Abstract
Background: Annotation ambiguities and annotation errors are a general challenge in genomics. While a reliable protein function assignment can be obtained by experimental characterization, this is expensive and time-consuming, and the number of such Gold Standard Proteins (GSP) with experimental support remains very low compared to proteins annotated by sequence homology, usually through automated pipelines. Even a GSP may give a misleading assignment when used as a reference: the homolog may be close enough to support isofunctionality, but the substrate of the GSP is absent from the species being annotated. In such cases, the enzymes cannot be isofunctional. Here, we examined a variety of such issues in halophilic archaea (class Halobacteria), with a strong focus on the model haloarchaeon Haloferax volcanii. Results: Annotated proteins of Hfx. volcanii were identified for which public databases tend to assign a function that is probably incorrect. In some cases, an alternative, probably correct, function can be predicted or inferred from the available evidence, but this has not been adopted by public databases because experimental validation is lacking. In other cases, a probably invalid specific function is predicted by homology, and while there is evidence that this assigned function is unlikely, the true function remains elusive. We listed 50 of those cases, each with detailed background information, so that a conclusion about the most likely biological function can be drawn. For reasons of brevity and comprehension, only the key aspects are listed in the main text, with detailed information being provided in a corresponding section of the Supplementary Materials. Conclusions: Compiling, describing and summarizing these open annotation issues and functional predictions will benefit the scientific community in the general effort to improve the evaluation of protein function assignments and more thoroughly detail them. By highlighting the gaps and likely annotation errors currently in the databases, we hope this study will provide a framework for experimentalists to systematically confirm (or disprove) our function predictions or to uncover yet more unexpected functions.
Keywords: haloarchaea, genome annotation, Gold Standard Protein, Haloferax volcanii, annotation error
1. Introduction
Haloferax volcanii is a model organism for halophilic archaea [1,2,3,4,5,6], for which an elaborate set of genetic tools has been developed [7,8,9]. Its genome has been sequenced and carefully annotated [1,10,11]. A plethora of biological aspects have been successfully tackled in this species, with examples including DNA replication [4]; cell division and cell shape [12,13,14,15,16]; metabolism [17,18,19,20,21,22,23,24,25]; protein secretion [26,27,28,29]; motility and biofilms [30,31,32,33,34,35]; mating [36]; signaling [37]; virus defense [38]; proteolysis [39,40,41,42,43,44]; posttranslational modification (N-glycosylation; SAMPylation) [45,46,47,48,49,50]; gene regulation [21,25,51,52,53,54,55]; microproteins [56,57,58] and small noncoding RNAs (sRNAs) [59,60,61,62].
Genome annotations are frequently compromised by annotation errors [11,63,64,65]. Many of these errors are caused by an invalid annotation transfer between presumed homologs, which, once introduced, are further spread by annotation robots. This problem can be partially overcome by using a Gold Standard Protein (GSP)-based annotation strategy [11]. Since the GSP has itself been subjected to an experimental analysis, its annotation cannot be caused by an invalid annotation transfer process. The GSP strategy was already applied to a detailed analysis of the metabolism of halophilic archaea [66]. However, with a decreasing level of sequence identity, the assumption of isofunctionality becomes increasingly uncertain. Although this may be counterbalanced by additional evidence, e.g., gene clustering, experimental confirmation would be the best option for validation of the annotation.
There are additional and much more subtle genome annotation problems. In some cases, GSPs are true homologs, and the annotated function in the database is correct. Nevertheless, the biological context in the query organism makes it unlikely that the homologs are isofunctional, e.g., when the substrate of the GSP is lacking in the query organism. Additionally, paralogs may have distinct but related functions that cannot be assigned by a sequence analysis but may be assigned based on phylogenetic considerations. Here, again, experimental confirmation is the preferred option for validation of the annotation. A lack of experimental confirmation may keep high-level databases like KEGG or the SwissProt section of UniProt from adopting assignments based on well-supported bioinformatic analyses, so that the database entries continue to provide information that is probably incorrect. We refer to annotation problems in these databases solely to underscore that the biological issues raised by us are far from trivial. There is no intention to question the exceedingly high quality of the SwissProt and KEGG databases [67,68] and their tremendous value for the scientific community. We have actively supported them by providing feedback and encourage others to do the same, e.g., with the recently implemented “Add a publication” functionality in the UniProt entries that allows users to connect a protein to a publication that describes its experimental characterization (https://community.uniprot.org/bbsub/bbsubinfo.html).
In this study, we describe a number of annotation issues for haloarchaea, with a strong emphasis on Hfx. volcanii. We denote such cases as “open annotation issues” with the hope of attracting members of the Haloferax community and other groups working with halophilic archaea to apply experimental analyses to elucidate the true function(s) of these proteins. This will increase the number of Gold Standard Proteins that originate from Hfx. volcanii or other haloarchaea, reduce genome annotation ambiguities and perhaps uncover novel metabolic processes.
2. Materials and Methods
2.1. Curation of Genome Annotation and Gold Standard Protein Identification
The Gold Standard Protein-based curation of haloarchaeal genomes has been described previously [11] (see, also, next paragraph). Since then, a systematic comparison to the KEGG data was performed for a subset of the curated genomes [69]. The Hfx. volcanii genome annotation is continuously scrutinized, especially when a closely related genome is annotated [70].
In brief, the core rule of Gold Standard Protein-based genome annotation is to assign a specific function only when a homologous protein has been confirmed experimentally to have this function. Two types of data must be available for that homolog: (a) a reference describing the experimental characterization and (b) an entry in a sequence database, so that the level of sequence similarity can be determined. The decision on whether isofunctionality can be assumed at this level of sequence similarity and, thus, if the annotation can be transferred represents an informed prediction by the annotator based on the available evidence. This decision may be taken only once for a set of closely related orthologs, such as those from halophilic archaea.
2.2. Additional Bioinformatics Tools
The key databases were UniProtKB/SwissProt [68], InterPro [71], KEGG [67] and OrthoDB [72]. The SyntTax server was used for inspecting the conservation of the gene neighborhood [73]. As general tools, the BLAST suite of programs [74,75] was used for sequence comparisons.
3. Results
The open issues are organized below under Section 3.1, the respiratory chain and oxidative decarboxylation; Section 3.2, amino acid metabolism; Section 3.3, heme and cobalamin biosynthesis; Section 3.4, coenzyme F420; Section 3.5, tetrahydrofolate as opposed to methanopterin; Section 3.6, NAD and riboflavin; Section 3.7, lipid metabolism; Section 3.8, genetic information processing and Section 3.9, stand-alone (miscellaneous) cases. We collected this set of open annotation issues during our continuous efforts to keep the Hfx. volcanii genome up-to-date since its initial publication in 2010 [1]. Not covered in this study are enigmatic reactions and pathways (e.g., archaeal signal peptidase II or the haloarchaeal O-glycosylation pathway) for which no support from experimentally characterized homologs (GSP proteins) is available.
3.1. The Respiratory Chain and Oxidative Decarboxylation
In the respiratory chain, the coenzymes that were reduced during catabolism (e.g., glycolysis) are reoxidized, with the energy being saved as an ion gradient. The textbook examples of a respiratory chain are the five mitochondrial complexes [76,77]: complex I (NADH dehydrogenase), complex II (succinate dehydrogenase), complex III (cytochrome bc1 complex), complex IV (cytochrome-c oxidase as a prototype for a terminal oxidase) and complex V (F-type ATP synthase). In mitochondria, a significant part of the NADH that feeds into the respiratory chain originates from oxidative decarboxylation: the conversion of pyruvate to acetyl-CoA by the pyruvate dehydrogenase complex and conversion of α-ketoglutarate to succinyl-CoA by the homologous 2-oxoglutarate dehydrogenase complex. While complexes I and II transfer reducing elements to a lipid-embedded two-electron carrier (ubiquinone), the bc1 complex transfers the electrons to the one-electron carrier cytochrome-c, a heme (and, thus, iron) protein, which then transfers electrons to the terminal oxidase.
Bacteria like Escherichia coli and Paracoccus denitrificans have related complexes and enzymes: NADH dehydrogenase (encoded by the nuo operon), succinate dehydrogenase (encoded by sdhABCD) and the related fumarate reductase (encoded by frdABCD) [78], several terminal oxidases (e.g., products of cyoABCDE and cydABC) and an F-type ATP synthase (encoded by atp genes). E. coli lacks a bc1 complex, which, however, occurs in Paracoccus denitrificans [79]. E. coli contains the canonical complexes of oxidative decarboxylation (the pyruvate dehydrogenase complex, encoded by aceEF+lpdA, and the 2-oxoglutarate dehydrogenase complex, encoded by sucAB+lpdA).
The respiratory chain of Hfx. volcanii and other haloarchaea deviates considerably from those of mitochondria and bacteria such as Paracoccus and E. coli (reviewed by [80]), and a number of questions remain unresolved. We focus on the equivalents of complexes I, III and IV, because these have unresolved issues. We also cover some aspects relevant for the NADH levels (oxidative decarboxylation enzymes and type II NADH dehydrogenase). We do not cover complexes that have already been studied in haloarchaea: complex II (succinate dehydrogenase) [81,82,83] and complex V (ATP synthase) [84,85].
(a) In haloarchaea, oxidative decarboxylation is not linked to the reduction of NAD to NADH but to the reduction of a ferredoxin (encoded by fdx, e.g., OE_4217R, HVO_2995), which has a redox potential similar to that of the NAD/NADH pair [86]. The enzymes for oxidative decarboxylation are pyruvate–ferredoxin oxidoreductase (porAB, e.g., OE_2623R/2622R and HVO_1305/1304) and 2-oxoglutarate–ferredoxin oxidoreductase (korAB, e.g., OE_1711R/1710R and HVO_0888/0887), and these have been characterized from Halobacterium salinarum [87,88,89].
(b) It is yet unresolved how ferredoxin Fdx is reoxidized, but this might be achieved by the Nuo complex. This ferredoxin may well be involved in additional metabolic processes. In Hfx. volcanii, ferredoxin Fdx (HVO_2995) plays an essential role in nitrate assimilation [90]. However, in Hbt. salinarum, this metabolic process for Fdx reoxidation does not exist.
(c) The nuo cluster of haloarchaea resembles that of E. coli, a type I NADH dehydrogenase, with the genes and gene order highly conserved and just a few domain fissions and fusions. However, haloarchaea lack NuoEFG [91], which is a subcomplex that mediates interaction with NADH [92,93]. Thus, the haloarchaeal nuo complex is unlikely to function as NADH dehydrogenase, despite its annotation as such in KEGG (as of April 2021).
(d) Other catabolic enzymes generate NADH, which must also be reoxidized. Based on inhibitor studies, NADH is not reoxidized by a type I but, rather, by a type II NADH dehydrogenase in Hbt. salinarum [82]. A tentative gene assignment has been made for Natronomonas pharaonis [66]. However, for reasons detailed in Supplementary Text S1 Section S1, this assignment is highly questionable, so this issue calls for an experimental analysis.
(e) About one-third of the haloarchaea, especially the Natrialbales, do not code for a complex III equivalent (the cytochrome bc1 complex encoded by petABC), according to OrthoDB analysis. The bc1 complex is required to transfer electrons from the lipid-embedded two-electron carrier (menaquinone in haloarchaea) to the one-electron carrier associated with terminal oxidases (probably halocyanin). How electrons flow in the absence of a complex III equivalent is currently unresolved.
The haloarchaeal petABC genes resemble those of the chloroplast b6-f complex rather than those of the mitochondrial bc1 complex (see Supplementary Text S1 Section S1 for more details).
(f) A bc cytochrome was purified from Nmn. pharaonis, but with an atypical 1:1 ratio between the b-type and c-type hemes [81]. The complex is heterodimeric, with subunits of 18 kDa and 14 kDa. The 18-kDa subunit carries the covalently attached heme group [81]. An attempt was made to identify the genes coding for these subunits [94] (for details, see Supplementary Text S1 Section S1). Two approaches were used to obtain protein sequence data, one being the N-terminal protein sequencing of the two subunits extracted from a SDS-polyacrylamide gel. In the other attempt, peptides from the purified complex were separated by HPLC, and a peptide which absorbed at 280 nm (protein), as well as 400 nm (heme), was isolated. Absorption at 400 nm clearly indicates that the isolated peptide contains a covalently attached heme group. The sequences from the two approaches overlapped and resulted in a contiguous sequence of 41 aa, with only the penultimate position remaining undefined [94]. Based on this information, a PCR probe was generated (designated “cyt-C Sonde”) that allowed the gene to be identified and sequenced, including its genomic neighborhood. It turned out that the genes coding for the four subunits of succinate dehydrogenase (sdhCDBA) were isolated. The obtained protein sequence corresponds to the N-terminal region of sdhD (with the initiator methionine cleaved off) and only two sequence discrepancies, in addition to the unresolved penultimate residue.
In the PhD thesis [94], this unambiguous result was rated to be a failure (and the data were never formally published). The reason is that SdhD is free of cysteine residues, while standard textbooks state that a pair of cysteines is required for covalent heme attachment [95]. The lack of the required cysteine pair was taken to indicate that the results were incorrect and that the identified genes did not encode the cytochrome bc that the study was seeking [94]. In contrast, we speculate that the results were completely correct, despite being in conflict with the cysteine pair paradigm. In our opinion, a paradigm shift is required. The obtained results call for a yet-unanticipated novel mode of covalent heme attachment, exemplified by the 18-kDa subunit of Natronomonas succinate dehydrogenase subunit SdhD. It should be noted that the 41-aa protein sequence, which was obtained, turned out to contain three histidine residues upon translation of the gene, but none of these were detected upon Edman degradation.
In Halobacterium, a small c-type cytochrome was purified (cytochrome c552, 14.1 kDa) [96]. Heme staining after SDS-PAGE indicated a covalent heme attachment, but no sequence or composition data were reported, so that it was not possible to identify the protein based on the available information. We speculate that the Halobacterium cytochrome c552 also represents SdhD (as detailed in Supplementary Text S1 Section S1). In that case, the proposed novel type of covalent heme attachment would not be restricted to Nmn. pharaonis but might be a general property of haloarchaea. This would also solve the “Halobacterium paradox” [95].
(g) The haloarchaeal one-electron carrier is the copper protein halocyanin rather than the iron-containing heme protein cytochrome-c. A halocyanin from Nmn. pharaonis (NP_3954A) was characterized, including its redox potential [97,98,99]. A gene fusion supports the close connection of a halocyanin with a subunit of a terminal oxidase. For further details, see Supplementary Text S1 Section S1.
(h) Terminal oxidases are highly diverse in haloarchaea, and we restricted our analysis to three species (Nmn. pharaonis, Hfx. volcanii and Hbt. salinarum), because in each of these, at least one terminal oxidase has been experimentally studied (Table 1). The details are described in Supplementary Text S1 Section S1 with subunits of all analyzed terminal oxidases listed in Supplementary Table S1.
Table 1.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.1).
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 1a | HVO_1305 HVO_1304 |
porAB | yes | 67% 80% |
OE2623R OE2622R |
B0R4 × 6 B0R4 × 5 |
[87] [88] [89] |
1555599 6266826 6266827 |
|
| 1a | HVO_0888 HVO_0887 |
korAB | yes | 77% 77% |
OE1711R OE1710R |
B0R3G0 B0R3F9 |
[88] [89] |
6266826 6266827 |
|
| 1a/1b | HVO_2995 | fdx | yes | 88% | OE4217R | B0R7I9 | [100] [101] [86] |
964365 188650 201489 |
role in oxidative decarboxylation |
| 1a/1b | HVO_2995 (cont.) |
self | D4GY89 | [90] | 22103537 | role in nitrate assimilation | |||
| 1c | HVO_0979 (complex) |
nuoB | possibly | 50% | tlr0705 | Q8DKZ4 | [102] [103] [104] |
15910282 30573545 32001694 |
reoxidizes ferredoxin |
| 1c | HVO_0979 (cont.) |
no | 48% | b2287 | P0AFC7 | [92] [93] |
7607227 9485311 |
reoxidizes NADH in E.coli | |
| 1d | NP_3508A |
ndh1 | special | 26% (N-term 140 aa) | - | Q7ZAG8 | function of Q7ZAG8 was reassigned (from ndh1 to sqr) after annotation transfer | ||
| 1d | NP_3508A (cont.) |
possibly | 30% | BpOF4_04810 | A7LKG4 | [105] | 18359284 | type II NADH dehydrogenase | |
| 1e | HVO_2620 HVO_0842 HVO_0841 |
petABD | yes | 39% | SYNPCC7002_ A0842 |
P28056 | [106] | 11245788 | HVO_0842 (petB) related to cytochrome b6 |
| 1f | HVO_2810 | sdhD | yes | 66% | NP_4268A | Q3INS7 | [81] [94] |
9109654 PhD_Mattar |
|
| 1g | HVO_0943 | cbaD | yes | 57% | NP_2966A | A0A1U7EWW4 | [107] | 9428682 | |
| HVO_0943 (cont.) |
- | 63% | OE_4073R (C-term) |
B0R7A9 | - | halocyanin/cbaD fusion protein, uncharacterized | |||
| 1g | HVO_2150 |
hcpG
|
- | 44% | OE_4073R (N-term) |
B0R7A9 | - | halocyanin/cbaD fusion protein, uncharacterized | |
| 1h | HVO_0945 (complex) |
cbaA | yes | 64% | NP_2966A | A0A1U7EWW4 | [107] | 9428682 | |
| 1h | HVO_0907 (complex) |
coxA1 | self | [108] | 11790755 | ||||
| 1h | HVO_0907 (cont.) |
yes | 70% | VNG_0657G (OE_1979R) | P33588 | [109] [110] |
2542239 1659810 |
||
| 1h | HVO_1645 (complex) |
coxAC2 | yes | 43% | APE_0793.1 | Q9YdX6 | [111] | 12471503 | |
| 1h | HVO_0462 HVO_0461 |
cydAB | yes | 32% 24% |
- - |
Q09049 Q05780 |
[112] | 1655703 | |
| 1h | HVO_0462 HVO_0461 (cont.) |
yes | 30% 27% |
b0733 b0734 |
P0ABJ9 P0ABK2 |
[113] | 6307994 | ||
| 1h | NP_4296A NP_4294A |
coxA3
coxB3 |
yes | 28% 33% |
TTHA1135 TTHA1134 |
Q5SJ79 Q5SJ80 |
[114] [115] |
2842747 7657607 |
|
| 1i | HVO_2958 HVO_2959 |
oadhAB1 | self | D4GY15 D4GY17 |
[116] | 19910413 | Ile indirectly assigned as substrate | ||
| 1i | HVO_2958 HVO_2959 (cont.) |
self | [117] [118] [119] |
10832633 17571210 17906130 |
no substrate was identified; pyruvate and alphaKG excluded | ||||
| 1i | HVO_2595 HVO_2596 |
oadhAB2 | self | [120] [119] [116] |
12003954 17906130 19910413 |
no substrate was identified; pyruvate and alphaKG excluded | |||
| 1i | HVO_0669 HVO_0668 |
oadhAB3 | self | [119] [116] |
17906130 19910413 |
no substrate was identified; pyruvate and alphaKG excluded | |||
| 1i | HVO_2209 | oadhA4 | self | not yet analyzed experimentally | |||||
| 1i | HVO_2958 HVO_2959 (cont.) |
yes/no | 38% 52% |
TA1438 TA1437 |
Q9HIA3 Q9HIA4 |
[121] | 17894823 | substrates are Ile, Leu, Val | |
| 1i | HVO_2595 HVO_2596 (cont.) |
no | 41% 41% |
- - |
Q57102 Q57041 |
[122] | 1898934 | substrate is acetoin | |
| 1i | HVO_2595 HVO_2596 (cont.) |
unknown | 40% 43% |
BSU08060 BSU08070 |
O31404 O34591 |
[123] | 10368162 | substrate is acetoin | |
| 1i | HVO_0669 HVO_0668 (cont.) |
unknown | 54% 47% |
BSU08060 BSU08070 |
O31404 O34591 |
[123] | 10368162 | substrate is acetoin | |
| 1i | HVO_0669 HVO_0668 (cont.) |
unknown | 49% 43% |
- - |
Q57102 Q57041 |
[122] | 1898934 | substrate is acetoin | |
| 1i | HVO_2209 (cont.) |
unknown | 38% | TA1438 | Q9HIA3 | [121] | 17894823 | substrates are Ile, Leu, Val | |
The column Section refers to the table listing the protein and to the section in the Results and in Supplementary Text S1. As an example, 2c covers topic (c) from the decimal-numbered Results Section 3.2. Amino Acid Biosynthesis. In Supplementary Text S1, this is covered under Section S2 subsection S2.c. The corresponding proteins are listed in Table 2. For a few proteins, two sections are indicated (e.g., 1a/1b). The column Code refers to a haloarchaeal protein by its locus tag, which is mainly from Haloferax volcanii (HVO) but, also, from Halobacterium salinarum (OE), Natronomonas pharaonis (NP) and Halohasta litchfieldiae (halTADL). When the reconstruction of a complete pathway is presented, the unassigned genes are indicated as a “pathway gap”. In one case, we indicate the absence of a haloarchaeal ortholog by a dash. In the case of a complex, we either list more than one code or we list only one subunit together with the term (complex). All subunits of these complexes are listed groupwise in Table S1. A protein may be shown in more than one row. From the 2nd row onwards, this is indicated by the term (cont.). The column Gene lists the assigned gene or a dash if no gene has been assigned. The assigned gene is only indicated in the first row of a protein. A set of four columns is used to relate a query protein to an experimentally characterized homolog, a GSP (Gold Standard Protein) (isofunc, %seq_id, Locus tag, UniProt). The column isofunc indicates if the query protein and its Gold Standard Protein homolog are isofunctional. The meanings of the terms used in this column in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6, Table 7, Table 8 and Table 9 (yes, no, yes/no, probably, possibly, unclear, unknown, prediction, special and “-“) are described at the end of this legend. The column %seq_id indicates the protein sequence identity between the query protein and the homologous GSP. The column Locus tag contains the locus tag, if assigned. The column UniProt contains the UniProt accession of the GSP. GSPs are experimentally characterized as described in a publication. The column Reference links to the reference list of the manuscript. The column PMID lists the PubMed ID of the publication, if available. Otherwise, this is indicated as “not in PubMed”. Additionally, one PhD thesis is indicated (PhD_Mattar). The column Comment provides various types of additional information. The terms used in the column isofunc in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6, Table 7, Table 8 and Table 9 have the following meanings: The term “yes” indicates that we consider the two proteins as isofunctional and annotate the query protein accordingly. The term “no” is used when we conclude that the proteins differ in function. Additional terms are used for more difficult cases. The term “yes/no” is used for GSPs that are multifunctional, and we assign only a subset of these functions to the query protein. The term “probably” is used when we consider it likely that the proteins are isofunctional and annotated the query protein accordingly (with the term probable added to the protein name). The term “possibly” is used when we see a good chance that the proteins are isofunctional but consider it too speculative to annotate the protein accordingly. The term “unclear” is used when we consider it likely that the same overall reaction is catalyzed but when reaction details, e.g., the energy-providing compound, are unresolved. The term “unknown” is used when it is not possible to predict the substrate of the query protein. The term “prediction” is used if a function assignment is based on bioinformatic analyses but not yet on an experimentally characterized homologous protein. The term “special” is used when multiple arguments have to be considered, with the full details provided in the corresponding section of Supplementary Text S1. Finally, a hyphen (“-“) is used when isofunctionality does not apply, e.g., when a homologous Gold Standard Protein could not be identified.
