Abstract
The emergence of CRISPR nucleases has transformed biotechnology by providing an easy, efficient and versatile platform for editing DNA. However, traditional CRISPR-based technologies initiate editing by activating DNA double-strand break repair pathways, which can cause adverse effects in cells and restrict certain therapeutic applications of the technology. To this end, several new CRISPR-based modalities have been developed that are capable of catalyzing editing without the requirement for a double-strand break. Here we review three of these technologies – base editors, prime editors and RNA-targeting Cas13 effectors. We discuss their strengths compared to traditional gene-modifying systems, we highlight their emerging therapeutic applications, and we examine challenges facing their safe and effective clinical implementation.
Keywords: CRISPR, Gene Therapy, Base Editing, Prime Editing, CRISPR-Cas13
Next-generation CRISPR technologies: the need to move beyond the DNA double-strand break for therapeutic gene-editing
The rise of programmable and sequence-specific technologies capable of correcting, replacing and deleting specific DNA sequences in cells has transformed modern biology, making the creation of designer cell lines and whole organisms a largely routine endeavor and providing a highly efficient means for implementing therapeutic gene correction. The core technology behind these advances are targeted nucleases [1], which stimulate the process of gene-editing by introducing a targeted DNA double-strand break (DSB, see Glossary) that then activates cellular DNA repair pathways [2–4] which, in turn, can be harnessed to modify the targeted genomic site. Though several platforms have been developed to facilitate this process, including zinc-finger nucleases [5] and transcription activator-like effector nucleases [6], technologies derived from naturally occurring CRISPR (Clustered Regularly Interspaced Short Palindromic Repeat) systems have emerged as the method of choice for most genome engineers [7–11], as CRISPR systems are versatile, efficient and easy-to-use.
Traditional CRISPR technologies typically consist of two components: a Cas protein (CRISPR-associated protein) that cleaves nucleic acids and a single guide RNA (sgRNA) that binds to the Cas protein and directs it to a specific nucleic acid sequence – always adjacent to a conserved and compatible protospacer adjacent motif (PAM) or protospacer flanking site (PFS) – for its cleavage [9]. Because of the general efficiency and the overall specificity with which they carry out this process, CRISPR technologies have emerged as promising therapeutic modalities. In fact, within only six years of the first reports demonstrating their ability to edit DNA in mammalian cells [8, 10, 11], CRISPR systems were used to successfully edit immune cells which were then safely transplanted to cancer patients [12, 13]. Clinical trials aimed at evaluating the ability of CRISPR nucleases to safely correct sickle cell disease (ClinicalTrials.gov Identifier: NCT03745287) and β-thalassemia (NCT03655678), two inherited blood disorders, as well as Leber congenital amaurosis type 10 (NCT03872479), an inherited retinal disorder, have also been initiated, which further demonstrates the therapeutic potential of the technology. However, despite the immense promise that CRISPR nucleases hold for treating a number of diseases, their reliance on DNA breaks to stimulate the gene-editing process could undermine their safety and, in some cases, their efficacy (Box 1). For example, it is well appreciated that DSBs induced by CRISPR nucleases have the potential to induce adverse effects in cells. These outcomes can include chromosomal translocations [14–16], genomic deletions [15, 17], reduced fitness [18], cell cycle dysfunction [19] and the activation of the tumor suppressor protein TP53 [20, 21]. Additionally, homology-directed repair (HDR), the DNA repair pathway most often used by CRISPR nucleases to faithfully introduce a specific change to a DNA sequence, is often limited by its efficiency in certain cell types [22–24]. For instance, HDR, which functions by incorporating genetic material from a co-delivered donor template into the target site, is typically only capable of achieving therapeutic rates of modification in dividing cells, which can prevent its implementation for therapeutic gene correction in post-mitotic cells [25]. Importantly, HDR can be further limited by its competition with non-homologous end joining (NHEJ), an error-prone DNA repair pathway that can create mutagenic base insertions and deletions (indels) at the target site [26, 27]. Recently however, several new technologies have emerged that hold the potential to overcome many of these fundamental limitations. These tools in particular possess the programmability and flexibility characteristic of CRISPR nucleases but carry expanded functional capabilities that enable editing without the traditional requirement of a DSB. Here we review three of these modalities: base editors, prime editors and Cas13 effectors. We discuss the individual strengths of these technologies compared to traditional gene-editing systems, and we examine their emerging applications for gene and cell therapy.
Box 1. Editing DNA with nickases.
Similar to DSBs, single-stranded breaks in DNA, known as nicks, can stimulate HDR. To this end, nickase enzymes, including Cas9 nickases, can be used to create targeted DNA nicks that can facilitate the integration of DNA but without a DSB [110–112]. However, this approach relies on HDR to mediate editing and thus remains restricted to dividing cells.
Base editors as a therapeutic gene-editing modality: early pre-clinical successes and future challenges
Mechanisms for DNA editing
CRISPR base editors are a technology that can facilitate the introduction of targeted point mutations in DNA, but without the requirement for a DSB. Most broadly, base editors consist of fusions of a Cas9 nickase (nCas9), a variant of the Cas9 nuclease that creates a single-strand rather than a double-strand break, with a nucleobase deaminase enzyme that initiates base editing by catalyzing a targeted deamination reaction. To date, several different base editors have been developed for use in mammalian cells (Fig. 1A,B), including cytosine base editors (CBEs), which can catalyze the deamination of a target cytosine to facilitate its conversion to a thymine [28, 29], adenosine base editors (ABEs), which can deaminate a target adenosine to facilitate its transition to a guanosine [30] and, most recently, guanosine base editors (GBEs), which can create C > G transversions in certain settings [31].
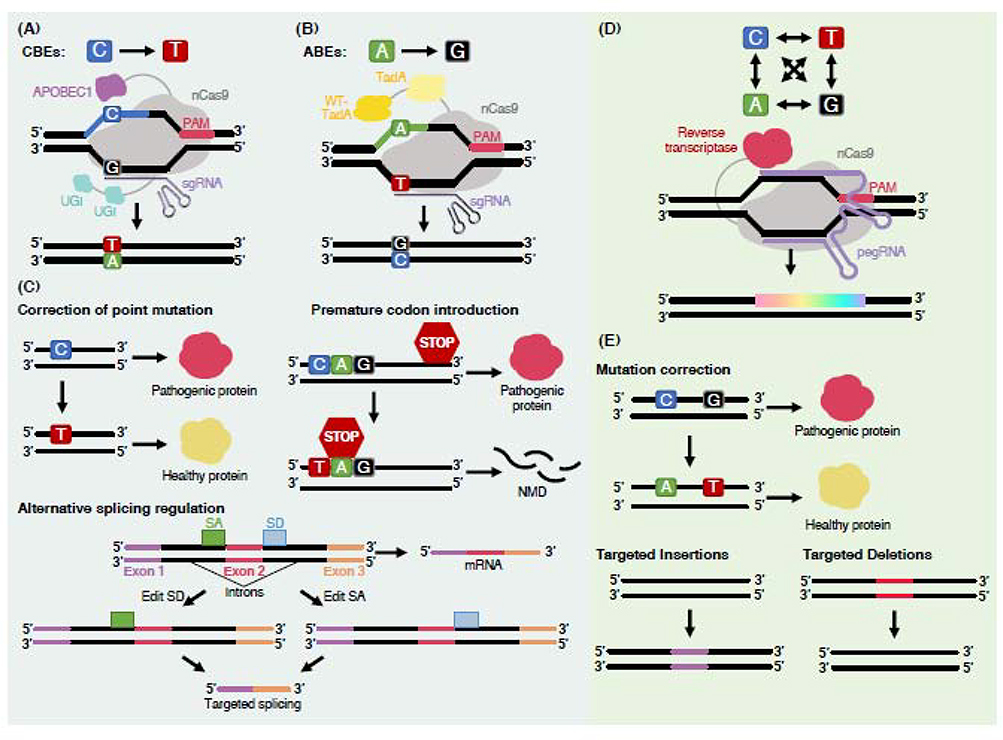
Figure 1. Overview of next-generation DNA editing technologies.
