Abstract
The ongoing SARS-CoV-2 pandemic highlights the importance of materials science in providing tools and technologies for antiviral research and treatment development. In this Review, we discuss previous efforts in materials science in developing imaging systems and microfluidic devices for the in-depth and real-time investigation of viral structures and transmission, as well as material platforms for the detection of viruses and the delivery of antiviral drugs and vaccines. We highlight the contribution of materials science to the manufacturing of personal protective equipment and to the design of simple, accurate and low-cost virus-detection devices. We then investigate future possibilities of materials science in antiviral research and treatment development, examining the role of materials in antiviral-drug design, including the importance of synthetic material platforms for organoids and organs-on-a-chip, in drug delivery and vaccination, and for the production of medical equipment. Materials-science-based technologies not only contribute to the ongoing SARS-CoV-2 research efforts but can also provide platforms and tools for the understanding, protection, detection and treatment of future viral diseases.
Subject terms: Virology, Diseases, Nanoscience and technology, Materials science
Materials science provides tools and technologies for the protection against viral infections, as well as for the understanding, diagnosis, treatment and prevention of viral diseases. This Review discusses present and future directions in antiviral materials-science research, with a focus on COVID-19.
Introduction
The novel coronavirus SARS-CoV-2 causes coronavirus disease 2019 (COVID-19) and has spread around the world, leading to a global pandemic1. The number of confirmed cases has reached 27,486,960 and the number of deaths has risen to 894,983 across 200 countries (as of September 9, 2020)2. The Center for Systems Science and Engineering at Johns Hopkins University3 has reported an even higher infection rate, with 27,699,112 confirmed cases and 900,203 deaths, indicating highly efficient transmission at an exponentially increasing rate1. The data from the World Health Organization (WHO) suggest that SARS-CoV-2 has a case fatality rate of about 3.3%, which is higher than that of the 1918 influenza pandemic (2%) and the 1957 influenza pandemic (0.6%)4. The rate of the spread of SARS-CoV-2 is 40-fold higher than that of SARS-CoV1, which makes it more difficult to control. Therefore, addressing the SARS-CoV-2 pandemic requires scientific collaborations across borders and disciplines5.
Viruses are able to adapt to changing environmental conditions and rapidly evolve6. Therefore, accurate, rapid and long-term detection strategies, efficient protection and frequent updating of treatments are needed to control the spread of viruses, which requires multidisciplinary solutions. In addition, only a global network can achieve coordinated detection, protection, containment and therapy. Materials science has long played a key role in the rapid response to emerging viral diseases by developing instruments that reveal the structures of viral proteins, by designing point-of-care devices for accurate and rapid detection and diagnosis 7,8, and by formulating therapeutic treatment and vaccination strategies9.
In this Review, we discuss the role of materials science in antiviral research and highlight the importance of collaborations with virologists, epidemiologists and clinicians in addressing the global pandemic caused by SARS-CoV-2 (ref.1). We examine viral transmission, infection and disease progression (Fig. 1a), and discuss viral protection, detection, treatment and vaccination from a materials-science viewpoint (Fig. 1b). Finally, we investigate the future role of materials science in studying infectious viruses and in the treatment of viral diseases. Indeed, the emergence of a new pandemic that is potentially more severe than the SARS-CoV-2 pandemic is not a matter of if but when1,10. Therefore, universal platform technologies offered by materials science will also play a crucial role in the understanding, protection, diagnosis, treatment and prevention of future viral outbreaks.
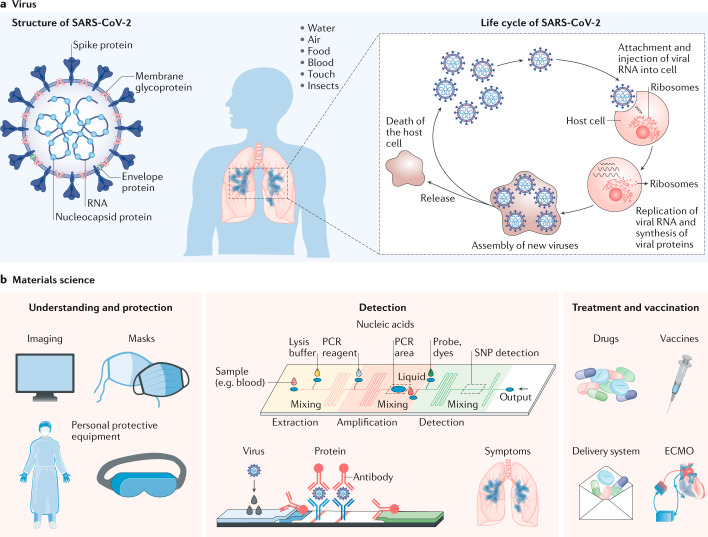
Fig. 1. SARS-CoV-2 and materials science.
a | The structure, transmission routes and replication cycles of SARS-CoV-2. b | Materials science contributes to the development and optimization of protective equipment and provides technologies and tools for the analysis of SARS-CoV-2, for example, high-resolution imaging, for sequencing (PCR) and protein analysis (immunoassays), for viral detection, for vaccine and treatment development and delivery, as well as by contributing advanced materials for clinical instruments, for example, filters for extracorporeal membrane oxygenation (ECMO) machines. SNP, single nucleotide polymorphism.
Materials science against viruses
Materials science is important in all areas of antiviral research, including investigation of viral structure and biology, protection, detection, treatment and vaccination.
Virus sequence and structure
The generation of efficient approaches to the protection, detection and treatment of viral diseases requires a thorough understanding of viral sequence, structure and transmission. Materials science offers important tools and technologies for viral research. For example, individual viral particles can be tracked in cells using confocal microscopy11,12 (Fig. 2a,b), allowing the dissection of the mechanism of infection. Gene sequencing provides the basis of antiviral-drug design and genomic-biomarker detection. Nanotechnology, for example, nanopore sequencing, has contributed to the speed and accuracy of next-generation gene-sequencing platforms13 (Fig. 2c). Based on the gene sequence, viral proteins can be synthesized and their 3D structure can be reconstructed at an atomic level using cryo-electron microscopy, X-ray crystallography14 or NMR spectroscopy15. However, X-ray crystallography requires a large number of small crystals, and large proteins are difficult to investigate by NMR spectroscopy, which also requires high protein concentration and stability. These limitations can be addressed by single-particle cryo-electron microscopy16, which has enabled the determination of the structure of many viruses17, including the 3.5-Å-resolution structure of the CoV spike (S) glycoprotein of SARS-CoV-2, as well as monoclonal antibody affinities18. Importantly, towards the understanding of viruses, materials science has played a key role in developing and optimizing instrumental components to ensure the accuracy and sensitivity of characterizations.
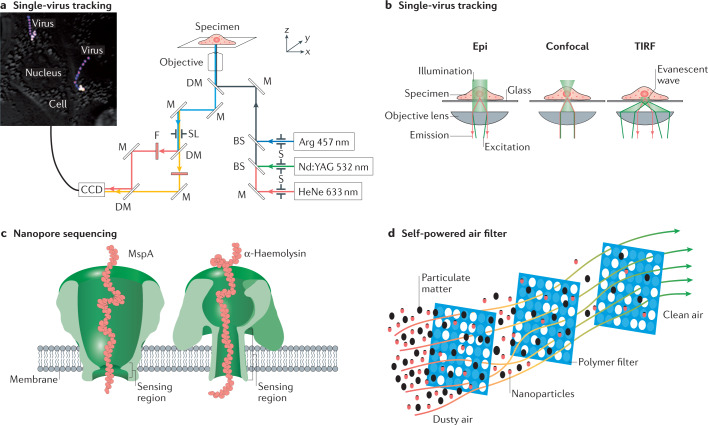
Fig. 2. Materials science in viral research and protection.
a | Single-virus tracking workflow using fluorescence microscopy. The representative image shows an influenza virus in Chinese hamster ovary cells (z-stacked time-lapse images, the colour code from pink/blue to yellow/white indicates the timescale from 0 s to 500 s). b | There are three possible imaging geometries in single-virus tracking, that is, epifluorescence geometry (Epi), confocal microscopy and total internal reflection fluorescence (TIRF) geometry. c | Nanopore sequencing using a nanosize pore and sensing regions in Mycobacterium smegmatis porin A (MspA) and α-haemolysin. d | A self-powered air filter can capture particulate matter and nanoparticles by surface adhesion. Arg, argon ion; BS, beam splitter; CCD, charge-coupled device; DM, dichroic mirror; F, filter; M, mirror; Nd:YAG, neodymium-doped yttrium aluminium garnet; S, shutters; SL, optical slits to control image size. Top image of panel a adapted with permission from ref.12, PNAS. Panels a and b reprinted from ref.11, Springer Nature Limited. Panel c reprinted from ref.13, Springer Nature Limited. Panel d reprinted from ref.34, CC BY 4.0.
Protection from viral infection
Viruses replicate inside living cells and are transmitted by direct contact19, saliva20, semen, vaginal fluid, anal mucus21, air22, blood-sucking insects23, food24, blood exposure25 or from pregnant women to newborns. SARS-CoV-2 first attaches to the host cell through binding of the S protein to the angiotensin-converting enzyme 2 (ACE2) receptor, allowing it to enter the cell and release its viral RNA (Fig. 1a). In the cell, the virus hijacks the machinery of the host cell to synthesize RNA and produce structural proteins, which assemble into new viruses that are released from the host cell. The replication cycle is then repeated26,27. Not only humans but also animals (cats, golden hamsters and ferrets)28,29 can be infected by SARS-CoV-2.
The most straightforward method for protection is physically isolating viruses by masks, gloves, face shields and protective suits. Surgical masks are made of three different fibre layers to prevent the entry of viruses; the outside layer is designed to stop liquid splash; the interlayer acts as a barrier against viruses and bacteria; and the inner coating absorbs moisture exhaled by the wearer30. Different materials can be synthesized to act as protective coatings, such as polyimide thin films with nanoporous membranes31 and carbon-based materials32. A polymer nanofibre filter, with the surface chemistry optimized to match that of particulate matter, allows both transparency to sunlight and sufficient airflow33. A self-powered air filter composed of an ionic liquid-polymer composite and a sponge network also enables efficient removal of particulate matter, including nanoparticles34 (Fig. 2d). Although these materials have not yet been used for viral protection owing to safety and cost concerns, they could be modified and introduced for virus protection in the foreseeable future, which will require research efforts to simplify synthesis procedures and guarantee safety.
Detection of infected patients
Rapid identification of infected patients helps limit the spread of viruses. Immunoassays and nucleic-acid-amplification tests using PCR are widely used and reliable methods for detecting viral proteins and nucleic acids, respectively. The accuracy of the results of immunoassays, such as radioimmunoassay35, enzyme-linked immunosorbent assay36 and the immune colloidal gold technique37, depends on the sensitivity of the instruments and materials. The immune colloidal gold technique is an immunoassay based on labelling the antibody with plasmonic gold nanostructures, enabling a fluorescent readout once the antigen is captured by the antibody. This technique can be used for the detection of viral antigens, for example, of the Zika and dengue viruses7. Compared with conventional singleplex approaches, this multiplexed immunoassay can amplify near-infrared fluorescence by up to 100 times, substantially improving the imaging of antibody binding. In addition, the immune colloidal gold technique requires only 1 μl of human serum, owing to its high sensitivity (Fig. 3a). Nanowire field-effect transistors can be combined with immune colloidal gold assays38 for real-time and direct electrical detection of single virus particles based on conductance changes after antigen–antibody binding (Fig. 3b). Similarly, an external electrical pulse can be applied to incorporate graphene quantum dots and gold-embedded polyaniline nanowires during the virus-accumulation step to increase the sensitivity of virus detection, for example, for the detection of the hepatitis E virus39 (Fig. 3c).