(i) NAD-dependent oxidative decarboxylation is a canonical reaction to convert pyruvate into acetyl-CoA and α-ketoglutarate into succinyl-CoA. In haloarchaea, the conversion of pyruvate to acetyl-CoA and α-ketoglutarate to succinyl-CoA is dependent on ferredoxin, not on NAD (see above). Nevertheless, most haloarchaeal genomes also code for homologs of enzymes catalyzing NAD-dependent oxidative decarboxylation, such as the E. coli pyruvate dehydrogenase complex. In most cases, the substrates could not be identified, an exception being a paralog involved in isoleucine catabolism [116]. In several cases, the enzymes were found not to show catalytic activity with pyruvate or α-ketoglutarate (see Supplementary Text S1 Section S1 for details). Additionally, a conditional lethal porAB mutant was unable to grow on glucose or pyruvate, thus excluding that alternative enzymes for the conversion of pyruvate to acetyl-CoA exist in Hfx. volcanii [22]. Nonetheless, despite experimental results to the contrary, pyruvate has been assigned as a substrate for some of the homologs of the pyruvate dehydrogenase complex in KEGG (as of April 2021).
3.2. Amino Acid Metabolism
While most amino acid biosynthesis and degradation pathways can be reliably reconstructed, a few open issues remain, which are discussed below.
(a) The first and last steps of arginine biosynthesis deal with blocking and unblocking of the α-amino group of the substrate (glutamate) and a product intermediate (ornithine). As detailed in Supplementary Text S1 Section S2, it is highly likely that glutamate is attached to the γ-carboxyl group of a carrier protein, and ornithine is released from that carrier protein. This is based on characterized proteins from Thermus thermophilus [124], Thermococcus kodakarensis [125] and Sulfolobus acidocaldarius [126]. The assignment is strongly supported by clustering of the arginine biosynthesis genes. Some of the homologs are bifunctional, being involved in arginine biosynthesis but, also, in lysine biosynthesis via the prokaryotic variant of the α-aminoadipate pathway. This ambiguity is not assumed to occur in haloarchaea, which use the diaminopimelate pathway for lysine biosynthesis [127] (see Supplementary Text S1 Section S2 for further discussion of this issue).
Expanding the above, we provided full details underlying our reconstruction of arginine and lysine biosynthesis in Hfx. volcanii in Table 2.
Table 2.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.2). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | isofunc | %seq_id | Locus tag | UniProt | Reference | PMID | Comment |
| 2a | HVO_0047 | argW | no | 54% | TT_C1544 | Q72HE5 | [128] | 25392000 | for Arg, not for Lys biosynthesis |
| 2a | HVO_0047 (cont.) |
yes/no | 39% | Saci_0753 | Q4JAQ0 | only for Arg, not for Lys biosynthesis | |||
| 2a | HVO_0047 (cont.) |
yes/no | 61% | TK0279 | Q5JFV9 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0046 | argX | no | 44% | TT_C1543 | Q72HE6 | [124] | 19620981 | for Arg, not for Lys biosynthesis |
| 2a | HVO_0046 (cont.) |
yes | 30% | Saci_1621 | Q4J8E7 | only for Arg, not for Lys biosynthesis | |||
| 2a | HVO_0046 (cont.) |
yes/no | 37% | TK0278 | Q5JFW0 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0044 | argB | no | 41% | TT_C1541 | O50147 | [124] [128] |
19620981 25392000 |
for Arg, not for Lys biosynthesis |
| 2a | HVO_0044 (cont.) |
yes/no | 33% | Saci_0751 | Q4JAQ2 | [126] | 23434852 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0044 (cont.) |
yes/no | 32% | TK0276 | Q5JFW2 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0045 | argC | no | 48% | TT_C1542 | O50146 | [124] [129] |
19620981 26966182 |
for Arg, not for Lys biosynthesis |
| 2a | HVO_0045 (cont.) |
yes/no | 42% | Saci_0750 | Q4JAQ3 | [126] | 23434852 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0045 (cont.) |
yes/no | 46% | TK0277 | Q5JFW1 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0043 | argD | no | 45% | TT_C1393 | Q93R93 | [130] | 11489859 | for Arg, not for Lys biosynthesis |
| 2a | HVO_0043 (cont.) |
yes/no | 40% | Saci_0755 | Q4JAP8 | [126] | 23434852 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0043 (cont.) |
yes/no | 42% | TK0275 | Q5JFW3 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0042 | argE | no | 36% | TT_C1396 | Q8VUS5 | [124] [131] |
19620981 28720495 |
for Arg, not for Lys biosynthesis |
| 2a | HVO_0042 (cont.) |
yes/no | 29% | Saci_0756 | Q4JAP7 | [126] | 23434852 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0042 (cont.) |
yes/no | 37% | TK0274 | Q5JFW4 | [125] | 27566549 | only for Arg, not for Lys biosynthesis | |
| 2a | HVO_0041 | argF | yes | 50% | P18186 | BSU11250 | [132] | 4216455 | |
| 2a | HVO_0041 (cont.) |
yes | 47% | OE_5205R | B0R9X3 | [133] | 7868583 | ||
| 2a | HVO_0049 | argG | yes | 35% | - | P00966 | [134] | 8792870 | human |
| 2a | HVO_0049 (cont.) |
yes | 23% | b3172 | P0A6E4 | [135] | 10666579 | E. coli | |
| 2a | HVO_0048 | argH | yes | 38% | MMP0013 | O74026 | [136] | 10220900 | |
| 2a | HVO_0008 | lysC | yes | 32% | BSU28470 | P08495 | [137] | 15033471 | |
| 2a | HVO_2487 | asd | yes | 51% | MJ0205 | Q57658 | [138] | 16225889 | |
| 2a/9e | HVO_1101 | dapA | yes | 45% | PA1010 | Q9I4W3 | [139] | 21396954 | |
| 2a | HVO_1100 | dapB | yes | 33% | b0031 | P04036 | [140] | 7893644 | |
| 2a | HVO_1099 | dapD | yes | 32% | b0166 | P0A9D8 | [141] | 6365916 | |
| 2a | HVO_1096 | dapE | yes | 29% | b2472 | P0AED7 | [142] | 3276674 | function supported by gene clustering |
| 2a | HVO_1097 | dapF | yes | 35% | b3809 | P0A6K1 | [143] | 6378903 | |
| 2a | HVO_1098 | lysA | yes | 38% | b2838 | P00861 | [144] | 14343156 | |
| 2a | HVO_A0634 | - | unknown | 25% | b2472 | P0AED7 | [142] | 3276674 | function assigned to HVO_1096 in dap cluster |
| 2b | HVO_0790 | fba2 | special | 67% | OE_1472F | B0R334 | [145] | 25216252 | EC 2.2.1.10 activity of OE_1472F not yet confirmed in vitro |
| 2b | HVO_0790 (cont.) |
special | 45% | MJ0400 | Q57843 | [146] | 15182204 | substrate uncertain | |
| 2b | HVO_0792 | aroB | yes | 69% | OE_1475F | B0R336 | [145] | 25216252 | OE_1475F only partially characterized |
| 2b | HVO_0792 (cont.) |
yes | 44% | MJ1249 | Q58646 | [146] | 15182204 | ||
| 2b | HVO_0602 | aroD1 | yes | 44% | OE_1477R | B0R338 | [145] | 25216252 | |
| 2b | HVO_0602 (cont.) |
yes | 31% | MMP1394 | Q6LXF7 | [147] | 15262931 | ||
| 2c | HVO_0009 | tnaA | yes | 41% | b3708 | P0A853 | [148] [149] |
2659590 14284727 |
|
| 2d | HVO_A0559 | hutH | yes | 42% | BSU39350 | P10944 | [150] [151] |
2454913 14066617 |
|
| 2d | HVO_A0562 | hutU | yes | 62% | BSU39360 | P25503 | [152] | 4990470 | |
| 2d | HVO_A0560 | hutI | yes | 42% | BSU39370 | P42084 | [153] | 16990261 | |
| 2d | HVO_A0561 | hutG | yes | 33% | BSU39380 | P42068 | [152] | 4990470 | |
| 2e | HVO_0431 | - | - | no GSP available | |||||
| 2e | HVO_0644 | leuA1 | yes/no | 47% | MJ1392 | Q58787 | [154] | 9864346 | HVO_0644 monofunc (CimA) or bifunc (CimA+LeuA); MJ1392 CimA |
| 2e | HVO_0644 (cont.) |
unclear | 44% | MJ1195 | Q58595 | [155] | 9665716 | HVO_0644 monofunc (CimA) or bifunc (CimA+LeuA); MJ1195 LeuA |
|
| 2e/2f | HVO_1510 | leuA2 | yes | 47% | MJ1195 | Q58595 | [155] | 9665716 | HVO_1510 LeuA; MJ1195 LeuA |
| 2e/2f | HVO_1510 (cont.) |
no | 41% | MJ1392 | Q58787 | [154] | 9864346 | HVO_1510 LeuA MJ1392 CimA |
|
| 2e | HVO_A0489 | - | no | 31% | MJ1392 | Q58787 | [154] | 9864346 | HVO_A0489 general function only; MJ1392 CimA |
| 2e | HVO_A0489 (cont.) |
no | 30% | MJ1195 | Q58595 | [155] | 9665716 | HVO_A0489 general function only; MJ1195 LeuA |
|
| 2e | HVO_1153 | - | - | function unassigned; no GSP |
|||||
(b) Archaea use a different precursor for aromatic amino acid biosynthesis than the classical pathway. This has been resolved for Methanocaldococcus jannaschii and for Methanococcus maripaludis [146,156]. However, the initial steps may differ from those reported for Methanocaldococcus in that fructose 1,6-bisphosphate, rather than 6-deoxy-5-ketofructose, might be a substrate [145]. Up to now, a clean deletion of the corresponding enzymes and confirmation with in vitro assays has not yet been achieved (for details, see Supplementary Text S1 Section S2).
(c) The gene for tryptophanase (tpa) is stringently regulated in Haloferax, which is the basis for using its promoter in the toolbox for regulated gene expression [157]. The shutdown of this gene avoids tryptophan degradation when supplies are scarce. Tryptophanase cleaves tryptophan into indole, pyruvate and ammonia. The fate of indole is, however, yet unresolved.
(d) A probable histidine utilization cluster exists, based on the characterized homologs from Bacillus subtilis, but has not yet been experimentally verified.
(e) Among the 16 auxotrophic mutants observed in a Hfx. volcanii transposon insertion library [9], some could grow only in the presence of one (or several) supplied amino acids. In many cases, the affected genes were known to be involved in the corresponding pathway, but the others may lead to novel function assignments. One affected gene resulted in histidine auxotrophy, and the product of this gene (HVO_0431) is an interesting candidate. The InterPro domain assignment (HAD family hydrolase) fits into the only remaining pathway gap in histidine biosynthesis (histidinol-phosphatase). In this context, it should be noted that the enzyme that catalyzes the preceding reaction (encoded by hisC) is part of a highly conserved three-gene operon involved in polar lipid biosynthesis (see below). For details, see Supplementary Text S1 Section S2. One affected gene resulted in isoleucine auxotrophy. The product of this gene (HVO_0644) is currently annotated to catalyze two reactions, one being an early step in isoleucine biosynthesis (EC 2.3.1.182) and the other being the first step after leucine biosynthesis branches off from valine biosynthesis (EC 2.3.3.13) (see below, (f)) (for details, see Supplementary Text S1 Section S2).
(f) Hfx. volcanii codes for two paralogs with an attributed function as 2-isopropylmalate synthase (EC 2.3.3.13). This is the first reaction specific to leucine biosynthesis when the pathway branches off valine biosynthesis. One paralog, HVO_0644, is annotated as bifunctional, also catalyzing a chemically similar reaction that is an early step in isoleucine biosynthesis (EC 2.3.1.182). When the gene encoding HVO_0644 is disrupted by transposon integration, cells cannot grow in the absence of isoleucine. It is unclear if the protein is really bifunctional and is really involved in leucine biosynthesis, catalyzing the reaction of EC 2.3.3.13. The other paralog, HVO_1510, belongs to an ortholog set with major problems concerning the start codon assignment. The ortholog set from the 16 genomes listed in Supplementary Table S2 was analyzed. When only canonical start codons are considered (ATG, GTG and TTG), the orthologs from Haloferax mediterranei, Nmn. pharaonis, Natronomonas moolapensis and Halohasta litchfieldiae either lack a long highly conserved N-terminal region or they are disrupted (pseudogenes), being devoid of a potential start codon. The gene from Hfx. volcanii has a start codon (GTG) that is consistent with that of Haloferax gibbonsii strain LR2-5 (but a GTA in Hfx. gibbonsii strain ARA6). In this region, the gene from Hfx. mediterranei is closely related but has in-frame stop codons. HVO_1510 is considerably longer than the orthologs from Haloquadratum walsbyi, Haloarcula hispanica and Natrialba magadii. The first alternative start codon for HVO_1510 codes for Met-93. This protein was proteomically identified in three ArcPP datasets [2], and peptides upstream of Met-93 were identified. This gene might be translated from an atypical start codon, either an in-frame CTG or an out-of-frame ATG, which would require ribosomal slippage (for details, see Supplementary Text S1 Section S2 and Supplementary Figure S1). It is tempting to speculate that translation occurs only when leucine is not available.
3.3. Coenzymes I: Cobalamin and Heme
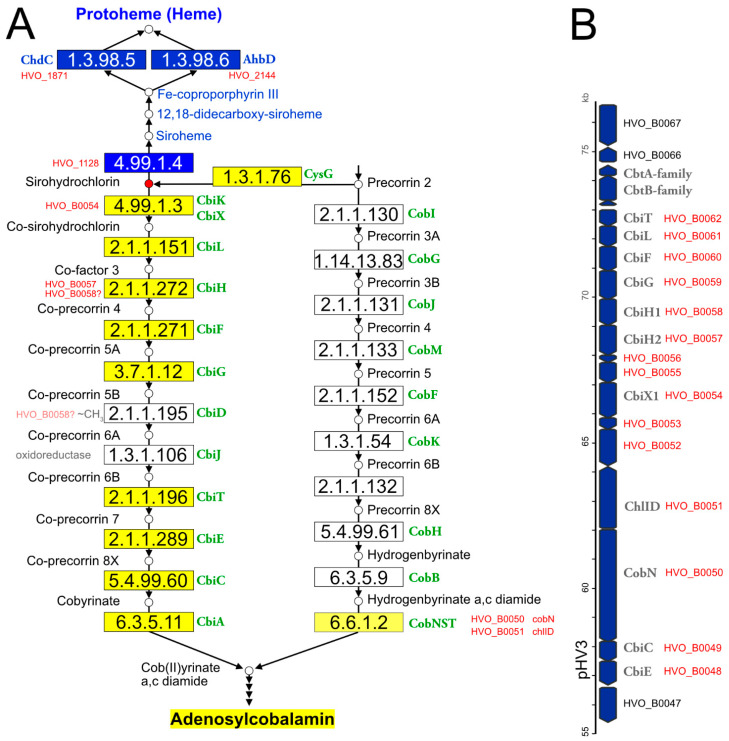
The classical heme biosynthesis pathway branches off cobalamin biosynthesis at the level of uroporphyrinogen III. A second pathway exists in bacteria (CPD pathway). Haloarchaea use the alternative heme biosynthesis pathway [158], which has an additional common step with cobalamin biosynthesis, the conversion of uroporphyrinogen III to precorrin-2. For heme biosynthesis, precorrin-2 is converted into siroheme. This pathway was reconstructed [159], except for the iron insertion step. For de novo cobalamin biosynthesis, haloarchaea use the cobalt-early pathway. A key reaction in this pathway variant, catalyzed by CbiG, is cobalt-dependent. Thus, cobalt must be inserted early and is present in all intermediates [160]. Several aspects of heme and cobalamin biosynthesis in haloarchaea have yet to be resolved. This is illustrated in Figure 1.
Figure 1.
Illustration of the haloarchaeal cobalamin and heme biosynthesis pathways and of the major cobalamin biosynthesis gene cluster. (A) Biosynthesis pathways. This illustration is based on the corresponding KEGG map 00860. Small circles represent pathway intermediates and have their names assigned. Pathway intermediates upstream of precorrin-2 are not displayed. The circle for sirohydrochlorin is highlighted in red, as this is the branchpoint for heme and cobalamin biosynthesis in haloarchaea. Enzymatic reactions are shown by arrows, the EC numbers being provided in rectangular boxes. Rectangles are colored when the enzyme has been reconstructed for haloarchaea (blue: heme biosynthesis; dark yellow: de novo cobalamin biosynthesis; light yellow: late cobaltochelatase, which may be a salvage reaction). Gene names in green are adopted from KEGG and represent those from bacterial model pathways. Consecutive arrowheads indicate reaction series that are not shown in detail for space reasons. Additionally, some enzymes of the heme biosynthesis pathway are omitted for space reasons. For enzymatic reactions that are considered to be open issues, Hfx. volcanii locus tags are provided. For two pathway gaps (white boxes in the cobalt-early pathway), the type of reaction is indicated (oxidoreductase and ~CH3, indicating a methylation reaction). The question mark after HVO_B0058 indicates that this protein, currently co-attributed to EC 2.1.1.272, is a candidate for the yet-unassigned EC 2.1.1.195 reaction. We note that haloarchaea might use a deviating biosynthesis pathway, e.g., by swapping the methylation and oxidoreductase reactions (not illustrated). (B) The major cobalamin cluster, encoded on megaplasmid pHV3. Arrows are used to indicate the coding strand and are roughly drawn to scale. If assigned, the gene name is provided in addition to the Hfx. volcanii locus tag. Locus tags in red indicate genes that are part of the cobalamin cluster.
