Abstract

The biological responses to dienone compounds with a 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophore have been studied extensively. Despite their expected general thiol reactivity, these compounds display considerable degrees of tumor cell selectivity. Here we review in vitro and preclinical studies of dienone compounds including b-AP15, VLX1570, RA-9, RA-190, EF24, HO-3867, and MCB-613. A common property of these compounds is their targeting of the ubiquitin–proteasome system (UPS), known to be essential for the viability of tumor cells. Gene expression profiling experiments have shown induction of responses characteristic of UPS inhibition, and experiments using cellular reporter proteins have shown that proteasome inhibition is associated with cell death. Other mechanisms of action such as reactivation of mutant p53, stimulation of steroid receptor coactivators, and induction of protein cross-linking have also been described. Although unsuitable as biological probes due to widespread reactivity, dienone compounds are cytotoxic to apoptosis-resistant tumor cells and show activity in animal tumor models.
Introduction
A considerable number of anticancer drugs have been discovered and developed using phenotypic screening.1 Large numbers (in the order of 103–106) of molecules can be screened for identification of compounds that induce a desired biological response.2 Screens can be set up using different strategies, using tumor cells that are resistant to conventional therapy, using cells with defined genetic makeups, using cellular reporter systems, etc.3 A disadvantage of this approach is that identified compounds may be associated with extensive polypharmacology. Furthermore, it is generally difficult to define precise mechanisms of action of screening hits. The revolution of molecular biology has, however, resulted in substantially improved possibilities for elucidation of targets and of molecular mechanisms of action. Thus, methods such as gene expression profiling4 (CMap), various proteomic techniques,5−7 and knock-down and gene deletion technologies can be used to generate hypotheses with regard to molecular mechanisms of action, to identify and validate potential targets.3
Investigators performing phenotypic screening are faced with the problem of how to manage projects where hit compounds display interesting features in terms of generating desired biological responses but are unattractive from a medicinal chemistry point of view. Should such projects be discontinued since, according to experience, such compounds can rarely be developed into useful drugs? Or is this a prejudiced view? Cancer is a disease characterized by the occurrence of a large number of genetic and epigenetic alterations.8 There is a considerable medical need for efficient drugs for improved treatment of diseases with dismal outcomes such as pancreatic cancer and glioblastoma. The clinical efficacy of many anticancer drugs is hampered by the development of clones expressing variant target proteins, resulting in treatment failure.9,10 Perhaps no stone should be left unturned?
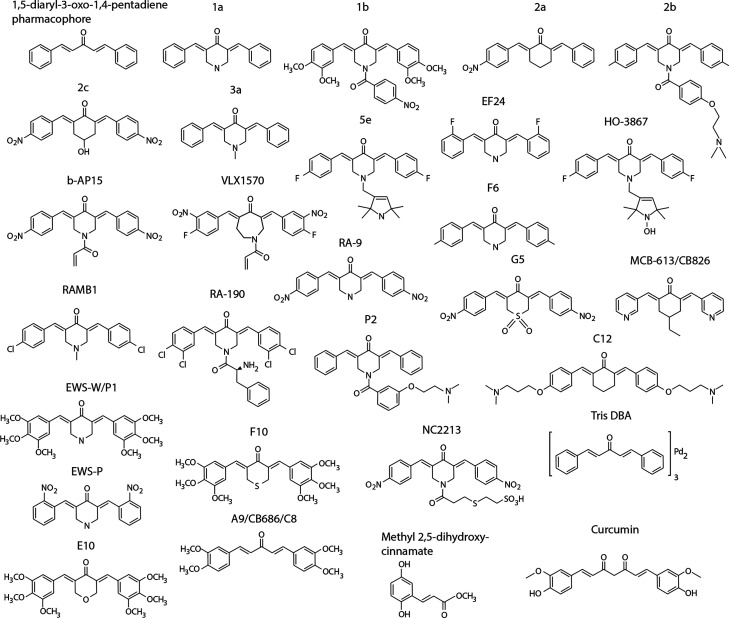
Here we review the reported biological activities of substances containing 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophores such as 3,5-diarylidene-4-piperidones and dibenzylidene-acetone (DBA) (Figure 1). These compounds are electrophiles, primarily reacting with thiols,11 and will be referred to as “dienones”. Dienone compounds with completely different structures such as prostaglandins of the J-series will generally not be considered here. The general assumption is that molecules containing 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophore will show similar mechanisms of action in biological systems by acting as Michael acceptors. This assumption is supported by the finding of common patterns of response in cells exposed to different compounds (see below).
Figure 1.
Chemical structures of some of the compounds discussed in the text.
Cancer Cells Are Selectively Sensitive to Dienone Compounds
Early studies of compounds containing the 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophore were performed by Dimmock and colleagues at the University of Saskatchewan.12−15 These investigators documented interesting biological responses to these compounds, stimulating further interest. One of the early compounds described by Dimmock et al. was 1a(13) (Figure 1), a compound that inhibited the proliferation of cancer cells in the low μM range. Dimmock and colleagues have over the years studied a respectable number of compounds belonging to this class and have established structure–activity relationships (reviewed in refs (11) and (16)). The dienone has been found to be required for optimal cytotoxicity, and as expected, electron drawing groups on the aryls increase the antiproliferative effect.17
An interesting property of the compounds described by Dimmock and co-workers is their preferential cytotoxic to tumor cells.18−21 The Dimmock group reported selective indices (sensitivity of tumor cells relative to noncancerous cells) of >10-fold19 (see Figure 2). Selective antiproliferative activity to tumor cells has been verified by other laboratories21−26 (Figure 2, Supplementary Table 1). Analyses have not been restricted only to slowly proliferating primary cells but also to established immortalized noncancerous cell lines such as MCF-10A (a human breast cell line with a reported population doubling time of ∼19 h 27) and hTERT-RPE1 (an immortalized human epithelial cell line with a doubling time of ∼14 h 28). The proliferation rates of these cells are in the same range as those of many of the cancer cells listed in Supplementary Table 1. It is therefore unlikely that the preferential cytotoxicities of dienones to tumor cells are solely due to their differences in proliferation rates.
Figure 2.
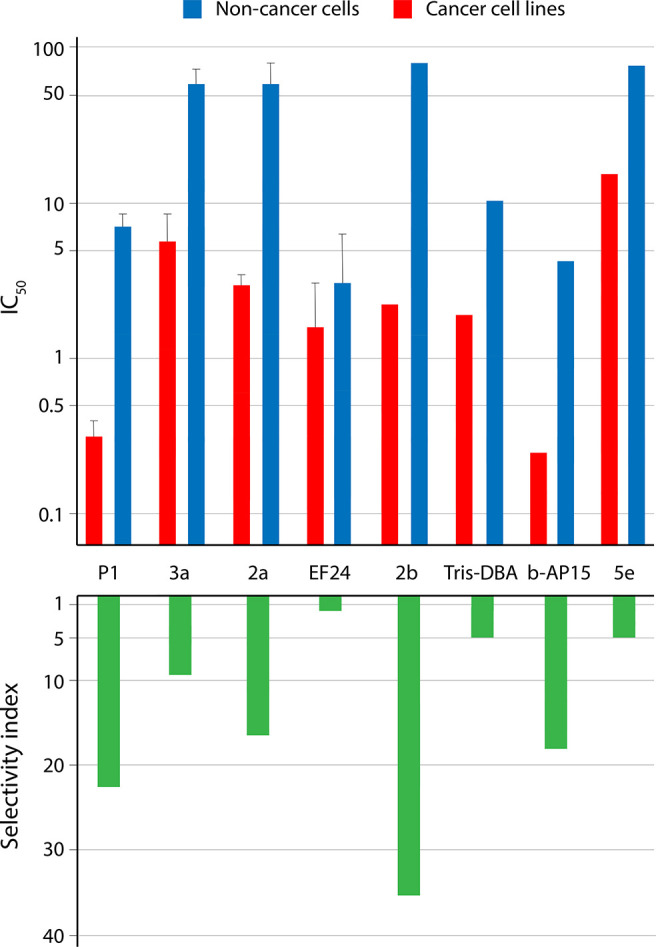
Tumor cell selectivity of dienone compounds. The log10 IC50 values are shown on the y-axis in the upper graph and selectivity indices on the y-axis in the lower graph. Shown are the mean ± SD values where possible to calculate (i.e., where data from three or more cell lines were reported). The figure is based on the data in presented Supplementary Table 1.
EF24 (3,5-bis(2-fluorobenzylidene)piperidin-4-one) (Figure 1) was identified in a screen of different α,β-unsaturated ketones for their activities against cancer cell lines.15 EF24 was also suggested to exhibit antiangiogenic properties based on inhibition of the proliferation of endothelial cells.15 EF24 induces apoptosis as evidenced by activation of caspase-3, externalization of phosphatidylserine, and depolarization of mitochondrial membranes.29 Inhibition of the proliferation of colon and gastric adenocarcinoma cells was observed at a concentration of 1 μM, whereas mouse embryo fibroblasts were not affected at 5 μM.26 Selective cytotoxicity to tumor cells by EF24 was also reported by Santiago-Vazquez and co-workers (Figure 2, Supplementary Table 1).20 Daily ip administration of EF24 suppressed the growth of colon cancer tumor xenografts in nude mice.26 Administration of 100 mg/kg EF24 did not cause harmful side effects to animals.15 These findings have been repeated in later studies of EF24 using hepatocellular carcinoma,30 adrenocortical tumors,31 colorectal cancer,32 cholangiocellular carcinoma,33 oral squamous cell carcinoma34 and pancreatic cancer.35
H-4073 is a para-fluorinated variant of EF24 described to be more potent to ovarian cancer cells than the parental compound.36 Yet another related compound, named HO-3867, was subsequently developed (Figure 1).37 HO-3867 showed a higher level of growth inhibitory activity to ovarian cancer cells compared to untransformed human ovarian surface epithelial cells.37 HO-3867 was effective in an ovarian cancer xenograft model using a schedule of oral administration. Further studies have shown that HO-3867 induces apoptosis of breast, colon, and liver cells whereas normal breast fibroblasts showed limited sensitivity.38 HO-3867 was also reported to show in vitro activity on pancreas cancer cell lines39,40 and endometrial cancer cell lines.41 An EF24 analogue was recently described that showed a similar antiproliferative activity toward cancer cells as the parental compound but lower activity to normal cells.42 For a recent review on the biological activities of EF24, see ref (43).
The group of Brancolini and co-workers at the University of Udine identified two compounds, F6 (NSC632839) and G5 (NSC144303) (Figure 1), as capable of triggering caspase activation in cells with a mutated caspase-9.44 A prodrug version of G5 (2c-PE) that can be activated by esterases was developed and shown to suppress the growth of A549 lung carcinoma xenografts in mice.45
The compound NSC687852 (later named b-AP15) was originally identified in a screen for compounds that induce apoptosis of colon cancer cells independently of p53.46 The compound was later found to elicit the lysosomal apoptosis pathway47 and to show antineoplastic activity in different tumor models including lung cancer and acute myelocytic leukemia.22 Nontransformed hTERT-RPE1 epithelial cells showed limited sensitivity to b-AP1522 (Figure 2). VLX1570 is an analogue of b-AP15 with a central azepane ring (Figure 1) that induces an indistinguishable cellular response from b-AP15 while having a slightly improved solubility and potency.48,49 VLX1570 has been shown to have antitumor activity in multiple myeloma,49 Ewings sarcoma,50 Waldenströms macroglobulinemia,51 and also other diseases (see52).
Bazzaro and colleagues53,54 demonstrated cytotoxicity of a series of 3,5-diarylidene-4-piperidones (RA-1 to RA-9, RAMB1 to RAMB4) toward three cervical cancer cell lines, whereas the viability of primary human keratinocytes was only marginally affected. The compound RA-9 (Figure 1) induced apoptosis of ovarian cancer cell lines but not of primary ovarian surface epithelial cells.24 A related compound named RA-190 was later shown to preferentially kill HPV-transformed cells.55 RA-190 reduced the growth of both multiple myeloma and ovarian cancer xenografts and the growth of HPV-16 positive cells in a syngeneic mouse model.55
Ewing sarcoma is a bone malignancy that affects children.56 Patients with metastatic disease have dismal outcomes despite aggressive treatment. Shukla et al. screened a library of >300 000 small molecules to identify compounds that selectively inhibit the growth of Ewing sarcoma cell lines.50 The screen identified two benzyl-4-piperidone compounds, EWS-P (NSC666038) and EWS-W (NSC669441) (Figure 1).
Wang et al.23 showed that the compound MCB-613 (Figure 1) induced ER stress and reactive oxygen species (ROS) in tumor cell lines. The potency of the compound in proliferation assays was 3–6 μM on tumor cell lines, whereas primary hepatocytes and mouse embryo fibroblasts were less sensitive. MCB-613 was also identified in a screen for UPS inhibitors (as CB826) and was shown to be cytotoxic to melanoma, colon cancer, and multiple myeloma cells.57
A number of additional laboratories have synthesized and examined the biological responses to various dienone compounds, generally 3,5-bis(arylidene)-4-piperidones. Two different groups have reported activity of dienones on estrogen receptor negative breast cancer cells.58,59 Stronger antiproliferative activities were observed toward breast cancer cells compared to the noncancerous human breast cell line MCF-10A. Kalai et al. reported the antiproliferative activities of a number of 3,5-bis(arylidene)-4-piperidones, including 5e, on two cancer cell lines and a noncancerous cardiomyoblast cell line60 (Figure 1, Figure 2, Supplementary Table 1). Phosphonate derivatives of 3,5-bis(arylidene)-4-piperidones were found to display inhibitory properties toward four different human cancer cell lines with IC50 values in the range of 2.5–8.5 μM.61 α,β-unsaturated piperidones were described that were active at <1 μM when tested on five cancer cell lines.62 One of the compounds (1b) (Figure 1) displayed marked inhibitory effects in a lung carcinoma xenograft model with no apparent toxicity. Wei et al. synthesized 61 symmetrical dienone compounds and examined their anticancer activity toward three different cancer cell lines.63 Two compounds containing sulfur or oxygen heteroatoms in the 4-position of the central six-carbon ring (E10, F10; Figure 1) exhibited antiproliferative activities at concentrations below 1 μM. Youssef et al. reported that amino acid conjugates of 3,5-bisarylmethylene-4-piperidones display improved cytostatic activities compared to compounds without N-substitutions.64 Finally, Cheng et al. showed that the 3,5-bis(arylidene)-4-piperidone 7g decreased PSA promoter activity and lowered viability of LNCaP prostate cancer cells.65
Molecules lacking the central ring structure (dibenzylideneacetone, DBA, and related molecules, Figure 1) have also been described to display antineoplastic activities. These compounds were shown to be less active than cycloketones.15 A possible explanation for this difference is the larger flexibility of acyclic dienones compared to compounds with rigid piperidine rings.16 The Arbiser laboratory at Emory University has examined the effects of Tris dibenzylideneacetone dipalladium (Tris DBA) (Figure 1) in a number of studies. Tris DBA showed significant antiproliferative activity against melanoma cells, and ip injection of Tris DBA resulted in a 97% decreased tumor volume in the B16 murine melanoma model.66 Tris DBA reduces cell proliferation and induces apoptosis in multiple myeloma cells while not exerting toxicity to normal peripheral blood mononuclear cells.25 Antineoplastic activity of Tris DBA was also documented in an uveal melanoma xenograft model.67 Finally and interestingly, considering the severity of the disease, Tris DBA was found to be effective in an orthotopic model of pancreatic cancer.68 Similar results have been reported using similar compounds.69,70 Human mucoepidermoid carcinoma cell lines and tumor xenografts were found to be sensitive to DBA without reported systemic toxicity.69
We conclude from this overview that there is considerable evidence from different laboratories that dienone compounds containing the 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophore are cytotoxic to tumor cells and that a number of these compounds are less cytotoxic to nontransformed cells (Figure 2).
Other Activities of Dienones
In addition to the cytotoxic/antiproliferative activities to tumor cells, dienone compounds have been reported to have other biological effects, including anti-inflammatory and antiparasitic activity.
Anti-Inflammatory and Antiviral Activity
Zhao et al. showed that compound C12 (Figure 1) showed low cytotoxicity to macrophages but potently inhibited the stimulation of proinflammatory cytokines such as TNF-α and IL-6 by LPS.71 The activity of C12 was associated with inhibition of ERK/JNK phosphorylation and NF-κB activation.72 C12 suppresses proinflammatory cytokine production in LPS-exposed mice. The compound was administered by iv injection prior to injection of LPS. EF24 has also been reported to display anti-inflammatory activities, related to NF-κB inhibition.73,74
3,5-Bis(arylidene)-4-piperidones have been reported to inhibit the Dengue virus NS3 protease.75 Only in vitro biochemical data were reported, and the results should be viewed with some caution due to the PAINS nature of this class of compounds.
