Abstract
Higher-order chromatin structure is tightly linked to gene expression and therefore cell identity. In recent years, the chromatin landscape of pluripotent stem cells has become better characterized, and unique features at various architectural levels have been revealed. However, the mechanisms that govern establishment and maintenance of these topological characteristics and the temporal and functional relationships with transcriptional or epigenetic features are still areas of intense study. Here, we will discuss progress and limitations of our current understanding regarding how the 3D chromatin topology of pluripotent stem cells is established during somatic cell reprogramming and maintained during cell division. We will also discuss evidence and theories about the driving forces of topological reorganization and the functional links with key features and properties of pluripotent stem cell identity.
Keywords: 3D chromatin organization, iPSC, ESC, reprogramming, mitosis, bookmarking, transcription factors, enhancer-promoter interaction
In this review article, Apostolou and colleagues discuss how the unique 3D chromatin architecture of pluripotent stem cells is established during somatic cell reprogramming and maintained during self-renewal. They analyze progress and gaps in our knowledge regarding the driving forces of 3D chromatin reorganization, the links to transcriptional changes and the impact on cell fate decisions.
Main Text
Principles and Mechanisms of 3D Chromatin Organization
The three-dimensional (3D) organization of the genome has been shown to have major implications for gene regulation and therefore cell identity. Over the last three decades, technological advancements in microscopy and genome-wide chromatin topology assays (Kempfer and Pombo, 2020) uncovered a hierarchical 3D genomic organization ranging from whole chromosome territories to (sub)megabase structures and fine-scale chromatin loops (Gibcus and Dekker, 2013; Yu and Ren, 2017). Individual chromosomes reside in discrete regions of the nucleus called chromosome territories (Cremer and Cremer, 2001). Within each chromosome, euchromatic and heterochromatic regions segregate into different subnuclear compartments named A and B, respectively (Lieberman-Aiden et al., 2009; Wang et al., 2016). The A compartments are gene rich, accessible, transcriptionally active, and largely occupy the nuclear interior, while B compartments are gene poor, inactive, and inaccessible regions, located at the nuclear periphery and overlapping with lamina-associated domains (LADs) (Guelen et al., 2008; Haarhuis et al., 2017). Due to their association with transcriptional activity, compartments are distinct across cell types (Dixon et al., 2012, 2015; Takebayashi et al., 2012). At (sub)megabase scale, we detect self-associating domains, known as topologically associating domains (TADs) (Dixon et al., 2016; Nora et al., 2017; Sexton et al., 2012) or contact domains (Tang et al., 2015). These are found in both A and B compartments and are demarcated by insulating boundaries, which prevent interactions with neighboring TADs. While TADs are largely conserved among cell types, the insulation strength of TAD boundaries and patterns of intra-TAD interactions can vary (Crane et al., 2015). Sub-TADs, or smaller insulated neighborhoods nested inside larger ones, have also been described (Phillips-Cremins et al., 2013; Rao et al., 2014). Although current nomenclature in the field is rather confusing (Dixon et al., 2016; Schoenfelder and Fraser, 2019), studies have provided strong evidence for the role of TADs and insulated neighborhoods in gene expression by restricting enhancer activity and specificity (Dowen et al., 2014; Lupiáñez et al., 2015; Sun et al., 2019). Finally, at the finest level of architectural organization there are pairwise chromatin contacts between distal genomic loci, ranging from few kilobase pairs (kb) to several megabases (Mb) of linear distance. Different types of long-range genomic interactions have been identified based on the mediating protein factors and their potential impact on gene expression (Cheutin and Cavalli, 2019; Schoenfelder and Fraser, 2019; van Steensel and Furlong, 2019). These include active contacts between cis-regulatory elements (enhancer-promoter), Polycomb-mediated repressive/poising interactions (Cheutin and Cavalli, 2019; Denholtz et al., 2013; Eagen et al., 2017; Schoenfelder et al., 2015b), and CTCF/Cohesin structural loops (Handoko et al., 2011; Robson et al., 2019). Although structural loops are largely conserved among cell types, activating and repressive/poised contacts can be reorganized during cell fate transitions (Penalosa-Ruiz et al., 2019).
Two main forces are thought to work together and against each other to shape the layers of the 3D genome: loop extrusion and compartmental segregation, as extensively reviewed recently (Beagan and Phillips-Cremins, 2020; Nuebler et al., 2018; Rada-Iglesias et al., 2018; Rowley et al., 2017). In loop extrusion, the loaded Cohesin complex keeps sliding/extruding DNA through its ring until it encounters convergently oriented CTCF sites (Fudenberg et al., 2016; Sanborn et al., 2015). Loop extrusion is thus responsible for the formation of CTCF/Cohesin-mediated loops, TADs, and sub-TADs (Beagan and Phillips-Cremins, 2020), as well as the recently described “stripes” or “flames” (Fudenberg et al., 2016; Hsieh et al., 2020; Rao et al., 2014; Vian et al., 2018). On the other hand, A and B compartmentalization occurs through increased “self-attraction”/affinity among chromatin loci with similar transcriptional and chromatin states, such as homotypic histone modifications and transcription factors (TFs)/cofactors. Liquid/liquid-phase separation is one mechanism of chromatin self-organization/segregation (Palikyras and Papantonis, 2019), and it results in the formation of subnuclear droplet-like condensates or membraneless organelles, such as nucleoli or nuclear speckles. A large number of TFs (Boija et al., 2018; Chong et al., 2018), epigenetic modulators (BRD4, Polycomb Repressive Complex 2 [PRC2]) (Sabari et al., 2018; Tatavosian et al., 2019), and transcriptional cofactors (Mediator, Pol II) (Cho et al., 2018; Zamudio et al., 2019) have been reported to form large nuclear condensates, which may mediate activating or repressive chromatin contacts in a loop-extrusion-independent manner. There is increasing evidence for the distinct but interconnected nature of these two guiding mechanisms, as knock-down or degradation of CTCF or Cohesin subunits disrupts TADs, stripes, and structural chromatin loops but has minimal effect on transcription and even reinforces compartmentalization (Haarhuis et al., 2017; Nora et al., 2017; Rao et al., 2017; Schwarzer et al., 2017; Wutz et al., 2017). Importantly, the expression levels and/or binding patterns of these critical factors (CTCF, Cohesin, TFs, cofactors, histone modifications) is often distinct across cell types and may thus be responsible for driving cell-type-specific chromatin architecture.
3D Chromatin (Re)Organization during Acquisition and Maintenance of Pluripotency
Pluripotent stem cells (PSCs) are endowed with the remarkable capacity to self-renew indefinitely in culture (self-renewal) while preserving their potential to differentiate into all somatic cell types upon proper stimulation in vitro or upon blastocyst injection in vivo (pluripotency) (Evans, 2011; Tabar and Studer, 2014). PSCs encompass both embryonic stem cells (ESCs), derived from early embryos, and induced PSCs (iPSCs), generated through somatic cell reprogramming (Takahashi and Yamanaka, 2006). The cellular properties of PSCs are supported by their unique epigenetic landscape, characterized by overall low levels of DNA methylation and heterochromatin marks and a prevalence of Polycomb-dependent repressive (H3K27me3) or bivalent (H3K4me3/H3K27me3) chromatin around lineage-related genes (extensively reviewed; see Di Giammartino and Apostolou, 2016; Harikumar and Meshorer, 2015; Papp and Plath, 2013). This “open” and permissive chromatin environment becomes progressively restrained during differentiation as potency decreases (Sexton and Cavalli, 2015), and must be reset during reprogramming. The 3D genomic organization of PSCs is also characterized by distinct architectural features, such as the prevalence of A versus B compartments and the presence of PRC1/2-driven long-range interactions (Denholtz et al., 2013; Joshi et al., 2015; Schoenfelder et al., 2015b). Importantly, the 3D chromatin features of PSCs have largely been characterized in established, asynchronously cycling PSCs and often in comparison with other steady-state somatic cell types. In the following sections we will discuss recent work that sheds light on the dynamic nature of these features and particularly on how PSC-specific 3D chromatin landscape is established upon acquisition of pluripotency and propagated during cell division (Figure 1). We will particularly focus on findings from mouse naive PSCs that represent a pre-implantation pluripotent state, unless otherwise specified. Our goal is to highlight key factors and forces involved in building and maintaining PSC fate through reprogramming and self-renewal, respectively.
Figure 1.
Reorganization of the 3D Genome during Self-Renewal and Cell Fate Change
Schematic illustration of the 3D genomic reorganization that occurs during the cell cycle in self-renewing cells as well as during cell fate changes, such as differentiation and reprogramming. Examples of Hi-C heatmaps at 250-kb resolution are shown to illustrate the topological rewiring between cell types and during mitosis.
3D Chromatin Reorganization upon Acquisition of Pluripotency during Somatic Cell Reprogramming
Since the first derivation of iPSCs by Takahashi and Yamanaka (2006), cellular reprogramming has proved to be a powerful system to study mechanisms that dictate cell fate changes (Takahashi and Yamanaka, 2006). Ectopic expression of four TFs (OCT4, SOX2, KLF4, and c-MYC) can convert somatic cells into iPSCs with the genetic and epigenetic characteristics of stem cells, capable of self-renewing and differentiating into all germ layers in vivo. As somatic cells reprogram, they undergo cellular, transcriptional, metabolic, and epigenetic changes that result in erasure of the somatic program and establishment of pluripotency-defining properties (Apostolou and Hochedlinger, 2013; Brumbaugh et al., 2019; Papp and Plath, 2013). 3D chromatin architecture is also drastically reorganized during reprogramming, as shown by comparing various somatic cells with the resultant iPSCs, through profiling long-range interactions around specific loci (4C-seq or 5C) (Apostolou et al., 2013; Beagan et al., 2016; Denholtz et al., 2013; Wei et al., 2013) or in a genome-wide fashion by Hi-C (Krijger et al., 2016). Although most of these studies examined 3D chromatin architecture between start and endpoint fates, our review will focus on recent work that has begun to characterize the dynamic topological reorganization during reprogramming as well as underlying mechanisms and temporal associations with transcriptional and epigenetic changes.
3D Chromatin Architecture Is Drastically Reorganized during Somatic Cell Reprogramming along with Epigenetic and Transcriptional Changes
Compartment Switching
Reprogramming of multiple different mouse somatic tissue types (pre-B cells, macrophages, neural stem cells, mouse embryonic fibroblasts (MEFs)) resulted in iPSCs with 99% compartment similarity to one another and to naive ESCs (Krijger et al., 2016). This highlights the tight link between 3D organization and cell identity, and demonstrates that reprogramming promotes a rather faithful re-establishment of the pluripotency-associated chromatin architecture. To understand the kinetics of chromatin changes during reprogramming, a study from the Graf group characterized reprogramming intermediates at a genome-wide scale. They captured the dynamic topological reorganization that mouse B cells undergo during reprogramming to iPSCs using a highly efficient and synchronized system (Stadhouders et al., 2018). They discovered that compartmentalization changes occur gradually during reprogramming, with up to 20% of the genome switching compartments between any two intermediate time points. Surprisingly, many loci experienced multiple compartment switches throughout the time course, suggesting the emergence of transient, intermediate topologies and states. Whether these represent byproducts or critical steps of the reprogramming process remains unknown. Importantly, compartmentalization changes are associated with, and often occurred prior to, robust transcriptional changes. About 30%–40% of genes related to either B-cell or PSC identities underwent compartment switches during the course of reprogramming. Interestingly, pluripotency genes that were initially residing in A compartments were upregulated early during reprogramming, in parallel with the silencing of somatic genes, while those that switched from B to A (e.g., Dppa3) were activated at later stages. The delayed activation of genes embedded into repressive B compartments likely reflects the requirement for extensive epigenetic and topological remodeling prior to transcriptional upregulation. In support of this, integrative analysis with chromatin immunoprecipitation sequencing (ChIP-seq) data for the active histone mark H3K4me2 revealed that epigenetic remodeling occurs either concomitant with or prior to compartment switches, indicating that histone marks and/or the associated epigenetic modulators play active roles in large-scale architectural reorganization.
TAD Reorganization
In concordance with the reported conservation of TAD boundaries across cell types (Dixon et al., 2012, 2015), the vast majority of TAD boundaries remained constant throughout B-cell reprogramming. Genes found near the few lost or gained boundaries experienced variable effects on expression, with no clear correlation between gain/loss and upregulation/downregulation. Despite minimal qualitative changes, 50%–80% of TADs experienced quantitative changes either in the insulation strength of their boundaries (frequency of interactions with neighboring TADs) or their self-connectivity (number of interactions within TAD). Interestingly, changes in TAD boundary insulation occurred early in reprogramming, broadly preceding epigenetic and compartmental reorganization. In agreement with reports in other cellular systems and during differentiation (Bonev et al., 2017; Crane et al., 2015; Van Bortle et al., 2014), insulation changes during reprogramming associated with transcriptional changes likely by facilitating or preventing enhancer-promoter communication. For example, both the Nanog and Sox2 loci underwent drastic TAD reorganization by losing or gaining a strong boundary, respectively. Weakening of the Nanog boundary enabled interaction with an upstream insulated enhancer, while the de novo boundary at the Sox2 locus generated a new insulated regulatory unit that included only Sox2 and its super-enhancer, potentially preventing aberrant/competing interactions. Importantly, the authors describe an inverse correlation between timing of transcriptional activation during reprogramming and degree of topological reorganization around the gene locus (Stadhouders et al., 2018). This suggests that the requirement for 3D chromatin reorganization may act as a limiting step for gene activation and, thus, as a reprogramming roadblock. Therefore, characterization of chromatin topology around critical genes may allow us to predict and overcome important architectural barriers during cell fate transition.
