Abstract
The COVID-19 pandemic has spread worldwide since the outbreak in Wuhan (China) in December 2019, currently infecting over 25 million people and causing more than 800.000 deaths. In Ecuador, up to the 30th of August 2020, overall 113.648 confirmed cases and 6.555 deaths have been declared. Besides overloading of hospital, capacity for molecular diagnosis of SARS-CoV-2 by the Ministry of Public Health was quickly overwhelmed. In this context, emergency authorization for SARS-CoV-2 RT-qPCR diagnosis was granted to other institutions, like the “Agencia de Regulación y Control para la Bioseguridad y Cuarentena de Galápagos” (ABG), that implemented “LabGal” with support of One Health Research Group from “Universidad de Las Américas” (UDLA). The previous experience of ABG and the One Health Research Group in conducting massive surveillance of zoonotic diseases on livestock was crucial to the success on the control of COVID-19 outbreak at Galapagos Islands by the end of May 2020, when Latin American countries were leading the spread of the pandemic.
1. Introduction
The world is facing a pandemic caused by a novel coronavirus named SARS-CoV-2 that has been called Coronavirus Disease 2019 (COVID-19) by the World Health Organization (WHO) [1,2]. From an initial outbreak in the Chinese province of Hubei (Wuhan city) in December 2019 [1], the virus spread worldwide in few weeks till a pandemic was officially declared by WHO on March 11th 2020. COVID-19 represents the worst public health crisis since the Spanish flu in 1918 with more than 25 million people infected and 800.000 deaths worldwide by mid-August 2020.
In Ecuador, up to the 30th of August 2020, overall 113.648 confirmed cases and 6.555 deaths have been confirmed (data source: Ministry of Public Health from Ecuador, MSP). The first SARS-CoV-2 infection was identified in Guayaquil city (most populous city in Ecuador) on February 29th. Within a few weeks, COVID-19 had challenged the Ecuadorian public healthcare system and overwhelmed it, particularly in Guayaquil city, where the worst outbreak took place, and from which dramatic images of dead bodies on its streets were broadcast worldwide. The Ecuadorian Government's declaration of a public health emergency came by March 14th 2020, coupled with the establishment of draconian measures, such as closure of all stores except for pharmacies, groceries, banking and other social essential services, and by restricting movement exclusively to activities related to these enterprises from 6 AM to 2 PM. Those measures slowed down the spread of the virus and somehow “flattened the curve” of new cases according to MSP. However, since the beginning of July 2020, step by step restoration towards a “new normality” has led to new outbreaks countrywide. Movement restrictions have been only partially lifted depending on the epidemiological situation of each canton and Ecuadorian public health authorities still urging people to wear masks and observe social distancing.
COVID-19 surveillance programs relies on RT-qPCR-based molecular diagnosis of the SARS-CoV-2 virus. This technique depends on qualified personnel and expensive equipment and supplies. As this diagnostic tool is mandatory for an accurate collection of epidemiological data to understand how the pandemic is progressing, the need of performing thousands of tests per day has challenged almost every public health system worldwide and overwhelmed testing capacity in developing countries like Ecuador. MSP only authorized public laboratories for SARS-CoV-2 diagnosis within the network of “Instituto Nacional de Salud Publica e Investigacion” located only in the three major cities of Ecuador (Quito, Guayaquil and Cuenca) at the beginning of the pandemic. However, once this approach was overwhelmed, MSP granted authorization for SARS-CoV2 diagnosis to new labs.
The first laboratory to obtain those credentials was “LabGal”, the laboratory for SARS-CoV-2 diagnosis at Galapagos Islands, located at Agencia de Regulacion y Control de la Bioseguridad y Cuarentena para Galapagos (ABG), that started to run SARS-CoV-2 molecular diagnosis tests from April 7th 2020, with equipment and personnel support for One Health Research Group from Universidad de Las Americas (UDLA, Quito, Ecuador). ABG is a public institution within the Ecuadorian Ministry of Environment that controls companion and livestock animal health in Galapagos Islands with a One Health scope to guarantee good quality animal products for human consumption and conservation of the unique biodiversity within Galapagos Islands. ABG and One Health Research Group from UDLA have been actively working on zoonotic diseases surveillance on Galapagos Islands prior to COVID-19 outbreak.