Antiparasitic Activity
Independent research groups have reported that dienones are active against the malaria parasite Plasmodium falciparum. The dibenzylidene-acetone A9 (Figure 1) was found to display an IC50 of 1.7 μM against a chloroquine-resistant isolate of P. falciparum.76 Similar results were reported using b-AP15, shown to inhibit the growth of intraerythrocytic P. falciparum parasites.77 b-AP15 induced the accumulation of lysine-48-linked polyubiquitinylated proteasome substrates in the treated parasites. The data suggested effects on the ubiquitin–proteasome system, although not at the level of the 20S proteasome or by general inhibition of deubiquitinases.77 DBA was reported to arrest the growth of Leishmania donovani parasites and induce dissipation of mitochondrial membrane potential.78
Targets Described for Dienone Compounds
3,5-Bis(arylidene)-4-piperidones and related compounds are often PAINS, and using these compounds in in vitro assays biochemical assays is potentially problematic.79 Reactive compounds will score positive also in different bioassays,80 but problems such as inhibition of enzyme activity due to coprecipitation with insoluble compounds are likely to be less common. In the following we have primarily focused on data that have been validated using cellular assays.
Inhibition of the Ubiquitin–Proteasome System (UPS)
A number of laboratories have reported that compounds with the 1,5-diaryl-3-oxo-1,4-pentadienyl pharmacophore induce the accumulation of lysine-48-linked polyubiquitinated proteins in cells.21−23,44,53,55 Lysine-48-linked polyubiquitinated proteins bind to, and are degraded by, the proteasome, and the accumulation of such proteins in cells is a characteristic feature of proteasome inhibitors. Experiments using glycerol gradient centrifugation49,57 and affinity purification of proteasomes22 have demonstrated that polyubiquitinated proteins are associated with proteasomes in exposed cells. Further evidence of proteasome inhibition includes the accumulation of a proteasome-degradable reporter (UbG76V-YFP) and proteins such as p53 and p21Cip1.22,30,37,45,55,81 These findings are interesting considering that the proteasome is a validated target of anticancer drugs and inhibitors of the 20S proteasome are in clinical use.82 The compounds discussed here generally do not affect 20S proteasome activity, and different mechanisms have been proposed to explain the phenomenon of accumulation of proteasome substrates.
Inhibition of Deubiquitinases (DUBs)
Brancolini et al. showed that compounds G5 and F6 (Figure 1) induced strong accumulation of polyubiquitinated proteins in exposed cancer cells.44 The same group83 used a biotin conjugate as a probe to identify the cellular targets of the dienone compound 2c (Figure 1). Importantly, the biotin modification did not alter the biological activity of 2c. The 2c-biotin probe was found to bind to a number of DUBs, USP1 and USP33 showing a higher affinity compared to USP14, USP18, and UCHL5.83 USP1 and USP33 were reported to be rare proteins, and their contribution to total cellular DUB activity may be limited. Other proteins were found to be modified by 2c-biotin, including AKT and a phosphorylated form of cofilin-1, and it was suggested that these targets contribute to the pharmacological effects of 2c.83 Thus, cells exposed to 2c show alterations of the cytoskeleton that may be functionally linked to cofilin-1.
Inhibition of Proteasome-Associated DUBs
D’Arcy et al. showed that b-AP15 (Figure 1) induces strong accumulation of polyubiquitin and inhibition of the degradation of a proteasome-degradable reporter in exposed cancer cells.22 Polyubiquitinated proteins were found to be associated with proteasomes22,57 (Figure 3). b-AP15 does not inhibit 20S proteasome activity but inhibits the activity of the proteasome-associated DUBs USP14 and UCHL5, both in vitro and in cells.22 Knock-down of both USP14 and UCHL5 is known to result in polyubiquitin accumulation and cytotoxicity,49,84,85 and inhibition of proteasome-associated DUBs represents an alternative strategy for inhibition of proteasome function.22 Finally, Didier et al. demonstrated that adenoviral-mediated overexpression of USP14 results in decreased sensitivity to b-AP15.81
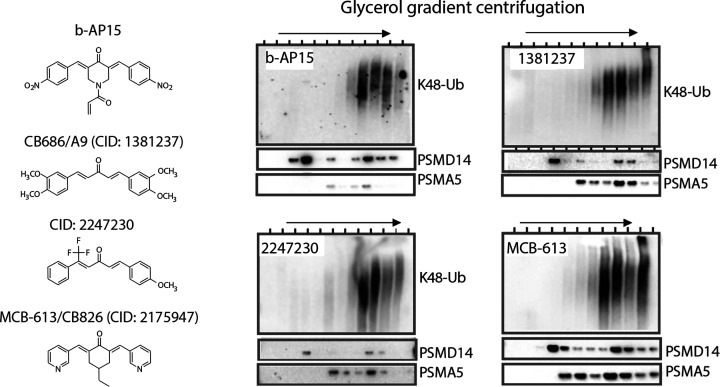
Figure 3.
b-AP15 and structurally related compound induce accumulation of polyubiquitinated proteins on proteasomes. Lysates from cells exposed to different compounds were fractionated by glycerol gradient centrifugation, and fractions were collected and processed for immunoblotting. Compounds CID1381237, CID2247230, and CID2175947 were identified in a cell-based screen for compounds stimulate accumulation of UbG76V-YFP in exposed cells.57 CID 2175947 (CB826 in ref (57)) is also known as MCB-613 and has been described to stimulate steroid receptor coactivator activity and to be preferentially cytotoxic to cancer cell lines.23 CID1381237 is also known as A976 (Figure 1) and C8.63 Data reproduced from ref (57), which licensed under CC BY 4.0.
Since the USP14 and UCHL5 DUBs are not structurally related, it would appear unlikely that a small molecule will inhibit both these enzymes. Inhibition of USP14 and UCHL5 has, however, been demonstrated in cells exposed to pharmacological concentrations of b-AP15.22,86 Most studies have shown that USP14 is inhibited at lower concentrations of b-AP15 (or VLX1570) than UCHL5.49,50,81,86,87 VLX1570 binds to USP14 and to proteasomes in surface plasmon resonance (SPR) experiments.49 Binding of VLX1570 to the 26S proteasome was inhibited by pretreatment with ubiquitin vinylsulfone (an active site-specific DUB probe88), consistent with binding to proteasome DUBs.49 Furthermore, as discussed above, USP14 binds to biotin-conjugated 2c.83 Incubation of recombinant USP14 with b-AP15 results in molecular weight shifts using MALDI.57 In these in vitro experiments, the degree of binding (i.e., one or two bound b-AP15 molecules) depends on incubation time. Somewhat surprisingly, an apparent thermal stabilization of USP14 at ∼53 °C was observed in cells exposed to b-AP15/VLX1570.49,50,57 The proteasome is known to unfold this temperature,89 and “thermal stabilization” is likely to result from a partial dissociation of USP14 from the proteasome, leading to an enrichment in the soluble fraction after heating. This effect has been observed under CETSA conditions (heating); USP14 does not dissociate from the proteasome at other conditions.49
UCHL5 appears to be less sensitive than USP14 to inhibition by dienones.49,81,87 Both G5 and RA-190 do, however, inhibit the activity of recombinant UCHL5.45,90 UCHL5 binds to a biotin-conjugated 2c probe,83 and RA-190 binds to the catalytic domain of UCHL5/UCH37.90 In the latter study, four adducts were observed after incubation of His-UCHL5 with 20-fold excess of RA-190 for 8 h at 37 °C.
RA-9 (Figure 1) was shown to induce accumulation of polyubiquitinated proteins in ovarian cancer and breast cancer cells.24,91 RA-9 inhibits proteasome-associated DUB activity, whereas total cellular DUB activity is not inhibited by it.24 Similar to b-AP15, RA-9 induces autophagic flux and inhibition of autophagy increases RA-9-mediated cytotoxicity.91
Since USP14 and UCHL5 are members of different families of DUB enzymes, it appears likely that compounds that inhibit the activities of both these enzymes will be general DUB inhibitors. Dienones have indeed been shown to inhibit DUB and also deSUMOylase activities.92 As discussed above, Ciotti et al. demonstrated widespread reactivity of compound 2c to cellular DUBs.83 Inhibition of UCHL1, UCHL5, USP2, and USP18 was demonstrated by G5 and 2c.45 These results are in apparent conflict with the results from other laboratories showing that total cellular DUB activity is not inhibited by b-AP15/RA-9 at concentrations where proteasome DUB activity is inhibited.22,24,77 A possible explanation for this discrepancy is that the contribution of different DUBs to the total cleavage activity in total cell lysates may be quite disproportionate: a limited number of DUBs may be responsible for most of the cleavage activity in the lysates, and these DUBs may be dienone-insensitive. However, at a concentration of 20 μM, VLX1570 did not show general DUB inhibitory activity on a panel of recombinant DUBs.48 It was recently reported that total DUB activity in cell lysates is inhibited by 10 and 50 μM b-AP15.93 The same authors reported inhibition of the activity of different recombinant DUBs using 100 μM b-AP15. This concentration is in the order of >100 times the IC50 and difficult to interpret. In general, results obtained using biochemical in vitro assays of enzyme activity are unreliable. Thus, findings of no inhibition of DUB enzyme activity can be incorrect due to limited compound solubility (a common problem encountered with these compounds). Conversely, findings of DUB inhibition may be incorrect due to the PAINS nature of these compounds. Considering the reported ability of dienones to inhibit DUBs of both the USP and UCH families in exposed cells, it seems reasonable to conclude that dienones are very likely to be partially selective DUB inhibitors.44
Given that dienones are partially selective DUB inhibitors, the question arises with regard to the importance of inhibition of proteasome-associated DUBs, in relation to other DUBs, for the cytotoxicity of dienones. CRISPR deletion screening has demonstrated that the majority of DUB enzymes are nonessential for cell viability.94,95 In contrast, co-transfection with siRNA to both USP14 and UCHL5 is known to result in decreased processing of proteasome substrates84,87 and decreased cell viability.85 We have, so far, been unable to generate USP14/UCHL5 double knock-out cells to test their sensitivity to b-AP15.
Binding to the Ubiquitin Receptor Rpn13
Anchoori et al.55 reported that, similar to other dienones, RA-190 (Figure 1) induces accumulation of polyubiquitin in exposed cancer cells. These authors found that RA-190 binds to Cys88 of the proteasome–ubiquitin receptor Rpn13 and suggested this to be the mechanism of action of the compound. A similar mechanism of action, involving Rpn13, has been suggested for EF24.96 Rpn13 is known to bind and to activate the DUB UCHL5, and it was subsequently shown that RA-190 does not affect the interaction of Rpn13 with UCHL5 but directly inactivates UCHL5.90
The studies of the role of Rpn13 and UCHL5 with regard to the mechanism of action of RA-190 are complicated by the fact that knock-out or knock-down of Rpn13 will lead to loss of UCHL5. In a recent study, Osei-Amponsa et al.97 showed that a cell line expressing an Rpn13 receptor lacking the Pru domain, necessary for proteasome and ubiquitin binding, is less sensitive to RA-190 compared to parental cells. Although these cells were not completely resistant to RA-190, the result demonstrates the involvement of the proteasome in conferring sensitivity to RA-190.
Organelle Stress
The robust induction of cell death by dienones (i.e., apoptosis-induction in Apaf-1 negative cells, BCL2 overexpressing cells, and p53null cells; see further below) can be interpreted in terms of major disruptions of cellular homeostasis. Many studies have shown that dienone compounds induce ER stress.98,23,99,39,57,81 This is expected considering the common finding of disturbancies of the ubiquitin–proteasome system observed by many investigators. Other types of organelle stress such as mitochondrial stress and lysosomal permeabilization have also been reported.
In their initial study of G5/F6, Brancolini and co-workers44 showed that these compounds induce mitochondrial fragmentation prior to induction of membrane permeabilization and cytochrome c release. The 20S proteasome inhibitor MG-132 induced a similar effect.44 Dienones have been suggested to induce direct mitochondrial membrane permeability transition by oxidation or cross-linking of membrane proteins.100 Zhang et al. showed that polyubiquitinated proteins become associated with mitochondrial outer membranes in cells exposed to b-AP15.52 The amounts of misfolded proteins on mitochondrial membranes correlated with decreases in oxidative phosphorylation.52 Dienone compounds, including b-AP15, were identified in a screen for compounds that induce the lysosomal pathway of apoptosis.47 Knock-down of the lysosomal enzyme cathepsin-D reduced caspase-cleavage in cells exposed to b-AP15 and also to other inhibitors of the UPS.47
An association between proteotoxic stress and organelle stress would offer an explanation for the selective sensitivity of tumor cells to dienones (due to high protein synthesis and a high production of misfolded proteins) and the robust induction of cell death.
Stimulation of Steroid Receptor Coactivators
MCB-613 (Figure 1) was identified in a cell-based screen for compounds that stimulate the activity of SRC-1 and SRC-3 steroid receptor coactivators.23 MCB-613 increases the interactions of SRC proteins with other coactivators and induces ER stress and generation of reactive oxygen species. MCB-613 was also found to induce cellular accumulation polyubiquitinated proteins but this was not considered to be the primary mechanism of action of the compound.23 MCB-613 (CB826) was subsequently identified in a screen for compounds that inhibit the processing of a proteasome-degradable reporter and was found to increase the levels of polyubiquitinated proteins on proteasomes57 (Figure 3).
Cross-Linking of Proteins
The dienone structure of 1,5-diaryl-3-oxo-1,4-pentadienes raises the possibility of protein cross-linking. Ward et al. recently raised the possibility of cross-linking being an important mechanism of action of b-AP15/VLX1570.93 The distance between the carbonyls is approximately 5.1 Å apart in 3,5-bis(arylidene)-4-piperidones,101 arguably a short distance for a chemical cross-linker. Ward et al. did show that b-AP15 can bind two molecules of GSH. The thiols of GSH would, however, be expected to be more sterically available than most cysteine thiols on larger protein structures. Ward et al. demonstrated the formation of protein dimers in cells exposed to VLX1570.93 Of particular interest was the finding of dimers of anamorsin/CIAPIN1, an apoptotic regulator, raising the possibility that functional depletion of this protein is important for the mechanism of action of VLX1570. The authors did, however, not detect direct cross-linking of anamorsin/CIAPIN1 involving thiols and the mechanism of dimer formation is unclear.
Cross-linked protein material, detected as slowly migrating material using by SDS–PAGE and Coomassie Blue staining, was observed using 10 μM b-AP15/1570.93 We also observe high MW protein complexes in AML cells exposed to 10 μM b-AP15 (Selvaraju et al., unpublished data). It should be noted that 10 μM b-AP15 is a quite high concentration (10- to 100-fold the IC50 in most cells) and that the cell death mode elicited by b-AP15 shifts from apoptosis to necrosis at concentrations of ≥5 μM.46 High molecular weight material is generally not observed using SDS–PAGE using IC50 concentrations of dienone compounds (e.g., RAMB1 (Figure 1), Figure 7A in Anchoori et al., 201154), and the molecular weights of various cellular proteins are not altered on immunoblots. However, the molecular weights of polyubiquitinated proteins gradually increase at pharmacologically more relevant concentrations of VLX1570 (0.5–1 μM, see Figure 4A). We initially interpreted the increases in molecular weight in terms of DUB inhibition leading to longer polyubiquitin chains being attached to proteins.22 An alternative interpretation is that cysteines in unfolded proteasome substrates will be more available to adduction by dienones. The formation of high molecular weight complexes and aggregation of proteins by Michael acceptors need to be further studied to evaluate mechanisms of aggregation and whether protein aggregation is coupled to cytotoxicity.
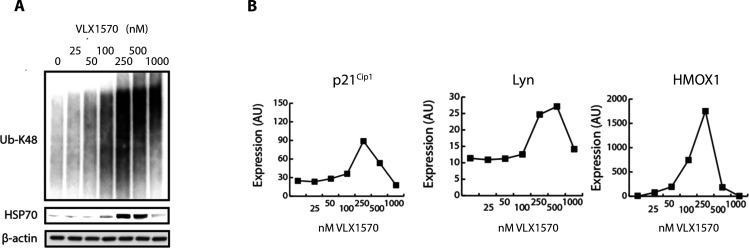
Figure 4.
Concentration-dependent responses to VLX1570. (A) Induction of polyubiquitinated proteins in RS4;11 ALL cells. Note strong increases of polyubiquitin and induction of Hsp70 at 250 nM VLX1570. (B) Increases in p21Cip1, LYN, and HMOX1. RS4;11 cells were exposed to VLX1570, and different markers were analyzed in the same extracts using a PCR-based multiplex method (OLINK, Uppsala, Sweden). Note the increases of HMOX1 at 50–100 nM VLX1570 and the decreases of all markers at high concentrations. Figure from ref (157), which licensed under CC BY 3.0.