(a) Hfx. volcanii contains two annotated cbiX genes. For the reasons detailed in Supplementary Text S1 Section S3, we predict that one is a cobaltochelatase, involved in cobalamin biosynthesis, while the other is a ferrochelatase, responsible for the conversion of precorrin-2 to siroheme in the alternative heme biosynthesis pathway.
(b) De novo cobalamin biosynthesis has been extensively reconstructed upon curation of the genome annotation [11]. All enzymes of the pathway and their associated GSPs are listed in Table 3. Only two pathway gaps remained, and because these are consecutive, it may be possible that the haloarchaeal pathway is noncanonical and proceeds via a novel biosynthetic intermediate. There are only four genes with yet-unassigned functions in the Hfx. volcanii cobalamin gene cluster, and their synteny is well-conserved in the majority of haloarchaeal genomes. Thus, these genes are obvious candidates for filling the pathway gaps (for details, see Supplementary Text S1 Section S3).
Table 3.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.3). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 3a | HVO_B0054 | cbiX1 | yes | 30% | - | O87690 | [163] | 12408752 | cobaltochelatase |
| 3a | HVO_B0054 (cont.) |
yes | 27% | MTH_1397 | O27448 | [164] | 12686546 | cobaltochelatase | |
| 3a | HVO_1128 | cbiX2 | no | 29% | AF0721 | O29537 | [165] | 16835730 | cobaltochelatase |
| 3a | HVO_1128 (cont.) |
no | 28% | MTH_1397 | O27448 | [164] | 12686546 | cobaltochelatase | |
| 3a | HVO_1128 (cont.) |
no | 29% | AF0721 | O29537 | [165] | 16835730 | cobaltochelatase | |
| 3a | NP_0734A | cbiX3 | - | function unassigned; no GSP; distantly related to paralogs |
|||||
| 3a | HVO_2312 | sirC | yes/no | 31% | Mbar_A1461 | Q46CH4 | [166] | 21197080 | precorrin-2 DH; no analysis for Fe-chelatase |
| 3a | HVO_2312 (cont.) |
yes/no | 29% | STM3477 | P25924 | [167] [168] |
14595395 32054833 |
matches to the N-term domain which is bifunctional as precorrin-2 DH and Fe-chelatase | |
| 3a | HVO_2312 (cont.) |
yes/no | 29% | - | P61818 | [163] [169] |
12408752 18588505 |
precorrin-2 DH; devoid of Fe-chelatase activity | |
| 3b | HVO_B0061 | cbiL | no | 32% | STM2024 | Q05593 | [170] | 1451790 | equivalent reaction on cobalt-free substrate |
| 3b | HVO_B0057 | cbiH2 | yes | 45% | - | O87689 | [160] | 23922391 | corresponds to N-term of O87689 which has a C-term extension |
| 3b | HVO_B0057 (cont.) |
no | 40% | STM2027 | Q05590 | [171] [172] |
9331403 16198574 |
equivalent reaction on cobalt-free substrate | |
| 3b | HVO_B0058 | cbiH1 | special | 32% | - | O87689 | [160] | 23922391 | corresponds to N-term of O87689 which has a C-term extension; more distant to O87689 than CbiH2 |
| 3b | HVO_B0058 (cont.) |
no | 30% | STM2027 | Q05590 | [171] [172] |
9331403 16198574 |
equivalent reaction on cobalt-free substrate | |
| 3b | HVO_B0060 | cbiF | no | 40% | STM2029 | P0A2G9 | [170] [173] |
1451790 16866557 |
equivalent reaction on cobalt-free substrate |
| 3b | HVO_B0060 (cont.) |
yes | 38% | - | O87686 | [160] | 23922391 | ||
| 3b | HVO_B0059 | cbiG | yes | 24% | - | O87687 | [160] | 23922391 | |
| 3b | pathway gap | EC 2.1.1.195 | |||||||
| 3b | pathway gap | EC 1.3.1.106 | |||||||
| 3b | HVO_B0062 | cbiT | yes | 36% | - | O87694 | [160] | 23922391 | corresponds to the C-term of bifunctional O87694 |
| 3b | HVO_B0048 | cbiE | yes | 28% | - | O87694 | [160] | 23922391 | corresponds to the N-term of bifunctional O87694 |
| 3b | HVO_B0049 | cbiC | yes | 33% | - | O87692 | [160] | 23922391 | |
| 3b | HVO_A0487 | cbiA | no | 37% | STM2035 | P29946 | [174] | 15311923 | equivalent reaction on cobalt-free substrate |
| 3b | HVO_B0052 | - | - | function unassigned; no GSP |
|||||
| 3b | HVO_B0053 | - | - | function unassigned; no GSP |
|||||
| 3b | HVO_B0055 | - | - | function unassigned; no GSP |
|||||
| 3b | HVO_B0056 | - | - | function unassigned; no GSP |
|||||
| 3c | HVO_A0488 | cobA | yes | 31% | MM_3138 | Q8PSE1 | [175] | 16672609 | |
| 3c | HVO_A0488 (cont.) |
yes | 30% | STM1718 | P31570 | [176] | 12080060 | ||
| 3c | HVO_2395 | pduO | yes | 37% | - |
Q9XDN2 |
[177] | 11160088 | PduO and CobA are isofunctional; In Q9XDN2, the PduO domain (N-term) is fused to a DUF336 domain |
| 3c | HVO_A0553 | cbiP | yes | 63% | VNG_1576G OE_3246F |
Q9HPL5 B0R5X2 |
[178] | 14645280 | |
| 3c | HVO_0587 | cbiB | yes | 58% | VNG_1578H OE_3253F |
Q9HPL3 B0R5X4 |
[178] | 14645280 | |
| 3c | HVO_0592 | cbiZ | yes | 57% | VNG_1583C OE_3261F |
Q9HPL3 B0R5X8 |
[179] | 14990804 | |
| 3c | HVO_0589 | cobY | yes | 47% | VNG_1581C OE_3257F |
Q9HPL1 B0R5X6 |
[180] | 12486068 | |
| 3c | HVO_0588 | cobS | yes | 30% | STM2017 | Q05602 | [181] | 17209023 | |
| 3c | - | STM0643 | P39701 | [182] | 7929373 | EC 3.1.3.73; CobC; no homolog in haloarchaea | |||
| 3c | HVO_0586 | - | prediction | - | - | - | [161] | 12869542 | EC 3.1.3.73; prediction for HSL01294 (VNG_1577C) |
| 3c | pathway gap | EC 2.7.1.177 | |||||||
| 3c | HVO_0591 | cobD1 | yes | 31% | STM0644 | P97084 | [183] | 9446573 | |
| 3c | HVO_0593 | cobD2 | yes | no GSP; 51% seq_id to HVO_0591 (cobD1) | |||||
| 3c | HVO_0590 | cobT | prediction | [161] | 12869542 | prediction for VNG_1572C | |||
| 3c | halTADL_3045 | cobT | yes | 39% | STM0644 | Q05603 | [184] | 8206834 | |
| 3d | HVO_B0051 | cobN | yes | 34% | - | P29929 | [185] | 1429466 | |
| 3d | HVO_B0051 (cont.) |
no | 29% | - | Q55284 | [186] [187] |
8663186 9716491 |
Mg chelatase | |
| 3d | HVO_B0050 | chlID | no | 46% | slr1030 | P51634 | [186] [187] |
8663186 9716491 |
match to N-term; Mg chelatase |
| 3d | HVO_B0050 (cont.) |
no | 33% | slr1777 | P52772 | [186] [187] |
8663186 9716491 |
match to complete sequence, incl distant match to N-term; Mg chelatase |
|
| 3e | HVO_2227 | ahbA | yes | 35% | - | I6UH61 | [158] | 21969545 | |
| 3e | HVO_2313 | ahbB | yes | 32% | - | I6UH61 | [158] | 21969545 | |
| 3f | HVO_1121 | ahbC | yes | 47% | Mbar_A1793 | Q46BK8 | [158] [188] |
21969545 24669201 |
|
| 3f | HVO_2144 | ahbD | self | [162] | 29284023 | EC 1.3.98.6 | |||
| 3f | HVO_2144 (cont.) |
yes | 42% | Mbar_A1458 | Q46CH7 | [188] | 24669201 | ||
| 3f | HVO_1871 | chdC | self | [162] | 29284023 | EC 1.3.98.5 | |||
| 3f | HVO_1871 (cont.) |
yes | 46% | BSU37670 | P39645 | [189] | 28123057 | ||
(c) The cobalamin biosynthesis and salvage reactions (those beyond ligand cobyrinate a,c diamide) involve “adenosylation of the corrin ring, attachment of the aminopropanol arm, and assembly of the nucleotide loop that bridges the lower ligand dimethylbenzimidazole and the corrin ring” [161]. The enzymes of these branches of cobalamin biosynthesis and their associated GSPs are listed in Table 3. Only two pathway gaps remain open. For one of these, a candidate was proposed upon a detailed bioinformatic analysis [161] (for further details, see Supplementary Text S1 Section S3).
(d) In the cobalt-late (aerobic) pathway variant, the intermediates are cobalt-free, and cobalt is inserted only late in the pathway. Even though haloarchaea do not use the cobalt-late pathway, so that a late cobaltochelatase is not required, they code for a homolog of the large subunit of a characterized heterotrimeric late cobaltochelatase. The adjacent gene is homologous to small subunits of other chelatases. We speculate that this late cobaltochelatase may be involved in cobalamin salvage. The chelatase has a mosaic subunit structure, as also reported previously [161] (see Supplementary Text S1 Section S3 for details).
(e) In the alternative heme biosynthesis pathway, siroheme is decarboxylated to 12,18-didecarboxysiroheme, which is attributed to the proteins encoded by ahbA and ahbB. These are homologous to each other and are organized as two two-domain proteins. It is unclear if AhbA and AhbB function independently or if they form a complex.
(f) Two of the three heme biosynthesis pathways (AHB and CPD) share a common last step (decarboxylation of Fe-coproporphyrin III to protoheme (heme b)). They use, however, distinct types of enzymes (AHB: ahbD, EC 1.3.98.6, adenosylmethionine-dependent heme synthase, a radical SAM enzyme; CPD: chdC, EC 1.3.98.5, peroxide-dependent heme synthase). Nearly all haloarchaea contain a chdC gene, and two-thirds also contain an ahbD gene. Hfx. volcanii was shown to use AhbD under anaerobic conditions and ChdC under aerobic conditions [162].
3.4. Coenzymes II: Coenzyme F420
Even though coenzyme F420 is predominantly associated with methanogenic archaea [190,191], it occurs also in bacteria, and a small amount of this coenzyme has been detected in non-methanogenic archaea, including halophiles [192]. The genes required for the biosynthesis of this coenzyme are encoded in haloarchaeal genomes, but the origin and attachment of the phospholactate moiety are not completely resolved (see below). To the best of our knowledge, only a single coenzyme F420-dependent enzymatic reaction has yet been reported for halophilic archaea [193]. Thus, the importance of this coenzyme in haloarchaeal biology is currently enigmatic and awaits experimental analysis.
(a) The pathway that creates the carbon backbone of this coenzyme has been reconstructed. We list the enzymes with their associated GSPs in Table 4. Coenzyme F420 contains a phospholactate moiety, which was reported to originate from 2-phospho-lactate [194], but this compound is metabolically not well-connected. As summarized in Supplementary Text S1 Section S4, there are various new insights regarding this pathway from recent studies in other prokaryotes [195,196]. To the best of our knowledge, the haloarchaeal coenzyme F420 biosynthesis pathway has never been experimentally analyzed.
Table 4.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.4). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 4a | HVO_2198 | cofH | yes | 35% | MJ1431 | Q58826 | [198] [199] |
14593448 25781338 |
|
| 4a | HVO_2201 | cofG | yes | 43% | MJ0446 | Q57888 | [198] [200] [199] |
14593448 23072415 25781338 |
|
| 4a | HVO_2202 | cofC | yes | 25% | MJ0887 | Q58297 | [194] [195] [196] |
18260642 30952857 31469543 |
|
| 4a | HVO_2479 | cofD | yes | 39% | MM_1874 | Q8PVT6 | [201] [196] |
18252724 31469543 |
|
| 4a | HVO_2479 (cont.) |
yes | 32% | MJ1256 | Q58653 | [202] | 11888293 | ||
| 4a | HVO_1936 | cofE | yes | 47% | AF_2256 | O28028 | [203] | 17669425 | |
| 4a | HVO_1936 (cont.) |
yes | 38% | MJ0768 | Q58178 | [204] | 12911320 | ||
| 4b | HVO_0433 | npdG | yes | 38% | AF_0892 | O29370 | [205] | not in PubMed | |
| 4b | HVO_B0113 | - | no | 27% | Rv0132c | P96809 | [206] | 24349169 | too distant to assume isofunctionality |
| 4b | HVO_B0342 | - | unknown | 29% | - | O93734 | [207] [208] |
8706724 15016352 |
too distant to assume isofunctionality |
| 4b | NP_1902A | - | no | 28% | - | Q9UXP0 | [209] [210] |
1735436 9933933 |
too distant to assume isofunctionality |
| 4b | NP_4006A | - | no | 27% | MJ0870 | Q58280 | [211] | 16048999 | too distant to assume isofunctionality |
| 4c/5c | HVO_1937 | mer | no | 38% | MTH_1752 | O27784 | [212] [213] [214] |
2298726 7649177 10891279 |
|
| 4d | HVO_2911 | phr2 | yes | 62% | VNG_1335G OE_2907R |
Q9HQ46 B0R5D6 |
[215] [216] |
2681164 12773185 |
|
| 4d | HVO_2843 | phr1 | no | 45% | sll1629 | P77967 | [217] | 12535521 | sll1629 implicated in transcription regulation |
| 4d | HVO_2843 (cont.) |
possibly | 45% | At5g24850 | Q84KJ5 | [218] [219] |
12834405 17062752 |
mediates photo-repair of ssDNA | |
| 4d | HVO_1234 | phr3 | possibly | 40% | Atu4765 | A9CH39 | [220] | 23589886 | |
(b) The prediction of coenzyme F420-specific oxidoreductases in Mycobacterium and actinobacteria has been reported [197], leading to patterns and domains that are also found in haloarchaea. Several such enzymes are described in Supplementary Text S1 Section S4.
(c) HVO_1937 might be a coenzyme F420-dependent 5,10-methylenetetrahydrofolate reductase (see, also, below: C1 metabolism, and Supplementary Text S1 Section S4).
(d) The precursor for coenzyme F420 may be used by a photolyase involved in DNA repair.
3.5. Coenzymes III: Coenzymes of C1 Metabolism: Tetrahydrofolate in Haloarchaea and Methanopterin in Methanogens
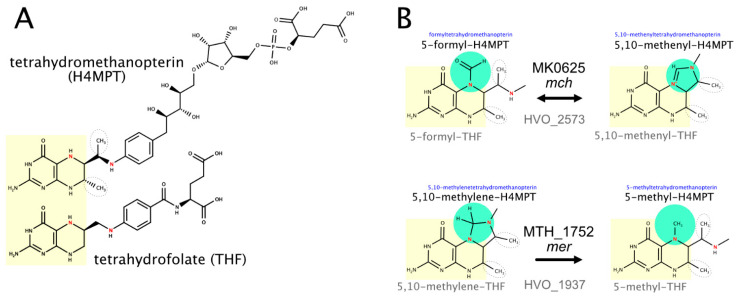
Halophilic and methanogenic archaea use distinct coenzymes as one-carbon carriers (C1 metabolism): tetrahydrofolate in haloarchaea and methanopterin in methanogens [221,222]. Several characterized methanogenic proteins that act on or with methanopterin have comparably close homologs in haloarchaea (Table 5), which results in the misannotation of haloarchaeal proteins (e.g., in SwissProt) as being involved in methanopterin biology. We assume that the haloarchaeal proteins function with the haloarchaeal one-carbon carrier tetrahydrofolate and that this shift in coenzyme specificity is possible due to the structural similarity between methanopterin and tetrahydrofolate (a near-identical core structure consisting of a pterin heterocyclic ring linked via a methylene bridge to a phenyl ring) (Figure 2). A detailed review on the many variants of the tetrahydrofolate biosynthetic pathway is available [223].
Table 5.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.5). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 5a | HVO_0709 | pabA | no | 47% | TTHA1843 | P05379 | [225] | 2844259 | Trp biosynthesis |
| 5a | HVO_0709 (cont.) |
yes/no | 39% | BSU00750 | P28819 | [226] | 2123867 | TrpG works with TrpE and with PabB | |
| 5a | HVO_0710 | pabB | no | 46% | TTHA1844 | P05378 | [225] | 2844259 | Trp biosynthesis |
| 5a | HVO_0710 (cont.) |
yes | 44% | BSU00740 | P28820 | [227] | 19275258 | PabB; para-aminobenzoate biosynthesis | |
| 5a | HVO_0708 | pabC | no | 36% | AF_0933 | O29329 | [228] | 30733943 | branched-chain amino acids |
| 5b | HVO_2348 | mptA | self | [229] | 19478918 | gene deletion phenotypes | |||
| 5b | HVO_2348 (cont.) |
yes | 41% | MJ0775 | Q58185 | [230] | 17497938 | common part of methanopterin and tetrahydrofolate biosynthesis | |
| 5b | HVO_A0533 | - | unknown | 27% | MJ0837 | Q58247 | [231] | 19746965 | if isofunctional would resolve a pathway gap |
| 5b | HVO_2628 | - | no | 31% | AF_2089 | O28190 | [232] | 12142414 | first committed step to methanopterin biosynthesis |
| 5b | HVO_2628 (cont.) |
no | 26% | MJ1427 | Q58822 | [233] | 15262968 | first committed step to methanopterin biosynthesis | |
| 5c | HVO_2573 | mch | no | 45% | MK0625 | P94954 | [234] | 9676239 | acts on a one-carbon attached to methanopterin |
| 4c/5c | HVO_1937 | mer | no | 38% | MTH_1752 | O27784 | [212] [213] [214] |
2298726 7649177 10891279 |
acts on a one-carbon compound attached to methanopterin |
Figure 2.
The structure of the C1 coenzymes tetrahydrofolate and methanopterin and two enzymes that act on the attached C1 compound. (A) The structures of tetrahydromethanopterin (top) and tetrahydrofolate (bottom) illustrate the similarities and differences between these C1 coenzymes. The common pteridine-based ring system is highlighted in yellow, and the initial biosynthesis step that generates this ring system is catalyzed by homologous enzymes (topic (b)). Two methanopterin-specific methyl groups are outlined by dashed ovals. N5 and N10, which are involved in the binding of the C1 compound, are colored red. (B) Two enzymatic reactions that alter the oxidation level of the C1 compound are illustrated. The methanogenic and haloarchaeal enzymes are homologous, even though they use distinct C1 coenzymes (topic (c)). It should be noted that MTH-1752 uses coenzyme F420 (not illustrated, Section 3.4, topic (c)), and this might also hold true for HVO_1937.
(a) Folate biosynthesis requires aminobenzoate. We proposed candidates for a pathway from chorismate to para-aminobenzoate [66,224] (for details, see Supplementary Text S1 Section S5). However, these predictions have not been adopted by KEGG (accessed April 2021), and without experimental confirmation, this is unlikely to ever happen.
(b) GTP cyclohydrolase MptA (HVO_2348) catalyzes a reaction in the common part of tetrahydrofolate and methanopterin biosynthesis. The enzymes specific for methanopterin biosynthesis are absent from haloarchaea, and thus, the assignment of HVO_2348 to the methanopterin biosynthesis pathway in UniProt is invalid (accessed March 2021).
The next common pathway step (EC 3.1.4.56) has been resolved in M. jannaschii (MJ0837) but is still a pathway gap in halophilic archaea. MJ0837 is very distantly related to HVO_A0533, which is a promising candidate for experimental analysis.