This aim of this manuscript is to describe the first 8 weeks of experience gained by the One Health Research team composed of ABG and UDLA in the diagnosis of SARS-CoV-2 in support of the MSP for COVID-19 surveillance at the Galapagos Islands in Ecuador, after the first case of COVID-19 in Galapagos Islands was recorded by mid march 2020 in a male patient travelling from Guayaquil city, till the COVID-19 outbreak was controlled and Galapagos Islands.
2. Materials and methods
Samples tested for the presence of SARS-CoV-2 RNA were collected from the respiratory tract of individuals which were either hospitalized, or screened due to contact history with SARS-CoV-2 positive individuals. Samples of hospitalized individuals were collected from the only two hospitals in Galapagos, located in Santa Cruz Island and San Cristobal Island (“Hospital República del Ecuador” and “Hospital Oskar Jandl”, respectively).
The workflow for SARS-CoV-2 RNA detection was composed by two steps. The first step includes virus collection from nasopharyngeal swabs on 0.5uL TRIS-EDTA pH 8, followed by RNA purification using several commercial kits that were validated prior to their clinical use, including brands like Invitrogen (USA), Quiagen (USA) and Bioneer (South Korea). Research experience of UDLA-ABG team was crucial to adapt RNA extraction protocols to several kits brands availability while maintaining diagnosis quality.
The second step consists of RNA detection by using the CDC FDA EUA 2019-nCoV Kit (IDT, USA), whose results are interpreted following the CDC protocol [3,4]. Briefly, this kit is based on N1 and N2 probes to detect SARS-CoV-2 that have received positive sensitivity and specificity evaluation on recent reports [[5], [6], [7]]; and RNase P as an RNA extraction quality control. A limit of detection (LOD) as low as 1 viral copy per microliter of RNA extraction solution was set. This low LOD was a fundamental tool for a successful control strategy implemented by the epidemiologist from Pan American Health Organization leading the COVID-19 outbreak surveillance on the Galapagos Islands.
The laboratory operated with a single biosafety hood for RNA extraction (BSL-2). The lack of a real time PCR thermo-cycler was solved with a donation of a CFX96 (BioRad) device from UDLA.
3. Results
Massive SARS-CoV-2 molecular diagnosis for a successful surveillance program at Galapagos Islands.
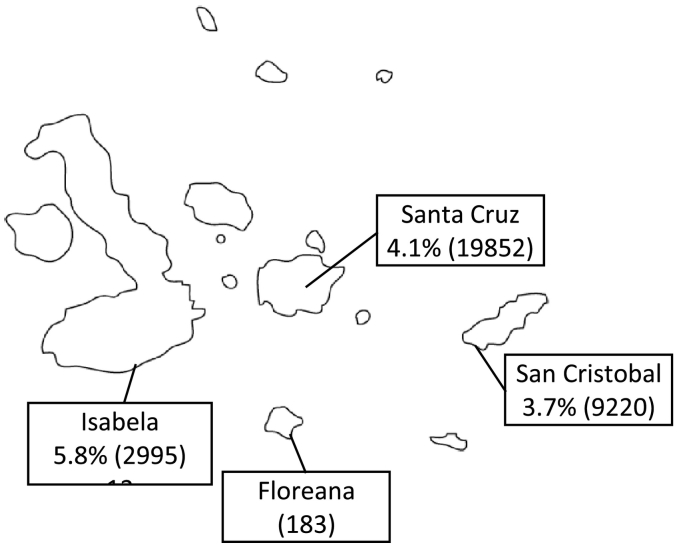
Starting from April 7th 2020 and up to May 1st 2020 a total 1691 samples were processed at “LabGal” (ABG). In the first week of testing, not more than 55 samples per day were tested, but in the following days the laboratory capacity was increased up to around 110 samples/day. There was a four days diagnosis disruption due to supplies shortage of RNA extraction kits (Fig. 1). Overall, 107 out of 1691 (6.3%) samples tested positive by RT-qPCR. So far, within a month, 5.41% of the total population of Galapagos Islands (31.280 according to “Instituto Nacional de Estadísticas y Censos” de Ecuador) was screened, while less than 0.25% of the continental Ecuadorian population was tested in the same period. Population distribution and percentage of population screened for SARS-CoV-2 by RT-PCR for each of the four inhabited islands of Santa Cruz, Isabela, San Cristobal and Floreana are described on Fig. 2. There was no testing at Floreana island, as the 183 inhabitants locked the island not allowing any medical brigade to access the island by the time this study was carried.
