Abstract
The human genome contains thousands of natural antisense transcripts (NAT) that can regulate epigenetic state, transcription, RNA stability, or translation of their overlapping genes1,2. We describe MAPT-AS1, a primate-conserved, brain-enriched NAT containing an embedded mammalian-wide interspersed repeat (MIR), which represses tau translation by competing with rRNA pairing to MAPT mRNA internal ribosome entry site (IRES)3. Tau, a neuronal intrinsically disordered protein (IDP), stabilises axonal microtubules. Hyperphosphorylated, aggregation-prone tau forms the hallmark inclusions of tauopathies4. MAPT mutations cause familial frontotemporal dementia (FTLD-tau), and common variation forming the MAPT H1 haplotype is a significant risk factor in many tauopathies5, and Parkinson’s disease. Notably, expression of MAPT-AS1 or its minimal essential sequences including MIR reduces, whereas silenced MAPT-AS1 increases neuronal tau, and is correlated with tau pathology in human brain. Moreover, we identified hundreds additional NATs with embedded MIRs (MIR-NATs), which are overrepresented at coding genes linked to neurodegeneration, and/or encoding IDPs, and confirmed MIR-NAT-mediated translational control of one such gene, PLCG1. Collectively, we present the importance of MAPT-AS1 for tauopathies, while also uncovering a potentially broad contribution of MIR-NATs to the tightly controlled translation of IDPs6, with particular relevance for proteostasis in neurodegeneration.
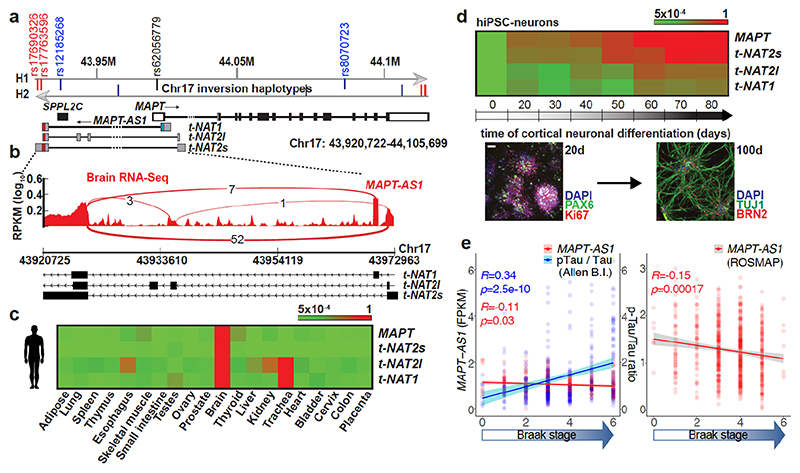
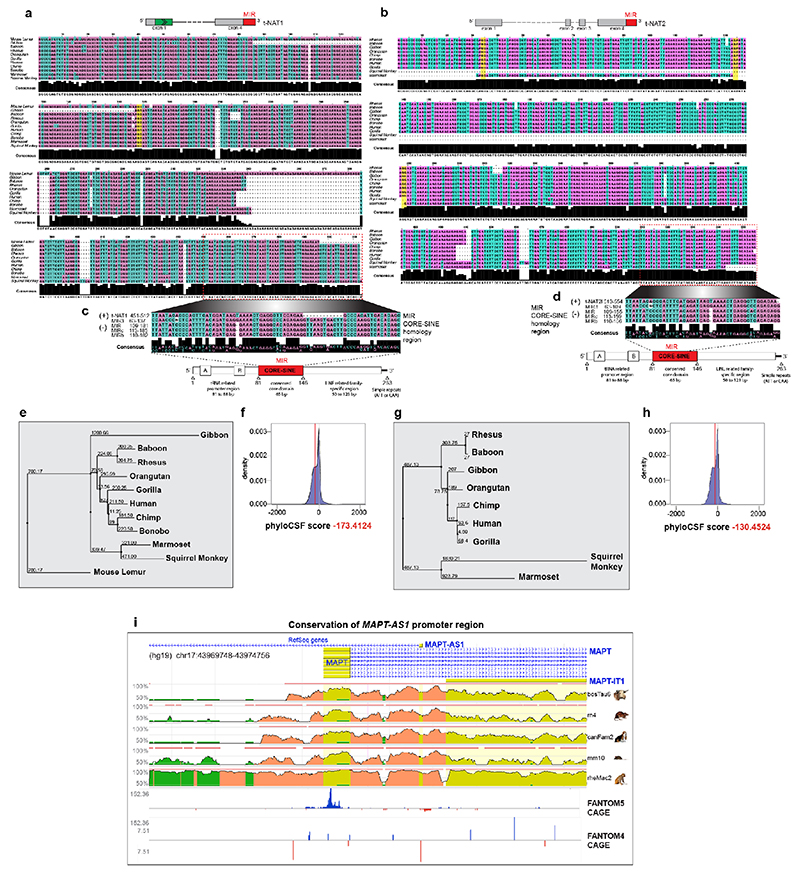
MAPT-AS1, overlapping head-to-head with MAPT 5’UTR, extends ~52 kilobases upstream from MAPT (Fig. 1a) into the linkage disequilibrium (LD)-region defining the H1/H2 haplotypes5 (Extended Data Fig. 1e). We identified three MAPT-AS1 isoforms (t-NAT1, t-NAT2s, t-NAT2l) (Fig. 1a), as bona-fide lncRNAs with negative PhyloCSF scores (Extended Data Fig. 2f,h) and no open reading frames (ORFs). Seventy-five nt of t-NAT1 exon1 overlaps MAPT 5’UTR, and exon1 of t-NAT2s and t-NAT2l overlaps an evolutionarily conserved region (Extended Data Fig. 2i). Alternative transcription start sites are supported by brain RNA-seq data (Fig. 1b and Extended Data Fig. 3a) and CAGE-clusters7 (Extended Data Fig. 2i). t-NAT2l includes two additional alternative exons. The first t-NAT1 splice junction is conserved in all primates (Extended Data Fig. 2a,e), whereas t-NAT2l alternative exons are conserved up to Old World primates (Extended Data Fig. 2b,g).
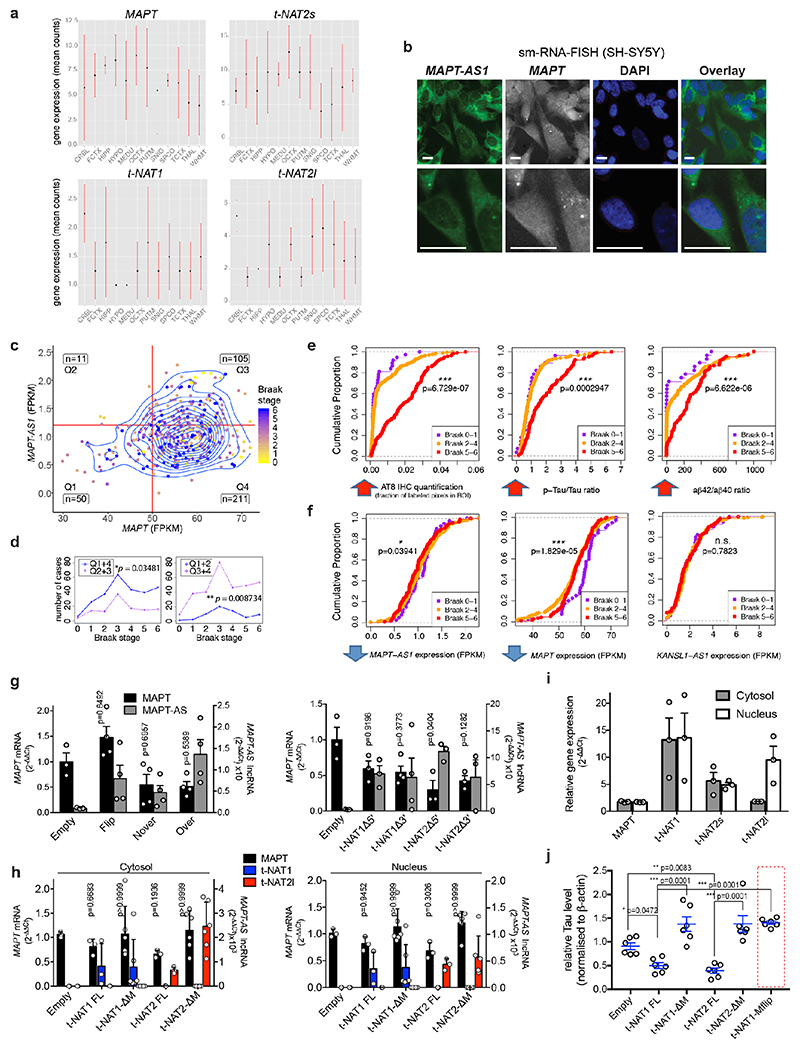
Fig. 1. MAPT-AS1 is brain-enriched, expressed during neuronal differentiation and inversely correlated to tau pathology.
a, MAPT-AS1 and MAPT genes (hg19). Grey arrows indicate inverted H1/H2 haplotypes, with haplotype-tagging SNPs (blue); PD-linked rs12185268; PSP and PD-associated rs8070723 SNPs in MAPT 5’UTR (black) and MAPT-AS1 (red). MAPT coding-exons are in black; UTRs in white; MAPT-AS1 exons in grey; MIR in red, AS exonic-overlap in blue. b, Sashimi-plot of brain RNA-seq (log10RPKM) with splice-junctions counts. c, MAPT and MAPT-AS1 relative expression by qRT-PCR (2–ΔΔCt/2–ΔΔCt max) in human tissues and (d) during iPSC differentiation into cortical neurons (0-80 days), scale bar =40 μm, n=3 independent experiments e, Linear regression: mean MAPT-AS1 expression from brain RNA-seq (red line) inversely correlates with mean tau pathology (blue line; phospho-tau(AT8):total-tau, Luminex-immunoassay) and Braak-stage in Allen (left) and ROS-MAP (right) cohorts, error bars:95%CI, R:Pearson’s correlation coefficient, (two-sided p-value, t-distribution with n-2 def).
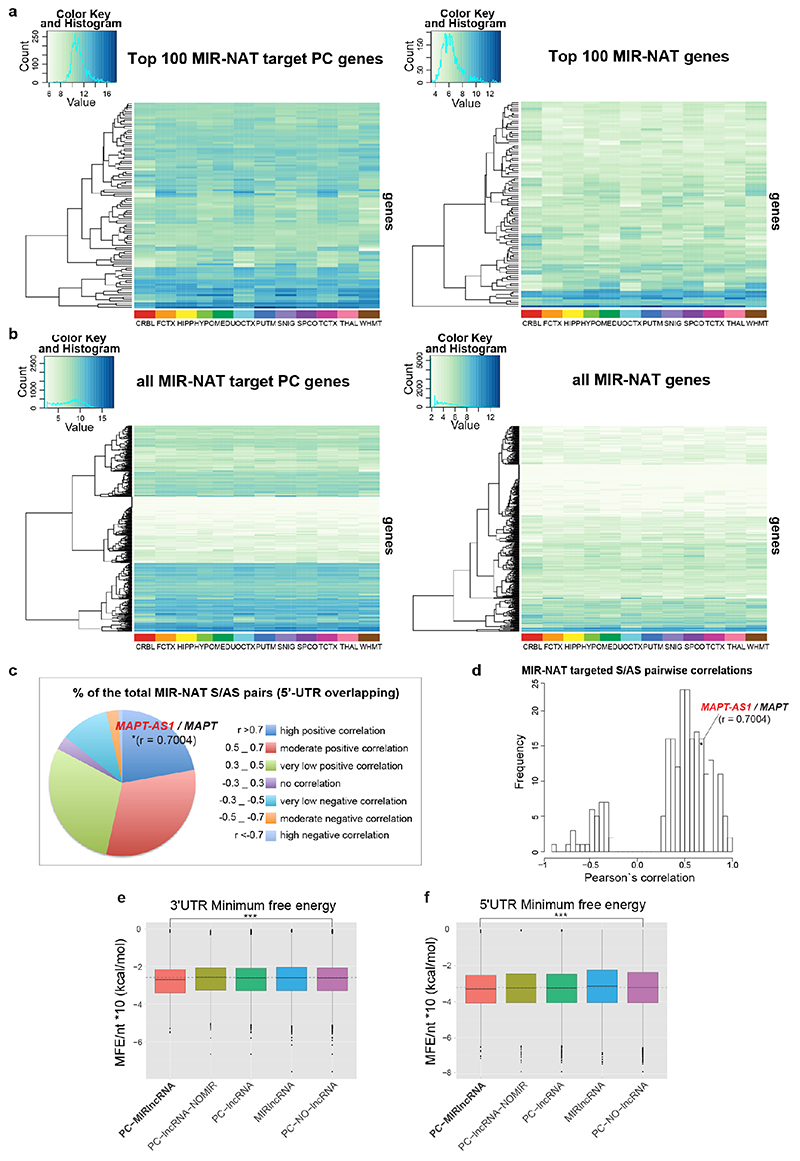
Tissue distribution of MAPT-AS1 expression is similar to MAPT, with highest levels in brain (Fig. 1c and Extended Data Fig. 3a). Human brain RNA-seq data8 showed positive correlation between MAPT-AS1 and MAPT (Pearson’s correlation coefficient 0.7004; Extended Data Fig. 3a, 6d), and their transcription concomitantly increases during cortical neuronal-differentiation of human induced pluripotent stem cells (hiPSC) (Fig. 1d Extended Data Fig. 4a,b). Single-molecule fluorescence RNA in situ hybridization with tiling-probes covering all transcripts (Supplementary Table1), showed mature MAPT-AS1 and MAPT RNAs both in nucleus and cytoplasm, with nuclear spots likely corresponding to transcription sites (Extended Data Fig. 3b). Localisation was confirmed by qRT-PCR of subcellular fractions (Extended Data Fig. 3h,i).
MAPT-AS1 and tau pathology progression
To assess MAPT-AS1 dysregulation in disease, we analysed recent multi-omics data9,10. Linear regression analysis of RNA-seq data from the Allen Brain Institute (http://aging.brain-map.org/)9 and the ROS-MAP10 cohorts (https://dx.doi.org/10.7303/syn3388564), showed that brain tau pathology (Braak-stage) inversely correlates with MAPT-AS1 levels (Fig. 1e), where high Braak-stages significantly associate with higher MAPT and lower MAPT-AS1 expression (Extended Data Fig. 3c,d). Similarly, the cumulative distribution of MAPT-AS1 and MAPT expression is significantly shifted towards smaller values at higher Braak-stages whereas the neighbouring KANSL1-AS1 is unchanged (Extended Data Fig. 3e,f). These data support a role of MAPT-AS1 in tau pathology progression. Interestingly, MAPT-AS1 expression is reduced in PD brains and substantia nigra11,12.
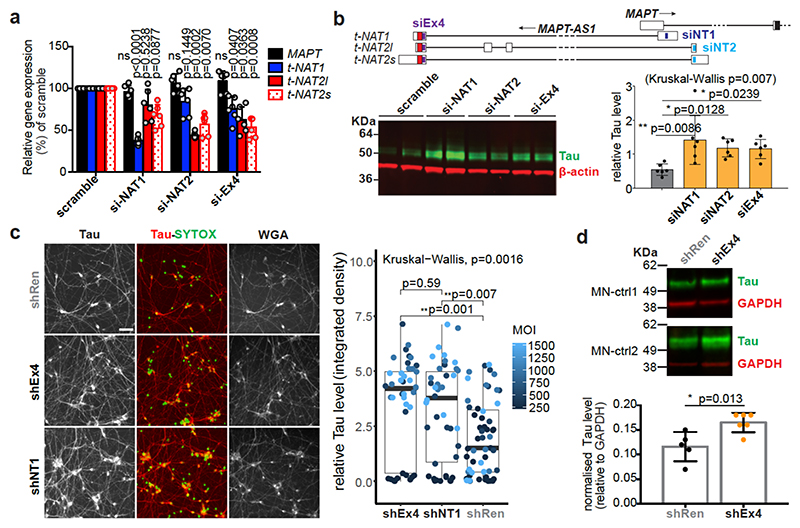
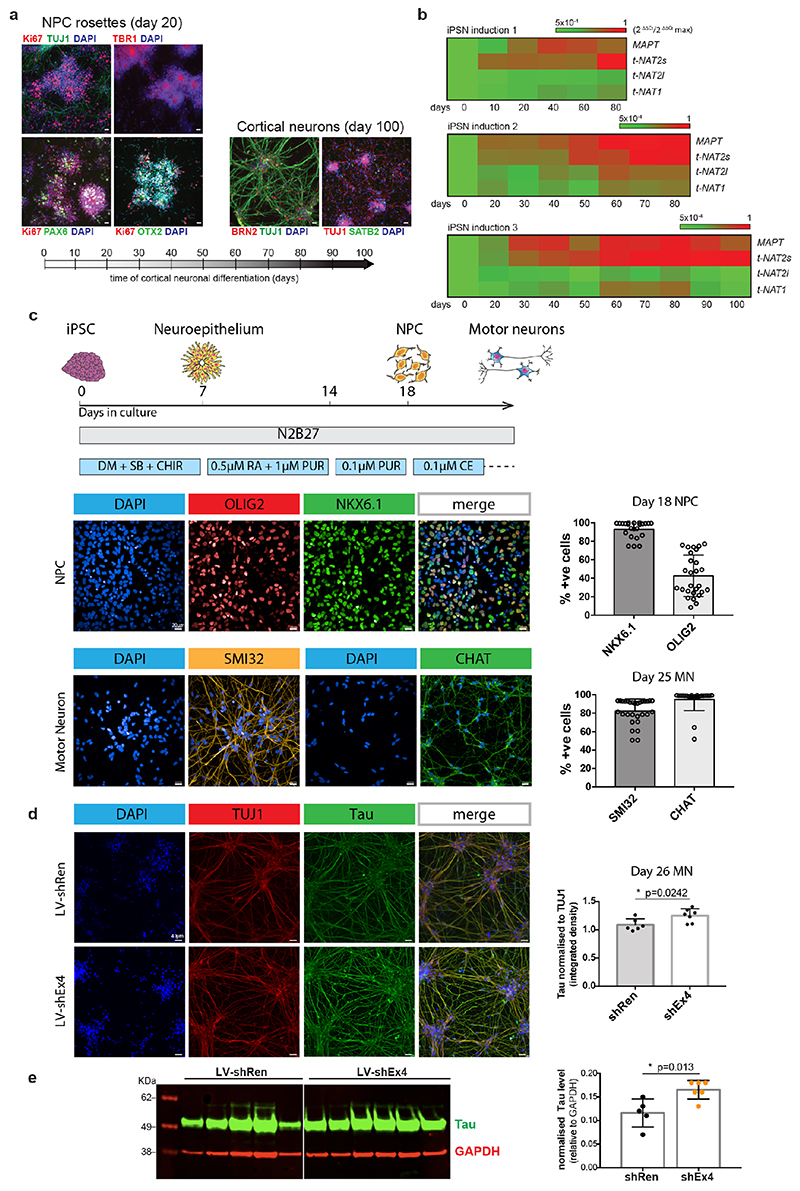
We silenced MAPT-AS1 in two neuronal models to test effects on MAPT expression. In SH-SY5Y cells, silencing with siRNAs targeting either t-NAT1 (siNAT1) or t-NAT2 (siNAT2) exon1, or the shared 3’-exon (siEx4), all caused significant tau increase (Fig. 2b) without affecting MAPT mRNA (Fig. 2a). Differentiated hiPSC-derived motor neurons (MN, Extended Data Fig. 4c) transduced with lentivirus (LV) expressing shRNAs targeting t-NAT1 exon1 (shNT1) or the 3’-exon (shEx4), showed significant dose-dependent increase of tau immunoreactivity (Fig. 2c), compared to negative control (shRen), 5-7 days-post-infection (Fig. 2c). To rule out lentiviral toxicity, we transduced MNs at much lower MOI (10) and observed a small but significant increase of tau immunoreactivity normalised to TUJ1 (Extended Data Fig. 4d). Significant tau increase by LV-shEx4 was confirmed by immunoblotting (Fig. 2d, Extended Data Fig. 4e), suggesting that despite its low expression, endogenous MAPT-AS1 tightly controls neuronal tau levels in a sub-stoichiometric manner. Overexpression of t-NAT1 or t-NAT2l (hereafter named t-NAT2) in SH-SY5Y cells, consistently reduced endogenous tau without affecting MAPT mRNA (Fig. 3a) or β-actin, TDP-43 or the neighbouring SPPL2C gene (Fig. 3b,c). These data indicate that MAPT-AS1 controls MAPT expression post-transcriptionally.