(A) Fourth generation CBEs (BE4) consist of fusions of nCas9 with a cytosine deaminase domain (APOBEC1) that catalyzes the deamination of cytosine and two uracil DNA glycosylase inhibitor (UGI) domains, which can prevent the unwanted excision of uracil by endogenous uracil N-glycosylase enzymes. To catalyze base editing, CBEs are directed to a specific DNA sequence via an sgRNA. Then, following nCas9 binding and denaturation of the target DNA sequence, the APOBEC1 enzyme binds to cytosines within a stretch of sequence in the exposed DNA strand and catalyzes a deamination reaction that results in the conversion of cytosine to uracil, which is recognized by cells as thymine during replication and/or repair. (B) ABE7.10 catalyzes A > G transitions and consists of a tandem fusion of a wild type tRNA adenosine deaminase (TadA) and an evolved TadA domain, engineered to accept DNA as a substrate, fused with nCas9. Deamination of adenosine yields inosine, which is recognized by cells as guanosine. (C) Therapeutic outcomes of base editing include: (1) the correction of single point mutations that encode for pathogenic proteins, (2) the introduction of premature stop codons to eliminate pathogenic transcripts via the nonsense-mediated decay (NMD) surveillance pathway, and (3) regulation of mRNA splicing by editing splicing motifs – SA and SD – to induce exon skipping. (D) Prime editors (PEs) consist of a prime editing guide RNA (pegRNA), which both specifies the target sequence and encodes the desired edits, and nCas9 fused to a reverse transcriptase (RT) domain, which promotes the incorporation of the edit encoded in the pegRNA template into the target DNA sequence. The RT fusion provides high levels of design versatility for achieving a range of editing outcomes. (E) Examples of therapeutic prime editing outcomes include: (1) the correction of single or multiple mutations that result in pathogenic proteins and (2) the insertion or deletion of pathogenic sequences. Additional abbreviations are as follows: SA, splice acceptor site; SD, splice donor site.
To catalyze a single-base modification, base editors, as directed by the sgRNA, first bind to a specific genomic target. Following the formation of a nCas9-sgRNA-DNA ternary complex that denatures the local target DNA sequence, the deaminase enzyme binds to its cognate base(s), typically within a narrow stretch or window of nucleotides in the exposed DNA strand, and then catalyzes a deamination reaction that, in the case of a CBE, results in the transition of cytosine to uracil, a nucleobase which is recognized by cells as thymidine. The resulting U-G base-mismatch is then resolved into the target T-A pair via cellular DNA replication or repair mechanisms that are activated in part by nCas9-induced nicking of the non-edited strand. Importantly, the enzymatic machinery and the mechanism(s) underlying this process vary by the base editor. For example, the most efficient ABEs rely on an engineered tRNA adenosine deaminase (TadA) enzyme that deaminates a target adenosine in an exposed DNA strand to yield an inosine intermediate, which is then recognized as guanosine during repair. However, unlike with HDR, the repair mechanisms underlying these processes appear to be sufficiently active in both dividing and non-dividing cells [32], which suggests that base editors could serve as a broadly applicable therapeutic modality.
Therapeutic applications of base editing
Given their considerable strengths and their potential to enable new opportunities for therapeutic editing, base editors have been rapidly deployed in vivo to treat a number of disorders (Table 1). Among the first successful examples demonstrating their potential was the finding that ABEs could be delivered to skeletal muscle to restore dystrophin expression in a mouse model of Duchenne muscular dystrophy (DMD) [33], a genetic disorder characterized by progressive muscle degeneration and weakness due to mutations in the dystrophin protein. Specifically, an ABE was used to revert a premature stop codon in a mutant dystrophin gene via A > G editing, which resulted in restored dystrophin expression. However, since most base editor proteins exceed the carrying capacity of a single adeno-associated virus (AAV) vector (Table 2), a particularly promising in vivo gene delivery vehicle [34], two AAV particles were needed to deliver the ABE in vivo. In this particular case, the base editor transgene was split in two halves and delivered across two AAV vectors that were subsequently joined together by a recombination reaction between the inverted terminal repeat (ITR) sequences present in each vector [35, 36] (Fig. 2A).
Table 1.
Pre-clinical gene and cell therapy applications of next-generation CRISPR technologies.