The cystic fibrosis transmembrane conductance regulator (CFTR) can be cross-linked into dimers after exposure of cells or microsomes to curcumin.102 Cross-linking of the CFTR was observed also in the presence of high concentrations of reducing agents and using a cysteine-free CFTR construct, showing that oligomerization was not due to disulfide cross-linking. The authors suggested that cross-linking was associated with an oxidative reaction.102 Protein cross-linking has been observed using also other compounds. Stanwell et al. tested the tyrosine kinase inhibitor erbstatin and related compounds for cross-linking of cellular proteins using an assay based on boiling cells for 10 min in 2% SDS/20 mM DTT and filtering through cellulose membranes.103 Methyl 2,5-dihydroxycinnamate (Figure 1), containing a single Michael acceptor, was found to generate SDS/DTT insoluble protein material in treated cells. Cross-linking was observed at concentrations ∼5-fold the IC50 in cell proliferation assays, suggesting that it was not associated with inhibition of cell proliferation. Cross-linking was observed at both +37 °C and +4 °C, and a mechanism involving oxidation of compounds to reactive quinones that cross-link protein nucleophiles was suggested.104 Pinto et al. reported that a benzophenazine photosensitizer, OR141, induced the formation of high molecular weight complexes.105 Interestingly, OR141 was shown to induce polyubiquitinated proteins and to inhibit USP14 and UCHL5 activity in vitro. OR141 was speculated to induce disulfide bond formation, leading to formation of high molecular weight complexes.
The findings of cross-link formation by dienone compounds lead to the possibility that (i) specific proteins of importance for cellular homeostasis may be depleted and/or that (ii) cross-linked protein complexes are formed in amounts sufficient to elicit cytotoxicity. The results showing depletion of anamorsin/CIAPIN1 support the first notion (further discussed below).93 We are, however, not aware of evidence supporting the notion that protein complexes formed at pharmacologically relevant concentrations of dienones lead to cytotoxicity.
Induction of Oxidative Stress
Induction of oxidative stress by dienone compounds has been reported in a number of studies.23,29,32,36,39,81,98,106 Some of these studies use N-acetyl cysteine (NAC) as a tool to demonstrate that oxidative stress is essential for apoptosis induction. This is problematic since NAC will bind directly to the compounds, leading to their inactivation.107 Other scavengers have, however, also been used, and the results showed that oxidative stress induction is involved in triggering apoptosis (but not cell death).98 Different mechanisms have been put forward to explain the induction of oxidative stress. The enzymatic activity of thioredoxin reductase (TrxR), an enzyme essential for cellular redox-homeostasis, is dependent on the motif Gly-Cys-Sec-Gly in which Sec is selenocysteine and particularly reactive to electrophiles.108 Both b-AP15 and EF24 inhibit TrxR,87,106 presumably by binding to the selenocysteine. An alternative mechanism for oxidative stress induction has been proposed involving mitochondria. Stimulation of oxidative stress by b-AP15 was diminshed in Rhonull cells depleted of mitochondrial DNA,109 suggesting that the oxidative stress observed in cells exposed to b-AP15 is partly of mitochondrial origin.
Inhibition of Translational Elongation
Bortezomib treatment results in the induction of the unfolded protein response (UPR), including induction of phosphorylation of eIF2α.110 Induction in eIF2α phosphorylation will decrease translation and the production of proteasome substrates, constituting a protective response.110 We observed that the induction of various markers such as p21Cip1 and LYN, proteins known to be proteasome substrates,111,112 peak at concentrations of 250–500 nM VLX1570 and then decrease (Figure 4B). The lower level of induction at higher drug concentrations would be consistent with induction of the UPR, leading to decreased translation. Interestingly, however, the same pattern was observed for Hsp70 (Figure 4A). Hsp70 translation is cap-independent due to the presence of an internal ribosome binding site,113 and translation should therefore not be affected by induction of the UPR. VLX1570 was indeed shown to inhibit translation by another mechanism, at the level of translational elongation, in a recent study.114
Inhibition of translation may be relevant to the mechanism of action of VLX1570. Tumor cells are sensitive to obstructions in protein homeostasis, and an inhibitor of translational elongation (omacetaxine mepesuccinate) has been approved for treatment of chronic myeloid leukemia.115 However, it should be noted that translational inhibition is observed at slightly higher concentrations compared to proteasome inhibition.
Inhibition of Intracellular Signaling Systems
Various investigators have examined the effects of dienones on important cellular signaling molecules such as STAT transcription factors, MAP, AKT, and Src kinases. HO-3867, Tris DBA, and b-AP15 have been described to inhibit STAT3 phosphorylation.36,41,66,86 Tris DBA and NC2213 (Figure 1) inhibit Src phosphorylation.25,116 AKT has been demonstrated to be targeted by 2c using mass spectrometry,83 and AKT phosphorylation is inhibited by b-AP15 in lymphoma cells,86 by EF24 in colon cancer cells,26 by the compounds RL90 and RL91 in breast cancer cells,58 by E10 in prostate cancer cells,63 and by Tris DBA in melanoma cells.66 Furthermore, ERK phosphorylation is inhibited by b-AP1549,86 and by DBA/Tris DBA34,66 and E1063 in different tumor cells. The basal levels of GTP-loaded RAS are decreased by b-AP15.49 Whether the effects on these signaling systems are instrumental to the mechanism(s) of action of various compounds or are secondary consequences of severe stress is not well understood.
Methionine Aminopeptidase
Methionine aminopeptidases (MetAPs) remove methionine from the amino terminus of some newly synthesized proteins, essential for co-translational protein maturation processes such as myristoylation.117 MetAP2 is essential for the growth of different types of cancer cells.118 MetAP2 expression decreases in HT29 cancer cells after exposure to NC2213 (Figure 1).116 Decreased levels of Src phosphorylation were observed in parallel to the decreases in MetAP2; Src is a myristoylated oncoprotein, and a causal relationship was suggested.116
Reactivation of Mutant p53
Much interest has been focused on the possibility of reactivating mutant p53 molecules to active forms.119 It was reported that HO-3867 (Figure 1) covalently binds to mutant p53, resulting in restoration of a wild-type p53-like response.38 This mechanism would explain the selective cytotoxicity toward cancer cells by this class of compounds. The result is, however, in apparent conflict with findings using b-AP15 where antiproliferative effects and/or apoptosis induction was found to be insensitive to p53 mutational status.22,81 Furthermore, the IC50 of p53wt cell lines in the NCI60 panel to b-AP15 is 0.18 μM, and the IC50 of p53mut cell lines is 0.15 μM (p = 0.70). The discrepancy could be explained by b-AP15 and HO-3867 having different mechanisms of action (HO-3867 has not been examined for possible inhibition of proteasome function).
Effects on DNA Repair Pathways
The ataxia telangiectasia mutated (ATM) gene product is activated in cells exposed to 2cPE (a pegylated derivative of 2c).120 ATM is known to be activated by oxidative stress due to disulfide-cross-linked dimerization,121 and the stimulation of ATM signaling may therefore be a consequence of oxidative stress. ATM inhibition was found to increase 2cPE-induced cell death.120 A similar finding has been reported for EF24, showing that ATM-deficient cells are more sensitive to this compound.122 EF24 was also found to inhibit the Fanconi anemia pathway, possibly due to inhibition of IκB kinase complex.122
Analyses of Gene Expression Profiles (CMap)
The connectivity map (CMap) is a freely available and quite useful resource for initial characterization of the cellular response to small molecules. The CMap method was originally described in 2006,4 and an updated database was presented more recently.123 In the original study by Lamb et al.4 it was demonstrated that compounds sharing molecular targets induce similar molecular signatures (i.e., alterations in gene expression patterns, determined by the microarray method). Conditions were standardized with regard to the concentration of small molecules, time of exposure, and the cell line used (usually the MCF-7 breast cancer cells). The authors were able to show that CMap could be used to elucidate the mechanism of action of a natural product.4
The responses to dienones have been characterized using gene expression profiling (CMap).21,22,48,120 These experiments show high correlation (connectivity) to known inhibitors of the 20S proteasome such as MG-132.22 Induction of HSPA6 mRNA transcripts (an inducible form of the Hsp70 family of molecular chaperones) after 6 h of exposure to b-AP15 or P1 was at the level of ∼1000-fold, stronger than that observed using standard 20S proteasome inhibitors21,98 (P1 is identical to EWS-W; Figure 1). Gene expression analyses have also been performed on cells exposed to 2c-PE and showed induction of expression of chaperones, particularly transcripts encoding Hsp70 proteins.120 The response to the compound MCB-613/CB826 has also been studied.23,57 Gene set enrichment analyses (GSEA)124 showed an enrichment score of >2 for genes encoding proteins involved in protein folding.57 The mechanism underlying the stronger induction of chaperone genes by dienones in comparison to bortezomib is unclear but may at least partly be explained by the limited formation of aggresomes.81,125 The formation of aggresomes represents a mechanism for cells to escape cytotoxic effects of unfolded cellular proteins,126 and ineffective formation of these structures may lead to increased proteotoxicity.
Compound F6 (Figure 1) is in the CMap database, and the connectivity score (tau-score) between F6 and loss of function of the proteasome pathway (deletion of proteasome subunits) is +99.55 (https://clue.io/query). Tau-scores of +90 or higher are considered as strong indicators of related mechanisms of action (https://clue.io/connectopedia/connectivity_scores). It should be noted, however, that compounds that affect protein homeostasis at other levels also show strong connectivity to loss of function of the proteasome pathway, including 17-DMAG (Hsp90 inhibitor, tau-score of +95.9) and puromycin (protein synthesis inhibitor, tau-score of +97.4). Furthermore, heat shock factor-1 (HSF1), essential for induction of chaperones, is activated by electrophiles127 and electrophiles will therefore show connectivity to UPS inhibitors. The extent of this latter effect may, however, be limited since the dienone curcumin shows only weak connectivity to loss of function of the proteasome pathway (tau-score of +19.5). We conclude that CMap data are consistent with the notion that dienone compounds are proteasome inhibitors but do not provide positive proof.
Association between Inhibition of Proteasome Function and Cell Death
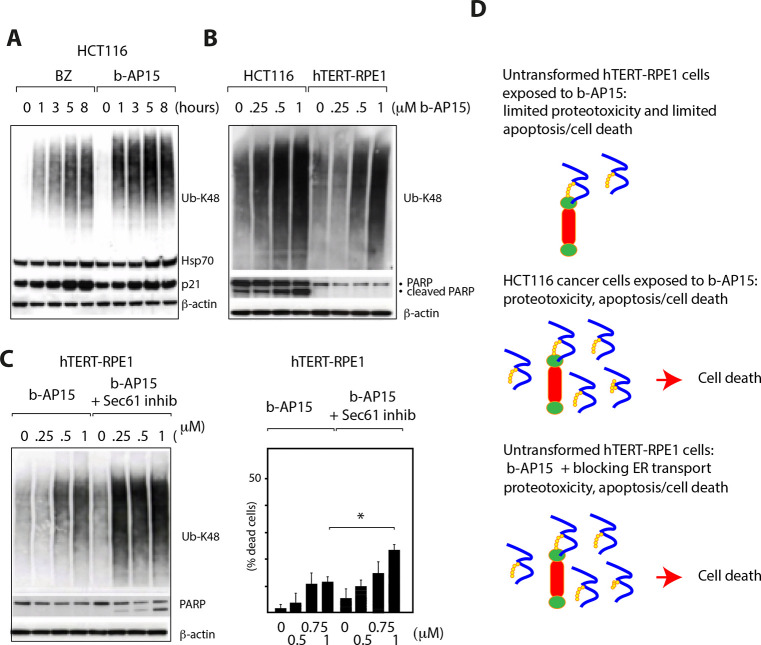
It is known that a considerable fraction of newly synthesized proteins is not folded correctly.128 High levels of protein synthesis will therefore result in an increased production of proteasome substrates.128 This would explain the therapeutic window of proteasome inhibitors, tumor cells being more active in protein synthesis, particularly the high sensitivity of myeloma cells that synthesize massive amounts of immunoglobulins. We previously reported that enhancing the levels of proteasome substrates by blocking anterograde transport of proteins through the ER increases the cytotoxic effect of b-AP15 (for details, see Figure 5).129 Interestingly, elevation of proteasome substrate levels increases the sensitivity of immortalized hTERT-RPE1 epithelial cells and normal human fibroblasts to b-AP15-induced apoptosis and to cell death (Figure 5C)129 and suggests that the difference in sensitivity of cancer cells and normal cells to b-AP15 is, at least in part, due to differences in the production of proteasome substrates (Figure 5D).
Figure 5.
Association between polyubiquitin association and cell death. (A) HCT116 colon cancer cells were exposed to 1 μM b-AP15 or 100 nM bortezomib and harvested at the indicated time points. Lysates were subjected to immunoblotting for K48-linked polyubiquitin, HSP-70B′, p21Cip1, PARP, or β-actin (loading control). (B) HCT116 cells or hTERT-RPE1 cells (immortalized epithelial cells) were exposed to different concentrations of b-AP15 for 1 h, followed by washing and incubation for 16 h in drug-free medium. Lysates were subjected to immunoblotting for K48-linked polyubiquitin, PARP and β-actin (loading control). Note the stronger accumulation of polyubiquitinated proteins and the induction of PARP cleavage in the colon cancer cells. (C) CpdA is an inhibitor of the Sec61 complex of anterograde protein export into the ER.130 Immortalized hTERT-RPE1 epithelial cells were exposed to increasing concentration of b-AP15 in the presence or absence of CpdA (“Sec61 inhib”). Note the increase in polyubiquitin accumulation, PARP cleavage, and cell death in the presence of CpdA (* p < 0.05). Parts A–C are reproduced from Sun et al.,129 which is licensed under CC BY 4.0. (D) Graphic illustration of the results. Shown are proteasomes (red, 20S; green, 19S) and polyubiquitinated proteins.
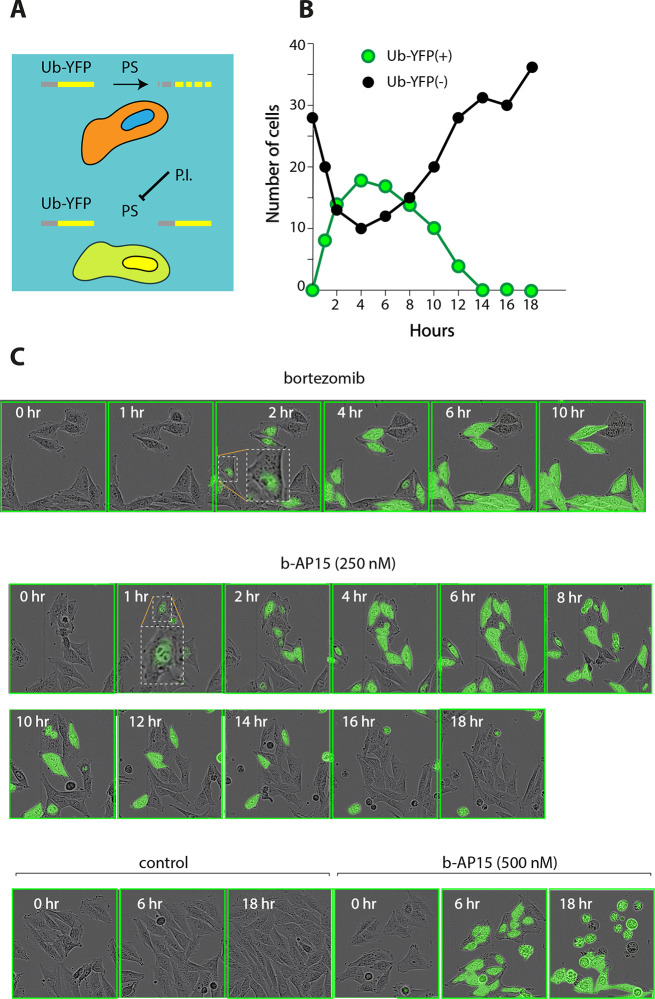
Evidence linking proteasome inhibition to cell death has been presented using cell lines expressing a proteasome-degradable reporter protein (Figure 6A).87,98 Cells can be followed after initiation of treatment and the fate of individual cells determined. Using concentrations close to IC50, it is found that cells where the proteasome reporter accumulates subsequently die (where loss of membrane integrity results in leakage of YFP from the cells), whereas cells where proteasome inhibition is not observed survive and continue to proliferate87,98 (Figure 6B). Bortezomib induces an increase in nuclear fluorescence after 2 h, followed by increases in cytoplasmic fluorescence (Figure 6C). This pattern is consistent with reports of predominant presence of 26S proteasomes in the nucleus.131 Exposure to b-AP15 results in the same pattern of initial nuclear fluorescence (Figure 6D), first observed after 1 h. The difference in onset can be explained by the more rapid uptake of b-AP15 into cells compared to bortezomib.87,132
Figure 6.
