ovarian, C1’s association with overall survival (adjusted p=0.065) was substantially attenuated. Thus, C1’s prognostic power is partially related to sampling location, which may in turn reflect tumor stage.
Discussion
We show that the proportion of stroma admixture in bulk tumors is a source of differential gene expression that impacts molecular subtyping and prognostic gene signature discovery in HGSOC.
Although adjacent stromal crosstalk can affect nearby epithelial cells’ gene expression patterns (16), this is a distinct phenomenon from changes in observed gene expression caused by the admixture of stromal and other cells in bulk tumors (tumor purity). In bulk gene expression profiling, the “observed” measure is an average of all cells in the admixture, which varies if the proportions of cell types change. We show that stromal genes differentiate the AOCS C1 molecular subtype or the TCGA mesenchymal subtype from other HGSOC molecular subtypes. This finding is supported by a recent immunohistochemical study of TCGA mesenchymal subtype tumors showing that several subtype marker genes were expressed exclusively or primarily in stroma (40). The dependence of the C1 subtype classifier on stromal genes may also explain why the TCGA mesenchymal subtype lacks robustness when applied to other datasets that may have variable tumor purity (8) and why few reproducible mutations or copy number alternations have been described for the subtype. In miRNA analysis, the mesenchymal TCGA molecular subtype clusters with the proliferative subtype (2). In contrast, the AOCS C5 and TCGA proliferative subtypes are reported to have genetic alterations, including copy number variation, in several genes (2,34), and consistently, these subtypes’ gene expression patterns were not well explained by stromal genes.
Small increases in stromal gene expression caused subtype misclassification in our computational model that mixed stroma and tumor gene expression profiles. The influence of non-cancer cell gene expression on subtype classification is potentially a source of systematic error in other solid tumors. The accuracy of signature classifiers that predict breast tumor molecular subtype (PAM50) or risk of recurrence (Oncotype DX) decrease considerably with decreasing proportion of tumor cells and increasing proportion of normal cells (33). Molecular subtype definitions developed from gene expression profiles of bulk ovarian, breast, and other carcinomas may need to be redefined such that molecular subtypes more precisely model the contributions of tumor, stroma, and other cell types. Thus far, at least two HGSOC stroma subtypes have been proposed (16) and their relationships to epithelial HGSOC molecular subtypes are yet to be described.
Deconvolution of cell mixtures in bulk gene expression studies is an active area of research (41), and we identified tissue specific marker genes of HGSOC epithelial carcinoma and stroma (Table S2) that may be applied to computational prediction of cell proportions and in the study of tumor/stroma interactions. These tissue-specific genes highlight the importance of studying both epithelial and stromal cell biology in cancer. CEP55 had the highest specificity and sensitivity of the tumor-specific genes, and it codes for a protein that interacts with BRCA2 and plays a critical role in regulating the final step of cytokinesis (42). Some stroma-specific were possibly related either to macrophage activity or to the recently described TGF-β-dependent and TGF-β-independent Smad signaling (16) in stroma of HGSOC. Two of the eleven stroma-specific genes (IFFO1, GLIPR1) were strong predictors both of pathologist’s stroma proportion scores in TCGA tumors (Pearson correlation 0.7 and 0.67, respectively, p<0.0001) and of a gene signature that predicts macrophage/monocyte activity (Figure S4, colony stimulating factor-1, CSF1 signature) (17,43). Three other stroma-specific genes were strongly correlated with TGCA immune landscape scores for TGF-β response (Figure S4) in TCGA tumors (17). These included Thrombospondin 1 (THBS1), which is associated with tumor growth and metastasis in gastric carcinoma (44), and biglycan (BGN), which has been shown to form a complex with either TGF-β1 or TGF-β1 type I receptor to intensify the phosphorylation of Smad2/3 in cultured endothelial cells (45).
Further understanding of stromal cell biology in HGSOC would be valuable, particularly since stromal gene expression was associated with a worse prognosis (Table 2) (12). Few studies provide pathologist estimates of stroma proportion, and we developed gene signature models for stroma proportion, which were weakly associated with worse survival in a meta-analysis of 2,527 tumors (HR 1.17) (20). We performed a systematic analysis of published prognostic signatures in HGSOC and discovered that most, including the BRCAness signature (35), do not predict stroma proportion. Several signatures were highly enriched in stromal genes and did not provide additional prognostic information beyond the stromal cell proportion, suggesting that adjusting for stromal proportions maybe useful in subtype and signature discovery.