| Disorder | Strategy | Delivery | Therapeutic outcomes | Ref. |
|---|---|---|---|---|
| DMD | Base editor-mediated correction of nonsense mutation | In vivo via dual AAV vectors | Restored dystrophin expression in myofibers in an adult DMD mouse model | [33] |
| PKU | Base editor-mediated correction of nonsense mutation | In vivo via dual AAV vectors | Restored physiological blood L-Phe levels, enhanced PAH enzyme activity, and reversed a light fur phenotype in PAHenu2 adult mice | [40] |
| Niemann-Pick disease type C | Base editor-mediated correction of loss of function mutation | In vivo via dual AAV vectors | Slowdown neurodegeneration and increased lifespan in Npc1I1061T homozygous mice | [41] |
| Deafness | Base editor-mediated correction of loss of function mutation | In vivo via dual AAV vectors | Restored inner hair cell sensory transduction and hair cell morphology and transiently rescued low-frequency hearing in neonatal mice carrying a loss-of-function point mutation in TMC1 | [42] |
| Coronary heart disease | Base editor-mediated gene silencing | In vivo via an adenoviral vector | Reduced plasma PCSK9 protein levels and reduced plasma cholesterol levels in adult mice | [53] |
| Base editor-mediated gene silencing | In utero via an adenoviral vector | Reduced plasma PCSK9 protein levels and reduced plasma cholesterol levels in | [52] | |
| Base editor-mediated gene silencing | In vivo via functionalized lipid-like nanoparticles | Reduced postnatal serum PCSK9 protein levels in mice | [113] | |
| ALS | Base editor-mediated gene silencing | In vivo via dual AAV vectors | Reduced the rate of muscle atrophy and muscle denervation, improved neuromuscular function, and slowed disease progression in the G93ASOD1 mouse model of ALS | [54] |
| HTI | Base editor-mediated gene silencing | In utero via an adenoviral vector | Rescue of the lethal phenotype of HTI, as indicated by weight gain and prolonged survival in Fah–/– mice, a model of HTI | [52] |
| Base editor-mediated splice site correction | In vivo via hydrodynamic injection of plasmid DNA | Partially restored splicing, rapid expansion of Fah+ hepatocytes in the liver and rescued weight loss in adult Fahmut/mut mice | [45] | |
| In vivo via lipid nanoparticle delivery | Edited hepatocytes in adult Fahmut/mut mice | |||
| Base editor-mediated correction of mutation | In vivo via lipid nanoparticle delivery | Rescued weight loss and restored Fah expression in liver tissue of mice carrying a mutant version of Fah | [44] | |
| Base editor-mediated generation of de novo in frame start codon | In vivo via dual AAV vectors | Restored Fah expression and halted weight loss in an HTI mouse model | [43] | |
| β-thalassemia and SCD | Base editing | Ex vivo via electroporation of RNP | Engraftment of edited HSCs in NBSGW mice resulted in HbF induction | [56] |
| Pancreatic cancer | Cas13-mediated transcript knockdown | In vivo via an AAV vector | Slowed tumor growth and reduced tumor cell proliferation in mice bearing subcutaneous AsPC-1 xenografts | [98] |
| Retinal injury | Cas13-mediated transcript knockdown | In vivo via an AAV vector | Induced the conversion of glia cells into retinal ganglion cells, which resulted in partially restored visual responses in a drug-induced retinal injury mouse model | [99] |
| PD | Cas13-mediated transcript knockdown | In vivo via an AAV vector | Induced the conversion of glia cells into dopaminergic neurons, which resulted in reduced motor dysfunction in a PD mouse model created by inducing lesions with 6-OHDA | [99] |
Additional abbreviations are as follows: DMD, Duchenne muscular dystrophy; PKU, phenylketonuria; L-Phe, phenylalanine; PAH, phenylalanine hydroxylase; TMC1, transmembrane channel-like 1; PCSK9, proprotein convertase subtilisin/kexin type 9; ALS, Amyotrophic lateral sclerosis; SOD1, superoxide dismutase 1; HTI, hereditary tyrosinemia type I; Fah, fumarylacetoacetate hydrolase; SCD, sickle cell disease; RNP, ribonucleoprotein; HSCs, hematopoietic stem cells; HbF, fetal hemoglobin; PD, Parkinson’s disease; 6-OHDA, 6-hydroxydopamine.
Table 2.
Delivery strategies for CRISPR technologies
| Viral methods | ||||||||
| Vector | Integrating / non-integrating | Pathogenicity | Packaging capacity | Duration of Expression | Cargo | Diameter | Primary setting for genome-editing | References |
| Lentivirus | Integrating* | Low to moderate | ~ 8 kb | Long-term | RNA | ~ 130 nm | Ex vivo | [114] |
| Adenovirus | Non-integrating | Moderate to high | ~ 8 kb | Long-term | DNA | ~ 100 nm | In vivo | [114] |
| AAV | Can integrate at low frequencies (0.1–1%) | Low | ~ 4.7 kb | Long-term in quiescent cells | DNA | ~ 25 nm | In vivo | [114] |
| Non-viral methods | ||||||||
| System | Composition | Pathogenicity | Distribution | Duration of Expression | Cargo | Size | Primary setting for genome-editing | References |
| Lipid nanoparticles | Ionizable lipids, cholesterol, PEG, DOPE, supplemental cationic components | Low | - Systemic administration (liver, lung)** - Local administration |
Short-term | Protein DNA, RNA | ~150–200 nm | In vivo | [115] |
| Lipid-like nanoparticles | TT derivatives, PEG, DOPE, | Low | - Systemic administration (liver and blood) | Short-term | RNA | ~100 nm | In vivo | [113, 116, 117] |
| Gold nanoparticles | Gold nanoparticles conjugated to thiol modified DNA | Low | Local administration | Short-term | Protein DNA, RNA | ~500 nm | In vivo | [118] |
| Electroporation | Wave pulse | Low | Local administration (skin, retina) | Short-term | Protein DNA, RNA | NA | In vivo, ex vivo | [119, 120] |
Integration-deficient lentiviruses also exist
Cationic component level-dependent.
Additional abbreviations are as follows: DOPE, 1,2-dioleyl-sn-glycero-3-phosphoethanolamine; TT, N1,N3,N5-tris(2-aminoethyl)benzene-1,3,5-tricarboxamide; NA, not applicable.
Figure 2. Methods of reconstituting large gene-editing proteins delivered by dual AAV vectors.
(A) ITR recombination can mediate the joining of two ITR-flanked viral genomes that each encode a portion of a base editor sequence. In the case of an ABE, the first vector encodes: (1) an sgRNA driven by the U6 promoter, (2) a promoter to drive expression of the NLS-tagged tandem TadA domains fused to the N-terminal (NT) half of a split nCas9 transgene and a splice donor (SD) sequence, while the second vector carries: (1) a splice acceptor (SA) signal sequence, (2) the C-terminal (CT) half of the split nCas9 fused to an NLS tag and an HA epitope tag and (3) a bGH polyadenylation (polyA) signal sequence. Once in the target cell, the two viral vectors are joined by a recombination reaction between the two ITRs and the full-length ABE pre-mRNA is transcribed. pre-mRNA processing removes the intron and the intervening ITR, resulting in the translation of a full-length ABE protein. (B) Intein-mediated protein trans-splicing of two separately expressed base editor halves can be used to assemble a full-length base editor protein. In the case of a CBE, the first vector encodes: (1) a promoter to drive expression of the NT of the base editor, (2) a V5 epitope and NLS tag followed by the APOBEC1 domain fused to the NT fragment of the split nCas9 half and a connected N-terminal intein domain, and (3) a bGH poly A signal sequence, while the second vector carries: (1) a promoter to drive the expression of the C-terminal intein domain fused to the CT half of nCas9 linked to two UGI domains, an NLS and an HA epitope tag, and (3) a poly A signal sequence. This vector also encodes an sgRNA expression cassette. Both vectors are packaged and co-delivered in vivo, where, once internalized by the cells, the separate intein-split base editor protein halves are expressed. Following their translation, the N and C intein fragments associate and catalyze a trans-splicing reaction that removes the intein moieties and results in the formation of the full-length CBE. Additional abbreviations are as follows: TadA, tRNA adenosine deaminase; NLS, nuclear localization signal; ITR, inverted terminal repeat; bGH, bovine growth hormone; APOBEC1, cytosine deaminase domain.