Fig. 1.
Temporal distribution of samples tested (total al positives) for SARS-CoV-2 RT-qPCR during the period from April 7th to May 1st 2020 at “LabGal”.
Fig. 2.
Galapagos Island Map showing the total population of each of the inhabited islands and the percentage of population tested for SARS-CoV-2. No tests were performed at Floreana due to the total lock of the island.
From the total 126 individuals that tested SARS-CoV-2 positive, 59 ones were detected at a single isolated location, a cruise boat located close to Santa Cruz Island. Although due to borders closure, no tourists were at the cruise at the time this study was performed, 61 crew members were set at quarantine inside the boat. This COVID-19 outbreak was successfully controlled with weekly testing of the whole crew with only one patient evacuation needed due to comorbidity background.
Excluding the SARS-CoV-2 positives cases from the cruise outbreak, we found 48 positives cases among population in Santa Cruz, Isabela and San Cristobal Islands, out of 1630 individuals tested. That means a positivity rate of 2.94%, making Galapagos Islands the only province of Ecuador to date achieving the WHO recommendations of a 5% positivity rate as a proxy for a successful surveillance program.
Once the COVID-19 outbreak was controlled by massive testing from April 7th till May 1st 2020, sentinel control of positive cases was follow up and by May 29th 2020, during a official visit of Vice President of Ecuador and Minister of Public Health, Galapagos Islands were declared free of SARS-CoV-2 [8].
3.1. Sample Pooling strategy to speed up SARS-CoV-2 diagnosis
After the first week of surveillance, once that preliminary data show a SARS-CoV-2 prevalence below 10%, a sample pooling protocol was implemented. As the CDC protocol requires three PCR reactions per sample for SARS-CoV2 diagnosis, we pooled 3 samples after RNA extraction into the same PCR reaction, showing that no shift on Ct values for N1 and N2 probes happened. This strategy allowed us to speed up diagnosis in our lab operating with a single 96 well thermo cycler, while keeping the same sensitivity that on single sample PCR reactions [9].
3.2. Cotton tipped plastic swabs for SARS-CoV-2 diagnosis to prevent supplies shortage
As we described above, we experienced a diagnosis disruption due to RNA extraction kits shortage. To prevent a new disruption due to the lack of swabs that Ecuador was experiencing, we carried out a validation study of cheap and locally produced cotton swabs for nasopharyngeal specimen collection prior to SARS-CoV-2 diagnosis [10]. Although CDC and WHO only endorse synthetic fiber swabs for SARS-CoV-2 diagnosis [11,12], our previous experience as ABG-One Health Research Group (UDLA) on zoonotic diseases surveillance supported our hypothesis that cotton swabs would be useful for sample collection prior to PCR based-diagnosis. These findings were not only useful to avoid new SARS-CoV-2 diagnosis disruptions at “LabGal” but also for other labs on continental Ecuador.
4. Discussion
Diagnosis of SARS-CoV-2 has been mainly performed in Ecuador following a traditional centralized schedule dependent on the network of laboratories from “Instituto Nacional de Salud Pública e Investigación” (MSP), with a capacity of approximately 3000–4000 samples processed daily for SARS-CoV-2 for the whole country (more than 17 million people). This diagnosis capacity has been mainly targeting symptomatic population attending public hospitals, leading to a positivity rate above 30%, clearly over WHO recommendations.
In Galapagos Islands, SARS-CoV-2 diagnosis and COVID-19 surveillance was carried out of by “LabGal”, a partnership between One Health Research Group for “Universidad de Las Americas” (UDLA) and “Agencia de Regulación y Control de la Bioseguridad y Cuarentena para Galápagos” (ABG), with support from MSP and WHO for surveillance strategy and samples collection. This scenario was possible for two main reasons: 1) ABG is a government agency, so coordination with MSP was an easy goal; 2) ABG is not only a regulatory but also a research institution actively collaborating in One Health research with UDLA. Moreover, the ABG-UDLA consortium has the technical and scientific capacities to support the extraordinary surge in demand for diagnostic testing of human samples not ever seen before COVID-19 pandemic. These skills for good biosafety and biosecurity performance, and high throughput and quality testing for the surveillance and control of infectious diseases is coming for our previous institutional collaboration on research projects related to surveillance of zoonotic diseases like brucellosis, Q-fever or leptospirosis in livestock and dogs at Galapagos Islands (several articles under preparation, currently delayed due to COVID-19), where thousands of animals were tested within few weeks in the second half of 2019.