Proteasome inhibition induced by b-AP15 is associated with cell death. (A) A reporter protein (UbG76V-YFP) is degraded by the proteasome (PS) in human melanoma cells. Proteasome inhibition (PI) leads to accumulation of the reporter protein in the cells.133 (B) Tracing individual cells expressing UbG76V-YFP following exposure to 250 nM b-AP15. Cells become UbG76V-YFP positive after 1–4 h, showing accumulation of the proteasome reporter. YFP-positive cells subsequently round up and the YFP leaks out from the cells after membrane disintegration. In contrast, cells that remain YFP-negative at 4 h of exposure survive and continue to divide. Results were obtained using an IncuCyte instrument (see refs (87) and (98)). (C) Time-lapse recordings of UbG76V-YFP-expressing human melanoma cells after addition of compounds. Upper panel: Human melanoma cells expressing UbG76V-YFP were exposed to 100 nM bortezomib and traced for 10 h. Notice the appearance of YFP fluorescence in cell nuclei at 2 h and later appearance in the cytoplasm (one cell highlighted at 2 h). Middle panel: Cells were exposed to 250 nM b-AP15. Note the similar pattern of initial nuclear and later cytoplasmic fluorescence (one cell highlighted at 1 h), similar to the pattern observed using bortezomib. Also note that cells either become positive and die or remain completely negative and survive. Lower panel, left: control. Vehicle-treated control cells remain YFP-negative and proliferate. Lower panel, right: >90% of the cells exposed to 500 nM b-AP15 become YFP-positive after 6 h, and most of these cells have rounded up at 18 h, some have lost membrane integrity and the YFP content. Original data were published in refs (87) and (98).
The accumulation of the UbG76V-YFP reporter in cells could be argued to be due to cross-linking and aggregation of the reporter protein and an inaccessibility of aggregated proteins to proteasomal degradation. We find this scenario unlikely for a number of reasons: (i) the same pattern of initial nuclear accumulation of UbG76V-YFP is observed in b-AP15- and bortezomib-treated cells; (ii) the phenomenon of “all-or-nothing” in terms of UbG76V-YFP positivity; it is difficult to understand how chemical cross-linking of UbG76V-YFP, leading to aggregation, will occur in some cells, whereas no (or undetectable) cross-linking and aggregation will occur in neighboring cells. This “all-or-nothing” phenomenon is identical in cells exposed to b-AP15 and bortezomib and is likely to reflect differences in the thresholds of proteasome degradation capacity and/or generation of proteasome substrates in different cells; (iii) we have not observed aggregation of UbG76V-YFP by immunoblotting in b-AP15-exposed cells using pharmacologically relevant concentrations;129 (iv) cross-linking of cellular proteins93 (see above) could be expected to result in general toxicity, not consistent with the tumor cell selectivity of dienones and the relative insensitivity of noncancerous cells to b-AP-15 and other dienones.
The notion that inhibition of the UPS is of importance for cytotoxicity of dienones is supported by experiments involving genetic manipulation: (i) deletion of Rpn13 and deletion of the Pru-domain of Rpn13 lead to a decreased (but not totally lost) sensitivity to RA-190;97 (ii) overexpression of USP14 results in decreased sensitivity to b-AP15;81 (iii) knock-out of SRC-3, the protein product of which has been suggested to be upstream of proteasome inhibition, leads to a decreased induction of DDIT3/CHOP and other stress-related markers.23
Studies of Binding and Degradation of Electrophile-Adducted Proteins
Some information is available with regard to the binding of dienones to the cellular proteome and the subsequent fate of adducted proteins. It was recently reported that 44 proteins were covalently adducted by VLX1570, including the anamorsin/CIAPIN1 apoptosis regulator.93 Studies have been performed on 4-hydroxy-2-nonenal (HNE), an α,β-unsaturated aldehyde that shows reactivity to proteomic cysteines,134 and the endogenous electrophile 15-deoxy-Δ12,14-prostaglandin-J2 (PG-J2, a cross-conjugated α,β-unsaturated dienone). HNE is known to be selective, and only a limited number of nucleophilic residues are carbonylated within each protein.135 The reactivities of different residues appear to be affected by the protein environment.135,136 PG-J2 has been demonstrated to bind ∼350 proteins, the largest group related to metabolic processes involved in protein modification, protein folding, translation, and proteolysis.137
Thioredoxin reductase (TrxR) is expected to be a common target of cytotoxic electrophiles due to its nucleophilic active site Sec residue, present in an unstructured portion of the enzyme.138 We recently evaluated 141 cytotoxic α,β-unsaturated compounds for their ability to inhibit proteasome function and their ability to inhibit TrxR. Surprisingly, whereas 28 compounds were found to inhibit proteasome function, only 20 could be shown to inhibit TrxR.57 One of the compounds that was found not to inhibit TrxR was the dienone compound CB826/MCB-613. The finding was unexpected both with regard to the presence of the nucleophilic Sec residue in the TrxR enzyme and with regard to the PAINS properties of many of the compounds. Furthermore, the findings did not show a direct relationship between electrophilicity of the compounds and their cytotoxicity.57
The inhibition of proteasome DUB activity and of USP14 was reported to be reversible under in vitro conditions.22,87 In surface plasmon resonance experiments (SPR), VLX1570 rapidly associates and subsequently dissociates from proteasomes.49 We have subsequently found that extended incubation of 19S proteasomes with b-AP15 (20 min at 37 °C) results in reduced recovery of (∼60%) of the DUB activity after dilution.57 We interpret these results to suggest that b-AP15 recognizes proteasome-associated DUBs by noncovalent interactions, increasing the residence time to facilitate subsequent covalent binding.
Is the strong and rapid accumulation of polyubiquitinated proteins in cells exposed to dienones a secondary consequence of adduct formation and aggregation? An early study suggested that HNE-modified proteins were significantly ubiquitinated and showed that proteasome inhibitors suppressed the elimination of such proteins.139 A later study showed that the elimination of HNE-modified proteins in cells was not stabilized by proteasome inhibitors, whereas the lysosome inhibitor chloroquine prevented their loss.140 In a study of HNE-induced degeneration of neurons, Zhang et al. observed an abnormal accumulation of both K48- and K63-linked polyubiquitinated proteins.141 The increases in K48-linked polyubiquitinated proteins were, however, quite modest, approximately 2-fold by 24 h. A particularly interesting study using mass spectrometry was published by Yang et al.134 These authors showed that the alkylation of some cysteine residues by HNE was stable over time, whereas and interestingly the modification of other cysteines decreased rapidly after removal of HNE. These findings suggest the existence of a repair mechanism for HNE adducts in cells but is not consistent with alkylated proteins being removed by mechanisms of global protein degradation. Although the literature is conflicting with regard to mechanisms of degradation of HNE-adducted proteins, we are unaware of any study demonstrating accumulation of K48-linked polyubiquitinated proteins similar to what is observed using b-AP15 (Figures 4 and 5). Since both the kinetics and amplitude of the increases in polyubiquitin are similar to those observed using the 20S proteasome inhibitor bortezomib, our interpretation is that they are likely due to a direct inhibitory effect of proteasome function.
Pharmacology and Toxicity
Measurement of HO-3867 uptake in tissue cultured cells showed maximal uptake within 60 min followed by a decline of intracellular levels.142 HO-3867 could still be detected after 72 h in cells exposed for 1 h followed by change of medium.142 We have reported very similar results using b-AP15, demonstrating a rapid uptake from the culture medium and enrichment of the compound in exposed cells.87 Intracellular enrichment of drugs is not uncommon and is also observed for the 20S proteasome inhibitor bortezomib.132 The underlying mechanism for the rapid uptake and enrichment of dienone compounds is unclear, but we have observed that cellular uptake was impaired by 30 min of pretreatment of cells with a relatively low concentration of N-ethylmaleimide (10 μM). One interpretation of this result is that the uptake is driven by the availability of intracellular thiol groups. Alternatively, uptake is dependent on a N-ethylmaleimide-sensitive transporter.
The higher potency of b-AP15 and VLX1570 in intact cells compared to its potency using in vitro assays for USP14 and UCHL5 has been recently discussed.93 Intracellular enrichment explains the findings by different investigators of USP14 activity being inhibited in cells exposed to relevant concentrations (0.1–1 μM) of b-AP15/VLX1570.22,48−50,81,86,87
The relevant drug concentrations that are to be used in in vitro studies have been an enigma in cancer drug research since compounds are often reactive and exhibit polypharmacology. Cisplatin can be used to illustrate this problem. Cisplatin is known to cross-link DNA, representing the major mechanism of action.143 The cisplatin molecule will also react with proteins, in particular with thiols, and a number of other targets will become relevant using suprapharmacological concentrations of the drug. Cisplatin-induced acute apoptotic cell death, studied by many groups (PubMed search September 2020, cisplatin + apoptosis + mechanism: ∼4000 hits), may be of limited importance for clinical drug sensitivity since concentrations of >10-fold the IC50 are required to induce this phenomenon.144
A number of investigators have demonstrated that dienone compounds display considerable higher cytotoxicity to tumor cells compared to noncancerous cells.18−26,38,42,53,59,60,129 Some of these studies have presented IC50 values, and these are summarized in Figure 2 and Supplementary Table 1. These results, using in vitro cultured cells, are interesting from a mechanistic point of view but are difficult to extrapolate to toxicity in living organisms. It was recently reported that b-AP15/VLX1570 shows similar cytotoxicities to cancer cell lines and HUVECs (human vascular endothelial cells) and, based on this observation, it was suggested that these compounds are generally cytotoxic.93 The sensitivity of HUVECs to b-AP15/VLX1570 is consistent with reports from other studies of dienones. In one study the IC50 of EF24 toward HUVECs was reported to be 2.6 μM, lower in comparison to two different liver cancer cell lines (3.8 and 4.4 μM).30 In another study, the IC50 of EF24 toward HUVECs was reported to be 1.1 μM, compared to 0.2 μM for leukemia cells, 0.3 μM for colon cancer cells, 0.4 μM for prostate cancer cells, and 1.4 μM for melanoma cells.15 MCB-613 showed IC50 values of 3–6 μM in cancer cell lines, whereas no effect was observed on the proliferation of primary hepatocytes or fibroblasts at these concentrations.23 HUVECs were, however, sensitive to high doses of MCB-613.23 On the basis of the sensitivity of endothelial cells, dienones have been advocated as angiogenesis inhibitors.145 Whether studies of the in vitro sensitivity of HUVECs predict the antiangiogenic potential of small molecules or whether HUVEC sensitivity can be used to argue that dienones are generally cytotoxic can be debated.
Multiple reports that demonstrated preclinical antitumor activity of dienones did not report animal toxicity (see above). In the specific case of VLX1570, preclinical use requires oil/detergents for formulation, leading to complications in the evaluation of toxicology due to species differences in the tolerance to such formulations.146 A phase 1 study has been performed using VLX1570 in a formulation based on polyethylene glycol, polyoxyethylated castor oil, and polysorbate 80 (PCT) formulation.147 The study used an accelerated protocol, and pulmonary toxicity was encountered at 1.2 mg/kg.147 Pulmonary toxicity has also been observed with bortezomib,148 and it was found that the PCT formulation enhances this type of adverse event by bortezomib and VLX1570.147 These problems will hopefully be possible to resolve using prodrugs that do not require oil/detergent for administration.
Cell Death Mode Induced by Dienones
A number of different cell death modes have been described,149 many of which may be induced by anticancer drugs. Describing the cell death mode being elicited by a particular compound is of importance both for understanding mechanisms of action and for understanding potential mechanisms of resistance.
The death mode induced by G5 has been studied in some detail.150−152 G5-induced apoptosis requires the Bcl-2 family proteins Bax and Bak.152 However, and in distinction to bortezomib, G5 is able to kill apoptosis-resistant cells deficient in Bax and Bak by activating a necrotic response.152 In fact, the kinetics of cell death observed in G5-exposed cultures is indistinguishable using wild-type cells and Bax/Bak defective cells.152 The morphology of cells exposed to G5 is characterized by massive cytoplasmic vacuolization.152 Interestingly, a similar phenomenon of vacuolization was reported in cells exposed to MCB-61323 and has also been demonstrated for b-AP15-treated cells.52 The death mode induced by MCB-613 was interpreted as paroptosis,23 a nonapoptotic form of cell death characterized by massive cytoplasmic vacuolization due to dilation of the ER and mitochondria,153 and it is possible that all dienone compounds induce this cell death mode.
b-AP15 induces various stress responses, including ER stress and oxidative stress.98 Inhibition of these stress responses impairs apoptosis but does not prevent cell death.98 It was recently demonstrated that VLX1570 induces dimerization of the apoptosis inhibitor anamorsin/CIAPIN1, leading to a depletion of the active monomeric form.93 This finding raises the possibility that inactivation of anamorsin contributes to apoptosis induction by VLX1570. However, the findings that dienones are able to kill apoptosis-resistant cells, possibly by paroptosis, lead to questions with regard to the importance of anamorsin depletion for cell death by VLX1570 and other dienones.
Conclusions
Dienones such as b-AP15/VLX1570 have obvious chemical liabilities leading to predictions of engagement of multiple targets and of general toxicity. Dienones have indeed been demonstrated to engage a number of target proteins in cells. These different targets will not be equivalent in terms of the pharmacological response that will be elicited and will not be engaged at the same drug concentrations (Figure 7A,B). Spliceosome, ribosome, and proteasome functions represent major vulnerabilities of tumor cells;154 evidence reviewed here suggests that effects on the UPS can be linked to the cytotoxicity of dienones. The exact mechanism(s) by which dienones elicit these effects, by inhibition of proteasome-associated DUBs, by effects on Rpn13, and/or by additional components, is (are) not clear at present.
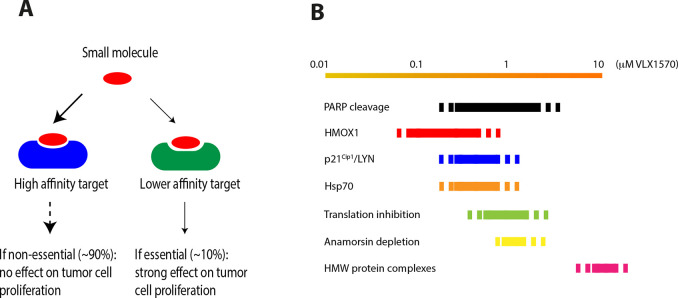
Figure 7.
(A) Target affinity does not necessarily correlate to mechanism of action. A major fraction of cellular genes (∼90%) have been found to be nonessential for proliferation of tumor cells in vitro.155 Small molecules that bind to the protein products of these genes are therefore unlikely to affect tumor cell proliferation. In contrast, low affinity binding to essential proteins may result in strong inhibition of proliferation. An example is microtubule dynamics, known to be sensitive to low concentrations of small molecules156 and for which off-target effects may occur for various compounds when used in screens at 5–10 μM. (B) Illustration of responses to VLX1570 reported in different studies.93,114,157 HMOX1 is induced at lower concentration of b-AP15/VLX1570 than other markers (see Figure 4). Nrf-2/KEAP is targeted by electrophiles, and the Nrf-2 system is likely to be particularly sensitive to electrophiles.158 Increases of polyubiquitinated proteins, Hsp70 chaperones, and proteasome substrates such p21Cip1 are observed at somewhat higher concentrations of VLX1570; increased levels of Hsp70 are also observed at these concentrations (Figure 4).157 Inhibition of translation is observed at increasing concentrations of VLX1570,114,157 leading to a blunted response of Hsp70, HMOX1, p21Cip1, and LYN expression (Figure 4). Depletion of anamorsin/CIAPIN1 was reported to occur at concentrations of 0.5–1 μM 93 and the formation of higher molecular weight protein complexes at 10 μM 93 (using HeLa cells).
Despite the chemical liabilities of the class of compounds reviewed here, a substantial number of studies have shown that these compounds show preferential toxicity to tumor cells and show activity in in vivo tumor models. The ability of dienones to kill apoptosis-resistant cells and the limited development of resistance have stimulated considerable interest in these molecules. These compounds are unsuitable as, and should not be used as, biological probes. They could possibly be developed into anticancer drugs, however, if soluble prodrugs such as 2c-PE will be found to be safe and effective.