Fig. 2. Loss of MAPT-AS1 increases neuronal tau.
a-b, Silencing MAPT-AS1 in SH-SY5Y cells with siRNAs (si-NAT1, si-NAT2, siEx4) unaffected MAPT expression by qRT-PCR but increased endogenous tau compared to scramble mean (n=6 independent treatments, mean±s.d., two-sided Kruskal-Wallis with Dunn’s test). c, (left) Representative immunostainings of MNs transduced at four multiplicities of infection (MOI) with negative control LV-shRen or MAPT-AS1-specific shNT1, shEx4. Nuclei labelled by SYTOX (green), total-Tau (red) normalised to wheat germ agglutinin (WGA), scale bar=40 μm. (right) ICC quantification (n=10±1 wells across 3 experiments, n=23 wells for shRen-250MOI, box-plots: midpoints, medians; boxes, 25th and 75th percentiles; whiskers, minima and maxima; two-sided Kruskal-Wallis with Dunn’s test). d, Immunoblots of MNs from two healthy donors (MN-ctrl1, MN-ctrl2) transduced with LV-shEx4 or LV-shRen, total-tau normalised to GAPDH (n=5 shRen, n=6 shEx4, independent transductions, mean±s.d. two-sided unpaired Wilcoxon-test).
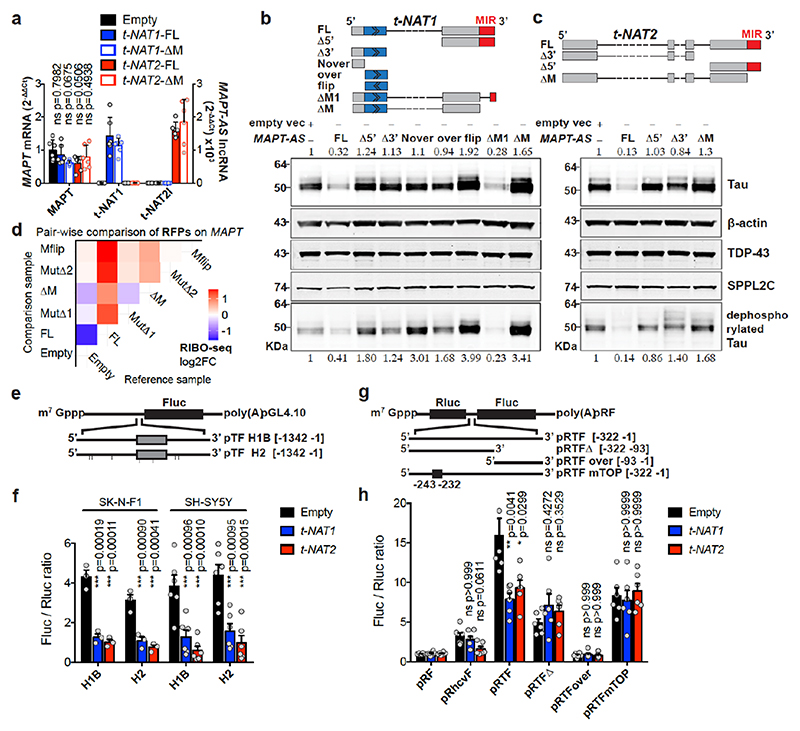
Fig. 3. MAPT-AS1 controls tau translation through embedded inverted MIR.
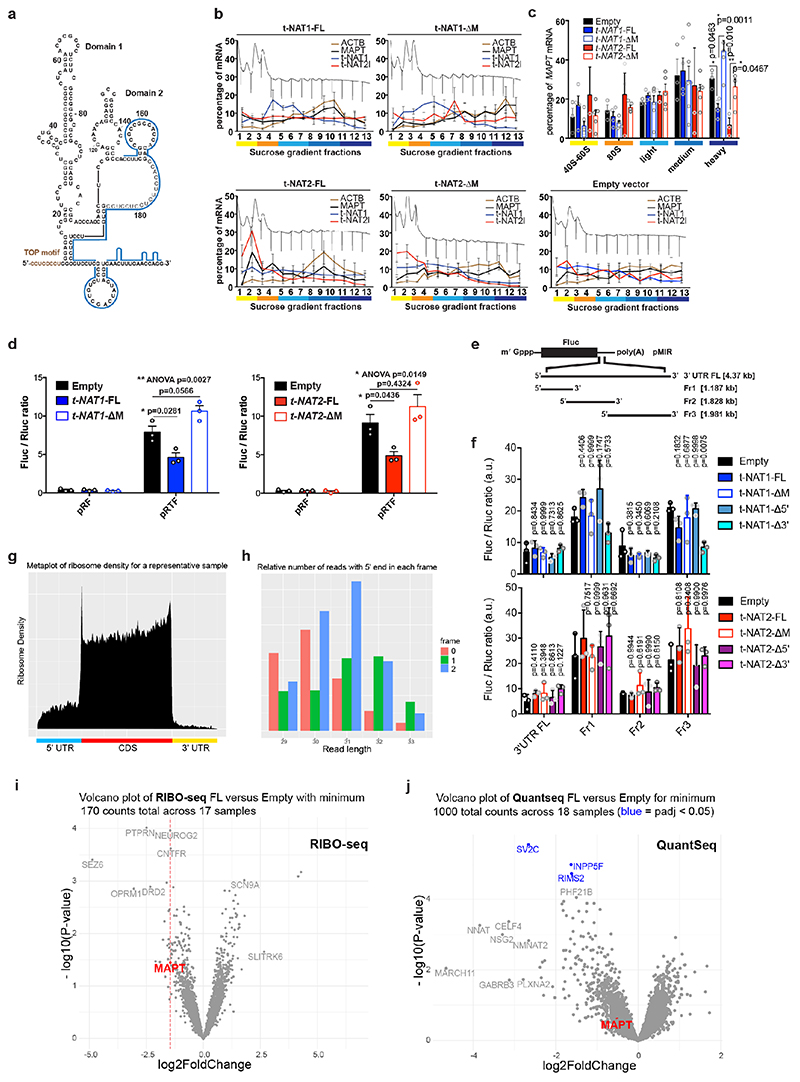
Stable expression in SH-SY5Y cells a, MAPT-AS1 and MAPT expression by qRT-PCR (2–ΔΔCt); Empty vector (Empty), full-length or mutant t-NAT1 (t-NAT1FL; t-NAT1ΔM), or t-NAT2l (t-NAT2FL; t-NAT2ΔM), MIR deletion (ΔM) (mean±s.e.m., n=6, 3 clones in 2 experiments, one-way ANOVA with Dunnett’s test), t-NAT1 (b) and t-NAT2 (c) with: full-length (FL), 5’-deletion (Δ5’); 3’-deletion (Δ3’); regions not-overlapping (Nover) or overlapping (over) with MAPT 5’UTR; flipped overlapping region (flip); partial (ΔM1) or full MIR deletion (ΔM). AS-region overlapping MAPT 5’UTR in blue; chevrons indicate orientation. t-NAT1-FL (b), t-NAT2-FL (c) reduce endogenous total- and dephosphorylated-tau (λ-phosphatase), suggesting regulation is independent of tau phosphorylation. Inverted MIR (red) is essential for controlling tau levels. Numbers above total-tau and below dephosphorylated-tau indicate levels normalised to β-actin, TDP-43 and SPPL2C geometric mean. d, Pairwise comparison heatmap of RIBO-seq ribosome footprints (RFPs) along MAPT from 3 independent SH-SY5Y clones expressing Empty-vector, t-NAT1 (FL), deletion of MIR motif-1 (MutΔ1) or motif-2 (MutΔ2) as in Fig.4a, MIR deletion (ΔM), MIR flipped (Mflip). FL significantly decreases MAPT RFPs compared to Empty (log2FC=-1.45, p=0.036, Wald test with Bonferroni correction). e, pTF reporters: a 1,342 nt genomic fragment spanning MAPT promoter, 5’UTR (grey box) and intron segment, upstream to firefly luciferase (Fluc) ORF. Haplotypes H1B and H2, (7 SNPs), were tested. f, FL t-NAT1 and t-NAT2 transient expression significantly repress Fluc translation normalised to Renilla luciferase (mean±s.e.m., one-way ANOVA with Dunnett’s test, n=3 SK-N-F1, n=6 SH-SY5Y independent experiments). g, Bicistronic reporters: MAPT 5’ UTR inserted between Renilla (Rluc) and Fluc ORFs in pRF vector62, resulted in pRTF. Truncations (pRTFΔ and pRTFover) or 5’TOP motif mutation (pRTFmTOP) reduced tau-IRES activity. Hepatitis C virus IRES (pRhcvF), positive control. h, SH-SY5Y cells stably expressing empty vector (Empty), t-NAT1 or t-NAT2, were transfected with constructs in (g) and capindependent translation (Fluc/Rluc ratio) measured. Control cells (Empty) transfected with pRTF showed a ~15- fold increase in Fluc/Rluc ratio over negative control pRF vector, and a ~3.7-fold increase over pRhcvF; FL t-NAT1 or t-NAT2 expression significantly reduced tau-IRES activity. (n=3 SH-SY5Y clones in 2 independent experiments, mean±s.e.m., two-sided Kruskal-Wallis with Dunn’s test).
To identify essential MAPT-AS1 sequences, with stably expressed full-length (FL) MAPT-AS1 or targeted deletions in SH-SY5Y cells, FL consistently inhibited tau protein production compared to control cells with empty-vector (Empty) (Fig. 3b,c). Deletion of 5’-exons (Δ5’) or the shared 3’-exon (Δ3’), completely abolished repression (Fig. 3b,c), showing both are functionally essential domains.
All MAPT-AS1 isoforms share a 3’-exon with an embedded MIR repeat, subclass MIRc, in inverse orientation (www.repeatmasker.org). MIR elements are ~260-nt non-autonomous tRNA-derived retrotransposons13 with a conserved central CORE-SINE14, found in all mammals and constitute about 2.54% of the human genome15. Notably, the 62-nt CORE-SINE within MAPT-AS1 is conserved in all primates (Extended Data Fig. 2c,d). Stable expression of t-NAT1 with partial MIR deletion retaining the CORE-SINE maintained capacity for translational repression, whereas this was lost with t-NAT1 or t-NAT2 lacking the MIR (ΔM) (Fig. 3b,c), or with MIR flipped (Mflip) (Extended Data Fig. 3j). Notably, flipped MIR increased tau levels, which might be due to its complementarity to rRNA. Moreover, stable expression of t-NAT1 5’-exon alone in either orientation had no effect on tau expression (Fig. 3b). This demonstrates that MAPT-AS1 acts in a modular fashion that requires the 5’AS-region overlapping MAPT conferring target specificity, and the 3’MIR CORE-SINE mediating translational repression.
cap- and IRES-mediated translation repression
We used polysome profiling of stably transfected cells, which showed that expression of FL t-NAT1 or t-NAT2 significantly shifted MAPT mRNA from heavy to lighter polysomes, where MAPT-AS1 transcripts are present, indicating its direct role in translational repression (Extended Data Fig. 5b-c). Conversely, ΔM transcripts did not affect MAPT mRNA polysome engagement (Extended Data Fig. 5c). To assess specificity of MAPT-AS1-mediated regulation we used RIBO-seq measuring genome-wide distribution of ribosome footprints (RFPs) (Extended Data Fig. 5g,h) comparing SH-SY5Y cells stably expressing MAPT-AS1 (FL, ΔM, MutΔl, MutΔ2, Mflip) to those with an empty vector (Empty). Despite its relatively low expression and thus few RFPs mapping to MAPT, we detected a significant (log2FC = -1.45, p = 0.036) decrease in RFPs on MAPT, ranked 22nd out of 4546 genes, for FL versus Empty (Fig. 3d, Extended Data Fig. 5i). Although a small number of other genes had similar fold-changes and p-values, none had a significant adjusted p-value, suggesting they are likely false positives (Extended Data Fig. 5i), with no shared sequence motifs. As expected, MAPT RFPs did not significantly change for cells stably expressing non-functional mutants of MAPT-AS1 (Fig. 3d, ΔM, MutΔl, MutΔ2, Mflip). Although we detected a small non-significant decrease in MAPT mRNA expression in FL versus Empty (log2FC = -0.52, p = 0.25), this does not account for the larger significant decrease in RFPs (Extended Data Fig. 5j). Only 3 genes were significantly downregulated and likely transcriptional off-targets when comparing FL versus Empty, (Extended Data Fig. 5j); none paired to MIR-NATs. To independently validate MAPT-AS1 translational effects, we cotransfected SH-SY5Y cells with pTF, a monocistronic luciferase reporter containing haplotype variants of a genomic fragment (1,342 bp) spanning MAPT core-promoter, 5’UTR, and part of downstream intron (Fig. 3e) and FL t-NAT1 or t-NAT2 expression plasmids. We detected significant reductions in relative luciferase activity with both MAPT-AS1 isoforms (Fig. 3f), confirming their role in controlling cap-dependent tau translation.
Tau translation, spatiotemporally controlled by the mTOR-p70S6K pathway via a 5’-terminal oligopyrimidine (TOP) sequence, promotes axonal tau accumulation16 and establishment of neuronal polarity. It can occur through both cap-dependent16 and IRES-mediated mechanisms3. MAPT 5’UTR folds into two domains forming an IRES, but factors controlling its efficiency remain unknown. We found that MAPT-AS1 overlaps by 75-nt with domain-2 of tau-IRES that binds to 40S ribosomes3 (Extended Data Fig. 5a). To examine effects of MAPT-AS1 on the tau-IRES, we generated bicistronic vectors expressing Renilla (Rluc) and firefly (Fluc) luciferase, translation of which is cap- or tau-IRES-dependent, respectively (Fig. 3g). While full-length tau-IRES produced high levels of firefly luciferase, tau-IRES deletions (pRTFover, pRTFΔ) or mutation (mTOP) caused significant reduction of expression, confirming that tau-IRES domains 1 and 2 are both required for maximal translation3 (Fig. 3h). Furthermore, normalised Fluc activity significantly decreased in cells expressing either t-NAT1 or t-NAT2 compared to negative control cells (pRF), but, neither mutant nor truncated tau-IRES was affected by either t-NAT (Fig. 3h). Furthermore, expression of t-NAT1 or t-NAT2 with deleted MIR failed to repress tau-IRES (Extended Data Fig. 5d). Collectively, these results corroborate MAPT-AS1 role in regulating both cap-dependent and IRES-mediated tau translation. Given that translation of mRNAs with long 5’UTRs, including MAPT, heavily depends on EIF4A helicase activity17, it is possible that MAPT-AS1 could increase tau helicase dependency when initiation factors become limiting.
Two essential MIR motifs
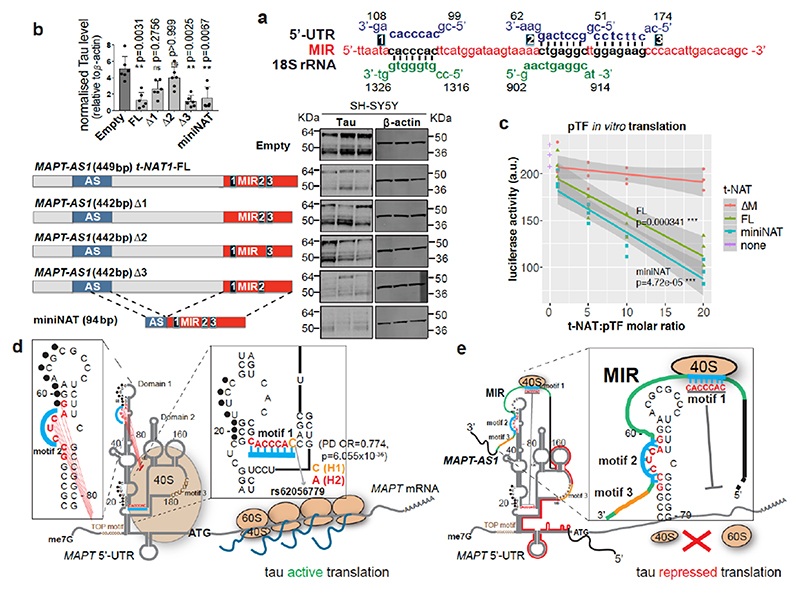
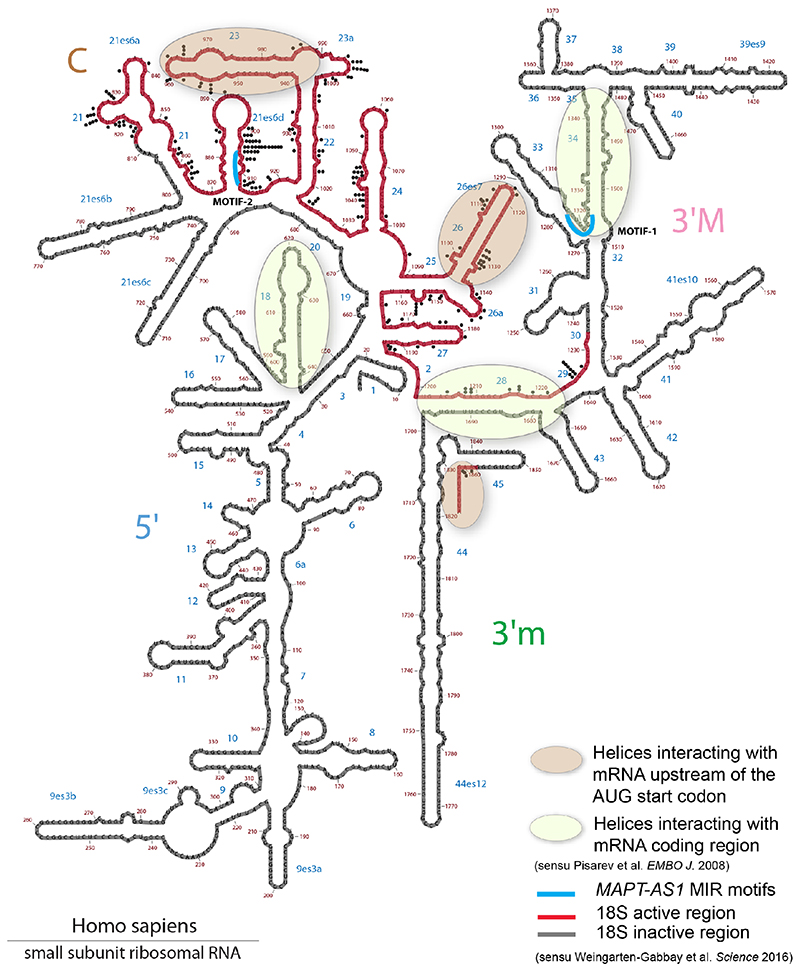
Cells co-transfected with full-length or truncated MAPT 3’UTR downstream to Fluc-ORF and wild-type or mutant MAPT-AS1 constructs showed no significant differences in luciferase activity (Extended Data Fig. 5e,f), suggesting that MAPT-AS1 function does not require MAPT 3’UTR. It is thus clear that MAPT 5’UTR mediates the effect, which could involve either cap-dependent16 and cap-independent3 translation. Therefore, we sought to identify essential motifs of MAPT-AS1 that could directly interfere with MAPT 5’UTR ribosome recruitment. A BLAST-search with 7-nt window, for similarities between MAPT-AS1, the 18S rRNA and MAPT5’UTR, uncovered two 7-mer motifs within the MIR: motif-1 (CACCCAC) complementary to position 1318-1324 of 18S rRNA helix 34, within the mRNA entry channel (Fig. 4a, Extended Data Fig. 6); motif-2 (CTGAGGC) identical to position 905-911 of 18S rRNA expansion segment 6, only present in eukaryotes (Fig. 4a, Extended Data Fig. 6). Strikingly, motif-1 and -2 are identical and complementary, respectively, to tau-IRES sequences interacting with 18S rRNA, suggesting the MIR competes with the first IRES motif for rRNA binding, and directly blocks the second IRES motif, impairing ribosomal recruitment. Another MIR motif-3 is only complementary to MAPT 5’UTR (Fig. 4a,d).
Fig. 4. Two essential MIR motifs for MAPT-AS1-mediated tau repression.
a, motif-1 and 2, (black) are identical or complementary to MAPT 5’UTR (blue) and 18S rRNA (green). Motif-3 is complementary to 5’UTR. b, FL-t-NAT1 stable expression significantly reduces total-tau in SH-SY5Y cells, compared to Empty. t-NAT1 motif-1 (Δ1) or -2 (Δ2) deletion unaffected tau. Deletion of motif-3 (Δ3) preserved t-NAT1-mediated repression. miniNAT composed of 32-nt AS-region (blue) complementary to MAPT 5’UTR, fused with inverted MIR (red) represses tau. (mean±s.d., n=6, 3 clones in 2 experiments; two-sided Kruskal-Wallis with Dunn’s test) c, in vitro transcribed t-NAT-FL and miniNAT repress dose-dependently in vitro translation of pTF luciferase compared to mutant ΔM (regression lines, mean with 95% CI, n=3 independent experiments; two-sided ANCOVA test; df=2, F=12.886, p=7.85x10–05ANOVA for slope; df=3, F=32.127, p=8.97x10–10ANOVA for t-NAT) d, MAPT mRNA with 5’UTR experimentally determined structure3. Tau-IRES recruits ribosomes (salmon ovals) by pairing with rRNA at two sites (motif-1, motif-2, turquoise). Complementary nucleotides 59-65 and 19-25 (black dots) form a kissing-hairpin, crucial for tau-IRES3. The PD-associated SNP rs62056779 (OR=0.774, p=6.055x10–36) is within motif-1 e MAPT-AS1 inhibits IRES- and cap-dependent tau translation through both 5’AS-region complementary to domain 2 (red line) and the inverted MIR (green line), containing motif-1 and -2 (turquoise). Motif-3 (orange) is dispensable.