Rewiring of Chromatin Loops and Hubs
In parallel with this large-scale 3D chromatin reorganization, long-range chromatin contacts also change drastically. Early 4C-seq analyses revealed a large number of PSC-specific chromatin interactions around critical stem cell genes, such as Nanog, Oct4, and Dppa2/3 (Apostolou et al., 2013; Denholtz et al., 2013; Wei et al., 2013). They also captured a dynamic rewiring of chromatin loops at different reprogramming stages, which associated with and often preceded transcriptional activation of linked genes. In contrast with these studies, the recent Hi-C analysis of B-cell reprogramming captured only a small fraction of dynamic chromatin contacts (<20% of total loops), which surprisingly did not associate with B-cell- or PSC-specific genes or with transcriptional changes (Stadhouders et al., 2018). This is likely due to technical limitations and biases of the Hi-C approach and analysis, which preferentially detects structural CTCF/Cohesin-mediated loops rather than enhancer-promoter contacts (Bonev et al., 2017; Hsieh et al., 2020; Mumbach et al., 2016; Rao et al., 2014). To overcome this limitation, a recent study employed H3K27ac HiChIP (Mumbach et al., 2016, 2017), an approach that combines Hi-C with chromatin immunoprecipitation (Di Giammartino et al., 2019) to generate 3D reference maps of active enhancers and promoter connectomes in MEFs and PSCs. This approach revealed tens of thousands of cell-type-specific regulatory contacts, strongly associated with the transcriptional activity of linked genes, suggesting that drastic 3D enhancer rewiring occurs during reprogramming. Importantly, 4C-seq analysis independently validated the abrogation of enhancer-promoter contacts around silenced somatic genes (e.g., Ets1, Jag1) and de novo establishment around activated enhancers and genes (e.g., Tbx3, Mycn, Sox2). This could support an instructive role of enhancer-promoter looping for gene activation. However, due to the biased nature of HiChIP, the authors cannot exclude that some differential HiChIP loops may simply reflect changes in H3K27ac strength in preexisting loops rather than actual 3D reorganization. Future high-resolution assays, such as Capture-C or Micro-C (Hsieh et al., 2020; Krietenstein et al., 2020), will further dissect the degree and kinetics of 3D rewiring during reprogramming and its association to gene expression.
Intriguingly, H3K27ac HiChIP analysis also revealed a number of highly interacting enhancers, which the authors call 3D enhancer hubs (Di Giammartino et al., 2019). PSC-specific enhancer hubs, which partly overlap with previously described super-enhancers (Whyte et al., 2013), interact with multiple genes that are coordinately activated during reprogramming at a much higher probability compared with gene pairs of similar linear distance or within the same TAD. Indeed, CRISPR-mediated genetic or epigenetic perturbation of enhancer hubs resulted in downregulation of multiple hub-connected genes, supporting the role of these elements in gene coregulation. Importantly, meta-analysis and advanced modeling of the B-cell reprogramming Hi-C data revealed the assembly of similar 3D chromatin hubs that associate with robust transcriptional activation (Di Stefano et al., 2020). The presence and importance of 3D chromatin hubs has been described in many other cellular contexts (originally in the context of globin locus; de Laat and Grosveld, 2003), usually in association with super-enhancers, genes critical for cell identity, and/or high transcriptional activity (recently reviewed in Di Giammartino et al., 2020). Although most of these multiconnected hubs likely represent different, high-frequency, dynamic contacts within the cell population, simultaneous multi-way contacts in the same allele and cell have been detected by imaging or other specialized chromatin topology assays, albeit at low frequency. Specifically, genome architecture mapping (GAM) in mouse PSCs captured a large number of three-way contacts, especially at highly transcribed regions and super-enhancers (Beagrie et al., 2017), further supporting a functional link between degree of 3D connectivity and levels of transcriptional activity.
Taken together, these results document drastic 3D chromatin rewiring during reprogramming, occurring alongside or prior to transcriptional changes (Figure 2). These studies also identify 3D enhancer hubs as critical architectural centers of cell identity, whose reorganization during cell fate transitions might coordinate large-scale transcriptional changes. More studies are needed to understand how 3D hubs assemble and function: Do they represent dynamic and/or heterogeneous pairwise interactions or large topological assemblies? To what degree do they represent competition or synergy among hub-connected genes and enhancers? Are they formed (or disrupted) in an all-or-nothing or a stepwise manner during cell fate change and what are the driving forces? Dissecting the organizational principles of such architectural features might allow us to interfere with their assembly and thus to promote or prevent cell fate change.
Figure 2.
Dynamic Changes in Chromatin Architecture during Somatic Cell Reprogramming
Changes in local chromatin state and TAD insulation are the earliest architectural changes that have been reported during iPSC reprogramming. Compartment switching begins to occur in tandem with new loop formation and continued changes in TAD insulation. Finally, transcriptional changes often occur concurrently or after topological reorganization.
TFs Are Major Drivers of 3D Chromatin Reorganization during Reprogramming
Somatic cell reprogramming is primarily driven by the activity of the ectopically expressed TFs, which bind the somatic genome and orchestrate many molecular changes, including transcriptional, post-transcriptional, and epigenetic remodeling, as summarized in recent reviews (Apostolou and Stadtfeld, 2018; Brumbaugh et al., 2019). The role of TFs in promoting/maintaining 3D chromatin topological features is increasingly appreciated and supported by computational and genomics analyses as well as imaging and functional experiments (recently reviewed in Di Giammartino et al., 2020; Kim and Shendure, 2019; Stadhouders et al., 2019). By combining this knowledge, we can envision various mechanisms of OKSM-driven architectural reorganization during reprogramming.
TF binding and subsequent recruitment of epigenetic modulators and chromatin remodelers can induce changes of chromatin accessibility and state, resulting in quantitative compartmental changes (e.g., strengthening or weakening) or compartment switches through self-association/segregation, as described in the introduction. Indeed, integration of Hi-C and ATAC-sequencing data during B-cell reprogramming showed that OCT4 and KLF4 binding (inferred from TF motifs around newly accessible sites) associated with early and stable B-to-A compartment switches (∼5% of the genome) even prior to transcriptional changes (Stadhouders et al., 2018). The pioneering properties of reprogramming TFs (Soufi et al., 2012; 2015) that enable binding on nucleosomal DNA followed by remodeling may contribute to early compartmental changes. However, dense heterochromatic regions were shown to be refractory to early OKSM binding (Soufi et al., 2012), suggesting that additional factors and remodeling events are required for opening and compartment switching, resulting in delayed transcriptional activation of associated genes. TF-induced silencing of the somatic program, either by direct displacement of somatic TFs, recruitment of repressive complexes (e.g., HDACs) (Chronis et al., 2017), and/or activation of critical co-repressors such as SIN3A (Li et al., 2017), may contribute to the opposite A-to-B switches.
OKS have been shown to directly interact and co-occupy chromatin with multiple architectural protein factors, including subunits of Cohesin, Mediator, and PRC1/2 complexes (Apostolou et al., 2013; Denholtz et al., 2013; Kagey et al., 2010; Pardo et al., 2010; Schoenfelder et al., 2015a; van den Berg et al., 2010; Wang et al., 2006; Wei et al., 2013). TFs and recruited coactivators at enhancer or promoter regions may mediate “stalling” of the extruding Cohesin ring resulting in de novo loop formation. The overlapping binding patterns among OKS and Cohesin subunits support this notion (Dowen et al., 2014; Kagey et al., 2010). Furthermore, highly active genes and (super)enhancers have been reported to localize within strong boundaries or on Hi-C stripes/flames (Fudenberg et al., 2016; Hsieh et al., 2020; Rao et al., 2014; Vian et al., 2018). This suggests that OKSM binding may directly induce changes in TAD organization, such as new or stronger boundaries (Stadhouders et al., 2018), and create de novo contacts (Di Giammartino et al., 2019). Obviously, TAD reorganization during reprogramming may also be induced by altered CTCF occupancy, due to DNA methylation changes (Doege et al., 2012; Gao et al., 2013; Polo et al., 2012; Sardina et al., 2018) around CTCF motifs (Bell and Felsenfeld, 2000; Hark et al., 2000; Maurano et al., 2015), or through its RNA-binding properties (Hansen et al., 2019; Saldaña-Meyer et al., 2019).
Reprogramming TFs are also involved at the finest level of topological reorganization as supported by their enriched binding at anchors of rewired chromatin contacts (Apostolou et al., 2013; Denholtz et al., 2013; Stadhouders et al., 2018). In addition, genetic perturbation experiments in established PSCs induced gain or abrogation of interactions around specific loci (de Wit et al., 2013; Novo et al., 2018; Wei et al., 2013). At a genome-wide scale, inducible depletion of KLF4 (and related KLF2 and KLF5) in PSCs resulted in a global loss of 3D enhancer connections with a preferential effect on 3D enhancer hubs (Di Giammartino et al., 2019). Similarly, CRISPR/Cas9-mediated mutation of a single KLF4 binding site within selected hubs was sufficient to induce partial disassembly of the hubs and downregulation of hub-connected genes (Di Giammartino et al., 2019). These experiments support an important role of KLF4 in the maintenance and function of 3D enhancer contacts. To also address its role in the de novo establishment/rewiring of long-range contacts, the authors performed an integrative analysis of KLF4 HiChIP, H3K27ac HiChIP, and transcriptional data at different reprogramming stages. This revealed that KLF4 binding associates both with silencing/disassembly of somatic enhancer-promoter contacts and with de novo formation of PSC enhancer-promoter contacts and hubs. KLF4 binding usually preceded or coincided with chromatin looping and enhancer/gene activation, but was not always sufficient to induce looping and/or transcription, highlighting the need for additional factors as described above. Another interesting finding was that KLF4 participates in two kinds of chromatin contacts: (1) activating enhancer-promoter loops and hubs, enriched in enhancer marks and transcriptional activators; and (2) repressive/pausing contacts enriched in H3K27me3 and PRC1/2 subunits, which were shown to promote long-range interactions among poised lineage-specific genes (de Wit et al., 2013; Denholtz et al., 2013; Pachano et al., 2019; Schoenfelder et al., 2015a; Schoenfelder and Fraser, 2019). Interestingly, many of these activating or repressive protein cofactors as well as the reprogramming TFs themselves are characterized by large intrinsically disordered regions (IDRs) (Xue et al., 2012), which were shown or presumed to mediate weak, low-affinity, multivalent interactions leading to the formation of subnuclear biomolecular condensates (Boija et al., 2018; Cho et al., 2018; Chong et al., 2018; Hnisz and Young, 2017; Sabari et al., 2018; Shrinivas et al., 2019). Such condensates may mediate not only pairwise chromatin interactions but also larger multiconnected active or repressive regulatory hubs, which are preferentially bound at high density by these protein factors (Beagrie et al., 2017; Di Giammartino et al., 2019; Hsieh et al., 2020; Rosencrance et al., 2020; Schmitt et al., 2016). Evidence that either supports or contradicts this notion has been extensively discussed in a recent review (Di Giammartino et al., 2020).
Future experiments will be critical to determine OKSM-dependent and OKSM-independent mechanisms of local and global topological reorganization during reprogramming. Another interesting direction would be to capture the dynamic formation of other PSC-characteristic structures, such as the reported extremely long-range chromatin contacts (Joshi et al., 2015; Novo et al., 2018) and the clustering of PRC-mediated bivalent genes (Denholtz et al., 2013; Schoenfelder et al., 2015b) and their significance in the establishment of pluripotency.
3D Chromatin Reorganization upon Maintenance of Pluripotency
Cell division poses a challenge for cell identity, since it is accompanied by transcriptional silencing (Gottesfeld and Forbes, 1997; Taylor, 1960), degradation or dissociation of most TFs and cofactors (Martínez-Balbás et al., 1995), widespread alterations in histone modifications (Wang and Higgins, 2013), and a dramatic reorganization of 3D genome architecture (Naumova et al., 2013) (Figure 3). PSCs are also characterized by a rapid cell cycle and short G1 phase compared with somatic cells (Savatier et al., 2002), necessitating efficient mechanisms for faithful re-establishment of PSC-specific transcriptional and chromatin landscapes upon mitotic exit. In this chapter, we will discuss our current understanding of the 3D chromatin changes that cells undergo during mitosis and G1 entry, the association with transcriptional resetting, and presumed mechanisms that ensure timely and faithful heritability of PSC fate.
Figure 3.