In ovarian cancer, the link between stroma and survival may sometimes be related to other variables, such the sampling methods. For example, the prognostic performance of the AOCS stroma-associated subtype, C1, is partly explained by stage or sampling location. This association with sampling location also suggests that stromal content may be variable within a single tumor or between studies. In Biade et al. (36), the benign rather than poor-prognosis tumors were enriched in stroma, and a recent analysis of tumor purity across several cancers showed that tumor purity’s relationship with survival is often confounded with other variables (46).
These issues of sampling heterogeneity in bulk tissue were recognized as early as 1999 when Golub et al. (47) remarked: “studies will require careful experimental design to avoid potential experimental artifacts – especially in the case of solid tumors. Biopsy specimens, for example, might have gross differences in the proportion of stromal cells. Furthermore, accumulating evidence suggests that the composition and gene expression of stroma may vary depending upon the specific anatomic location of the specimens. Blind application of class discovery could result in identifying classes reflecting the proportion of stromal contamination in the samples, rather than underlying tumor biology. Such ‘classes’ would be real and reproducible but would not be of biological or clinical interest.”
Our results strongly support single-cell analysis or microdissection of tumor samples in gene expression studies so that the features of the epithelial tumor cells, stromal, and immune components can be each analyzed. Retrospective analysis of bulk gene expression patterns in light of emerging single-cell sequencing data are needed to refine molecular subtypes in ovarian cancer and other carcinomas and enable a more comprehensive search for molecular targets amenable to therapeutic intervention.
Supplementary Material
Acknowledgments
This study was supported in part by grants from the National Cancer Institute at the National Institutes of Health (RC4CA156551 to M. J. Birrer, P30CA006516 and R01CA174206 to G. Parmigiani, R01CA133057 to S.C. Mok, R03CA191447 to L. Waldron, U19CA148065 and P01CA087969 to J. Quackenbush) and the National Institute on Minority Health and Health Disparities at the National Institutes of Health (G12MD007599 to L. Waldron) and by the Assistant Secretary of Defense Health Program, through the Breast Cancer Research Program (W81XWH-15–1-0013 to A. C. Culhane). Opinions, interpretations, conclusions and recommendations are those of the author and are not necessarily endorsed by the Department of Defense.
Footnotes
The authors disclose no potential conflicts of interest.
References
- 1.Tothill RW, Tinker AV, George J, Brown R, Fox SB, Lade S, et al. Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin Cancer Res Off J Am Assoc Cancer Res. 2008;14:5198–208. [DOI] [PubMed] [Google Scholar]
- 2.Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Waldron L, Haibe-Kains B, Culhane AC, Riester M, Ding J, Wang XV, et al. Comparative meta-analysis of prognostic gene signatures for late-stage ovarian cancer. J Natl Cancer Inst. 2014;106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Riester M, Wei W, Waldron L, Culhane AC, Trippa L, Oliva E, et al. Risk prediction for late-stage ovarian cancer by meta-analysis of 1525 patient samples. J Natl Cancer Inst. 2014;106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Waldron L, Riester M, Birrer M. Molecular subtypes of high-grade serous ovarian cancer: the holy grail? J Natl Cancer Inst. 2014;106. [DOI] [PubMed]
- 6.Konecny GE, Wang C, Hamidi H, Winterhoff B, Kalli KR, Dering J, et al. Prognostic and therapeutic relevance of molecular subtypes in high-grade serous ovarian cancer. J Natl Cancer Inst. 2014;106. [DOI] [PMC free article] [PubMed]
- 7.Verhaak RGW, Tamayo P, Yang J-Y, Hubbard D, Zhang H, Creighton CJ, et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J Clin Invest. 2013;123:517–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen GM, Kannan L, Geistlinger L, Kofia V, Safikhani Z, Gendoo DMA, et al. Consensus on Molecular Subtypes of High-Grade Serous Ovarian Carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res. 2018; [DOI] [PMC free article] [PubMed]