With mutually exclusive complementarity of MIR motifs with 18S rRNA or the MAPT 5’UTR, we tested how MIR influences MAPT-AS1 function. With stable expression in SH-SY5Y cells, deletion of either motif-1 (Δ1) or -2 (Δ2), but not motif-3 (Δ3), significantly impaired the capacity to repress tau compared to FL MAPT-AS1 (Fig. 4b). To further support our conclusions, we stably expressed a miniNAT, containing a fusion of 32-nt AS region overlapping with MAPT 5’UTR and the inverted MIR (62-nt), which retained full capacity to inhibit tau translation (Fig. 4b). Similarly, in vitro transcribed FL and miniNAT, but not ΔM, significantly repressed in vitro translation of pTF luciferase reporter in a dose-dependent manner (Fig. 4c). This is compelling evidence that the AS-domain together with the inverted MIR are essential and sufficient for tau repression. Based on this, we propose a model (Fig. 4d,e) whereby MIR motif-1 and -2 repress both IRES- and cap-dependent translation by competing with MAPT 5’UTR for pairing with 18S rRNA. This model is in line with the ‘ribosome filter hypothesis’, which proposed that differential binding of mRNAs to 40S ribosomes might selectively affect translation rates via mRNA-rRNA complementarity that could be modulated by ribosomal heterogeneity or competitive pairing with ncRNAs18.
In vivo effects on MAPT proteostasis
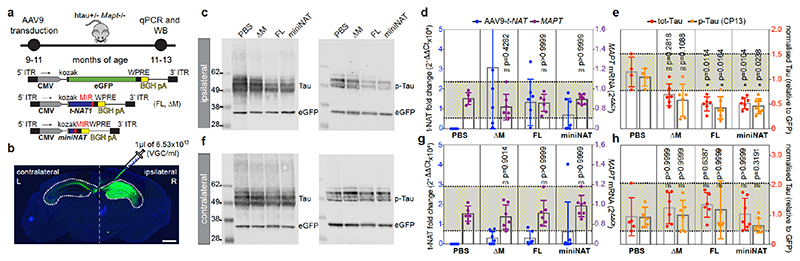
htau transgenic mice carrying 4-5 copies of the human MAPT gene including promoter and UTRs19 express all six CNS isoforms of human tau in the absence of murine tau (MAPT +/- Mapt -/-), displaying age-dependent tau pathology and late-onset behavioural impairments19. Adult (9-11 mo) mice were unilaterally injected in the hippocampus with adeno-associated virus serotype-9 (AAV9)-CMV vectors expressing full-length MAPT-AS1 (FL), a MIR deletion mutant (ΔM), miniNAT, eGFP or PBS as vehicle control (Fig. 5a). Eight weeks post-injection, AAV9-eGFP-transduced mice showed robust ipsilateral labelling and limited contralateral spread (Fig. 5b) with qRT-PCR showing similar distribution of MAPT-AS1-FL and miniNAT (Fig. 5d,g). AAV9-MAPT-AS1-FL or miniNAT transduced brains showed significantly reduced (~50%) ipsilateral levels of total- and phospho-tau compared to PBS-injected mice (Fig. 5c,e). These differences did not extend to contralateral hemisphere (Fig. 5f,h). AAV9-MAPT-AS1-ΔM injection did not significantly reduce tau despite higher transduction efficiency (Fig. 5e). Crucially, robust tau reduction in vivo caused by miniNAT confirmed functionality of only the MIR together with AS-domain.
Fig. 5. MAPT-AS1 represses tau translation in vivo in a MIR-dependent manner.
a, AAV9 expressing eGFP or MAPT-AS1 (FL, ΔM, miniNAT), for unilateral hippocampal transduction of htau+/- Mapt-/- mice (9-11 mo). b, Coronal section of AAV9-eGFP transduced htau mouse (n=4), showing robust ipsilateral (R) and limited contralateral (L) labelling; scale bar=900μm. Representative immunoblots of ipsilateral (c) and contralateral (f) brain hemispheres injected with PBS or AAV9-MAPT-AS1 (FL, ΔM, miniNAT), immunolabeled for total-tau (DAKO), pSer202-tau (CP13) and eGFP. AAV9-MAPT-AS1 and MAPT quantitative expression (relative to PBS) from transduced ipsilateral (d) and contralateral (g) hemispheres. Quantification (normalised to eGFP) of total-tau and p-tau from ipsilateral (e) and contralateral (h) hemispheres. Dashed lines delimit minima-maxima in PBS-injected mice (tau), or across all samples (MAPT); means, grey bars. (mean±s.d., n=4 PBS, n=6 ΔM, n=6 FL, n=7 miniNAT in c-d-e, n=5 PBS, n=6 ΔM, n=6 FL, n=7 miniNAT in f-g-h; two-sided Kruskal-Wallis with Dunn’s test, experiments repeated 3 times).
Our results link two previously observed mechanisms for PD pathogenesis. Firstly, MAPT-AS1 but not MAPT levels are significantly reduced in PD brains11,12. Secondly, a single-nucleotide polymorphism (SNP), rs62056779, is located within motif-1 of tau-IRES (Fig. 4d)3, and based on the PDGene database (13,708 PD cases and 95,282 healthy controls)20, significantly influences PD risk (OR=0.774, p=6.055x10-36). The risk (C) allele of H1 haplotype, among the strongest genetic risk factors for PD20, favours base-pairing with 18S rRNA, whereas the protective H2 (A) allele does not, and thus decreases tau-IRES activity3. Hence, the combined decreased MAPT-AS1 levels and presence of the H1 haplotype could jointly enable high tau-IRES activity and drive PD risk by disrupting tau proteostasis.
MIR-NATs provide RBSs
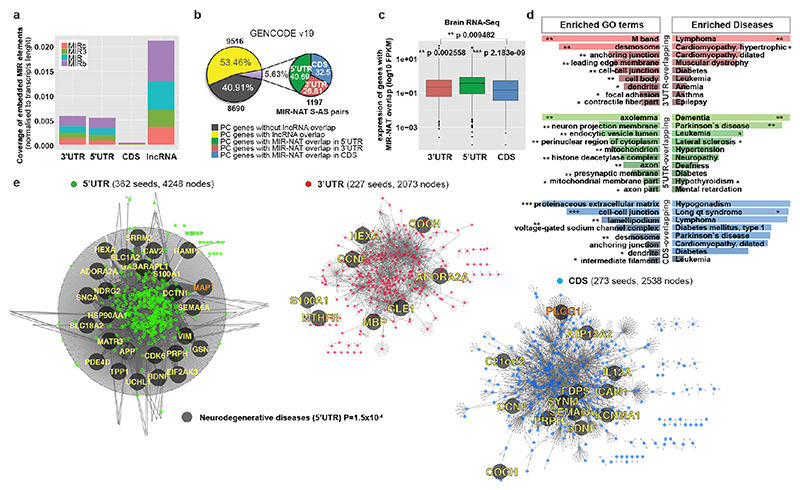
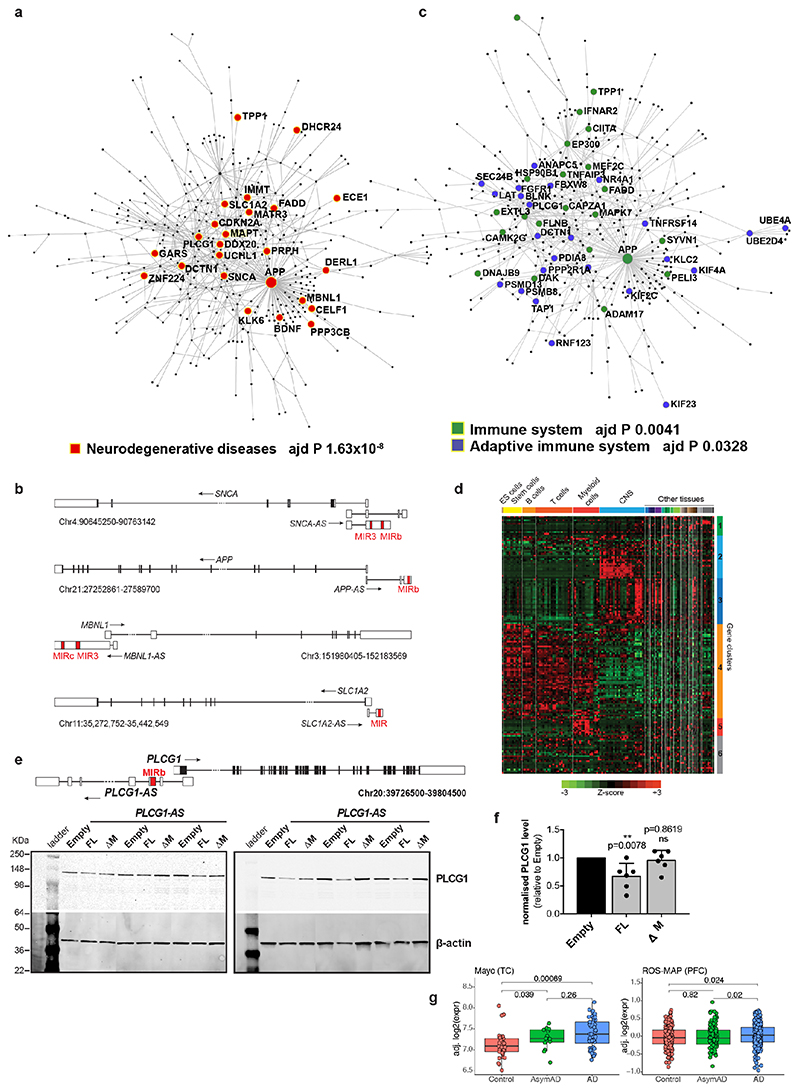
Transposable elements (TEs) are present in over two-thirds of mature lncRNAs21, thus contributing to lineage-specific diversification of vertebrate lncRNA repertoires22. However, there is scant data on the functionality of specific TEs in lncRNAs. We therefore evaluated the genomewide prevalence of MIR-NATs from GENCODE annotations (Supplementary Table2a-e). Considering the CORE-SINE conservation in all subclasses (MIR, MIR3, MIRb, MIRc)14, all MIRs were included in both orientations. MIR coverage within each transcript was normalized to their lengths. In line with a general enrichment of TEs in lncRNAs22, all MIR subclasses are enriched in lncRNAs compared to protein-coding mRNAs (Extended Data Fig. 8a). Next, we systematically examined features of protein-coding genes paired with MIR-NATs. In GENCODE v19, 5.63% of NAT-lncRNAs are MIR-NATs (n=1,197), 40.69% overlap with 5’UTR, 32.50% with 3’UTRs and 26.81% span coding sequences (CDS) (Extended Data Fig. 8b). Interestingly, genes with different extents of MIR-NAT overlap encode proteins enriched in different cellular components and diseases (Supplementary Table2b). Coding genes with 5’UTR-overlapping MIR-NATs are significantly more expressed in brain and associated with dementia, PD or amyotrophic lateral sclerosis and localise mainly to neuronal projections (Extended Data Fig. 8c,d,e, Supplementary Table2b).
Notably, genes targeted by MIR-NATs have significantly more structured 5’UTRs (Extended Data Fig. 7e,f), suggesting they could be sensitive to EIF4A helicase inhibition, and more prone to IRES-mediated translation, which is common in neuronal mRNAs23. Moreover, as with MAPT-AS1, most embedded MIRs are enriched for short motifs complementary to “active region” sequences of 18S rRNA (Supplementary Table4, Extended Data Fig. 6) thus providing potential ribosome binding sites (RBS)24. These RBSs could similarly compete with cognate mRNAs for rRNA access, inhibiting their translation initiation by impairing cellular IRESs, affecting ribosome scanning of long structured 5’UTRs and/or impeding start codon definition by RNA looping25.
To determine if additional MIR-NATs might repress translation of paired genes, we selected PLCG1-AS based on similar topology to MAPT-AS1, including an inverted MIRb with a 9-mer motif (positions 104-112), complementary to the 5’UTR of the phospholipase-C gamma 1 (PLCG1, positions 158-166, Extended Data Fig. 9e) as well as 18S rRNA (positions 305-313). Furthermore, PLCG1 5’UTR (positions 139-174) is complementary to another 18S rRNA site (positions 722-763), suggesting the inverted MIRb in PLCG1-AS could similarly compete with PLCG1 5’UTR for recruiting ribosomes. Stable PLCG1-AS expression in SH-SY5Y cells caused robust reduction of PLCG1 protein, whereas deletion of the inverted MIRb (ΔM) abolished this repression (Extended Data Fig. 9e,f). Notably PLCG1 is dysregulated in AD (Extended Data Fig. 9g).
MIR-NATs overlap with NDD and IDP genes
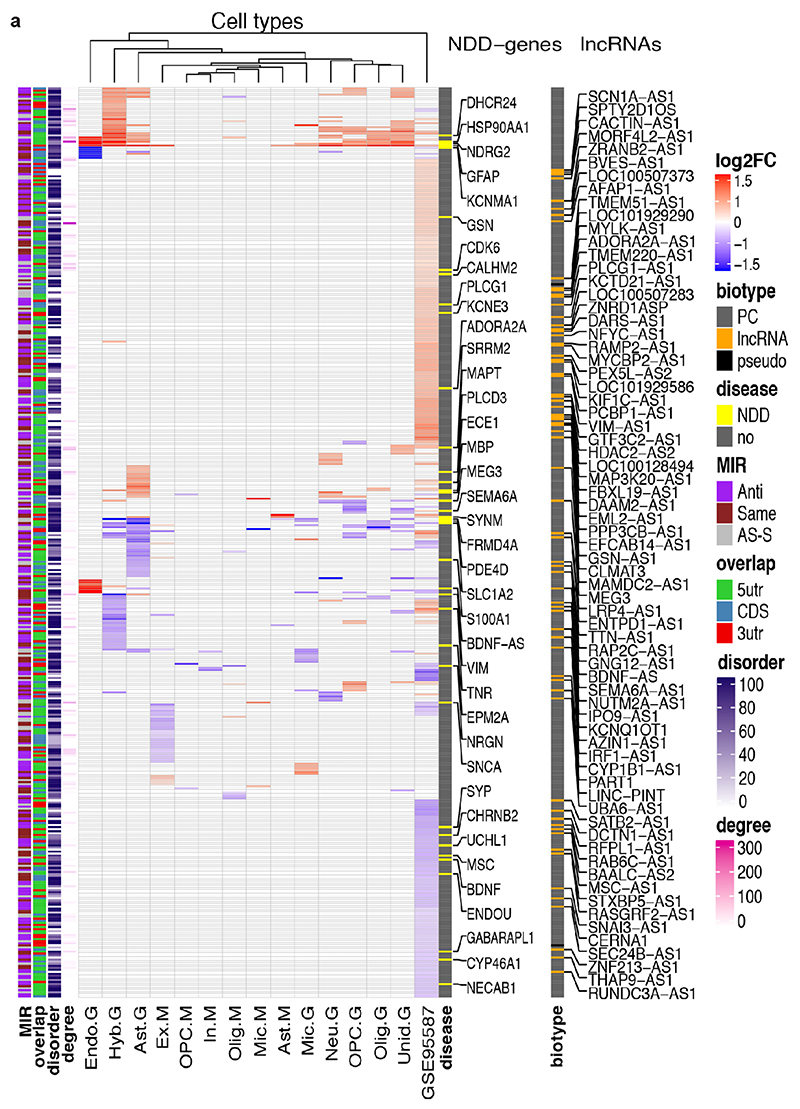
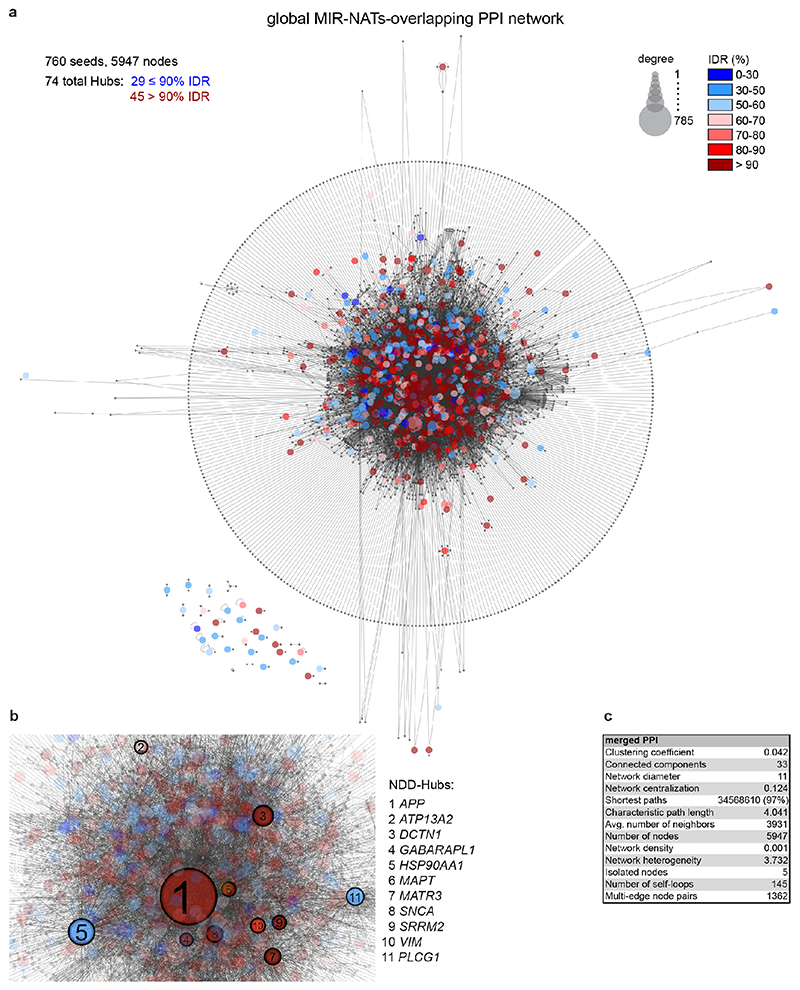
To understand the broader relationships of MIR-NATs to disease, we performed a transcriptomic meta-analysis across three large datasets from post-mortem brains of AD patients, including single nucleus RNA-seq from different cell-types of prefrontal26 and entorhinal cortex27, and bulk RNA-seq from fusiform gyrus28, and identified 446 differentially expressed MIR-NAT S-AS pairs in AD compared to healthy controls. Over 40% of these paired with genes encoding highly intrinsically disordered proteins (IDPs; containing >90% intrinsically disordered regions) (Extended Data Fig. 10). To explore the possibility of functional relationships between proteins encoded by genes paired to MIR-NATs, using PINOT29, we identified an extensive protein-protein interaction (PPI)-network where 95.7% of the 760 seeds share interactors (Extended Data Fig. 11a,c), with 5,947 nodes accounting for 31.2% of the human proteome (19,074 genes), and with 4.04 degrees of separation, lower than 6 observed for large scale-free networks. Mining the human-proteome disorder annotations from D2P2 database30 (http://d2p2.pro), based on 9 predictors, we found 399 seeds (40.3%) of the extended PPI-network are significantly enriched for highly unstructured IDPs, with >90% predicted IDRs (Extended Data Fig. 11a, p=0.0096, 100,000 random simulations, Bonferroni, Supplementary Table3). Depending on position of S-AS overlap of MIR-NAT with coding-gene (5’UTR, CDS, 3’UTR), protein seeds are clustered in 3 PPI-subnetworks (Extended Data Fig. 8e).
Our brain RNA-seq data showed that coding genes with 5’UTR-overlapping MIR-NATs are significantly more expressed in brain compared to genes with 3’UTR or CDS overlaps (Extended Data Fig. 8c), and are strikingly enriched for those involved in neurodegenerative disorders (NDD) (Extended Data Figs. 8e, 9a,b) or in immune functions (Extended Data Fig. 9c,d). Interestingly, the extended PPI-network contains a prevalence of hub proteins (degree ≥40) enriched for IDPs (45/74 hubs with >90% IDR, p=0.0012, Fisher’s exact test). NDD-genes are significantly overrepresented among these hub-IDPs (mean degree=177.8±224; p=0.0029, Fisher’s exact test), and preferentially overlap with MIR-NATs over 5’UTRs (Extended Data Fig. 11b). These data suggest a widespread potential for MIR-NATs in post-transcriptional regulation of many IDPs and in neuronal proteostasis, particularly for NDD-associated genes. IDPs form promiscuous complexes with multiple partners subject to conformational selection upon binding and are metastable, aggregation-prone, dosage-sensitive, often supersaturated and implicated in neurodegeneration31. To avoid prolonged availability and aggregation of surplus IDPs, their expression is tightly regulated at multiple levels, including enrichment of microRNA binding sites and destabilizing PEST sequences6,32. Our data present MIR-NATs as an additional regulatory layer that might contribute to the tightly controlled translation of IDPs6, and present their regulation as new therapeutic opportunities in neurodegeneration.