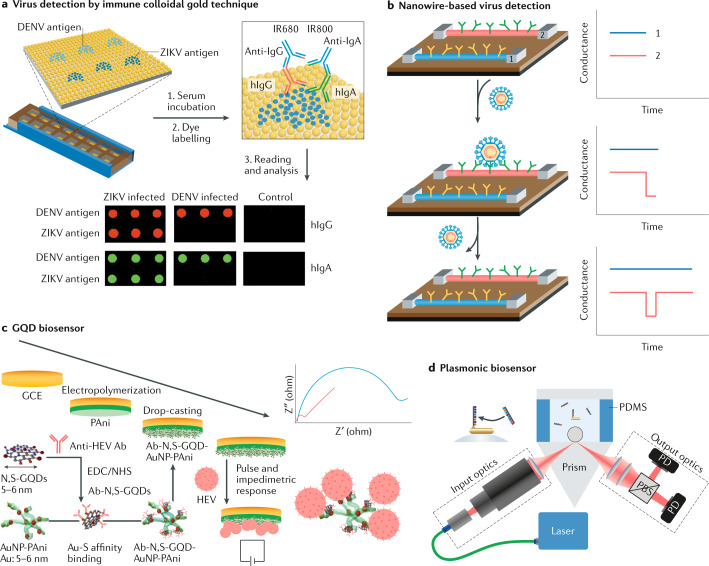
Fig. 3. Materials science in virus detection.
a | Multiplexed Zika virus/dengue virus (ZIKV/DENV) antigen microarray combining nanostructured plasmonic gold and near-infrared fluorescence molecules. Antibodies against ZIKV and DENV antigens in human serum are first captured by the microarray and then labelled with anti-human immunoglobulin G-infrared fluorescent dye 680 (IgG-IRDye680) and immunoglobulin A-infrared fluorescent dye 800 (IgA-IRDye800). Binding between IgG and IgA with antigens is evaluated by measuring the fluorescence intensities of the two dyes. b | Nanowire-based detection of single viruses. Binding of the virus to a specific antibody (Ab) leads to a change in conductance. c | An external electrical pulse and biosensors based on graphene quantum dots (GQDs) and gold-embedded polyaniline nanowires (AuNP-PAni) can be used for the detection of hepatitis E virus (HEV). The biosensor electrode, which is based on anti-HEV Ab-conjugated to nitrogen and sulfur codoped graphene quantum dots and gold-embedded polyaniline nanowires (Ab-N,S-GQDs@AuNP-PAni), can capture HEV. The HEV concentration is determined from the pulse-induced impedimetric response. d | A single-molecule whispering gallery mode biosensor platform using plasmonic gold nanorods can be used to detect single nucleic acid molecules. EDC, 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride; GCE, glass carbon electrode; NHS, N-hydroxysuccinimide; N,S-GQDs, nitrogen and sulfur codoped graphene quantum dots; PBS, polarizing beam splitter; PD, photodetector; PDMS, polydimethylsiloxane. Panel a reprinted from ref.7, Springer Nature Limited. Panel b reprinted with permission from ref.38, PNAS. Panel c reprinted from ref.39, CC BY 4.0. Panel d reprinted from ref.47, Springer Nature Limited.
Real-time PCR with reverse transcription (rRT-PCR) is commonly used for the detection of viral DNA or RNA40, and is the recommended technique for the detection of the single-stranded RNA virus SARS-CoV-2 (ref.41). However, PCR-based techniques for nucleic-acid detection come with trade-offs in terms of efficacy, cost, simplicity and speed. Alternatively, the CRISPR effectors Cas12 (refs42,43) and Cas13a44 can be applied for nucleic-acid detection. For example, specific high-sensitivity enzymatic reporter unlocking (SHERLOCK) is a fluorescence-based or lateral-flow-based assay using CRISPR nucleases for nucleic-acid detection45. Interestingly, a prophylactic antiviral CRISPR strategy using Cas13 has also been proposed to inhibit SARS-CoV-2 in human cells46. Single-molecule nucleic-acid sensitivity and selectivity can also be achieved using a biosensing platform based on microcavities. In this platform, the hybridization kinetics of matching and non-matching strands are detected by monitoring microcavity resonance wavelength shifts caused by interaction of the molecules with nanorods47 (Fig. 3d). This label-free technology has the advantage of transiently monitoring molecular interactions, allowing statistical analysis of interaction kinetics and increasing sensor lifetime. Nanopore-based platforms can also be used for targeted sequencing, and have already been applied for SARS-CoV-2 virus detection48.
The immune system responds to viral infections and induces detectable symptoms, such as high body temperature. Non-contact detection of body temperature is mainly performed to preliminarily detect infected patients by using infrared thermal-imaging systems, for example, hand-held infrared temperature guns. However, the temperature gun is inaccurate at cold ambient temperatures, because the temperature is calculated based on a Seebeck coefficient49 close to the body temperature, which is not available at low temperatures. Although laboratory assays have been developed for identifying infected patients, the detection and diagnosis of severe viral infections, such as COVID-19, may also require clinical diagnostic methods, such as computed tomography (CT), ultrasonic imaging, magnetic resonance imaging, single-photon emission computerized tomography and positron emission tomography. For example, CT can be used to detect interstitial changes and small patchy shadows in the lung, as well as extrapulmonary bands, in patients infected with SARS-CoV-2 (ref.50). However, instruments and professional medical staff are limited, and, therefore, portable diagnosis instruments51,52 and artificial intelligence (AI)53 should be further developed and combined with clinical diagnosis to improve the detection process and guarantee the accuracy of results.
Treatment and vaccination
Currently available antiviral drugs can be broadly divided based on their mechanism of action into penetration and husking inhibitors, DNA polymerase inhibitors, reverse transcriptase inhibitors, protein inhibitors, neuraminidase inhibitors and broad-spectrum antiviral drugs54–56. Indeed, the majority of antiviral drugs currently used in the clinic are polymerase inhibitors, which are specific for a given viral enzyme. However, owing to their specificity, polymerase inhibitors cannot be applied for the treatment of infections with new viruses, such as SARS-CoV-2.
In contrast to bacteria, which can often be treated with antibiotics owing to their similar structure and replication mechanisms, the structure and replication of viruses greatly differ between types and species. Therefore, only few broad-spectrum antiviral drugs exist57,58. For example, cyclodextrins modified with mercaptoundecane sulfonic acids mimic the key viricidal effects of heparan sulfates without causing systemic toxicity59. Alternatively, antiviral nanoparticles have been developed with flexible and long linkers that mimic heparan sulfate proteoglycans. The nanoparticles generate high forces (~190 pN), which irreversibly deform the virus60 (Fig. 4a). Based on the spatially defined arrangement of ligands for the viral spike protein, bacteriophage capsids have also been designed that carry a multivalent binder. The bacteriophages can cover the entire virus envelope and, thus, prevent its binding to the host cell61. In addition, nanosponges derived from the plasma membranes of human lung epithelial type II cells or of human macrophages can neutralize SARS-CoV-2 by displaying the protein receptors necessary for SARS-CoV-2 entry into host cells62.
Fig. 4. Materials science in the treatment and vaccination of viral diseases.
a | Gold nanoparticles (AuNPs) coated with long and flexible moieties of undecanesulfonic acid (MUS) show viricidal activity against heparan sulfate proteoglycan (HSPG)-binding viruses, owing to the generation of high forces (~190 pN), which irreversibly deform the virus; by contrast, 3-mercaptoethylsulfonate (MES)-coated AuNPs are not antiviral because of the short molecule length. b | Nasal delivery of inactivated H1N1 influenza virus and pulmonary surfactant guanosine monophosphate–adenosine monophosphate (PS-GAMP; an activator of stimulator of interferon genes (STING)) leads to the stimulation of dendritic cell (DC) maturation, antibody generation and, subsequently, to a CD8+ T cell and tissue-resident memory T (TRM) cell response, generating broad protection against seasonal influenza B virus (IBV), H3N2, H5N1 and H7N9 influenza viruses. c | Lipid nanoparticles can be used for the delivery of a Zika virus pre-membrane and envelope (ZIKV prM-E)-encoding mRNA vaccine against the Zika virus. Delivering a ZIKV prM-E fusion loop mutant-encoding mRNA diminishes the generation of cross-reactive antibodies that promote Dengue virus infection. d | During extracorporeal membrane oxygenation, venous blood is drained from the body, oxygenated by fresh gas (the blender modulates the ratio between air and oxygen) using a gas-exchange device and then returned to the body. AEC, alveolar epithelial cell. Panel a reprinted from ref.60, Springer Nature Limited. Panel b from ref.84, Herold, S. & Sander, L.-E. Toward a universal flu vaccine. Science 367, 852–853 (2020). Redrawn with permission from AAAS. Panel c reprinted with permission from ref.89, Elsevier. Panel d from ref.95, N. Engl. J. Med. Brodie, D. & Bacchetta, M. Extracorporeal membrane oxygenation for ARDS in adults. 365, 1905–1914. Copyright © (2011) Massachusetts Medical Society. Reprinted with permission from Massachusetts Medical Society.
Similarly, biomimetic cell membrane vesicles can act as delivery platforms. Cell membrane vesicles have excellent biocompatibility and possess the inherited functions of the source cells, such as long circulation time (red blood cells)63, specific targeting moieties and receptors62,64. For example, virus-mimetic nanovesicles can deliver antiviral antigens and mimic the viral property to trigger specific immunogenicity65. Nanodecoys composed of cell membranes of the Zika virus and gelatin nanoparticles can be administered to capture the Zika virus and inhibit Zika virus entry and infection66. Nanomaterials may also be applied to control the cytokine storm associated with pneumonia, which is a common symptom in patients with severe COVID-19. For example, neutrophil membrane-coated poly(lactic-co-glycolic acid) nanoparticles can neutralize pro-inflammatory cytokines and suppress inflammation in inflammatory arthritis, which might also be applicable for pneumonia67.
The development of new antiviral drugs is challenging and requires thorough knowledge of the infection and replication mechanisms of the specific virus. For example, based on the genome sequence similarity between SARS-CoV and SARS-CoV-2, the serine protease TMPRSS2 was identified to be crucial for S protein priming and, thus, for SARS-CoV-2 cell entry via ACE2. Therefore, a TMPRSS2 inhibitor may provide a treatment strategy to prevent SARS-CoV-2 infection68. Remdesivir, which is a nucleotide analogue prodrug that inhibits RNA-dependent RNA synthetase, has also been explored for COVID-19 treatment69. Preliminary data of the Randomised Evaluation of COVID-19 Therapy (RECOVERY) trial, testing dexamethasone as a treatment strategy for hospitalized COVID-19 patients, shows a lower 28-day mortality rate of treated patients than patients in the standard-care group, supporting a potential treatment effect of dexamethasone in patients who need respiratory support70. Injection or inhalation of dexamethasone nanoformulations may promote the treatment efficacy on COVID-19 through targeting of the drug to hyperactivated immune cells, potentiating its antioedema activity and antifibrotic effects71. Moreover, neutralizing antibodies72–74 and nanobodies (antibody fragments)75,76 have been proposed for COVID-19 treatment. A recent clinical trial (NCT04313127), testing neutralizing antibodies to prevent COVID-19, had shown satisfactory results and is in further evaluation77,78. However, achieving a high concentration of neutralizing antibodies is crucial for efficacy and, thus, aluminium-based adjuvants, such as those used for PiCoVacc79,80, may be helpful.