- 9.Wang Y, Xia X-Q, Jia Z, Sawyers A, Yao H, Wang-Rodriquez J, et al. In silico estimates of tissue components in surgical samples based on expression profiling data. Cancer Res. 2010;70:6448–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Downey CL, Simpkins SA, White J, Holliday DL, Jones JL, Jordan LB, et al. The prognostic significance of tumour-stroma ratio in oestrogen receptor-positive breast cancer. Br J Cancer. 2014;110:1744–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Huijbers A, Tollenaar R a. EM, v Pelt GW, Zeestraten ECM, Dutton S, McConkey CC, et al. The proportion of tumor-stroma as a strong prognosticator for stage II and III colon cancer patients: validation in the VICTOR trial. Ann Oncol Off J Eur Soc Med Oncol. 2013;24:179–85. [DOI] [PubMed] [Google Scholar]
- 12.Chen Y, Zhang L, Liu W, Liu X. Prognostic Significance of the Tumor-Stroma Ratio in Epithelial Ovarian Cancer. BioMed Res Int. 2015;2015:589301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Joyce JA, Pollard JW. Microenvironmental regulation of metastasis. Nat Rev Cancer. 2009;9:239–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bentink S, Haibe-Kains B, Risch T, Fan J-B, Hirsch MS, Holton K, et al. Angiogenic mRNA and microRNA gene expression signature predicts a novel subtype of serous ovarian cancer. PloS One. 2012;7:e30269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhang S, Jing Y, Zhang M, Zhang Z, Ma P, Peng H, et al. Stroma-associated master regulators of molecular subtypes predict patient prognosis in ovarian cancer. Sci Rep. 2015;5:16066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Yeung T-L, Sheng J, Leung CS, Li F, Kim J, Ho SY, et al. Systematic Identification of Druggable Epithelial-Stromal Crosstalk Signaling Networks in Ovarian Cancer. J Natl Cancer Inst. 2018; [DOI] [PMC free article] [PubMed]
- 17.Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang T-H, et al. The Immune Landscape of Cancer. Immunity. 2018;48:812–830.e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mok SC, Bonome T, Vathipadiekal V, Bell A, Johnson ME, Wong K, et al. A gene signature predictive for outcome in advanced ovarian cancer identifies a survival factor: microfibril-associated glycoprotein 2. Cancer Cell. 2009;16:521–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Leung CS, Yeung T-L, Yip K-P, Pradeep S, Balasubramanian L, Liu J, et al. Calcium-dependent FAK/CREB/TNNC1 signalling mediates the effect of stromal MFAP5 on ovarian cancer metastatic potential. Nat Commun. 2014;5:5092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ganzfried BF, Riester M, Haibe-Kains B, Risch T, Tyekucheva S, Jazic I, et al. curatedOvarianData: clinically annotated data for the ovarian cancer transcriptome. Database J Biol Databases Curation. 2013;2013:bat013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Boersma BJ, Reimers M, Yi M, Ludwig JA, Luke BT, Stephens RM, et al. A stromal gene signature associated with inflammatory breast cancer. Int J Cancer. 2008;122:1324–32. [DOI] [PubMed] [Google Scholar]
- 22.Casey T, Bond J, Tighe S, Hunter T, Lintault L, Patel O, et al. Molecular signatures suggest a major role for stromal cells in development of invasive breast cancer. Breast Cancer Res Treat. 2009;114:47–62. [DOI] [PubMed] [Google Scholar]
- 23.McCall MN, Bolstad BM, Irizarry RA. Frozen robust multiarray analysis (fRMA). Biostat Oxf Engl. 2010;11:242–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostat Oxf Engl. 2003;4:249–64. [DOI] [PubMed] [Google Scholar]
- 25.Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003;31:e15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J R Stat Soc Ser B Methodol. 1995;57:289–300. [Google Scholar]
- 27.Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3. [DOI] [PubMed] [Google Scholar]
- 28.Culhane AC, Thioulouse J, Perrière G, Higgins DG. MADE4: an R package for multivariate analysis of gene expression data. Bioinforma Oxf Engl. 2005;21:2789–90. [DOI] [PubMed] [Google Scholar]