Methods
Oligonucleotides
The complete list of oligonucleotides used for cloning and for quantitative real-time PCR experiments is included in Supplementary Table1. Oligonucleotides were designed using Primer3Web 4.1.0.
Plasmids
cDNA sequence of human antisense t-NAT1 and t-NAT2l were amplified from a sample of human brain total RNA (Clontech, 636530) with the primers NT1-5’F, NT1-3’R and TOPO2-F, TOPO2-R respectively.
The antisense t-NAT1 5’deletion mutant (Δ5’) was generated by PCR using the oligonucleotides forward NT1Δ5-BamHI and reverse NT1Δ5-XhoI. PCR fragment was cloned directionally in the unique BamHI and XhoI sites in pcDNA3.1V5 (Invitrogen). Similarly, the antisense t-NAT2l 5’ deletion mutant (Δ5’) was generated by PCR using the forward NT2Δ5-BamHI and reverse NT2Δ5-XhoI primers and cloned in the same sites in pcDNA3.1V5.
The antisense t-NAT1 3’ deletion mutant (Δ3’) was generated by PCR using the forward NT1Δ3-BamHI and reverse NT1Δ3-XhoI primers and cloned in the unique BamHI and XhoI sites in pcDNA3.1V5. Similarly, the antisense t-NAT2l 3’ deletion mutant (Δ3’) was generated by PCR using the forward NT2Δ3-BamHI and reverse NT2Δ3-XhoI primers and cloned in the same sites in pcDNA3.1V5.
The antisense t-NAT1 (ΔM1) (partial ΔMir, 386-433) mutant was obtained by cloning of a PCR fragment amplified using the primers (NT1Δ3-BamHI and NT1Δmir1-XhoI) into the BamHI-XhoI sites of pcDNA3.1V5.
The antisense t-NAT1 (ΔM) (total ΔMir, 386-449) mutant was obtained by cloning of a PCR fragment amplified using the primers (NT1Δ3-BamHI and NT1Δmir2-XhoI) into the BamHI-XhoI sites of pcDNA3.1V5.
The antisense t-NAT2l (ΔM) (ΔMir, 498-532) mutant was obtained by cloning of a PCR fragment amplified using the primers (NT2Δ3-BamHI and NT2Δmir-XhoI) into the BamHI-XhoI sites of pcDNA3.1V5.
The antisense t-NAT1 (over) (S/AS overlapping region, 93–168) fragment was generated by direct ligation of in vitro annealed oligonucleotides, with reconstituted 5’-end overhangs, forward NT1overS and reverse NT1overAS (75 nt) onto BamHI and XhoI sites of pcDNA3.1V5. Similarly, the antisense t-NAT1 (Flip) (S/AS overlapping region in a Flipped orientation, 168–93) fragment was generated by direct ligation of in vitro annealed oligonucleotides forward NT1overFlipS and reverse NT1overFlipAS (75 nt) onto BamHI and XhoI sites of pcDNA3.1V5.
The antisense t-NAT1 (Nover) (non-overlapping region, 4–93) mutant was obtained with a similar strategy to antisense t-NAT1 (over). Oligonucleotides forward NT1nonoverS and reverse NT1nonoverAS were annealed in vitro and directionally ligated onto BamHI and XhoI sites of pcDNA3.1V5.
The antisense t-NAT1 (Mflip) (MIR repeat flipped) mutant was obtained as a gene synthesis construct (GENEWIZ) and subcloned into pcDNA3.1V5 using BamHI and XhoI restriction sites. Similarly, antisense t-NAT1 (MIRΔ1, MIRΔ2, MIRΔ3) deleted of motif-1, -2 or -3 respectively were obtained as a gene synthesis construct (GENEWIZ) and subcloned into pcDNA3.1V5 using BamHI and XhoI restriction sites. The miniNAT, consisting only of the AS domain (32 nt) fused together with the full-length MIR element (62 nt), was also obtained as a gene synthetic construct (GENEWIZ) and subcloned using the BamHI/XhoI sites. Fulllength PLCG1-AS lncRNA (ENST00000454626.1, 1,459nt) was designed as a gene synthetic construct (GENEWIZ) and subcloned into pcDNA3.1V5 using BamHI and EcoRV restriction enzymes. Similarly, an PLCG1-AS lncRNA deleted of the inverted MIRb repeat in its third exon (PLCG1-AS ΔM, 1333 nt) was also generated by gene synthesis (GENEWIZ) subcloned into pcDNA3.1V5 using BamHI and EcoRV restriction enzymes.
Cells
SH-SY5Y (ECACC 94030304); SK-N-F1 (ECACC 94092304); HEK-293T (ECACC 12022001) were purchased from Sigma-Aldrich, provided with an authentication certificate by using STR PCR genotyping. All cell lines in culture were regularly tested for mycoplasma using the Lonza Mycoalert detection kit (LT07-318) running in parallel the Mycoalert assay control set (LT07-518) and all lines used for experiments were free of any mycoplasma. Cells were seeded in 75-cm2 flasks in complete medium containing 44% Minimum Essential Medium Eagle (MEME), 44% Ham’s nutrient mixture (F12), 10% fetal bovine serum (Sigma) supplemented with 1% non-essential aminoacids (Sigma), 1% L-glutamine (Sigma), 0.1% Amphotericin B (Gibco), penicillin (50 units ml-1) and streptomycin (50 units ml-1), and maintained at 37°C with 5% CO2. For experiments, 60% confluent cells were plated in 6-well plates (VWR), grown overnight before transfection and harvested 48 hours post-transfection. Transient transfections were done with TransFast (Promega). For establishing the stable cell lines (Empty pcDNA 3.1, t-NAT1FL, t-NAT1Δ5’, t-NAT1Δ3’, t-NAT1over, t-NAT1Flip, t-NAT1Nover, t-NAT1ΔM1, t-NAT1ΔM, t-NAT2FL, t-NAT2Δ5’, t-NAT2Δ3’, t-NAT2ΔM), SH-SY5Y cells were seeded in 10-cm Petri dishes and transfected with TransFast (Promega) and 7.5μg plasmid DNA according to the manufacturer’s instruction. Stable clones were selected by 500 μM G418 sulfate (345810, Millipore). For each type of stable cell line, at least 6 independent clones were isolated using glass cloning cylinders (C1059, Sigma), expanded in 6-well plates and screened individually by Western Blot and qRT-PCR.
Induced pluripotent stem cells (iPSC) and cortical neuron cultures
The control induced pluripotent stem cells (iPSCs) from a healthy male donor used in this study have been described previously33. Ethical permission for this study was obtained from the National Hospital for Neurology and Neurosurgery and the Institute of Neurology joint research ethics committee (study reference 09/H0716/64). iPSCs were authenticated by STR profiling and karyotyping. iPSCs were cultured in feeder-free conditions on Geltrex-coated plates in Essential 8 medium (Thermo Scientific). Media was replaced daily and iPSCs were passaged every 5-6 days with 0.5mM EDTA (Thermo Scientific). iPSCs were subsequently differentiated into cortical neurons, as previously described34, using dual SMAD inhibition followed by in vitro neurogenesis. Briefly, iPSCs were plated at 100% confluency and the media was switched to neural induction media (1:1 mixture of N-2 and B-27-containing media supplemented with the SMAD inhibitors Dorsomorphin and SB431452 (Tocris). N-2 medium consists of DMEM/F-12 GlutaMAX, 1× N-1 insulin, 1 mM l-amino acids, β-mercaptoethanol, 50 U ml–1 penicillin and 50 mg ml–1 streptomycin. B-27 medium consists of Neurobasal, 1× B-27, 200 mM l-glutamine, 50 U ml–1 penicillin and 50 mg ml–1 streptomycin) (Thermo Scientific). At the end of the 10-day induction period, the converted neuroepithelium was replated onto laminin-coated plates using dispase (Thermo Scientific) and maintained in a 1:1 mix of the described N-2 and B-27 media which was replaced every 2-3 days. At around days 25-35, neuronal precursors were passaged further with accutase (Thermo Scientific) and plated for the final time at day 35 onto poly-ornithine and laminin coated plates (Sigma).
iPSC-derived motor neuron cultures
iPSC-derived motor neuron cultures were differentiated from three healthy control lines using a previously established and validated protocol35. The iPSC were derived from three healthy donors: control-1 (age 78, male), control-2 (age 64, male), control-3 (age unknown, female). Two of the control lines used (control-2 and control-3) are commercially available and were purchased from Coriell (cat. number ND41866*C) and ThermoFisher Scientific (cat. number A18945) respectively. Informed consent was obtained from all healthy controls for human iPSC work in this study. Experimental protocols were all undertaken in compliance with approved regulations and guidelines set by UCLH’s National Hospital for Neurology and Neurosurgery and UCL’s Institute of Neurology joint research ethics committee (09/0272). All hiPSC lines were authenticated by STR profiling and karyotyping. At day 14 spinal cord MN precursors were treated with 0.1 μM Purmorphamine for further 4 days before being terminally differentiated for >10 days in 0.1 μM Compound E (Enzo Life Sciences) to promote cell-cycle exit. At relevant timepoints, cells were harvested for Western blot analysis or fixed in 4% paraformaldehyde for immunolabelling.
Lentiviral-shRNAs vectors cloning, preparation and titration
29-mer shRNA sequences were designed by the RNAi-Central shRNA retriever online tool (http://cancan.cshl.edu/RNAi_central/RNAi.cgi?type=shRNA) to target either MAPT-AS1 exon-4 or exon-1 in the non-overlapping region of tNAT1 were ordered as complementary DNA oligonucleotides (IDT) including terminal BbsI restriction sites. An shRNA targeting Renilla luciferase ORF was used as negative control. shRNA oligonucleotides were denatured for 10 min ant 95 °C and annealed in a thermoblock. Their sequences were the following:
shRenilla-S
caccggGTACAAACGCTCTCATCGACAAGGACGGCTtcaagagAGCCGTCCTTGTCGATGAGAGCGTTTGTATTTTTTTGATATCgt
shRenilla-AS
taaaacGATATCAAAAAAATACAAACGCTCTCATCGACAAGGACGGCTctcttgaAGCCGTCCTTGTCGATGAGAGCGTTTGTACcc
shNTIS
caccggGGACGGCGAGGCAGATTTCGGAGCCTtcaagagAGGCTCCGAAATCTGCCTCGCCGTCCTTTTTTTGATATCgt
shNTIAS
taaaacGATATCAAAAAAAGGACGGCGAGGCAGATTTCGGAGCCTctcttgaAGGCTCCGAAATCTGCCTCGCCGTCCcc
shEx4S
caccggGGAGGACAATGTCCTAAGGAATGGAGAGGtcaagagCCTCTCCATTCCTTAGGACATTGTCCTCCTTTTTTTGATATCgt
shEx4AS
taaaacGATATCAAAAAAAGGAGGACAATGTCCTAAGGAATGGAGAGGctcttgaCCTCTCCATTCCTTAGGACATTGTCCTCCcc
All shRNAs were cloned downstream of the U6 promoter into the lentiviral vector pKLV-U6gRNA(BbsI)-PGKpuro2A-BFP, using the BbsI sites. The plasmid was a kind gift of Gabriele Lignani and Eleonora Lugara (UCL, UK). Third-generation LVs were produced by transient four-plasmid co-transfection of 80% confluent HEK293T cells using the Lipofectamine 2000 transfection reagent (Thermofisher Scientific). For each T225 flask the plasmids were co-transfected using the following ratios (pKLV-shRNA 38 μg, pVSVG 13.5 μg, pMDL 18.75 μg pREV 9.3 μg). For each LV-shRNA, supernatants were collected from 2 fully confluent T225 flasks 48h-post transfection, passed through a 0.45 μm Millex-HV filter (Merck Millipore) and purified by ultracentrifugation as previously described36. Viral vectors were titrated by quantitative real-time PCR as previously detailed37. Briefly genomic DNA was extracted from iPS-derived motor neurons (3 div) infected with 5 serial 1:5 dilutions of each LV-shRNA and subjected to qPCR with primers targeting the LV psi packaging region or the BFP ORF (psi-F CTCTCTCGACGCAGGACTC; psi-R TTTGGCGTACTCACCAGTCG; BFP-F GCCTGGCGTCTACTATGTGG; BFP-R TGCTAGGGAGGTCGCAGTAT) and normalised with primers against GAPDH (GAPDH-F TGCACCACCAACTGCTTAGC; GAPDH-R GGCATGGACTGTGGTCATGAG). Viral Integration units per ml, (IU ml–1) were calculated according to the following formula: IU ml–1= (C x N x D x 1,000)/V, where C = proviral copies per genome, N = number of cells at time of transduction (corresponding to about 5 x 105 seeded neurons per well), D = dilution of vector preparation, V = volume of diluted vector added in each well for transduction. LVs concentrations were ranging from 4.5 × 1011 to 7.67 × 1011 transducing units/ml and were normalised to the same concentration of 4.5 x 1011. MN cultures were infected at 3 div by using 250– 2,700 multiplicity of infection (MOI), and neurons were checked for positive transduction at 8–10 div. The efficiency of transduction was estimated by counting neurons expressing BFP protein respect to the total number of SytoX-stained cells, was > 90%.
AAV vectors
Both full-length (FL) and delta-MIR (ΔM) tNAT1 constructs were subcloned from the pcDNA3.1-V5 vector into the pZAC2.1-eGFP adeno-associated virus serotype 9 (AAV9) vector using the PstI and HindIII restriction sites, by removing the SV40 intron and the eGFP ORF. This resulted in AAV9-CMV-tNAT1FL-BGHpA and AAV9-CMV-tNAT1 ΔM-BGHpA vectors respectively. A pair of complementary oligonucleotides bearing the 94-nt long miniNAT sequence flanked by the PstI and HindIII sites were denatured, annealed and ligated into pZAC2.1-eGFP giving rise to the AAV9-CMV-miniNAT-BGHpA vector. Purification and titration of all packaged AAVs were performed by UPenn Vector Core.
Animals, AAV injections and brain tissue processing
All animal studies were performed in agreement with the European Communities Council and approved by the Stockholm North Ethical Committee (reference numbers N166-14 and N1525-2017). All mice used were htau+/- Mapt -/- purchased originally from Jackson laboratory (B6.Cg-Mapttm1(EGFP)Klt Tg(MAPT)8cPdav/J) and bred in house. Mice were housed, maximum five per cage at Astrid Fagreus Laboratory (Karolinska Institutet) with an ambient temperature of 22±1°C and a relative humidity of 50±5%, on a reverse 12-h light/12-h dark cycle, with standard mouse chow and water provided ad libitum throughout the duration of the study. Optimal sample size was determined using G*power v3.1 assuming 4 different groups injected with different AAVs to be compared at a 0.05 significance level. Animals were assigned randomly to experimental and control groups, and within- animal controls were performed wherever possible like in the case of contralateral hemisphere opposite to the site of AAV injection, used as internal negative controls. different groups of stable cell lines and AAV-injected mice were given an alpha-numeric code to blinding investigators soon after sample harvesting. A different blinded investigator prepared and processed samples. Codes correspondence were revealed after quantification for data analysis, and quantifications were repeated in most cases by two independent investigators. A total of 25 htau transgene positive mice were used for randomised stereotactic injections: 5 (3 males, 2 females, mean age 11.89 mo) were injected with 1x PBS buffer, 6 (5 males, 1 female, mean age 12.54mo) with AAV9-CMV-tNAT1-ΔM-BGHpA, 7 (5 males, 2 females, mean age 11.74mo) with AAV9-CM-tNAT1FL-BGHpA and 7 (5 males, 2 females, mean age 12.25mo) with AAV9-CM-miniNAT-BGHpA. All stereotactic surgical procedures were performed on 10–13 months old mice under isoflurane anaesthesia. After the induction of anaesthesia, the animals were placed into a stereotactic frame (David Kopf Instruments). A total of 1 μl of 1.14 × 1014 genome copies/mL of AAV9 vectors or 1x PBS was injected unilaterally into the right hippocampus at the coordinates: AP -0.145 cm, ML -0.15 cm, DV -0.16 cm relative to dura, according to the mouse brain atlas38. All infusions were performed using a 5-μL Hamilton syringe with a 33-gauge needle at a rate of 0.2 μL/30 sec. To prevent reflux, after completion of the infusion the needle was left at the position for an additional 5 min, then slowly retracted a short distance, left in the new position for few seconds and then withdrawn completely. Animals were sacrificed 8 weeks post-injection by cervical dislocation. Brains were quickly dissected, snap-frozen in 2-methyl butane on dry ice and stored at -80°C for RNA and protein extraction following a modified version of a previously published two-steps method39. To avoid introducing positional biases both RNA and proteins were extracted sequentially from the same tissue blocks. Brains were kept on a Petri dish on dry ice for 2 min before cutting first along the sagittal axis to separate ipsilateral (R) and contralateral (L) hemispheres and then coronally, so that the AAV injection site would be spanned by the most anterior right block (R1). For each brain, the resulting 4 blocks (R1, L1, R2, L2) were homogenised in 1ml of TRIzol solution (Life Technologies) on ice using a Tissue Ruptor (Qiagen). After mixing with 200 μl of chloroform, samples were centrifuged at 12,000 g at 4°C, the upper aqueous phase was transferred into new Eppendorf tubes for RNA extraction. The intermediate phase containing proteins and DNA was subjected to DNA precipitation by addition of 100% ethanol and centrifugation at 2,000 g at 4°C. The DNA pellet was stored and 2x the sample volume of isopropanol was added to the phenol-ethanol solution. The samples were incubated at room temperature for 10 min and centrifuged at 12,000 g at 4°C to precipitate the proteins. The protein pellet was washed twice with 95% ethanol, centrifuged at 7,600 g at 4°C, air dried at room temperature for 10 min and solubilised over-night at 50°C with an optimised lysis buffer (40 mM NaCl, 20 mM EDTA, 5% SDS, 100 mM Tris pH 8). To avoid SDS precipitation lysis buffer was pre-incubated at 37°C and supplemented with Complete Protease Inhibitor tablet (Roche Diagnostics) just before usage. To avoid bias introduced by random precipitation induced by high SDS concentration, all protein samples were subjected to dialysis against PBS or TBS using Slide-A-Lyzer MINI dialysis devices (7,000 kDa cut-off, ThermoFisher Scientific) for 2 hours at 4°C. Protein lysates were quantified using DC protein assay (Bio-Rad) and 15 μg were run on Bis-Tris SDS-PAGE 4-12% gels (Bio-Rad) using 3-(N-morpholino)-propanesulfonic acid (MOPS) running buffer (Bio-Rad) and transferred to 0.2 μm nitrocellulose membranes (Bio-Rad).
Double Immunofluorescence
Neurons were fixed in 4% PFA for 25 minutes at room temperature, followed by 10 min permeabilisation in 0.25% Triton-X100/PBS and 30 min blocking in 3% BSA and 0.1% Triton-X100/PBS and incubation with primary antibody overnight at 4°C. The following primary antibodies were used: anti-PAX6 (PRB-278P Covance, Rabbit, 1:500); anti-OTX2 (AB9566-1 Merck-Millipore, Rabbit, 1:500); anti-Ki67 (550609 BD, Mouse, 1:500); anti-TBR1 (ab31940 Abcam, Rabbit, 1:300); anti-SATB2 (ab51502 Abcam, Mouse, 1:100); anti-BRN2 (C-20, sc-6029 SantaCruz, Goat, 1:400); anti-TUJ1 (βIII-tubulin) (Biolegend, 801202 Mouse and 802001 Rabbit, 1:2000). Incubation with secondary Alexa Fluor 488 and 568-conjugated secondary antibodies, (Thermo Scientific) both diluted 1:200 in 3% BSA in 0.1% Triton-X100/PBS, was performed for 1 h at room temperature. Nuclei were stained using DAPI and cells were mounted on slides with Prolong Gold Antifade Reagent (Thermo Scientific). Images were obtained using a Zeiss LSM 710 confocal microscope and the Zeiss ZEN software v2.1.
High Content Imaging (HCI) of motor neurons
Motor neurons were fixed in 4% PFA for 10 minutes at room temperature, followed by 10 min incubation with Wheat Germ Agglutinin CF®680-WGA (Botium, 1:1000), 10 min permeabilisation in 0.25% Triton-X100/PBS and 30 min blocking in 3% BSA and 0.1% Triton-X100/PBS. Neurons were incubated with primary antibody overnight at 4°C. The following primary antibodies were used: anti-tau (DAKO, Rabbit, 1:2000); Anti-NKX6.1 antibody (AF5857, R&D Systems, Goat, 1:1000); Anti-OLIG2 antibody (AB9610, Merck Millipore, Rabbit, 1:500); Anti-ChAT antibody (AB144P, Merck Millipore, Goat, 1:100); Anti-SMI32 antibody (801701, BioLegend, mouse, 1:1000); anti-TUJ1 (βIII-tubulin) (801202, Biolegend, Mouse, 1:2000). The following secondary antibodies were used: Alexa Fluor Donkey anti-goat IgG 488 (A-11055, Invitrogen); Donkey anti-mouse IgG 568 (A-10037, Invitrogen); Donkey anti-rabbit IgG 647 (A-31573, Invitrogen); Donkey anti-rabbit IgG 594, (Thermo Scientific), diluted 1:1000 in 3% BSA in 0.1% Triton-X100/PBS, and incubated for 1 h at room temperature. Nuclei were stained either with 167mM SYTOX™-Green (Invitrogen) or alternatively with 200 mM DAPI (Invitrogen) and cells were imaged in PBS 1x. For each condition 5-15 wells were taken, with a minimum of 5 fields acquired from each well, using an OPERA-Phoenix high-content screening platform (PerkinElmer). Images were analysed using Harmony 4.5 and Fiji 2.040 or alternatively Columbus v2.8.0.138890.