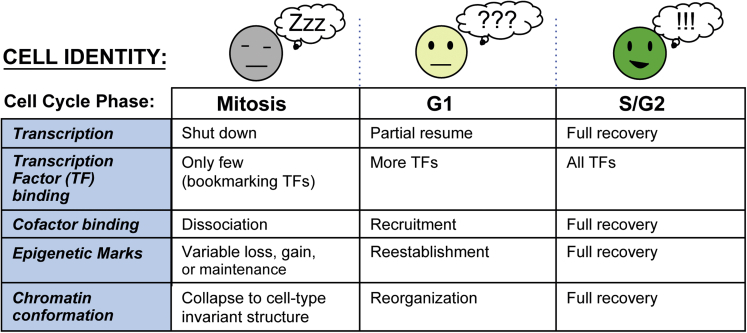
Disruption of Cell-Type-Specifying Features Causes Temporary “Identity Crisis” during Mitosis
Summary of the molecular changes that challenge cell identity during cell division. Transcription largely shuts down and most TFs and cofactors are dissociated from mitotic chromatin, except for a select few mitotic bookmarking factors. Histone modifications are variably lost (acetylation), gained (phosphorylation), or selectively maintained (bookmarked). Finally, the 3D chromatin architecture collapses from a cell-type-specific structure to a cell-type-invariant metaphase state. These features then must all be faithfully reset during G1 in the daughter cells in order to maintain proper cell identity.
Reorganization of Chromatin Architecture during Mitotic Entry and Exit
The complex hierarchical and cell-type-specific 3D chromatin topology described in the previous chapters has been shown to drastically change during mitosis. Compartments, TADs, and loops collapse and chromatin acquires a cell-type-invariant conformation that consists of a compressed array of consecutive loops (Dekker, 2014; Naumova et al., 2013) (see Figure 1). This metaphase folding state is formed through a discrete series of steps: in prophase, formation of a consecutive loop array; in prometaphase, production of a helical, spiral-staircase-like structure with a condensin II scaffold; and in metaphase, further compaction into smaller clusters by condensin I and self-interactions (Batty and Gerlich, 2019; Gibcus et al., 2018). This mitosis-specific structure has been extensively characterized by microscopy (Earnshaw and Laemmli, 1983; Liang et al., 2015b; Marsden and Laemmli, 1979; Ou et al., 2017) and 3C-based methods (Dekker, 2014; Oomen and Dekker, 2017). The metaphase chromatin structure must unfold and reset during G1 entry, in concert with recruitment of TFs, cofactors, and transcriptional reactivation.
Several recent studies have begun characterizing the dynamic resetting of chromatin architecture upon mitotic exit in different cell types (Abramo et al., 2019; Kang et al., 2020; Naumova et al., 2013; Zhang et al., 2019), including mouse naive PSCs (Nagano et al., 2017; Pelham-Webb et al., 2020). Nagano et al. (2017) utilized single-cell Hi-C in ∼3,000 asynchronous PSCs and performed cell cycle phasing to align them to a trajectory along the cell cycle. They observed a nonsynchronous manner of resetting upon mitotic exit, in which different levels of chromatin architecture (compartments, TADs, loops) displayed distinct reformation kinetics. Overall, compartments began to reform in G1, as shown previously by multiplexed 4C-seq (Dileep et al., 2015), and continued to increase in strength through S phase, reaching their maximum in G2 before dissolution in mitosis (Nagano et al., 2017). Some spatial nuclear organization (B toward the periphery and A in the nuclear interior) was already present at the beginning of G1 phase, although further compartmental segregation continued throughout G1. TAD boundaries were also rapidly reset by G1, with boundary insulation reaching its maximum strength globally at G1 (Nagano et al., 2017), as previously observed (Naumova et al., 2013).
Nagano et al. (2017) also assessed chromatin loop dynamics during cell cycle, focusing on a small number (∼2,000) of contacts with convergently oriented CTCF-bound motifs between 200 kb and 1 Mb apart. Although these few structural loops appeared stable throughout interphase, global analysis showed that cells in G1 had a lower percentage of short-range interactions than cells in S or G2, suggesting variability and asynchrony of contact reformation during mitotic exit (Nagano et al., 2017). However, the low resolution of Hi-C in this study and small number of cells captured in M-to-G1 transition make it difficult to dissect the kinetics of loop re-establishment and leave many questions unanswered. What types of chromatin contacts are established earlier or later during mitotic exit and what determines their differential kinetics? How does architectural reorganization associate with resetting of the PSC transcriptional program and identity?
More recent studies have shed light on these remaining questions in various mouse and human cell lines (Abramo et al., 2019; Kang et al., 2020; Zhang et al., 2019), including mouse PSCs (Pelham-Webb et al., 2020), by performing bulk Hi-C in various time points after mitotic arrest and release. Despite the technical differences, these reports also observed a rapid but heterogeneous resetting of genome architecture, with many similarities to the single-cell Hi-C study (Nagano et al., 2017). Compartments were detectable by ana-/telophase, but increasingly strengthened and expanded during G1, gaining longer-range A-A and B-B interactions (Abramo et al., 2019; Zhang et al., 2019). TAD boundaries were reset early, again reaching maximum insulation in early G1 (Pelham-Webb et al., 2020). Interestingly, Zhang et al. (2019) observed a “bottom-up” re-building of TADs in which small sub-TADs form first before merging into larger structures.
The increased resolution of Hi-C in these studies enabled better tracking of chromatin loop reformation kinetics, which appeared more heterogeneous than TADs or compartments (Abramo et al., 2019; Pelham-Webb et al., 2020; Zhang et al., 2019). Although many chromatin contacts were detected before G1 entry, they did not reach their half maximums until much later (Abramo et al., 2019). Short-range chromatin contacts reformed before longer ones (Pelham-Webb et al., 2020; Zhang et al., 2019) and Hi-C stripes gradually lengthened during G1 entry (Zhang et al., 2019), suggesting that CTCF/Cohesin-mediated loop extrusion (Fudenberg et al., 2016; Sanborn et al., 2015) is a major mechanism of chromatin reorganization upon mitotic exit. Interestingly, enhancer-promoter contacts were generally reset prior to structural loops in mouse G1E erythroblasts and PSCs, which could be a universal principle of architectural resetting or specific to stem and progenitor cells whose chromatin is more active and plastic. The slower kinetics of structural loops were partly due to the requirement for Cohesin reloading upon mitotic exit (Zhang et al., 2019), suggesting that the faster formation of enhancer-promoter loops might be driven by Cohesin-independent affinity forces. Importantly, chromatin contacts involving PSC super-enhancers were among the first to be detected (Pelham-Webb et al., 2020), highlighting their critical role in cell identity maintenance. These studies also reported transient conformational changes upon mitotic exit. Abramo et al. (2019) reported a temporary lack of loops during the transition from mitotic condensin-driven to interphase Cohesin-driven folding, while Zhang et al. (2019) identified a small group of transient contacts in telophase that were often disrupted by new boundaries or loops forming in G1 (Abramo et al., 2019; Zhang et al., 2019). The degree to which these transient contacts have any implications on genome function and cell identity remains to be shown.
Overall, these studies begin to shape a model of architectural resetting during mitotic exit (Figure 4). Boundary insulation is immediately reset, with some sub-TADs building up to larger TADs later on. Compartments are designated early (A or B), but gradually expand and gain strength. Loops follow more heterogeneous dynamics, with enhancer-promoter contacts reforming prior to structural ones and a few transiently established contacts. Future investigation could dissect the relative kinetics of 3D reorganization around cell-type-specific genes, enhancers, and hubs and their importance in cell fate determination.
Figure 4.
Resetting of Chromatin Architecture during Mitotic Exit
Schematic showing the resetting of different layers of chromatin architecture during the cell cycle. All of these features are disrupted in prometaphase (prometa), yet they reform with variable kinetics during mitotic exit. Compartments begin reorganizing during anaphase and telophase (ana/telo) and continue strengthening throughout S and G2. TADs are also rapidly reset, with smaller TADs in ana/telo building up to larger TADs in G1. TAD boundary insulation is strongest in G1 before weakening in S phase with the onset of DNA replication. Loops reform gradually and heterogeneously, with enhancer-promoter (Enh-Prom) contacts forming prior to CTCF/Cohesin-mediated structural loops. A few transient enhancer-promoter contacts are also observed during mitotic exit that are disrupted or lost later in the cell cycle.
Coordination of Transcriptional Reactivation with 3D Chromatin Reorganization
Alongside the chromatin reorganization during mitosis, transcription is largely halted (Prescott and Bender, 1962), as poised RNA polymerase II (Pol II) is cleared from DNA and blocked from re-initiation (Liang et al., 2015a). A recent study showed that Cohesin unloading from chromosome arms plays a role in the dissociation of Pol II during mitosis, and that suppression and reactivation of transcription is important for resetting the transcriptional landscape in G1 (Perea-Resa et al., 2020). Given the complex relationship between chromatin architecture and gene expression in steady state, how is the resetting of these processes coordinated during mitotic exit?
Recent studies have begun to characterize transcriptional reactivation during mitotic exit on a genome-wide scale, although few have explored its coordination with 3D chromatin reorganization. Here we focus on those that measured nascent transcriptional activity rather than steady-state RNA levels, by assays such as PRO-seq (precision nuclear run on sequencing) (Core et al., 2008; Kwak et al., 2013) and EU-RNA-seq (ethyl uridine-labeled RNA sequencing) (Palozola et al., 2017). These studies consistently showed that transcription of genes and enhancers is dramatically downregulated during mitosis but rapidly recovered during G1 entry (Hsiung et al., 2016; Kang et al., 2020; Palozola et al., 2017; Pelham-Webb et al., 2020; Teves et al., 2018). Studies in somatic cells also saw a global, transient spike in transcription during mitotic exit, although groups of genes showed distinct kinetics (Hsiung et al., 2016; Palozola et al., 2017). A similar hypertranscriptional spike during G1 was independently detected in mouse PSCs (Pelham-Webb et al., 2020), indicating that this is a general feature of transcriptional resetting. However, the relative timing of gene reactivation differed among cell types, with cell identity genes being slowly reactivated in somatic cells, such as hepatocytes (Kang et al., 2020; Palozola et al., 2017), while PSC-associated genes and enhancers were among the first reactivated in stem cells (Pelham-Webb et al., 2020), indicating different requirements for propagation/resetting of the PSC program. Interestingly, these studies also described a significant correlation in the reactivation kinetics between enhancers and their target genes (assigned by linear proximity or looping) (Hsiung et al., 2016; Palozola et al., 2017; Pelham-Webb et al., 2020), suggesting that transcriptional reactivation is coordinated locally and/or through looping. However, there were many examples of early reactivated genes prior to enhancer activation and/or physical association (Hsiung et al., 2016; Pelham-Webb et al., 2020), arguing for both enhancer-dependent and -independent mechanisms.
To explore the coordination of transcriptional and architectural resetting during mitotic exit on a genome-wide scale, two recent studies integrated Hi-C with either Pol II ChIP-seq in erythroblasts (Zhang et al., 2019) or PRO-seq in PSCs (Pelham-Webb et al., 2020). Zhang et al. (2019) found that transcriptional kinetics were comparable within early- and late-forming TADs, indicating a minimal effect of these structures on gene reactivation. In contrast, Pelham-Webb et al., 2020 showed that rapid transcriptional reactivation correlated with faster compartmentalization and stronger boundary insulation in early G1, providing a link between transcriptional activity and large-scale architectural reorganization after cell division. Interestingly, although transcriptional activity positively correlated with loop strength over time, both studies reported no significant difference in the reactivation kinetics between genes involved in early or late regulatory loops (Pelham-Webb et al., 2020; Zhang et al., 2019). Another recent study using EU-RNA-seq and Hi-C in U2OS cells found that transcription is largely reset prior to loop reformation, supporting that transcriptional reactivation is independent from long-range interactions during mitotic exit (Kang et al., 2020). Furthermore, in mouse PSCs, depletion of Pol II during mitosis and mitotic exit had no effect on large-scale chromatin (TAD, compartment) reformation (Jiang et al., 2020). Together, these studies indicate a surprisingly weak correlation between transcriptional and architectural resetting. However, given that Hi-C enriches for structural over regulatory loops (Bonev et al., 2017; Di Giammartino et al., 2019; Hsieh et al., 2020; Mumbach et al., 2016), perhaps Hi-C is simply an insufficient method to explore this coordination. Indeed, Capture-C in erythroblasts successfully tracked enhancer-promoter contact reformation upon mitotic exit (Hsiung et al., 2016), and 4C-seq in PSCs detected a significantly faster resetting of chromatin contacts around early-activated enhancers (including super-enhancers) compared with all other regions (Pelham-Webb et al., 2020). In addition, despite the heterogeneity/asynchrony in loop dynamics around viewpoints, this analysis detected at least one early enhancer-promoter contact around every rapidly reactivated gene. Future experiments utilizing high-resolution genome-wide assays, such as Micro-C, Capture-C, or H3K27ac HiChIP, to detect re-establishment of enhancer-promoter contacts upon mitotic exit will be required to more concretely understand the interplay between enhancer-promoter activation and physical communication.