HVO_2628 shows 30% protein sequence identity with the enzyme catalyzing the first committed step to methanopterin biosynthesis. As detailed in Supplementary Text S1 Section S5, we consider it likely that it does not catalyze that reaction.
(c) Two enzymes that alter the oxidation level of the coenzyme-attached one-carbon compound probably function with tetrahydrofolate, even though their methanogenic homologs function with methanopterin. In contrast to their assignments in KEGG and UniProt (as of March 2021), their probable functions are thus methenyltetrahydrofolate cyclohydrolase (HVO_2573) and 5,10-methylenetetrahydrofolate reductase (HVO_1937) (see Figure 2 and Supplementary Text S1 Section S5).
3.6. Coenzymes IV: NAD and FAD (Riboflavin)
(a) The energy source for NAD kinase may be ATP or polyphosphate. This is unresolved for the two paralogs of probable NAD kinase (HVO_2363, nadK1 and HVO_0837, nadK2). These show only 25% protein sequence identity to each other (see Supplementary Text S1 Section S6). Polyphosphate was not found in exponentially growing Hfx. volcanii cells [235], and thus ATP is the more likely energy source.
(b) HVO_0781 is encoded in nearly all haloarchaeal genomes, according to OrthoDB, and shows very strong syntenic coupling with the adjacent gene, HVO_0782, according to SyntTax analysis. Characterized homologs to HVO_0781 cleave S-adenosyl-methionine into methionine and adenosine, a reaction that seems wasteful. If so, then this gene would not be expected to be retained in most species and neither would it maintain a strongly conserved gene clustering (see Supplementary Text S1 Section S6). HVO_0782 is an enzyme involved in NAD biosynthesis, which is encoded in most haloarchaeal and archaeal genomes. Thus, HVO_0781 is also a candidate for being involved in NAD biosynthesis.
(c) We described the reconstruction of riboflavin biosynthesis based on a detailed bioinformatic reconstruction [236]. The enzymes and their associated GSPs are listed in Table 6. Three pathway gaps remain, with candidate genes predicted for two of these [236] (for details, see Supplementary Text S1 Section S6).
Table 6.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.6). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 6a | HVO_2363 | nadK1 | unclear | 37% | Rv1695 | P9WHV7 | [237] | 11006082 | can use ATP and PP |
| 6a | HVO_2363 (cont.) |
unclear | 31% | AF_2373 | O30297 | ATP or PP usage unresolved | |||
| 6a | HVO_0837 | nadK2 | unclear | 28% | Rv1695 | P9WHV7 | can use ATP and PP | ||
| 6a | HVO_0837 (cont.) |
unclear | partial | AF_2373 | O30297 | ATP or PP usage unresolved | |||
| 6b | HVO_0782 | nadM | yes | 53% | MJ0541 | Q57961 | [238] [239] |
9401030 10331644 |
|
| 6b | HVO_0781 | - | unknown | 42% | Sare_1364 | A8M783 | [240] | 18720493 | |
| 6b | HVO_0781 (cont.) |
unknown | 35% | PH0463 | O58212 | [241] | 18551689 | ||
| 6c | HVO_0327 | ribB | yes | 43% | MJ0055 | Q60364 | [242] | 12200440 | |
| 6c | HVO_0974 | ribH | yes | 45% | MJ0303 | Q57751 | [243] | 12603336 | |
| 6c | HVO_1284 | arfA | self | [244] | 21999246 | gene deletion leads to riboflavin auxotrophy | |||
| 6c | HVO_1284 (cont.) |
yes | 44% | MJ0145 | Q57609 | [245] | 12475257 | ||
| 6c | HVO_1235 | - | prediction | [236] | 28073944 | arfB candidate | |||
| 6c | HVO_1341 | arfC | yes | 36% | MJ0671 | Q58085 | [246] [247] |
11889103 18671734 |
|
| 6c | HVO_2483 | - | prediction | 34% | MJ0699 | Q58110 | [236] | 28073944 | also predicted for MJ0699 |
| 6c | pathway gap | EC 3.1.3.104 | |||||||
| 6c | HVO_0326 | rbkR | yes | 37% | TA1064 | Q9HJA6 | [236] | 28073944 | bifunctional as gene regulator and enzyme |
| 6c | HVO_0326 (cont.) |
yes/no | 32% | MJ0056 | Q60365 | [248] | 18073108 | enzyme only; lacks an N-terminal HTH domain | |
| 6c | HVO_1015 | ribL | yes | 50% | MJ1179 | Q58579 | [249] | 20822113 | |
3.7. Biosynthesis of Membrane Lipids, Bacterioruberin and Menaquinone
Archaeal membrane lipids contain ether-linked isoprenoid side chains (see [250] and the references cited therein). The isoprenoid precursor isopentenyl diphosphate is synthesized in haloarchaea by a modified version of the mevalonate pathway [251]. Isoprenoid units are then linearly condensed into the C20 compound geranylgeranyl diphosphate. The haloarchaeal core lipid, archaeol, consists of 2,3-sn-glycerol with two C20 isoprenoid side chains attached by ether linkages. In some archaea, especially alkaliphiles, C25 isoprenoids are also found (see, e.g., [252,253]). Additionally, a number of distinct headgroups are found in polar lipids (phospholipids) (reviewed in [250]) (Figure 3). Even though polar lipids are used as important taxonomic markers [254], their biosynthetic pathways are not completely resolved.
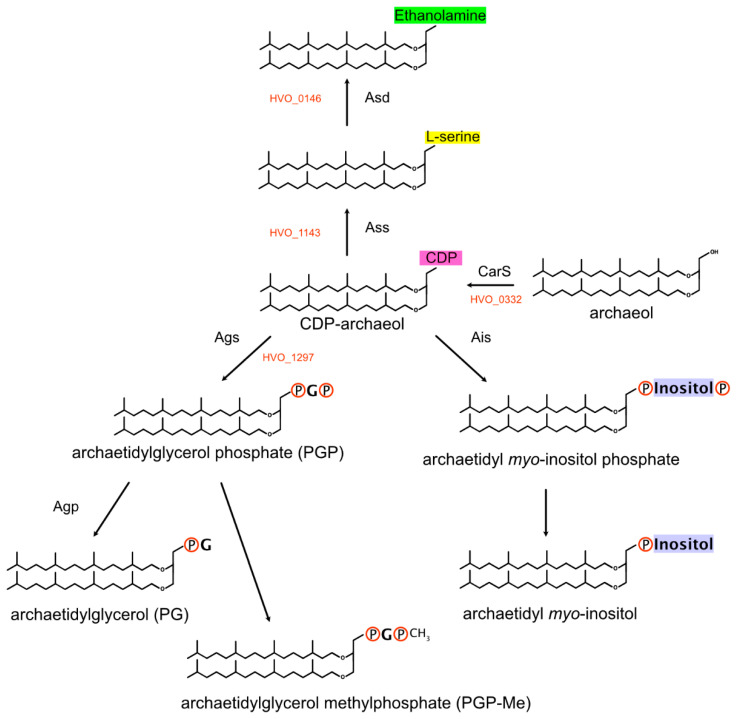
Figure 3.
Biosynthesis of polar lipids. A key intermediate is CDP-archaeol, which is generated from archaeol (displayed as fully saturated) by CarS. Members of the InterPro:IPR000462 family then transfer the CDP-archaeol to the hydroxyl group (alcohol group) of the target molecule (backbone: serine, glycerol and myo-inositol). Subsequent modifications contribute to the diversity of polar lipids.
Haloarchaea typically have a red color, which is due to carotenoids, mainly the C50 carotenoid bacterioruberin [255,256,257]. For carotenoid biosynthesis, two molecules of geranylgeranyl diphosphate, a C20 compound, are linked head-to-head to generate phytoene, which is desaturated to lycopene [66,258]. The pathway from lycopene to the C50 compound bacterioruberin has been experimentally characterized [257,259].
(a) We assigned HVO_2725 (idsA1, paralog of NP_3696A) and HVO_0303 (idsA2, paralog of NP_0604A) for the linear isoprenoid condensation reactions, resulting in a C20 isoprenoid (EC 2.5.1.10 and EC 2.5.1.29, short-chain isoprenyl diphosphate synthase) (see, also, Supplementary Text S1 Section S7). Some archaea, mainly haloalkaliphiles, also contain C25 isoprenoid side chains. Geranylfarnesyl diphosphate synthase, the enzyme that generates the C25 isoprenoids, has been purified and enzymatically characterized from Nmn. pharaonis [260], but data required for the assignment to a specific gene have not been collected. Three paralogous genes from Nmn. pharaonis are candidates for this function (NP_0604A, NP_3696A and NP_4556A). Since NP_0604A and NP_3696A have orthologs in Hfx. volcanii, a species devoid of C25 lipids, we assigned the synthesis of C25 isoprenoids (geranylfarnesyl diphosphate synthase activity) to the third paralog, NP_4556A. UniProt assigned C25 biosynthesis activity to NP_3696A for undescribed reasons (as of April 2021), and KEGG does not make this assignment for any of the three paralogs (as of April 2021). Our assignments are supported by an analysis of the key residues that determine the length of the isoprenoid chain [261]. These authors labeled the cluster containing NP_3696A (WP011323557.1) as “C15/C20” and the cluster containing NP_4556A (WP011323984.1) as “C20->C25->C30?”.
(b) Typical polar lipids in haloarchaea (Figure 3) are phosphatidylglycerophosphate methyl ester (PGP-Me) and phosphatidylglycerol (PG) but, also, phosphatidylglycerosulfate (PGS) [261,262,263]. Other polar lipids are archaetidylserine and its decarboxylation product archaetidylethanolamine, both of which are found in rather low quantities in Haloferax [264]. A third group of polar lipids has a headgroup derived from myo-inositol. The biosynthetic pathway of the headgroup is only partially resolved. One CDP-archaeol 1-archaetidyltransferase that belongs to a highly conserved three-gene operon may attach either glycerol phosphate or myo-inositol phosphate. In Supplementary Text S1 Section S7, we summarize the arguments in favor of each of these candidates, but the true function can only be decided by experimental analysis.
(c) Carotenoid biosynthesis involves the head-to-head condensation of the C20 isoprenoid geranylgeranyl diphosphate to phytoene, which is desaturated to lycopene [66,258]. The crtB gene product (e.g., HVO_2524) catalyzes the head-to-head condensation. It is yet uncertain which gene product is responsible for the desaturation of phytoene to lycopene. The further pathway from lycopene to bacterioruberin has been experimentally characterized in Haloarcula japonica [257]. A three-gene cluster (crtD-lyeJ-cruF) codes for the three enzymes of this pathway. The synteny of this three gene cluster is strongly conserved, according to SyntTax analysis. Several genes that are certainly or possibly involved in carotenoid biosynthesis are encoded in the vicinity of this cluster (for details, see Supplementary Text S1 Section S7).
(d) Halophilic archaea contain menaquinone as a lipid-based two-electron carrier of the respiratory chain [264,265]. We described the reconstruction of the menaquinone biosynthesis pathway (Table 7), with two pathway gaps remaining open (see Supplementary Text S1 Section S7 for details).
Table 7.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.7). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 7a | NP_0604A | idsA2 | yes | 32% | GACE_1337 | A0A0A7GEY4 | [266] | 30062607 | ortholog of HVO_0303 (66%); produces a C20 isoprenoid (same assignment for NP_0604A) |
| 7a | NP_0604A (cont.) |
idsA2 | no | 30% | APE_1764 |
Q9YB31 Q9UWR6 |
[267] | 10632701 | produces a C25 isoprenoid (C20 assigned to NP_0604A) |
| 7a | NP_3996A | idsA3 | yes | 44% | GACE_1337 | A0A0A7GEY4 | [266] | 30062607 | ortholog of HVO_2725 (67%); produces a C20 isoprenoid (same assignment for NP_3996A) |
| 7a | NP_3996A (cont.) |
idsA2 | no | 36% | APE_1764 |
Q9YB31 Q9UWR6 |
[267] | 10632701 | produces a C25 isoprenoid (C20 assigned to NP_3996A) |
| 7a | NP_4556A | idsA1 | no | 34% | GACE_1337 | A0A0A7GEY4 | [266] | 30062607 | no ortholog in Hfx. volcanii; produces a C20 isoprenoid (C25 assigned to NP_4556A) |
| 7a | NP_4556A (cont.) |
idsA1 | yes | 29% | APE_1764 |
Q9YB31 Q9UWR6 |
[267] | 10632701 | produces a C25 isoprenoid (same assignment for NP_4556A) |
| 7b | HVO_0332 | carS | yes | 45% | AF_1740 | O28537 | [268] | 25219966 | |
| 7b | HVO_1143 | assA | yes | 32% | MTH_1027 | O27106 | [269] | 12562787 | gene synonym: pgsA3 |
| 7b | HVO_1297 | aisA | yes | 25% | MTH_1691 | O27726 | [270] | 19740749 | gene synonym: pgsA2 |
| 7b | HVO_1136 | pgsA1 | - | only distant partial matches to GSPs | |||||
| 7b | HVO_1971 | pgsA4 | unclear | 26% | MTH_1027 | O27106 | [269] | 12562787 | MTH_1027 is less distant to HVO_1143 |
| 7b | HVO_0146 | asd | no | 39% | SMc00551 | Q9FDI9 | [271] | 18708506 | equivalent function for the bacterial lipid |
| 7b | HVO_1295 | hisC | self | [272] | 2345144 | complements a His auxotrophy mutant | |||
| 7b | HVO_1295 (cont.) |
yes | 31% | b2021 | P06986 | [273] | 2999081 | weak support, see text | |
| 7b | HVO_1296 | adk2 | unclear | 34% | PAB0757 | Q9UZK4 | [274] | 24823650 | Pyrococcus: involved in ribosome biogenesis |
| 7b | HVO_1296 (cont.) |
unclear | 32% | - | Q9Y3D8 | [275] | 15630091 | human: adenylate kinase; HVO_1296 may be inositol kinase | |
| 7b | HVO_2496 | adk1 | yes | 45% | BSU01370 | P16304 | [276] | 31111079 | Bacillus: adenylate kinase |
| 7b | HVO_B0213 | - | yes | 43% | AF_1794 | O28480 | [277] [278] |
11015222 22261071 |
Archaeoglobus: adenylate kinase |
| 7b | HVO_1135 | - | - | a SAM-dependent methyltransferase | |||||
| 7c | HVO_2524 | crtB | self | [9] [279] |
25488358 29038254 |
crtB mutants are colorless | |||
| 7c | HVO_2524 (cont.) |
yes | 32% | Synpcc7942 _1984 |
P37269 | [280] | 1537409 | ||
| 7c | HVO_2527 | lyeJ | self | [259] | 21840984 | ||||
| 7c | HVO_2527 (cont.) |
yes | 65% | VNG_1682C OE_3380R |
Q9HPD9 B0R651 |
[259] | 21840984 | ||
| 7c | HVO_2527 (cont.) |
yes | 61% | C444_12922 | M0L7V9 | [257] | 25712483 | ||
| 7c | HVO_2528 | crtD | self | [279] | 29038254 | a HVO_2528 mutant was white | |||
| 7c | HVO_2528 (cont.) |
yes | 71% | C444_12917 | A0A0A1GKA2 | [257] | 25712483 | ||
| 7c | HVO_2526 | cruF | yes | 59% | C444_12927 | A0A0A1GNF2 | [257] | 25712483 | |
| 7d | HVO_1470 | menF | yes | 38% | PA4231 | Q51508 | [281] | 7500944 | |
| 7d | HVO_1469 | menD | yes | 37% | BSU30820 | P23970 | [282] | 20600129 | |
| 7d | pathway gap | EC 4.2.99.20 | |||||||
| 7d | HVO_1461 | menC | no | 29% | BSU12980 | O34508 | [283] | 11747447 | Ala/Glu epimerase |
| 7d | HVO_1461 (cont.) |
yes | 24% | BSU30780 | O34514 | [284] | 10194342 | o-succinylbenzoate synthase | |
| 7d | HVO_1375 | menE | yes | 36% | BSU30790 | P23971 | [285] | 27933791 | |
| 7d | HVO_1465 | menB | yes | 66% | Rv0548c | P9WNP5 | [286] | 20643650 | |
| 7d | pathway gap | EC 3.1.2.28 | |||||||
| 7d | HVO_1462 | menA | yes | 37% | b3930 | P32166 | [287] | 9573170 | |
| 7d | HVO_0309 | menG | yes/no | 44% | At3g63410 | Q9LY74 | [288] | 14508009 | A. thaliana enzyme also involved in tocopherol biosynthesis |
| 7d | HVO_0309 (cont.) |
yes | 27% | - | O86169 | [289] | 9139683 | ||
3.8. Issues Concerning RNA Polymerase, Protein Translation Components and Signal Peptide Degradation
(a) Haloarchaeal RNA polymerase consists of a set of canonical subunits (encoded by rpoA1A2B1B2DEFHKLNP). Hbt. salinarum and a subset of other haloarchaea contain an additional subunit called epsilon [290,291]. Purified RNA polymerase containing the epsilon subunit transcribes native templates efficiently, in contrast to the RNA polymerase devoid of this subunit [291]. The biological relevance of this subunit is enigmatic (see Supplementary Text S1 Section S8).
(b) Two distant paralogs are found for haloarchaeal ribosomal protein S10 (uS10) in nearly all haloarchaeal genomes. It is uncertain if both occur in the ribosome, whether they occur together or are mutually exclusive. The latter distribution would result in heterogeneity of the ribosomes. Alternatively, one of the paralogs may exclusively have a non-ribosomal function.
In a subset of archaea, two distant paralogs are found for haloarchaeal ribosomal protein S14 (uS14) (ca 20% of the genomes, e.g., in Nmn. pharaonis). For more details, see Supplementary Text S1 Section S8.
(c) The ribosomal protein L43e (eL43) shows heterogeneity with respect to the presence of the C2–C2-type zinc finger motif. This zinc finger is found in L43e from all Halobacteriales and all euryarchaeal proteins outside the order Halobacteria but is not found in Haloferacales and is very rare in Natrialbales. Eukaryotic orthologs (e.g., from rat and yeast) contain this zinc finger, and its biological importance has been experimentally shown for the yeast protein [292] (for details, see Supplementary Text S1 Section S8).
(d) Diphthamide is a complex covalent modification of a histidine residue of translation elongation factor a-EF2. This pathway has been reconstructed (Table 8) based on distant homologs (enzymes encoded by dph2 and dph5) and by a detailed bioinformatic analysis (enzyme encoded by dph6) [293] (for details, see Supplementary Text S1 Section S8). These uncertain function assignments await experimental confirmation.
Table 8.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.8). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 8a | OE_1279R | rpoeps | self | [290] [291] |
2495365 6852054 |
||||
| 8b | HVO_0360 | rps10a | yes | 94% | rrnAC2405 | P23357 | [296] | 1764513 | |
| 8b | HVO_1392 | rps10b | - | no GSP; 24% seq_id to HVO_0360 (rps10a) | |||||
| 8b | NP_4882A | rps14a | yes | 72% | rrnAC1597.1 | P26816 | [297] | 1832208 | full-length similarity; Haloarcula protein was not isolated or characterized |
| 8b | NP_4882A (cont.) |
yes | 57% | YDL061C | P41058 | [298] | 18782943 | yeast YS29B; N-term 20 aa divergent |
|
| 8b | NP_1768A | rps14b | unclear | 80% | rrnAC1597.1 | P26816 | [297] | 1832208 | N-term 20 aa divergent |
| 8c | OE_1373R | rpl43e | yes | 69% | rrnAC1669 | P60619 | [299] | 10937989 | |
| 8c | OE_1373R (cont.) |
yes | 39% | YPR043W | P0CX25 | [292] [300] |
10588896 11866512 |
||
| 8c | HVO_0654 | rpl43e | yes | 54% | rrnAC1669 | P60619 | [299] | 10937989 |
Haloarcula: has zinc finger; Haloferax; lacks zinc finger |
| 8d | HVO_1631 | dph2 | yes | 35% | PH1105 | O58832 | [301] | 20931132 | |
| 8d | HVO_0916 | dph5 | yes | 39% | PH0725 | O58456 | [302] | 20873788 | |
| 8d | HVO_1077 | dph6 | yes | 31% | YLR143W | Q12429 | [303] [304] |
23169644 23468660 |
|
| 8e | HVO_0881 | sppA1 | yes | 33% | BSU19530 | O34525 | [305] [306] |
10455123 22472423 |
|
| 8e | HVO_1987 | sppA2 | probably | 23% | BSU19530 | O34525 | [305] [306] |
10455123 22472423 |
|
| 8e | HVO_1107 | - | prediction | no GSP | |||||
(e) N-terminal signal sequences target proteins to the secretion machinery. Subsequent to membrane insertion or transmembrane transfer, the signal sequence is cleaved off by a signal peptidase. After cleavage, the signal peptide must be degraded to avoid clogging of the membrane. Degradation is catalyzed by signal peptide peptidase. Candidates for this activity have been predicted from two protein families [294,295] (for details, see Supplementary Text S1 Section S8).