Splinkerette PCR
Sites of integration of individual clones of stable cell lines were determined following a method previously described41. Approximately 1x106 stable cells were used for each clone and genomic DNA was extracted using the Gentra Puregene kit (Qiagen) according to the manufacturer’s instruction. DNA integrity was assessed on 1% agarose gel and DNA purity and concentration were measured by UV spectrophotometer (Eppendorf). For each clone, 1μg DNA was digested with 10 units of BstYI restriction enzyme (New England Biolabs) in 35 μl volume, at 60°C overnight, followed by heat-inactivation at 80°C for 20 min. 6 μl of annealed double stranded splinkerette linkers:
(SPLNK-TOP: GATCCCACTAGTGTCGACACCAGTCTCTAATTTTTTTTTTCAAAAAAA SPLNK-BOT:
CGAAGAGTAACCGTTGCTAGGAGAGACCGTGGCTGAATGAGACTGGTGTCGACACTAGTGG) were ligated onto the ends of genomic DNA fragments using 600 units of T4 DNA ligase (New England Biolabs) in 50 μl volume, incubating for 3 h at room temperature. Fragments containing the integrated target DNA were amplified by PCR with Phusion Taq polymerase (Fynnzymes) for 32 cycles using a forward primer (SPLINKF2 GGGAGGATTGGGAAGACAATAGC) annealing to the target gene and a reverse primer specific to the splinkerette linker (SPLNK#1 CGAAGAGTAACCGTTGCTAGGAGAGACC). A nested PCR was performed using the primers (SPLINKF3 CTATGGCTTCTGAGGCGGAAAGAA, SPLNK#2 GTGGCTGAATGAGACTGGTGTCGAC). The first-round reaction was heated to 98°C for 75 seconds, followed by two cycles of 98°C for 20 seconds and 64°C for 15 seconds. A further 30 cycles of 98°C for 20 seconds, 68°C for 15 seconds and 72°C for 2 minutes was followed by a final extension at 72°C for 7 minutes. The round 2 reaction was heated to 98°C for 75 seconds, followed by 30 cycles of 98°C for 20 seconds, 68°C for 15 seconds and 72°C for 90 seconds. Final extension occurred at 72°C for 7 minutes. A 5μl aliquot of the round 2 PCR product was resolved by agarose electrophoresis to confirm the presence of a single band. The remaining PCR product was purified using the QIAquick PCR purification kit (Qiagen) and sequenced using a primer that anneals to the pcDNA3.1V5 vector (SplinkSeq: CCCTGTAGCGGCGCATTA). The resulting sequence was aligned to the human genome using the Blat tool of UCSC genome browser hg19.
RNA-seq library preparation and sequencing
Brain samples for analysis were provided by the Medical Research Council Sudden Death Brain and Tissue Bank (Edinburgh, UK). Post-mortem human tissue transcriptomic analysis was approved by the National Hospital for Neurology and Neurosurgery & Institute of Neurology Joint Research Ethics Committee, UK (REC reference number 10/H0716/3). All four individuals sampled were of European descent, neurologically normal during life and confirmed to be neuropathologically normal by a consultant neuropathologist using histology performed on sections prepared from paraffin-embedded tissue blocks. Twelve central nervous system regions were sampled from each individual. The regions studied were: cerebellar cortex, frontal cortex, temporal cortex, occipital cortex, hippocampus, the inferior olivary nucleus (sub-dissected from the medulla), putamen, substantia nigra, thalamus, hypothalamus, intralobular white matter and cervical spinal cord.
RNA was extracted using Qiagen tissue kits (Qiagen, US), and quality controlled as detailed previously42. Libraries were prepared by the UK Brain Expression Consortium in conjunction with AROS Applied Biotechnology A/S (Aarhus, Denmark). In brief, 100 ng total RNA was used as input for cDNA generation using NuGen’s Ovation RNA-seq System V2 (NuGen Technologies, US). The RNA was processed according to the manufacturer’s protocol resulting in amplified cDNA from total RNA and concomitant de-selection of rRNA. Importantly, reverse transcription in this protocol is carried out using both oligo dT and random primers. Total RNA profile patterns were assessed with the latter and locations of splicing were inferred. 1μg of the cDNA was fragmented using a Covaris S220 Ultrasonicator and the fragmented cDNA was used as the starting point for Illumina’s TruSeq DNA library preparation (Illumina, US). Finally, library molecules containing adapter molecules on both ends were amplified through 10 cycles of PCR. The libraries were sequenced using Illumina’s TruSeq V3 chemistry / HiSeq2000 and 100 base pair paired-end reads. The sequencing data was converted to fastq-files using Illumina’s CASAVA 1.8.2 Software.
qRT–PCR
Total RNA was extracted from cells and human post-mortem brain tissue samples (temporal cortex, occipital cortex, caudate) using Trizol reagent (Invitrogen) according to the manufacturer’s instruction. A panel of RNA from 20 different normal human tissues (each consisting of pools of three tissue donors with full documentation on age, sex, race, cause of death) was obtained from Ambion (AM6000). All RNA samples were subjected to DNase I treatment (Roche). A total of 1 μg of RNA was subjected to retrotranscription using SuperScript III cDNA synthesis kit (Invitrogen) and Real Time qRT–PCR was carried out using the SYBR green fluorescence dye (Power SYBR Green Master mix, Applied Biosystems) on a Stratagene Mx3000P thermo-cycler. TATA-binding protein (TBP) and Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) were used as housekeeping genes to normalize different samples as tested by the GeNorm program, incorporated into qbase+ (http://medgen.ugent.be/~jvdesomp/genorm/). All experiments were analysed in Microsoft excel 2011. The amplified transcripts were quantified using the comparative Ct method and the differences in gene expression were presented as normalized fold expression (ΔΔCt). All of the experiments were performed in triplicate. A heat map graphical representation of rescaled normalized fold expression (ΔΔCt/ΔΔCtmax) was obtained by using Matrix2png (http://www.chibi.ubc.ca/matrix2png/). A list of oligonucleotides used for qRT–PCR experiments is in Supplementary Table1.
Two-colour single-molecule RNA fluorescent in situ hybridization (sm-FISH)
A set of 48 antisense 20bp-long DNA tiling probes complementary to 3 alternative splicing isoforms of human t-NAT transcripts (t-NAT1, t-NAT2s, t-NAT2l) were designed by using Stellaris Probe designer 2015 (http://www.biosearchtech.com/stellarisdesigner/), and were labeled at 3’ -end with the fluorescent dye Quasar 670. Another set of 26 antisense DNA tiling probes complementary to the exons of human MAPT transcript (NM_005910) were labeled at the 3’-end with the dye Quasar 570. All FISH probes (as reported at the end of the Methods section) were 19 to 20 bp long, designed with a stringency factor 2, checked using BLAST 2.2.28, and obtained from Biosearch technologies. Fluorescent in situ hybridization was performed as previously described4. Briefly, cells were fixed with 3.7% formaldehyde (Pierce) in PBS for 10 min at room temperature, washed twice in PBS and permeabilized with 70% ethanol at 4 °C for 1 h. Probes were resuspended in hybridization buffer containing 100 mg ml–1 dextrane sulphate (Sigma), 10% deionized formamide (Ambion), 5% BSA (Roche), 0.1 mg ml–1 yeast tRNA (Sigma) in 2x SSC (Sigma). Hybridization was performed with probes at a concentration of 125 nM at 37 °C for 16 h in a humidified chamber. Cells were then washed twice in wash buffer containing 10% formamide (Ambion), 2x SSC (Sigma) counterstained with 5 ng ml–1 DAPI, washed once in SSC2x and mounted with Vectashield (Vector labs) mounting medium. Images were obtained with a fluorescence microscope (Leica DM5500-B) using the Leica Application Suite X v3.6.0.20104.
siRNA Knockdown
SH-SY5Y cells were seeded at 70% of confluence in 6-well plates, and after 24 h were transfected with 75 μl of 2μM siRNAs, using RNAiMax (Invitrogen) transfection reagent following manufacturer’s instructions. After 48 h cells were harvested for protein and RNA extraction. Three independent pools of siRNAs (Ambion) were used to target different MAPT-AS1 exons as follows:
siNT1nover (S, CGGCGAGGCAGAUUUCGGAtt; AS, UCCGAAAUCUGCCUCGCCGtc);
siNT2nover (S, GCCGCCGAGUCCGUCCACAtt; AS, UGUGGACGGACUCGGCGGCcg);
siEx4-n268302 (S, AGGACAAUGUCCUAAGGAAtt; AS, UUCCUUAGGACAUUGUCCUcc);
siEx4-n268298 (S, GAUUUGUCAUGAGUCUCUUtt; AS, AAGAGACUCAUGACAAAUCaa).
A scrambled sequence #2 was also used as negative control. Both pre-designed and custom-designed were LNA-modified as Silencer® Select siRNAs (Ambion).
Protein dephosphorylation
After lysing cells in RIPA lysis buffer supplemented with complete EDT A-free protease inhibitor cocktail (Roche Diagnostics) and determining protein concentration by the DC protein assay (Bio-Rad), lambda protein phosphatase (NEB, USA) was used to dephosphorylate protein lysates. Approximately 400 units of enzyme dephosphorylates ~40μg of lysate. The dephosphorylation mixture was prepared using 40μg of lysate with 8μL of 10x PMP buffer (50 mM HEPES, 100 mM NaCl, 2 mM DTT, 0.01% Brij 35, pH7.5), 8μL of MnCL2 (2mM) and 400 units of enzyme (1μl). The mixture was incubated in a water bath at 30°C for 3 hours for optimal dephosphorylation. The lambda phosphatase was inactivated by adding 4x XT samples buffer (Bio-Rad) and 10x NuPAGE reducing agent (ThermoFisher), denatured at 95°C for 10 min and half volume (corresponding to ~20ug) separated in 4-12% SDS–polyacrylamide gel (Bio-Rad) in MOPS buffer.
Western blots
Cells were lysed in cold RIPA lysis buffer supplemented with complete EDTA-free protease inhibitor cocktail (Roche Diagnostics). Protein lysate concentrations were measured by the DC protein assay (Bio-Rad). For each sample 20 μg proteins were separated in 4-12% SDS–polyacrylamide gel (Criterion XT Bis-Tris, Bio-Rad) in MOPS buffer and transferred to 0.2 μm nitrocellulose membrane (Trans-Blot Turbo Transfer pack, 1704159 Bio-Rad) for 10 min at 2.5A constant, using the Trans-Blot Turbo Transfer system (Bio-Rad). Immunoblotting of neuroblastoma cells and motor neurons was performed with the following primary antibodies: anti-tau (T-1308-1, rPeptide 1:60,000 and A0024 DAKO rabbit polyclonal 1:15,000), anti-β-actin (A2228, Sigma 1:2,000), anti-SPPL2C polyclonal antibody (12664-1-AP, Proteintech 1:1,000) and anti-TDP43 (10782-2-AP, Proteintech, 1:1,000), anti-PLCG1 (D9H10, rabbit monoclonal, Cell Signaling, 1:1,000), anti-GAPDH (G8795 Sigma, 1:10,000). Mouse brain lysates were additionally immunoblotted with the CP13 mouse monoclonal antibody (kind gift from Prof. Peter Davies, Einstein College-USA) to detect p-Ser202 phosphorylated-tau, and with chicken anti-GFP antibody (GFP-1020, Aves Labs, 1:3,000). Secondary antibodies (1:15,000) were as follows: infrared IRDye®-800CW goat anti-rabbit (P/N 926-32211), donkey anti-mouse (P/N 926-32212), donkey anti-chicken (P/N 926-32218) or IRDye®-680RD donkey anti-rabbit (P/N 926-68073), goat anti-mouse (P/N 926-68072) or donkey anti-goat (P/N 926-68074), IgG (Li-COR Bioscience). Signals were digitally acquired by using an Odyssey Fc infrared scanner (Li-COR Bioscience) and quantified using Fiji version 2.0.0-rc-39/1.50d59 or Image Studio 5.2 (Li-COR Bioscience).
Cellular fractionation
Nucleo-cytoplasmic fractionation was performed using Nucleo-Cytoplasmic separation kit (Norgen) according to the manufacturer’s instruction. RNA was eluted and treated with RNAse-free DNase I (Roche). RNA concentrations were measured by NanoDrop spectrophotometer. The purity of the cytoplasmic fraction was confirmed by qRT–PCR on pre-ribosomal RNA.
Luciferase reporter vectors
Firefly luciferase reporter plasmids were constructed by inserting the human MAPT core promoter (CP, 1,342bp) amplified using the primers (CP-F GAGCTCCAAATGCTCTGCGATGTGTT, CP-R GCTAGCGGACAGCGGATTTCAGATTC) between the SacI and NheI sites into pGL4.10 vector (Promega) to create pGL4-CP vector. A 901bp fragment of genomic DNA spanning the t-NAT promoter (NP) was amplified using the primers (NP-F gaGCTAGCTGCCGCTGTTCGCCATCAG, NP-R gtGCTAGCACCCTCAGAATAAAAGCCAG) and inserted into NheI site either of pGL4-CP or pGL4.10 vectors to create pGL4-CNP and pGL4-NP respectively. The full-length 322bp-long human MAPT 5’UTR was amplified with primers (pRTF-EcoRI, pRTF-NcoI) and ligated onto EcoRI and NcoI sites of the pRF vector (a kind gift from Prof. Anne Willis, Leicester University, UK) to create the pRTF vector. A fragment of MAPT 5’UTR devoid of t-NAT overlapping region was amplified using the primers (pRTF-EcoRI, pRTFDover-NcoI) and inserted between same sites into pRF, to generate the pRTF-Delta vector. pRTFover vector was constructed in the same way using the primers (pRTF-Dover-EcoRI, pRTF-NcoI). A pRhcvF, used as a positive control viral IRES, was a kind gift of Prof. Anne E. Willis and was constructed as described previously43. Mutant reporter plasmids were created using the QuickChange lightning multisite-directed mutagenesis kit (Agilent) according to the manufacturer’s instructions. The following mutagenic oligonucleotides (pRTF-mTOP) were annealed to the pRTF vector, extended by PCR, and the parental methylated plasmid DNA was digested with DpnI enzyme to obtain the correspondent mutant bicistronic luciferase vector. The full-length human MAPT 3’UTR and 3 partially overlapping fragments were amplified from brain cDNA with the primers (Fr1-F, Fr1-R, Fr2-F, Fr2-R, Fr3-F, Fr3-R) and cloned individually into SacI and HindIII sites of pMIR-REPORT vector (Invitrogen).
Dual Luciferase Reporter Assay
SH-SY5Y cells or t-NAT-stably expressing cells were seeded in Greiner 96-well plates overnight and then cotransfected using TransFast (Promega) with the bicistronic reporter vector pRF, pRhcvF, pRTF or pRTF deletion mutants and either a pcDNA3.1 empty vector or each of the t-NAT expression vectors. 48 h after transfection cap-dependent translation (Renilla luciferase activity) and IRES-mediated translation (firefly luciferase activity) were measured with the DualGlo Luciferase Assay kit (Promega) according to the manufacturer’s instructions. Luminescence on Greiner 96-well plates was quantified using a Spark 10M microplate reader (Tecan) and the software SPARKCONTROL v1.2. Firefly to Renilla ratios were normalized to a common pMIR-Report vector used to account for transfection efficiency in each experiment and results are represented as mean ± s.d. Experiments were done in triplicate.
In Vitro Transcription-Translation (IVTT) Assay
PCR amplicons for t-NAT (FL, ΔM, miniNAT) and the pTF reporter, containing Firefly luciferase ORF downstream of MAPT 5’UTR, were amplified from 2 ng of their plasmids using Platinum II Taq hot start DNA polymerase (Invitrogen), with the following conditions: 2min 94°C, 32 cycles (15s 94°C, 15s 60°C, 50s 72°C), 7min 72°C. T7fwd (TAATACGACTCACTATAGG) and BGHrev (CCTCGACTGTGCCTTCTA) oligonucleotides were used for amplifying t-NAT constructs.
T7MAPT5utr-Fwd (TAATACGACTCACTATAGCGGACGGCCGAGCG) and 3FlucPolyA-Rev (TTTTTTTTTTTTTTTTTTTTCGCCCCGACTCTAGAATTACAC) primers were used to amplify pTF. PCR products were purified on 1% agarose gel using the Qiaquick gel extraction kit (Qiagen) and quantified by spectrophotometry. 200 ng of each amplicon were used as template for in vitro transcription using the mMESSAGE-mMACHINE T7 Transcription kit (Invitrogen, AM1344), and incubated at 37°C for 2 hr following manufacturer’s instructions. In vitro transcribed (IVT) RNAs were purified with the MEGAclear Transcription Clean-Up kit (Invitrogen, AM1908) and quantified by spectrophotometry. 100 ng of pTF-luciferase reporter m7G capped-RNA (155.7 fmoles) were mixed with t-NAT FL, ΔM or miniNAT IVT RNAs in 1:0, 1, 5, 10, 20 molar ratio, in the presence of 1x Translation Mix (-Met), 50μM unlabelled Metionine (Sigma), 17μl Reticulocyte lysate in 25μl volume using the ReticLysate kit (Invitrogen, AM1200). Reactions were incubated at 30°C for 3hr in a water bath. In vitro translation of the pTF reporter was measured with DualGlo Luciferase Assay kit (Promega), according to manufacturer’s instructions. Luminescence of each sample in triplicate on Greiner 96-well plates was quantified using a Spark 10M microplate reader (Tecan) and the software SPARKCONTROL v1.2. Results of three independent experiments were subject to linear regression and ANCOVA analysis using the car package v3.0-3 (R3.5.3), to assess differences in slope and intercept for each t-NAT construct.
Polysomal fractionation
1×106 cells were seeded in two 10 cm2 dishes and collected for polysomal fractionation after 48 h. All the experiments were run in biological triplicate. Cells were incubated for 4 min with 100 μg/ml cycloheximide at 37°C to block translational elongation. Cells were washed with PBS supplemented with 10 μg/ml cycloheximide, scraped into 300 μl lysis buffer (10 mM NaCl, 10 mM MgCl2, 10 mM Tris-HCl, pH 7.5, 1% Triton X-100, 1% sodium deoxycholate, 0.2 U/μl RNase inhibitor (Fermentas Burlington, CA), 100 μg/ml cycloheximide and 1 mM DTT) and transferred to a microfuge tube. Nuclei and cellular debris were removed by centrifugation at 13,000g for 5 min at 4°C. The supernatant was layered on a linear sucrose gradient (15-50% sucrose (w/v) in 30 mM Tris–HCl at pH 7.5, 100 mM NaCl, 10 mM MgCl2) and centrifuged in a SW41Ti rotor (Beckman Coulter, Indianapolis, IN) at 180,000g for 100 min at 4°C. Ultracentrifugation separates polysomes by the sedimentation coefficient of macromolecules: gradients are then fractionated and mRNAs in active translation (polysome-containing fractions) are separated from untranslated mRNAs (subpolysomal fractions). Fractions of 1 ml volume were collected with continuous absorbance monitoring at 254 nm.
qRT-PCR of polysomal fractions and statistical analysis
Total RNA was extracted from each polysomal fraction using 1ml of Trizol (Invitrogen) following manufacturer’s instructions. After DNAse I treatment, equal volumes of RNA were retrotranscribed in the presence of an equimolar mixture of oligo dT and random hexamer, using SuperScript III (Invitrogen). For the statistics of polysome fractionation qRT-PCR analyses, the raw Ct value for each of the individual fractions was transformed to 2–Ct and normalized to the sum total for all fractions, generating a percentage of total transcript within each fraction. Each fraction’s values were aggregated into different categories corresponding to different phases of polysome assembly on a total RNA absorbance curve. For qRT-PCR analysis we followed a previously published method44. Briefly: fractions 1 and 2 were summed into “40S–60S”; fractions 3 and 4 were summed into “80S”; fractions 5-7 were summed into “light”; fractions 8-10 were summed into “medium” and fractions 11–13 were summed into “heavy”—corresponding to peaks on total RNA absorbance curves monitored during fractionation. For significance testing of qRT-PCR data, t tests were conducted between Empty vector and t-NAT-expressing cells in each category, with p < 0.05 considered significant.