Role of Mitotic Bookmarking Factors in Architectural Resetting during G1 Entry
The heterogeneous kinetics of topological and transcriptional resetting during mitotic exit suggest the presence of distinct architectural factors or combinations of factors that drive reorganization. We have discussed these driving forces in the previous chapters, but it is important to note that many relevant factors are dramatically altered during mitosis and G1 entry, as the epigenetic landscape and TF binding patterns of mitotic cells are distinct from interphase cells. In addition to the global topological changes during mitosis, local chromatin features, including chromatin accessibility, nucleosome positioning, and histone modifications, are also perturbed (Kelly et al., 2010; Liang et al., 2015a; Wang and Higgins, 2013). Furthermore, TFs and cofactors are largely degraded or evicted from mitotic chromatin (Martínez-Balbás et al., 1995). What chromatin features are maintained during mitosis and are they important to instruct the resetting of stem cell fate?
Recent studies using advanced proteomics, imaging, and genomics technologies have revealed milder chromatin changes during mitosis than originally reported or presumed. Despite the global compaction of chromatin, DNA accessibility is largely retained, especially at promoters of active genes (Blythe and Wieschaus, 2016; Hsiung et al., 2015; Teves et al., 2016; Xu et al., 2017). Although histone tails are globally hypoacetylated in mitosis, there is widespread retention of acetylation at certain residues (Behera et al., 2019; Hsiung et al., 2016; Javasky et al., 2018; Liu et al., 2017). This selective, partial maintenance of local interphase structure in mitosis is consistent with the concept of mitotic bookmarking (Festuccia et al., 2017; Kadauke and Blobel, 2013; Palozola et al., 2019). Recent work has explored mitotic bookmarking in mouse PSCs, identifying master regulator TFs such as ESRRB (Festuccia et al., 2016), SOX2 (Deluz et al., 2016; Liu et al., 2017; Teves et al., 2016), OCT4 (Deluz et al., 2016; Liu et al., 2017), and KLF4 (Liu et al., 2017), retained at selected genomic regions. Other mitotically retained factors and marks in PSCs include CTCF (Owens et al., 2019), TBP (Teves et al., 2018), and H3K27ac (Liu et al., 2017).
The potential bookmarking role of CTCF is particularly interesting due to its critical role in loop extrusion together with Cohesin. Owens et al. (2019) showed that while CTCF was retained during mitosis in mouse PSCs, it was lost in somatic cell lines (NIH3T3 and C2C12), which could explain incongruent results from prior studies (Burke et al., 2005; Oomen et al., 2019; Sekiya et al., 2017; Shen et al., 2015). Although CTCF is retained during mitosis, Cohesin is lost from all interphase sites (Nishiyama et al., 2013; Owens et al., 2019), arguing that CTCF bookmarking alone is likely insufficient to drive loop reformation. In support of this, while CTCF was either mitotically retained or rapidly re-associated with chromatin upon mitotic exit in G1E erythroblasts, loop recovery kinetics correlated instead with the timing of Cohesin re-recruitment (Zhang et al., 2019). Mitotic retention of CTCF may, however, promote faster boundary resetting by sustaining local chromatin accessibility and nucleosome positioning (Owens et al., 2019) at critical sites, such as TAD boundaries (Hsieh et al., 2020). On the other hand, mitotic loss of CTCF may allow formation of transient contacts during telophase (Zhang et al., 2019) or G1, resulting in aberrant gene activation.
Additional factors involved in extrusion-independent 3D chromatin organization (e.g., phase separation) and transcriptional regulation have also been identified as bookmarking factors in mouse PSCs. For example, TBP, a component of the RNA Pol II transcription complex, was widely retained at active promoters and its degradation during mitosis abrogated transcriptional reactivation globally (Teves et al., 2018). ESRRB maintained interphase nucleosome positioning during mitosis at bookmarked sites (Festuccia et al., 2019), which are enriched for stem cell identity genes (Festuccia et al., 2016). Degradation of either SOX2 (Deluz et al., 2016) or OCT4 (Liu et al., 2017) during M-to-G1 affected stemness, highlighting their importance during this window. Whether mitotic retention of any of these factors is involved in faster resetting of chromatin contacts and regulatory hubs remains to be tested.
Mitotic bookmarking by H3K27ac, which preferentially marks enhancers and genes associated with cell identity (Hsiung et al., 2016; Liu et al., 2017), was positively correlated with robust post-mitotic transcriptional reactivation in multiple cell types, including PSCs (Behera et al., 2019; Hsiung et al., 2016; Kang et al., 2020; Pelham-Webb et al., 2020), and its depletion during mitosis resulted in aberrant transcription in G1 (Kang et al., 2020; Pelham-Webb et al., 2020). Interestingly, post-mitotic recovery rate of enhancer-promoter contacts in erythroblasts positively correlated with levels of active histone marks, including H3K27ac (Zhang et al., 2019). Mitotic retention of H3K27ac was also identified as the best predictive mark for faster loop reformation and stronger TAD boundary insulation in PSCs exiting mitosis (Pelham-Webb et al., 2020). Together, these results support the role of this bookmark in faster transcriptional and architectural resetting of PSC identity after cell division (Figure 5), although further experimental evidence is required to dissect its function and underlying mechanisms. One possibility is that mitotic retention of H3K27ac promotes faster assembly of A compartments and active hubs through increased self-attraction with other homotypic chromatin regions. Alternatively, retention of H3K27ac may enable rapid recruitment of TFs/cofactors and reassembly of multiprotein complexes and condensates. One such candidate cofactor could be the chromatin reader BRD4, which plays critical roles in transcriptional activation and chromatin architecture in interphase (reviewed in Devaiah et al., 2016). Although BRD4 is partly retained on mitotic chromatin (Dey et al., 2009), its inhibition by JQ1 during mitosis did not alter transcriptional reactivation in G1 (Behera et al., 2019). This argues against the direct bookmarking function of BRD4 and suggests instead that mitotically retained histone acetylation marks, such as H3K27ac, are sufficient for rapid recruitment of BRD4 and proper genome activation during mitotic exit (Mochizuki et al., 2008; Yang et al., 2008). Future experiments depleting H3K27ac or other candidate cofactors during mitosis and mitotic exit will be required to validate this model.
Figure 5.
Role of Mitotic Bookmarking in Transcriptional and Architectural Resetting during G1 Entry
Model describing the role of mitotic bookmarking TFs and histone marks in the rapid resetting of pluripotency genes and the transient activation of lineage-specific genes. In PSCs, bookmarking factors (blue circles) are preferentially retained at stem cell-specific genes and enhancers during mitosis. This allows for fast recruitment of cofactors and transcriptional machinery (yellow ovals), resetting of enhancer-promoter contacts, and rapid transcriptional reactivation in G1 (arrows). Non-bookmarking TFs (green circles) must be recruited back to the chromatin during G1, resulting in a slower resetting of these sites. Lineage-specifying genes, often marked by PRC2 and bivalent histone marks (H3K4me3/H3K27me3) (pink circles), are transiently activated during G1. This could be caused by fluctuation of PRC2 components over the cell cycle, slower recruitment of other repressive factors (maroon ovals), and/or transient miswiring of chromatin loops.
Together, these results indicate that mitotic bookmarking factors contribute to a faster and more faithful molecular resetting of cell identity in daughter cells. It is plausible that the extent, nature, and significance of bookmarking in epigenetic inheritance of cell identity varies among cell types, depending on their cell cycle properties and vulnerability to cell fate change. Mitotic bookmarking by various histone marks and protein factors is quite widespread in PSCs (Liu et al., 2017; Teves et al., 2016), although direct side-by-side comparison with self-renewing progenitors or terminally differentiated cells is still missing. Future investigation will be required to determine the extent to which this distinct mitotic landscape of PSCs relates to their characteristic cell cycle and/or increased epigenetic plasticity and how it contributes to their ability to self-renew while maintaining pluripotency.
M-to-G1 as a Window for Cell Fate Change
The balance between self-renewal and pluripotency is especially important during mitotic exit, as G1 is the critical window for PSCs to “decide” either to self-renew or respond to differentiation cues toward defined lineages (Boward et al., 2016; Dalton and Coverdell, 2015; Soufi and Dalton, 2016). Although the concept of G1 as a window for cell fate change has been around for decades (Mummery et al., 1987; Pierce et al., 1984; Wells, 1982), only recently it was shown, for both human and mouse PSCs, that G1-sorted cells respond more rapidly to differentiation cues than S- or G2-sorted cells, which do not respond until the next G1 phase (Calder et al., 2013; Coronado et al., 2013; Pauklin and Vallier, 2013; Sela et al., 2012). Studies have also reported “noisy” expression of developmental genes during G1 (Asenjo et al., 2020; Singh et al., 2013), which could represent a temporal priming step that enables PSCs to tip the balance from self-renewal to lineage specification. This finding was independently validated using PRO-seq analysis in mouse PSCs, which revealed a weak but consistent activation of various lineage-related genes and enhancers during early and late G1 phase (Pelham-Webb et al., 2020). What is unique about the M-to-G1 phase that primes PSCs to respond to differentiation cues? And what factors are responsible for closing this window after G1 to ensure self-renewal?
Recent studies have attempted to address this question, supporting different mechanistic models for transient activation of developmental genes during G1. Studies in human PSCs have implicated signaling molecules (Smads, Cyclins), which naturally fluctuate during cell cycle, concluding that varying levels of these factors allow for differentiation to endoderm in early G1 and neuroectoderm in late G1 (Pauklin et al., 2016; Pauklin and Vallier, 2013). Others consider that transient relaxation of repressive mechanisms during M-to-G1, such as the recently described cell-cycle-dependent fluctuations of PRC2 subunits (Asenjo et al., 2020), could be sufficient to allow for gene de-repression. In agreement, promoters of transiently G1-activated genes predominantly enriched for binding of PRC2 components (Pelham-Webb et al., 2020). The dissociation of other factors, such as pluripotency TFs, specifically from repressive sites has also been observed during mitosis, which could contribute to this behavior (Festuccia et al., 2016; Liu et al., 2017).
A few recent studies support a functional link between 3D chromatin reorganization during the M-to-G1 window and transient gene activation and re-repression. For example, late formation of a TAD boundary could allow for temporary enhancer-promoter “miswiring”, as observed during telophase in erythroblasts (Zhang et al., 2019). Alternatively, slower re-establishment of repressive loops (e.g., PRC1/2-mediated contacts) could enable transient de-repression of poised developmental genes. Both categories of temporarily mis-wired or late-established contacts have been detected by 4C-seq around the transiently activated, lineage-specific genes Gata6 and Neurod1 (Pelham-Webb et al., 2020). Interestingly, this study also captured a strong contact between the Gata6 promoter and an endoderm enhancer, which was pre-established/maintained in mitosis (Pelham-Webb et al., 2020). This suggests that developmentally relevant chromatin structures may be already established in PSCs, as shown before (Cruz-Molina et al., 2017; Rubin et al., 2017; Schoenfelder et al., 2015a), but also selectively maintained during mitosis to enable rapid G1 activation and differentiation upon proper stimulation. The degree to which such permissive and “hard-wired” chromatin loops exist on a genome-wide scale and their functional role for transcriptional priming toward successful differentiation remain to be tested. Follow-up experiments in single cells will be critical to determine whether transient activation occurs in all or a subset of cells during G1 and the functional impact on lineage preference.
Conclusions and Future Perspectives
PSCs are considered a powerful system for disease modeling and regenerative medicine. Their unique cell identity is determined and maintained by a specific gene expression program, unique epigenetic landscape, and characteristic 3D genomic organization. Delineating the complex interplay among these regulatory layers is crucial for understanding and modulating PSC fate and harnessing their full biomedical potential. As discussed above, significant progress has been made by characterizing the dynamic changes that result in the (re)establishment of PSC identity starting either from a defined somatic identity (reprogramming) or a mitotic state (during self-renewal). A deeper understanding of these process by constructing high-resolution 4D molecular roadmaps that integrate multiple -omics analyses will help address the following important questions: what are the essential transcriptional and architectural milestones and bottlenecks toward establishment of PSC identity? Are there common or unique regulatory hierarchies associated with each process (e.g., different order of establishment of PSC-associated features) and what are the critical driving forces? Can we identify specific vulnerabilities that may enable modulation of the self-renewal versus differentiation balance and a controlled pluripotency exit toward new fates?