3.9. Miscellaneous Metabolic Enzymes and Proteins with Other Functions
Here, we list a few other enzymatic or nonenzymatic functions for which candidate genes have been assigned but without experimental validation.
(a) Ketohexokinase from Haloarcula vallismortis has been experimentally characterized [307]. However, the activity was not assigned to a gene. Detailed bioinformatic analyses have been made [308,309] and point to a small set of orthologs represented by Hmuk_2662, the ortholog of HVO_1812 (for further details, see Supplementary Text S1 Section S9).
(b) The assignment of fructokinase activity to the Hht. litchfieldiae candidate gene halTADL_1913 (UniProt:A0A1H6QYL4) is based on a differential proteomic analysis [309] (see Supplementary Text S1 Section S9 for details). Very close homologs are rare in haloarchaea. For this protein family (carbohydrate kinase), it is unclear if more distant homologs (with about 50% protein sequence identity) are isofunctional.
(c) A candidate gene for glucoamylase is HVO_1711 for the reasons described in Supplementary Text S1 Section S9. The enzyme from Halorubrum sodomense has been characterized [310], but the activity has not yet been assigned to a gene.
(d) A strong candidate for having glucose-6-phosphate isomerase activity is Hfx. volcanii HVO_1967 (pgi), based on 36% protein sequence identity to the characterized enzyme from M. jannaschii (MJ1605) [311] (Table 9).
Table 9.
Proteins with open annotation issues and their Gold Standard Protein homologs (Section 3.9). For a description of this table, see the legend to Table 1.
| Gold Standard Protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Section | Code | Gene | Isofunc | %seq_id | Locus Tag | UniProt | Reference | PMID | Comment |
| 9a | HVO_1812 | - | prediction | no GSP | |||||
| 9b | halTADL_1913 | - | yes | 37% | - | P26984 | [312] | 1809835 | |
| 9b | halTADL_1913 (cont.) |
- | yes | 31% | OCC_03567 |
Q7LYW8 H3ZP68 |
[313] | 15138858 | |
| 9c | HVO_1711 | - | probably | 33% | - | P29761 | [314] | 1633799 | P29761 matches to C-term half of HVO_1711 |
| 9c | HVO_1711 (cont.) |
- | probably | 51% | SAMN 04487937_ 2677 |
A0A1I6HD35 | [310] | 8305855 | correlation between PMID:8305855 and A0A1I6HD35 likely (see text) |
| 9d | HVO_1967 | pgi | yes | 36% | MJ1605 | Q59000 | [311] | 14655001 | |
| 9e | OE_1665R | kdgA | no | 31% | PA1010 | Q9I4W3 | [139] | 21396954 | GSP for dapA (see under 2a) |
| 9e | OE_1665R (cont.) |
probably | 30% | TTX_1156.1 TTX_1156a |
G4RJQ2 | [315] | 15869466 | ||
| 9e | OE_1665R (cont.) |
probably | 25% | SSO3197 | Q97U28 | [315] | 15869466 | ||
| 9f | HVO_1692 | ludB | self | [21] | 30707467 | ||||
| 9f | HVO_1692 (cont.) |
probably | 35% | BSU34040 | O07021 | [316] | 19201793 | matches up to HVO_1692 pos 490 of 733 | |
| 9f | HVO_1692 (cont.) |
probably | 35% | PST_3338 | O4VPR6 | [317] | 25917905 | matches up to HVO_1692 pos 400 of 733 | |
| 9f | HVO_1693 | ludC | self | [21] | 30707467 | ||||
| 9f | HVO_1693 (cont.) |
probably | 30% | BSU34030 | O32259 | [316] | 19201793 | ||
| 9f | HVO_1693 (cont.) |
probably | 33% | PST_3339 | O4VPR7 | [317] | 25917905 | partial match | |
| 9f | HVO_1697 | - | unclear | 24% | PST_3340 | O4VPR8 | [317] | 25917905 | |
| 9f | HVO_1696 | lctP | probably | 44% | PST_3336 | O4VPR4 | [317] | 25917905 | |
| 9g | HVO_B0300 | pucL1 | yes | 49% | BSU32450 | O32141 | [318] | 20168977 | Bacillus: bifunctional, matches to C-term |
| 9g | HVO_B0299 | pucM | yes | 43% | BSU32460 | O32142 | [319] | 16098976 | |
| 9g | HVO_B0301 | pucL2 | yes | 43% | BSU32450 | O32141 | [320] | 17567580 | Bacillus: bifunctional, matches to N-term |
| 9g | HVO_B0302 | pucH1 | no | 33% | - | Q8VTT5 | [321] | 12148274 | paper in Chinese, abstract in English; pyrimidine degradation |
| 9g | HVO_B0302 (cont.) |
yes | 30% | STM0523 | Q7CR08 | [322] | 23287969 | purine degradation | |
| 9g | HVO_B0302 (cont.) |
yes | 29% | BSU32410 | O32137 | [323] | 11344136 | purine degradation | |
| 9g | HVO_B0306 | amaB4 | no | 39% | - | Q53389 | [324] | 22904279 | carbamoyl-AA hydrolysis |
| 9g | HVO_B0306 (cont.) |
yes | 34% | At5g43600 | Q8VXY9 | [325] [326] |
19935661 23940254 |
purine degradation | |
| 9g | HVO_B0308 | coxS | no | 46% | Saci_2270 | Q4J6M5 | [327] | 10095793 | GAPDH |
| 9g | HVO_B0308 (cont.) |
no | 41% | - | P19915 | [328] | 10482497 | CO-DH | |
| 9g | HVO_B0308 (cont.) |
yes | 39% | b2868 | Q46801 | [329] | 10986234 | xanthine DH | |
| 9g | HVO_B0309 | coxL | yes | 33% | b2866 | Q46799 | [329] | 10986234 | xanthine DH |
| 9g | HVO_B0309 (cont.) |
no | 28% | - | P19913 | [328] | 10482497 | CO-DH | |
| 9g | HVO_B0309 (cont.) |
no | 26% | Saci_2271 | Q4J6M3 | [327] | 10095793 | GAPDH | |
| 9g | HVO_B0310 | coxM | no | 31% | Saci_2269 | Q4J6M6 | [327] | 10095793 | GAPDH |
| 9g | HVO_B0310 (cont.) |
no | 31% | - | P19914 | [328] | 10482497 | CO-DH | |
| 9g | HVO_B0310 (cont.) |
yes | 25% | b2867 | Q46800 | [329] | 10986234 | xanthine DH | |
| 9g | HVO_B0303 | uraA4 | yes | 38% | b3654 | P0AGM9 | [330] | 16096267 | |
| 9h | HVO_0197 | - | possibly | 39% | lp_0105 | F9UST0 | [331] | 27114550 | LarB family protein |
| 9h | HVO_2381 | - | possibly | 31% | lp_0106/ lp_0107 |
F9UST1 | [331] | 27114550 | LarC family protein |
| 9h | HVO_0190 | - | possibly | 34% | lp_0109 | F9UST4 | [331] | 27114550 | LarE family protein |
| 9i | HVO_1660 | dacZ | self | [37] | 30884174 | ||||
| 9i | HVO_0756 | - | prediction | [332] | 32095817 | ||||
| 9i | HVO_0990 | - | prediction | [332] | 32095817 | ||||
| 9i | HVO_1690 | - | prediction | [332] | 32095817 | ||||
| 9j | HVO_2763 | - | self | [333] | 22350204 | no function could be assigned | |||
| 9j | HVO_2763 (cont.) |
no | 27% | HVO_0144 | D4GZ88 | [334] | 18437358 | Rnase Z | |
| 9k | HVO_2410 | dabA | yes | 33% | Hneap_0211 | D0KWS7 | [335] | 31406332 | |
| 9k | HVO_2411 | dabB | yes | 31% | Hneap_0212 | D0KWS8 | [335] | 31406332 | |
(e) A candidate gene for specifying an enzyme with 2-dehydro-3-deoxy-(phospho)gluconate aldolase activity is Hbt. salinarum kdgA (OE_1665R). It is rather closely related (36% protein sequence) to Hfx. volcanii HVO_1101 (encoded by dapA), which is involved in lysine biosynthesis, a biosynthetic pathway that is absent from Hbt. salinarum. The function assignment is based on distant homologs from Saccharolobus (Sulfolobus) solfataricus and Thermoproteus tenax, which have been characterized [315] (for details, see Supplementary Text S1 Section S9).
(f) Haloarchaea may contain an NAD-independent L-lactate dehydrogenase, LudBC (HVO_1692 and HVO_1693). The deletion of this gene pair impairs growth on rhamnose, which is catabolized to pyruvate and lactate [21]. There is a very distant relationship (for details, see Supplementary Text S1 Section S9) to the LldABC subunits of the characterized L-lactate dehydrogenase from Pseudomonas stutzeri A1501 [317] and to the LutABC proteins from B. subtilis, which have been shown to be involved in lactate utilization [316].
(g) Hfx. volcanii may be able to convert urate into allantoin using the gene cluster HVO_B0299-HVO_B0302. This could be part of a complete degradation pathway for purines, but this has to be considered highly speculative (see Supplementary Text S1 Section S9 and Supplementary Figure S2).
(h) Hfx. volcanii may contain an enzyme having a “nickel-pincer cofactor”. The biogenesis of this cofactor may be catalyzed by larBCE (as detailed in Supplementary Text S1 Section S9).
(i) Cyclic di-AMP (c-di-AMP) is an important nucleotide signaling molecule in bacteria and archaea. It is generated from two molecules of ATP by diadenylate cyclase (encoded by dacZ) and is degraded to pApA by phosphodiesterases [336]. The level of this signaling molecule is strictly controlled [337,338], thus requiring a sophisticated interplay of cyclase and phosphodiesterase. DacZ from Hfx. volcanii has been characterized, and it was shown that the c-di-AMP levels must be tightly regulated [37]. The degrading enzyme, however, has not yet been identified in Haloferax, but candidates have been proposed [332,336,339] (see Supplementary Text S1 Section S9).
(j) HVO_2763 is distantly related to RNase Z (HVO_0144, rnz). The experimental characterization of HVO_2763 [333] excluded activity as an exonuclease but did not reveal its physiological function. Upon transcriptome analysis, the downregulation of several genes was detected. Several of these were uncharacterized at the time of the experiment but have since been shown to be involved in the minor N-glycosylation pathway that was initially detected under low-salt conditions (see Supplementary Text S1 Section S9 for further details).
(k) A pair of genes (dabAB, HVO_2410 and HVO_2411) is predicted to function as a carbon dioxide transporter, based on the identification of such transporters in Halothiobacillus neapolitanus [335]. Being a member of the proton-conducting membrane transporter family, this protein may be misannotated as a subunit of the nuo or mrp complexes (see Supplementary Text S1 Section S9 for further details).
4. Conclusions
We described a large number of cases where the protein function cannot be correctly predicted when restricting considerations to the computational analyses without taking the biological contexts into account. An example was the switch from methanopterin to tetrahydrofolate as a C1 carrier in haloarchaea. Homologous enzymes, inherited from the common ancestor, have adapted to the new C1 carrier, rather than being replaced by non-homologous proteins. Function prediction tools may misannotate haloarchaeal proteins to work with methanopterin. Another example was the nuo complex and its misannotation as a type I NADH dehydrogenase. In other cases, even a distant sequence similarity may allow a valid function prediction if additional evidence (e.g., from a gene neighborhood analysis or from a detailed evaluation of the metabolic pathway gaps) is taken into account. Examples include cobalamin cluster proteins, which probably close the two residual pathway gaps, and the predicted degradation pathway for purines. In all these cases, we presented reasonable hypotheses based on the current knowledge, and in many cases, these were so well-supported as to be compelling, but to be certain, experimental data are required. With this overview, we attempted to arouse the curiosity of our colleagues, hoping that they will confirm or disprove our speculations and, thus, advance the knowledge about haloarchaeal biology. Hfx. volcanii is a model species for halophilic archaea, and the more complete and correctly its genome is annotated, the higher will be its value for system biology analyses (modeling) and for synthetic biology (metabolic engineering) and biotechnology.
Acknowledgments
We thank all members of the Haloferax community for more than a decade of fruitful cooperation and for sharing their deep knowledge about the biology of this model species. We thank our colleagues for kindly reading and providing thoughtful comments on the manuscript before submission: Sonja-Verena Albers, Thorsten Allers, Maria Jose Bonete, Rosana de Castro, Sebastien Ferreira-Cerca, Anita Marchfelder, Mechthild Pohlschroder and Joerg Soppa. We thank Birgit Scharf for the insightful discussions on the Natronomonas respiratory chain.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/genes12070963/s1: Text S1: Detailed background information for all open annotation issues. Figure S1: Alignment of the 5’ end of leuA2 (HVO_1510) from Hfx. volcanii DS2 with homologs from other species of Haloferax. Figure S2: The proposed purine degradation pathway. Table S1: Listing of all proteins mentioned in the text and in Supplementary Text S1. Table S2: Listing of the genomes which are manually curated and kept up to date.
Author Contributions
Conceptualization, F.P.; Data curation, F.P. Project administration, F.P. Formal analysis, F.P. and M.D.-S.; Investigation F.P. and M.D.-S. Validation, F.P. and M.D.-S.; Visualization, F.P. and M.D.-S. Writing—original draft, F.P. and M.D.-S. and Writing—review and editing, F.P. and M.D.-S. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no specific grants from any funding agency in the public, commercial or non-for-profit sectors.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
Footnotes
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Hartman A.L., Norais C., Badger J.H., Delmas S., Haldenby S., Madupu R., Robinson J., Khouri H., Ren Q., Lowe T.M., et al. The complete genome sequence of Haloferax volcanii DS2, a model archaeon. PLoS ONE. 2010;5:e9605. doi: 10.1371/journal.pone.0009605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schulze S., Adams Z., Cerletti M., De Castro R., Ferreira-Cerca S., Fufezan C., Gimenez M.I., Hippler M., Jevtic Z., Knuppel R., et al. The Archaeal Proteome Project advances knowledge about archaeal cell biology through comprehensive proteomics. Nat. Commun. 2020;11:3145. doi: 10.1038/s41467-020-16784-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Leigh J.A., Albers S.V., Atomi H., Allers T. Model organisms for genetics in the domain Archaea: Methanogens, halophiles, Thermococcales and Sulfolobales. FEMS Microbiol. Rev. 2011;35:577–608. doi: 10.1111/j.1574-6976.2011.00265.x. [DOI] [PubMed] [Google Scholar]
- 4.Perez-Arnaiz P., Dattani A., Smith V., Allers T. Haloferax volcanii-a model archaeon for studying DNA replication and repair. Open Biol. 2020;10:200293. doi: 10.1098/rsob.200293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Soppa J. Functional genomic and advanced genetic studies reveal novel insights into the metabolism, regulation, and biology of Haloferax volcanii. Archaea. 2011;2011:602408. doi: 10.1155/2011/602408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Haque R.U., Paradisi F., Allers T. Haloferax volcanii for biotechnology applications: Challenges, current state and perspectives. Appl Microbiol Biotechnol. 2020;104:1371–1382. doi: 10.1007/s00253-019-10314-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Allers T., Barak S., Liddell S., Wardell K., Mevarech M. Improved strains and plasmid vectors for conditional overexpression of His-tagged proteins in Haloferax volcanii. Appl Environ Microbiol. 2010;76:1759–1769. doi: 10.1128/AEM.02670-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Allers T., Mevarech M. Archaeal genetics—the third way. Nat. Rev. Genet. 2005;6:58–73. doi: 10.1038/nrg1504. [DOI] [PubMed] [Google Scholar]
- 9.Kiljunen S., Pajunen M.I., Dilks K., Storf S., Pohlschroder M., Savilahti H. Generation of comprehensive transposon insertion mutant library for the model archaeon, Haloferax volcanii, and its use for gene discovery. BMC Biol. 2014;12:103. doi: 10.1186/s12915-014-0103-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pfeiffer F., Broicher A., Gillich T., Klee K., Mejia J., Rampp M., Oesterhelt D. Genome information management and integrated data analysis with HaloLex. Arch. Microbiol. 2008;190:281–299. doi: 10.1007/s00203-008-0389-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pfeiffer F., Oesterhelt D. A manual curation strategy to improve genome annotation: Application to a set of haloarchael genomes. Life (Basel) 2015;5:1427–1444. doi: 10.3390/life5021427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Turkowyd B., Schreiber S., Wortz J., Segal E.S., Mevarech M., Duggin I.G., Marchfelder A., Endesfelder U. Establishing live-cell single-molecule localization microscopy imaging and single-particle tracking in the archaeon Haloferax volcanii. Front. Microbiol. 2020;11:583010. doi: 10.3389/fmicb.2020.583010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Walsh J.C., Angstmann C.N., Bisson-Filho A.W., Garner E.C., Duggin I.G., Curmi P.M.G. Division plane placement in pleomorphic archaea is dynamically coupled to cell shape. Mol. Microbiol. 2019;112:785–799. doi: 10.1111/mmi.14316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.de Silva R.T., Abdul-Halim M.F., Pittrich D.A., Brown H.J., Pohlschroder M., Duggin I.G. Improved growth and morphological plasticity of Haloferax volcanii. Microbiology (Reading) 2021;167:001012. doi: 10.1099/mic.0.001012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Duggin I.G., Aylett C.H., Walsh J.C., Michie K.A., Wang Q., Turnbull L., Dawson E.M., Harry E.J., Whitchurch C.B., Amos L.A., et al. CetZ tubulin-like proteins control archaeal cell shape. Nature. 2015;519:362–365. doi: 10.1038/nature13983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Liao Y., Ithurbide S., Evenhuis C., Lowe J., Duggin I.G. Cell division in the archaeon Haloferax volcanii relies on two FtsZ proteins with distinct functions in division ring assembly and constriction. Nat. Microbiol. 2021;6:594–605. doi: 10.1038/s41564-021-00894-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brasen C., Schonheit P. Mechanisms of acetate formation and acetate activation in halophilic archaea. Arch. Microbiol. 2001;175:360–368. doi: 10.1007/s002030100273. [DOI] [PubMed] [Google Scholar]
- 18.Johnsen U., Dambeck M., Zaiss H., Fuhrer T., Soppa J., Sauer U., Schonheit P. D-xylose degradation pathway in the halophilic archaeon Haloferax volcanii. J. Biol. Chem. 2009;284:27290–27303. doi: 10.1074/jbc.M109.003814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pickl A., Johnsen U., Schonheit P. Fructose degradation in the haloarchaeon Haloferax volcanii involves a bacterial type phosphoenolpyruvate-dependent phosphotransferase system, fructose-1-phosphate kinase, and class II fructose-1,6-bisphosphate aldolase. J. Bacteriol. 2012;194:3088–3097. doi: 10.1128/JB.00200-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sutter J.M., Tastensen J.B., Johnsen U., Soppa J., Schonheit P. Key enzymes of the semiphosphorylative Entner-Doudoroff pathway in the haloarchaeon Haloferax volcanii: Characterization of glucose dehydrogenase, gluconate dehydratase and 2-keto-3-deoxy-6-phosphogluconate aldolase. J. Bacteriol. 2016;198:2251–2262. doi: 10.1128/JB.00286-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Reinhardt A., Johnsen U., Schonheit P. L-Rhamnose catabolism in archaea. Mol. Microbiol. 2019;111:1093–1108. doi: 10.1111/mmi.14213. [DOI] [PubMed] [Google Scholar]