Vaccines remain the most efficient strategy to protect against viral infections, because they generate specific antibodies81. Following infection, the virus causes inflammation and stimulates the release of chemokines, which attract leucocytes to the infected area to destroy the virus82. At the same time, phagocytes engulf viruses, disassemble them, deliver them to nearby lymph nodes and present the antigens to other immune cells, such as B and T cells, to initiate the production of antibodies that specifically recognize viral antigens82. However, upon first contact with a virus, the immune response is slow, and can only help patients to recover, but not to prevent the disease. Once antigen-specific T and B cells are activated, they proliferate and generate memory cells. Therefore, upon second contact with the same type of virus, the immune system can respond quickly and efficiently. Pulmonary surfactant biomimetic liposomes can be applied to encapsulate and deliver an agonist of the interferon gene inducer, that is, stimulator of interferon genes, to increase the immune response against influenza viruses83,84 (Fig. 4b).
The purpose of a vaccine is to protect the body against a specific virus, capitalizing on the memory of the immune system, so that the immune system can eliminate the virus upon first contact. Vaccine design includes the identification of an antigen and adjuvant, which boosts the immune response caused by the antigen85, as well as an appropriate delivery platform. A variety of vaccines have been developed, including live-attenuated vaccines, inactivated vaccines, toxoid vaccines, subunit vaccines, conjugate vaccines, DNA vaccines, mRNA vaccines and recombinant vector vaccines. mRNA vaccines86, which directly induce antigen expression, provide an interesting alternative for RNA viruses, such as the Zika virus and SARS-CoV-2, which are susceptible to mutation and, therefore, to drug resistance6. mRNA vaccines require nanocarriers for encapsulation and stabilization of the mRNA87,88; for example, mRNA can be encapsulated in lipid nanoparticles to protect mice against Zika89 virus infection (Fig. 4c).
Although there is no mRNA vaccine yet approved for any disease, several clinical trials are under way. For example, Pfizer Inc., in a collaboration with BioNTech AG, is working on the phase II/III clinical trial of BNT162 mRNA vaccines for COVID-19 (NCT04368728)90 and has just announced data from preclinical studies91. Moderna, Inc. is currently conducting phase II clinical trials (NCT04232280) with an mRNA vaccine encoding a monoclonal antibody against cytomegalovirus92. In addition, the mRNA-1273 vaccine, developed by Moderna, Inc., against SARS-CoV-2 is currently in phase III clinical trials (NCT04470427)93. These mRNA vaccines are all delivered by lipid nanoparticles.
Finally, medical devices, such as ventilators, are important for the treatment of infected patients, to provide oxygen and ensure airflow in the lungs. Alternatively, extracorporeal membrane oxygenation (ECMO) can be applied in patients with severe respiratory failure to promote their recovery process94. In this procedure, venous blood is drained from the body, oxygenated using a gas-exchange device and then returned to the body95 (Fig. 4d). ECMO has also been applied for the treatment of COVID-19 patients94.
Possibilities for materials science
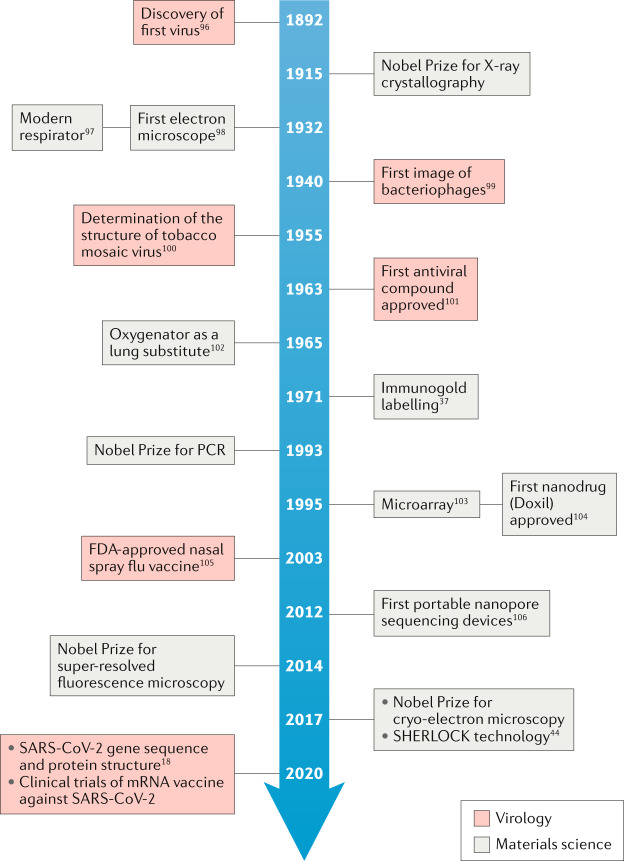
Materials science has long played an important role in the study of viruses and in the treatment of viral infections18,37,44,96–106 (Fig. 5). In the future, materials science and engineering will certainly continue to contribute to confronting viral outbreaks by providing platform technologies and tools for virus research, ensuring efficient and comfortable protection, performing rapid, accurate and low-cost detection, and enabling timely and effective treatment and vaccination.
Fig. 5. Timeline of key contributions of materials science to virology.
FDA, US Food and Drug Administration; SHERLOCK, specific high-sensitivity enzymatic reporter unlocking.
Physical protection
Physical protection provides a key strategy to prevent viral infection. Materials science can be applied to optimize physical-protection platforms in terms of preparation, manufacturing, material performance and structure, and to balance comfort and practicality. Commercial face masks can reduce virus emission from infected patients by providing a physical barrier with a certain cut-off size30. However, masks could also be designed to incorporate drugs or physical obstacles for killing the virus and stopping the transmission; for example, the virus could be damaged by heat generated by photothermal materials, such as graphene32, or by producing reactive oxygen species through a photodynamic process107–109. The design of reusable masks could further address personal protective equipment shortages, as experienced during the COVID-19 pandemic. Reusable masks would need to retain filtration ability and remain safe after decontamination, such as ultraviolet irradiation, moist heat and vaporous hydrogen peroxide110. To address this challenge, fibres, such as washable polypropylene fibres coated with assembled metal-organic frameworks111 or thermally stable electrospun polyimide/zeolitic imidazolate framework-8 nanofibres112, could be applied that tolerate heat, ultraviolet irradiation113 and hydrogen-peroxide treatment. In addition, a self-cleaning capability could be implemented by changing the surface chemistry of the material or by coating materials with photothermal materials32,109 or hydrophobic membranes31 to capture, kill and release the virus; for example, likewise, using filters composed of metal-organic frameworks with photodynamic ability109 or graphene with photothermal ability32. Functional nanofibres, for example, fibres modified with amino-functionalized silica particles114 or nanodecoys66 capable of capturing viruses through charge interactions or by active recognition, could also be used for masks and manufactured by nanospinning115 or 3D printing116,117. To address the problem of fogging when wearing safety glasses and masks, a layer of non-ionic surfactant could be added to the surface of protective equipment. Environmental contamination may also play a role in SARS-CoV-2 transmission118. Therefore, long-lasting and sprayable antiviral materials (for example, functional nanogels can multivalently interact with viruses and have adhesive effects on the surface)119 could be developed that can be applied on surfaces to keep the environment and protective equipment clean.
Virus detection
Materials science offers tools and techniques to improve the speed, sensitivity, accuracy and portability of virus-detection methods, in particular, for the isolation of nucleic acids and proteins from viruses. For example, magnetic particles can be used to capture nucleic acids or proteins in a sample, which can then be separated by application of a magnetic field120. Integrating microfluidics with isolation techniques can reduce the amount of reagents, increase isolation speed and allow automation121. Moreover, microfluidics can be introduced to rapidly mass-synthesize uniform magnetic particles122 and to efficiently isolate viruses using magnetic-particle-based microfluidics123. In addition, DNA124 or RNA nanostructures125 that contain complementary oligonucleotides to virus DNA or RNA could be combined with microfluidics for virus isolation. Although current nasal swabs are effective in collecting virus samples, nasal swabs with a controlled porous architecture could be functionalized with materials that can capture viruses through multivalent interactions119, which would enable the collection of viruses even at very low concentrations.
Similarly, aggregation-induced-emission (AIE) technology or quantum dots could be integrated with immune colloidal gold techniques to increase viral-detection sensitivity126. In addition, materials-science approaches, such as ionic-liquid-based materials127,128, could be applied to improve the stability of extracted nucleic acids or proteins for further investigation. Improving the sensitivity, specificity and simplicity of virus-detection methods is important to ensure accurate and rapid testing of samples. For example, an Fe–Au nanoparticle-coupling approach, combining PCR amplification with the quantitative measurement of Au nanoparticles using inductively coupled plasma mass spectrometry129, which was originally developed for the detection of circulating tumour DNA, could also be applied for the sensitive and specific detection of SARS-CoV-2 RNA. Microarrays, which have been developed for the precise control and manipulation of fluids, combine sample preparation, reaction, separation and detection on one chip of a few square centimetres130, and could be used in combination with surface-enhanced Raman spectroscopy for rapid virus detection using nanoparticle aggregates or arrays that can generate surface-enhanced Raman spectroscopy signals131,132. Low-cost point-of-care devices, designed with microfluidics, could also be adapted for viral detection, requiring only small sample volumes. Microfluidics can further be used for the production of designed nanoparticles133,134, which are used for viral detection, treatment and vaccination.
Plasma protein profiles have been investigated as disease-related biomarkers using mass-spectroscopy-based proteomics; similarly, viral protein biomarkers could be detected in blood135. However, the accuracy of protein identification by mass-spectrometry-based proteomics is limited, owing to the concentration range (from pg ml−1 to mg ml−1) of the >10,000 proteins in plasma. The protein corona, that is, the shell of biomolecules formed on a nanoparticle upon contact with a biofluid136, can be exploited to increase the concentration of proteins for mass spectrometry to overcome the bottleneck of blood-sample-based proteomics and to enable the detection of low-abundance proteins137. Therefore, investigating the protein corona138,139 may also provide a strategy for virus detection and/or to study the host response. For example, nanoparticles could be designed to adsorb viral protein biomarkers or host response proteins (that is, cytokines) in plasma, serum or other biological samples. The nanoparticles could then be isolated for downstream mass-spectrometry-based proteomics analysis, enabling fully automated, high-throughput viral detection, including nanoparticle-plasma incubation, protein-corona purification using magnetism, protein digestion and liquid chromatography–tandem mass spectrometry139.
Bioelectronic systems140 enable the real-time and remotely controlled monitoring of biological information in the body; for example, wearable sensors for the wireless monitoring of uric acid and tyrosine in sweat141, glucose levels during diabetes therapy142 and skin temperature143. Bioelectronic devices could also be applied for the long-term monitoring of symptoms of viral infections, such as an increase in temperature or respiratory syndrome. Protective equipment could also be equipped with a sensor that can sense the entry of viral particles and translate the signal to a smartphone. For example, cellular-binding-like events between the virus and host-cell-mimicking sensors (that is, sensors coated with host-cell membranes or multivalent binders) can be detected by thermal-shift measurements based on protein stabilization induced by a ligand144,145, which can be translated into a digital signal recorded by a smartphone. Similarly, interactions between viral particles and specific molecules could be detected in protective masks.