RIBO-seq
Ribosome footprints were isolated as previously described45 from SH-SY5Y cells stably expressing different MAPT-AS1 constructs or an empty vector (n=3 independent clones for each construct), except with a lowered concentration of RNase I (Thermo Scientific, EN0601) to reduce rRNA contamination (10 U/50 μg of total RNA). From each sample, an aliquot of ~ 1 μg total RNA was taken before RNase I treatment for QuantSeq. rRNA was depleted using an RNase H-based methodology46, but with standard RNase H (NEB) and at a lowered incubation temperature of 37°C. Library preparation followed the irCLIP protocol47 and the libraries were sequenced using Illumina PE150 by BGI Genomics. The 3’ adapters were trimmed using Cutadapt v2.1048, then reads were demultiplexed using a custom script, pre-mapped to common RNA contaminant sequences using Bowtie 249, and aligned to the human hg38 genome using STAR v2.350 and PCR duplicates were removed using UMI-Tools 1.0.051. Differential translation analysis was performed using DESeq252.
QuantSeq
QuantSeq FWD53 libraries (Lexogen) were generated from RNA isolated from the aforementioned aliquots. Reads were aligned to the human hg38 genome using STAR v2.350 and differential expression was analysed using DESeq252, filtering for genes with at least 1,000 counts across 18 samples. All scripts are on Github.
Bioinformatic analyses
Bedtools v2.254, Python 2.7.5 (http://www.python.org) and R v.3.1.1 (https://www.r-project.org) were used extensively during analysis unless otherwise specified. All plots were produced using R package ggplot2 v3.2.055 and data processing was done using dplyr v0.8.3 (http://CRAN.R-project.org/package=dplyr) and tidyr v0.8.3 (https://cran.r-project.org/web/packages/tidyr/index.html). The following open source R packages were also used: car3.0-3, ComplexHeatmap v1.20.0, circlize v0.4.6, reshape2 v1.4.3, RColorBrewer v1.1-2, grid v3.5.3, fastcluster 1.1.25, gtools v3.8.1, ggpubr v0.2.1, ggsignif v0.5.0, ggpmisc v0.3.1., DESeq2 v1.22.2.
MAPT-AS1 evolutionary conservation across primates
Multiple sequence alignment of the human t-NAT1 and t-NAT2l transcript with the genomic sequences of 10 non-human primates (baboon, bonobo, chimp, gibbon, gorilla, marmoset, mouse lemur, orangutan, rhesus, squirrel monkey). Sequences were aligned using MUSCLE 3.856, and displayed using Jalview 257. MAPT-AS1 protein-coding potential were scored by PhyloCSF 1.0.1-058 (https://github.com/mlin/PhyloCSF/wiki). Evolutionary conservation of MAPT-AS1 promoter region across 6 distant species (Homo sapiens, Macaca mulatta Mus musculus, Rattus norvegicus, Canis familiaris, Bos taurus), was computed using the ECR browser59 (https://ecrbrowser.dcode.org). CAGE and nanoCAGE60 tag clusters from FANTOM4 and FANTOM5 datasets were retrieved from ZENBU genome browser v2.1161 (https://fantom.gsc.riken.jp/zenbu/).
Combining all transcript exons into single gene annotations
For each gene a single non-overlapping list of exons was created, by merging exons from all transcripts. Each exon was defined as either 5’UTR, 3’UTR or CDS using GENCODE v19 comprehensive (hg19 build) annotations (http://www.gencodegenes.org/releases/19.html). All exons with multiple annotations were preferentially defined as either 5’UTR or 3’UTR. All further analysis utilized this annotation.
Identifying overlapping lncRNA – protein-coding gene S-AS pairs and defining gene groups
For the identification of additional translational repressor candidates, we searched for GENCODE v19 transcripts that were non-coding RNAs and overlap the 5’ UTR, CDS or 3’UTR of coding transcripts in a head-to-head configuration. All protein-coding genes were intersected with lncRNAs from GENCODE v19 and these lncRNAs were then checked for overlaps with MIR elements from RepeatMasker v4.0.5 (www.repeatmasker.org). These intersections were used to create the following groups:
All protein coding genes
Protein coding genes without lncRNA overlap
Protein coding genes with lncRNA overlap
Protein coding genes that overlap lncRNA that include MIR elements
Protein coding genes that overlap lncRNA that do not include MIR elements
Various analyses were applied to these groups, namely:
Calculating an estimate of gene feature length relative to exon number
From the non-overlapping exon annotations we were able to calculate a normalized number of exons per gene region (5’UTR, 3’UTR or CDS) by dividing the total number of exons within all gene transcripts by the sum of transcripts. This value was used to divide by the total length of gene region to estimate the length of feature compared to the number of exons. A one-way ANOVA followed by Bonferroni’s multiple comparisons test was performed on the different gene groups to determine if the distributions between groups were significantly different.
Predicting secondary structures for protein-coding gene UTRs
For each gene the longest 5’UTR and 3’UTR were selected as representative for the gene. RNAfold v2.1.9 from the ViennaRNA package62 was used to predict the minimum free energy (mfe) of the secondary structure (kcal/mol). A one-way ANOVA followed by Bonferroni’s multiple comparisons test was performed on the different gene groups to determine if the distributions between groups were significantly different.
Calculating the MIR element nucleotide overlap per transcript
The non-overlapping length of each gene feature or lncRNA transcript was divided by the number of base pairs overlapping a RepeatMasker (v4.0.5) defined MIR repeat element. This provided an indication of relative abundance of MIR elements across the human transcriptome.
Gene expression analysis of postmortem brain tissue
Post-mortem, total RNA sequence data was aligned using the STAR50 aligner v2.3 with default settings and GENCODE v21 annotations. Gene counts and FPKM values were calculated based on the non-overlapping annotation for each gene using Bedtools v2.254 and custom python scripts. Counts across splice-junctions were quantified by MISO v1.0 (https://github.com/yarden/MISO/blob/fastmiso/docs/source/sashimi.rst). All regions were merged into a single mean value to describe whole brain expression of protein-coding genes.
Linear regression analysis of postmortem brain RNA-seq and tau pathology (Braak-stage)
RNA-seq data together with Luminex-immunoassay and tau-IHC data from the Allen Brain Institute cohort were retrieved from (http://aging.brain-map.org/)9. To have more statistical power, data from hippocampus, temporal and parietal cortex, and frontal white matter were aggregated. Linear regression of MAPT-AS1 expression (normalised FPKM) against different Braak-stages was performed using the stat_cor() function of ggpubr R package, computing the Pearson’s correlation coefficients and p-values. The same linear regression analysis was performed for correlating tau pathology (phospho-tau(AT8):total-tau ratio, Luminex-immunoassay) to Braak-stage. Similarly, bulk RNA-seq data of dorsolateral prefrontal cortex from the ROS-MAP cohort (https://dx.doi.org/10.7303/syn3388564)10 was used for linear regression analysis to correlate MAPT-AS1 expression (normalised FPKM) with Braak-stage. Results are plotted in Fig.1e.
Transcriptomic meta-analysis of snRNA-seq and bulk RNA-seq of AD brain cell types
Single-nucleus RNA-seq expression data was obtained from two different studies that compared AD cases with controls. In the first case (Mathys)26 the data was obtained from the supplementary data in their publication, in the second case (Grubman)27, data was retrieved from their interactive webpage (http://adsn.ddnetbio.com/). These data were already processed. For Mathys dataset, the pathology vs no pathology set was selected, and the genes that were indicated as differentially expressed (DEG) according to their criteria (log2FC > 0.25, fdr-adjusted p-value < 0.01 and fdr-adjusted p-value of the mixed Poisson model < 0.05) were selected. For the Grubman data, log2FC > 0.5 (the lowest cutoff available) and adjusted p-value < 0.05 were used.
Bulk RNA-seq data of AD and controls from Friedman dataset28 (GEO accession GSE95587) was also used. Raw counts were selected and the standard limma-voom63,64 pipeline was followed to obtain the resulting DEGs, setting Diagnosis as the variable of interest (with AD vs Control as the contrast matrix) and RNA Integrity Number (RIN) and Sex as covariates. Log2FC > 0.25 and adjusted p-value < 0.05 were set as cutoffs for the GSE95587 data. In all three cases, the genes that matched our gene-set and were differentially expressed according to these filters were selected. DEGs were plotted using ComplexHeatmap v1.20.0 (http://bioconductor.org/packages/release/bioc/html/ComplexHeatmap.html)65.
Gene Ontology (GO)-terms enrichment analysis
1,045 MIR-NAT protein-coding target genes were divided by the type of their exonic overlap into three groups. Genes in each category were analyzed for GO terms enrichment using Enrichr (Sept-2014)66 (https://amp.pharm.mssm.edu/Enrichr/), and the results for the top 10 most enriched terms are reported in the form of bar plot, with the length of each bar being proportional to a combined score c=log(p)z, where p represents the p-value computed using the Fisher exact test, and z is the z-score computed by assessing the deviation from the expected rank. The same groups of genes were also tested separately using WebGestalt 201367 (http://www.webgestalt.org/2013/), obtaining similar results.
Gene Network analysis and representation
1,045 MIR-NAT protein-coding target genes were analysed for their potential interactions. Protein-protein interactions (PPIs) for the 3 gene-lists of interest (3’UTR, 5’UTR and CDS) have been extracted from literature using the scripts now embedded in the Protein Interaction Network Online Tool (PINOT 1.1)29 (scripts are freely downloadable from http://www.reading.ac.uk/bioinf/PINOT/PINOT_form.html). PPIs were downloaded from the following PPI repositories: APID Interactomes, BioGrid, bhf-ucl, InnateDB, InnateDB-All, IntAct, mentha, MINT, UniProt and MBInfo through the PSICQUIC platform on April 2018. The interactions were then processed to remove poorly annotated entries and duplicated annotations across databases. Interactions were finally scored based on the number of different methods and different publications reporting them. A threshold was applied to retain PPIs that have been experimentally replicated at least twice (final score > 2). More information regarding QC and scoring can be found in 29. The seeds have been then overlapped with a panel of neurodegenerative genes (Neurodegenerative Disorders, C0524851 DisGenNet v6.0 -April 2019, https://www.disgenet.org) to evaluate a possible overrepresentation of genes involved in neurodegenerative conditions. The overlap of the 3 lists of seeds revealed the presence of ~6.6% genes previously linked to neurodegeneration for the 5’UTR (p=1.497x10–5), while for the 3’UTR and the CDS the overlap was ~3% (Fig.7a). To verify whether this result was statistically significant, highlighting a true enrichment instead of just being driven by chance, 100,000 random simulations have been run matching the genes from DisGenNet to a number of randomly selected genes (matching numbers of seeds). The comparison between the simulated versus real experiments gave us the confidence value associated with the enrichment of neurodegenerative genes within the interactomes (p-value were calculated with the pnorm R function). Graphs have been obtained through Cytoscape 3.7.168, scripts were written in R 3.5.3. To limit the number of nodes within the network, PPI interactions only first-degree interactors were computed. Similar enrichment for neurodegenerative disease genes was observed in a global PPI network computed using NetworkAnalyst 3.069 (https://www.networkanalyst.ca/), starting from a total of 392 seed proteins searched within the InnateDB PPI database (https://www.innatedb.com) (Extended Data Fig. 7a,c). Genes within the PPI network were then searched for enriched diseases and pathways, using the default options of NetworkAnalyst 3.0. Neurodegenerative disease associated genes were additionally annotated using WebGestalt 201367 disease enrichment analysis.
Instrinsically disordered proteins predictions and overrepresentation analysis
1,045 MIR-NAT protein-coding target genes were analysed for the presence of predicted intrinsically disordered regions (IDR). 989 out 1,045 genes were mapped to ENSG IDs (Ensembl v63), and missing IDs were discarded from the rest of the analysis. IDR predictions by 9 different predictors (Espritz-D, Espritz-N, Espritz-X, IUPred-L, IUPred-S, PrDOS, PV2, VLXT, VSL2b) for the whole human-proteome were downloaded from the D2P2 database30 (http://d2p2.pro). The browse function of the D2P2 was used to retrieve all annotated genes to have a predicted %IDR over a specified threshold lower value, and with at least 75% concordance among all predictors, resulting in 8 lists of genes according to the following %IDR thresholds (0%, 30%, 50%, 60%, 70%, 80%, 90% and 100%). 19,074 gene IDs retrieved using the 0% IDR threshold and 75% concordance among all predictors were used as representative of the whole human proteome (Ensembl v63-D2P2 annotations). Gene IDs from pairs of lists were then matched and filtered using Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/) according to the percentage of IDR covering their ORF, in the following bins (0-30%, 30-50%, 50-60%, 60-70%, 70-80%, 80-90% and >90%). We compared %-30%IDR, 50%-90%IDR, and >90%IDR groups. Statistical overrepresentation of the >90%IDR IDPs was computed by 100,000 randomized simulations of PPI-networks with the same number of seeds using a custom R script.
k-mer enrichment analysis and complementarity to 18S rRNA
Each antisense-MIR-NAT was mined for 7-mers that appeared within the overlapping protein coding targets 5’ UTR and showed complementarity to the active region within the 18S rRNA (nt 812 to 1233, 1859-1871), as defined previously24. These 7-mers were then checked to ensure they resulted from the MIR element within the antisense-MIR-NAT. This approach was implemented using a custom python script.
MAPT SNP genetic data analysis and linkage disequilibrium analysis of MAPT-AS1 region
Frequency data for the two SNPs (rs62056779 and rs11575895) within the MAPT 5’UTR were obtained from publicly available data from GWAS-meta-analyses of 13,708 PD cases and 95,282 healthy control subjects, deposited in the PDGene database (http://www.pdgene.org). To test for genetic association with Parkinson’s disease, allele frequencies were analysed according to the guidelines reported by Nalls and colleagues20. SNPs within MAPT-AS1 genomic region that are linked (R2≥0.5) to tagging SNPs from the NHGRI GWAS catalog are reported. The specific trait associated to each tagging SNP together with the p-value from the GWAS study. All p-values ≤5x10–8 were considered to be significant. Linkage disequilibrium (LD) correlations (R2) were calculated using LDlink1.170 for different populations. Pairwise linkage disequilibrium heatmap created using LDmatrix (https://ldlink.nci.nih.gov/?tab=ldmatrix).
Statistical analysis
Statistical analyses were performed using GraphPad Prism 7, R 3.5.3 and RStudio 1.2.1335 unless otherwise specified. Unpaired two-tailed Student’s t-test or Wilcoxon rank-sum test was performed when comparing two categories. When more than two groups were compared, one-way ANOVA followed by a Dunnett’s multiple comparisons test or the Kruskal-Wallis nonparametric equivalent with Dunn’s multiple comparisons test were used. Results are mean (n ≥ 3) ± standard deviation (s.d.) unless otherwise stated.
Extended Data
Extended Data Figure 1. Linkage disequilibrium analysis of MAPT-AS1 region.
(a) SNPs within MAPT-AS1 genomic region that are linked (R2≥0.5) to tagging SNPs from the NHGRI GWAS catalog are reported. The specific trait associated to each tagging SNP together with the p-value from the GWAS study and their cited publications PubMed ID are shown. All p-values ≤5x10–8 were considered to be significant. Linkage disequilibrium (LD) correlations (R2) were calculated using LDlink1.170 for different populations. ASW: Americans of African Ancestry in SW USA; CEU: Utah Residents (CEPH) with Northern and Western European Ancestry; CHB: Han Chinese in Beijing, China; CHD: Chinese in Metropolitan Denver, Colorado; GIH: Gujarati Indians in Houston, Texas; JPT: Japanese in Tokyo, Japan; LWK: Luhya in Webuye, Kenya; MXL: Mexican ancestry in Los Angeles, California; MKK: Maasai in Kinyawa, Kenya; TSI: Toscani in Italy; YRI: Yoruba in Ibadan, Nigeria. (b) For each linked SNP listed in (a), the minor allele frequency (MAF) from the 1000 Genomes Project is given, together with the exon/intron location. (c) Pairwise linkage disequilibrium heatmap created using LDmatrix (https://ldlink.nci.nih.gov/?tab=ldmatrix). Red squares of increasing hue indicate increasing LD correlation between SNPs. A physical map of the genomic region is reported together with annotated RefSeq transcripts for each gene. (d) Enlarged view of the MAPT-AS1 3’-exon (in grey) containing the inverted MIRc element (in green), with two exonic linked SNPs downstream (rs17690326, rs17763596). (e) Detailed scheme of the H1/H2 inversion haplotypes (hg19). All major annotated genes in the linkage disequilibrium (LD) region are coloured in blue for the H1 haplotype, and in orange for the H2 inversion haplotype, with a white arrow representing their relative orientation. Arrays of Low Copy Repeats (LCRs), delimiting the inversion region, are represented by tandem arrows. MAPT-AS1 gene is coloured in yellow.
Extended Data Figure 2. Evolutionary conservation of t-NAT1 and -2 isoforms and MAPT-AS1 promoter region across primates.
(a-b) Scheme of human t-NAT1 and t-NAT2l transcript isoforms, exons (grey), with the region of overlap with MAPT (green) and the inverted MIR element in 3’-end (red). Multiple sequence alignment of the human t-NAT1 and t-NAT2l transcripts with the genomic sequences of 10 non-human primates (baboon, bonobo, chimp, gibbon, gorilla, marmoset, mouse lemur, orangutan, rhesus, squirrel monkey). Sequences were aligned using MUSCLE 3.856, and graphically displayed using Jalview 257. Pyrimidines in cyan and purines in magenta; splice junction is highlighted in yellow. A consensus sequence is at the base of multialignment with bar plot representing percentage sequence identity. (e, g) Phylogenetic trees associated to t-NAT1 and t-NAT2l multi-alignment represented in (a-b), obtained with the neighbour joining method using Jalview 2. Numbers reported on each connecting line in the tree represent Jaccard distances based on pairwise sequence similarity. (f, h) Negative PhyloCSF score58 (https://github.com/mlin/PhyloCSF/wiki) showing low protein-coding potential of t-NAT1 and t-NAT2l. The plots represent distribution of scores for each codon in each frame within each t-NAT isoform, across 29 mammals. (c-d) Multi-alignment showing sequence similarity between 3’-ends of human t-NAT1 (388-449) and t-NAT2l (510-554) and consensus MIR elements of different subfamilies (MIR3, MIR, MIRb, MIRc), as annotated by RepeatMasker. Homology regions of 62 and 45 nt respectively, are shared with the CORE-SINE, a 65 nt evolutionarily conserved domain at the centre of each MIR repeat element, schematically represented here and originally described by14. (i) Evolutionary conservation of MAPT-AS1 promoter region across 6 distant species (Homo sapiens, Macaca mulatta Mus musculus, Rattus norvegicus, Canis familiaris, Bos taurus), computed using the ECR browser59. Exons: yellow, introns: orange and repeat elements: green. Peaks represent percentage of identity to the human sequence. At bottom, CAGE and nanoCAGE60 tag clusters from FANTOM4 and FANTOM5 datasets retrieved from the ZENBU genome browser61, mapped to MAPT-AS1 promoter region, on sense (blue) or antisense strand (red). Values on the y-axis represent CAGE counts normalized per million tags (tpm).
Extended Data Figure 3. Expression of MAPT and MAPT-AS1 across brain regions and inverse correlation to tau pathology; levels and localization of endogenous MAPT mRNA is unaffected by stable expression of MAPT-AS1, whereas tau protein is increased by MAPT-AS1 with a flipped-MIR.