As an alternative to vector-mediated recombination, intein-mediated protein trans-splicing, an approach that harnesses self-splicing intein domains to catalyze the ligation of two separately expressed polypeptides into a single protein chain, has also been used to reassemble two base editor halves into the full-length protein [37–39] (Fig. 2B). The potential of intein-mediated trans-splicing for in vivo gene correction was in fact first demonstrated in a mouse model of phenylketonuria, a recessive liver disease that is caused by mutations in the phenylalanine hydroxylase (PAH) gene. In this particular case, dual vector delivery to the liver of an intein-containing split CBE led to the correction of up to 25% of mutant PAH alleles [40]. Similar intein-mediated trans-splicing approaches have been employed to deliver base editors to the brain to correct loss-of-function point mutations in the Npc1 gene, which gives rise to the lysosomal storage disorder Niemann-Pick disease type C [41] and to the inner ear to correct a mutation in the TMC1 gene in a neonatal mouse model of hereditary deafness, which resulted in partially restored auditory function [42]. Additionally, intein-mediated trans-splicing has been utilized to create an in-frame start codon in the fumarylacetoacetate hydrolase (Fah) gene to restore its expression and mitigate a pathogenic phenotype in a murine model of hereditary tyrosinemia type I (HTI) [43], further solidifying the potential of dual vector delivery for enabling therapeutic base editing. Notably, non-viral approaches (Table 2) involving lipid nanoparticles (LNPs) have also been used to deliver base editor-encoding mRNA and have enabled correction in a mouse model of HTI [44, 45]. Importantly, LNP-delivered mRNA is only transiently maintained, which can reduce the risk for accumulating off-target effects from persistently expressing a base editor protein in cells from a viral vector.
In addition to correcting loss-of-function mutations, base editors can also be used to silence gene expression in a manner that overcomes a key limitation underlying the use of CRISPR-Cas9 nucleases for the same application, namely their reliance on NHEJ. Nuclease-induced NHEJ, in particular, carries a risk for creating in-frame mutations in the target gene that could lead to the production of unique mutant protein isoforms, which can not only diminish the therapeutic efficacy of this approach but, in some cases, also potentially cause adverse effects [46–48]. In contrast, by catalyzing targeted C > T transitions at CGA, CAG, CAA and TGG triplets, CBEs can create premature stop codons that can result in the more uniform degradation of the target mRNA by nonsense-mediated decay (NMD) [49, 50], a surveillance mechanism used by cells to prevent the formation of truncated proteins (Fig. 1C). Along similar lines, ABEs can be used to prevent gene expression by altering the ATG start codon [51].
The therapeutic potential of this approach has now been demonstrated in multiple studies. Specifically, adenoviral (Ad) vectors, which, unlike AAV, can accommodate the delivery of a full-length base editor protein within a single viral particle (Table 2), have been used to deliver base editors to the liver to silence PCSK9, a protein that can regulate LDL receptor degradation and has emerged as a potential target for hypercholesterolemia, in both adult mice [52] and prenatal mice [53]. Similarly, a prenatal strategy was undertaken in mice to silence the HPD gene to rescue hereditary tyrosinemia type 1 [53], a metabolic disorder characterized by elevated blood levels of the amino acid tyrosine. These studies altogether demonstrated the potential of base editors to not only silence gene expression in vivo by introducing nonsense mutations, but also the potential of in utero gene therapy for correcting inherited genetic disorders.
In addition to reprogramming metabolic flux in vivo, base editors can be used to silence the expression of genes with toxic gain-of-function mutations. Specifically, dual AAV particle delivery of a split-intein CBE engineered to disable a mutant gene causative for a form of amyotrophic lateral sclerosis, a currently incurable disorder that involves the selective loss of motor neurons in the spinal cord and brain, prolonged survival and slowed the deterioration of motor function in a mouse model of the disease [54], further reinforcing the potential of dual vector delivery approaches for achieving meaningful therapeutic effects in preclinical rodent models.
Similar to CRISPR nucleases, base editors also have the potential to advance cell therapy. For instance, base editors have been used to simultaneously disable the expression of multiple targets known to boost the potency of allogeneic CAR-T cells, including the TRAC locus, the β-2 microglobulin gene, and the programmed cell death 1 gene, which altogether resulted in not only a more efficient CAR-T cell therapy but also higher quality cells, as measured by a reduction in DSB-induced translocations compared to Cas9-edited cells [55]. Along similar lines, base editors have been used to correct inherited blood disorders by ex vivo gene therapy. This was achieved by electroporating purified ribonucleoproteins (RNPs) to cells, which, unlike base editors expressed from viral vectors, exist in cells only transiently. In this case, C > T editing in blood progenitor cells was used to activate fetal hemoglobin by mutating a key transcription factor binding site within the BCL11A enhancer, a repressor of fetal hemoglobin [56]. Thus, base editing technologies can be used to tune gene expression by modifying functional elements within non-coding sequences.
Moving forward, the finding that base editors can modulate alternative splicing by modifying conserved intronic motifs in genes [57, 58], which can include both donor and acceptor signals, should extend their use to therapeutic exon skipping, as demonstrated against the mutant dystrophin gene in patient-derived induced pluripotent stem cells (iPSCs) [58] (Fig. 1C).
Challenges for therapeutic base editing
While the therapeutic potential for base editing has been demonstrated now in multiple studies, the clinical implementation of the technology, like CRISPR-Cas9 nucleases and other gene-editing modalities before them, must overcome a number of hurdles, particularly with respect to safety and in vivo efficacy. In particular, it is now well appreciated that, in addition to both Cas9:sgRNA-dependent [28, 59–62] and independent [63] off-target effects in DNA, base editors can non-specifically modify RNA [64]. Therefore, a number of protein engineering strategies have been implemented to create base editor scaffolds with improved specificity, including variants with not only decreased off-target DNA editing activity [62, 65–69] but also reduced RNA targeting capabilities [64, 69–71]. Additionally, base editors can sometimes create an incorrect edit at the target base or produce bystander mutations via their editing of non-target bases within the targeted sequence. As a result, base editor variants with improved on-target editing capabilities have been developed. These proteins in fact have been reported to possess not only improved on-target editing frequencies [72–77] but also lower rates of unwanted bystander mutations [66, 68, 77, 78] and fewer indel byproducts [73, 75].
Additionally, as AAV vectors have emerged as a highly promising therapeutic gene delivery vehicle [34], it will be important to evaluate the effectiveness and safety of dual vector strategies for delivering split-intein-containing base editors in vivo, particularly within large animal models, which can better mimic many aspects of human physiology relevant to vector delivery than rodent models. Such models will also be important for studying the potential immunogenicity and long-term safety of base editor proteins. Finally, considering recent observations indicating their integration into the genome when delivering Cas9 [79, 80], it will be important to determine at what frequency base editor-encoding AAV vectors are also integrated into a host cells’ genome.
Prime editors: a newly emerged gene-editing technology with broad therapeutic potential
Though base editors have expanded the range of therapeutic applications possible for gene editing, the technology remains limited in part by the functional capabilities of current deaminase domains, which, while enabling transition mutations and C > G transversions, cannot currently insert or delete DNA sequences. However, a recently reported alternative gene-editing technology named ‘prime editing’ [81] holds the potential to overcome current functional limitations.