Glossary
Abbreviations Used
- AKT
protein kinase B
- ATM
ataxia telangiectasia mutated
- CETSA
cellular thermal shift assay
- CFTR
cystic fibrosis transmembrane conductance regulator
- CIAPIN1
cytokine-induced inhibitor of apoptosis
- CMap
connectivity map
- DBA
dibenzylidene-acetone
- DDIT3/CHOP
DNA damage-inducible transcript 3/C/EBP homologous protein
- DUB
deubiquitinase
- DTT
dithiotreitol
- ERK
extracellular signal-regulated
- HMOX1
heme oxygenase 1
- HNE
4-hydroxy-2-nonenal
- HPV
human papilloma virus
- Hsp
heat shock protein
- hTERT
human telomerase reverse transcriptase
- HUVEC
human vascular endothelial cell
- JNK
c-Jun N-terminal kinase
- mRNA
messenger RNA
- LYN
Src family tyrosine kinase LYN
- MALDI
matrix-assisted laser desorption/ionization
- MAP kinase
mitogen activated protein kinase
- NF-κB
nuclear factor κB
- PAINS
pan-assay interference compounds
- PARP
poly ADP-ribose polymerase
- PG-J2
prostaglandin-J2
- Rpn
regulatory particle of non-ATPase
- SDS–PAGE
sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- Sec61
human secretory translocase
- Src
sarcoma oncogene
- SRC
steroid receptor coactivator
- STAT
signal transducers and activators of transcription
- TNF
tumor necrosis factor
- TrxR
thioredoxin reductase
- UPR
unfolded protein response
- USP
ubiquitin specific peptidase
- UCH
ubiquitin C-terminal hydrolase
- UPS
ubiquitin proteasome system
- YFP
yellow fluorescent protein
Biographies
Martina Bazzaro is an Associate Professor at the University of Minnesota. She received her Ph.D. in Pharmaceutical Chemistry from the University of Ferrara, Italy, became a Visiting Fellow at the University of Lausanne, Switzerland, and carried out postdoctoral research at the Karolinska Institute in Sweden and Johns Hopkins Medical School. She has 15 years’ experience in cancer biology, with a focus on metabolic pathways including deubiquitinases and the proteasome. She has published extensively in the area of medicinal chemistry and cancer biology and serves as Editor in Chief of the Journal of Gynecology and Obstetrics and as Topic Editor for Cancers. She is funded by and serves as reviewer for national and international grant agencies including the National Institutes of Health and the Department of Defense.
Stig Linder is an Emeritus Professor in Pharmacology at the Department of Biomedical and Clinical Sciences, Linköping, Sweden, and in Cancer Pharmacology at the Department of Oncology and Pathology, Karolinska Institute, Sweden. He received his Ph.D. degree in Cell Biology from the Karolinska Institute and carried out postdoctoral research at Yale University School of Medicine, CT. He is an Adjunct Professor (Docent) in Medical Microbiology at Uppsala University and in Experimental Oncology at the Karolinska Institute. He has published original research papers, reviews, and book chapters in various fields of biology. He has been a scientific advisor for enterprises in the biotechnology area and holds several patents. He is a member of the research board of the Swedish Cancer Society (Cancerfonden).
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jmedchem.0c00812.
Table 1 listing selective antiproliferative activity to tumor cells, as verified by other laboratories (PDF)
This work was supported by Department of Defense Ovarian Cancer Research Program Grant OC160377, the Minnesota Ovarian Cancer Alliance, the Randy Shaver Cancer Research Funds and the NIH grant NIGMS R01-GM130800 to Martina Bazzaro.
The authors declare no competing financial interest.
Supplementary Material
References
- Swinney D. C.; Anthony J. How were new medicines discovered?. Nat. Rev. Drug Discovery 2011, 10, 507–519. 10.1038/nrd3480. [DOI] [PubMed] [Google Scholar]
- Mayr L. M.; Bojanic D. Novel trends in high-throughput screening. Curr. Opin. Pharmacol. 2009, 9, 580–588. 10.1016/j.coph.2009.08.004. [DOI] [PubMed] [Google Scholar]
- Schenone M.; Dancik V.; Wagner B. K.; Clemons P. A. Target identification and mechanism of action in chemical biology and drug discovery. Nat. Chem. Biol. 2013, 9, 232–240. 10.1038/nchembio.1199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamb J.; Crawford E. D.; Peck D.; Modell J. W.; Blat I. C.; Wrobel M. J.; Lerner J.; Brunet J. P.; Subramanian A.; Ross K. N.; Reich M.; Hieronymus H.; Wei G.; Armstrong S. A.; Haggarty S. J.; Clemons P. A.; Wei R.; Carr S. A.; Lander E. S.; Golub T. R. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. 10.1126/science.1132939. [DOI] [PubMed] [Google Scholar]
- Jafari R.; Almqvist H.; Axelsson H.; Ignatushchenko M.; Lundback T.; Nordlund P.; Martinez Molina D. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 2014, 9, 2100–2122. 10.1038/nprot.2014.138. [DOI] [PubMed] [Google Scholar]
- Lee R. F. S.; Chernobrovkin A.; Rutishauser D.; Allardyce C. S.; Hacker D.; Johnsson K.; Zubarev R. A.; Dyson P. J. Expression proteomics study to determine metallodrug targets and optimal drug combinations. Sci. Rep. 2017, 7, 1590. 10.1038/s41598-017-01643-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaetani M.; Sabatier P.; Saei A. A.; Beusch C. M.; Yang Z.; Lundstrom S. L.; Zubarev R. A. Proteome integral solubility alteration: a high-throughput proteomics assay for target deconvolution. J. Proteome Res. 2019, 18, 4027–4037. 10.1021/acs.jproteome.9b00500. [DOI] [PubMed] [Google Scholar]
- da Silva-Diz V.; Lorenzo-Sanz L.; Bernat-Peguera A.; Lopez-Cerda M.; Munoz P. Cancer cell plasticity: impact on tumor progression and therapy response. Semin. Cancer Biol. 2018, 53, 48–58. 10.1016/j.semcancer.2018.08.009. [DOI] [PubMed] [Google Scholar]
- Wagle N.; Emery C.; Berger M. F.; Davis M. J.; Sawyer A.; Pochanard P.; Kehoe S. M.; Johannessen C. M.; Macconaill L. E.; Hahn W. C.; Meyerson M.; Garraway L. A. Dissecting therapeutic resistance to RAF inhibition in melanoma by tumor genomic profiling. J. Clin. Oncol. 2011, 29, 3085–3096. 10.1200/JCO.2010.33.2312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Wekken A. J.; Saber A.; Hiltermann T. J.; Kok K.; van den Berg A.; Groen H. J. Resistance mechanisms after tyrosine kinase inhibitors afatinib and crizotinib in non-small cell lung cancer, a review of the literature. Crit. Rev. Oncol./Hematol. 2016, 100, 107–116. 10.1016/j.critrevonc.2016.01.024. [DOI] [PubMed] [Google Scholar]
- Hossain M.; Das U.; Dimmock J. R. Recent advances in alpha, beta-unsaturated carbonyl compounds as mitochondrial toxins. Eur. J. Med. Chem. 2019, 183, 111687. 10.1016/j.ejmech.2019.111687. [DOI] [PubMed] [Google Scholar]
- Dimmock J. R.; Arora V. K.; Wonko S. L.; Hamon N. W.; Quail J. W.; Jia Z.; Warrington R. C.; Fang W. D.; Lee J. S. 3,5-Bis-benzylidene-4-piperidones and related compounds with high activity towards P388 leukemia cells. Drug Des. Delivery 1990, 6, 183–194. [PubMed] [Google Scholar]
- Dimmock J. R.; Arora V. K.; Quail J. W.; Pugazhenthi U.; Allen T. M.; Kao G. Y.; De Clercq E. Cytotoxic evaluation of some 3,5-diarylidene-4-piperidones and various related quaternary ammonium compounds and analogs. J. Pharm. Sci. 1994, 83, 1124–1130. 10.1002/jps.2600830811. [DOI] [PubMed] [Google Scholar]
- El-Subbagh H. I.; Abu-Zaid S. M.; Mahran M. A.; Badria F. A.; Al-Obaid A. M. Synthesis and biological evaluation of certain alpha, beta-unsaturated ketones and their corresponding fused pyridines as antiviral and cytotoxic agents. J. Med. Chem. 2000, 43, 2915–2921. 10.1021/jm000038m. [DOI] [PubMed] [Google Scholar]
- Adams B. K.; Ferstl E. M.; Davis M. C.; Herold M.; Kurtkaya S.; Camalier R. F.; Hollingshead M. G.; Kaur G.; Sausville E. A.; Rickles F. R.; Snyder J. P.; Liotta D. C.; Shoji M. Synthesis and biological evaluation of novel curcumin analogs as anti-cancer and anti-angiogenesis agents. Bioorg. Med. Chem. 2004, 12, 3871–3883. 10.1016/j.bmc.2004.05.006. [DOI] [PubMed] [Google Scholar]
- Das U.; Sharma R. K.; Dimmock J. R. 1,5-diaryl-3-oxo-1,4-pentadienes: a case for antineoplastics with multiple targets. Curr. Med. Chem. 2009, 16, 2001–2020. 10.2174/092986709788682218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel M. R.; Dimmock J. R.; Talele T. T. CoMFA and CoMSIA studies on 1,3-bis(benzylidene)-3,4-dihydro-1H-naphthalen-2-one, 2,6-bis(benzylidene)cyclohexanone, and 3,5-bis(benzylidene)-4-piperidone series of cytotoxic compounds. J. Chem. Inf. Model. 2007, 47, 2110–2123. 10.1021/ci700210z. [DOI] [PubMed] [Google Scholar]
- Pati H. N.; Das U.; Das S.; Bandy B.; De Clercq E.; Balzarini J.; Kawase M.; Sakagami H.; Quail J. W.; Stables J. P.; Dimmock J. R. The cytotoxic properties and preferential toxicity to tumour cells displayed by some 2,4-bis(benzylidene)-8-methyl-8-azabicyclo[3.2.1] octan-3-ones and 3,5-bis(benzylidene)-1-methyl-4-piperidones. Eur. J. Med. Chem. 2009, 44, 54–62. 10.1016/j.ejmech.2008.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das U.; Doroudi A.; Inci Gul H.; Pati H. N.; Kawase M.; Sakagami H.; Chu Q.; Stables J. P.; Dimmock J. R. Cytotoxic 2-benzylidene-6-(nitrobenzylidene)cyclohexanones which display substantially greater toxicity for neoplasms than non-malignant cells. Bioorg. Med. Chem. 2010, 18, 2219–2224. 10.1016/j.bmc.2010.01.069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santiago-Vazquez Y.; Das U.; Varela-Ramirez A.; Baca S. T.; Ayala-Marin Y.; Lema C.; Das S.; Baryyan A.; Dimmock J. R.; Aguilera R. J. Tumor-selective cytotoxicity of a novel pentadiene analogue on human leukemia/ lymphoma cells. Clin. Cancer Drugs 2016, 3, 138–146. 10.2174/2212697X03666160830165250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Contreras L.; Calderon R. I.; Varela-Ramirez A.; Zhang H. Y.; Quan Y.; Das U.; Dimmock J. R.; Skouta R.; Aguilera R. J. Induction of apoptosis via proteasome inhibition in leukemia/lymphoma cells by two potent piperidones. Cell. Oncol. 2018, 41, 623–636. 10.1007/s13402-018-0397-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Arcy P.; Brnjic S.; Olofsson M. H.; Fryknäs M.; Lindsten K.; De Cesare M.; Perego P.; Sadeghi B.; Hassan M.; Larsson R.; Linder S. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat. Med. 2011, 17, 1636–1640. 10.1038/nm.2536. [DOI] [PubMed] [Google Scholar]
- Wang L.; Yu Y.; Chow D. C.; Yan F.; Hsu C. C.; Stossi F.; Mancini M. A.; Palzkill T.; Liao L.; Zhou S.; Xu J.; Lonard D. M.; O’Malley B. W. Characterization of a steroid receptor coactivator small molecule stimulator that overstimulates cancer cells and leads to cell stress and death. Cancer Cell 2015, 28, 240–252. 10.1016/j.ccell.2015.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coughlin K.; Anchoori R.; Iizuka Y.; Meints J.; MacNeill L.; Vogel R. I.; Orlowski R. Z.; Lee M. K.; Roden R. B.; Bazzaro M. Small-molecule RA-9 inhibits proteasome-associated DUBs and ovarian cancer in vitro and in vivo via exacerbating unfolded protein responses. Clin. Cancer Res. 2014, 20, 3174–3186. 10.1158/1078-0432.CCR-13-2658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de la Puente P.; Azab F.; Muz B.; Luderer M.; Arbiser J.; Azab A. K. Tris DBA palladium overcomes hypoxia-mediated drug resistance in multiple myeloma. Leuk. Lymphoma 2016, 57, 1677–1686. 10.3109/10428194.2015.1099645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramaniam D.; May R.; Sureban S. M.; Lee K. B.; George R.; Kuppusamy P.; Ramanujam R. P.; Hideg K.; Dieckgraefe B. K.; Houchen C. W.; Anant S. Diphenyl difluoroketone: a curcumin derivative with potent in vivo anticancer activity. Cancer Res. 2008, 68, 1962–1969. 10.1158/0008-5472.CAN-07-6011. [DOI] [PubMed] [Google Scholar]
- Starcevic S. L.; Diotte N. M.; Zukowski K. L.; Cameron M. J.; Novak R. F. Oxidative DNA damage and repair in a cell lineage model of human proliferative breast disease (PBD). Toxicol. Sci. 2003, 75, 74–81. 10.1093/toxsci/kfg154. [DOI] [PubMed] [Google Scholar]
- Gabriel M.; Balle D.; Bigault S.; Pornin C.; Getin S.; Perraut F.; Block M. R.; Chatelain F.; Picollet-D’hahan N.; Gidrol X.; Haguet V. Time-lapse contact microscopy of cell cultures based on non-coherent illumination. Sci. Rep. 2015, 5, 14532. 10.1038/srep14532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams B. K.; Cai J.; Armstrong J.; Herold M.; Lu Y. J.; Sun A.; Snyder J. P.; Liotta D. C.; Jones D. P.; Shoji M. EF24, a novel synthetic curcumin analog, induces apoptosis in cancer cells via a redox-dependent mechanism. Anti-Cancer Drugs 2005, 16, 263–275. 10.1097/00001813-200503000-00005. [DOI] [PubMed] [Google Scholar]
- Liu H.; Liang Y.; Wang L.; Tian L.; Song R.; Han T.; Pan S.; Liu L. In vivo and in vitro suppression of hepatocellular carcinoma by EF24, a curcumin analog. PLoS One 2012, 7, e48075 10.1371/journal.pone.0048075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertazza L.; Barollo S.; Mari M. E.; Faccio I.; Zorzan M.; Redaelli M.; Rubin B.; Armanini D.; Mian C.; Pezzani R. Biological effects of EF24, a curcumin derivative, alone or combined with mitotane in adrenocortical tumor cell lines. Molecules 2019, 24, 2202. 10.3390/molecules24122202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He G.; Feng C.; Vinothkumar R.; Chen W.; Dai X.; Chen X.; Ye Q.; Qiu C.; Zhou H.; Wang Y.; Liang G.; Xie Y.; Wu W. Curcumin analog EF24 induces apoptosis via ROS-dependent mitochondrial dysfunction in human colorectal cancer cells. Cancer Chemother. Pharmacol. 2016, 78, 1151–1161. 10.1007/s00280-016-3172-x. [DOI] [PubMed] [Google Scholar]
- Yin D. L.; Liang Y. J.; Zheng T. S.; Song R. P.; Wang J. B.; Sun B. S.; Pan S. H.; Qu L. D.; Liu J. R.; Jiang H. C.; Liu L. X. EF24 inhibits tumor growth and metastasis via suppressing NF-kappaB dependent pathways in human cholangiocarcinoma. Sci. Rep. 2016, 6, 32167. 10.1038/srep32167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin C.; Tu C.; Ma Y.; Ye P.; Shao X.; Yang Z.; Fang Y. Curcumin analog EF24 induces apoptosis and downregulates the mitogen activated protein kinase/extracellular signal-regulated signaling pathway in oral squamous cell carcinoma. Mol. Med. Rep. 2017, 16, 4927–4933. 10.3892/mmr.2017.7189. [DOI] [PubMed] [Google Scholar]