A fundamental and actively debated question regarding cell fate control is the functional interplay between 3D genome architecture and transcriptional regulation (Beagan and Phillips-Cremins, 2020; Gibcus and Dekker, 2013; Gorkin et al., 2014; Robson et al., 2019; van Steensel and Furlong, 2019). The integrative reprogramming studies described above demonstrated that architectural changes (compartment switches, TAD insulation changes, and loop reorganization) often precede or coincide with transcriptional changes. Although this could support a cause-and-effect relationship, additional experimental evidence is required. In vivo imaging of chromatin reorganization, by advanced CRISPR-based approaches such as LiveFISH (Wang et al., 2019) or CLING (Maass et al., 2018), will enable tracking of long-range chromatin contacts and their association to gene activation. More importantly, breaking or engineering the chromatin loops/hubs predicted to be critical for PSC identity with technologies such as CRIPSR-GO (Wang et al., 2018) or LADL (Kim et al., 2019) will directly test their functional relevance. Previous studies in different systems have supported an instructive role of 3D chromatin architecture on gene regulation, showing that topological reorganization (altered insulation and/or looping) caused transcriptional changes of linked or nearby genes (Deng et al., 2012; Lupiáñez et al., 2015; Schoenfelder and Fraser, 2019). Other studies suggest a permissive role, where preexisting chromatin loops enabled gene activation only upon recruitment of protein factors (Cruz-Molina et al., 2017; Ghavi-Helm et al., 2014; Jin et al., 2013; Melo et al., 2013; Rubin et al., 2017; Schoenfelder et al., 2015a). Finally, there is evidence for a reciprocal interplay and positive feedback between transcription and topological organization (van Steensel and Furlong, 2019). For example, transcriptional inhibition was shown to alter enhancer-promoter contacts (Hsieh et al., 2020) and to weaken CTCF-mediated loops that require its RNA-binding domain (Saldaña-Meyer et al., 2019), while genomic insertion of a transcription start site (TSS) was sufficient to induce local chromatin folding (Zhang et al., 2020). Moreover, RNA per se has been extensively documented to shape various subnuclear compartments and chromatin interactions (Quinodoz et al., 2020; Rinn and Guttman, 2014). Importantly, in most studies, perturbation of candidate factors and features happened in asynchronous cells and for variable time periods, often over many cell divisions. Therefore, it is hard to uncouple direct from indirect effects as well as their role in maintenance versus establishment of a molecular state (e.g., gene activation or loop formation). We argue that systematic interrogation of candidate factors specifically during mitosis or M-to-G1 may help provide more definitive answers on the role of these regulators. To this direction, advanced technologies that enable temporal and reversible protein degradation (Kadauke et al., 2012; Nabet et al., 2018) or genetic/epigenetic engineering combined with reporters that enable prospective isolation or tracking of cells at defined cell cycle stages (Sakaue-Sawano et al., 2008; Zerjatke et al., 2017) will be critical. Finally, an important consideration when performing or interpreting such studies is the degree of cell-type specificity of the molecular principles that are uncovered. Therefore, systematic comparison of multiple cell types or dynamic cell fate transitions are extremely valuable for dissecting universal and cell-type-specific mechanisms.
Acknowledgments
We thank members of the Apostolou and the Stadtfeld groups for input on our manuscript. We apologize to colleagues whose work was not cited due to space constraints. B.P.W. is supported by the NICHD (F30HD097926) and a Medical Scientist Training Program grant by the NIGMS (T32GM007739), and D.M. is supported by a Training Grant in Developmental and Stem Cell Biology by the NICHD (T32HD060600). E.A. is a recipient of the NIH Director's New Innovator award (DP2DA043813).
References
- Abramo K., Valton A.-L., Venev S.V., Ozadam H., Fox A.N., Dekker J. A chromosome folding intermediate at the condensin-to-cohesin transition during telophase. Nat. Cell Biol. 2019;21:1393–1402. doi: 10.1038/s41556-019-0406-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apostolou E., Ferrari F., Walsh R.M., Bar-Nur O., Stadtfeld M., Cheloufi S., Stuart H.T., Polo J.M., Ohsumi T.K., Borowsky M.L. Genome-wide chromatin interactions of the Nanog locus in pluripotency, differentiation, and reprogramming. Cell Stem Cell. 2013;12:699–712. doi: 10.1016/j.stem.2013.04.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apostolou E., Hochedlinger K. Chromatin dynamics during cellular reprogramming. Nature. 2013;502:462–471. doi: 10.1038/nature12749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apostolou E., Stadtfeld M. Cellular trajectories and molecular mechanisms of iPSC reprogramming. Curr. Opin. Genet. Dev. 2018;52:77–85. doi: 10.1016/j.gde.2018.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asenjo H.G., Gallardo A., López-Onieva L., Tejada I., Martorell-Marugán J., Carmona-Sáez P., Landeira D. Polycomb regulation is coupled to cell cycle transition in pluripotent stem cells. Sci. Adv. 2020;6:eaay4768. doi: 10.1126/sciadv.aay4768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batty P., Gerlich D.W. Mitotic chromosome mechanics: how cells segregate their genome. Trends Cell Biol. 2019;29:717–726. doi: 10.1016/j.tcb.2019.05.007. [DOI] [PubMed] [Google Scholar]
- Beagan J.A., Gilgenast T.G., Kim J., Plona Z., Norton H.K., Hu G., Hsu S.C., Shields E.J., Lyu X., Apostolou E. Local genome topology can exhibit an incompletely rewired 3D-folding state during somatic cell reprogramming. Cell Stem Cell. 2016;18:611–624. doi: 10.1016/j.stem.2016.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beagan J.A., Phillips-Cremins J.E. On the existence and functionality of topologically associating domains. Nat. Genet. 2020;52:8–16. doi: 10.1038/s41588-019-0561-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beagrie R.A., Scialdone A., Schueler M., Kraemer D.C., Chotalia M., Xie S.Q., Barbieri M., de Santiago I., Lavitas L.M., Branco M.R. Complex multi-enhancer contacts captured by genome architecture mapping. Nature. 2017;543:519–524. doi: 10.1038/nature21411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behera V., Stonestrom A.J., Hamagami N., Hsiung C.C., Keller C.A., Giardine B., Sidoli S., Yuan Z.-F., Bhanu N.V., Werner M.T. Interrogating histone acetylation and BRD4 as mitotic bookmarks of transcription. Cell Rep. 2019;27:400–415.e5. doi: 10.1016/j.celrep.2019.03.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell A.C., Felsenfeld G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature. 2000;405:482–485. doi: 10.1038/35013100. [DOI] [PubMed] [Google Scholar]
- Blythe S.A., Wieschaus E.F. Establishment and maintenance of heritable chromatin structure during early Drosophila embryogenesis. eLife. 2016;5:e20148. doi: 10.7554/eLife.20148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boija A., Klein I.A., Sabari B.R., Dall'Agnese A., Coffey E.L., Zamudio A.V., Li C.H., Shrinivas K., Manteiga J.C., Hannett N.M. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell. 2018;175:1842–1855.e16. doi: 10.1016/j.cell.2018.10.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonev B., Cohen N.M., Szabo Q., Fritsch L., Papadopoulos G.L., Lubling Y., Xu X., Lv X., Hugnot J.-P., Tanay A. Multiscale 3D genome rewiring during mouse neural development. Cell. 2017;171:557–572.e24. doi: 10.1016/j.cell.2017.09.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boward B., Wu T., Dalton S. Concise review: control of cell fate through cell cycle and pluripotency networks. Stem Cells. 2016;34:1427–1436. doi: 10.1002/stem.2345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brumbaugh J., Di Stefano B., Hochedlinger K. Reprogramming: identifying the mechanisms that safeguard cell identity. Development. 2019;146:dev182170. doi: 10.1242/dev.182170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burke L.J., Zhang R., Bartkuhn M., Tiwari V.K., Tavoosidana G., Kurukuti S., Weth C., Leers J., Galjart N., Ohlsson R. CTCF binding and higher order chromatin structure of the H19 locus are maintained in mitotic chromatin. EMBO J. 2005;24:3291–3300. doi: 10.1038/sj.emboj.7600793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calder A., Roth-Albin I., Bhatia S., Pilquil C., Lee J.H., Bhatia M., Levadoux-Martin M., McNicol J., Russell J., Collins T. Lengthened G1 phase indicates differentiation status in human embryonic stem cells. Stem Cells Dev. 2013;22:279–295. doi: 10.1089/scd.2012.0168. [DOI] [PubMed] [Google Scholar]
- Cheutin T., Cavalli G. The multiscale effects of polycomb mechanisms on 3D chromatin folding. Crit. Rev. Biochem. Mol. Biol. 2019;54:399–417. doi: 10.1080/10409238.2019.1679082. [DOI] [PubMed] [Google Scholar]
- Cho W.-K., Spille J.-H., Hecht M., Lee C., Li C., Grube V., Cisse I.I. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science. 2018:eaar4199. doi: 10.1126/science.aar4199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chong S., Dugast-Darzacq C., Liu Z., Dong P., Dailey G.M., Cattoglio C., Heckert A., Banala S., Lavis L., Darzacq X. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science. 2018:eaar2555. doi: 10.1126/science.aar2555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chronis C., Fiziev P., Papp B., Butz S., Bonora G., Sabri S., Ernst J., Plath K. Cooperative binding of transcription factors orchestrates reprogramming. Cell. 2017;168:442–459.e20. doi: 10.1016/j.cell.2016.12.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Core L.J., Waterfall J.J., Lis J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science. 2008;322:1845–1848. doi: 10.1126/science.1162228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coronado D., Godet M., Bourillot P.-Y., Tapponnier Y., Bernat A., Petit M., Afanassieff M., Markossian S., Malashicheva A., Iacone R. A short G1 phase is an intrinsic determinant of naïve embryonic stem cell pluripotency. Stem Cell Res. 2013;10:118–131. doi: 10.1016/j.scr.2012.10.004. [DOI] [PubMed] [Google Scholar]
- Crane E., Bian Q., McCord R.P., Lajoie B.R., Wheeler B.S., Ralston E.J., Uzawa S., Dekker J., Meyer B.J. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature. 2015;523:240–244. doi: 10.1038/nature14450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cremer T., Cremer C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2001;2:292–301. doi: 10.1038/35066075. [DOI] [PubMed] [Google Scholar]
- Cruz-Molina S., Respuela P., Tebartz C., Kolovos P., Nikolic M., Fueyo R., van Ijcken W.F.J., Grosveld F., Frommolt P., Bazzi H. PRC2 facilitates the regulatory topology required for poised enhancer function during pluripotent stem cell differentiation. Cell Stem Cell. 2017;20:689–705.e9. doi: 10.1016/j.stem.2017.02.004. [DOI] [PubMed] [Google Scholar]
- Dalton S., Coverdell P. Linking the cell cycle to cell fate decisions. Trends Cell Biol. 2015;25:592–600. doi: 10.1016/j.tcb.2015.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Laat W., Grosveld F. Spatial organization of gene expression: the active chromatin hub. Chromosome Res. 2003;11:447–459. doi: 10.1023/a:1024922626726. [DOI] [PubMed] [Google Scholar]
- de Wit E., Bouwman B.A., Zhu Y., Klous P., Splinter E., Verstegen M.J., Krijger P.H., Festuccia N., Nora E.P., Welling M. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature. 2013;501:227–231. doi: 10.1038/nature12420. [DOI] [PubMed] [Google Scholar]