Mobile technologies offer the possibility of track-and-trace systems that provide information on COVID-19-infected patients, including age, gender, symptoms and geographical areas. These systems can alert users when they are in high-risk areas via an app to slow down the spread of SARS-CoV-2. Mobile technology can also be used to collect data for analysis and to inform guidelines for government policies. For example, a COVID-19 symptoms tracker app has been developed. The app has already collected data from over 2.8 million participants, providing information about risk factors, symptoms, clinical results and geographic hotspots. The data are publicly available and important in the response to a pandemic; however, privacy concerns and ethical considerations should also be considered146. In addition, AI, such as machine learning, can be applied to large datasets of clinical results to help clinicians in their diagnosis. For example, an AI system has been developed based on the CT database of COVID-19 patients, enabling rapid diagnosis147.
Finally, point-of-care devices are particularly important in areas with low resources; for example, low-cost and portable gene-sequencing devices that can be operated without the need for trained medical staff. Accurate, real-time monitoring of viral transmission can further help to understand the infection process; for example, bioluminescent microscopy can be used to image the transmission of a virus with an inserted luciferase gene in mice148,149 or ferrets150. Similarly, fluorescent protein genes such as green fluorescent proteins and red fluorescent proteins genes151–153 could be integrated into the SARS-CoV-2 genome for tracking transmission in vitro and in animal models.
Drug discovery and delivery
Materials science can also contribute to the development of new technologies, such as thermal proximity coaggregation144 and time-resolved wide-angle X-ray scattering154, which allow the dynamic observation of structural protein changes in host cells in response to a virus, and virus–drug interactions, to accelerate and improve the process of drug discovery. In addition, drug development relies heavily on studies in animal models, which cannot entirely recapitulate the human response and often inaccurately predict the safety and efficacy in humans. Alternatively, microphysiological systems, such as organ-on-a-chip platforms, in which engineered human tissues are connected by physiologically relevant flows, can be applied for drug development, as well as for toxicity and efficacy evaluation. Similarly, organoids derived from stem cells can provide macroscale models of human tissues featuring the complexity and spatial heterogeneity of human organs. However, compared with animal models, organs-on-a-chip and organoids would fail in discovering systemic immunity-modulation drugs, because they cannot recapitulate the organ microenvironment155,156. Moreover, the lifespan of these platforms would need to be prolonged to allow the generation of mature tissues undergoing homeostasis156. Importantly, organoids can be used for disease modelling and for the evaluation of antiviral drug toxicity and treatment effects157–159. For example, bat and human intestinal organoids were used to confirm that bat intestinal cells can be infected by SARS-CoV-2 and that the human intestinal tract is a possible transmission route of SARS-CoV-2 (ref.160). Furthermore, it could be demonstrated in SARS-CoV-2-infected kidney and human blood vessel organoids that soluble human ACE2 can inhibit SARS-CoV-2 infection161. Materials science can improve organoid cultures by developing chemically defined, tunable, synthetic scaffolds that guide the assembly of human cells into tissues and that support organoid maturation162,163.
Nanoparticles can enable the effective delivery of drugs, improving circulation time, enhancing targeting efficiency and reducing potential adverse side effects. Nanocarriers can also be used to deliver antiviral therapeutics into cells to intervene with the viral-replication cycle at the molecular level or to deliver plasmid DNA164 and mRNA165 for vaccination. Vaccine efficiency and safety are of the utmost importance, and, therefore, US Food and Drug Administration (FDA)-approved materials should be used as delivery platforms; for example, a lipid-nanoparticle-encapsulated mRNA, encoding the receptor binding domain of SARS-CoV-2, has been reported as a vaccine candidate166. The FDA-approved lipid nanoparticle improves the stability of the mRNA and the transfection efficiency in vivo.
Lessons learned from cancer vaccines can benefit the development of antiviral vaccines. Biocompatible materials have been explored for drug delivery in cancer, to improve the stability and pharmacokinetics of payloads and to prevent premature release or degradation. In addition, nanomaterials can be modified to target the delivery to antigen-presenting cells and to enable co-delivery of viral antigens and adjuvants to the lymph nodes167, for example, by using an albumin-hitchhiking approach168 or by targeting phagocytic cells to promote lymph-node transportation169. Nanomaterials can also be designed to respond to a specific environment or to external stimuli, allowing the control of payload release and preventing undesired immune reactions, such as immune toxicities in normal tissues caused by payload leakage or an immune cytokine storm caused by rapid payload release87, for example, pH-responsive nanoparticles170 or light-responsive nanoparticles171. Cancer vaccines can be designed to integrate diagnosis and treatment using nanomaterials with imaging abilities172, for example, to monitor the delivery of antigens and adjuvants in vivo, which may also be relevant for antiviral vaccines. Real-time data on the location and treatment responses could be generated by introducing NIR-II bioluminescence173, NIR-II chemiluminescence174 or NIR-II fluorescence175 to enable the rational design of nanoscale materials and bench-to-bedside translation of vaccines. Materials science also enables the design of self-administration platforms; for example, vaccines formulated as microneedle patches176, which slowly release payloads upon self-administration. Reproducibility and scalability also play major roles for the translation of nanomaterials for drug delivery. Microfluidic systems can be used for the controlled, reproducible and high-throughput synthesis of libraries of nanoparticles. Nanomaterials also play a role in virus treatment, for example, to provide cofactors, such as Zn2+, which inhibits RNA polymerase and, thus, transcription177. Zn-based nanomaterials that bind RNA polymerase could also be designed and administered to inhibit virus replication. In contrast to dendrimers or polymers, the valency and spatial structures of DNA nanostructures can be easily designed. In addition, DNA nanomaterials are non-toxic and stable. Therefore, DNA nanostructures with particular shapes, modified with virus-targeting aptamers, can be applied for virus detection and inhibition124. Owing to their customized spatial structure, such DNA nanostructures precisely match the viral surface, enabling efficient virus capture.
Inhalation as a route of drug and vaccine administration is particularly relevant for viruses causing respiratory symptoms, such as SARS-CoV-2, because it allows small-molecule drugs178 and vaccines179 to directly target the lungs, in which the virus resides180. Inhalation is particularly advantageous in resource-poor regions, because no medical staff is needed and self-administration of drugs or vaccines improves patient compliance. Inspired by the approved intranasal spray vaccine FluMist105, similar inhaled drugs or vaccines could be encapsulated using smart nanoparticles to promote their stability and to realize long-term and controlled release. In addition, inhaled nanoparticles encapsulating drugs or vaccines can be modified with specific antibodies targeting the cells in which the viruses reside to kill the viruses, or immune cells to trigger an immune response. This strategy could also decrease potential side effects of drugs and vaccines by avoiding drug delivery to healthy tissues. For example, nanomaterials could be coated with a specific ligand that binds to receptors on host cells of the virus in the lungs. Therefore, following inhalation, the nanoparticles would inhibit virus–host cell interactions. In the case of SARS-CoV-2, the ACE2 receptor or the serine protease TMPRSS2 could be targeted68,181. Nanoparticles can also be modified with elements of host-cell membranes to inhibit viral entry64,66; for example, nanomaterials can be decorated with host-cell membranes derived from human lung epithelial type II cells or human macrophages62 to capture SARS-CoV-2 and inhibit infection. Similarly, using nanomaterials that can generate free radicals56 or heat32 could improve the antiviral efficacy of such approaches.
Nanoscale materials have a similar size to viruses, which allows them to interact with the virus at the same size level. Cyclodextrins modified with mercaptoundecane sulfonic acids59, nanoparticles modified with long linkers that mimic heparan sulfate proteoglycans60, nanodecoys64, cellular nanosponges62 or DNA nanostructures124 could bind the virus to inhibit infection by inhibiting virus entry56.
Viral infections, such as SARS-CoV182 and dengue183 virus, can cause cytokine storms as a result of an excessive immune response184. SARS-CoV-2 infection can also induce a cytokine storm in some patients, which contributes to inflammation and pneumonia185. Lessons from nanotechnology-based cancer immunotherapy can inform treatment designs to modulate the cytokine storm by delivering anti-inflammatory drugs or inhibitors to inflammation sites. In particular, bioresponsive nanomaterials186–188 that target inflammation sites can deliver anti-inflammatory drugs and inhibitors. However, the targeting efficacy and payload leakage still need to be optimized to decrease side effects. In addition, the fabrication of clinical-grade materials with the ability of modulating cytokine storms remains a challenge.
Medical equipment and instruments
Materials science can be applied to optimize filters in ventilators with self-cleaning and virus-inhibition ability. 3D printing of ventilators could be introduced and optimized to fabricate ventilators with more functions. Similarly, ECMO machines, which are applied as life-saving treatment for severe respiratory diseases, such as COVID-19, suffer from high complexity, the need for highly trained staff, low portability and concerns about potential harm189, such as changes in thrombotic190 and immune-related factors189. A small device could be developed that combines the blood pump and the oxygenator to improve portability and reduce thrombotic issues, by decreasing the time that blood spends outside the body. Indeed, portable ECMO devices, such as the Maquet Cardiohelp device, which combines the pump and oxygenator in one unit191, have been developed. To decrease the size of such devices, wireless technology and energy-converting materials could be applied. In addition, bioelectronic systems and wireless mobile technology could be integrated in ECMO devices to monitor blood-status changes in real time. The gas-exchange membranes, which are a key component of ECMO machines, are usually made of hollow fibres, such as silicone rubber fibres192 or polymethylpentene materials193. Their resistance to blood flow and their gas-exchange capacity could be further improved by using 3D-printable biocompatible materials, such as respiratory-membrane-derived materials.
Outlook
Viral-disease outbreaks require a rapid response to obtain basic knowledge, but also continuous antiviral research. Materials science plays an important role in many aspects of antiviral research, including the study of viral biology and transmission, protection, identification of infected patients, treatment and vaccination. The investigation of new viral strains requires virus isolation and purification, size, structure and component determination, and analysis of infection, replication and spread. To prepare for new viral outbreaks and to accelerate and coordinate our response, portable and inexpensive devices for viral isolation and purification from samples such as blood or saliva are needed; for example, a hand-powered ultra-low-cost centrifuge194,195. To investigate virus structure and composition, electron microscopy, cryo-electron microscopy, confocal microscopy and sequencing technologies need to be applied. Additionally, the real-time dynamics of the replication of a single virus in host cells and tissues, including RNA replication, protein formation and cell-to-cell transmission, need to be assessed, for example, using super-resolution imaging or time-resolved wide-angle X-ray scattering. Advanced materials-enabled technologies, such as thermal-proximity coaggregation144 and time-resolved wide-angle X-ray scattering154, could further enable the observation of interactions between drugs and viruses. These technologies will require more powerful instruments with high temporal and spatial resolution, as well as high sensitivity. However, spatial resolution should not be achieved at the expense of time resolution, for example, in super-resolution microscopy. The development of portable and inexpensive imaging equipment, such as the Octopi196, BiteOscope197 and scale-free vertical tracking microscopy198, are particularly important in low-resource settings. However, the nanoscale settings in viral research provide a challenge for portable optical-imaging systems, owing to the diffraction limit. Thus, the time resolution might be more important than the spatial resolution in this case.
During viral outbreaks, large amounts of masks and personal protective equipment need to be rapidly produced, for example, using 3D printing117 and nano-electrospinning for the fabrication of nanofibres to compose filters. Virus detection further needs to be quick and done with high accuracy and at a low price, for example, by distributing self-testing kits, which have a sample-collection tool for viral capture (enabled by a specific surface chemistry, for example, a coating that can capture viruses by multivalent interactions119 at low concentrations), and the results are shown on a strip43,199. Sharing of results and data is also an important aspect in antiviral research, in particular, during viral-disease outbreaks.