- Xie X.; Tu J.; You H.; Hu B. Design, synthesis, and biological evaluation of novel EF24 and EF31 analogs as potential I-kappaB kinase beta inhibitors for the treatment of pancreatic cancer. Drug Des., Dev. Ther. 2017, 11, 1439–1451. 10.2147/DDDT.S133172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selvendiran K.; Ahmed S.; Dayton A.; Kuppusamy M. L.; Tazi M.; Bratasz A.; Tong L.; Rivera B. K.; Kalai T.; Hideg K.; Kuppusamy P. Safe and targeted anticancer efficacy of a novel class of antioxidant-conjugated difluorodiarylidenyl piperidones: differential cytotoxicity in healthy and cancer cells. Free Radical Biol. Med. 2010, 48, 1228–1235. 10.1016/j.freeradbiomed.2010.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selvendiran K.; Tong L.; Bratasz A.; Kuppusamy M. L.; Ahmed S.; Ravi Y.; Trigg N. J.; Rivera B. K.; Kalai T.; Hideg K.; Kuppusamy P. Anticancer efficacy of a difluorodiarylidenyl piperidone (HO-3867) in human ovarian cancer cells and tumor xenografts. Mol. Cancer Ther. 2010, 9, 1169–1179. 10.1158/1535-7163.MCT-09-1207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madan E.; Parker T. M.; Bauer M. R.; Dhiman A.; Pelham C. J.; Nagane M.; Kuppusamy M. L.; Holmes M.; Holmes T. R.; Shaik K.; Shee K.; Kiparoidze S.; Smith S. D.; Park Y. A.; Gomm J. J.; Jones L. J.; Tomas A. R.; Cunha A. C.; Selvendiran K.; Hansen L. A.; Fersht A. R.; Hideg K.; Gogna R.; Kuppusamy P. The curcumin analog HO-3867 selectively kills cancer cells by converting mutant p53 protein to transcriptionally active wildtype p53. J. Biol. Chem. 2018, 293, 4262–4276. 10.1074/jbc.RA117.000950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu Y.; Zhao C.; Zheng H.; Lu K.; Shi D.; Liu Z.; Dai X.; Zhang Y.; Zhang X.; Hu W.; Liang G. A novel STAT3 inhibitor HO-3867 induces cell apoptosis by reactive oxygen species-dependent endoplasmic reticulum stress in human pancreatic cancer cells. Anti-Cancer Drugs 2017, 28, 392–400. 10.1097/CAD.0000000000000470. [DOI] [PubMed] [Google Scholar]
- Mast J. M.; Tse D.; Shee K.; Lakshmi Kuppusamy M.; Kmiec M. M.; Kalai T.; Kuppusamy P. Diarylidenylpiperidones, H-4073 and HO-3867, induce G2/M cell-cycle arrest, apoptosis and inhibit STAT3 phosphorylation in human pancreatic cancer cells. Cell Biochem. Biophys. 2019, 77, 109–119. 10.1007/s12013-019-00873-6. [DOI] [PubMed] [Google Scholar]
- Tierney B. J.; McCann G. A.; Naidu S.; Rath K. S.; Saini U.; Wanner R.; Kuppusamy P.; Suarez A.; Goodfellow P. J.; Cohn D. E.; Selvendiran K. Aberrantly activated pSTAT3-Ser727 in human endometrial cancer is suppressed by HO-3867, a novel STAT3 inhibitor. Gynecol. Oncol. 2014, 135, 133–141. 10.1016/j.ygyno.2014.07.087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen L.; Li Q.; Zheng Z.; Xie J.; Lin X.; Jiang C.; Xu H.; Wu X.; Wu J.; Zhang H. Design and optimize N-substituted EF24 as effective and low toxicity NF-kappaB inhibitor for lung cancer therapy via apoptosis-to-pyroptosis switch. Chem. Biol. Drug Des. 2019, 94, 1368–1377. 10.1111/cbdd.13514. [DOI] [PubMed] [Google Scholar]
- He Y.; Li W.; Hu G.; Sun H.; Kong Q. Bioactivities of EF24, a novel curcumin analog: a review. Front. Oncol. 2018, 8, 614. 10.3389/fonc.2018.00614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aleo E.; Henderson C. J.; Fontanini A.; Solazzo B.; Brancolini C. Identification of new compounds that trigger apoptosome-independent caspase activation and apoptosis. Cancer Res. 2006, 66, 9235–9244. 10.1158/0008-5472.CAN-06-0702. [DOI] [PubMed] [Google Scholar]
- Cersosimo U.; Sgorbissa A.; Foti C.; Drioli S.; Angelica R.; Tomasella A.; Picco R.; Semrau M. S.; Storici P.; Benedetti F.; Berti F.; Brancolini C. Synthesis, characterization, and optimization for in vivo delivery of a nonselective isopeptidase inhibitor as new antineoplastic agent. J. Med. Chem. 2015, 58, 1691–1704. 10.1021/jm501336h. [DOI] [PubMed] [Google Scholar]
- Erdal H.; Berndtsson M.; Castro J.; Brunk U.; Shoshan M. C.; Linder S. Induction of lysosomal membrane permeabilization by compounds that activate p53-independent apoptosis. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 192–197. 10.1073/pnas.0408592102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berndtsson M.; Beaujouin M.; Rickardson L.; Havelka A. M.; Larsson R.; Westman J.; Liaudet-Coopman E.; Linder S. Induction of the lysosomal apoptosis pathway by inhibitors of the ubiquitin-proteasome system. Int. J. Cancer 2009, 124, 1463–1469. 10.1002/ijc.24004. [DOI] [PubMed] [Google Scholar]
- Wang X.; D’Arcy P.; Caulfield T. R.; Paulus A.; Chitta K.; Mohanty C.; Gullbo J.; Chanan-Khan A.; Linder S. Synthesis and evaluation of derivatives of the proteasome deubiquitinase inhibitor b-AP15. Chem. Biol. Drug Des. 2015, 86, 1036–1048. 10.1111/cbdd.12571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X.; Mazurkiewicz M.; Hillert E. K.; Olofsson M. H.; Pierrou S.; Hillertz P.; Gullbo J.; Selvaraju K.; Paulus A.; Akhtar S.; Bossler F.; Khan A. C.; Linder S.; D’Arcy P. The proteasome deubiquitinase inhibitor VLX1570 shows selectivity for ubiquitin-specific protease-14 and induces apoptosis of multiple myeloma cells. Sci. Rep. 2016, 6, 26979. 10.1038/srep26979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shukla N. N.; Somwar R.; Smith R. S.; Ambati S. R.; Munoz S.; Merchant M. S.; D’Arcy P.; Wang X.; Kobos R.; Antczak C.; Bhinder B.; Shum D.; Radu C.; Yang G.; Taylor B. S.; Ng C. K.; Weigelt B.; Khodos I.; de Stanchina E.; Reis-Filho J. S.; Ouerfelli O.; Linder S.; Djaballah H.; Ladanyi M. Proteasome addiction defined in Ewing’s sarcoma is effectively targeted by a novel class of 19S proteasome inhibitors. Cancer Res. 2016, 76, 4525–4534. 10.1158/0008-5472.CAN-16-1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulus A.; Akhtar S.; Caulfield T. R. K; Samuel K.; Yousaf H.; Bashir Y.; Paulus S. M.; Tran D.; Hudec R.; Cogen D.; Jiang J.; Edenfield B.; Novak A.; Ansell S. M.; Witzig T.; Martin P.; Coleman M.; Roy V.; Ailawadhi S.; Chitta K.; Linder S.; Chanan-Khan A. Coinhibition of the deubiquitinating enzymes, USP14 and UCHL5, with VLX1570 is lethal to ibrutinib- or bortezomib-resistant Waldenstrom macroglobulinemia tumor cells. Blood Cancer J. 2016, 6, e492 10.1038/bcj.2016.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X.; Pellegrini P.; Saei A. A.; Hillert E. K.; Mazurkiewicz M.; Olofsson M. H.; Zubarev R. A.; D’Arcy P.; Linder S. The deubiquitinase inhibitor b-AP15 induces strong proteotoxic stress and mitochondrial damage. Biochem. Pharmacol. 2018, 156, 291–301. 10.1016/j.bcp.2018.08.039. [DOI] [PubMed] [Google Scholar]
- Bazzaro M.; Anchoori R. K.; Mudiam M. K.; Issaenko O.; Kumar S.; Karanam B.; Lin Z.; Isaksson Vogel R.; Gavioli R.; Destro F.; Ferretti V.; Roden R. B.; Khan S. R. alpha, beta-Unsaturated carbonyl system of chalcone-based derivatives is responsible for broad inhibition of proteasomal activity and preferential killing of human papilloma virus (HPV) positive cervical cancer cells. J. Med. Chem. 2011, 54, 449–456. 10.1021/jm100589p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anchoori R. K.; Khan S. R.; Sueblinvong T.; Felthauser A.; Iizuka Y.; Gavioli R.; Destro F.; Isaksson Vogel R.; Peng S.; Roden R. B.; Bazzaro M. Stressing the ubiquitin-proteasome system without 20S proteolytic inhibition selectively kills cervical cancer cells. PLoS One 2011, 6, e23888 10.1371/journal.pone.0023888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anchoori R. K.; Karanam B.; Peng S.; Wang J. W.; Jiang R.; Tanno T.; Orlowski R. Z.; Matsui W.; Zhao M.; Rudek M. A.; Hung C. F.; Chen X.; Walters K. J.; Roden R. B. A bis-benzylidine piperidone targeting proteasome ubiquitin receptor RPN13/ADRM1 as a therapy for cancer. Cancer Cell 2013, 24, 791–805. 10.1016/j.ccr.2013.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gargallo P.; Juan A.; Yanez Y.; Dolz S.; Segura V.; Castel V.; Canete A. Precision medicine in Ewing sarcoma: a translational point of view. Clin. Transl. Oncol. 2020, 22, 1440–1454. 10.1007/s12094-020-02298-7. [DOI] [PubMed] [Google Scholar]
- Selvaraju K.; Mofers A.; Pellegrini P.; Salomonsson J.; Ahlner A.; Morad V.; Hillert E. K.; Espinosa B.; Arner E. S. J.; Jensen L.; Malmstrom J.; Turkina M. V.; D’Arcy P.; Walters M. A.; Sunnerhagen M.; Linder S. Cytotoxic unsaturated electrophilic compounds commonly target the ubiquitin proteasome system. Sci. Rep. 2019, 9, 9841. 10.1038/s41598-019-46168-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Somers-Edgar T. J.; Taurin S.; Larsen L.; Chandramouli A.; Nelson M. A.; Rosengren R. J. Mechanisms for the activity of heterocyclic cyclohexanone curcumin derivatives in estrogen receptor negative human breast cancer cell lines. Invest. New Drugs 2011, 29, 87–97. 10.1007/s10637-009-9339-0. [DOI] [PubMed] [Google Scholar]
- Robles-Escajeda E.; Das U.; Ortega N. M.; Parra K.; Francia G.; Dimmock J. R.; Varela-Ramirez A.; Aguilera R. J. A novel curcumin-like dienone induces apoptosis in triple-negative breast cancer cells. Cell. Oncol. 2016, 39, 265–277. 10.1007/s13402-016-0272-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalai T.; Kuppusamy M. L.; Balog M.; Selvendiran K.; Rivera B. K.; Kuppusamy P.; Hideg K. Synthesis of N-substituted 3,5-bis(arylidene)-4-piperidones with high antitumor and antioxidant activity. J. Med. Chem. 2011, 54, 5414–5421. 10.1021/jm200353f. [DOI] [PubMed] [Google Scholar]
- Makarov M. V.; Rybalkina E. Y.; Anikina L. V.; Pukhov S. A.; Klochkov S. G.; Mischenko D. V.; Neganova M. E.; Khrustalev V. N.; Klemenkova Z. S.; Brel V. K. 1,5-diaryl-3-oxo-1,4-pentadienes based on (4-oxopiperidin-1-yl)(aryl)methyl phosphonate scaffold: synthesis and antitumor properties. Med. Chem. Res. 2017, 26, 140–152. 10.1007/s00044-016-1726-4. [DOI] [Google Scholar]
- Sun J.; Wang S.; Li H.; Jiang W.; Hou G.; Zhao F.; Cong W. Synthesis, antitumor activity evaluation of some new N-aroyl-alpha, beta-unsaturated piperidones with fluorescence. J. Enzyme Inhib. Med. Chem. 2016, 31, 495–502. 10.3109/14756366.2015.1043296. [DOI] [PubMed] [Google Scholar]
- Wei X.; Du Z. Y.; Zheng X.; Cui X. X.; Conney A. H.; Zhang K. Synthesis and evaluation of curcumin-related compounds for anticancer activity. Eur. J. Med. Chem. 2012, 53, 235–245. 10.1016/j.ejmech.2012.04.005. [DOI] [PubMed] [Google Scholar]
- Youssef D.; Potter E.; Jha M.; De Clercq E.; Balzarini J.; Stables J. P.; Jha A. Design, synthesis and bioevaluation of novel maleamic amino acid ester conjugates of 3,5-bisarylmethylene-4-piperidones as cytostatic agents. Bioorg. Med. Chem. Lett. 2009, 19, 6364–6367. 10.1016/j.bmcl.2009.09.069. [DOI] [PubMed] [Google Scholar]
- Cheng D.; Valente S.; Castellano S.; Sbardella G.; Di Santo R.; Costi R.; Bedford M. T.; Mai A. Novel 3,5-bis(bromohydroxybenzylidene)piperidin-4-ones as coactivator-associated arginine methyltransferase 1 inhibitors: enzyme selectivity and cellular activity. J. Med. Chem. 2011, 54, 4928–4932. 10.1021/jm200453n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhandarkar S. S.; Bromberg J.; Carrillo C.; Selvakumar P.; Sharma R. K.; Perry B. N.; Govindarajan B.; Fried L.; Sohn A.; Reddy K.; Arbiser J. L. Tris (dibenzylideneacetone) dipalladium, a N-myristoyltransferase-1 inhibitor, is effective against melanoma growth in vitro and in vivo. Clin. Cancer Res. 2008, 14, 5743–5748. 10.1158/1078-0432.CCR-08-0405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musi E.; Schwartz G. K.; Yoo J. H.; Odelberg S. J.; Li D. Y.; Bonner M. Y.; Selvakumar P.; Rao S.; Gilbert L. C.; Elsey J.; Arbiser J. L. Tris DBA palladium is an orally available inhibitor of GNAQ mutant uveal melanoma in vivo. Oncotarget 2019, 10, 4424–4436. 10.18632/oncotarget.27040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz B.; Ostapoff K. T.; Toombs J. E.; Lo J.; Bonner M. Y.; Curatolo A.; Adsay V.; Brekken R. A.; Arbiser J. L. Tris DBA palladium is highly effective against growth and metastasis of pancreatic cancer in an orthotopic model. Oncotarget 2016, 7, 51569–51580. 10.18632/oncotarget.10514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee H. E.; Choi E. S.; Jung J. Y.; You M. J.; Kim L. H.; Cho S. D. Inhibition of specificity protein 1 by dibenzylideneacetone, a curcumin analogue, induces apoptosis in mucoepidermoid carcinomas and tumor xenografts through Bim and truncated Bid. Oral Oncol. 2014, 50, 189–195. 10.1016/j.oraloncology.2013.11.006. [DOI] [PubMed] [Google Scholar]
- Quincoces Suarez J. A.; Rando D. G.; Santos R. P.; Goncalves C. P.; Ferreira E.; de Carvalho J. E.; Kohn L.; Maria D. A.; Faiao-Flores F.; Michalik D.; Marcucci M. C.; Vogel C. New antitumoral agents I: in vitro anticancer activity and in vivo acute toxicity of synthetic 1,5-bis(4-hydroxy-3-methoxyphenyl)-1,4-pentadien-3-one and derivatives. Bioorg. Med. Chem. 2010, 18, 6275–6281. 10.1016/j.bmc.2010.07.026. [DOI] [PubMed] [Google Scholar]
- Zhao C.; Yang J.; Wang Y.; Liang D.; Yang X.; Li X.; Wu J.; Wu X.; Yang S.; Li X.; Liang G. Synthesis of mono-carbonyl analogues of curcumin and their effects on inhibition of cytokine release in LPS-stimulated RAW 264.7 macrophages. Bioorg. Med. Chem. 2010, 18, 2388–2393. 10.1016/j.bmc.2010.03.001. [DOI] [PubMed] [Google Scholar]