Prime editors (PEs) consist of nCas9 tethered to a reverse transcriptase (RT) domain and a modified sgRNA, referred to as a prime editing guide RNA (pegRNA), that not only specifies the target site for nCas9 but also serves as a template for the RT domain during DNA repair (Fig. 1D). To catalyze editing, PEs, as directed by the pegRNA, first bind to a specific target DNA sequence, which then facilitates nCas9-induced nicking of the target strand and hybridization of the nicked 3’ DNA strand with a complementary “primer binding site” encoded in the pegRNA. Following heteroduplexation, the RT domain then catalyzes the synthesis of a new DNA strand, which encodes the edit, from the template specified by the pegRNA. This new strand is then introduced into the target site following DNA repair. Currently the most efficient version of this system, termed PE3, consists of the PE platform described above and a second sgRNA that facilitates nicking of the non-edited strand, which can enhance the rate of editing.
Compared to base editors, PEs offer increased targeting flexibility, since they do not have as restricted an editing window, and the potential to generate all possible base transitions and transversions, thereby offering a means to correct a wide range of possible gene mutation (Fig. 1D), as evidenced by proof-of-principle studies demonstrating their ability to correct mutations known to cause sickle cell disease and Tay-Sachs disease in cell culture [81]. However, compared to similarly active CBEs and ABEs, PE3, the prime editing system currently found to be the most efficient for editing DNA, has been reported to generate increased rates of indel byproducts [81]. Because high product purity and a relative lack of indel mutations are both defining features of many single-base editing technologies, refining prime editors to generate fewer collateral indels will be a high priority moving forward. Additionally, streamlining the design of an optimal pegRNA configuration to minimize laboratory screening efforts is also likely a high priority, as recently demonstrated by two studies [82, 83]. Along these lines, machine learning strategies, which have proven effective for not only predicting Cas9-specific indel patterns [84, 85] but also base editor-mediated editing outcomes [86–88], could be used to improve pegRNA design and enable more efficient prime editing. Nonetheless, given their additional capacity for inserting and deleting new DNA sequences, prime editing technologies hold potential therapeutic editing applications beyond base correction.
However, while the versatility and the reported efficiency of PEs makes them an attractive platform for correcting potential disease-causing mutations, additional studies are required to comprehensively interrogate their genome-wide specificities, though it is likely that the combination of pegRNA-mediated targeting specificity and RT-based templating requirements will ensure similar, if not greater, specificity than observed using base editors. Nonetheless, the RT domain itself could induce off-target effects. Finally, similar to base editor proteins, efficiently delivering PEs via AAV could prove challenging given their large size. However, established dual vector delivery strategies that rely on ITR recombination [33] and intein-mediated protein trans-splicing [32, 40, 41, 54, 89] to reassemble the full-length PE protein should be compatible with the technology and enable its in vivo delivery for further preclinical study.
Targeting, skipping, and editing RNA using CRISPR-Cas13 effectors: expanding the CRISPR toolbox beyond DNA editing
Though CRISPR technology is most commonly associated with DNA editing, the discovery of Cas13 enzymes [90–95], which cleave a target RNA via an intrinsic RNase activity that is activated by the binding of a CRISPR RNA (crRNA) guide molecule, has facilitated the creation of a flexible and programmable toolbox capable for targeting RNA (Fig. 3A).
Figure 3. Overview of RNA-targeting CRISPR systems.
(A, left) To catalyze RNA cleavage, a CRISPR RNA (crRNA) molecule binds to Cas13 and directs it to a target RNA site, which stimulates the RNase catalytic activity of the HEPN domains of Cas13, triggering its cleavage. Cas13 systems can also mediate targeted splicing utilizing dCas13 and a single or multiple crRNA molecules that bind to splice site motifs – SA and SD – and likely block them from recognition by the splicing machinery. (A, right) Additionally, Cas13 effectors can serve as scaffolds to enable targeted RNA editing. RNA editors consist of fusions of dCas13 to adenosine deaminases acting on RNA (ADAR) to stimulate A to I edits and cytidine deaminase domains to mediate C to U conversions. (B) Examples of therapeutic RNA targeting outcomes include the use of Cas13 protein to cleave pathogenic transcripts to prevent the translation of mutated proteins and the use of dCas13 to bind splice donor and acceptor sites to block spliceosome protein complexes from triggering exon inclusion that may encode for pathogenic isoforms. Additionally, RNA editing can be used to correct single-point mutations in mRNA that encode for pathogenic proteins. Abbreviations as follows: E1, exon 1; E2, exon2; E3, exon 3; SA, splice acceptor site; SD, splice donor site.
From a therapeutic perspective, similar to antisense oligonucleotides (ASOs) and RNA interference (RNAi), CRISPR-Cas13 effectors offer a means to suppress gene expression without the risk for inducing DNA damage to cells, as Cas13 proteins have been demonstrated to generally lack DNase activity [96]. Cas13 targeting is also, in theory, reversible. For example, because Cas13 engages with RNA, its expression could potentially be terminated by an engineered kill-switch, which, if implemented, could return the cell’s transcriptome back to its original state. Cas13 proteins may also offer distinct targeting advantages to RNAi, a clinically promising platform capable of efficiently knocking down target gene expression: for example, side-by-side comparisons with spacer-matching short-hairpin RNAs (shRNAs) have revealed that Cas13d proteins induce fewer off-target effects in cultured cells, indicating that they may have superior targeting abilities [90, 96, 97], though additional studies comparing Cas13 to pre-clinically optimized gene-silencing technologies will be needed to establish this.
Given these advantages, Cas13 proteins have been deployed in vivo to facilitate therapeutic outcomes (Table 1). For instance, repeated injections of a Cas13 RNP designed to target a mutant KRAS transcript that can trigger cancer development was demonstrated to shrink the mass of a tumor xenograft [98]. However, because of their compact size, certain Cas13 effectors, such as the Cas13 protein from Ruminococcus flavefaciens (RfxCas13), can be packaged into a single AAV vector particle to enable continuous in vivo targeting. In fact, RfxCas13 has been delivered in vivo by AAV to knockdown PTBP1, a suppressor of neuronal differentiation, to mediate the conversion of retinal glial cells into ganglion cells, a type of neuron located in the retina, to restore visual responses in a retinal injury mouse model [99]. A similar strategy was undertaken to induce the expression of dopaminergic neurons in the brain to slow motor dysfunction in a mouse model of Parkinson’s disease [99]. In both cases, Cas13 targeting was found to induce few off-target effects.
In addition to knocking down a target mRNA, Cas13 proteins can effectively modulate alternative splicing (Fig. 3B). In particular, catalytically inactivated Cas13 protein variants (dubbed dCas13) can be directed by a crRNA to bind to – and thus interfere with – positive and negative splicing factors to mediate the inclusion or exclusion of specific exon(s) [96, 100]. The therapeutic potential of this approach was demonstrated by multiplexed targeting of a splice acceptor site and two putative exonic splice enhancer sites in MAPT pre-mRNA, which facilitated correction of a mis-splicing event in a patient-derived neuronal model of dementia [96], and the restoration of exon 7 splicing within the survival motor neuron 2 gene in spinal muscular atrophy (SMA) patient fibroblasts [100].