(a) RNA-Seq read counts from8, for MAPT mRNA and MAPT-AS1 lncRNA transcripts (t-NAT2s, t-NAT1, t-NAT2l) across 12 different regions of four independent human brains. Values represent mean counts ± s.d. CBRL, Cerebellum; FCTX, frontal cortex; HIPP, hippocampus; HYPO, hypothalamus; MEDU, medulla; OCTX, occipital cortex; PUTM, putamen; SNIG, substantia nigra; SPCO, spinal cord; TCTX temporal cortex; THAL, thalamus; WHMT white matter. (b) single-molecule RNA fluorescent in situ hybridization (smRNA-FISH) showing MAPT-AS1 (green) and MAPT (grey) transcripts expressed both in nucleus (DAPI, blue) and cytoplasm of SH-SY5Y neuroblastoma cells. Representative images of n=3 independent experiments. Scale bars represent 10 μm. (c) 2ddensity scatter plot of MAPT-AS1 and MAPT expression (FPKM) from post-mortem brains (Allen Brain Institute) coloured by Braak-stage. Red lines delimit middle points. Inset numbers represent samples. (d) Braak-stage distributions within upper (Q2+3), lower (Q1+4), left (Q1+2) or right (Q3+4) hemi-plot as in (c) are significantly different (two-sided unpaired Wilcoxon Rank-Sum test). (e) Cumulative proportion (y-axis) of phospho-tau immunohistochemistry (AT8-IHC, fraction of labelled pixels in ROI), phospho-tau to total-tau ratio (p-Tau/Tau ratio) and Aß42 to Aß40 ratio (aß42/aß40 ratio) (x-axis) for different Braak-stages (0-1, 2-4, 5-6). (f) Cumulative proportion (y-axis) of MAPT, MAPT-AS1 and KANSL1-AS1 gene expression levels (normalised FPKM, x-axis) for different Braak-stages (0-1, 2-4, 5-6). For data in (e-f) *P<0.05, ***P<0.001 two-sided Kolmogorov-Smirnov (KS) test, n=377 human post-mortem brains. RNA-seq, IHC and Illuminex-immunoassay data in this analysis are from the Allen Brain Institute’s Dementia, Ageing and Traumatic Brain Injury study (http://aging.brain-map.org/)9. (g) Normalized MAPT and MAPT-AS1 RNA expression levels (fold-changes) detected by qRT-PCR from SH-SY5Y cells stably expressing different deletion mutants of MAPT-AS1: t-NAT1 with flipped overlapping region (Flip), t-NAT1 with region not-overlapping with 5’UTR (Nover), t-NAT1 with overlapping region (Over), tNAT1 with deleted 5’-exon (t-NAT1Δ5’), tNAT1 with deleted 3’-exon (t-NAT1Δ3’), t-NAT2l with deleted 5’-exon (t-NAT2Δ5’), t-NAT2l with deleted 3’-exon (t-NAT2Δ3’). Values are normalized to cells stably transfected with an empty vector (Empty). Data represent independent SH-SY5Y clones stably expressing each construct (n=3 for Empty, n=4 for Flip, Nover and Over, mean ± s.d.; two-sided Kruskal-Wallis with Dunn’s multiple comparison test). (h) Both full-length (FL) and mutants with deleted MIR element (ΔM) of MAPT-AS1 localise to both cytosol and nucleus without altering the nucleo-cytoplasmic distribution of MAPT mRNA as detected by qRT-PCR. (data represent independent SH-SY5Y clones stably expressing each construct: n=3 Empty, n=3 t-NAT1-FL, n=6 t-NAT1-ΔM, n=3 t-NAT2-FL, n=6 t-NAT2-ΔM, mean ± s.d.; two-sided Kruskal-Wallis with Dunn’s multiple comparison test). (i) Quantitative expression of human MAPT-AS1 and MAPT transcripts measured by qRT-PCR (2–ΔΔCt) in sub-cellular fractions of SH-SY5Y cells, (n=3 independent experiments, mean ± s.d.). (j) Quantification of immunoblots probed with anti-tau and anti-β-actin antibodies. Protein lysates (20μg) from independent clones of SH-SY5Y cells stably expressing different MAPT-AS1 splice-isoforms, either full-length (t-NAT1-FL, t-NAT2-FL), with deleted MIR (t-NAT1-ΔM, t-NAT2-ΔM) or with a flipped MIR repeat (t-NAT1-Mflip). For each construct, total tau was normalized to β-actin levels quantified using ImageJ (n=6 independent stable clones, mean ± s.d.; one-way ANOVA with Dunnett’s test). As with the whole deletion of MIR (t-NAT1-ΔM), flipped MIR (t-NAT1-Mflip, delimited by red lines) increases tau protein.
Extended Data Figure 4. Characterization of human induced pluripotent stem cell-derived cortical and motor neurons.
(a) Control-1 (male) human iPSCs (hiPSCs) differentiated into cortical neurons using dual SMAD inhibition followed by specification of both deep- and upper-layer cortical excitatory neurons34. Neural rosettes at 20 days in vitro (DIV) express cortical progenitor markers PAX6 and OTX2, proliferation marker ki67 and neuronal marker TUJ1. By 100DIV, terminally differentiated neurons express βIII-tubulin, and later-born upper-layer neurons express SATB2 and BRN2. Scale bars=20μm, n=3 independent experiments. (b) Quantitative expression of MAPT and MAPT-AS1 (t-NAT1, t-NAT2s, t-NAT2l) in 3 independent inductions of hiPSC-derived cortical neurons (from 0 to 100DIV, one male healthy donor) measured by qRT-PCR (2–ΔΔCt/2–ΔΔCt max). (c) hiPSCs (control-1 and control-3), differentiated into motor neurons (MNs) using a previous established protocol71, were immunostained for NPC and MN markers and imaged by the Opera-Phenix (PerkinElmer). Images were acquired and quantified using Columbus v2.8.0.138890. NPCs at 18DIV express OLIG2 and NKX6.1, whereas 25DIV MNs express SMI32 and choline acetyltransferase (ChAT), bar graphs on the right (mean ± sem, n=23 (NKX6.1), n=27 (OLIG2), n=29 (SMI32), n=22 (ChAT) imaged wells across 3 different lines, scale bars: 20μm). (d) ICC images of MNs (26DIV), immunolabeled with the TUJ1, total-tau and DAPI after transduction with lentivirus (MOI 10), expressing shRNAs targeting either the exon-4 of MAPT-AS1 (shEx4) or Renilla luciferase ORF as a negative control (shRen) (mean ± s.d. n=3 for control-1 and control-2 iPSC-MNs, scale bars: 40μm). Relative tau levels normalised to TUJ1 measured as ratio of integrated densities is compared between the two groups as reported in bar graph on right (unpaired two-tailed t test). (e) Western blot of MNs (26-28DIV) from two healthy controls, transduced with LV-shRen (n=5) or LV-shEx4 (n=6), probed with anti-total-tau and anti-GAPDH antibodies. Quantification is shown on the right (mean ± s.d. *p<0.05, two-sided Wilcoxon-test).
Extended Data Figure 5. -MAPT-AS1 represses tau IRES-mediated translation in a MIR-dependent manner, with no effect on MAPT 3’-UTR and no major off-targets.
a. Reported secondary structure of MAPT 5’UTR (-242 to -1 relative to AUG)3. Domains 1 and 2 and 5’-TOP motif of tau-IRES are indicated and a blue line denotes overlap with t-NAT1 (5’-exon position 88-163). b, Relative abundance of MAPT-AS1, MAPT and β-actin mRNAs in polysomal fractions from cells stably expressing FL or ΔM MAPT-AS1 isoforms (mean±s.d.). Absorbance profiles (254 nm) are in background. c, Relative abundance of MAPT mRNA in fraction pools corresponding to 40-60S, 80S, light, medium or heavy polysomes. FL but not ΔM t-NAT1 or t-NAT2 significantly reduced MAPT mRNA association with heavy polysomes (n=3 Empty, n=4 t-NAT1FL, n=6 t-NAT1ΔM, n=3 t-NAT2FL, n=5 t-NAT2ΔM in b-c) (mean±s.e.m., one-way ANOVA with Holm-Sidak’s test; two points outside of axes in c). d pRTF or pRF construct with pcDNA3.1 empty vector, t-NAT1 full-length (FL) or with deleted MIR (t-NAT1-ΔM) were co-transfected into SH-SY5Y cells and relative luciferase levels measured after 48 hours. Significant reduction of tau-IRES activity (Fluc/Rluc ratio) was detected in cells expressing t-NAT1-FL, but not t-NAT1-ΔM, resulting in significant increase in MAPT IRES-mediated cap-independent translation. Similarly, t-NAT2l-FL repressed MAPT IRES activity, whereas t-NAT2l-ΔM with deleted MIR, had no such effect. Data in d represent mean± s.d., n=3 independent experiments (**P<0.01, *P<0.05, one-way ANOVA with Dunnett’s test). e. Schematic representation of luciferase constructs (pMIR-reporter) to study MAPT-AS1 effects on MAPT 3’-UTR following co-transfection in SH-SY5Y cells. Either the full-length (FL) or 3 partially overlapping fragments (Fr1, Fr2, Fr3) of MAPT 3’-UTR were cloned downstream to the Firefly luciferase ORF. (f, upper) firefly luciferase (Fluc) normalized to Renilla luciferase (Rluc) was quantified in SH-SY5Y cells co-transfected with either an empty pcDNA3.1 vector or different variants of t-NAT1 lncRNA (n = 3 independent experiments). (f, lower) Fluc/Rluc ratio was quantified in SH-SY5Y cells co-transfected with either empty vector or different variants of t-NAT2l lncRNA (n = 3 independent experiments). In all cases differences were not statistically significant except for t-NAT1 -Δ3’ (one-way ANOVA with Dunnet’s test). (g) Representative genome-wide metaplot of ribosome density over protein-coding mRNAs; a large majority of reads align as expected with 5’UTR and CDS, with a minority at 3’UTRs. RIBO-seq libraries were from 3 independent SH-SY5Y clones stably expressing each MAPT-AS1 variant or an empty vector (n=17). (h) Bar plot of the relative number of RIBO-seq reads with 5’-end in each reading frame, showing periodicity of ribosome footprints (RFPs) (n=17). (i) RIBO-seq volcano plot showing differentially translated genes in SH-SY5Y cells stably expressing full-length t-NAT1 (FL) compared to those with empty vector (Empty). Vertical red line in correspondence of MAPT (Log2FC=-1.45, p=0.036, Wald test with Bonferroni correction) shows that few other genes are similarly depleted of RFPs, with only 6 (gene symbols in grey) having at least 170 counts in all 17 libraries (a sample was excluded due to barcode cross-contamination with an unrelated CLIP library on the same sequencing run), but none with an adjusted significant p-value. (j) QuantSeq volcano plot showing differentially expressed genes in SH-SY5Y cells stably expressing full-length t-NAT1 (FL) compared to cells with empty vector (Empty). MAPT (red) mRNA levels not significantly different. Only genes with at least 1,000 read counts across 18 samples are named by their symbol (grey), although their adjusted p-values were not significant. Only three genes show a significant downregulation at the mRNA level (in blue, adjusted p-value <0.05), likely representing transcriptional off-targets. P-values in i-j were computed by DESeq2 using the Wald test with Bonferroni multiple comparison correction.
Extended Data Figure 6. Distribution of 7-mer MIR-complementary motifs along the human 18S rRNA secondary structure.
Human 18S ribosomal RNA secondary structure as retrieved from (http://apollo.chemistry.gatech.edu/RibosomeGallery/) is divided into an “active region” (red) and an “inactive region” (grey). As described24, active region is enriched for motifs able to mediate 40S ribosome recruitment through direct mRNA-rRNA interactions with 5’-UTRs of about 10% of human genes. Here, the 18S rRNA secondary structure is superimposed with 7-mers of complementary motifs (black dots) contained within each MIR embedded in MIR-NATs overlapping with 5’-UTRs of PC genes. Only 7-mers complementary to the 18S active region are shown. The 7-mer motifs represented here map to both the MIR elements within antisense MIR-NATs and the 5’-UTRs of the respective target genes, as reported in detail in Supplementary Table4. Matching positions of MIR motif-1 and -2 from MAPT-AS1 are reported (blue lines). 18S rRNA helices previously reported by Pisarev et al.72 to interact with mRNA regions upstream (yellow ovals) or downstream (salmon ovals) to the AUG start codon are indicated.
Extended Data Figure 7. Brain RNA-seq co-expression analysis. Genes paired with antisense MIR-NATs have significantly more structured 5’- and 3’-UTRs.
(a) Co-expression heatmaps representing distribution of RNA-seq read counts for 100 most abundant MIR-NAT target protein-coding genes (left panel) and 100 most abundant MIR-NAT genes (right panel), both hierarchically clustered based on their expression level in 12 different regions of 4 independent post-mortem brains from healthy human donors. Genes are clustered on y-axis. Brain regions on x-axis (CBRL, Cerebellum; FCTX, frontal cortex; HIPP, hippocampus; HYPO, hypothalamus; MEDU, medulla; OCTX, occipital cortex; PUTM, putamen; SNIG, substantia nigra; SPCO, spinal cord; TCTX temporal cortex; THAL, thalamus; WHMT, white matter). For each brain region, 4 independent brain samples are represented in each column. A colour key with histogram relative to each heatmap, have z-values associated to each color on the x-axis and RNA-seq counts on the y-axis. The histogram represents distribution of the RNA-seq counts for each z-value. (b) Similar co-expression heatmaps, as in (a), representing 1,045 MIR-NAT target protein-coding genes (on the left side) and 1,197 antisense MIR-NAT genes (on the right side). (c) Pie chart showing the percentage of MIR-NAT S-AS pairs annotated in GENCODE v19 and with 5’-UTR overlap, sorted by their Pearson’s correlation coefficient. The majority of S-AS pairs show positive correlations. (d) Histogram representing frequency of occurrence for 1,197 MIR-NAT S-AS pairs in bins of Pearson’s correlation (from -1 to + 1 in bins of 0.05). All MIR-NAT S-AS are visualized together, irrespective of their pattern of overlapping. MAPT-AS1-MAPT correlation coefficient is indicated. 3’-UTR (e) or 5’-UTR (f) minimum free energy (MFE), normalized by its length was computed using RNAfold 2.1.9 for each protein-coding gene in the human genome (hg19), and sorted based on their respective type of lncRNA overlap. Box plot presents median, upper and lower quartile boundaries for each group of protein-coding (PC) genes. PC genes pairing with MIR-NATs have both 3’-UTR and 5’-UTR significantly more structured than PC genes without lncRNA overlap (***, p < 0.0001 one-way ANOVA followed by Dunnett’s test). PC gene groups are as follows: PC genes overlapping antisense with MIR-NAT, ‘PC-MIRlncRNA’; PC genes overlapping with any lncRNA without embedded MIR repeat, ‘PC-lncRNA-NOMIR’; all PC genes with any overlapping lncRNA, ‘PC-lncRNA’; MIR-NATs, ‘MIRlncRNA’; PC genes without lncRNA overlap, ‘PC-NO-lncRNA’.
Extended Data Figure 8. MIR-NATs S-AS pairs within networks of interacting proteins, enriched for NDD-genes.
a, MIRs are more frequent in lncRNAs than mRNAs (5’UTR, 3’UTR, CDS). b, 1,197 GENCODE v19 MIR-NATs form S-AS pairs with 1,045 protein-coding (PC) genes: 40.69% overlap 5’UTR, 32.50% overlap CDS and 26.81% overlap 3’UTR. c, PC-genes with 5’UTR-overlapping MIR-NATs (n=630) are more expressed in human brain (log10FPKM) compared to genes with 3’UTR (n=392) or CDS (n=474) overlaps. Box plot: median with upper and lower quartiles; whiskers, values outside of interquartile range; points represent outliers (Welch two-sample t-test; one-way ANOVA across all gene-regions p=0.0214). d, Enriched cellular components and disease GO-terms ranked by Enrichr. 5’UTR-overlapping genes significantly associate with dementia (one-sided Fisher’s exact test p-values combined with z-scores, Supplementary Table 2b). e, MIR-NATs cognate PC-genes sorted by their overlap (3’UTR, 5’UTR, CDS) form networks of interacting proteins (coloured seeds), computed using PINOT29, and are associated with neurodegenerative diseases, enriched within 5’UTR network (p=1.5x10–4, 100,000 random simulations pnorm) f, PLCG1 and PLCG1-AS genes. g, Immunoblot quantification of SH-SY5Y cells stably expressing empty vector (Empty), full-length (FL) or MIR deleted (ΔM)-PLCG1-AS. PLCG1 is reduced in cells expressing FL but not ΔM-PLCG1-AS (n=6 clones stably expressing each construct, mean ± s.d., one-way ANOVA with Dunnett’s test).
Extended Data Figure 9. Majority of genes targeted by antisense MIR-NATs interact in a PPI network and are enriched for neurodegenerative disease-associated and immune system-associated genes.
(a) Protein-protein interaction (PPI)-network obtained from literature-curated interaction data from InnateDB database, using 392 seed proteins participating in S-AS pairs with MIR-NATs. Genes coding proteins associated with neurodegenerative diseases, represented as red-filled circles, are significantly enriched in network (p=1.63x10–8, Benjamini-Hochberg FDR using WebGestalt). Only primary interactions are represented in a zerodegree interaction network generated with NetworkAnalyst tool69. Self-interactions are not considered. (b) Schematic structures of representative genes pairing with antisense MIR-NATs and involved in different neurodegenerative diseases. GENCODE v19 annotated isoforms of the human SNCA, APP, MBNL1 and SLC1A2 genes and respective overlapping antisense MIR-NAT. MIR elements within each lncRNA are indicated (red). (c) Protein-protein interaction (PPI)-network obtained from literature-curated interaction data from InnateDB database, using 392 seed proteins participating in S-AS pairs with MIR-NATs. Genes encoding proteins associated with either the immune system (green) or innate immune system (blue), are significantly enriched into the network (respectively p=0.0041, p=0.0328, Benjamini-Hochberg FDR using NetworkAnalyst). Only primary interactions are represented in a zero-degree network generated using NetworkAnalyst tool69. Self-interactions are not considered. (d) Gene expression heatmap for 487 protein-coding genes with 5’-UTR overlapping with antisense MIR-NATs in 126 normal human tissues, from 557 publicly available microarray datasets, retrieved from the Enrichment Profiler Database (http://xavierlab2.mgh.harvard.edu/EnrichmentProfiler/index.html). Genes are clustered on y-axis and tissues are clustered on x-axis. Scale bar at bottom indicates colours associated to each z-score in the expression heatmap. (e) Scheme of the PLCG1 and PLCG1-AS genes is reported (hg19); the inverted MIRb is in red. Immunoblots of 6 independent SH-SY5Y clones stably expressing either empty vector (Empty), PLCG1-AS full-length (FL) or with whole inverted MIRb deleted (ΔM), probed with anti-PLCG1 and β-actin antibodies. (f) PLCG1 protein level is reduced in cells expressing FL- but not ΔM-PLCG1-AS as quantified in the graph (n=6 independent stable SH-SY5Y clones for each construct, mean± s.d., *p<0.05; one-way ANOVA with Dunnett’s test). (g) PLCG1 mRNA expression level from bulk RNA-seq of temporal cortex (TC) and prefrontal cortex (PFC) from the Mayo Clinic (n=160) and ROS-MAP (n=632) datasets respectively, is significantly increased in AD patients (AD) compared to asymptomatic AD (AsymAD) and healthy controls (Control), (box-plots: midpoints, medians; boxes, 25th and 75th percentiles; whiskers, minima and maxima; two-sided Wilcoxon-test) (data from http://swaruplab.bio.uci.edu:3838/bulkRNA/). Control samples were classified as Braak stage 0-I. Early-stage pathology samples were defined as Braak stage II-IV and CERAD score of possible AD, while late-stage pathology samples were Braak stage V-VI and CERAD score of probable and definite AD.
Extended Data Figure 10. 446 genes targeted by MIR-NATs contribute to the transcriptional signature of Alzheimer’s disease.
a, meta-analysis of snRNA-seq from Mathys (M), Grubman (G) and bulk RNA-seq from Friedman (GSE95587) datasets: rows are 446 MIR-NAT differentially expressed genes (DEG): 38 NDD-genes and 69 lncRNAs. DEGs across datasets partially overlap with 65 (27.7% up, 72.3% down) within Mathys, 160 (48.1% up, 51.9% down) within Grubman and 307 (58% up, 42% down) within Friedman datasets. Cell types: excitatory neurons (Ex), inhibitory neurons (In), neurons (Neu), astrocytes (Ast), oligodendrocytes (Olig), oligodendrocyte precursors (OPC), microglia (Mic), hybrid cells (Hyb), endothelial (Endo), unidentified cells (Unid). DEG counts are log2(mean gene expression in AD-pathology/mean gene expression in no-pathology) > 0.25 (two-sided Wilcoxon rank-sum test FDR<0.01 and Poisson mixed-model FDR<0.05, Mathys; two-sided Wilcoxon rank-sum test, FDR<0.05, Grubman and GSE95587). Annotations: gene-type (biotype), NDD-genes in DisGeNET database (disease), MIR orientation (MIR), S-AS region (overlap), percentage of protein IDRs by 75% of D2P2 predictors (disorder), number of protein-protein interactors (degree).
Extended Data Figure 11. Majority of genes targeted by MIR-NATs are enriched for interacting intrinsically disordered proteins (IDPs).
(a) Extended protein-protein interaction (PPI)-network from experimentally validated interaction data from various databases mined by PINOT29, using 760 nonredundant seed proteins participating in S-AS pairs with MIR-NATs. 399 seeds (40.3%) are genes encoding for IDPs with more than 90% IDRs, represented as red-filled circles, are significantly enriched into the network (p=0.0096, 100,000 random simulations in R, Bonferroni, details in Supplementary Table3). Only first-degree interactions are represented. Percentage of sequence predicted to span intrinsically disordered regions (IDRs) by at least 75% of the 9 algorithms from the D2P2 database30 is colour coded from blue (0-30%) to red (>90%) (b), 11 NDD-hub proteins in the above network are presented in this zoom-in view: (APP, ATP13A2, DCTN1, GABARAPL1, HSP90AA1, MAPT, MATR3, PLCG1, SNCA, SRRM2, VIM) (c), Topological properties of extended PPI network, computed by Cytoscape68.