- 22.Kuprat T., Ortjohann M., Johnsen U., Schonheit P. Glucose metabolism and acetate switch in Archaea: The enzymes in Haloferax volcanii. J. Bacteriol. 2021;203:e00690-20. doi: 10.1128/JB.00690-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kuprat T., Johnsen U., Ortjohann M., Schonheit P. Acetate metabolism in Archaea: Characterization of an acetate transporter and of enzymes involved in acetate activation and gluconeogenesis in Haloferax volcanii. Front. Microbiol. 2020;11:604926. doi: 10.3389/fmicb.2020.604926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sutter J.M., Johnsen U., Reinhardt A., Schonheit P. Pentose degradation in archaea: Halorhabdus species degrade D-xylose, L-arabinose and D-ribose via bacterial-type pathways. Extremophiles. 2020;24:759–772. doi: 10.1007/s00792-020-01192-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tästensen J.B., Johnsen U., Reinhardt A., Ortjohann M., Schonheit P. D-galactose catabolism in archaea: Operation of the DeLey-Doudoroff pathway in Haloferax volcanii. FEMS Microbiol. Lett. 2020;367:fnaa029. doi: 10.1093/femsle/fnaa029. [DOI] [PubMed] [Google Scholar]
- 26.Abdul-Halim M.F., Schulze S., DiLucido A., Pfeiffer F., Bisson Filho A.W., Pohlschroder M. Lipid anchoring of archaeosortase substrates and midcell growth in haloarchaea. mBio. 2020;11:e00349-20. doi: 10.1128/mBio.00349-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Abdul Halim M.F., Rodriguez R., Stoltzfus J.D., Duggin I.G., Pohlschroder M. Conserved residues are critical for Haloferax volcanii archaeosortase catalytic activity: Implications for convergent evolution of the catalytic mechanisms of non-homologous sortases from archaea and bacteria. Mol. Microbiol. 2018;108:276–287. doi: 10.1111/mmi.13935. [DOI] [PubMed] [Google Scholar]
- 28.Abdul Halim M.F., Pfeiffer F., Zou J., Frisch A., Haft D., Wu S., Tolic N., Brewer H., Payne S.H., Pasa-Tolic L., et al . Haloferax volcanii archaeosortase is required for motility, mating, and C-terminal processing of the S-layer glycoprotein. Mol. Microbiol. 2013;88:1164–1175. doi: 10.1111/mmi.12248. [DOI] [PubMed] [Google Scholar]
- 29.Storf S., Pfeiffer F., Dilks K., Chen Z.Q., Imam S., Pohlschroder M. Mutational and bioinformatic analysis of haloarchaeal lipobox-containing proteins. Archaea. 2010;2010:410975. doi: 10.1155/2010/410975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schiller H., Schulze S., Mutan Z., de Vaulx C., Runcie C., Schwartz J., Rados T., Bisson Filho A.W., Pohlschroder M. Haloferax volcanii immersed liquid biofilms develop independently of known biofilm machineries and exhibit rapid honeycomb pattern formation. mSphere. 2020;5:e00976-20. doi: 10.1128/mSphere.00976-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pohlschroder M., Esquivel R.N. Archaeal type IV pili and their involvement in biofilm formation. Front. Microbiol. 2015;6:190. doi: 10.3389/fmicb.2015.00190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Li Z., Rodriguez-Franco M., Albers S.V., Quax T.E.F. The switch complex ArlCDE connects the chemotaxis system and the archaellum. Mol. Microbiol. 2020;114:468–479. doi: 10.1111/mmi.14527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Collins M., Afolayan S., Igiraneza A.B., Schiller H., Krespan E., Beiting D.P., Dyall-Smith M., Pfeiffer F., Pohlschroder M. Mutations affecting HVO_1357 or HVO_2248 cause hypermotility in Haloferax volcanii, suggesting roles in motility regulation. Genes (Basel) 2020;12:58. doi: 10.3390/genes12010058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Quax T.E.F., Altegoer F., Rossi F., Li Z., Rodriguez-Franco M., Kraus F., Bange G., Albers S.V. Structure and function of the archaeal response regulator CheY. Proc. Natl. Acad. Sci. USA. 2018;115:E1259–E1268. doi: 10.1073/pnas.1716661115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nussbaum P., Ithurbide S., Walsh J.C., Patro M., Delpech F., Rodriguez-Franco M., Curmi P.M.G., Duggin I.G., Quax T.E.F., Albers S.V. An oscillating MinD protein determines the cellular positioning of the motility machinery in Archaea. Curr. Biol. 2020;30:4956–4972. doi: 10.1016/j.cub.2020.09.073. [DOI] [PubMed] [Google Scholar]
- 36.Shalev Y., Turgeman-Grott I., Tamir A., Eichler J., Gophna U. Cell surface glycosylation is required for efficient mating of Haloferax volcanii. Front. Microbiol. 2017;8:1253. doi: 10.3389/fmicb.2017.01253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Braun F., Thomalla L., van der Does C., Quax T.E.F., Allers T., Kaever V., Albers S.V. Cyclic nucleotides in archaea: Cyclic di-AMP in the archaeon Haloferax volcanii and its putative role. Microbiologyopen. 2019;8:e00829. doi: 10.1002/mbo3.829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Maier L.K., Stachler A.E., Brendel J., Stoll B., Fischer S., Haas K.A., Schwarz T.S., Alkhnbashi O.S., Sharma K., Urlaub H., et al. The nuts and bolts of the Haloferax CRISPR-Cas system I-B. RNA Biol. 2019;16:469–480. doi: 10.1080/15476286.2018.1460994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Reuter C.J., Maupin-Furlow J.A. Analysis of proteasome-dependent proteolysis in Haloferax volcanii cells, using short-lived green fluorescent proteins. Appl. Environ. Microbiol. 2004;70:7530–7538. doi: 10.1128/AEM.70.12.7530-7538.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Reuter C.J., Uthandi S., Puentes J.A., Maupin-Furlow J.A. Hydrophobic carboxy-terminal residues dramatically reduce protein levels in the haloarchaeon Haloferax volcanii. Microbiology (Reading) 2010;156:248–255. doi: 10.1099/mic.0.032995-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Prunetti L., Reuter C.J., Hepowit N.L., Wu Y., Barrueto L., Miranda H.V., Kelly K., Maupin-Furlow J.A. Structural and biochemical properties of an extreme ’salt-loving’ proteasome activating nucleotidase from the archaeon Haloferax volcanii. Extremophiles. 2014;18:283–293. doi: 10.1007/s00792-013-0615-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cerletti M., Paggi R., Troetschel C., Ferrari M.C., Guevara C.R., Albaum S., Poetsch A., De Castro R. LonB protease is a novel regulator of carotenogenesis controlling degradation of phytoene synthase in Haloferax volcanii. J. Proteome Res. 2018;17:1158–1171. doi: 10.1021/acs.jproteome.7b00809. [DOI] [PubMed] [Google Scholar]
- 43.Cerletti M., Martinez M.J., Gimenez M.I., Sastre D.E., Paggi R.A., De Castro R.E. The LonB protease controls membrane lipids composition and is essential for viability in the extremophilic haloarchaeon Haloferax volcanii. Environ. Microbiol. 2014;16:1779–1792. doi: 10.1111/1462-2920.12385. [DOI] [PubMed] [Google Scholar]
- 44.Costa M.I., Cerletti M., Paggi R.A., Trotschel C., De Castro R.E., Poetsch A., Gimenez M.I. Haloferax volcanii proteome response to deletion of a rhomboid protease gene. J. Proteome Res. 2018;17:961–977. doi: 10.1021/acs.jproteome.7b00530. [DOI] [PubMed] [Google Scholar]
- 45.Cao S., Hepowit N., Maupin-Furlow J.A. Ubiquitin-like protein SAMP1 and JAMM/MPN+ metalloprotease HvJAMM1 constitute a system for reversible regulation of metabolic enzyme activity in Archaea. PLoS ONE. 2015;10:e0128399. doi: 10.1371/journal.pone.0128399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kaminski L., Eichler J. Haloferax volcanii N-glycosylation: Delineating the pathway of dTDP-rhamnose biosynthesis. PLoS ONE. 2014;9:e97441. doi: 10.1371/journal.pone.0097441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tripepi M., You J., Temel S., Onder O., Brisson D., Pohlschroder M. N-Glycosylation of Haloferax volcanii flagellins requires known Agl proteins and Is essential for biosynthesis of stable flagella. J. Bacteriol. 2012;194:4876–4887. doi: 10.1128/JB.00731-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gribaldo S., Schulze S., Pfeiffer F., Garcia B.A., Pohlschroder M. Comprehensive glycoproteomics shines new light on the complexity and extent of glycosylation in archaea. PLoS Biol. 2021;19:e3001277. doi: 10.1371/journal.pbio.3001277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shalev Y., Soucy S.M., Papke R.T., Gogarten J.P., Eichler J., Gophna U. Comparative analysis of surface layer glycoproteins and genes involved in protein glycosylation in the genus Haloferax. Genes (Basel) 2018;9:172. doi: 10.3390/genes9030172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kandiba L., Lin C.W., Aebi M., Eichler J., Guerardel Y. Structural characterization of the N-linked pentasaccharide decorating glycoproteins of the halophilic archaeon Haloferax volcanii. Glycobiology. 2016;26:745–756. doi: 10.1093/glycob/cww014. [DOI] [PubMed] [Google Scholar]
- 51.Qi Q., Ito Y., Yoshimatsu K., Fujiwara T. Transcriptional regulation of dimethyl sulfoxide respiration in a haloarchaeon, Haloferax volcanii. Extremophiles. 2016;20:27–36. doi: 10.1007/s00792-015-0794-6. [DOI] [PubMed] [Google Scholar]
- 52.Rawls K.S., Yacovone S.K., Maupin-Furlow J.A. GlpR represses fructose and glucose metabolic enzymes at the level of transcription in the haloarchaeon Haloferax volcanii. J. Bacteriol. 2010;192:6251–6260. doi: 10.1128/JB.00827-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hattori T., Shiba H., Ashiki K., Araki T., Nagashima Y.K., Yoshimatsu K., Fujiwara T. Anaerobic growth of haloarchaeon Haloferax volcanii by denitrification is controlled by the transcription regulator NarO. J. Bacteriol. 2016;198:1077–1086. doi: 10.1128/JB.00833-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hwang S., Cordova B., Abdo M., Pfeiffer F., Maupin-Furlow J.A. ThiN as a versatile domain of transcriptional repressors and catalytic enzymes of thiamine biosynthesis. J. Bacteriol. 2017;199:e00810–e00816. doi: 10.1128/JB.00810-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Johnsen U., Sutter J.M., Schulz A.C., Tastensen J.B., Schonheit P. XacR—A novel transcriptional regulator of D-xylose and L-arabinose catabolism in the haloarchaeon Haloferax volcanii. Environ. Microbiol. 2015;17:1663–1676. doi: 10.1111/1462-2920.12603. [DOI] [PubMed] [Google Scholar]
- 56.Zahn S., Kubatova N., Pyper D.J., Cassidy L., Saxena K., Tholey A., Schwalbe H., Soppa J. Biological functions, genetic and biochemical characterization, and NMR structure determination of the small zinc finger protein HVO_2753 from Haloferax volcanii. FEBS J. 2021;288:2042–2062. doi: 10.1111/febs.15559. [DOI] [PubMed] [Google Scholar]
- 57.Nagel C., Machulla A., Zahn S., Soppa J. Several one-domain zinc finger micro-proteins of Haloferax volcanii are important for stress adaptation, biofilm formation, and swarming. Genes (Basel) 2019;10:361. doi: 10.3390/genes10050361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kubatova N., Jonker H.R.A., Saxena K., Richter C., Vogel V., Schreiber S., Marchfelder A., Schwalbe H. Solution Structure and Dynamics of the Small Protein HVO_2922 from Haloferax volcanii. ChemBioChem. 2020;21:149–156. doi: 10.1002/cbic.201900085. [DOI] [PubMed] [Google Scholar]
- 59.Straub J., Brenneis M., Jellen-Ritter A., Heyer R., Soppa J., Marchfelder A. Small RNAs in haloarchaea: Identification, differential expression and biological function. RNA Biol. 2009;6:281–292. doi: 10.4161/rna.6.3.8357. [DOI] [PubMed] [Google Scholar]
- 60.Heyer R., Dorr M., Jellen-Ritter A., Spath B., Babski J., Jaschinski K., Soppa J., Marchfelder A. High throughput sequencing reveals a plethora of small RNAs including tRNA derived fragments in Haloferax volcanii. RNA Biol. 2012;9:1011–1018. doi: 10.4161/rna.20826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Babski J., Maier L.K., Heyer R., Jaschinski K., Prasse D., Jager D., Randau L., Schmitz R.A., Marchfelder A., Soppa J. Small regulatory RNAs in Archaea. RNA Biol. 2014;11:484–493. doi: 10.4161/rna.28452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Wyss L., Waser M., Gebetsberger J., Zywicki M., Polacek N. mRNA-specific translation regulation by a ribosome-associated ncRNA in Haloferax volcanii. Sci. Rep. 2018;8:12502. doi: 10.1038/s41598-018-30332-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Schnoes A.M., Brown S.D., Dodevski I., Babbitt P.C. Annotation error in public databases: Misannotation of molecular function in enzyme superfamilies. PLoS Comput. Biol. 2009;5:e1000605. doi: 10.1371/journal.pcbi.1000605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Promponas V.J., Iliopoulos I., Ouzounis C.A. Annotation inconsistencies beyond sequence similarity-based function prediction—Phylogeny and genome structure. Stand Genomic Sci. 2015;10:108. doi: 10.1186/s40793-015-0101-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Danchin A., Ouzounis C., Tokuyasu T., Zucker J.D. No wisdom in the crowd: Genome annotation in the era of big—Current status and future prospects. Microb. Biotechnol. 2018;11:588–605. doi: 10.1111/1751-7915.13284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Falb M., Muller K., Konigsmaier L., Oberwinkler T., Horn P., von Gronau S., Gonzalez O., Pfeiffer F., Bornberg-Bauer E., Oesterhelt D. Metabolism of halophilic archaea. Extremophiles. 2008;12:177–196. doi: 10.1007/s00792-008-0138-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Kanehisa M., Sato Y., Furumichi M., Morishima K., Tanabe M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 2019;47:D590–D595. doi: 10.1093/nar/gky962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.UniProt C. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021;49:D480–D489. doi: 10.1093/nar/gkaa1100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Pfeiffer F., Losensky G., Marchfelder A., Habermann B., Dyall-Smith M. Whole-genome comparison between the type strain of Halobacterium salinarum (DSM 3754(T) ) and the laboratory strains R1 and NRC-1. Microbiologyopen. 2020;9:e974. doi: 10.1002/mbo3.974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Tittes C., Schwarzer S., Pfeiffer F., Dyall-Smith M., Rodriguez-Franco M., Oksanen H.M., Quax T.E.F. Cellular and genomic properties of Haloferax gibbonsii LR2-5, the host of euryarchaeal virus HFTV1. Front. Microbiol. 2021;12:625599. doi: 10.3389/fmicb.2021.625599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hunter S., Apweiler R., Attwood T.K., Bairoch A., Bateman A., Binns D., Bork P., Das U., Daugherty L., Duquenne L., et al. InterPro: The integrative protein signature database. Nucleic Acids Res. 2009;37:D211–D215. doi: 10.1093/nar/gkn785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Kriventseva E.V., Kuznetsov D., Tegenfeldt F., Manni M., Dias R., Simao F.A., Zdobnov E.M. OrthoDB v10: Sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res. 2019;47:D807–D811. doi: 10.1093/nar/gky1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Oberto J. SyntTax: A web server linking synteny to prokaryotic taxonomy. BMC Bioinformatics. 2013;14:4. doi: 10.1186/1471-2105-14-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Johnson M., Zaretskaya I., Raytselis Y., Merezhuk Y., McGinnis S., Madden T.L. NCBI BLAST: A better web interface. Nucleic Acids Res. 2008;36:W5–W9. doi: 10.1093/nar/gkn201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Altschul S.F., Madden T.L., Schaffer A.A., Zhang J., Zhang Z., Miller W., Lipman D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Rich P.R., Marechal A. The mitochondrial respiratory chain. Essays Biochem. 2010;47:1–23. doi: 10.1042/bse0470001. [DOI] [PubMed] [Google Scholar]
- 77.Guo R., Gu J., Zong S., Wu M., Yang M. Structure and mechanism of mitochondrial electron transport chain. Biomed. J. 2018;41:9–20. doi: 10.1016/j.bj.2017.12.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Crofts A.R., Hong S., Wilson C., Burton R., Victoria D., Harrison C., Schulten K. The mechanism of ubihydroquinone oxidation at the Qo-site of the cytochrome bc1 complex. Biochim. Biophys. Acta. 2013;1827:1362–1377. doi: 10.1016/j.bbabio.2013.01.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Kaila V.R.I., Wikstrom M. Architecture of bacterial respiratory chains. Nat. Rev. Microbiol. 2021;19:319–330. doi: 10.1038/s41579-020-00486-4. [DOI] [PubMed] [Google Scholar]
- 80.Schafer G., Engelhard M., Muller V. Bioenergetics of the Archaea. Microbiol. Mol. Biol. Rev. 1999;63:570–620. doi: 10.1128/MMBR.63.3.570-620.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Scharf B., Wittenberg R., Engelhard M. Electron transfer proteins from the haloalkaliphilic archaeon Natronobacterium pharaonis: Possible components of the respiratory chain include cytochrome bc and a terminal oxidase cytochrome ba3. Biochemistry. 1997;36:4471–4479. doi: 10.1021/bi962312d. [DOI] [PubMed] [Google Scholar]
- 82.Sreeramulu K., Schmidt C.L., Schafer G., Anemuller S. Studies of the electron transport chain of the euryarcheon Halobacterium salinarum: Indications for a type II NADH dehydrogenase and a complex III analog. J. Bioenerg. Biomembr. 1998;30:443–453. doi: 10.1023/A:1020538129400. [DOI] [PubMed] [Google Scholar]
- 83.Gradin C.H., Hederstedt L., Baltscheffsky H. Soluble succinate dehydrogenase from the halophilic archaebacterium, Halobacterium halobium. Arch. Biochem. Biophys. 1985;239:200–205. doi: 10.1016/0003-9861(85)90827-6. [DOI] [PubMed] [Google Scholar]
- 84.Steinert K., Wagner V., Kroth Pancic P.G., Bickel Sandkoetter S. Characterization and subunit structure of the ATP synthase of the halophilic archaeon Haloferax volcanii and organization of the ATP synthase genes. J. Biol. Chem. 1997;272:6261–6269. doi: 10.1074/jbc.272.10.6261. [DOI] [PubMed] [Google Scholar]
- 85.Nanba T., Mukohata Y. A membrane-bound ATPase from Halobacterium halobium: Purification and characterization. J. Biochem. (Tokyo) 1987;102:591–598. doi: 10.1093/oxfordjournals.jbchem.a122092. [DOI] [PubMed] [Google Scholar]