- Wang Y.; Yu C.; Pan Y.; Li J.; Zhang Y.; Ye F.; Yang S.; Zhang H.; Li X.; Liang G. A novel compound C12 inhibits inflammatory cytokine production and protects from inflammatory injury in vivo. PLoS One 2011, 6, e24377 10.1371/journal.pone.0024377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vilekar P.; Awasthi S.; Natarajan A.; Anant S.; Awasthi V. EF24 suppresses maturation and inflammatory response in dendritic cells. Int. Immunol. 2012, 24, 455–464. 10.1093/intimm/dxr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vilekar P.; Rao G.; Awasthi S.; Awasthi V. Diphenyldifluoroketone EF24 suppresses pro-inflammatory interleukin-1 receptor 1 and toll-like receptor 4 in lipopolysaccharide-stimulated dendritic cells. J. Inflammation 2015, 12, 55. 10.1186/s12950-015-0096-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osman H.; Idris N. H.; Kamarulzaman E. E.; Wahab H. A.; Hassan M. Z. 3,5-Bis(arylidene)-4-piperidones as potential dengue protease inhibitors. Acta Pharm. Sin. B 2017, 7, 479–484. 10.1016/j.apsb.2017.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aher R. B.; Wanare G.; Kawathekar N.; Kumar R. R.; Kaushik N. K.; Sahal D.; Chauhan V. S. Dibenzylideneacetone analogues as novel Plasmodium falciparum inhibitors. Bioorg. Med. Chem. Lett. 2011, 21, 3034–3036. 10.1016/j.bmcl.2011.03.037. [DOI] [PubMed] [Google Scholar]
- Wang L.; Delahunty C.; Fritz-Wolf K.; Rahlfs S.; Helena Prieto J.; Yates J. R.; Becker K. Characterization of the 26S proteasome network in Plasmodium falciparum. Sci. Rep. 2015, 5, 17818. 10.1038/srep17818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chauhan I. S.; Rao G. S.; Shankar J.; Chauhan L. K. S.; Kapadia G. J.; Singh N. Chemoprevention of Leishmaniasis: in-vitro antiparasitic activity of dibenzalacetone, a synthetic curcumin analog leads to apoptotic cell death in Leishmania donovani. Parasitol. Int. 2018, 67, 627–636. 10.1016/j.parint.2018.06.004. [DOI] [PubMed] [Google Scholar]
- Baell J.; Walters M. A. Chemistry: Chemical con artists foil drug discovery. Nature 2014, 513, 481–483. 10.1038/513481a. [DOI] [PubMed] [Google Scholar]
- Yang J. J.; Ursu O.; Lipinski C. A.; Sklar L. A.; Oprea T. I.; Bologa C. G. Badapple: promiscuity patterns from noisy evidence. J. Cheminf. 2016, 8, 29. 10.1186/s13321-016-0137-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Didier R.; Mallavialle A.; Ben Jouira R.; Domdom M. A.; Tichet M.; Auberger P.; Luciano F.; Ohanna M.; Tartare-Deckert S.; Deckert M. Targeting the proteasome-associated deubiquitinating enzyme USP14 impairs melanoma cell survival and overcomes resistance to MAPK-targeting therapies. Mol. Cancer Ther. 2018, 17, 1416–1429. 10.1158/1535-7163.MCT-17-0919. [DOI] [PubMed] [Google Scholar]
- Richardson P. G.; Schlossman R.; Hideshima T.; Anderson K. C. New treatments for multiple myeloma. Oncology 2005, 19, 1781–1792. [PubMed] [Google Scholar]
- Ciotti S.; Sgarra R.; Sgorbissa A.; Penzo C.; Tomasella A.; Casarsa F.; Benedetti F.; Berti F.; Manfioletti G.; Brancolini C. The binding landscape of a partially-selective isopeptidase inhibitor with potent pro-death activity, based on the bis(arylidene)cyclohexanone scaffold. Cell Death Dis. 2018, 9, 184. 10.1038/s41419-017-0259-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koulich E.; Li X.; DeMartino G. N. Relative structural and functional roles of multiple deubiquitylating proteins associated with mammalian 26S proteasome. Mol. Biol. Cell 2008, 19, 1072–1082. 10.1091/mbc.e07-10-1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian Z.; D’Arcy P.; Wang X.; Ray A.; Tai Y. T.; Hu Y.; Carrasco R. D.; Richardson P.; Linder S.; Chauhan D.; Anderson K. C. A novel small molecule inhibitor of deubiquitylating enzyme USP14 and UCHL5 induces apoptosis in multiple myeloma and overcomes bortezomib resistance. Blood 2014, 123, 706–716. 10.1182/blood-2013-05-500033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang L.; Sun Y.; Wang J.; He Q.; Chen X.; Lan X.; Chen J.; Dou Q. P.; Shi X.; Liu J. Proteasomal cysteine deubiquitinase inhibitor b-AP15 suppresses migration and induces apoptosis in diffuse large B cell lymphoma. J. Exp. Clin. Cancer Res. 2019, 38, 453. 10.1186/s13046-019-1446-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X.; Stafford W.; Mazurkiewicz M.; Fryknas M.; Brjnic S.; Zhang X.; Gullbo J.; Larsson R.; Arner E. S.; D’Arcy P.; Linder S. The 19S Deubiquitinase inhibitor b-AP15 is enriched in cells and elicits rapid commitment to cell death. Mol. Pharmacol. 2014, 85, 932–945. 10.1124/mol.113.091322. [DOI] [PubMed] [Google Scholar]
- Borodovsky A.; Kessler B. M.; Casagrande R.; Overkleeft H. S.; Wilkinson K. D.; Ploegh H. L. A novel active site-directed probe specific for deubiquitylating enzymes reveals proteasome association of USP14. EMBO J. 2001, 20, 5187–5196. 10.1093/emboj/20.18.5187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagner B. J.; Margolis J. W. Thermal stability and activation of bovine lens multicatalytic proteinase complex (proteasome). Arch. Biochem. Biophys. 1993, 307, 146–152. 10.1006/abbi.1993.1573. [DOI] [PubMed] [Google Scholar]
- Lu X.; Nowicka U.; Sridharan V.; Liu F.; Randles L.; Hymel D.; Dyba M.; Tarasov S. G.; Tarasova N. I.; Zhao X. Z.; Hamazaki J.; Murata S.; Burke T. R. Jr.; Walters K. J. Structure of the Rpn13-Rpn2 complex provides insights for Rpn13 and Uch37 as anticancer targets. Nat. Commun. 2017, 8, 15540. 10.1038/ncomms15540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogel R. I.; Coughlin K.; Scotti A.; Iizuka Y.; Anchoori R.; Roden R. B.; Marastoni M.; Bazzaro M. Simultaneous inhibition of deubiquitinating enzymes (DUBs) and autophagy synergistically kills breast cancer cells. Oncotarget 2015, 6, 4159–4170. 10.18632/oncotarget.2904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson B.; Leach C. A.; Goldenberg S. J.; Francis D. M.; Kodrasov M. P.; Tian X.; Shanks J.; Sterner D. E.; Bernal A.; Mattern M. R.; Wilkinson K. D.; Butt T. R. Characterization of ubiquitin and ubiquitin-like-protein isopeptidase activities. Protein Sci. 2008, 17, 1035–1043. 10.1110/ps.083450408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward J. A.; Pinto-Fernandez A.; Cornelissen L.; Bonham S.; Diaz-Saez L.; Riant O.; Huber K. V. M.; Kessler B. M.; Feron O.; Tate E. W. Re-evaluating the mechanism of action of alpha, beta-unsaturated carbonyl DUB inhibitors b-AP15 and VLX1570: a paradigmatic example of unspecific protein cross-linking with Michael acceptor motif-containing drugs. J. Med. Chem. 2020, 63, 3756–3762. 10.1021/acs.jmedchem.0c00144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers R. M.; Bryan J. G.; McFarland J. M.; Weir B. A.; Sizemore A. E.; Xu H.; Dharia N. V.; Montgomery P. G.; Cowley G. S.; Pantel S.; Goodale A.; Lee Y.; Ali L. D.; Jiang G.; Lubonja R.; Harrington W. F.; Strickland M.; Wu T.; Hawes D. C.; Zhivich V. A.; Wyatt M. R.; Kalani Z.; Chang J. J.; Okamoto M.; Stegmaier K.; Golub T. R.; Boehm J. S.; Vazquez F.; Root D. E.; Hahn W. C.; Tsherniak A. Computational correction of copy number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells. Nat. Genet. 2017, 49, 1779–1784. 10.1038/ng.3984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin T. D.; Cook D. R.; Choi M. Y.; Li M. Z.; Haigis K. M.; Elledge S. J. A role for mitochondrial translation in promotion of viability in K-Ras mutant cells. Cell Rep. 2017, 20, 427–438. 10.1016/j.celrep.2017.06.061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao G.; Nkepang G.; Xu J.; Yari H.; Houson H.; Teng C.; Awasthi V. Ubiquitin receptor RPN13 mediates the inhibitory interaction of diphenyldihaloketones CLEFMA and EF24 with the 26S proteasome. Front. Chem. 2018, 6, 392. 10.3389/fchem.2018.00392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osei-Amponsa V.; Sridharan V.; Tandon M.; Evans C. N.; Klarmann K.; Cheng K. T.; Lack J.; Chari R.; Walters K. J. Impact of losing hRpn13 Pru or UCHL5 on proteasome clearance of ubiquitinated proteins and RA190 cytotoxicity. Mol. Cell. Biol. 2020, 40, e00122 10.1128/MCB.00122-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brnjic S.; Mazurkiewicz M.; Fryknas M.; Sun C.; Zhang X.; Larsson R.; D’Arcy P.; Linder S. Induction of tumor cell apoptosis by a proteasome deubiquitinase inhibitor is associated with oxidative stress. Antioxid. Redox Signaling 2014, 21, 2271–2285. 10.1089/ars.2013.5322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan K. L.; Ali A.; Du Y.; Fu H.; Jin H. X.; Chin T. M.; Khan M.; Go M. L. Synthesis and evaluation of bisbenzylidenedioxotetrahydrothiopranones as activators of endoplasmic reticulum (ER) stress signaling pathways and apoptotic cell death in acute promyelocytic leukemic cells. J. Med. Chem. 2014, 57, 5904–5918. 10.1021/jm401352a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Addala E.; Rafiei H.; Das S.; Bandy B.; Das U.; Karki S. S.; Dimmock J. R. 3,5-Bis(3-dimethylaminomethyl-4-hydroxybenzylidene)-4-piperidone and related compounds induce glutathione oxidation and mitochondria-mediated cell death in HCT-116 colon cancer cells. Bioorg. Med. Chem. Lett. 2017, 27, 3669–3673. 10.1016/j.bmcl.2017.07.018. [DOI] [PubMed] [Google Scholar]
- Khalid M.; Ali A.; Rehman M. F. U.; Mustaqeem M.; Ali S.; Khan M. U.; Asim S.; Ahmad N.; Saleem M. Exploration of noncovalent interactions, chemical reactivity, and nonlinear optical properties of piperidone derivatives: a concise theoretical approach. ACS omega 2020, 5, 13236–13249. 10.1021/acsomega.0c01273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernard K.; Wang W.; Narlawar R.; Schmidt B.; Kirk K. L. Curcumin cross-links cystic fibrosis transmembrane conductance regulator (CFTR) polypeptides and potentiates CFTR channel activity by distinct mechanisms. J. Biol. Chem. 2009, 284, 30754–30765. 10.1074/jbc.M109.056010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stanwell C.; Burke T. R. Jr.; Yuspa S. H. The erbstatin analogue methyl 2,5-dihydroxycinnamate cross-links proteins and is cytotoxic to normal and neoplastic epithelial cells by a mechanism independent of tyrosine kinase inhibition. Cancer Res. 1995, 55, 4950–4956. [PubMed] [Google Scholar]
- Stanwell C.; Ye B.; Yuspa S. H.; Burke T. R. Jr. Cell protein cross-linking by erbstatin and related compounds. Biochem. Pharmacol. 1996, 52, 475–480. 10.1016/0006-2952(96)00250-X. [DOI] [PubMed] [Google Scholar]
- Pinto A.; Mace Y.; Drouet F.; Bony E.; Boidot R.; Draoui N.; Lobysheva I.; Corbet C.; Polet F.; Martherus R.; Deraedt Q.; Rodriguez J.; Lamy C.; Schicke O.; Delvaux D.; Louis C.; Kiss R.; Kriegsheim A. V.; Dessy C.; Elias B.; Quetin-Leclercq J.; Riant O.; Feron O. A new ER-specific photosensitizer unravels O-driven protein oxidation and inhibition of deubiquitinases as a generic mechanism for cancer PDT. Oncogene 2016, 35, 3976–3985. 10.1038/onc.2015.474. [DOI] [PubMed] [Google Scholar]
- Zou P.; Xia Y.; Chen W.; Chen X.; Ying S.; Feng Z.; Chen T.; Ye Q.; Wang Z.; Qiu C.; Yang S.; Liang G. EF24 induces ROS-mediated apoptosis via targeting thioredoxin reductase 1 in gastric cancer cells. Oncotarget 2016, 7, 18050–18064. 10.18632/oncotarget.7633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skoupa N.; Dolezel P.; Ruzickova E.; Mlejnek P. Apoptosis induced by the curcumin analogue EF-24 is neither mediated by oxidative stress-related mechanisms nor affected by expression of main drug transporters ABCB1 and ABCG2 in human leukemia cells. Int. J. Mol. Sci. 2017, 18, 2289. 10.3390/ijms18112289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rundlof A. K.; Arner E. S. Regulation of the mammalian selenoprotein thioredoxin reductase 1 in relation to cellular phenotype, growth, and signaling events. Antioxid. Redox Signaling 2004, 6, 41–52. 10.1089/152308604771978336. [DOI] [PubMed] [Google Scholar]
- Zhang X.; Espinosa B.; Saei A. A.; D’Arcy P.; Zubarev R. A.; Linder S. Oxidative stress induced by the deubiquitinase Inhibitor b-AP15 is associated with mitochondrial impairment. Oxid. Med. Cell. Longevity 2019, 2019, 1659468. 10.1155/2019/1659468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McConkey D. J. The integrated stress response and proteotoxicity in cancer therapy. Biochem. Biophys. Res. Commun. 2017, 482, 450–453. 10.1016/j.bbrc.2016.11.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheaff R. J.; Singer J. D.; Swanger J.; Smitherman M.; Roberts J. M.; Clurman B. E. Proteasomal turnover of p21Cip1 does not require p21Cip1 ubiquitination. Mol. Cell 2000, 5, 403–410. 10.1016/S1097-2765(00)80435-9. [DOI] [PubMed] [Google Scholar]
- Eldeeb M. A.; Fahlman R. P. The anti-apoptotic form of tyrosine kinase Lyn that is generated by proteolysis is degraded by the N-end rule pathway. Oncotarget 2014, 5, 2714–2722. 10.18632/oncotarget.1931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez G.; Vazquez-Pianzola P.; Sierra J. M.; Rivera-Pomar R. Internal ribosome entry site drives cap-independent translation of reaper and heat shock protein 70 mRNAs in Drosophila embryos. RNA 2004, 10, 1783–1797. 10.1261/rna.7154104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pellegrini P.; Selvaraju K.; Faustini E.; Mofers A.; Zhang X.; Ternerot J.; Schubert A.; Linder S.; D’Arcy P. Induction of ER stress in acute lymphoblastic leukemia cells by the deubiquitinase inhibitor VLX1570. Int. J. Mol. Sci. 2020, 21, 4757. 10.3390/ijms21134757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khoury H. J.; Cortes J.; Baccarani M.; Wetzler M.; Masszi T.; Digumarti R.; Craig A.; Benichou A. C.; Akard L. Omacetaxine mepesuccinate in patients with advanced chronic myeloid leukemia with resistance or intolerance to tyrosine kinase inhibitors. Leuk. Lymphoma 2015, 56, 120–127. 10.3109/10428194.2014.889826. [DOI] [PubMed] [Google Scholar]
- Selvakumar P.; Lakshmikuttyamma A.; Das U.; Pati H. N.; Dimmock J. R.; Sharma R. K. NC2213: a novel methionine aminopeptidase 2 inhibitor in human colon cancer HT29 cells. Mol. Cancer 2009, 8, 65. 10.1186/1476-4598-8-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giglione C.; Fieulaine S.; Meinnel T. N-terminal protein modifications: Bringing back into play the ribosome. Biochimie 2015, 114, 134–146. 10.1016/j.biochi.2014.11.008. [DOI] [PubMed] [Google Scholar]
- Shimizu H.; Yamagishi S.; Chiba H.; Ghazizadeh M. Methionine aminopeptidase 2 as a potential therapeutic target for human Non-Small-Cell Lung Cancers. Adv. clin exp med 2016, 25, 117–128. 10.17219/acem/60715. [DOI] [PubMed] [Google Scholar]
- Brown C. J.; Cheok C. F.; Verma C. S.; Lane D. P. Reactivation of p53: from peptides to small molecules. Trends Pharmacol. Sci. 2011, 32, 53–62. 10.1016/j.tips.2010.11.004. [DOI] [PubMed] [Google Scholar]