- 29.McCall MN, Uppal K, Jaffee HA, Zilliox MJ, Irizarry RA. The Gene Expression Barcode: leveraging public data repositories to begin cataloging the human and murine transcriptomes. Nucleic Acids Res. 2011;39:D1011–1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sotiriou C, Wirapati P, Loi S, Harris A, Fox S, Smeds J, et al. Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst. 2006;98:262–72. [DOI] [PubMed] [Google Scholar]
- 31.Culhane AC, Schröder MS, Sultana R, Picard SC, Martinelli EN, Kelly C, et al. GeneSigDB: a manually curated database and resource for analysis of gene expression signatures. Nucleic Acids Res. 2012;40:D1060–1066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Haibe-Kains B, Desmedt C, Loi S, Culhane AC, Bontempi G, Quackenbush J, et al. A three-gene model to robustly identify breast cancer molecular subtypes. J Natl Cancer Inst. 2012;104:311–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Elloumi F, Hu Z, Li Y, Parker JS, Gulley ML, Amos KD, et al. Systematic bias in genomic classification due to contaminating non-neoplastic tissue in breast tumor samples. BMC Med Genomics. 2011;4:54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Helland Å, Anglesio MS, George J, Cowin PA, Johnstone CN, House CM, et al. Deregulation of MYCN, LIN28B and LET7 in a molecular subtype of aggressive high-grade serous ovarian cancers. PloS One. 2011;6:e18064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Konstantinopoulos PA, Spentzos D, Karlan BY, Taniguchi T, Fountzilas E, Francoeur N, et al. Gene expression profile of BRCAness that correlates with responsiveness to chemotherapy and with outcome in patients with epithelial ovarian cancer. J Clin Oncol Off J Am Soc Clin Oncol. 2010;28:3555–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Biade S, Marinucci M, Schick J, Roberts D, Workman G, Sage EH, et al. Gene expression profiling of human ovarian tumours. Br J Cancer. 2006;95:1092–100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Bignotti E, Tassi RA, Calza S, Ravaggi A, Bandiera E, Rossi E, et al. Gene expression profile of ovarian serous papillary carcinomas: identification of metastasis-associated genes. Am J Obstet Gynecol. 2007;196:245.e1–11. [DOI] [PubMed] [Google Scholar]
- 38.Bonome T, Levine DA, Shih J, Randonovich M, Pise-Masison CA, Bogomolniy F, et al. A gene signature predicting for survival in suboptimally debulked patients with ovarian cancer. Cancer Res. 2008;68:5478–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Spentzos D, Levine DA, Ramoni MF, Joseph M, Gu X, Boyd J, et al. Gene expression signature with independent prognostic significance in epithelial ovarian cancer. J Clin Oncol Off J Am Soc Clin Oncol. 2004;22:4700–10. [DOI] [PubMed] [Google Scholar]
- 40.Zhang Q, Wang C, Cliby WA. Cancer-associated stroma significantly contributes to the mesenchymal subtype signature of serous ovarian cancer. Gynecol Oncol. 2018; [DOI] [PMC free article] [PubMed]
- 41.Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Mondal G, Rowley M, Guidugli L, Wu J, Pankratz VS, Couch FJ. BRCA2 localization to the midbody by filamin A regulates cep55 signaling and completion of cytokinesis. Dev Cell. 2012;23:137–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Beck AH, Espinosa I, Edris B, Li R, Montgomery K, Zhu S, et al. The Macrophage Colony Stimulating Factor-1 Response Signature in Breast Carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res. 2009;15:778–87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lin X-D, Chen S-Q, Qi Y-L, Zhu J-W, Tang Y, Lin J-Y. Overexpression of thrombospondin-1 in stromal myofibroblasts is associated with tumor growth and nodal metastasis in gastric carcinoma. J Surg Oncol. 2012;106:94–100. [DOI] [PubMed] [Google Scholar]
- 45.Hara T, Yoshida E, Shinkai Y, Yamamoto C, Fujiwara Y, Kumagai Y, et al. Biglycan Intensifies ALK5-Smad2/3 Signaling by TGF-β1 and Downregulates Syndecan-4 in Cultured Vascular Endothelial Cells. J Cell Biochem. 2017;118:1087–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Aran D, Sirota M, Butte AJ. Systematic pan-cancer analysis of tumour purity. Nat Commun. 2015;6:8971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science. 1999;286:531–7. [DOI] [PubMed] [Google Scholar]
- 48.Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43:D447–452. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.