- 86.Kerscher L., Oesterhelt D. Ferredoxin is the coenzyme of α-ketoacid oxidoreductases in Halobacterium halobium. FEBS Lett. 1977;83:197–201. doi: 10.1016/0014-5793(77)81004-1. [DOI] [PubMed] [Google Scholar]
- 87.Plaga W., Lottspeich F., Oesterhelt D. Improved purification, crystallization and primary structure of pyruvate:ferredoxin oxidoreductase from Halobacterium halobium. Eur. J. Biochem. 1992;205:391–397. doi: 10.1111/j.1432-1033.1992.tb16792.x. [DOI] [PubMed] [Google Scholar]
- 88.Kerscher L., Oesterhelt D. Purification and properties of two 2-oxoacid:ferredoxin oxidoreductases from Halobacterium halobium. Eur. J. Biochem. 1981;116:587–594. doi: 10.1111/j.1432-1033.1981.tb05376.x. [DOI] [PubMed] [Google Scholar]
- 89.Kerscher L., Oesterhelt D. The catalytic mechanism of 2-oxoacid:ferredoxin oxidoreductases from Halobacterium halobium. One-electron transfer at two distinct steps of the catalytic cycle. Eur. J. Biochem. 1981;116:595–600. doi: 10.1111/j.1432-1033.1981.tb05377.x. [DOI] [PubMed] [Google Scholar]
- 90.Zafrilla B., Martinez-Espinosa R.M., Bonete M.J., Butt J.N., Richardson D.J., Gates A.J. A haloarchaeal ferredoxin electron donor that plays an essential role in nitrate assimilation. Biochem. Soc. Trans. 2011;39:1844–1848. doi: 10.1042/BST20110709. [DOI] [PubMed] [Google Scholar]
- 91.Falb M., Pfeiffer F., Palm P., Rodewald K., Hickmann V., Tittor J., Oesterhelt D. Living with two extremes: Conclusions from the genome sequence of Natronomonas pharaonis. Genome Res. 2005;15:1336–1343. doi: 10.1101/gr.3952905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Leif H., Sled V.D., Ohnishi T., Weiss H., Friedrich T. Isolation and characterization of the proton-translocating NADH: Ubiquinone oxidoreductase from Escherichia coli. Eur. J. Biochem. 1995;230:538–548. doi: 10.1111/j.1432-1033.1995.tb20594.x. [DOI] [PubMed] [Google Scholar]
- 93.Braun M., Bungert S., Friedrich T. Characterization of the overproduced NADH dehydrogenase fragment of the NADH:ubiquinone oxidoreductase (complex I) from Escherichia coli. Biochemistry. 1998;37:1861–1867. doi: 10.1021/bi971176p. [DOI] [PubMed] [Google Scholar]
- 94.Mattar S. Ph.D. Thesis. Ruhr-Universität Bochum; Bochum, Germany: 1996. Molekularbiologische und Biochemische Charakterisierung zweier Komplexe der Atmungskette von Natronobacterium pharaonis. [Google Scholar]
- 95.Kletzin A., Heimerl T., Flechsler J., van Niftrik L., Rachel R., Klingl A. Cytochromes c in Archaea: Distribution, maturation, cell architecture, and the special case of Ignicoccus hospitalis. Front. Microbiol. 2015;6:439. doi: 10.3389/fmicb.2015.00439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Sreeramulu K. Purification and partial characterization of cytochrome c552 from Halobacterium salinarium. Indian J. Biochem. Biophys. 2003;40:274–277. [PubMed] [Google Scholar]
- 97.Mattar S., Scharf B., Kent S.B., Rodewald K., Oesterhelt D., Engelhard M. The primary structure of halocyanin, an archaeal blue copper protein, predicts a lipid anchor for membrane fixation. J. Biol. Chem. 1994;269:14939–14945. doi: 10.1016/S0021-9258(17)36556-0. [DOI] [PubMed] [Google Scholar]
- 98.Scharf B., Engelhard M. Halocyanin, an archaebacterial blue copper protein (type I) from Natronobacterium pharaonis. Biochemistry. 1993;32:12894–12900. doi: 10.1021/bi00210a043. [DOI] [PubMed] [Google Scholar]
- 99.Hildebrandt P., Matysik J., Schrader B., Scharf B., Engelhard M. Raman spectroscopic study of the blue copper protein halocyanin from Natronobacterium pharaonis. Biochemistry. 1994;33:11426–11431. doi: 10.1021/bi00204a004. [DOI] [PubMed] [Google Scholar]
- 100.Kerscher L., Oesterhelt D. A ferredoxin from halobacteria. FEBS Lett. 1976;67:320–322. doi: 10.1016/0014-5793(76)80556-X. [DOI] [PubMed] [Google Scholar]
- 101.Kerscher L., Oesterhelt D., Cammack R., Hall D.O. A new plant-type ferredoxin from halobacteria. Eur. J. Biochem. 1976;71:101–107. doi: 10.1111/j.1432-1033.1976.tb11094.x. [DOI] [PubMed] [Google Scholar]
- 102.Zhang P., Battchikova N., Paakkarinen V., Katoh H., Iwai M., Ikeuchi M., Pakrasi H.B., Ogawa T., Aro E.M. Isolation, subunit composition and interaction of the NDH-1 complexes from Thermosynechococcus elongatus BP-1. Biochem. J. 2005;390:513–520. doi: 10.1042/BJ20050390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Schuller J.M., Birrell J.A., Tanaka H., Konuma T., Wulfhorst H., Cox N., Schuller S.K., Thiemann J., Lubitz W., Setif P., et al. Structural adaptations of photosynthetic complex I enable ferredoxin-dependent electron transfer. Science. 2019;363:257–260. doi: 10.1126/science.aau3613. [DOI] [PubMed] [Google Scholar]
- 104.Pan X., Cao D., Xie F., Xu F., Su X., Mi H., Zhang X., Li M. Structural basis for electron transport mechanism of complex I-like photosynthetic NAD(P)H dehydrogenase. Nat. Commun. 2020;11:610. doi: 10.1038/s41467-020-14456-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Liu J., Krulwich T.A., Hicks D.B. Purification of two putative type II NADH dehydrogenases with different substrate specificities from alkaliphilic Bacillus pseudofirmus OF4. Biochim. Biophys. Acta. 2008;1777:453–461. doi: 10.1016/j.bbabio.2008.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Lee T.-X., Metzger S.U., Cho Y.S., Whitmarsh J., Kallas T. Modification of inhibitor binding sites in the cytochrome bf complex by directed mutagenesis of cytochrome b6 in Synechococcus sp. PCC 7002. Biochim. Biophys. Acta. 2001;1504:235–247. doi: 10.1016/S0005-2728(00)00253-X. [DOI] [PubMed] [Google Scholar]
- 107.Mattar S., Engelhard M. Cytochrome ba3 from Natronobacterium pharaonis—An archaeal four-subunit cytochrome-c-type oxidase. Eur. J. Biochem. 1997;250:332–341. doi: 10.1111/j.1432-1033.1997.0332a.x. [DOI] [PubMed] [Google Scholar]
- 108.Tanaka M., Ogawa N., Ihara K., Sugiyama Y., Mukohata Y. Cytochrome aa(3) in Haloferax volcanii. J. Bacteriol. 2002;184:840–845. doi: 10.1128/JB.184.3.840-845.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Fujiwara T., Fukumori Y., Yamanaka T. Purification and properties of Halobacterium halobium “cytochrome aa3” which lacks CuA and CuB. J. Biochem. (Tokyo) 1989;105:287–292. doi: 10.1093/oxfordjournals.jbchem.a122655. [DOI] [PubMed] [Google Scholar]
- 110.Denda K., Fujiwara T., Seki M., Yoshida M., Fukumori Y., Yamanaka T. Molecular cloning of the cytochrome aa3 gene from the archaeon (Archaebacterium) Halobacterium halobium. Biochem. Biophys. Res. Commun. 1991;181:316–322. doi: 10.1016/S0006-291X(05)81420-2. [DOI] [PubMed] [Google Scholar]
- 111.Ishikawa R., Ishido Y., Tachikawa A., Kawasaki H., Matsuzawa H., Wakagi T. Aeropyrum pernix K1, a strictly aerobic and hyperthermophilic archaeon, has two terminal oxidases, cytochrome ba3 and cytochrome aa3. Arch. Microbiol. 2002;179:42–49. doi: 10.1007/s00203-002-0496-1. [DOI] [PubMed] [Google Scholar]
- 112.Moshiri F., Chawla A., Maier R.J. Cloning, characterization, and expression in Escherichia coli of the genes encoding the cytochrome d oxidase complex from Azotobacter vinelandii. J. Bacteriol. 1991;173:6230–6241. doi: 10.1128/jb.173.19.6230-6241.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Miller M.J., Gennis R.B. The purification and characterization of the cytochrome d terminal oxidase complex of the Escherichia coli aerobic respiratory chain. J. Biol. Chem. 1983;258:9159–9165. doi: 10.1016/S0021-9258(17)44645-X. [DOI] [PubMed] [Google Scholar]
- 114.Zimmermann B.H., Nitsche C.I., Fee J.A., Rusnak F., Munck E. Properties of a copper-containing cytochrome ba3: A second terminal oxidase from the extreme thermophile Thermus thermophilus. Proc. Natl. Acad. Sci. USA. 1988;85:5779–5783. doi: 10.1073/pnas.85.16.5779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Keightley J.A., Zimmermann B.H., Mather M.W., Springer P., Pastuszyn A., Lawrence D.M., Fee J.A. Molecular genetic and protein chemical characterization of the cytochrome ba3 from Thermus thermophilus HB8. J. Biol. Chem. 1995;270:20345–20358. doi: 10.1074/jbc.270.35.20345. [DOI] [PubMed] [Google Scholar]
- 116.Sisignano M., Morbitzer D., Gatgens J., Oldiges M., Soppa J. A 2-oxoacid dehydrogenase complex of Haloferax volcanii is essential for growth on isoleucine but not on other branched-chain amino acids. Microbiology. 2010;156:521–529. doi: 10.1099/mic.0.033449-0. [DOI] [PubMed] [Google Scholar]
- 117.Jolley K.A., Maddocks D.G., Gyles S.L., Mullan Z., Tang S.L., Dyall-Smith M.L., Hough D.W., Danson M.J. 2-Oxoacid dehydrogenase multienzyme complexes in the halophilic Archaea? Gene sequences and protein structural predictions. Microbiology (Reading) 2000;146(Pt 5):1061–1069. doi: 10.1099/00221287-146-5-1061. [DOI] [PubMed] [Google Scholar]
- 118.Al-Mailem D.M., Hough D.W., Danson M.J. The 2-oxoacid dehydrogenase multienzyme complex of Haloferax volcanii. Extremophiles. 2008;12:89–96. doi: 10.1007/s00792-007-0091-0. [DOI] [PubMed] [Google Scholar]
- 119.van Ooyen J., Soppa J. Three 2-oxoacid dehydrogenase operons in Haloferax volcanii: Expression, deletion mutants and evolution. Microbiology. 2007;153:3303–3313. doi: 10.1099/mic.0.2007/008508-0. [DOI] [PubMed] [Google Scholar]
- 120.Wanner C., Soppa J. Functional role for a 2-oxo acid dehydrogenase in the halophilic archaeon Haloferax volcanii. J. Bacteriol. 2002;184:3114–3121. doi: 10.1128/JB.184.11.3114-3121.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Heath C., Posner M.G., Aass H.C., Upadhyay A., Scott D.J., Hough D.W., Danson M.J. The 2-oxoacid dehydrogenase multi-enzyme complex of the archaeon Thermoplasma acidophilum—Recombinant expression, assembly and characterization. FEBS J. 2007;274:5406–5415. doi: 10.1111/j.1742-4658.2007.06067.x. [DOI] [PubMed] [Google Scholar]
- 122.Oppermann F.B., Schmidt B., Steinbuchel A. Purification and characterization of acetoin:2,6-dichlorophenolindophenol oxidoreductase, dihydrolipoamide dehydrogenase, and dihydrolipoamide acetyltransferase of the Pelobacter carbinolicus acetoin dehydrogenase enzyme system. J. Bacteriol. 1991;173:757–767. doi: 10.1128/jb.173.2.757-767.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Huang M., Oppermann-Sanio F.B., Steinbuchel A. Biochemical and molecular characterization of the Bacillus subtilis acetoin catabolic pathway. J. Bacteriol. 1999;181:3837–3841. doi: 10.1128/JB.181.12.3837-3841.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Horie A., Tomita T., Saiki A., Kono H., Taka H., Mineki R., Fujimura T., Nishiyama C., Kuzuyama T., Nishiyama M. Discovery of proteinaceous N-modification in lysine biosynthesis of Thermus thermophilus. Nat. Chem. Biol. 2009;5:673–679. doi: 10.1038/nchembio.198. [DOI] [PubMed] [Google Scholar]
- 125.Yoshida A., Tomita T., Atomi H., Kuzuyama T., Nishiyama M. Lysine biosynthesis of Thermococcus kodakarensis with the capacity to function as an ornithine biosynthetic system. J. Biol. Chem. 2016;291:21630–21643. doi: 10.1074/jbc.M116.743021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Ouchi T., Tomita T., Horie A., Yoshida A., Takahashi K., Nishida H., Lassak K., Taka H., Mineki R., Fujimura T., et al. Lysine and arginine biosyntheses mediated by a common carrier protein in Sulfolobus. Nat. Chem. Biol. 2013;9:277–283. doi: 10.1038/nchembio.1200. [DOI] [PubMed] [Google Scholar]
- 127.Hochuli M., Patzelt H., Oesterhelt D., Wuthrich K., Szyperski T. Amino acid biosynthesis in the halophilic archaeon Haloarcula hispanica. J. Bacteriol. 1999;181:3226–3237. doi: 10.1128/JB.181.10.3226-3237.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Yoshida A., Tomita T., Fujimura T., Nishiyama C., Kuzuyama T., Nishiyama M. Structural insight into amino group-carrier protein-mediated lysine biosynthesis: Crystal structure of the LysZ.LysW complex from Thermus thermophilus. J. Biol. Chem. 2015;290:435–447. doi: 10.1074/jbc.M114.595983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Shimizu T., Tomita T., Kuzuyama T., Nishiyama M. Crystal structure of the LysY.LysW complex from Thermus thermophilus. J. Biol. Chem. 2016;291:9948–9959. doi: 10.1074/jbc.M115.707034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Miyazaki J., Kobashi N., Nishiyama M., Yamane H. Functional and evolutionary relationship between arginine biosynthesis and prokaryotic lysine biosynthesis through alpha-aminoadipate. J. Bacteriol. 2001;183:5067–5073. doi: 10.1128/JB.183.17.5067-5073.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Fujita S., Cho S.H., Yoshida A., Hasebe F., Tomita T., Kuzuyama T., Nishiyama M. Crystal structure of LysK, an enzyme catalyzing the last step of lysine biosynthesis in Thermus thermophilus, in complex with lysine: Insight into the mechanism for recognition of the amino-group carrier protein, LysW. Biochem. Biophys. Res. Commun. 2017;491:409–415. doi: 10.1016/j.bbrc.2017.07.088. [DOI] [PubMed] [Google Scholar]
- 132.Issaly I.M., Issaly A.S. Control of ornithine carbamoyltransferase activityby arginase in Bacillus subtilis. Eur. J. Biochem. 1974;49:485–495. doi: 10.1111/j.1432-1033.1974.tb03853.x. [DOI] [PubMed] [Google Scholar]
- 133.Ruepp A., Muller H.N., Lottspeich F., Soppa J. Catabolic ornithine transcarbamylase of Halobacterium halobium (salinarium)—Purification, characterization, sequence determination, and evolution. J. Bacteriol. 1995;177:1129–1136. doi: 10.1128/jb.177.5.1129-1136.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Shaheen N., Kobayashi K., Terazono H., Fukushige T., Horiuchi M., Saheki T. Characterization of human wild-type and mutant argininosuccinate synthetase proteins expressed in bacterial cells. Enzyme Protein. 1994;48:251–264. doi: 10.1159/000474998. [DOI] [PubMed] [Google Scholar]
- 135.Lemke C., Yeung M., Howell P.L. Expression, purification, crystallization and preliminary X-ray analysis of Escherichia coli argininosuccinate synthetase. Acta Crystallogr. D Biol. Crystallogr. 1999;55:2028–2030. doi: 10.1107/S0907444999011816. [DOI] [PubMed] [Google Scholar]
- 136.Cohen-Kupiec R., Kupiec M., Sandbeck K., Leigh J.A. Functional conservation between the argininosuccinate lyase of the archaeon Methanococcus maripaludis and the corresponding bacterial and eukaryal genes. FEMS Microbiol. Lett. 1999;173:231–238. doi: 10.1111/j.1574-6968.1999.tb13507.x. [DOI] [PubMed] [Google Scholar]
- 137.Kato C., Kurihara T., Kobashi N., Yamane H., Nishiyama M. Conversion of feedback regulation in aspartate kinase by domain exchange. Biochem. Biophys. Res. Commun. 2004;316:802–808. doi: 10.1016/j.bbrc.2004.02.122. [DOI] [PubMed] [Google Scholar]
- 138.Faehnle C.R., Ohren J.F., Viola R.E. A new branch in the family: Structure of aspartate-beta-semialdehyde dehydrogenase from Methanococcus jannaschii. J. Mol. Biol. 2005;353:1055–1068. doi: 10.1016/j.jmb.2005.09.027. [DOI] [PubMed] [Google Scholar]
- 139.Kaur N., Gautam A., Kumar S., Singh A., Singh N., Sharma S., Sharma R., Tewari R., Singh T.P. Biochemical studies and crystal structure determination of dihydrodipicolinate synthase from Pseudomonas aeruginosa. Int. J. Biol. Macromol. 2011;48:779–787. doi: 10.1016/j.ijbiomac.2011.03.002. [DOI] [PubMed] [Google Scholar]
- 140.Reddy S.G., Sacchettini J.C., Blanchard J.S. Expression, purification, and characterization of Escherichia coli dihydrodipicolinate reductase. Biochemistry. 1995;34:3492–3501. doi: 10.1021/bi00011a002. [DOI] [PubMed] [Google Scholar]
- 141.Simms S.A., Voige W.H., Gilvarg C. Purification and characterization of succinyl-CoA: Tetrahydrodipicolinate N-succinyltransferase from Escherichia coli. J. Biol. Chem. 1984;259:2734–2741. doi: 10.1016/S0021-9258(17)43207-8. [DOI] [PubMed] [Google Scholar]
- 142.Lin Y.K., Myhrman R., Schrag M.L., Gelb M.H. Bacterial N-succinyl-L-diaminopimelic acid desuccinylase. Purification, partial characterization, and substrate specificity. J. Biol. Chem. 1988;263:1622–1627. doi: 10.1016/S0021-9258(19)77924-1. [DOI] [PubMed] [Google Scholar]
- 143.Wiseman J.S., Nichols J.S. Purification and properties of diaminopimelic acid epimerase from Escherichia coli. J. Biol. Chem. 1984;259:8907–8914. doi: 10.1016/S0021-9258(17)47241-3. [DOI] [PubMed] [Google Scholar]
- 144.White P.J., Kelly B. Purification and properties of diaminopimelate decarboxylase from Escherichia coli. Biochem. J. 1965;96:75–84. doi: 10.1042/bj0960075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Gulko M.K., Dyall-Smith M., Gonzalez O., Oesterhelt D. How do haloarchaea synthesize aromatic amino acids? PLoS ONE. 2014;9:e107475. doi: 10.1371/journal.pone.0107475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.White R.H. L-Aspartate semialdehyde and a 6-deoxy-5-ketohexose 1-phosphate are the precursors to the aromatic amino acids in Methanocaldococcus jannaschii. Biochemistry. 2004;43:7618–7627. doi: 10.1021/bi0495127. [DOI] [PubMed] [Google Scholar]
- 147.Porat I., Waters B.W., Teng Q., Whitman W.B. Two biosynthetic pathways for aromatic amino acids in the archaeon Methanococcus maripaludis. J. Bacteriol. 2004;186:4940–4950. doi: 10.1128/JB.186.15.4940-4950.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Phillips R.S., Gollnick P.D. Evidence that cysteine 298 Is in the active site of tryptophan indole-lyase. J. Biol. Chem. 1989;264:10627–10632. doi: 10.1016/S0021-9258(18)81667-2. [DOI] [PubMed] [Google Scholar]
- 149.Newton W.A., Morino Y., Snell E.E. Properties of crystalline tryptophanase. J. Biol. Chem. 1965;240:1211–1218. doi: 10.1016/S0021-9258(18)97562-9. [DOI] [PubMed] [Google Scholar]
- 150.Oda M., Sugishita A., Furukawa K. Cloning and nucleotide sequences of histidase and regulatory genes in the Bacillus subtilis hut operon and positive regulation of the operon. J. Bacteriol. 1988;170:3199–3205. doi: 10.1128/jb.170.7.3199-3205.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Hartwell L.H., Magasanik B. The molecular basis of histidase induction in Bacillus subtilis. J. Mol. Biol. 1963;7:401–420. doi: 10.1016/S0022-2836(63)80033-9. [DOI] [PubMed] [Google Scholar]