Islands at remote oceanic locations offer an extraordinary scenario for infectious diseases control and erradication. For instance, ABG has previous experience declaring Galapagos Islands as a free territory of Classical Swine Fever certified by World Organization for Animal Health [13], and ABG and UDLA are currently working on the brucellosis free status for the islands. Moreover, in the context of COVID-19 pandemic, the last territories remaining free of SARS-CoV-2 or where the outbreaks were quickly controlled are the Pacific Small Islands Developing States [14]. Iceland is another example of successful surveillance and control of COVID-19 [15]. As we have described on the results, flights were banned to Galapagos Islands by the time this study was carried out and also restriction to travel among islands was implemented. Under this epidemiological scenario, by May 29th 2020, during an official visit of Vice President of Ecuador and Minister of Public Health, Galapagos Islands were declared free of SARS-CoV-2 [8].
Since July 2020th, restriction to flights were partially lifted for Galapagos Islands, and this scenario has lead to a new outbreak of around 70 new cases of SARS-CoV-2 in the islands. The experience described in this manuscript for April and May 2020 has been fundamental to control the new outbreak.
Moreover, future strategies for SARS-CoV-2 containment are being designed by ABG and One Health Research Group (UDLA), including not only RT-qPCR but also massive serological testing as long as flights and touristic activities are recovered. Again, our previous experience as One Health Research Group (UDLA)-ABG consortium, processing a large number of livestock serum samples for zoonotic diseases like brucellosis, tuberculosis, leptospirosis or Q-fever, will be extremely helpful for the ongoing challenge and threat of COVID-19 to our beloved World Heritage Site of the “enchanted” Galapagos Islands.
Authorship contribution statement
All authors contributed to study conceptualization, experimental procedures and revision and approval of final version of the manuscript.
Byron Freire-Paspuel and Miguel Angel García Bereguiain analyzed the data and wrote the manuscript.
Declaration of Competing Interest
The authors declare no conflict of interest.
Acknowledgements
We thank “Consejo de Gobierno de las Islas Galápagos”, “Ministerio de Salud Pública”, “Ministerio del Ambiete”, World Health Organization and authorities from “Universidad de Las Américas” for their support to make SARS-CoV-2 diagnosis possible at “LabGal” at Galápagos Islands. We also thank “Gobierno Autónomo Descentralizado de Santa Cruz”, “Gobierno Autónomo Descentralizado de San Cristobal” and“Gobierno Autónomo Descentralizado de Isabela” for their logistic support during the COVID19 surveillance.
References
- 1.Zhou P., Yang X.L., Wang X.G., Hu B., Zhang L., Zhang W., Si H.R., Zhu Y., Li B., Huang C.L., Chen H.D., Chen J., Luo Y., Guo H., Jiang R.D., Liu M.Q., Chen Y., Shen X.R., Wang X., Zheng X.S., Zhao K., Chen Q.J., Deng F., Liu L.L., Yan B., Zhan F.X., Wang Y.Y., Xiao G.F., Shi Z.L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020 doi: 10.1038/s41586-020-2012-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Gorbalenya A.E., Baker S.C., Baric R.S., de Groot R.J., Drosten C., Gulyaeva A.A., Haagmans B.L., Lauber C., Leontovich A.M., Neuman B.W., Penzar D., Perlman S., Poon L.L.M., Samborskiy D.V., Sidorov I.A., Sola I., Ziebuhr J. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020;4:536–544. doi: 10.1038/s41564-020-0695-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lu Xiaoyan, Wang Lijuan, Sakthivel Senthilkumar K., Whitaker Brett, Murray Janna, Kamili Shifaq, Lynch Brian, Malapati Lakshmi, Burke Stephen A., Harcourt Jennifer, Azaibi Tamin Natalie J., Thornburg Julie M. Villanueva and Stephen Lindstrom. US CDC real-time reverse transcription PCR panel for detection of severe acute respiratory syndrome Coronavirus 2. Emerg. Infect. Dis. 2020;26:8. doi: 10.3201/eid2608.201246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Interim Guidelines for Collecting, Handling, and Testing Clinical Specimens from Persons for Coronavirus Disease 2019 (COVID-19). Center for Diseases Control and Prevention, USA. https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html (last access 04/20/20).