Additionally, akin to current base editing technologies, Cas13 can also serve as a scaffold to direct RNA editing in cells (Fig. 3A). Specifically, dCas13 proteins have been fused to adenosine deaminases acting on RNA (ADAR) domains to mediate A > I conversions in a target RNA [97] and an evolved ADAR domain that instead acts as a cytidine deaminase to create C > U edits [101]. Though the full therapeutic potential of Cas13-based RNA editing has yet to be harnessed, RNA editing systems have been used to correct a point mutation in MECP2 mRNA, which enabled rescue of certain phenotypes associated with Rett syndrome in a mouse model of the disease [102].
Moving forward however, the clinical implementation of Cas13 effectors could face several obstacles. In particular, similar to ASOs and RNAi, Cas13 proteins must continuously engage with a target RNA to sustain their therapeutic effect. Given the finding that many individuals may have pre-existing antibodies and/or reactive T cells to CRISPR proteins [103, 104], the persistent expression of Cas13 proteins could pose a risk for stimulating a specific immune response (though the surface of Cas13 proteins could be re-engineered to potentially evade common neutralizing responses [105]). Cas13 proteins may carry additional risks owing to the fact that, in certain situations, they have been reported to cleave non-target RNAs after activation [91, 106]. Thus, long-term study of the immunogenicity and targeting specificity of Cas13 proteins will be important for determining their potential as a therapeutic.
Concluding Remarks and Future Perspectives
The past several years has seen a rapid rise in the number of technologies capable of editing DNA and RNA with both high precision and with high frequency. Owing to their potential to correct disease-causing mutations via repair mechanisms that are generally available in most cell types, these technologies hold immense potential as therapeutics for a range of disorders. It should be noted though that many of these platforms have overlapping capabilities. For instance, both base editors and prime editors are capable of creating C > T and A > G transitions. Thus, when identifying a target editor, multiple factors, including on-target editing efficiency and non-target editing byproducts should all be considered and weighed appropriately. Additionally, despite the rapid emergence of base editor scaffolds with improved targeting capabilities, it likely will be necessary to thoroughly optimize individual base editor proteins for their specific therapeutic targets in order to ensure maximally efficient formation of the on-target product and to further minimize potentially counterproductive non-target editing outcomes.
In addition to developing methods to further refine the capabilities of these increasingly sophisticated gene-editing machines, special attention must be paid to their delivery [107]. In particular large animal studies will be needed to determine the overall effectiveness for dual AAV-based strategies for delivering base editors and potentially prime editors. Additionally, any optimization of a gene-editing protein should go hand-in-hand with optimization of: (1) the viral capsid to ensure efficient delivery to the most relevant cell populations, and (2) the promoter sequence to enable efficient and also preferably target cell type specific editing. Effectively implementing next-generation CRISPR technologies will also require extensively assessing their immunogenicity. Specifically, while DNA-editing technologies have, in theory, the capacity to self-inactivate their own expression, RNA-targeting technologies must be continuously expressed in order to sustain a therapeutic effect, which could increase their risk for eliciting an immune response. Studies in large animal models will therefore be important for helping to establish whether continuously expressing Cas13 or any other next-generation CRISPR technology causes long-term adverse effects (see Outstanding Questions). Finally, in addition to the single-base DNA and RNA editing technologies described here, targetable technologies for inserting large transgenes into DNA through RNA-guided transposition mechanisms [108, 109] are actively being developed and could soon enable a new means of accomplishing therapeutic gene integration into safe-harbor sites, among other possible applications.
Outstanding Questions.
Can the expression of next-generation CRISPR technologies be effectively controlled in vivo to prevent adverse effects?
Can base editing technologies be engineered to possess the specificity and efficiency needed to safely edit DNA in vivo?
Will prime editing technology emerge as an effective gene therapy agent?
Which delivery method(s) will prove most effective for enabling in vivo base editing and prime editing?
Will the long-term expression of Cas13 proteins cause adverse effects?
In summary, next-generation DNA and RNA editing technologies have enormous potential to advance gene and cell therapies.
Highlights.
Next-generation CRISPR technologies hold the potential to expand the capabilities of therapeutic editing.
Advanced CRISPR modalities harness the versatility and programmability of traditional CRISPR systems but can catalyze highly precise editing outcomes without a DNA double-strand break.
Next-generation CRISPR technologies include base editors, prime editors and RNA-targeting Cas13 effectors.
The continued refinement of these technologies could enable their safe and effective implementation to treat a range of disorders.
Acknowledgements
M.A.Z.C. was supported by the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health (T32EB019944), the Mavis Future Faculty Fellows Program and by a University of Illinois Aspire Fellowship. This work was also supported by the Muscular Dystrophy Association (MDA602798).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- 1.Kim H and Kim JS (2014) A guide to genome engineering with programmable nucleases. Nat Rev Genet 15 (5), 321–34. [DOI] [PubMed] [Google Scholar]
- 2.Lukacsovich T et al. (1994) Repair of a specific double-strand break generated within a mammalian chromosome by yeast endonuclease I-SceI. Nucleic Acids Res 22 (25), 5649–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Rouet P et al. (1994) Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc Natl Acad Sci U S A 91 (13), 6064–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wyman C and Kanaar R (2006) DNA double-strand break repair: all’s well that ends well. Annu Rev Genet 40, 363–83. [DOI] [PubMed] [Google Scholar]
- 5.Urnov FD et al. (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11 (9), 636–46. [DOI] [PubMed] [Google Scholar]
- 6.Joung JK and Sander JD (2013) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14 (1), 49–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cho SW et al. (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31 (3), 230–2. [DOI] [PubMed] [Google Scholar]
- 8.Cong L et al. (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339 (6121), 819–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jinek M et al. (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337 (6096), 816–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Jinek M et al. (2013) RNA-programmed genome editing in human cells. Elife 2, e00471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mali P et al. (2013) RNA-guided human genome engineering via Cas9. Science 339 (6121), 823–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Stadtmauer EA et al. (2020) CRISPR-engineered T cells in patients with refractory cancer. Science 367 (6481). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lu Y et al. (2020) Safety and feasibility of CRISPR-edited T cells in patients with refractory non-small-cell lung cancer. Nat. Med. [DOI] [PubMed] [Google Scholar]
- 14.Brunet E and Jasin M (2018) Induction of chromosomal translocations with CRISPR-Cas9 and other nucleases: understanding the repair mechanisms that give rise to translocations. Adv Exp Med Biol 1044, 15–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kosicki M et al. (2018) Repair of double-strand breaks induced by CRISPR-Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol 36 (8), 765–771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rayner E et al. (2019) CRISPR-Cas9 causes chromosomal instability and rearrangements in cancer cell lines, detectable by cytogenetic methods. CRISPR J. 2 (6), 406–416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cullot G et al. (2019) CRISPR-Cas9 genome editing induces megabase-scale chromosomal truncations. Nat. Commun. 10 (1), 1136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chen X et al. (2020) Expanding the editable genome and CRISPR-Cas9 versatility using DNA cutting-free gene targeting based on in trans paired nicking. Nucleic Acids Res 48 (2), 974–995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.van den Berg J et al. (2018) A limited number of double-strand DNA breaks is sufficient to delay cell cycle progression. Nucleic Acids Res. 46 (19), 10132–10144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Haapaniemi E et al. (2018) CRISPR-Cas9 genome editing induces a p53-mediated DNA damage response. Nat Med. [DOI] [PubMed] [Google Scholar]