The delivery of antiviral drugs and vaccines will greatly benefit from the use of vehicles based on biocompatible materials to reduce systemic toxicity, to improve circulation time, for the co-delivery of multiple components, to increase drug or vaccine stability and to target specific cells or tissues. Drugs and vaccines need to be preclinically tested in cells and virus-infected animal models. Alternatively, engineered ex vivo systems can be applied, for example, organoids and organs-on-a-chip, which allow the assessment of drug effects in real time. The production of drugs or vaccines may further require the use of robots and microfluidic technologies to enable scalability and to be able to respond to high demand. To tackle future challenges, the collaboration between different scientific fields, clinicians and industry will be required.
Acknowledgements
This study is supported by the US METAvivor Early Career Investigator Award (No. 2018A020560, W.Tao), Harvard Medical School/Brigham and Women’s Hospital Department of Anaesthesiology Basic Scientist Grant (No. 2420 BPA075, W.Tao) and Center for Nanomedicine Research Fund (No.2019A014810, W.Tao). W.Tao is a recipient of the Khoury Innovation Award (No. 2020A003219) and American Heart Association (AHA) Collaborative Science Award (No. 2018A004190). W.Tao also received a start-up package (for 3 years) from the Department of Anesthesiology,Perioperative and Pain Medicine to establish his independent research laboratory at Harvard Medical School and Brigham and Women’s Hospital. We thank our department for this generous support. We also thank J. Chen from University of California, Los Angeles, for his proofreading and correction before the online publication of this Review.
Author contributions
All authors contributed to the discussion and writing of this manuscript.
Competing interests
O.C.F. has financial interests in Selecta Biosciences, Tarveda Therapeutics and Seer. R.L. receives licensing fees (to patents on which he was an inventor) from, invested in, consults (or was on scientific advisory boards or boards of directors) for, lectured (and received a fee) or conducts sponsored research at MIT for which he was not paid for the following entities: 7th Sense, Abpro Labs, Abpro Korea, Acorda (formerly Civitas Therapeutics), Aleph Farms, Alivio Therapeutics, Alkermes, Allevi, Alnylam Pharmaceuticals, Apotex, Arcadia Biosciences, Arsenal Medical, Artificial Cells, Avalon GloboCare, BASF, BioInnovation Institute, Blackstone, Boston Children’s Hospital, Celero, Cellomics, Cellular Biomedical, Charles River, Clontech, Combined Therapeutics, Conference Forum, Cornell University, CRISPR Therapeutics AG, Crown Bioscience, Cygnal Therapeutics, Daros, Daré Bioscience, DeepBiome, Dewpoint Therapeutics, Dispendix/Cellink, Domain, Eagle, EdiGene Biotechnology, Editas Medicine, Eli Lily, Eisai, Entrega, Everlywell, Evox Therapeutics, Flagship Pioneering, Frequency Therapeutics, GeneMedicine, GenScript USA, GENUV, GlaxoSmithKline, Glycobia, Glympse Bio, GreenLight Biosciences, HCR, Hopewell Therapeutics, Horizon Discovery, Humacyte, IBEX Pharmaceuticals, ImmuneXcite, Indivior, Inovio, Institute of Immunology, Integrated DNA Technologies, InVivo Therapeutics, IxBio, J.R. Simplot, Jnana Therapeutics, Kala Pharmaceuticals, Kallyope, Kensa, Kodikaz Therapeutics, KSQ Therapeutics, Landsdowne Labs, LikeMinds, Luminopia, Luye, Lyndra Therapeutics, Lyra Therapeutics, McGovern Institute, Medikinetics, Merck, MGH Ragon Institute, Micelle, Moderna Therapeutics, Momenta, Monsanto, Muse Biotechnologies, Mylan, Nanobiosym, Nanobiotix, NewBridge Ventures, Noveome Biotherapeutics, Novo Nordisk, Particles for Humanity, Pfizer, Pioneer Hi-Bred International, Polaris Partners, Portal Instruments, Preceres, Pulmatrix, PureTech, ReLive, ReproCELL USA, Rubius Therapeutics, Secant Medical, Seer, Selecta Biosciences, Senses Biosciences, Senses, Setsuro Tech, Seventh Sense Biosystems, Shenzhen Rice Life Technology, Shire AG, Shiseido, Siglion, Sigma-Aldrich, SiO2, SKE S.R.L., Soil Culture Solutions, SQZ Biotechnologies, StemBioSys, Suono Bio, T2 Biosystems, Taconic Biosciences, TARA, Tarveda Therapeutics, Tesio Pharmaceuticals, Third Rock, Tiba Biotech, TISSIUM, TransGen, Translate Bio, TriLink BioTechnologies, Unilever, VasoRx, Verseau Therapeutics, Vivtex, Whitehead Institute, Wiki Foods, YZ Biosciences and Zenomics.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Zhongmin Tang, Na Kong, Xingcai Zhang.
Contributor Information
Omid C. Farokhzad, Email: ofarokhzad@bwh.harvard.edu
Wei Tao, Email: wtao@bwh.harvard.edu.
References
- 1.Gates B. Responding to Covid-19 — a once-in-a-century pandemic? N. Engl. J. Med. 2020;382:1677–1679. doi: 10.1056/NEJMp2003762. [DOI] [PubMed] [Google Scholar]
- 2.World Health Organization. Weekly operational update on COVID-19. 9 September 2020 (WHO, 2020).
- 3.Johns Hopkins University. COVID-19 dashboard by the center for systems science and engineering (CSSE) at Johns Hopkins University (JHU). JHUhttps://coronavirus.jhu.edu/map.html (2020).
- 4.The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team. Vital surveillances: the epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) — China, 2020. China CDC Wkly2, 113–122 (2020). [PMC free article] [PubMed]
- 5.Thorp HH. Time to pull together. Science. 2020;367:1282. doi: 10.1126/science.abb7518. [DOI] [PubMed] [Google Scholar]
- 6.Lauring AS, Frydman J, Andino R. The role of mutational robustness in RNA virus evolution. Nat. Rev. Microbiol. 2013;11:327–336. doi: 10.1038/nrmicro3003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhang B, et al. Diagnosis of Zika virus infection on a nanotechnology platform. Nat. Med. 2017;23:548–550. doi: 10.1038/nm.4302. [DOI] [PubMed] [Google Scholar]
- 8.Ferguson BS, et al. Genetic analysis of H1N1 influenza virus from throat swab samples in a microfluidic system for point-of-care diagnostics. J. Am. Chem. Soc. 2011;133:9129–9135. doi: 10.1021/ja203981w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nguyen DN, Green JJ, Chan JM, Langer R, Anderson DG. Polymeric materials for gene delivery and DNA vaccination. Adv. Mater. 2009;21:847–867. doi: 10.1002/adma.200801478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gates B. The next epidemic — lessons from Ebola. N. Engl. J. Med. 2015;372:1381–1384. doi: 10.1056/NEJMp1502918. [DOI] [PubMed] [Google Scholar]
- 11.Brandenburg B, Zhuang X. Virus trafficking–learning from single-virus tracking. Nat. Rev. Microbiol. 2007;5:197–208. doi: 10.1038/nrmicro1615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lakadamyali M, Rust MJ, Babcock HP, Zhuang X. Visualizing infection of individual influenza viruses. Proc. Natl Acad. Sci. USA. 2003;100:9280–9285. doi: 10.1073/pnas.0832269100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Deamer D, Akeson M, Branton D. Three decades of nanopore sequencing. Nat. Biotechnol. 2016;34:518–524. doi: 10.1038/nbt.3423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Drenth, J. Principles of Protein X-ray Crystallography (Springer, 2007).
- 15.Wuthrich K. Protein structure determination in solution by nuclear magnetic resonance spectroscopy. Science. 1989;243:45–50. doi: 10.1126/science.2911719. [DOI] [PubMed] [Google Scholar]
- 16.Cheng Y, Grigorieff N, Penczek PA, Walz T. A primer to single-particle cryo-electron microscopy. Cell. 2015;161:438–449. doi: 10.1016/j.cell.2015.03.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sirohi D, et al. The 3.8 Å resolution cryo-EM structure of Zika virus. Science. 2016;352:467–470. doi: 10.1126/science.aaf5316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wrapp D, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367:1260–1263. doi: 10.1126/science.abb2507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Killingley B, Nguyen-Van-Tam J. Routes of influenza transmission. Influenza Other Respir. Viruses. 2013;7:42–51. doi: 10.1111/irv.12080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Abe K, Inchauspe G. Transmission of hepatitis C by saliva. Lancet. 1991;337:248. doi: 10.1016/0140-6736(91)92222-n. [DOI] [PubMed] [Google Scholar]
- 21.Royce RA, Sena A, Cates W, Jr, Cohen MS. Sexual transmission of HIV. N. Engl. J. Med. 1997;336:1072–1078. doi: 10.1056/NEJM199704103361507. [DOI] [PubMed] [Google Scholar]
- 22.Cowling BJ, et al. Aerosol transmission is an important mode of influenza A virus spread. Nat. Commun. 2013;4:1935–1940. doi: 10.1038/ncomms2922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu Y, et al. Evolutionary enhancement of Zika virus infectivity in Aedes aegypti mosquitoes. Nature. 2017;545:482–486. doi: 10.1038/nature22365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Acheson D, Fiore AE. Hepatitis A transmitted by food. Clin. Infect. Dis. 2004;38:705–715. doi: 10.1086/381671. [DOI] [PubMed] [Google Scholar]
- 25.Ward JW, et al. Transmission of human immunodeficiency virus (HIV) by blood transfusions screened as negative for HIV antibody. N. Engl. J. Med. 1988;318:473–478. doi: 10.1056/NEJM198802253180803. [DOI] [PubMed] [Google Scholar]
- 26.Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB. Pharmacologic treatments for coronavirus disease 2019 (COVID-19): a review. JAMA. 2020;323:1824–1836. doi: 10.1001/jama.2020.6019. [DOI] [PubMed] [Google Scholar]
- 27.Shang J, et al. Cell entry mechanisms of SARS-CoV-2. Proc. Natl Acad. Sci. USA. 2020;117:11727–11734. doi: 10.1073/pnas.2003138117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shi J, et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2. Science. 2020;368:1016–1020. doi: 10.1126/science.abb7015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sia SF, et al. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature. 2020;583:834–838. doi: 10.1038/s41586-020-2342-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chua MH, et al. Face masks in the new COVID-19 normal: materials, testing, and perspectives. Research. 2020;2020:7286735. doi: 10.34133/2020/7286735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.El-Atab N, Qaiser N, Badghaish HS, Shaikh SF, Hussain MM. A flexible nanoporous template for the design and development of reusable anti-COVID-19 hydrophobic face masks. ACS Nano. 2020;14:7659–7665. doi: 10.1021/acsnano.0c03976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhong H, et al. Reusable and recyclable graphene masks with outstanding superhydrophobic and photothermal performances. ACS Nano. 2020;14:6213–6221. doi: 10.1021/acsnano.0c02250. [DOI] [PubMed] [Google Scholar]
- 33.Liu C, et al. Transparent air filter for high-efficiency PM2.5 capture. Nat. Commun. 2015;6:6205. doi: 10.1038/ncomms7205. [DOI] [PubMed] [Google Scholar]
- 34.Zhang G-H, et al. High-performance particulate matter including nanoscale particle removal by a self-powered air filter. Nat. Commun. 2020;11:1653. doi: 10.1038/s41467-020-15502-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Panet A, Baltimore D, Hanafusa T. Quantitation of avian RNA tumor virus reverse transcriptase by radioimmunoassay. J. Virol. 1975;16:146–152. doi: 10.1128/jvi.16.1.146-152.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Clark MF, Adams A. Characteristics of the microplate method of enzyme-linked immunosorbent assay for the detection of plant viruses. J. Gen. Virol. 1977;34:475–483. doi: 10.1099/0022-1317-34-3-475. [DOI] [PubMed] [Google Scholar]
- 37.Faulk WP, Taylor GM. Communication to the editors: an immunocolloid method for the electron microscope. Immunochemistry. 1971;8:1081–1083. doi: 10.1016/0019-2791(71)90496-4. [DOI] [PubMed] [Google Scholar]
- 38.Patolsky F, et al. Electrical detection of single viruses. Proc. Natl Acad. Sci. USA. 2004;101:14017–14022. doi: 10.1073/pnas.0406159101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chowdhury AD, Takemura K, Li T-C, Suzuki T, Park EY. Electrical pulse-induced electrochemical biosensor for hepatitis E virus detection. Nat. Commun. 2019;10:3737. doi: 10.1038/s41467-019-11644-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Carr MJ, et al. Development of a real-time RT-PCR for the detection of swine-lineage influenza A (H1N1) virus infections. J. Med. Virol. 2009;45:196–199. doi: 10.1016/j.jcv.2009.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Centers for Disease Control and Prevention. Interim guidelines for collecting, handling, and testing clinical specimens from persons for coronavirus disease 2019 (COVID-19) (CDC, 2020).