- 5.Daniel D. Rhoads, Sree S. Cherian, Katharine Roman, Lisa M. Stempak, Christine L. Schmotzer, Navid Sadri. Comparison of Abbott ID Now, Diasorin Simplexa, and CDC FDA EUA methods for the detection of SARS-CoV-2 from nasopharyngeal and nasal swabs from individuals diagnosed with COVID-19. Accepted Manuscript Posted Online 17 April 2020. J. Clin. Microbiol. doi: 10.1128/JCM.00760-20. [DOI] [PMC free article] [PubMed]
- 6.Arun K. Nallaa, Amanda M. Castob, Meei-Li W. Huanga, Garrett A. Perchettia, Reigran Sampoleoa, Lasata Shresthaa, Yulun Weia, Haiying Zhua, Keith R. Jeromea, Alexander L. Greningera. Comparative Performance of SARS-CoV-2 Detection Assays using Seven Different Primer/Probe Sets and One Assay Kit. JCM Accepted Manuscript Posted Online 8 April 2020. J. Clin. Microbiol. doi: 10.1128/JCM.00557-20. [DOI] [PMC free article] [PubMed]
- 7.Freire-Paspuel Byron, Vega-Mariño Patricio, Velez Alberto, Castillo Paulina, Cruz Marilyn, Garcia-Bereguiain Miguel Angel. Evaluation of nCoV-QS (MiCo BioMed) for RT-qPCR Detection of SARS-CoV-2 From Nasopharyngeal Samples Using CDC FDA EUA qPCR Kit as a Gold Standard: An Example of the Need of Validation Studies. J Clin Virol. 2020 May 22;128:104454. doi: 10.1016/j.jcv.2020.104454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.https://www.elcomercio.com/actualidad/vicepresidente-actividades-turisticas-galapagos-julio.html
- 9.Freire-Paspuel Byron, Vega-Mariño Patricio, Velez Alberto, Castillo Paulina, Cruz Marilyn. Miguel Angel Garcia-Bereguiain. Sample pooling of RNA extracts to speed up SARS-CoV-2 diagnosis using CDC FDA EUA RT-qPCR kit. Virus Research. 2020;2020:198173. doi: 10.1016/j.virusres.2020.198173. Volume 290, December. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Freire-Paspuel B., Vega-Mariño P., Velez A., Castillo P., Gomez-Santos E.E. Cruz M and Garcia-Bereguiain MA (2020) cotton-tipped plastic swabs for SARS-CoV-2 RT-qPCR diagnosis to prevent supply shortages. Front. Cell. Infect. Microbiol. 2020;10:356. doi: 10.3389/fcimb.2020.00356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Center for Diseases Control and Prevention (2020). Interim Guidelines for Collecting, Handling, and Testing Clinical Specimens from Persons for Coronavirus Disease 2019 (COVID-19). Center for Diseases Control and Prevention. Available online at: https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html (accessed April 20, 2020).
- 12.WHO . World Health Organization; 2020. Laboratory Testing for Coronavirus Disease (COVID-19) in Suspected Human Cases. Interim Guidance. [Google Scholar]
- 13.https://www.oie.int/en/animal-health-in-the-world/official-disease-status/classical-swine-fever/list-of-csf-free-member-countries/
- 14.Leal Filho W., Lütz J.M., Sattler D.N., Coronavirus Nunn P.D. COVID-19 transmission in Pacific Small Island developing states. Int. J. Environ. Res. Public Health. 2020;17:5409. doi: 10.3390/ijerph17155409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gudbjartsson D.F. Humoral Immune Response to SARS-CoV-2 in Iceland. New England Journal of Medicine. 2020 doi: 10.1056/NEJMoa2026116. [DOI] [PMC free article] [PubMed] [Google Scholar]