- 21.Ihry RJ et al. (2018) p53 inhibits CRISPR-Cas9 engineering in human pluripotent stem cells. Nat Med. [DOI] [PubMed] [Google Scholar]
- 22.Cox DB et al. (2015) Therapeutic genome editing: prospects and challenges. Nat Med 21 (2), 121–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nami F et al. (2018) Strategies for In Vivo Genome Editing in Nondividing Cells. Trends Biotechnol 36 (8), 770–786. [DOI] [PubMed] [Google Scholar]
- 24.Rothkamm K et al. (2003) Pathways of DNA double-strand break repair during the mammalian cell cycle. Mol Cell Biol 23 (16), 5706–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lin S et al. (2014) Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. Elife 3, e04766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Howden SE et al. (2015) Simultaneous Reprogramming and Gene Correction of Patient Fibroblasts. Stem Cell Reports 5 (6), 1109–1118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Merkle FT et al. (2015) Efficient CRISPR-Cas9-mediated generation of knockin human pluripotent stem cells lacking undesired mutations at the targeted locus. Cell Rep 11 (6), 875–883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Komor AC et al. (2016) Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533 (7603), 420–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nishida K et al. (2016) Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353 (6305). [DOI] [PubMed] [Google Scholar]
- 30.Gaudelli NM et al. (2017) Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kurt IC et al. (2020) CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat Biotechnol. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yeh WH et al. (2018) In vivo base editing of post-mitotic sensory cells. Nat. Commun. 9 (1), 2184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ryu SM et al. (2018) Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat Biotechnol 36 (6), 536–539. [DOI] [PubMed] [Google Scholar]
- 34.Wang D et al. (2019) Adeno-associated virus vector as a platform for gene therapy delivery. Nat Rev Drug Discov 18 (5), 358–378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lai Y et al. (2005) Efficient in vivo gene expression by trans-splicing adeno-associated viral vectors. Nat Biotechnol 23 (11), 1435–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sun L et al. (2000) Overcoming adeno-associated virus vector size limitation through viral DNA heterodimerization. Nat Med 6 (5), 599–602. [DOI] [PubMed] [Google Scholar]
- 37.Fine EJ et al. (2015) Trans-spliced Cas9 allows cleavage of HBB and CCR5 genes in human cells using compact expression cassettes. Sci Rep 5, 10777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li J et al. (2008) Protein trans-splicing as a means for viral vector-mediated in vivo gene therapy. Hum Gene Ther 19 (9), 958–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Truong DJ et al. (2015) Development of an intein-mediated split-Cas9 system for gene therapy. Nucleic Acids Res 43 (13), 6450–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Villiger L et al. (2018) Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat Med 24 (10), 1519–1525. [DOI] [PubMed] [Google Scholar]
- 41.Levy JM et al. (2020) Cytosine and adenine base editing of the brain, liver, retina, heart and skeletal muscle of mice via adeno-associated viruses. Nat Biomed Eng 4 (1), 97–110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yeh WH et al. (2020) In vivo base editing restores sensory transduction and transiently improves auditory function in a mouse model of recessive deafness. Sci Transl Med 12 (546). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yang L et al. (2020) Amelioration of an Inherited Metabolic Liver Disease through Creation of a De Novo Start Codon by Cytidine Base Editing. Mol Ther 28 (7), 1673–1683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Jiang T et al. (2020) Chemical modifications of adenine base editor mRNA and guide RNA expand its application scope. Nat Commun 11 (1), 1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Song CQ et al. (2020) Adenine base editing in an adult mouse model of tyrosinaemia. Nat Biomed Eng 4 (1), 125–130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lalonde S et al. (2017) Frameshift indels introduced by genome editing can lead to in-frame exon skipping. PLoS One 12 (6), e0178700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mou H et al. (2017) CRISPR/Cas9-mediated genome editing induces exon skipping by alternative splicing or exon deletion. Genome Biology 18 (1). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Tuladhar R et al. (2019) CRISPR-Cas9-based mutagenesis frequently provokes on-target mRNA misregulation. Nat. Commun. 10 (1), 4056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Billon P et al. (2017) CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol Cell 67 (6), 1068–1079 e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kuscu C et al. (2017) CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations. Nat Methods 14 (7), 710–712. [DOI] [PubMed] [Google Scholar]
- 51.Wang X et al. (2020) Efficient gene silencing by adenine base editor-mediated start codon mutation. Mol Ther 28 (2), 431–440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Rossidis AC et al. (2018) In utero CRISPR-mediated therapeutic editing of metabolic genes. Nat Med 24 (10), 1513–1518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chadwick AC et al. (2017) In vivo base editing of PCSK9 (Proprotein Convertase Subtilisin/Kexin Type 9) as a therapeutic alternative to genome editing. Arterioscler Thromb Vasc Biol 37 (9), 1741–1747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Lim CKW et al. (2020) Treatment of a mouse model of ALS by in vivo base editing. Mol Ther 28 (4), 1177–1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Webber BR et al. (2019) Highly efficient multiplex human T cell engineering without double-strand breaks using Cas9 base editors. Nat Commun 10 (1), 5222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zeng J et al. (2020) Therapeutic base editing of human hematopoietic stem cells. Nat Med 26 (4), 535–541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gapinske M et al. (2018) CRISPR-SKIP: programmable gene splicing with single base editors. Genome Biol 19 (1), 107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Yuan J et al. (2018) Genetic modulation of RNA splicing with a CRISPR-guided cytidine deaminase. Mol Cell 72 (2), 380–394 e7. [DOI] [PubMed] [Google Scholar]
- 59.Gaudelli NM et al. (2017) Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551 (7681), 464–471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kim D et al. (2017) Genome-wide target specificities of CRISPR RNA-guided programmable deaminases. Nat Biotechnol 35 (5), 475–480. [DOI] [PubMed] [Google Scholar]
- 61.Liang P et al. (2019) Genome-wide profiling of adenine base editor specificity by EndoV-seq. Nat Commun 10 (1), 67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Rees HA et al. (2017) Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat Commun 8, 15790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zuo E et al. (2019) Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364 (6437), 289–292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Grunewald J et al. (2019) Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors. Nature 569 (7756), 433–437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Doman JL et al. (2020) Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors. Nat Biotechnol 38 (5), 620–628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Gehrke JM et al. (2018) An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat Biotechnol 36 (10), 977–982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hu JH et al. (2018) Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556 (7699), 57–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kim YB et al. (2017) Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat Biotechnol. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Yu Y et al. (2020) Cytosine base editors with minimized unguided DNA and RNA off-target events and high on-target activity. Nat Commun 11 (1), 2052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Grunewald J et al. (2019) CRISPR DNA base editors with reduced RNA off-target and self-editing activities. Nat Biotechnol 37 (9), 1041–1048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Rees HA et al. (2019) Analysis and minimization of cellular RNA editing by DNA adenine base editors. Sci Adv 5 (5), eaax5717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Jiang W et al. (2018) BE-PLUS: a new base editing tool with broadened editing window and enhanced fidelity. Cell Res 28 (8), 855–861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Komor AC et al. (2017) Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Science Advances 3 (8). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wang L et al. (2017) Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor. Cell Res 27 (10), 1289–1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Gaudelli NM et al. (2020) Directed evolution of adenine base editors with increased activity and therapeutic application. Nat Biotechnol 38 (7), 892–900. [DOI] [PubMed] [Google Scholar]