- 42.East-Seletsky A, et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature. 2016;538:270–273. doi: 10.1038/nature19802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Broughton JP, et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020;38:870–874. doi: 10.1038/s41587-020-0513-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gootenberg JS, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356:438–442. doi: 10.1126/science.aam9321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat. Protoc. 2019;14:2986–3012. doi: 10.1038/s41596-019-0210-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Abbott TR, et al. Development of CRISPR as an antiviral strategy to combat SARS-CoV-2 and influenza. Cell. 2020;181:856–876. doi: 10.1016/j.cell.2020.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Baaske MD, Foreman MR, Vollmer F. Single-molecule nucleic acid interactions monitored on a label-free microcavity biosensor platform. Nat. Nanotechnol. 2014;9:933–939. doi: 10.1038/nnano.2014.180. [DOI] [PubMed] [Google Scholar]
- 48.Wang M, et al. Nanopore targeted sequencing for the accurate and comprehensive detection of SARS-CoV-2 and other respiratory viruses. Small. 2020;16:2002169. doi: 10.1002/smll.202002169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Van Herwaarden A, Sarro P. Thermal sensors based on the Seebeck effect. Sens. Actuators. 1986;10:321–346. [Google Scholar]
- 50.Chung M, et al. CT imaging features of 2019 novel coronavirus (2019-nCoV) Radiology. 2020;295:202–207. doi: 10.1148/radiol.2020200230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Chen S-C, Tsai T-T, Luo C-H. Portable and programmable clinical EOG diagnostic system. J. Med. Eng. Technol. 2000;24:154–162. doi: 10.1080/03091900050163436. [DOI] [PubMed] [Google Scholar]
- 52.Sands VM, Auerbach PS, Birnbaum J, Green M. Evaluation of a portable clinical blood analyzer in the emergency department. Acad. Emerg. Med. 1995;2:172–178. doi: 10.1111/j.1553-2712.1995.tb03190.x. [DOI] [PubMed] [Google Scholar]
- 53.Li L, et al. Artificial intelligence distinguishes COVID-19 from community acquired pneumonia on chest CT. Radiology. 2020;296:E65–E71. doi: 10.1148/radiol.2020200905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.De Clercq, E. & Herdewijn, P. in Pharmaceutical Sciences Encyclopedia: Drug Discovery, Development, and Manufacturing (Wiley, 2010).
- 55.De Clercq E. Antivirals and antiviral strategies. Nat. Rev. Microbiol. 2004;2:704–720. doi: 10.1038/nrmicro975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Vigant F, Santos NC, Lee B. Broad-spectrum antivirals against viral fusion. Nat. Rev. Microbiol. 2015;13:426–437. doi: 10.1038/nrmicro3475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Wang M, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30:269–271. doi: 10.1038/s41422-020-0282-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.De Clercq E, Li G. Approved antiviral drugs over the past 50 years. Clin. Microbiol. Rev. 2016;29:695–747. doi: 10.1128/CMR.00102-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Jones ST, et al. Modified cyclodextrins as broad-spectrum antivirals. Sci. Adv. 2020;6:eaax9318. doi: 10.1126/sciadv.aax9318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Cagno V, et al. Broad-spectrum non-toxic antiviral nanoparticles with a virucidal inhibition mechanism. Nat. Mater. 2018;17:195–203. doi: 10.1038/nmat5053. [DOI] [PubMed] [Google Scholar]
- 61.Lauster D, et al. Phage capsid nanoparticles with defined ligand arrangement block influenza virus entry. Nat. Nanotechnol. 2020;15:373–379. doi: 10.1038/s41565-020-0660-2. [DOI] [PubMed] [Google Scholar]
- 62.Zhang Q, et al. Cellular nanosponges inhibit SARS-CoV-2 infectivity. Nano Lett. 2020;20:5570–5574. doi: 10.1021/acs.nanolett.0c02278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Fang RH, Kroll AV, Gao W, Zhang L. Cell membrane coating nanotechnology. Adv. Mater. 2018;30:1706759. doi: 10.1002/adma.201706759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Rao L, Tian R, Chen X. Cell-membrane-mimicking nanodecoys against infectious diseases. ACS Nano. 2020;14:2569–2574. doi: 10.1021/acsnano.0c01665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhang P, et al. Virus-mimetic nanovesicles as a versatile antigen-delivery system. Proc. Natl Acad. Sci. USA. 2015;112:E6129–E6138. doi: 10.1073/pnas.1505799112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Rao L, et al. A biomimetic nanodecoy traps zika virus to prevent viral infection and fetal microcephaly development. Nano Lett. 2018;19:2215–2222. doi: 10.1021/acs.nanolett.8b03913. [DOI] [PubMed] [Google Scholar]
- 67.Zhang Q, et al. Neutrophil membrane-coated nanoparticles inhibit synovial inflammation and alleviate joint damage in inflammatory arthritis. Nat. Nanotechnol. 2018;13:1182–1190. doi: 10.1038/s41565-018-0254-4. [DOI] [PubMed] [Google Scholar]
- 68.Hoffmann M, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271–280. doi: 10.1016/j.cell.2020.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Grein J, et al. Compassionate use of remdesivir for patients with severe Covid-19. N. Engl. J. Med. 2020;384:2327–2336. doi: 10.1056/NEJMoa2007016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.RECOVERY Collaborative Group et al. Dexamethasone in hospitalized patients with Covid-19 — preliminary report. N. Engl. J. Med.10.1056/NEJMoa2021436 (2020).
- 71.Lammers T, et al. Dexamethasone nanomedicines for COVID-19. Nat. Nanotechnol. 2020;15:622–624. doi: 10.1038/s41565-020-0752-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Wu Y, et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science. 2020;368:1274–1278. doi: 10.1126/science.abc2241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Cao Y, et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients’ B cells. Cell. 2020;181:73–84. doi: 10.1016/j.cell.2020.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Shi R, et al. A human neutralizing antibody targets the receptor binding site of SARS-CoV-2. Nature. 2020;584:120–124. doi: 10.1038/s41586-020-2381-y. [DOI] [PubMed] [Google Scholar]
- 75.Walter, J. D. et al. Synthetic nanobodies targeting the SARS-CoV-2 receptor-binding domain. Preprint at BioRxiv10.1101/2020.04.16.045419 (2020).
- 76.Wrapp D, et al. Structural basis for potent neutralization of betacoronaviruses by single-domain camelid antibodies. Cell. 2020;181:1004–1015. doi: 10.1016/j.cell.2020.04.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Zhu F-C, et al. Safety, tolerability, and immunogenicity of a recombinant adenovirus type-5 vectored COVID-19 vaccine: a dose-escalation, open-label, non-randomised, first-in-human trial. Lancet. 2020;395:1845–1854. doi: 10.1016/S0140-6736(20)31208-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.US National Library of Medicine. ClinicalTrials.govhttps://clinicaltrials.gov/ct2/show/NCT04313127 (2020).
- 79.Gao Q, et al. Development of an inactivated vaccine candidate for SARS-CoV-2. Science. 2020;369:77–81. doi: 10.1126/science.abc1932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Hotez PJ, Corry DB, Strych U, Bottazzi ME. COVID-19 vaccines: neutralizing antibodies and the alum advantage. Nat. Rev. Immunol. 2020;20:399–400. doi: 10.1038/s41577-020-0358-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Burton DR. Antibodies, viruses and vaccines. Nat. Rev. Immunol. 2002;2:706–713. doi: 10.1038/nri891. [DOI] [PubMed] [Google Scholar]
- 82.Ryan KJ, Ray CG. Medical microbiology. McGraw Hill. 2004;4:370. [Google Scholar]
- 83.Wang J, et al. Pulmonary surfactant–biomimetic nanoparticles potentiate heterosubtypic influenza immunity. Science. 2020;367:eaau0810. doi: 10.1126/science.aau0810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Herold S, Sander L-E. Toward a universal flu vaccine. Science. 2020;367:852–853. doi: 10.1126/science.aba2754. [DOI] [PubMed] [Google Scholar]
- 85.Coffman RL, Sher A, Seder RA. Vaccine adjuvants: putting innate immunity to work. Immunity. 2010;33:492–503. doi: 10.1016/j.immuni.2010.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 2018;17:261–279. doi: 10.1038/nrd.2017.243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Riley RS, June CH, Langer R, Mitchell MJ. Delivery technologies for cancer immunotherapy. Nat. Rev. Drug Discov. 2019;18:175–196. doi: 10.1038/s41573-018-0006-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Kong N, et al. Synthetic mRNA nanoparticle-mediated restoration of p53 tumor suppressor sensitizes p53-deficient cancers to mTOR inhibition. Sci. Transl. Med. 2019;11:eaaw1565. doi: 10.1126/scitranslmed.aaw1565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Richner JM, et al. Modified mRNA vaccines protect against Zika virus infection. Cell. 2017;168:1114–1125. e1110. doi: 10.1016/j.cell.2017.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.US National Library of Medicine. ClinicalTrials.govhttps://clinicaltrials.gov/ct2/show/NCT04368728 (2020).
- 91.Vogel, A. et al. A prefusion SARS-CoV-2 spike RNA vaccine is highly immunogenic and prevents lung infection in non-human primates. Preprint at bioRxiv10.1101/2020.09.08.280818 (2020).
- 92.US National Library of Medicine. Clinicaltrials.govhttps://clinicaltrials.gov/ct2/show/NCT04232280 (2020).
- 93.US National Library of Medicine. ClinicalTrials.govhttps://clinicaltrials.gov/ct2/show/NCT04470427 (2020).