- Dekker J. Two ways to fold the genome during the cell cycle: insights obtained with chromosome conformation capture. Epigenetics Chromatin. 2014;7:25. doi: 10.1186/1756-8935-7-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deluz C., Friman E.T., Strebinger D., Benke A., Raccaud M., Callegari A., Leleu M., Manley S., Suter D.M. A role for mitotic bookmarking of SOX2 in pluripotency and differentiation. Genes Dev. 2016;30:2538–2550. doi: 10.1101/gad.289256.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng W., Lee J., Wang H., Miller J., Reik A., Gregory P.D., Dean A., Blobel G.A. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell. 2012;149:1233–1244. doi: 10.1016/j.cell.2012.03.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denholtz M., Bonora G., Chronis C., Splinter E., de Laat W., Ernst J., Pellegrini M., Plath K. Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell Stem Cell. 2013;13:602–616. doi: 10.1016/j.stem.2013.08.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devaiah B.N., Gegonne A., Singer D.S. Bromodomain 4: a cellular Swiss army knife. J. Leukoc. Biol. 2016;100:679–686. doi: 10.1189/jlb.2RI0616-250R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dey A., Nishiyama A., Karpova T., McNally J., Ozato K. Brd4 marks select genes on mitotic chromatin and directs postmitotic transcription. Mol. Biol. Cell. 2009;20:4899–4909. doi: 10.1091/mbc.E09-05-0380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Giammartino D.C., Apostolou E. The chromatin signature of pluripotency: establishment and maintenance. Curr. Stem Cell Rep. 2016;2:255–262. doi: 10.1007/s40778-016-0055-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Giammartino D.C., Kloetgen A., Polyzos A., Liu Y., Kim D., Murphy D., Abuhashem A., Cavaliere P., Aronson B., Shah V. KLF4 is involved in the organization and regulation of pluripotency-associated three-dimensional enhancer networks. Nat. Cell Biol. 2019;21:1179–1190. doi: 10.1038/s41556-019-0390-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Giammartino D.C., Polyzos A., Apostolou E. Transcription factors: building hubs in the 3D space. Cell Cycle. 2020:1–16. doi: 10.1080/15384101.2020.1805238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Stefano M., Stadhouders R., Farabella I., Castillo D., Serra F., Graf T., Marti-Renom M.A. Transcriptional activation during cell reprogramming correlates with the formation of 3D open chromatin hubs. Nat. Commun. 2020;11:2564. doi: 10.1038/s41467-020-16396-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dileep V., Ay F., Sima J., Vera D.L., Noble W.S., Gilbert D.M. Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication-timing program. Genome Res. 2015;25:1104–1113. doi: 10.1101/gr.183699.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dixon J.R., Gorkin D.U., Ren B. Chromatin domains: the unit of chromosome organization. Mol. Cell. 2016;62:668–680. doi: 10.1016/j.molcel.2016.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dixon J.R., Jung I., Selvaraj S., Shen Y., Antosiewicz-Bourget J.E., Lee A.Y., Ye Z., Kim A., Rajagopal N., Xie W. Chromatin architecture reorganization during stem cell differentiation. Nature. 2015;518:331–336. doi: 10.1038/nature14222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dixon J.R., Selvaraj S., Yue F., Kim A., Li Y., Shen Y., Hu M., Liu J.S., Ren B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012;485:376–380. doi: 10.1038/nature11082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doege C.A., Inoue K., Yamashita T., Rhee D.B., Travis S., Fujita R., Guarnieri P., Bhagat G., Vanti W.B., Shih A. Early-stage epigenetic modification during somatic cell reprogramming by Parp1 and Tet2. Nature. 2012;488:652–655. doi: 10.1038/nature11333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dowen J.M., Fan Z.P., Hnisz D., Ren G., Abraham B.J., Zhang L.N., Weintraub A.S., Schujiers J., Lee T.I., Zhao K. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell. 2014;159:374–387. doi: 10.1016/j.cell.2014.09.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eagen K.P., Aiden E.L., Kornberg R.D. Polycomb-mediated chromatin loops revealed by a subkilobase-resolution chromatin interaction map. Proc. Natl. Acad. Sci. U S A. 2017;114:8764–8769. doi: 10.1073/pnas.1701291114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Earnshaw W.C., Laemmli U.K. Architecture of metaphase chromosomes and chromosome scaffolds. J. Cell Biol. 1983;96:84–93. doi: 10.1083/jcb.96.1.84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans M. Discovering pluripotency: 30 years of mouse embryonic stem cells. Nat. Rev. Mol. Cell Biol. 2011;12:680–686. doi: 10.1038/nrm3190. [DOI] [PubMed] [Google Scholar]
- Festuccia N., Dubois A., Vandormael-Pournin S., Tejeda E.G., Mouren A., Bessonnard S., Mueller F., Proux C., Cohen-Tannoudji M., Navarro P. Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat. Cell Biol. 2016;18:1139–1148. doi: 10.1038/ncb3418. [DOI] [PubMed] [Google Scholar]
- Festuccia N., Gonzalez I., Owens N., Navarro P. Mitotic bookmarking in development and stem cells. Development. 2017;144:3633–3645. doi: 10.1242/dev.146522. [DOI] [PubMed] [Google Scholar]
- Festuccia N., Owens N., Papadopoulou T., Gonzalez I., Tachtsidi A., Vandoermel-Pournin S., Gallego E., Gutierrez N., Dubois A., Cohen-Tannoudji M. Transcription factor activity and nucleosome organization in mitosis. Genome Res. 2019;29:250–260. doi: 10.1101/gr.243048.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fudenberg G., Imakaev M., Lu C., Goloborodko A., Abdennur N., Mirny L.A. formation of chromosomal domains by loop extrusion. Cell Rep. 2016;15:2038–2049. doi: 10.1016/j.celrep.2016.04.085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao L., Smit M.A., van den Oord J.J., Goeman J.J., Verdegaal E.M., van der Burg S.H., Stas M., Beck S., Gruis N.A., Tensen C.P. Genome-wide promoter methylation analysis identifies epigenetic silencing of MAPK13 in primary cutaneous melanoma. Pigment Cell Melanoma Res. 2013;26:542–554. doi: 10.1111/pcmr.12096. [DOI] [PubMed] [Google Scholar]
- Ghavi-Helm Y., Klein F.A., Pakozdi T., Ciglar L., Noordermeer D., Huber W., Furlong E.E. Enhancer loops appear stable during development and are associated with paused polymerase. Nature. 2014;512:96–100. doi: 10.1038/nature13417. [DOI] [PubMed] [Google Scholar]
- Gibcus J.H., Dekker J. The hierarchy of the 3D genome. Mol. Cell. 2013;49:773–782. doi: 10.1016/j.molcel.2013.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibcus J.H., Samejima K., Goloborodko A., Samejima I., Naumova N., Nuebler J., Kanemaki M.T., Xie L., Paulson J.R., Earnshaw W.C. A pathway for mitotic chromosome formation. Science. 2018;359:eaao6135. doi: 10.1126/science.aao6135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorkin D.U., Leung D., Ren B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell. 2014;14:762–775. doi: 10.1016/j.stem.2014.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottesfeld J.M., Forbes D.J. Mitotic repression of the transcriptional machinery. Trends Biochem. Sci. 1997;22:197–202. doi: 10.1016/s0968-0004(97)01045-1. [DOI] [PubMed] [Google Scholar]
- Guelen L., Pagie L., Brasset E., Meuleman W., Faza M.B., Talhout W., Eussen B.H., de Klein A., Wessels L., de Laat W. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature. 2008;453:948–951. doi: 10.1038/nature06947. [DOI] [PubMed] [Google Scholar]
- Haarhuis J.H.I., van der Weide R.H., Blomen V.A., Yanez-Cuna J.O., Amendola M., van Ruiten M.S., Krijger P.H.L., Teunissen H., Medema R.H., van Steensel B. The cohesin release factor WAPL restricts chromatin loop extension. Cell. 2017;169:693–707.e14. doi: 10.1016/j.cell.2017.04.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Handoko L., Xu H., Li G., Ngan C.Y., Chew E., Schnapp M., Lee C.W.H., Ye C., Ping J.L.H., Mulawadi F. CTCF-mediated functional chromatin interactome in pluripotent cells. Nat. Genet. 2011;43:630–638. doi: 10.1038/ng.857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen A.S., Hsieh T.S., Cattoglio C., Pustova I., Saldana-Meyer R., Reinberg D., Darzacq X., Tjian R. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell. 2019;76:395–411.e13. doi: 10.1016/j.molcel.2019.07.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harikumar A., Meshorer E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 2015;16:1609–1619. doi: 10.15252/embr.201541011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hark A.T., Schoenherr C.J., Katz D.J., Ingram R.S., Levorse J.M., Tilghman S.M. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature. 2000;405:486–489. doi: 10.1038/35013106. [DOI] [PubMed] [Google Scholar]
- Hnisz D., Young R.A. New insights into genome structure: genes of a feather stick together. Mol. Cell. 2017;67:730–731. doi: 10.1016/j.molcel.2017.08.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh T.-H.S., Cattoglio C., Slobodyanyuk E., Hansen A.S., Rando O.J., Tjian R., Darzacq X. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell. 2020;78:539–553.e8. doi: 10.1016/j.molcel.2020.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiung C.C.S., Bartman C.R., Huang P., Ginart P., Stonestrom A.J., Keller C.A., Face C., Jahn K.S., Evans P., Sankaranarayanan L. A hyperactive transcriptional state marks genome reactivation at the mitosis–G1 transition. Genes Dev. 2016;30:1423–1439. doi: 10.1101/gad.280859.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiung C.C.S., Morrissey C.S., Udugama M., Frank C.L., Keller C.A., Baek S., Giardine B., Crawford G.E., Sung M.-H., Hardison R.C. Genome accessibility is widely preserved and locally modulated during mitosis. Genome Res. 2015;25:213–225. doi: 10.1101/gr.180646.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javasky E., Shamir I., Gandhi S., Egri S., Sandler O., Rothbart S.B., Kaplan N., Jaffe J.D., Goren A., Simon I. Study of mitotic chromatin supports a model of bookmarking by histone modifications and reveals nucleosome deposition patterns. Genome Res. 2018;28:1455–1466. doi: 10.1101/gr.230300.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang Y., Huang J., Lun K., Li B., Zheng H., Li Y., Zhou R., Duan W., Feng Y., Yao H. Genome-wide analyses of chromatin interactions after the loss of Pol I, Pol II, and Pol III. Genome Biol. 2020;21:158–186. doi: 10.1186/s13059-020-02067-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin F., Li Y., Dixon J.R., Selvaraj S., Ye Z., Lee A.Y., Yen C.-A., Schmitt A.D., Espinoza C.A., Ren B. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature. 2013;503:290–294. doi: 10.1038/nature12644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joshi O., Wang S.Y., Kuznetsova T., Atlasi Y., Peng T., Fabre P.J., Habibi E., Shaik J., Saeed S., Handoko L. Dynamic reorganization of extremely long-range promoter-promoter interactions between two states of pluripotency. Cell Stem Cell. 2015;17:748–757. doi: 10.1016/j.stem.2015.11.010. [DOI] [PubMed] [Google Scholar]
- Kadauke S., Blobel G.A. Mitotic bookmarking by transcription factors. Epigenetics & Chromatin. 2013;6:6. doi: 10.1186/1756-8935-6-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadauke S., Udugama M.I., Pawlicki J.M., Achtman J.C., Jain D.P., Cheng Y., Hardison R.C., Blobel G.A. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell. 2012;150:725–737. doi: 10.1016/j.cell.2012.06.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kagey M.H., Newman J.J., Bilodeau S., Zhan Y., Orlando D.A., van Berkum N.L., Ebmeier C.C., Goossens J., Rahl P.B., Levine S.S. Mediator and cohesin connect gene expression and chromatin architecture. Nature. 2010;467:430–435. doi: 10.1038/nature09380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang H., Shokhirev M.N., Xu Z., Chandran S., Dixon J.R., Hetzer M.W. Dynamic regulation of histone modifications and long-range chromosomal interactions during postmitotic transcriptional reactivation. Genes Dev. 2020;34:913–920. doi: 10.1101/gad.335794.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelly T.K., Miranda T.B., Liang G., Berman B.P., Lin J.C., Tanay A., Jones P.A. H2A.Z maintenance during mitosis reveals nucleosome shifting on mitotically silenced genes. Mol. Cell. 2010;39:901–911. doi: 10.1016/j.molcel.2010.08.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kempfer R., Pombo A. Methods for mapping 3D chromosome architecture. Nat. Rev. Genet. 2020;21:207–226. doi: 10.1038/s41576-019-0195-2. [DOI] [PubMed] [Google Scholar]
- Kim J.H., Rege M., Valeri J., Dunagin M.C., Metzger A., Titus K.R., Gilgenast T.G., Gong W., Beagan J.A., Raj A. LADL: light-activated dynamic looping for endogenous gene expression control. Nat. Methods. 2019;16:633–639. doi: 10.1038/s41592-019-0436-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S., Shendure J. Mechanisms of interplay between transcription factors and the 3D genome. Mol. Cell. 2019;76:306–319. doi: 10.1016/j.molcel.2019.08.010. [DOI] [PubMed] [Google Scholar]
- Krietenstein N., Abraham S., Venev S.V., Abdennur N., Gibcus J., Hsieh T.-H.S., Parsi K.M., Yang L., Maehr R., Mirny L.A. Ultrastructural details of mammalian chromosome architecture. Mol. Cell. 2020;78:554–565. doi: 10.1016/j.molcel.2020.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krijger P.H.L., Di Stefano B., de Wit E., Limone F., van Oevelen C., de Laat W., Graf T. Cell-of-Origin-Specific 3D genome structure acquired during somatic cell reprogramming. Cell Stem Cell. 2016;18:597–610. doi: 10.1016/j.stem.2016.01.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwak H., Fuda N.J., Core L.J., Lis J.T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science. 2013;339:950–953. doi: 10.1126/science.1229386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D., Liu J., Yang X., Zhou C., Guo J., Wu C., Qin Y., Guo L., He J., Yu S. Chromatin accessibility dynamics during iPSC reprogramming. Cell Stem Cell. 2017;21:819–833.e6. doi: 10.1016/j.stem.2017.10.012. [DOI] [PubMed] [Google Scholar]