- 76.Richter MF et al. (2020) Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat Biotechnol 38 (7), 883–891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Zhang X et al. (2020) Increasing the efficiency and targeting range of cytidine base editors through fusion of a single-stranded DNA-binding protein domain. Nat Cell Biol. [DOI] [PubMed] [Google Scholar]
- 78.Liu Z et al. (2020) Efficient base editing with high precision in rabbits using YFE-BE4max. Cell Death Dis 11 (1), 36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Hanlon KS et al. (2019) High levels of AAV vector integration into CRISPR-induced DNA breaks. Nat Commun 10 (1), 4439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Nelson CE et al. (2019) Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat Med 25 (3), 427–432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Anzalone AV et al. (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576 (7785), 149–157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Chow RD et al. (2020) A web tool for the design of prime-editing guide RNAs. Nat Biomed Eng. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Kim HK et al. (2020) Predicting the efficiency of prime editing guide RNAs in human cells. Nat Biotechnol. [DOI] [PubMed] [Google Scholar]
- 84.Chakrabarti AM et al. (2019) Target-specific precision of CRISPR-mediated genome editing. Mol Cell 73 (4), 699–713 e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Shen MW et al. (2018) Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563 (7733), 646–651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Arbab M et al. (2020) Determinants of Base Editing Outcomes from Target Library Analysis and Machine Learning. Cell. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Dandage R et al. (2019) beditor: A computational workflow for designing libraries of guide RNAs for CRISPR-mediated base editing. Genetics 212 (2), 377–385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Hwang GH et al. (2018) Web-based design and analysis tools for CRISPR base editing. BMC Bioinformatics 19 (1), 542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Winter J et al. (2019) Targeted exon skipping with AAV-mediated split adenine base editors. Cell Discov 5, 41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Abudayyeh OO et al. (2017) RNA targeting with CRISPR-Cas13. Nature 550 (7675), 280–284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Abudayyeh OO et al. (2016) C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353 (6299), aaf5573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.East-Seletsky A et al. (2016) Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538 (7624), 270–273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Meeske AJ et al. (2019) Cas13-induced cellular dormancy prevents the rise of CRISPR-resistant bacteriophage. Nature 570 (7760), 241–245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Smargon AA et al. (2017) Cas13b Is a Type VI-B CRISPR-Associated RNA-Guided RNase Differentially Regulated by Accessory Proteins Csx27 and Csx28. Mol Cell 65 (4), 618–630 e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Yan WX et al. (2018) Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol Cell 70 (2), 327–339 e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Konermann S et al. (2018) Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173 (3), 665–676 e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Cox DBT et al. (2017) RNA editing with CRISPR-Cas13. Science 358 (6366), 1019–1027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Zhao X et al. (2018) A CRISPR-Cas13a system for efficient and specific therapeutic targeting of mutant KRAS for pancreatic cancer treatment. Cancer Lett 431, 171–181. [DOI] [PubMed] [Google Scholar]
- 99.Zhou H et al. (2020) Glia-to-neuron conversion by CRISPR-CasRx alleviates symptoms of neurological disease in mice. Cell 181 (3), 590–603 e16. [DOI] [PubMed] [Google Scholar]
- 100.Du M et al. (2020) CRISPR artificial splicing factors. Nat Commun 11 (1), 2973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Abudayyeh OO et al. (2019) A cytosine deaminase for programmable single-base RNA editing. Science 365 (6451), 382–386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Sinnamon JR et al. (2017) Site-directed RNA repair of endogenous Mecp2 RNA in neurons. Proc Natl Acad Sci U S A 114 (44), E9395–E9402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Charlesworth CT et al. (2019) Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat Med 25 (2), 249–254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Simhadri VL et al. (2018) Prevalence of pre-existing antibodies to CRISPR-associated nuclease Cas9 in the USA population. Mol Ther Methods Clin Dev 10, 105–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Ferdosi SR et al. (2019) Multifunctional CRISPR-Cas9 with engineered immunosilenced human T cell epitopes. Nat Commun 10 (1), 1842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Varble A and Marraffini LA (2019) Three new Cs for CRISPR: collateral, communicate, cooperate. Trends Genet 35 (6), 446–456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.van Haasteren J et al. (2020) The delivery challenge: fulfilling the promise of therapeutic genome editing. Nat Biotechnol 38 (7), 845–855. [DOI] [PubMed] [Google Scholar]
- 108.Klompe SE et al. (2019) Transposon-encoded CRISPR-Cas systems direct RNA-guided DNA integration. Nature 571 (7764), 219–225. [DOI] [PubMed] [Google Scholar]
- 109.Strecker J et al. (2019) RNA-guided DNA insertion with CRISPR-associated transposases. Science 365 (6448), 48–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Chen X et al. (2017) In trans paired nicking triggers seamless genome editing without double-stranded DNA cutting. Nat Commun 8 (1), 657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Hyodo T et al. (2020) Tandem Paired Nicking Promotes Precise Genome Editing with Scarce Interference by p53. Cell Rep 30 (4), 1195–1207 e7. [DOI] [PubMed] [Google Scholar]
- 112.Nakajima K et al. (2018) Precise and efficient nucleotide substitution near genomic nick via noncanonical homology-directed repair. Genome Res 28 (2), 223–230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Zhang X et al. (2020) Functionalized lipid-like nanoparticles for in vivo mRNA delivery and base editing. Sci Adv 6 (34). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Chen X and Goncalves MA (2016) Engineered Viruses as Genome Editing Devices. Mol Ther 24 (3), 447–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Yin H et al. (2017) Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat Biotechnol 35 (12), 1179–1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Jiang C et al. (2017) A non-viral CRISPR/Cas9 delivery system for therapeutically targeting HBV DNA and pcsk9 in vivo. Cell Res 27 (3), 440–443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Wei T et al. (2020) Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing. Nat Commun 11 (1), 3232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Lee K et al. (2017) Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat Biomed Eng 1, 889–901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Latella MC et al. (2016) In vivo editing of the human mutant rhodopsin gene by electroporation of plasmid-based CRISPR/Cas9 in the mouse retina. Mol Ther Nucleic Acids 5 (11), e389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Wu W et al. (2017) Efficient in vivo gene editing using ribonucleoproteins in skin stem cells of recessive dystrophic epidermolysis bullosa mouse model. Proc Natl Acad Sci U S A 114 (7), 1660–1665. [DOI] [PMC free article] [PubMed] [Google Scholar]