- 94.Mustafa AK, et al. Extracorporeal membrane oxygenation for patients with COVID-19 in severe respiratory failure. JAMA Surg. 2020 doi: 10.1001/jamasurg.2020.3950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Brodie D, Bacchetta M. Extracorporeal membrane oxygenation for ARDS in adults. N. Engl. J. Med. 2011;365:1905–1914. doi: 10.1056/NEJMct1103720. [DOI] [PubMed] [Google Scholar]
- 96.Ivanovsky D. Über die Mosaikkrankheit der Tabakspflanze. St. Petersb. Acad. Imp. Sci. Bull. 1892;35:67–70. [Google Scholar]
- 97.Drinker P, Shaw LA. The prolonged administration of artificial respiration. J. Franklin Inst. 1932;213:355–372. [Google Scholar]
- 98.Knoll M, Ruska E. Das elektronenmikroskop. Z. Phys. 1932;78:318–339. [Google Scholar]
- 99.Ruska HDie. Sichtbarmachung der bakteriophagen lyse im übermikroskop. Naturwissenschaften. 1940;28:45–46. [Google Scholar]
- 100.Franks A. An optically focusing X-ray diffraction camera. Proc. Phys. Soc. B. 1955;68:1054. [Google Scholar]
- 101.US Food and Drug Administration. Drugs@FDA: FDA-approved drugs. FDAhttps://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=reportsSearch.process (1963).
- 102.Rashkind W, Freeman A, Klein D, Toft R. Evaluation of a disposable plastic, low volume, pumpless oxygenator as a lung substitute. J. Pediatr. 1965;66:94–102. doi: 10.1016/s0022-3476(65)80342-0. [DOI] [PubMed] [Google Scholar]
- 103.Schena M, Shalon D, Davis RW, Brown PO. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995;270:467–470. doi: 10.1126/science.270.5235.467. [DOI] [PubMed] [Google Scholar]
- 104.US Food and Drug Administration. Highlights of prescribing information Doxil (FDA, 1995).
- 105.US Food and Drug Administration. FDA information regarding FluMist quadrivalent vaccine. FDAhttps://www.fda.gov/vaccines-blood-biologics/vaccines/fda-information-regarding-flumist-quadrivalent-vaccine (2018).
- 106.Eisenstein M. Oxford Nanopore announcement sets sequencing sector abuzz. Nat. Biotechnol. 2012;30:295–296. doi: 10.1038/nbt0412-295. [DOI] [PubMed] [Google Scholar]
- 107.Murray BK, et al. Virion disruption by ozone-mediated reactive oxygen species. J. Virol. Methods. 2008;153:74–77. doi: 10.1016/j.jviromet.2008.06.004. [DOI] [PubMed] [Google Scholar]
- 108.Almeida A, Faustino MAF, Neves MG. Antimicrobial photodynamic therapy in the control of COVID-19. Antibiotics. 2020;9:320. doi: 10.3390/antibiotics9060320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Li P, et al. Metal-organic frameworks with photocatalytic bactericidal activity for integrated air cleaning. Nat. Commun. 2019;10:2177. doi: 10.1038/s41467-019-10218-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Centers for Disease Control and Prevention. Implementing filtering facepiece respirator (FFR) reuse, including reuse after decontamination, when there are known shortages of N95 respirators (CDC, 2020).
- 111.Koo W-T, et al. Hierarchical metal–organic framework-assembled membrane filter for efficient removal of particulate matter. ACS Appl. Mater. Interfaces. 2018;10:19957–19963. doi: 10.1021/acsami.8b02986. [DOI] [PubMed] [Google Scholar]
- 112.Hao Z, Wu J, Wang C, Liu J. Electrospun polyimide/metal-organic framework nanofibrous membrane with superior thermal stability for efficient PM2.5 capture. ACS Appl. Mater. Interfaces. 2019;11:11904–11909. doi: 10.1021/acsami.8b22415. [DOI] [PubMed] [Google Scholar]
- 113.Liao L, et al. Can N95 respirators be reused after disinfection? How many times? ACS Nano. 2020;14:6348–6356. doi: 10.1021/acsnano.0c03597. [DOI] [PubMed] [Google Scholar]
- 114.Chen Z, Hsu F-C, Battigelli D, Chang H-C. Capture and release of viruses using amino-functionalized silica particles. Anal. Chim. Acta. 2006;569:76–82. [Google Scholar]
- 115.Badrossamay MR, McIlwee HA, Goss JA, Parker KK. Nanofiber assembly by rotary jet-spinning. Nano Lett. 2010;10:2257–2261. doi: 10.1021/nl101355x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Murphy SV, Atala A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 2014;32:773–785. doi: 10.1038/nbt.2958. [DOI] [PubMed] [Google Scholar]
- 117.Choong YYC, et al. The global rise of 3D printing during the COVID-19 pandemic. Nat. Rev. Mater. 2020;5:637–639. doi: 10.1038/s41578-020-00234-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Ong SWX, et al. Air, surface environmental, and personal protective equipment contamination by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) from a symptomatic patient. JAMA. 2020;323:1610–1612. doi: 10.1001/jama.2020.3227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Dey P, et al. Multivalent flexible nanogels exhibit broad-spectrum antiviral activity by blocking virus entry. ACS Nano. 2018;12:6429–6442. doi: 10.1021/acsnano.8b01616. [DOI] [PubMed] [Google Scholar]
- 120.Berensmeier S. Magnetic particles for the separation and purification of nucleic acids. Appl. Microbiol. Biotechnol. 2006;73:495–504. doi: 10.1007/s00253-006-0675-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Reinholt SJ, Baeumner AJ. Microfluidic isolation of nucleic acids. Angew. Chem. Int. Ed. 2014;53:13988–14001. doi: 10.1002/anie.201309580. [DOI] [PubMed] [Google Scholar]
- 122.Taddei C, et al. Fabrication of polystyrene-encapsulated magnetic iron oxide nanoparticles via batch and microfluidic-assisted production. Colloid Polym. Sci. 2019;297:861–870. [Google Scholar]
- 123.Unni M, et al. Engineering magnetic nanoparticles and their integration with microfluidics for cell isolation. J. Colloid Interface Sci. 2020;564:204–215. doi: 10.1016/j.jcis.2019.12.092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Kwon PS, et al. Designer DNA architecture offers precise and multivalent spatial pattern-recognition for viral sensing and inhibition. Nat. Chem. 2020;12:26–35. doi: 10.1038/s41557-019-0369-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Geary C, Rothemund PW, Andersen ES. A single-stranded architecture for cotranscriptional folding of RNA nanostructures. Science. 2014;345:799–804. doi: 10.1126/science.1253920. [DOI] [PubMed] [Google Scholar]
- 126.Nankai University. Fast testing only Takes 15 minutes: Nankai University’s team developed antibody test kit for novel coronavirus. Nankai Univ.http://en.nankai.edu.cn/2020/0219/c14577a265299/page.htm (2020)
- 127.Singh N, Sharma M, Mondal D, Pereira MM, Prasad K. Very high concentration solubility and long-term stability of DNA in an ammonium-based ionic liquid: A suitable medium for nucleic acid packaging and preservation. ACS Sustain. Chem. Eng. 2017;5:1998–2005. [Google Scholar]
- 128.Zhu C, Varona M, Anderson JL. Magnetic ionic liquids as solvents for RNA extraction and preservation. ACS Omega. 2020;5:11151–11159. doi: 10.1021/acsomega.0c01098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Hu P, et al. Fe–Au nanoparticle-coupling for ultrasensitive detections of circulating tumor DNA. Adv. Mater. 2018;30:1801690. doi: 10.1002/adma.201801690. [DOI] [PubMed] [Google Scholar]
- 130.Wang D, et al. Microarray-based detection and genotyping of viral pathogens. Proc. Natl Acad. Sci. USA. 2002;99:15687–15692. doi: 10.1073/pnas.242579699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Lim D-K, et al. Highly uniform and reproducible surface-enhanced Raman scattering from DNA-tailorable nanoparticles with 1-nm interior gap. Nat. Nanotechnol. 2011;6:452–460. doi: 10.1038/nnano.2011.79. [DOI] [PubMed] [Google Scholar]
- 132.Di Zhang LH, Liu B, Ge Q, Dong J, Zhao X. Rapid and ultrasensitive quantification of multiplex respiratory tract infection pathogen via lateral flow microarray based on SERS nanotags. Theranostics. 2019;9:4849–4859. doi: 10.7150/thno.35824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Rao L, et al. A microfluidic electrostatic separator based on pre-charged droplets. Sens. Actuators B. Chem. 2015;210:328–335. [Google Scholar]
- 134.Rao L, et al. Microfluidic electroporation-facilitated synthesis of erythrocyte membrane-coated magnetic nanoparticles for enhanced imaging-guided cancer therapy. ACS Nano. 2017;11:3496–3505. doi: 10.1021/acsnano.7b00133. [DOI] [PubMed] [Google Scholar]
- 135.Sánchez-Purrà M, et al. Surface-enhanced Raman spectroscopy-based sandwich immunoassays for multiplexed detection of Zika and Dengue viral biomarkers. ACS Infect. Dis. 2017;3:767–776. doi: 10.1021/acsinfecdis.7b00110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Bertrand N, et al. Mechanistic understanding of in vivo protein corona formation on polymeric nanoparticles and impact on pharmacokinetics. Nat. Commun. 2017;8:777. doi: 10.1038/s41467-017-00600-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Hadjidemetriou M, Al-Ahmady Z, Buggio M, Swift J, Kostarelos K. A novel scavenging tool for cancer biomarker discovery based on the blood-circulating nanoparticle protein corona. Biomaterials. 2019;188:118–129. doi: 10.1016/j.biomaterials.2018.10.011. [DOI] [PubMed] [Google Scholar]
- 138.Pattipeiluhu R, et al. Unbiased identification of the liposome protein corona using photoaffinity-based chemoproteomics. ACS Cent. Sci. 2020;6:535–545. doi: 10.1021/acscentsci.9b01222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Blume JE, et al. Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona. Nat. Commun. 2020;11:3662. doi: 10.1038/s41467-020-17033-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Schmidt C. Bioelectronics: The bionic material. Nature. 2012;483:S37. doi: 10.1038/483S37a. [DOI] [PubMed] [Google Scholar]
- 141.Yang Y, et al. A laser-engraved wearable sensor for sensitive detection of uric acid and tyrosine in sweat. Nat. Biotechnol. 2020;38:217–224. doi: 10.1038/s41587-019-0321-x. [DOI] [PubMed] [Google Scholar]
- 142.Lee H, et al. A graphene-based electrochemical device with thermoresponsive microneedles for diabetes monitoring and therapy. Nat. Nanotechnol. 2016;11:566–572. doi: 10.1038/nnano.2016.38. [DOI] [PubMed] [Google Scholar]
- 143.Nakata S, Arie T, Akita S, Takei K. Wearable, flexible, and multifunctional healthcare device with an ISFET chemical sensor for simultaneous sweat pH and skin temperature monitoring. ACS Sens. 2017;2:443–448. doi: 10.1021/acssensors.7b00047. [DOI] [PubMed] [Google Scholar]
- 144.Tan CSH, et al. Thermal proximity coaggregation for system-wide profiling of protein complex dynamics in cells. Science. 2018;359:1170–1177. doi: 10.1126/science.aan0346. [DOI] [PubMed] [Google Scholar]
- 145.Molina DM, et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science. 2013;341:84–87. doi: 10.1126/science.1233606. [DOI] [PubMed] [Google Scholar]
- 146.Drew DA, et al. Rapid implementation of mobile technology for real-time epidemiology of COVID-19. Science. 2020;368:1362–1367. doi: 10.1126/science.abc0473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Zhang K, et al. Clinically applicable AI system for accurate diagnosis, quantitative measurements and prognosis of COVID-19 pneumonia using computed tomography. Cell. 2020;181:1423–1433. doi: 10.1016/j.cell.2020.04.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Pan W, et al. Visualizing influenza virus infection in living mice. Nat. Commun. 2013;4:2369–2377. doi: 10.1038/ncomms3369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Rameix-Welti M-A, et al. Visualizing the replication of respiratory syncytial virus in cells and in living mice. Nat. Commun. 2014;5:5104. doi: 10.1038/ncomms6104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Karlsson EA, et al. Visualizing real-time influenza virus infection, transmission and protection in ferrets. Nat. Commun. 2015;6:6378. doi: 10.1038/ncomms7378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Finke S, Brzózka K, Conzelmann K-K. Tracking fluorescence-labeled rabies virus: enhanced green fluorescent protein-tagged phosphoprotein P supports virus gene expression and formation of infectious particles. J. Virol. 2004;78:12333–12343. doi: 10.1128/JVI.78.22.12333-12343.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Lux K, et al. Green fluorescent protein-tagged adeno-associated virus particles allow the study of cytosolic and nuclear trafficking. J. Virol. 2005;79:11776–11787. doi: 10.1128/JVI.79.18.11776-11787.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Klingen Y, Conzelmann K-K, Finke S. Double-labeled rabies virus: live tracking of enveloped virus transport. J. Virol. 2008;82:237–245. doi: 10.1128/JVI.01342-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Cammarata M, et al. Tracking the structural dynamics of proteins in solution using time-resolved wide-angle X-ray scattering. Nat. Methods. 2008;5:881–886. doi: 10.1038/nmeth.1255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Junttila MR, de Sauvage FJ. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature. 2013;501:346–354. doi: 10.1038/nature12626. [DOI] [PubMed] [Google Scholar]
- 156.Rossi G, Manfrin A, Lutolf MP. Progress and potential in organoid research. Nat. Rev. Genet. 2018;19:671–687. doi: 10.1038/s41576-018-0051-9. [DOI] [PubMed] [Google Scholar]
- 157.Zhou T, et al. High-content screening in hPSC-neural progenitors identifies drug candidates that inhibit Zika virus infection in fetal-like organoids and adult brain. Cell Stem Cell. 2017;21:274–283.e5. doi: 10.1016/j.stem.2017.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Watanabe M, et al. Self-organized cerebral organoids with human-specific features predict effective drugs to combat Zika virus infection. Cell Rep. 2017;21:517–532. doi: 10.1016/j.celrep.2017.09.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Duan, X. et al. Identification of drugs blocking SARS-CoV-2 infection using human pluripotent stem cell-derived colonic organoids. Preprint at bioRxiv10.1101/2020.05.02.073320 (2020).