- Liang K., Woodfin A.R., Slaughter B.D., Unruh J.R., Box A.C., Rickels R.A., Gao X., Haug J.S., Jaspersen S.L., Shilatifard A. Mitotic transcriptional activation: clearance of actively engaged Pol II via transcriptional elongation control in mitosis. Mol. Cell. 2015;60:435–445. doi: 10.1016/j.molcel.2015.09.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang Z., Zickler D., Prentiss M., Chang F.S., Witz G., Maeshima K., Kleckner N. Chromosomes progress to metaphase in multiple discrete steps via global compaction/expansion cycles. Cell. 2015;161:1124–1137. doi: 10.1016/j.cell.2015.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lieberman-Aiden E., van Berkum N.L., Williams L., Imakaev M., Ragoczy T., Telling A., Amit I., Lajoie B.R., Sabo P.J., Dorschner M.O. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–293. doi: 10.1126/science.1181369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y., Pelham-Webb B., Di Giammartino D.C., Li J., Kim D., Kita K., Saiz N., Garg V., Doane A., Giannakakou P. Widespread mitotic bookmarking by histone marks and transcription factors in pluripotent stem cells. Cell Rep. 2017;19:1283–1293. doi: 10.1016/j.celrep.2017.04.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lupiáñez D.G., Kraft K., Heinrich V., Krawitz P., Brancati F., Klopocki E., Horn D., Kayserili H., Opitz J.M., Laxova R. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161:1012–1025. doi: 10.1016/j.cell.2015.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maass P.G., Barutcu A.R., Weiner C.L., Rinn J.L. Inter-chromosomal contact properties in live-cell imaging and in Hi-C. Mol. Cell. 2018;69:1039–1045.e3. doi: 10.1016/j.molcel.2018.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsden M.P.F., Laemmli U.K. Metaphase chromosome structure: evidence for a radial loop model. Cell. 1979;17:849–858. doi: 10.1016/0092-8674(79)90325-8. [DOI] [PubMed] [Google Scholar]
- Martínez-Balbás M.A., Dey A., Rabindran S.K., Ozato K., Wu C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell. 1995;83:29–38. doi: 10.1016/0092-8674(95)90231-7. [DOI] [PubMed] [Google Scholar]
- Maurano M.T., Wang H., John S., Shafer A., Canfield T., Lee K., Stamatoyannopoulos J.A. Role of DNA methylation in modulating transcription factor occupancy. Cell Rep. 2015;12:1184–1195. doi: 10.1016/j.celrep.2015.07.024. [DOI] [PubMed] [Google Scholar]
- Melo C.A., Drost J., Wijchers P.J., van de Werken H., de Wit E., Oude Vrielink J.A., Elkon R., Melo S.A., Léveillé N., Kalluri R. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell. 2013;49:524–535. doi: 10.1016/j.molcel.2012.11.021. [DOI] [PubMed] [Google Scholar]
- Mochizuki K., Nishiyama A., Jang M.K., Dey A., Ghosh A., Tamura T., Natsume H., Yao H., Ozato K. The bromodomain protein Brd4 stimulates G1 gene transcription and promotes progression to S phase. J. Biol. Chem. 2008;283:9040–9048. doi: 10.1074/jbc.M707603200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mumbach M.R., Rubin A.J., Flynn R.A., Dai C., Khavari P.A., Greenleaf W.J., Chang H.Y. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods. 2016;13:919–922. doi: 10.1038/nmeth.3999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mumbach M.R., Satpathy A.T., Boyle E.A., Dai C., Gowen B.G., Cho S.W., Nguyen M.L., Rubin A.J., Granja J.M., Kazane K.R. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 2017;49:1602–1612. doi: 10.1038/ng.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mummery C.L., van Rooijen M.A., van den Brink S.E., de Laat S.W. Cell cycle analysis during retinoic acid induced differentiation of a human embryonal carcinoma-derived cell line. Cell Differ. 1987;20:153–160. doi: 10.1016/0045-6039(87)90429-5. [DOI] [PubMed] [Google Scholar]
- Nabet B., Roberts J.M., Buckley D.L., Paulk J., Dastjerdi S., Yang A., Leggett A.L., Erb M.A., Lawlor M.A., Souza A. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 2018;14:431–441. doi: 10.1038/s41589-018-0021-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagano T., Lubling Y., Várnai C., Dudley C., Leung W., Baran Y., Cohen N.M., Wingett S., Fraser P., Tanay A. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature. 2017;547:61–67. doi: 10.1038/nature23001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naumova N., Imakaev M., Fudenberg G., Zhan Y., Lajoie B.R., Mirny L.A., Dekker J. Organization of the mitotic chromosome. Science. 2013;342:948–953. doi: 10.1126/science.1236083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishiyama T., Sykora M.M., Huis in ’t Veld P.J., Mechtler K., Peters J.-M. Aurora B and Cdk1 mediate Wapl activation and release of acetylated cohesin from chromosomes by phosphorylating Sororin. Proc. Natl. Acad. Sci. U S A. 2013;110:13404. doi: 10.1073/pnas.1305020110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nora E.P., Goloborodko A., Valton A.L., Gibcus J.H., Uebersohn A., Abdennur N., Dekker J., Mirny L.A., Bruneau B.G. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell. 2017;169:930–944.e22. doi: 10.1016/j.cell.2017.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novo C.L., Javierre B.-M., Cairns J., Segonds-Pichon A., Wingett S.W., Freire-Pritchett P., Furlan-Magaril M., Schoenfelder S., Fraser P., Rugg-Gunn P.J. Long-range enhancer interactions are prevalent in mouse embryonic stem cells and are reorganized upon pluripotent state transition. Cell Rep. 2018;22:2615–2627. doi: 10.1016/j.celrep.2018.02.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nuebler J., Fudenberg G., Imakaev M., Abdennur N., Mirny L.A. Chromatin organization by an interplay of loop extrusion and compartmental segregation. Proc. Natl. Acad. Sci. U S A. 2018;115:E6697–E6706. doi: 10.1073/pnas.1717730115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oomen M.E., Dekker J. Epigenetic characteristics of the mitotic chromosome in 1D and 3D. Crit. Rev. Biochem. Mol. Biol. 2017;52:185–204. doi: 10.1080/10409238.2017.1287160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oomen M.E., Hansen A.S., Liu Y., Darzacq X., Dekker J. CTCF sites display cell cycle-dependent dynamics in factor binding and nucleosome positioning. Genome Res. 2019;29:236–249. doi: 10.1101/gr.241547.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ou H.D., Phan S., Deerinck T.J., Thor A., Ellisman M.H., O’Shea C.C. ChromEMT: visualizing 3D chromatin structure and compaction in interphase and mitotic cells. Science. 2017;357:eaag0025. doi: 10.1126/science.aag0025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Owens N., Papadopoulou T., Festuccia N., Tachtsidi A., Gonzalez I., Dubois A., Vandormael-Pournin S., Nora E.P., Bruneau B., Cohen-Tannoudji M. CTCF confers local nucleosome resiliency after DNA replication and during mitosis. eLife. 2019;8:e47898. doi: 10.7554/eLife.47898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pachano T., Crispatzu G., Rada-Iglesias A. Polycomb proteins as organizers of 3D genome architecture in embryonic stem cells. Brief. Funct. Genomics. 2019;18:358–366. doi: 10.1093/bfgp/elz022. [DOI] [PubMed] [Google Scholar]
- Palikyras S., Papantonis A. Modes of phase separation affecting chromatin regulation. Open Biol. 2019;9:190167. doi: 10.1098/rsob.190167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palozola K.C., Donahue G., Liu H., Grant G.R., Becker J.S., Cote A., Yu H., Raj A., Zaret K.S. Mitotic transcription and waves of gene reactivation during mitotic exit. Science. 2017;358:119–122. doi: 10.1126/science.aal4671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palozola K.C., Lerner J., Zaret K.S. A changing paradigm of transcriptional memory propagation through mitosis. Nat. Rev. Mol. Cell Biol. 2019;20:55–64. doi: 10.1038/s41580-018-0077-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papp B., Plath K. Epigenetics of reprogramming to induced pluripotency. Cell. 2013;152:1324–1343. doi: 10.1016/j.cell.2013.02.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pardo M., Lang B., Yu L., Prosser H., Bradley A., Babu M.M., Choudhary J. An expanded Oct4 interaction network: implications for stem cell biology, development, and disease. Cell Stem Cell. 2010;6:382–395. doi: 10.1016/j.stem.2010.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pauklin S., Madrigal P., Bertero A., Vallier L. Initiation of stem cell differentiation involves cell cycle-dependent regulation of developmental genes by Cyclin D. Genes Dev. 2016;30:421–433. doi: 10.1101/gad.271452.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pauklin S., Vallier L. The cell-cycle state of stem cells determines cell fate propensity. Cell. 2013;155:135–147. doi: 10.1016/j.cell.2013.08.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelham-Webb B., Polyzos A., Wojenski L., Kloetgen A., Li J., Di Giammartino D.C., Core L., Tsirigos A., Apostolou E. Mitotic retention of H3K27 acetylation promotes rapid topological and transcriptional resetting of stem cell-related genes and enhancers upon G1 entry. bioRxiv. 2020 doi: 10.1101/2020.06.02.130104. [DOI] [Google Scholar]
- Penalosa-Ruiz G., Bright A.R., Mulder K.W., Veenstra G.J.C. The interplay of chromatin and transcription factors during cell fate transitions in development and reprogramming. Biochim. Biophys. Acta Gene Regul. Mech. 2019;1862:194407. doi: 10.1016/j.bbagrm.2019.194407. [DOI] [PubMed] [Google Scholar]
- Perea-Resa C., Bury L., Cheeseman I.M., Blower M.D. Cohesin removal reprograms gene expression upon mitotic entry. Mol. Cell. 2020;78:127–140.e7. doi: 10.1016/j.molcel.2020.01.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips-Cremins J.E., Sauria M.E., Sanyal A., Gerasimova T.I., Lajoie B.R., Bell J.S., Ong C.T., Hookway T.A., Guo C., Sun Y. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell. 2013;153:1281–1295. doi: 10.1016/j.cell.2013.04.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pierce G.B., Aguilar D., Hood G., Wells R.S. Trophectoderm in control of murine embryonal carcinoma. Cancer Res. 1984;44:3987–3996. [PubMed] [Google Scholar]
- Polo J.M., Anderssen E., Walsh R.M., Schwarz B.A., Nefzger C.M., Lim S.M., Borkent M., Apostolou E., Alaei S., Cloutier J. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell. 2012;151:1617–1632. doi: 10.1016/j.cell.2012.11.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prescott D.M., Bender M.A. Synthesis of RNA and protein during mitosis in mammalian tissue culture cells. Exp. Cell Res. 1962;26:260–268. doi: 10.1016/0014-4827(62)90176-3. [DOI] [PubMed] [Google Scholar]
- Quinodoz S.A., Bhat P., Ollikainen N., Jachowicz J.W., Banerjee A.K., Chovanec P., Blanco M.R., Chow A., Markaki Y., Plath K. RNA promotes the formation of spatial compartments in the nucleus. bioRxiv. 2020 doi: 10.1101/2020.08.25.267435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rada-Iglesias A., Grosveld F.G., Papantonis A. Forces driving the three-dimensional folding of eukaryotic genomes. Mol. Syst. Biol. 2018:e8214. doi: 10.15252/msb.20188214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao S.S.P., Huang S.-C., Glenn St Hilaire B., Engreitz J.M., Perez E.M., Kieffer-Kwon K.-R., Sanborn A.L., Johnstone S.E., Bascom G.D., Bochkov I.D. Cohesin loss eliminates all loop domains. Cell. 2017;171:305–320.e24. doi: 10.1016/j.cell.2017.09.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao S.S.P., Huntley M.H., Durand N.C., Stamenova E.K., Bochkov I.D., Robinson J.T., Sanborn A.L., Machol I., Omer A.D., Lander E.S. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159:1665–1680. doi: 10.1016/j.cell.2014.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinn J., Guttman M. RNA Function. RNA and dynamic nuclear organization. Science. 2014;345:1240–1241. doi: 10.1126/science.1252966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robson M.I., Ringel A.R., Mundlos S. Regulatory landscaping: how enhancer-promoter communication is sculpted in 3D. Mol. Cell. 2019;74:1110–1122. doi: 10.1016/j.molcel.2019.05.032. [DOI] [PubMed] [Google Scholar]
- Rosencrance C.D., Ammouri H.N., Yu Q., Ge T., Rendleman E.J., Marshall S.A., Eagen K.P. Chromatin hyperacetylation impacts chromosome folding by forming a nuclear subcompartment. Mol. Cell. 2020;78:112–126 e112. doi: 10.1016/j.molcel.2020.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowley M.J., Nichols M.H., Lyu X., Ando-Kuri M., Rivera I.S.M., Hermetz K., Wang P., Ruan Y., Corces V.G. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell. 2017;67:837–852.e7. doi: 10.1016/j.molcel.2017.07.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin A.J., Barajas B.C., Furlan-Magaril M., Lopez-Pajares V., Mumbach M.R., Howard I., Kim D.S., Boxer L.D., Cairns J., Spivakov M. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat. Genet. 2017;49:1522–1528. doi: 10.1038/ng.3935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabari B.R., Dall’Agnese A., Boija A., Klein I.A., Coffey E.L., Shrinivas K., Abraham B.J., Hannett N.M., Zamudio A.V., Manteiga J.C. Coactivator condensation at super-enhancers links phase separation and gene control.10.1126/science.aar3958. Science. 2018;361:eaar3958. doi: 10.1126/science.aar3958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakaue-Sawano A., Kurokawa H., Morimura T., Hanyu A., Hama H., Osawa H., Kashiwagi S., Fukami K., Miyata T., Miyoshi H. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell. 2008;132:487–498. doi: 10.1016/j.cell.2007.12.033. [DOI] [PubMed] [Google Scholar]