- Tomasella A.; Picco R.; Ciotti S.; Sgorbissa A.; Bianchi E.; Manfredini R.; Benedetti F.; Trimarco V.; Frezzato F.; Trentin L.; Semenzato G.; Delia D.; Brancolini C. The isopeptidase inhibitor 2cPE triggers proteotoxic stress and ATM activation in chronic lymphocytic leukemia cells. Oncotarget 2016, 7, 45429–45443. 10.18632/oncotarget.9742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo Z.; Kozlov S.; Lavin M. F.; Person M. D.; Paull T. T. ATM activation by oxidative stress. Science 2010, 330, 517–521. 10.1126/science.1192912. [DOI] [PubMed] [Google Scholar]
- Landais I.; Hiddingh S.; McCarroll M.; Yang C.; Sun A.; Turker M. S.; Snyder J. P.; Hoatlin M. E. Monoketone analogs of curcumin, a new class of Fanconi anemia pathway inhibitors. Mol. Cancer 2009, 8, 133. 10.1186/1476-4598-8-133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A.; Narayan R.; Corsello S. M.; Peck D. D.; Natoli T. E.; Lu X.; Gould J.; Davis J. F.; Tubelli A. A.; Asiedu J. K.; Lahr D. L.; Hirschman J. E.; Liu Z.; Donahue M.; Julian B.; Khan M.; Wadden D.; Smith I. C.; Lam D.; Liberzon A.; Toder C.; Bagul M.; Orzechowski M.; Enache O. M.; Piccioni F.; Johnson S. A.; Lyons N. J.; Berger A. H.; Shamji A. F.; Brooks A. N.; Vrcic A.; Flynn C.; Rosains J.; Takeda D. Y.; Hu R.; Davison D.; Lamb J.; Ardlie K.; Hogstrom L.; Greenside P.; Gray N. S.; Clemons P. A.; Silver S.; Wu X.; Zhao W. N.; Read-Button W.; Wu X.; Haggarty S. J.; Ronco L. V.; Boehm J. S.; Schreiber S. L.; Doench J. G.; Bittker J. A.; Root D. E.; Wong B.; Golub T. R. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 2017, 171, 1437–1452. 10.1016/j.cell.2017.10.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A.; Tamayo P.; Mootha V. K.; Mukherjee S.; Ebert B. L.; Gillette M. A.; Paulovich A.; Pomeroy S. L.; Golub T. R.; Lander E. S.; Mesirov J. P. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 15545–15550. 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hillert E. K.; Brnjic S.; Zhang X.; Mazurkiewicz M.; Saei A. A.; Mofers A.; Selvaraju K.; Zubarev R.; Linder S.; D’Arcy P. Proteasome inhibitor b-AP15 induces enhanced proteotoxicity by inhibiting cytoprotective aggresome formation. Cancer Lett. 2019, 448, 70–83. 10.1016/j.canlet.2019.02.003. [DOI] [PubMed] [Google Scholar]
- Rodriguez-Gonzalez A.; Lin T.; Ikeda A. K.; Simms-Waldrip T.; Fu C.; Sakamoto K. M. Role of the aggresome pathway in cancer: targeting histone deacetylase 6-dependent protein degradation. Cancer Res. 2008, 68, 2557–2560. 10.1158/0008-5472.CAN-07-5989. [DOI] [PubMed] [Google Scholar]
- Cajone F.; Crescente M. In vitro activation of heat shock transcription factor by 4-hydroxynonenal. Chem.-Biol. Interact. 1992, 84, 97–112. 10.1016/0009-2797(92)90071-R. [DOI] [PubMed] [Google Scholar]
- Schubert U.; Anton L. C.; Gibbs J.; Norbury C. C.; Yewdell J. W.; Bennink J. R. Rapid degradation of a large fraction of newly synthesized proteins by proteasomes. Nature 2000, 404, 770–774. 10.1038/35008096. [DOI] [PubMed] [Google Scholar]
- Sun C.; Roboti P.; Puumalainen M. R.; Fryknas M.; Wang X.; D’Arcy P.; Hult M.; High S.; Linder S.; Swanton E. Elevation of proteasomal substrate levels sensitizes cells to apoptosis induced by inhibition of proteasomal deubiquitinases. PLoS One 2014, 9, e108839 10.1371/journal.pone.0108839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Besemer J.; Harant H.; Wang S.; Oberhauser B.; Marquardt K.; Foster C. A.; Schreiner E. P.; de Vries J. E.; Dascher-Nadel C.; Lindley I. J. Selective inhibition of cotranslational translocation of vascular cell adhesion molecule 1. Nature 2005, 436, 290–293. 10.1038/nature03670. [DOI] [PubMed] [Google Scholar]
- Enam C.; Geffen Y.; Ravid T.; Gardner R. G. Protein quality control degradation in the nucleus. Annu. Rev. Biochem. 2018, 87, 725–749. 10.1146/annurev-biochem-062917-012730. [DOI] [PubMed] [Google Scholar]
- Clemens J.; Seckinger A.; Hose D.; Theile D.; Longo M.; Haefeli W. E.; Burhenne J.; Weiss J. Cellular uptake kinetics of bortezomib in relation to efficacy in myeloma cells and the influence of drug transporters. Cancer Chemother. Pharmacol. 2015, 75, 281–291. 10.1007/s00280-014-2643-1. [DOI] [PubMed] [Google Scholar]
- Menendez-Benito V.; Verhoef L. G.; Masucci M. G.; Dantuma N. P. Endoplasmic reticulum stress compromises the ubiquitin-proteasome system. Hum. Mol. Genet. 2005, 14, 2787–2799. 10.1093/hmg/ddi312. [DOI] [PubMed] [Google Scholar]
- Yang J.; Tallman K. A.; Porter N. A.; Liebler D. C. Quantitative chemoproteomics for site-specific analysis of protein alkylation by 4-hydroxy-2-nonenal in cells. Anal. Chem. 2015, 87, 2535–2541. 10.1021/ac504685y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mol M.; Regazzoni L.; Altomare A.; Degani G.; Carini M.; Vistoli G.; Aldini G. Enzymatic and non-enzymatic detoxification of 4-hydroxynonenal: methodological aspects and biological consequences. Free Radical Biol. Med. 2017, 111, 328–344. 10.1016/j.freeradbiomed.2017.01.036. [DOI] [PubMed] [Google Scholar]
- Maisonneuve E.; Ducret A.; Khoueiry P.; Lignon S.; Longhi S.; Talla E.; Dukan S. Rules governing selective protein carbonylation. PLoS One 2009, 4, e7269 10.1371/journal.pone.0007269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcone S.; Fitzgerald D. J. Proteomic identification of the candidate target proteins of 15-deoxy-delta12,14-prostaglandin J2. Proteomics 2013, 13, 2135–2139. 10.1002/pmic.201200289. [DOI] [PubMed] [Google Scholar]
- Fang J.; Lu J.; Holmgren A. Thioredoxin reductase is irreversibly modified by curcumin: a novel molecular mechanism for its anticancer activity. J. Biol. Chem. 2005, 280, 25284–25290. 10.1074/jbc.M414645200. [DOI] [PubMed] [Google Scholar]
- Okada K.; Wangpoengtrakul C.; Osawa T.; Toyokuni S.; Tanaka K.; Uchida K. 4-Hydroxy-2-nonenal-mediated impairment of intracellular proteolysis during oxidative stress. Identification of proteasomes as target molecules. J. Biol. Chem. 1999, 274, 23787–23793. 10.1074/jbc.274.34.23787. [DOI] [PubMed] [Google Scholar]
- Marques C.; Pereira P.; Taylor A.; Liang J. N.; Reddy V. N.; Szweda L. I.; Shang F. Ubiquitin-dependent lysosomal degradation of the HNE-modified proteins in lens epithelial cells. FASEB J. 2004, 18, 1424–1426. 10.1096/fj.04-1743fje. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang S.; Eitan E.; Mattson M. P. Early involvement of lysosome dysfunction in the degeneration of cerebral cortical neurons caused by the lipid peroxidation product 4-hydroxynonenal. J. Neurochem. 2017, 140, 941–954. 10.1111/jnc.13957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dayton A.; Selvendiran K.; Kuppusamy M. L.; Rivera B. K.; Meduru S.; Kalai T.; Hideg K.; Kuppusamy P. Cellular uptake, retention and bioabsorption of HO-3867, a fluorinated curcumin analog with potential antitumor properties. Cancer Biol. Ther. 2010, 10, 1027–1032. 10.4161/cbt.10.10.13250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D.; Lippard S. J. Cellular processing of platinum anticancer drugs. Nat. Rev. Drug Discovery 2005, 4, 307–320. 10.1038/nrd1691. [DOI] [PubMed] [Google Scholar]
- Berndtsson M.; Hagg M.; Panaretakis T.; Havelka A. M.; Shoshan M. C.; Linder S. Acute apoptosis by cisplatin requires induction of reactive oxygen species but is not associated with damage to nuclear DNA. Int. J. Cancer 2007, 120, 175–180. 10.1002/ijc.22132. [DOI] [PubMed] [Google Scholar]
- Robinson T. P.; Ehlers T.; Hubbard R. B. IV; Bai X.; Arbiser J. L.; Goldsmith D. J.; Bowen J. P. Design, synthesis, and biological evaluation of angiogenesis inhibitors: aromatic enone and dienone analogues of curcumin. Bioorg. Med. Chem. Lett. 2003, 13, 115–117. 10.1016/S0960-894X(02)00832-6. [DOI] [PubMed] [Google Scholar]
- Thackaberry E. A.; Kopytek S.; Sherratt P.; Trouba K.; McIntyre B. Comprehensive investigation of hydroxypropyl methylcellulose, propylene glycol, polysorbate 80, and hydroxypropyl-beta-cyclodextrin for use in general toxicology studies. Toxicol. Sci. 2010, 117, 485–492. 10.1093/toxsci/kfq207. [DOI] [PubMed] [Google Scholar]
- Rowinsky E. K.; Paner A.; Berdeja J. G.; Paba-Prada C.; Venugopal P.; Porkka K.; Gullbo J.; Linder S.; Loskog A.; Richardson P. G.; Landgren O. Phase 1 study of the protein deubiquitinase inhibitor VLX1570 in patients with relapsed and/or refractory multiple myeloma. Invest. New Drugs 2020, 38, 1448–1453. 10.1007/s10637-020-00915-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kharel P.; Uprety D.; Chandra A. B.; Hu Y.; Belur A. A.; Dhakal A. Bortezomib-Induced Pulmonary Toxicity: A case report and review of literature. Case Rep. Med. 2018, 2018, 2913124. 10.1155/2018/2913124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galluzzi L.; Vitale I.; Aaronson S. A.; Abrams J. M.; Adam D.; Agostinis P.; Alnemri E. S.; Altucci L.; Amelio I.; Andrews D. W.; Annicchiarico-Petruzzelli M.; Antonov A. V.; Arama E.; Baehrecke E. H.; Barlev N. A.; Bazan N. G.; Bernassola F.; Bertrand M. J. M.; Bianchi K.; Blagosklonny M. V.; Blomgren K.; Borner C.; Boya P.; Brenner C.; Campanella M.; Candi E.; Carmona-Gutierrez D.; Cecconi F.; Chan F. K.-M.; Chandel N. S.; Cheng E. H.; Chipuk J. E.; Cidlowski J. A.; Ciechanover A.; Cohen G. M.; Conrad M.; Cubillos-Ruiz J. R.; Czabotar P. E.; D’Angiolella V.; Dawson T. M.; Dawson V. L.; De Laurenzi V.; De Maria R.; Debatin K.-M.; DeBerardinis R. J.; Deshmukh M.; Di Daniele N.; Di Virgilio F.; Dixit V. M.; Dixon S. J.; Duckett C. S.; Dynlacht B. D.; El-Deiry W. S.; Elrod J. W.; Fimia G. M.; Fulda S.; Garcia-Saez A. J.; Garg A. D.; Garrido C.; Gavathiotis E.; Golstein P.; Gottlieb E.; Green D. R.; Greene L. A.; Gronemeyer H.; Gross A.; Hajnoczky G.; Hardwick J. M.; Harris I. S.; Hengartner M. O.; Hetz C.; Ichijo H.; Jaattela M.; Joseph B.; Jost P. J.; Juin P. P.; Kaiser W. J.; Karin M.; Kaufmann T.; Kepp O.; Kimchi A.; Kitsis R. N.; Klionsky D. J.; Knight R. A.; Kumar S.; Lee S. W.; Lemasters J. J.; Levine B.; Linkermann A.; Lipton S. A.; Lockshin R. A.; Lopez-Otin C.; Lowe S. W.; Luedde T.; Lugli E.; MacFarlane M.; Madeo F.; Malewicz M.; Malorni W.; Manic G.; Marine J.-C.; Martin S. J.; Martinou J.-C.; Medema J. P.; Mehlen P.; Meier P.; Melino S.; Miao E. A.; Molkentin J. D.; Moll U. M.; Munoz-Pinedo C.; Nagata S.; Nunez G.; Oberst A.; Oren M.; Overholtzer M.; Pagano M.; Panaretakis T.; Pasparakis M.; Penninger J. M.; Pereira D. M.; Pervaiz S.; Peter M. E.; Piacentini M.; Pinton P.; Prehn J. H. M.; Puthalakath H.; Rabinovich G. A.; Rehm M.; Rizzuto R.; Rodrigues C. M. P.; Rubinsztein D. C.; Rudel T.; Ryan K. M.; Sayan E.; Scorrano L.; Shao F.; Shi Y.; Silke J.; Simon H.-U.; Sistigu A.; Stockwell B. R.; Strasser A.; Szabadkai G.; Tait S. W. G.; Tang D.; Tavernarakis N.; Thorburn A.; Tsujimoto Y.; Turk B.; Vanden Berghe T.; Vandenabeele P.; Vander Heiden M. G.; Villunger A.; Virgin H. W.; Vousden K. H.; Vucic D.; Wagner E. F.; Walczak H.; Wallach D.; Wang Y.; Wells J. A.; Wood W.; Yuan J.; Zakeri Z.; Zhivotovsky B.; Zitvogel L.; Melino G.; Kroemer G. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. 10.1038/s41418-017-0012-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foti C.; Demarchi F.; Brancolini C. Inhibitors of the ubiquitin-proteasome system are not all alike: identification of a new necrotic pathway. Autophagy 2009, 5, 543–545. 10.4161/auto.5.4.8169. [DOI] [PubMed] [Google Scholar]
- Foti C.; Florean C.; Pezzutto A.; Roncaglia P.; Tomasella A.; Gustincich S.; Brancolini C. Characterization of caspase-dependent and caspase-independent deaths in glioblastoma cells treated with inhibitors of the ubiquitin-proteasome system. Mol. Cancer Ther. 2009, 8, 3140–3150. 10.1158/1535-7163.MCT-09-0431. [DOI] [PubMed] [Google Scholar]
- Fontanini A.; Foti C.; Potu H.; Crivellato E.; Maestro R.; Bernardi P.; Demarchi F.; Brancolini C. The isopeptidase inhibitor G5 triggers a caspase-independent necrotic death in cells resistant to apoptosis: a comparative study with the proteasome inhibitor bortezomib. J. Biol. Chem. 2009, 284, 8369–8381. 10.1074/jbc.M806113200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sperandio S.; de Belle I.; Bredesen D. E. An alternative, nonapoptotic form of programmed cell death. Proc. Natl. Acad. Sci. U. S. A. 2000, 97, 14376–14381. 10.1073/pnas.97.26.14376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nijhawan D.; Zack T. I.; Ren Y.; Strickland M. R.; Lamothe R.; Schumacher S. E.; Tsherniak A.; Besche H. C.; Rosenbluh J.; Shehata S.; Cowley G. S.; Weir B. A.; Goldberg A. L.; Mesirov J. P.; Root D. E.; Bhatia S. N.; Beroukhim R.; Hahn W. C. Cancer vulnerabilities unveiled by genomic loss. Cell 2012, 150, 842–854. 10.1016/j.cell.2012.07.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang T.; Birsoy K.; Hughes N. W.; Krupczak K. M.; Post Y.; Wei J. J.; Lander E. S.; Sabatini D. M. Identification and characterization of essential genes in the human genome. Science 2015, 350, 1096–1101. 10.1126/science.aac7041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang W. S.; Shimada K.; Delva D.; Patel M.; Ode E.; Skouta R.; Stockwell B. R. Identification of simple compounds with microtubule-binding activity that inhibit cancer cell growth with high potency. ACS Med. Chem. Lett. 2012, 3, 35–38. 10.1021/ml200195s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazurkiewicz M.; Hillert E. K.; Wang X.; Pellegrini P.; Olofsson M. H.; Selvaraju K.; D’Arcy P.; Linder S. Acute lymphoblastic leukemia cells are sensitive to disturbances in protein homeostasis induced by proteasome deubiquitinase inhibition. Oncotarget 2017, 8, 21115–21127. 10.18632/oncotarget.15501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kobayashi A.; Kang M. I.; Okawa H.; Ohtsuji M.; Zenke Y.; Chiba T.; Igarashi K.; Yamamoto M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol. 2004, 24, 7130–7139. 10.1128/MCB.24.16.7130-7139.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.