- 160.Zhou J, et al. Infection of bat and human intestinal organoids by SARS-CoV-2. Nat. Med. 2020;26:1077–1083. doi: 10.1038/s41591-020-0912-6. [DOI] [PubMed] [Google Scholar]
- 161.Monteil V, et al. Inhibition of SARS-CoV-2 infections in engineered human tissues using clinical-grade soluble human ACE2. Cell. 2020;181:905–913. doi: 10.1016/j.cell.2020.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Aisenbrey EA, Murphy WL. Synthetic alternatives to Matrigel. Nat. Rev. Mater. 2020;5:539–551. doi: 10.1038/s41578-020-0199-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Kratochvil MJ, et al. Engineered materials for organoid systems. Nat. Rev. Mater. 2019;4:606–622. doi: 10.1038/s41578-019-0129-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Wang C, et al. Molecularly engineered poly(ortho ester) microspheres for enhanced delivery of DNA vaccines. Nat. Mater. 2004;3:190–196. doi: 10.1038/nmat1075. [DOI] [PubMed] [Google Scholar]
- 165.Miao L, et al. Delivery of mRNA vaccines with heterocyclic lipids increases anti-tumor efficacy by STING-mediated immune cell activation. Nat. Biotechnol. 2019;37:1174–1185. doi: 10.1038/s41587-019-0247-3. [DOI] [PubMed] [Google Scholar]
- 166.Zhang N-N, et al. A thermostable mRNA vaccine against COVID-19. Cell. 2020;182:1271–1283. doi: 10.1016/j.cell.2020.07.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.John ALS, Chan CY, Staats HF, Leong KW, Abraham SN. Synthetic mast-cell granules as adjuvants to promote and polarize immunity in lymph nodes. Nat. Mater. 2012;11:250–257. doi: 10.1038/nmat3222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Liu H, et al. Structure-based programming of lymph-node targeting in molecular vaccines. Nature. 2014;507:519–522. doi: 10.1038/nature12978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Schudel A, Francis DM, Thomas SN. Material design for lymph node drug delivery. Nat. Rev. Mater. 2019;4:415–428. doi: 10.1038/s41578-019-0110-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Wilson JT, et al. pH-Responsive nanoparticle vaccines for dual-delivery of antigens and immunostimulatory oligonucleotides. ACS Nano. 2013;7:3912–3925. doi: 10.1021/nn305466z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Zhang C, et al. A light responsive nanoparticle-based delivery system using pheophorbide A graft polyethylenimine for dendritic cell-based cancer immunotherapy. Mol. Pharm. 2017;14:1760–1770. doi: 10.1021/acs.molpharmaceut.7b00015. [DOI] [PubMed] [Google Scholar]
- 172.Zhong Y, et al. In vivo molecular imaging for immunotherapy using ultra-bright near-infrared-IIb rare-earth nanoparticles. Nat. Biotechnol. 2019;37:1322–1331. doi: 10.1038/s41587-019-0262-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Lu L, et al. NIR-II bioluminescence for in vivo high contrast imaging and in situ ATP-mediated metastases tracing. Nat. Commun. 2020;11:4192. doi: 10.1038/s41467-020-18051-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Yang Y, et al. NIR-II chemiluminescence molecular sensor for in vivo high-contrast inflammation imaging. Angew. Chem. Int. Ed. 2020;59:18380. doi: 10.1002/anie.202007649. [DOI] [PubMed] [Google Scholar]
- 175.Li B, et al. Organic NIR-II molecule with long blood half-life for in vivo dynamic vascular imaging. Nat. Commun. 2020;11:3102. doi: 10.1038/s41467-020-16924-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.McHugh KJ, et al. Biocompatible near-infrared quantum dots delivered to the skin by microneedle patches record vaccination. Sci. Transl. Med. 2019;11:eaay7162. doi: 10.1126/scitranslmed.aay7162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Kaushik N, et al. Zinc salts block hepatitis E virus replication by inhibiting the activity of viral RNA-dependent RNA polymerase. J. Virol. 2017;91:e00754–00717. doi: 10.1128/JVI.00754-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Boivin G, et al. Rapid antiviral effect of inhaled zanamivir in the treatment of naturally occurring influenza in otherwise healthy adults. J. Infect. Dis. 2000;181:1471–1474. doi: 10.1086/315392. [DOI] [PubMed] [Google Scholar]
- 179.Garcia-Contreras L, Awashthi S, Hanif S, Hickey A. Inhaled vaccines for the prevention of tuberculosis. J. Mycobac Dis. 2012;S1:002. [Google Scholar]
- 180.Willis L, Hayes D, Mansour HM. Therapeutic liposomal dry powder inhalation aerosols for targeted lung delivery. Lung. 2012;190:251–262. doi: 10.1007/s00408-011-9360-x. [DOI] [PubMed] [Google Scholar]
- 181.Elmezayen, A. D., Al-Obaidi, A., Şahin, A. T. & Yelekçi, K. Drug repurposing for coronavirus (COVID-19): in silico screening of known drugs against coronavirus 3CL hydrolase and protease enzymes. J. Biomol. Struct. Dyn. 10.1080/07391102.2020.1758791 (2020). [DOI] [PMC free article] [PubMed]
- 182.Huang KJ, et al. An interferon-γ-related cytokine storm in SARS patients. J. Med. Virol. 2005;75:185–194. doi: 10.1002/jmv.20255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Rothman AL. Immunity to dengue virus: a tale of original antigenic sin and tropical cytokine storms. Nat. Rev. Immunol. 2011;11:532–543. doi: 10.1038/nri3014. [DOI] [PubMed] [Google Scholar]
- 184.Tisoncik JR, et al. Into the eye of the cytokine storm. Microbiol. Mol. Biol. Rev. 2012;76:16–32. doi: 10.1128/MMBR.05015-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Pedersen SF, Ho Y-C. SARS-CoV-2: a storm is raging. J. Clin. Invest. 2020;130:2202–2205. doi: 10.1172/JCI137647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Dormont F, et al. Squalene-based multidrug nanoparticles for improved mitigation of uncontrolled inflammation in rodents. Sci. Adv. 2020;6:eaaz5466. doi: 10.1126/sciadv.aaz5466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Zhang CY, Gao J, Wang Z. Bioresponsive nanoparticles targeted to infectious microenvironments for sepsis management. Adv. Mater. 2018;30:e1803618. doi: 10.1002/adma.201803618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Ma Q, et al. Calming cytokine storm in pneumonia by targeted delivery of TPCA-1 using platelet-derived extracellular vesicles. Matter. 2020;3:287–301. doi: 10.1016/j.matt.2020.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Henry BM. COVID-19, ECMO, and lymphopenia: a word of caution. Lancet Respir. Med. 2020;8:e24. doi: 10.1016/S2213-2600(20)30119-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Chlebowski MM, Baltagi S, Carlson M, Levy JH, Spinella PC. Clinical controversies in anticoagulation monitoring and antithrombin supplementation for ECMO. Crit. Care. 2020;24:19–30. doi: 10.1186/s13054-020-2726-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Tulman DB, et al. Veno-venous ECMO: a synopsis of nine key potential challenges, considerations, and controversies. BMC Anesthesiol. 2014;14:65. doi: 10.1186/1471-2253-14-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Motomura T, et al. Development of silicone rubber hollow fiber membrane oxygenator for ECMO. Artif. Organs. 2003;27:1050–1053. doi: 10.1046/j.1525-1594.2003.07077.x. [DOI] [PubMed] [Google Scholar]
- 193.Agati S, et al. DIDECMO: A new polymethylpentene oxygenator for pediatric extracorporeal membrane oxygenation. ASAIO J. 2006;52:509–512. doi: 10.1097/01.mat.0000237693.09432.13. [DOI] [PubMed] [Google Scholar]
- 194.Bhamla MS, et al. Hand-powered ultralow-cost paper centrifuge. Nat. Biomed. Eng. 2017;1:0009. [Google Scholar]
- 195.Li, E., Larson, A., Kothari, A. & Prakash, M. Handyfuge-LAMP: low-cost and electricity-free centrifugation for isothermal SARS-CoV-2 detection in saliva. Preprint at medRxiv10.1101/2020.06.30.20143255 (2020).
- 196.Li, H., Soto-Montoya, H., Voisin, M., Valenzuela, L. F. & Prakash, M. Octopi: open configurable high-throughput imaging platform for infectious disease diagnosis in the field. Preprint at bioRxiv10.1101/684423 (2019).
- 197.Hol, F. J., Lambrechts, L. & Prakash, M. BiteOscope, an open platform to study mosquito biting behavior. eLife9, e56829 (2020). [DOI] [PMC free article] [PubMed]
- 198.Krishnamurthy D, et al. Scale-free vertical tracking microscopy. Nat. Methods. 2020;17:1040–1051. doi: 10.1038/s41592-020-0924-7. [DOI] [PubMed] [Google Scholar]
- 199.Zhang, F., Abudayyeh, O. O. & Gootenberg, J. S. A protocol for detection of COVID-19 using CRISPR diagnostics (Broad Institute, 2020).