- Saldaña-Meyer R., Rodriguez-Hernaez J., Escobar T., Nishana M., Jácome-López K., Nora E.P., Bruneau B.G., Tsirigos A., Furlan-Magaril M., Skok J. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cel. 2019;76:412–422.e15. doi: 10.1016/j.molcel.2019.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanborn A.L., Rao S.S., Huang S.C., Durand N.C., Huntley M.H., Jewett A.I., Bochkov I.D., Chinnappan D., Cutkosky A., Li J. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. U S A. 2015;112:E6456–E6465. doi: 10.1073/pnas.1518552112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sardina J.L., Collombet S., Tian T.V., Gomez A., Di Stefano B., Berenguer C., Brumbaugh J., Stadhouders R., Segura-Morales C., Gut M. Transcription factors drive tet2-mediated enhancer demethylation to reprogram cell fate. Cell Stem Cell. 2018;23:727–741.e9. doi: 10.1016/j.stem.2018.08.016. [DOI] [PubMed] [Google Scholar]
- Savatier P., Lapillonne H., Jirmanova L., Vitelli L., Samarut J. Analysis of the cell cycle in mouse embryonic stem cells. Methods Mol. Biol. 2002;185:27–33. doi: 10.1385/1-59259-241-4:27. [DOI] [PubMed] [Google Scholar]
- Schmitt A.D., Hu M., Jung I., Xu Z., Qiu Y., Tan C.L., Li Y., Lin S., Lin Y., Barr C.L. A compendium of chromatin contact maps reveals spatially active regions in the human genome. Cell Rep. 2016;17:2042–2059. doi: 10.1016/j.celrep.2016.10.061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoenfelder S., Fraser P. Long-range enhancer–promoter contacts in gene expression control. Nat. Rev. Genet. 2019;55:1. doi: 10.1038/s41576-019-0128-0. [DOI] [PubMed] [Google Scholar]
- Schoenfelder S., Furlan-Magaril M., Mifsud B., Tavares-Cadete F., Sugar R., Javierre B.M., Nagano T., Katsman Y., Sakthidevi M., Wingett S.W. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 2015;25:582–597. doi: 10.1101/gr.185272.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoenfelder S., Sugar R., Dimond A., Javierre B.M., Armstrong H., Mifsud B., Dimitrova E., Matheson L., Tavares-Cadete F., Furlan-Magaril M. Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat. Genet. 2015;47:1179–1186. doi: 10.1038/ng.3393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarzer W., Abdennur N., Goloborodko A., Pekowska A., Fudenberg G., Loe-Mie Y., Fonseca N.A., Huber W., Haering C., Mirny L. Two independent modes of chromatin organization revealed by cohesin removal. Nature. 2017;551:51–56. doi: 10.1038/nature24281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekiya T., Murano K., Kato K., Kawaguchi A., Nagata K. Mitotic phosphorylation of CCCTC-binding factor (CTCF) reduces its DNA binding activity. FEBS Open Biol. 2017;7:397–404. doi: 10.1002/2211-5463.12189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sela Y., Molotski N., Golan S., Itskovitz-Eldor J., Soen Y. Human embryonic stem cells exhibit increased propensity to differentiate during the G1 phase prior to phosphorylation of retinoblastoma protein. Stem Cells. 2012;30:1097–1108. doi: 10.1002/stem.1078. [DOI] [PubMed] [Google Scholar]
- Sexton T., Cavalli G. The role of chromosome domains in shaping the functional genome. Cell. 2015;160:1049–1059. doi: 10.1016/j.cell.2015.02.040. [DOI] [PubMed] [Google Scholar]
- Sexton T., Yaffe E., Kenigsberg E., Bantignies F., Leblanc B., Hoichman M., Parrinello H., Tanay A., Cavalli G. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell. 2012;148:458–472. doi: 10.1016/j.cell.2012.01.010. [DOI] [PubMed] [Google Scholar]
- Shen W., Wang D., Ye B., Shi M., Zhang Y., Zhao Z. A possible role of Drosophila CTCF in mitotic bookmarking and maintaining chromatin domains during the cell cycle. Biol. Res. 2015;48:27. doi: 10.1186/s40659-015-0019-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shrinivas K., Sabari B.R., Coffey E.L., Klein I.A., Boija A., Zamudio A.V., Schuijers J., Hannett N.M., Sharp P.A., Young R.A. Enhancer features that drive formation of transcriptional condensates. Mol. Cell. 2019;75:549–561.e7. doi: 10.1016/j.molcel.2019.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh A.M., Chappell J., Trost R., Lin L., Wang T., Tang J., Matlock B.K., Weller K.P., Wu H., Zhao S. Cell-cycle control of developmentally regulated transcription factors accounts for heterogeneity in human pluripotent cells. Stem Cell Rep. 2013;1:532–544. doi: 10.1016/j.stemcr.2013.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soufi A., Dalton S. Cycling through developmental decisions: how cell cycle dynamics control pluripotency, differentiation and reprogramming. Development. 2016;143:4301–4311. doi: 10.1242/dev.142075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soufi A., Donahue G., Zaret K.S. Facilitators and impediments of the pluripotency reprogramming factors' initial engagement with the genome. Cell. 2012;151:994–1004. doi: 10.1016/j.cell.2012.09.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soufi A., Garcia M.F., Jaroszewicz A., Osman N., Pellegrini M., Zaret K.S. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell. 2015;161:555–568. doi: 10.1016/j.cell.2015.03.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stadhouders R., Filion G.J., Graf T. Transcription factors and 3D genome conformation in cell-fate decisions. Nature. 2019;569:345–354. doi: 10.1038/s41586-019-1182-7. [DOI] [PubMed] [Google Scholar]
- Stadhouders R., Vidal E., Serra F., Di Stefano B., Le Dily F., Quilez J., Gomez A., Collombet S., Berenguer C., Cuartero Y. Transcription factors orchestrate dynamic interplay between genome topology and gene regulation during cell reprogramming. Nat. Genet. 2018;50:238–249. doi: 10.1038/s41588-017-0030-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun F., Chronis C., Kronenberg M., Chen X.F., Su T., Lay F.D., Plath K., Kurdistani S.K., Carey M.F. Promoter-enhancer communication occurs primarily within insulated neighborhoods. Mol. Cell. 2019;73:250–263 e255. doi: 10.1016/j.molcel.2018.10.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabar V., Studer L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat. Rev. Genet. 2014;15:82–92. doi: 10.1038/nrg3563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takahashi K., Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–676. doi: 10.1016/j.cell.2006.07.024. [DOI] [PubMed] [Google Scholar]
- Takebayashi S.-i., Dileep V., Ryba T., Dennis J.H., Gilbert D.M. Chromatin-interaction compartment switch at developmentally regulated chromosomal domains reveals an unusual principle of chromatin folding. Proc. Natl. Acad. Sci. U S A. 2012;109:12574–12579. doi: 10.1073/pnas.1207185109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang Z., Luo O.J., Li X., Zheng M., Zhu J.J., Szalaj P., Trzaskoma P., Magalska A., Wlodarczyk J., Ruszczycki B. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell. 2015;163:1611–1627. doi: 10.1016/j.cell.2015.11.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatavosian R., Kent S., Brown K., Yao T., Duc H.N., Huynh T.N., Zhen C.Y., Ma B., Wang H., Ren X. Nuclear condensates of the Polycomb protein chromobox 2 (CBX2) assemble through phase separation. J. Biol. Chem. 2019;294:1451–1463. doi: 10.1074/jbc.RA118.006620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor J.H. Nucleic acid synthesis in relation to the cell division cycle. Ann. N. Y Acad. Sci. 1960;90:409–421. doi: 10.1111/j.1749-6632.1960.tb23259.x. [DOI] [PubMed] [Google Scholar]
- Teves S.S., An L., Bhargava-Shah A., Xie L., Darzacq X., Tjian R. A stable mode of bookmarking by TBP recruits RNA polymerase II to mitotic chromosomes. eLife. 2018;7:e35621. doi: 10.7554/eLife.35621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teves S.S., An L., Hansen A.S., Xie L., Darzacq X., Tjian R. A dynamic mode of mitotic bookmarking by transcription factors. eLife. 2016;5:e22280. doi: 10.7554/eLife.22280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Bortle K., Nichols M.H., Li L., Ong C.T., Takenaka N., Qin Z.S., Corces V.G. Insulator function and topological domain border strength scale with architectural protein occupancy. Genome Biol. 2014;15:R82. doi: 10.1186/gb-2014-15-5-r82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Berg D.L., Snoek T., Mullin N.P., Yates A., Bezstarosti K., Demmers J., Chambers I., Poot R.A. An Oct4-centered protein interaction network in embryonic stem cells. Cell Stem Cell. 2010;6:369–381. doi: 10.1016/j.stem.2010.02.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Steensel B., Furlong E.E.M. The role of transcription in shaping the spatial organization of the genome. Nat. Rev. Mol. Cell Biol. 2019;20:327–337. doi: 10.1038/s41580-019-0114-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vian L., Pekowska A., Rao S.S.P., Kieffer-Kwon K.R., Jung S., Baranello L., Huang S.C., El Khattabi L., Dose M., Pruett N. The energetics and physiological impact of cohesin extrusion. Cell. 2018;173:1165–1178.e20. doi: 10.1016/j.cell.2018.03.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Higgins J.M.G. Histone modifications and mitosis: countermarks, landmarks, and bookmarks. Trends Cell Biol. 2013;23:175–184. doi: 10.1016/j.tcb.2012.11.005. [DOI] [PubMed] [Google Scholar]
- Wang H., Nakamura M., Abbott T.R., Zhao D., Luo K., Yu C., Nguyen C.M., Lo A., Daley T.P., La Russa M. CRISPR-mediated live imaging of genome editing and transcription. Science. 2019;365:1301. doi: 10.1126/science.aax7852. [DOI] [PubMed] [Google Scholar]
- Wang H., Xu X., Nguyen C.M., Liu Y., Gao Y., Lin X., Daley T., Kipniss N.H., La Russa M., Qi L.S. CRISPR-mediated programmable 3D genome positioning and nuclear organization. Cell. 2018;175:1405–1417.e14. doi: 10.1016/j.cell.2018.09.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J., Rao S., Chu J., Shen X., Levasseur D.N., Theunissen T.W., Orkin S.H. A protein interaction network for pluripotency of embryonic stem cells. Nature. 2006;444:364–368. doi: 10.1038/nature05284. [DOI] [PubMed] [Google Scholar]
- Wang S., Su J.H., Beliveau B.J., Bintu B., Moffitt J.R., Wu C.T., Zhuang X. Spatial organization of chromatin domains and compartments in single chromosomes. Science. 2016;353:598–602. doi: 10.1126/science.aaf8084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei Z., Gao F., Kim S., Yang H., Lyu J., An W., Wang K., Lu W. Klf4 organizes long-range chromosomal interactions with the oct4 locus in reprogramming and pluripotency. Cell Stem Cell. 2013;13:36–47. doi: 10.1016/j.stem.2013.05.010. [DOI] [PubMed] [Google Scholar]
- Wells R.S. An in vitro assay for growth regulation of embryonal carcinoma by the blastocyst. Cancer Res. 1982;42:2736–2741. [PubMed] [Google Scholar]
- Whyte W.A., Orlando D.A., Hnisz D., Abraham B.J., Lin C.Y., Kagey M.H., Rahl P.B., Lee T.I., Young R.A. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–319. doi: 10.1016/j.cell.2013.03.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wutz G., Várnai C., Nagasaka K., Cisneros D.A., Stocsits R.R., Tang W., Schoenfelder S., Jessberger G., Muhar M., Hossain M.J. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 2017;36:3573–3599. doi: 10.15252/embj.201798004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu J., Carter A.C., Gendrel A.-V., Attia M., Loftus J., Greenleaf W.J., Tibshirani R., Heard E., Chang H.Y. Landscape of monoallelic DNA accessibility in mouse embryonic stem cells and neural progenitor cells. Nat. Genet. 2017;49:377–386. doi: 10.1038/ng.3769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xue B., Oldfield C.J., Van Y.Y., Dunker A.K., Uversky V.N. Protein intrinsic disorder and induced pluripotent stem cells. Mol. Biosyst. 2012;8:134–150. doi: 10.1039/c1mb05163f. [DOI] [PubMed] [Google Scholar]
- Yang Z., He N., Zhou Q. Brd4 recruits P-TEFb to chromosomes at late mitosis to promote G1 gene expression and cell cycle progression. Mol. Cell Biol. 2008;28:967–976. doi: 10.1128/MCB.01020-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu M., Ren B. The three-dimensional organization of mammalian genomes. Annu. Rev. Cell Dev. Biol. 2017;33:265–289. doi: 10.1146/annurev-cellbio-100616-060531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zamudio A.V., Dall'Agnese A., Henninger J.E., Manteiga J.C., Afeyan L.K., Hannett N.M., Coffey E.L., Li C.H., Oksuz O., Sabari B.R. Mediator condensates localize signaling factors to key cell identity genes. Mol. Cell. 2019;76:753–766.e6. doi: 10.1016/j.molcel.2019.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zerjatke T., Gak I.A., Kirova D., Fuhrmann M., Daniel K., Gonciarz M., Müller D., Glauche I., Mansfeld J. Quantitative cell cycle analysis based on an endogenous all-in-one reporter for cell tracking and classification. Cell Rep. 2017;19:1953–1966. doi: 10.1016/j.celrep.2017.05.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang D., Huang P., Sharma M., Keller C.A., Giardine B., Zhang H., Gilgenast T.G., Phillips-Cremins J.E., Hardison R.C., Blobel G.A. Alteration of genome folding via contact domain boundary insertion. Nat. Genet. 2020;52:1076–1087. doi: 10.1038/s41588-020-0680-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang H., Emerson D.J., Gilgenast T.G., Titus K.R., Lan Y., Huang P., Zhang D., Wang H., Keller C.A., Giardine B. Chromatin structure dynamics during the mitosis-to-G1 phase transition. Nature. 2019;576:158–162. doi: 10.1038/s41586-019-1778-y. [DOI] [PMC free article] [PubMed] [Google Scholar]