Supplementary Material
Acknowledgements
We thank Dr. L. Wilson and Prof. A. Willis (University of Leicester, UK) for providing pRF and pRhcvF luciferase reporter vectors. We thank P. Fratta and A. Isaacs for suggestions and comments on the manuscript, and remaining members of the UK Brain Expression Consortium: S. Guelfi, K. D’Sa, M. Matarin, J. Vandrovcova, A. Ramasamy, J. A. Botia, C. Smith and P. Forabosco. This work was supported by the Reta Lila Weston Trust for Medical Research for funding to T.T.W. R.dS. and R.S.; CBD Solutions for funding to R.dS, R.S. and P.S.); the Medical Research Council (G0501560 to R.dS.), Parkinson’s UK (K1212 to R.dS.), PSP Association (R.dS.), CurePSP (R.dS.), Brain Research UK (R.dS.), Alzheimer’s Research UK to R.dS.; BBSRC LiDo PhD studentship to F.J.; AgeUK PhD Studentship to V.A.K.; the NIHR Queen Square Dementia BRU to S.W., E.P. and J.H.; the Italian Ministry of Education, University and Research “Futuro in Ricerca” (RBFR-0895DC) “Mechanisms of post-transcriptional regulation of gene expression in dementias”, to M.A.D.; University of Trento PhD studentship and an IBRO InEurope Short Stay grant to K.S.; and the MRC Sudden Death Brain Bank. This work was supported by the Francis Crick Institute which receives its core funding from Cancer Research UK (FC001002), the UK Medical Research Council (FC001002), and the Wellcome Trust (FC001002). This research was funded in part by the Wellcome Trust (4 Year Wellcome Trust Studentship to O.G.W.) and by the European Research Council under the European Union’s Seventh Framework Programme (617837-Translate) and under the European Union’s Horizon 2020 research and innovation programme (835300-RNPdynamics). This work was also supported by the UK Dementia Research Institute which receives its funding from DRI Ltd, funded by the UK Medical Research Council, Alzheimer’s Society and Alzheimer’s Research UK; Medical Research Council (award number MR/N026004/1 to J.H.), Wellcome Trust (award number 202903/Z/16/Z to J.H.), Dolby Family Fund to J.H., National Institute for Health Research University College London Hospitals Biomedical Research Centre funding to J.H.
Footnotes
Author Contributions
R.S., and R.dS. conceived and designed the project with contributions from J.U.; R.S., F.J., O.G.W., M.E., P.Z., A.M., G.H., V.A.K., G.V., F.L.A., J.Z.P. and R.dS. performed experiments; D.T. and M.R. contributed to provide brain RNA-seq data; R.S. and O.W. generated and analysed RIBO-seq and QuantSeq data. S.W. and E.P. contributed iPSC-derived cortical neurons; J.S.M., J.H and R.P. contributed iPSC-derived motor neurons; J.Z.P. performed all mice AAV-injections and F.LA. processed and analysed in vivo AAV-transduced brain samples; D.K. and A.P. analysed SNP data from PDGene dataset and estimated association to Parkinson’s disease; R.F. and C.M. contributed to PPI network analysis; R.S, W.E., O.G.W. and N.B.T. performed bioinformatics; R.S., F.J., W.E., O.G.W., J.U. and R.dS. analysed data and interpreted results with contributions from M.A.D., and K.S.; R.S. wrote the manuscript with contributions from W.E., O.G.W., J.U. and R.dS.
Competing Interests
The authors (R.S. and R.dS.) declare the following competing interest: Patent WO2017199041A1
Data availability
The RIBO-seq and QuantSeq data generated as part of this work can be retrieved from ArrayExpress (E-MTAB-9921). For correlation analysis of MAPT-AS1 FPKM levels and tau Braak-stages, RNA-seq and Luminex data and clinical metadata from the Allen Brain Institute can be retrieved at (http://aging.brain-map.org/)9, whereas bulk RNA-seq and clinical metadata from the ROS-MAP project10 can be retrieved on Synapse at https://www.synapse.org/#!Synapse:syn3388564 and https://www.synapse.org/#!Synapse:syn3219045 respectively); access was granted under the signed MTA-20-011. Bulk RNA-seq from twelve brain regions of four healthy subjects published by Sibley et al.8 source data can be retrieved from ArrayExpress (E-MTAB-3534). For Alzheimer’s disease meta-transcriptomic analysis, the following publicly deposited data were used: snRNA-seq data published by Mathys et al.26 can be accessed at (https://www.radc.rush.edu/docs/omics.htm or at Synapse https://www.synapse.org/#!Synapse:syn18485175 under the doi 10.7303/syn18485175). snRNA-seq data released by Grubman et al.27 can be accessed here (http://adsn.ddnetbio.com/) and are also available through Gene Expression Omnibus (GSE138852). Bulk RNA-seq published by Friedman et al. 28 has been deposited at Gene Expression Omnibus under the following identifier (GSE95587).
Gene annotations were retrieved from GENCODE v19 (https://www.gencodegenes.org). MIR repeats chromosomal coordinates were retrieved using RepeatMasker v4.0.5 (www.repeatmasker.org). For the PPI networks, we used publicly accessible interactions data retrieved using either PINOT v1.1 (http://www.reading.ac.uk/bioinf/PINOT/PINOT_form.html) or NetworkAnalyzer v3.0 (https://www.networkanalyst.ca/) and InnateDB (https://www.innatedb.com). For protein disorder predictions we used the D2P2 database (Ensembl v63) (http://d2p2.pro). For TSS mapping CAGE/nanoCAGE data were retrieved from ZENBU Genome Browser v2.11 (https://fantom.gsc.riken.jp/zenbu/). NDD annotations for enrichment analysis were retrieved from DisGeNET database v6.0 (https://www.disgenet.org). Genetic SNP data analysis was performed using data retrieved from PDGene database (2016) (http://www.pdgene.org) and linkage disequilibrium analysis around MAPT-AS1 locus was performed using LDmatrix (https://ldlink.nci.nih.gov/?tab=ldmatrix). GO-terms enrichment using Enrichr (Sept-2014) (https://amp.pharm.mssm.edu/Enrichr/). PLCG1 gene expression data in AD were also retrieved from (http://swaruplab.bio.uci.edu:3838/bulkRNA/). Source data are provided with this manuscript.
Code availability
Customised code used throughout this study can be found at (https://github.com/robertosimone-ucl/scripts_RIBOseq_QuantSeq) and (https://github.com/robertosimone-ucl/scripts_DEG_in_AD). The code used by PINOT can be found here (https://www.reading.ac.uk/bioinf/downloads/PINOT_scripts/). Further details are available upon reasonable request from the corresponding authors. In all other cases software tools used for specific analyses are reported and cited in the Methods.
References
- 1.Pelechano V, Steinmetz LM. Gene regulation by antisense transcription. Nat Rev Genet. 2013;14:880–893. doi: 10.1038/nrg3594. [DOI] [PubMed] [Google Scholar]
- 2.Statello L, Guo C-J, Chen L-L, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118. doi: 10.1038/s41580-020-00315-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Veo BL, Krushel LA. Secondary RNA structure and nucleotide specificity contribute to internal initiation mediated by the human tau 5’ leader. RNA Biol. 2012;9:1344–1360. doi: 10.4161/rna.22181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Spillantini MG, Goedert M. Tau pathology and neurodegeneration. Lancet Neurol. 2013;12:609–622. doi: 10.1016/S1474-4422(13)70090-5. [DOI] [PubMed] [Google Scholar]
- 5.Pittman AM, et al. Linkage disequilibrium fine mapping and haplotype association analysis of the tau gene in progressive supranuclear palsy and corticobasal degeneration. J Med Genet. 2005;42:837–846. doi: 10.1136/jmg.2005.031377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Gsponer J, Futschik ME, Teichmann SA, Babu MM. Tight regulation of unstructured proteins: from transcript synthesis to protein degradation. Science. 2008;322:1365–1368. doi: 10.1126/science.1163581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zucchelli S, et al. Antisense Transcription in Loci Associated to Hereditary Neurodegenerative Diseases. Mol Neurobiol. 2019 doi: 10.1007/s12035-018-1465-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sibley CR, et al. Recursive splicing in long vertebrate genes. Nature. 2015;521:371–375. doi: 10.1038/nature14466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Miller JA, et al. Neuropathological and transcriptomic characteristics of the aged brain. Elife. 2017;6 doi: 10.7554/eLife.31126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bennett DA, et al. Religious Orders Study and Rush Memory and Aging Project. J Alzheimers Dis. 2018;64:S161–S189. doi: 10.3233/JAD-179939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Coupland KG, et al. Role of the Long Non-Coding RNA MAPT-AS1 in Regulation of Microtubule Associated Protein Tau (MAPT) Expression in Parkinson’s Disease. PLoS ONE. 2016;11:e0157924. doi: 10.1371/journal.pone.0157924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Elkouris M, et al. Long Non-coding RNAs Associated With Neurodegeneration-Linked Genes Are Reduced in Parkinson’s Disease Patients. Front Cell Neurosci. 2019;13:58. doi: 10.3389/fncel.2019.00058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Smit AF, Riggs AD. MIRs are classic, tRNA-derived SINEs that amplified before the mammalian radiation. Nucleic Acids Res. 1995;23:98–102. doi: 10.1093/nar/23.1.98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gilbert N, Labuda D. CORE-SINEs: eukaryotic short interspersed retroposing elements with common sequence motifs. Proc Natl Acad Sci USA. 1999;96:2869–2874. doi: 10.1073/pnas.96.6.2869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 16.Morita T, Sobue K. Specification of neuronal polarity regulated by local translation of CRMP2 and Tau via the mTOR-p70S6K pathway. J Biol Chem. 2009;284:27734–27745. doi: 10.1074/jbc.M109.008177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bottley A, Phillips NM, Webb TE, Willis AE, Spriggs KA. eIF4A inhibition allows translational regulation of mRNAs encoding proteins involved in Alzheimer’s disease. PLoS ONE. 2010;5 doi: 10.1371/journal.pone.0013030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mauro VP, Edelman GM. The ribosome filter hypothesis. Proc Natl Acad Sci USA. 2002;99:12031–12036. doi: 10.1073/pnas.192442499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Andorfer C, et al. Hyperphosphorylation and aggregation of tau in mice expressing normal human tau isoforms. J Neurochem. 2003;86:582–590. doi: 10.1046/j.1471-4159.2003.01879.x. [DOI] [PubMed] [Google Scholar]
- 20.Nalls MA, et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat Genet. 2014;46:989–993. doi: 10.1038/ng.3043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hon C-C, et al. An atlas of human long non-coding RNAs with accurate 5’ ends. Nature. 2017 doi: 10.1038/nature21374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kapusta A, et al. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet. 2013;9:e1003470. doi: 10.1371/journal.pgen.1003470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Holcik M. Internal Ribosome Entry Site-Mediated Translation in Neuronal Protein Synthesis. In: Sossin WS, editor. The Oxford Handbook of Neuronal Protein Synthesis. Oxford University Press; 2018. [DOI] [Google Scholar]
- 24.Weingarten-Gabbay S, et al. Comparative genetics. Systematic discovery of cap-independent translation sequences in human and viral genomes. Science. 2016;351 doi: 10.1126/science.aad4939. [DOI] [PubMed] [Google Scholar]
- 25.Paek KY, et al. Translation initiation mediated by RNA looping. Proc Natl Acad Sci U S A. 2015;112:1041–1046. doi: 10.1073/pnas.1416883112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mathys H, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature. 2019;570:332–337. doi: 10.1038/s41586-019-1195-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Grubman A, et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci. 2019;22:2087–2097. doi: 10.1038/s41593-019-0539-4. [DOI] [PubMed] [Google Scholar]
- 28.Friedman BA, et al. Diverse Brain Myeloid Expression Profiles Reveal Distinct Microglial Activation States and Aspects of Alzheimer’s Disease Not Evident in Mouse Models. Cell Rep. 2018;22:832–847. doi: 10.1016/j.celrep.2017.12.066. [DOI] [PubMed] [Google Scholar]
- 29.Tomkins JE, et al. PINOT: an intuitive resource for integrating protein-protein interactions. Cell Commun Signal. 2020;18:92. doi: 10.1186/s12964-020-00554-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Oates ME, et al. D2P2 database of disordered protein predictions. Nucleic Acids Res. 2013;41:D508–516. doi: 10.1093/nar/gks1226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ciryam P, Tartaglia GG, Morimoto RI, Dobson CM, Vendruscolo M. Widespread aggregation and neurodegenerative diseases are associated with supersaturated proteins. Cell Rep. 2013;5:781–790. doi: 10.1016/j.celrep.2013.09.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Edwards YJK, Lobley AE, Pentony MM, Jones DT. Insights into the regulation of intrinsically disordered proteins in the human proteome by analyzing sequence and gene expression data. Genome Biol. 2009;10:R50. doi: 10.1186/gb-2009-10-5-r50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sposito T, et al. Developmental regulation of tau splicing is disrupted in stem cell-derived neurons from frontotemporal dementia patients with the 10 + 16 splice-site mutation in MAPT. Hum Mol Genet. 2015;24:5260–5269. doi: 10.1093/hmg/ddv246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Shi Y, Kirwan P, Livesey FJ. Directed differentiation of human pluripotent stem cells to cerebral cortex neurons and neural networks. Nat Protoc. 2012;7:1836–1846. doi: 10.1038/nprot.2012.116. [DOI] [PubMed] [Google Scholar]
- 35.Hall CE, et al. Progressive Motor Neuron Pathology and the Role of Astrocytes in a Human Stem Cell Model of VCP-Related ALS. Cell Rep. 2017;19:1739–1749. doi: 10.1016/j.celrep.2017.05.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.De Palma M, Naldini L. Transduction of a gene expression cassette using advanced generation lentiviral vectors. Meth Enzymol. 2002;346:514–529. doi: 10.1016/s0076-6879(02)46074-0. [DOI] [PubMed] [Google Scholar]
- 37.Kutner RH, Zhang X-Y, Reiser J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat Protoc. 2009;4:495–505. doi: 10.1038/nprot.2009.22. [DOI] [PubMed] [Google Scholar]
- 38.Paxinos G, F K. The mouse brain in stereotaxic coordinates. Academic; 2004. [Google Scholar]
- 39.Kopec AM, Rivera PD, Lacagnina MJ, Hanamsagar R, Bilbo SD. Optimized solubilization of TRIzol-precipitated protein permits Western blotting analysis to maximize data available from brain tissue. J Neurosci Methods. 2017;280:64–76. doi: 10.1016/j.jneumeth.2017.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Schindelin J, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–682. doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Potter CJ, Luo L. Splinkerette PCR for mapping transposable elements in Drosophila. PLoS ONE. 2010;5:e10168. doi: 10.1371/journal.pone.0010168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Trabzuni D, et al. Quality control parameters on a large dataset of regionally dissected human control brains for whole genome expression studies. J Neurochem. 2011;119:275–282. doi: 10.1111/j.1471-4159.2011.07432.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Stoneley M, Paulin FE, Le Quesne JP, Chappell SA, Willis AE. C-Myc 5’ untranslated region contains an internal ribosome entry segment. Oncogene. 1998;16:423–428. doi: 10.1038/sj.onc.1201763. [DOI] [PubMed] [Google Scholar]
- 44.Kraushar ML, et al. Temporally defined neocortical translation and polysome assembly are determined by the RNA-binding protein Hu antigen R. Proc Natl Acad Sci USA. 2014;111:E3815–3824. doi: 10.1073/pnas.1408305111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.McGlincy NJ, Ingolia NT. Transcriptome-wide measurement of translation by ribosome profiling. Methods. 2017;126:112–129. doi: 10.1016/j.ymeth.2017.05.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Adiconis X, et al. Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat Methods. 2013;10:623–629. doi: 10.1038/nmeth.2483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Blazquez L, et al. Exon Junction Complex Shapes the Transcriptome by Repressing Recursive Splicing. Mol Cell. 2018;72:496–509.:e9. doi: 10.1016/j.molcel.2018.09.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet j. 2011;17:10. [Google Scholar]
- 49.Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–359. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dobin A, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Smith T, Heger A, Sudbery I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 2017;27:491–499. doi: 10.1101/gr.209601.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Moll P, Ante M, Seitz A, Reda T. QuantSeq 3’ mRNA sequencing for RNA quantification. Nat Methods. 2014;11 i-iii. [Google Scholar]
- 54.Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–842. doi: 10.1093/bioinformatics/btq033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wickham H. Ggplot2: elegant graphics for data analysis. Springer; 2009. [Google Scholar]
- 56.Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ. Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics. 2009;25:1189–1191. doi: 10.1093/bioinformatics/btp033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Lin MF, Jungreis I, Kellis M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics. 2011;27:i275–282. doi: 10.1093/bioinformatics/btr209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Ovcharenko I, Nobrega MA, Loots GG, Stubbs L. ECR Browser: a tool for visualizing and accessing data from comparisons of multiple vertebrate genomes. Nucleic Acids Res. 2004;32:W280–286. doi: 10.1093/nar/gkh355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Plessy C, et al. Linking promoters to functional transcripts in small samples with nanoCAGE and CAGEscan. Nat Methods. 2010;7:528–534. doi: 10.1038/nmeth.1470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Lizio M, et al. Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol. 2015;16:22. doi: 10.1186/s13059-014-0560-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Lorenz R, et al. ViennaRNA Package 2.0. Algorithms Mol Biol. 2011;6:26. doi: 10.1186/1748-7188-6-26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ritchie ME, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Law CW, Chen Y, Shi W, Smyth GK. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15:R29. doi: 10.1186/gb-2014-15-2-r29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–2849. doi: 10.1093/bioinformatics/btw313. [DOI] [PubMed] [Google Scholar]
- 66.Chen EY, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics. 2013;14:128. doi: 10.1186/1471-2105-14-128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wang J, Duncan D, Shi Z, Zhang B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 2013;41:W77–83. doi: 10.1093/nar/gkt439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Shannon P, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Xia J, Benner MJ, Hancock REW. NetworkAnalyst--integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res. 2014;42:W167–174. doi: 10.1093/nar/gku443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Machiela MJ, Chanock SJ. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics. 2015;31:3555–3557. doi: 10.1093/bioinformatics/btv402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Luisier R, et al. Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS. Nat Commun. 2018;9 doi: 10.1038/s41467-018-04373-8. 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Pisarev AV, Kolupaeva VG, Yusupov MM, Hellen CUT, Pestova TV. Ribosomal position and contacts of mRNA in eukaryotic translation initiation complexes. EMBO J. 2008;27:1609–1621. doi: 10.1038/emboj.2008.90. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The RIBO-seq and QuantSeq data generated as part of this work can be retrieved from ArrayExpress (E-MTAB-9921). For correlation analysis of MAPT-AS1 FPKM levels and tau Braak-stages, RNA-seq and Luminex data and clinical metadata from the Allen Brain Institute can be retrieved at (http://aging.brain-map.org/)9, whereas bulk RNA-seq and clinical metadata from the ROS-MAP project10 can be retrieved on Synapse at https://www.synapse.org/#!Synapse:syn3388564 and https://www.synapse.org/#!Synapse:syn3219045 respectively); access was granted under the signed MTA-20-011. Bulk RNA-seq from twelve brain regions of four healthy subjects published by Sibley et al.8 source data can be retrieved from ArrayExpress (E-MTAB-3534). For Alzheimer’s disease meta-transcriptomic analysis, the following publicly deposited data were used: snRNA-seq data published by Mathys et al.26 can be accessed at (https://www.radc.rush.edu/docs/omics.htm or at Synapse https://www.synapse.org/#!Synapse:syn18485175 under the doi 10.7303/syn18485175). snRNA-seq data released by Grubman et al.27 can be accessed here (http://adsn.ddnetbio.com/) and are also available through Gene Expression Omnibus (GSE138852). Bulk RNA-seq published by Friedman et al. 28 has been deposited at Gene Expression Omnibus under the following identifier (GSE95587).
Gene annotations were retrieved from GENCODE v19 (https://www.gencodegenes.org). MIR repeats chromosomal coordinates were retrieved using RepeatMasker v4.0.5 (www.repeatmasker.org). For the PPI networks, we used publicly accessible interactions data retrieved using either PINOT v1.1 (http://www.reading.ac.uk/bioinf/PINOT/PINOT_form.html) or NetworkAnalyzer v3.0 (https://www.networkanalyst.ca/) and InnateDB (https://www.innatedb.com). For protein disorder predictions we used the D2P2 database (Ensembl v63) (http://d2p2.pro). For TSS mapping CAGE/nanoCAGE data were retrieved from ZENBU Genome Browser v2.11 (https://fantom.gsc.riken.jp/zenbu/). NDD annotations for enrichment analysis were retrieved from DisGeNET database v6.0 (https://www.disgenet.org). Genetic SNP data analysis was performed using data retrieved from PDGene database (2016) (http://www.pdgene.org) and linkage disequilibrium analysis around MAPT-AS1 locus was performed using LDmatrix (https://ldlink.nci.nih.gov/?tab=ldmatrix). GO-terms enrichment using Enrichr (Sept-2014) (https://amp.pharm.mssm.edu/Enrichr/). PLCG1 gene expression data in AD were also retrieved from (http://swaruplab.bio.uci.edu:3838/bulkRNA/). Source data are provided with this manuscript.
Customised code used throughout this study can be found at (https://github.com/robertosimone-ucl/scripts_RIBOseq_QuantSeq) and (https://github.com/robertosimone-ucl/scripts_DEG_in_AD). The code used by PINOT can be found here (https://www.reading.ac.uk/bioinf/downloads/PINOT_scripts/). Further details are available upon reasonable request from the corresponding authors. In all other cases software tools used for specific analyses are reported and cited in the Methods.