- 152.Kaminskas E., Kimhi Y., Magasanik B. Urocanase and N-formimino-L-glutamate formiminohydrolase of Bacillus subtilis, two enzymes of the histidine degradation pathway. J. Biol. Chem. 1970;245:3536–3544. doi: 10.1016/S0021-9258(18)62959-X. [DOI] [PubMed] [Google Scholar]
- 153.Yu Y., Liang Y.H., Brostromer E., Quan J.M., Panjikar S., Dong Y.H., Su X.D. A catalytic mechanism revealed by the crystal structures of the imidazolonepropionase from Bacillus subtilis. J. Biol. Chem. 2006;281:36929–36936. doi: 10.1074/jbc.M607703200. [DOI] [PubMed] [Google Scholar]
- 154.Howell D.M., Xu H., White R.H. (R)-citramalate synthase in methanogenic archaea. J. Bacteriol. 1999;181:331–333. doi: 10.1128/JB.181.1.331-333.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Howell D.M., Harich K., Xu H.M., White R.H. Alpha-keto acid chain elongation reactions involved in the biosynthesis of coenzyme b (7-mercaptoheptanoyl threonine phosphate) in methanogenic archaea. Biochemistry. 1998;37:10108–10117. doi: 10.1021/bi980662p. [DOI] [PubMed] [Google Scholar]
- 156.Porat I., Sieprawska-Lupa M., Teng Q., Bohanon F.J., White R.H., Whitman W.B. Biochemical and genetic characterization of an early step in a novel pathway for the biosynthesis of aromatic amino acids and p-aminobenzoic acid in the archaeon Methanococcus maripaludis. Mol. Microbiol. 2006;62:1117–1131. doi: 10.1111/j.1365-2958.2006.05426.x. [DOI] [PubMed] [Google Scholar]
- 157.Large A., Stamme C., Lange C., Duan Z., Allers T., Soppa J., Lund P.A. Characterization of a tightly controlled promoter of the halophilic archaeon Haloferax volcanii and its use in the analysis of the essential cct1 gene. Mol. Microbiol. 2007;66:1092–1106. doi: 10.1111/j.1365-2958.2007.05980.x. [DOI] [PubMed] [Google Scholar]
- 158.Bali S., Lawrence A.D., Lobo S.A., Saraiva L.M., Golding B.T., Palmer D.J., Howard M.J., Ferguson S.J., Warren M.J. Molecular hijacking of siroheme for the synthesis of heme and d1 heme. Proc. Natl. Acad. Sci. USA. 2011;108:18260–18265. doi: 10.1073/pnas.1108228108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Siddaramappa S., Challacombe J.F., Decastro R.E., Pfeiffer F., Sastre D.E., Gimenez M.I., Paggi R.A., Detter J.C., Davenport K.W., Goodwin L.A., et al. A comparative genomics perspective on the genetic content of the alkaliphilic haloarchaeon Natrialba magadii ATCC 43099T. BMC Genomics. 2012;13:165. doi: 10.1186/1471-2164-13-165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Moore S.J., Lawrence A.D., Biedendieck R., Deery E., Frank S., Howard M.J., Rigby S.E., Warren M.J. Elucidation of the anaerobic pathway for the corrin component of cobalamin (vitamin B12) Proc. Natl. Acad. Sci. USA. 2013;110:14906–14911. doi: 10.1073/pnas.1308098110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Rodionov D.A., Vitreschak A.G., Mironov A.A., Gelfand M.S. Comparative genomics of the vitamin B12 metabolism and regulation in prokaryotes. J. Biol. Chem. 2003;278:41148–41159. doi: 10.1074/jbc.M305837200. [DOI] [PubMed] [Google Scholar]
- 162.Kosugi N., Araki T., Fujita J., Tanaka S., Fujiwara T. Growth phenotype analysis of heme synthetic enzymes in a halophilic archaeon, Haloferax volcanii. PLoS ONE. 2017;12:e0189913. doi: 10.1371/journal.pone.0189913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Raux E., Leech H.K., Beck R., Schubert H.L., Santander P.J., Roessner C.A., Scott A.I., Martens J.H., Jahn D., Thermes C., et al. Identification and functional analysis of enzymes required for precorrin-2 dehydrogenation and metal ion insertion in the biosynthesis of sirohaem and cobalamin in Bacillus megaterium. Biochem. J. 2003;370:505–516. doi: 10.1042/bj20021443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Brindley A.A., Raux E., Leech H.K., Schubert H.L., Warren M.J. A story of chelatase evolution: Identification and characterization of a small 13-15-kDa "ancestral" cobaltochelatase (CbiXS) in the archaea. J. Biol. Chem. 2003;278:22388–22395. doi: 10.1074/jbc.M302468200. [DOI] [PubMed] [Google Scholar]
- 165.Yin J., Xu L.X., Cherney M.M., Raux-Deery E., Bindley A.A., Savchenko A., Walker J.R., Cuff M.E., Warren M.J., James M.N. Crystal structure of the vitamin B12 biosynthetic cobaltochelatase, CbiXS, from Archaeoglobus fulgidus. J. Struct. Funct. Genomics. 2006;7:37–50. doi: 10.1007/s10969-006-9008-x. [DOI] [PubMed] [Google Scholar]
- 166.Storbeck S., Rolfes S., Raux-Deery E., Warren M.J., Jahn D., Layer G. A novel pathway for the biosynthesis of heme in Archaea: Genome-based bioinformatic predictions and experimental evidence. Archaea. 2010;2010:175050. doi: 10.1155/2010/175050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Stroupe M.E., Leech H.K., Daniels D.S., Warren M.J., Getzoff E.D. CysG structure reveals tetrapyrrole-binding features and novel regulation of siroheme biosynthesis. Nat. Struct. Biol. 2003;10:1064–1073. doi: 10.1038/nsb1007. [DOI] [PubMed] [Google Scholar]
- 168.Pennington J.M., Kemp M., McGarry L., Chen Y., Stroupe M.E. Siroheme synthase orients substrates for dehydrogenase and chelatase activities in a common active site. Nat. Commun. 2020;11:864. doi: 10.1038/s41467-020-14722-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Schubert H.L., Rose R.S., Leech H.K., Brindley A.A., Hill C.P., Rigby S.E., Warren M.J. Structure and function of SirC from Bacillus megaterium: A metal-binding precorrin-2 dehydrogenase. Biochem. J. 2008;415:257–263. doi: 10.1042/BJ20080785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Roessner C.A., Warren M.J., Santander P.J., Atshaves B.P., Ozaki S.-i., Stolowich N.J., Iida K., Scott A.I. Expression of 9 Salmonella typhimurium enzymes for cobinamide synthesis. Identification of the 11-methyl and 20-methyl transferases of corrin biosynthesis. FEBS Lett. 1992;301:73–78. doi: 10.1016/0014-5793(92)80213-Z. [DOI] [PubMed] [Google Scholar]
- 171.Santander P.J., Roessner C.A., Stolowich N.J., Holderman M.T., Scott A.I. How corrinoids are synthesized without oxygen: Nature’s first pathway to vitamin B12. Chem. Biol. 1997;4:659–666. doi: 10.1016/S1074-5521(97)90221-0. [DOI] [PubMed] [Google Scholar]
- 172.Santander P.J., Kajiwara Y., Williams H.J., Scott A.I. Structural characterization of novel cobalt corrinoids synthesized by enzymes of the vitamin B12 anaerobic pathway. Bioorg. Med. Chem. 2006;14:724–731. doi: 10.1016/j.bmc.2005.08.062. [DOI] [PubMed] [Google Scholar]
- 173.Kajiwara Y., Santander P.J., Roessner C.A., Perez L.M., Scott A.I. Genetically engineered synthesis and structural characterization of cobalt-precorrin 5A and -5B, two new intermediates on the anaerobic pathway to vitamin B12: Definition of the roles of the CbiF and CbiG enzymes. J. Am. Chem. Soc. 2006;128:9971–9978. doi: 10.1021/ja062940a. [DOI] [PubMed] [Google Scholar]
- 174.Fresquet V., Williams L., Raushel F.M. Mechanism of cobyrinic acid a,c-diamide synthetase from Salmonella typhimurium LT2. Biochemistry. 2004;43:10619–10627. doi: 10.1021/bi048972x. [DOI] [PubMed] [Google Scholar]
- 175.Buan N.R., Rehfeld K., Escalante-Semerena J.C. Studies of the CobA-type ATP:Co(I)rrinoid adenosyltransferase enzyme of Methanosarcina mazei strain Go1. J. Bacteriol. 2006;188:3543–3550. doi: 10.1128/JB.188.10.3543-3550.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Fonseca M.V., Buan N.R., Horswill A.R., Rayment I., Escalante-Semerena J.C. The ATP:Co(I)rrinoid adenosyltransferase (CobA) enzyme of Salmonella enterica requires the 2’-OH group of ATP for function and yields inorganic triphosphate as its reaction byproduct. J. Biol. Chem. 2002;277:33127–33131. doi: 10.1074/jbc.M203893200. [DOI] [PubMed] [Google Scholar]
- 177.Johnson C.L., Pechonick E., Park S.D., Havemann G.D., Leal N.A., Bobik T.A. Functional genomic, biochemical, and genetic characterization of the Salmonella pduO gene, an ATP:cob(I)alamin adenosyltransferase gene. J. Bacteriol. 2001;183:1577–1584. doi: 10.1128/JB.183.5.1577-1584.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Woodson J.D., Zayas C.L., Escalante-Semerena J.C. A new pathway for salvaging the coenzyme B12 precursor cobinamide in archaea requires cobinamide-phosphate synthase (CbiB) enzyme activity. J. Bacteriol. 2003;185:7193–7201. doi: 10.1128/JB.185.24.7193-7201.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Woodson J.D., Escalante-Semerena J.C. CbiZ, an amidohydrolase enzyme required for salvaging the coenzyme B12 precursor cobinamide in archaea. Proc. Natl. Acad. Sci. USA. 2004;101:3591–3596. doi: 10.1073/pnas.0305939101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Woodson J.D., Peck R.F., Krebs M.P., Escalante-Semerena J.C. The cobY gene of the archaeon Halobacterium sp. strain NRC-1 is required for de novo cobamide synthesis. J. Bacteriol. 2003;185:311–316. doi: 10.1128/JB.185.1.311-316.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Zayas C.L., Escalante-Semerena J.C. Reassessment of the late steps of coenzyme B12 synthesis in Salmonella enterica: Evidence that dephosphorylation of adenosylcobalamin-5’-phosphate by the CobC phosphatase is the last step of the pathway. J. Bacteriol. 2007;189:2210–2218. doi: 10.1128/JB.01665-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.O’Toole G.A., Trzebiatowski J.R., Escalante-Semerena J.C. The cobC gene of Salmonella typhimurium codes for a novel phosphatase involved in the assembly of the nucleotide loop of cobalamin. J. Biol. Chem. 1994;269:26503–26511. doi: 10.1016/S0021-9258(18)47223-7. [DOI] [PubMed] [Google Scholar]
- 183.Brushaber K.R., O’Toole G.A., Escalante-Semerena J.C. CobD, a novel enzyme with L-threonine-O-3-phosphate decarboxylase activity, is responsible for the synthesis of (R)-1-amino-2-propanol O-2-phosphate, a proposed new intermediate in cobalamin biosynthesis in Salmonella typhimurium LT2. J. Biol. Chem. 1998;273:2684–2691. doi: 10.1074/jbc.273.5.2684. [DOI] [PubMed] [Google Scholar]
- 184.Trzebiatowski J.R., O’Toole G.A., Escalante-Semerena J.C. The cobT gene of Salmonella typhimurium encodes the NaMN: 5,6-dimethylbenzimidazole phosphoribosyltransferase responsible for the synthesis of N1-(5-phospho-alpha-D-ribosyl)-5,6-dimethylbenzimidazole, an intermediate in the synthesis of the nucleotide loop of cobalamin. J. Bacteriol. 1994;176:3568–3575. doi: 10.1128/jb.176.12.3568-3575.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Debussche L., Couder M., Thibaut D., Cameron B., Crouzet J., Blanche F. Assay, purification, and characterization of cobaltochelatase, a unique complex enzyme catalyzing cobalt insertion in hydrogenobyrinic acid a,c-diamide during coenzyme B12 biosynthesis in Pseudomonas denitrificans. J. Bacteriol. 1992;174:7445–7451. doi: 10.1128/jb.174.22.7445-7451.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Jensen P.E., Gibson L.C., Henningsen K.W., Hunter C.N. Expression of the chlI, chlD, and chlH genes from the Cyanobacterium Synechocystis PCC6803 in Escherichia coli and demonstration that the three cognate proteins are required for magnesium-protoporphyrin chelatase activity. J. Biol. Chem. 1996;271:16662–16667. doi: 10.1074/jbc.271.28.16662. [DOI] [PubMed] [Google Scholar]
- 187.Jensen P.E., Gibson L.C., Hunter C.N. Determinants of catalytic activity with the use of purified I, D and H subunits of the magnesium protoporphyrin IX chelatase from Synechocystis PCC6803. Biochem. J. 1998;334(Pt 2):335–344. doi: 10.1042/bj3340335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Kuhner M., Haufschildt K., Neumann A., Storbeck S., Streif J., Layer G. The alternative route to heme in the methanogenic archaeon Methanosarcina barkeri. Archaea. 2014;2014:327637. doi: 10.1155/2014/327637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Dailey H.A., Dailey T.A., Gerdes S., Jahn D., Jahn M., O’Brian M.R., Warren M.J. Prokaryotic heme biosynthesis: Multiple pathways to a common essential product. Microbiol. Mol. Biol. Rev. 2017;81:e00048-16. doi: 10.1128/MMBR.00048-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Eirich L.D., Vogels G.D., Wolfe R.S. Distribution of coenzyme F420 and properties of its hydrolytic fragments. J. Bacteriol. 1979;140:20–27. doi: 10.1128/jb.140.1.20-27.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Jaenchen R., Schonheit P., Thauer R.K. Studies on the biosynthesis of coenzyme F420 in methanogenic bacteria. Arch. Microbiol. 1984;137:362–365. doi: 10.1007/BF00410735. [DOI] [PubMed] [Google Scholar]
- 192.Lin X.L., White R.H. Occurrence of coenzyme F420 and its gamma-monoglutamyl derivative in nonmethanogenic archaebacteria. J. Bacteriol. 1986;168:444–448. doi: 10.1128/jb.168.1.444-448.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.de Wit L.E.A., Eker A.P.M. 8-Hydroxy-5-deazaflavin-dependent electron transfer in the extreme halophile Halobacterium cutirubrum. FEMS Microbiol. Lett. 1987;48:121–125. doi: 10.1111/j.1574-6968.1987.tb02527.x. [DOI] [Google Scholar]
- 194.Grochowski L.L., Xu H., White R.H. Identification and characterization of the 2-phospho-L-lactate guanylyltransferase involved in coenzyme F420 biosynthesis. Biochemistry. 2008;47:3033–3037. doi: 10.1021/bi702475t. [DOI] [PubMed] [Google Scholar]
- 195.Bashiri G., Antoney J., Jirgis E.N.M., Shah M.V., Ney B., Copp J., Stuteley S.M., Sreebhavan S., Palmer B., Middleditch M., et al. A revised biosynthetic pathway for the cofactor F420 in prokaryotes. Nat. Commun. 2019;10:1558. doi: 10.1038/s41467-019-09534-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Braga D., Last D., Hasan M., Guo H., Leichnitz D., Uzum Z., Richter I., Schalk F., Beemelmanns C., Hertweck C., et al. Metabolic pathway rerouting in Paraburkholderia rhizoxinica evolved long-overlooked derivatives of coenzyme F420. ACS Chem. Biol. 2019;14:2088–2094. doi: 10.1021/acschembio.9b00605. [DOI] [PubMed] [Google Scholar]
- 197.Selengut J.D., Haft D.H. Unexpected abundance of coenzyme F(420)-dependent enzymes in Mycobacterium tuberculosis and other actinobacteria. J. Bacteriol. 2010;192:5788–5798. doi: 10.1128/JB.00425-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Graham D.E., Xu H., White R.H. Identification of the 7,8-didemethyl-8-hydroxy-5-deazariboflavin synthase required for coenzyme F(420) biosynthesis. Arch. Microbiol. 2003;180:455–464. doi: 10.1007/s00203-003-0614-8. [DOI] [PubMed] [Google Scholar]
- 199.Philmus B., Decamps L., Berteau O., Begley T.P. Biosynthetic versatility and coordinated action of 5’-deoxyadenosyl radicals in deazaflavin biosynthesis. J. Am. Chem. Soc. 2015;137:5406–5413. doi: 10.1021/ja513287k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Decamps L., Philmus B., Benjdia A., White R., Begley T.P., Berteau O. Biosynthesis of F0, precursor of the F420 cofactor, requires a unique two radical-SAM domain enzyme and tyrosine as substrate. J. Am. Chem. Soc. 2012;134:18173–18176. doi: 10.1021/ja307762b. [DOI] [PubMed] [Google Scholar]
- 201.Forouhar F., Abashidze M., Xu H., Grochowski L.L., Seetharaman J., Hussain M., Kuzin A., Chen Y., Zhou W., Xiao R., et al. Molecular insights into the biosynthesis of the F420 coenzyme. J. Biol. Chem. 2008;283:11832–11840. doi: 10.1074/jbc.M710352200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Graupner M., Xu H., White R.H. Characterization of the 2-phospho-L-lactate transferase enzyme involved in coenzyme F(420) biosynthesis in Methanococcus jannaschii. Biochemistry. 2002;41:3754–3761. doi: 10.1021/bi011937v. [DOI] [PubMed] [Google Scholar]
- 203.Nocek B., Evdokimova E., Proudfoot M., Kudritska M., Grochowski L.L., White R.H., Savchenko A., Yakunin A.F., Edwards A., Joachimiak A. Structure of an amide bond forming F(420):gamma-glutamyl ligase from Archaeoglobus fulgidus—A member of a new family of non-ribosomal peptide synthases. J. Mol. Biol. 2007;372:456–469. doi: 10.1016/j.jmb.2007.06.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Li H., Graupner M., Xu H., White R.H. CofE catalyzes the addition of two glutamates to F420-0 in F420 coenzyme biosynthesis in Methanococcus jannaschii. Biochemistry. 2003;42:9771–9778. doi: 10.1021/bi034779b. [DOI] [PubMed] [Google Scholar]
- 205.Kunow J., Schwoërer B., Stetter K.O., Thauer R.K. A F420-dependent NADP reductase in the extremely thermophilic sulfate-reducing Archaeoglobus fulgidus. Arch. Microbiol. 1993;160:199–205. [Google Scholar]
- 206.Purwantini E., Mukhopadhyay B. Rv0132c of Mycobacterium tuberculosis encodes a coenzyme F420-dependent hydroxymycolic acid dehydrogenase. PLoS ONE. 2013;8:e81985. doi: 10.1371/journal.pone.0081985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Klein A.R., Berk H., Purwantini E., Daniels L., Thauer R.K. Si-face stereospecificity at C5 of coenzyme F420 for F420-dependent glucose-6-phosphate dehydrogenase from Mycobacterium smegmatis and F420-dependent alcohol dehydrogenase from Methanoculleus thermophilicus. Eur. J. Biochem. 1996;239:93–97. doi: 10.1111/j.1432-1033.1996.0093u.x. [DOI] [PubMed] [Google Scholar]
- 208.Aufhammer S.W., Warkentin E., Berk H., Shima S., Thauer R.K., Ermler U. Coenzyme binding in F420-dependent secondary alcohol dehydrogenase, a member of the bacterial luciferase family. Structure. 2004;12:361–370. doi: 10.1016/j.str.2004.02.010. [DOI] [PubMed] [Google Scholar]
- 209.Haase P., Deppenmeier U., Blaut M., Gottschalk G. Purification and characterization of F420H2-dehydrogenase from Methanolobus tindarius. Eur. J. Biochem. 1992;203:527–531. doi: 10.1111/j.1432-1033.1992.tb16579.x. [DOI] [PubMed] [Google Scholar]
- 210.Westenberg D.J., Braune A., Ruppert C., Muller V., Herzberg C., Gottschalk G., Blaut M. The F420H2-dehydrogenase from Methanolobus tindarius: Cloning of the ffd operon and expression of the genes in Escherichia coli. FEMS Microbiol. Lett. 1999;170:389–398. doi: 10.1111/j.1574-6968.1999.tb13399.x. [DOI] [PubMed] [Google Scholar]
- 211.Johnson E.F., Mukhopadhyay B. A new type of sulfite reductase, a novel coenzyme F420-dependent enzyme, from the methanarchaeon Methanocaldococcus jannaschii. J. Biol. Chem. 2005;280:38776–38786. doi: 10.1074/jbc.M503492200. [DOI] [PubMed] [Google Scholar]
- 212.te Brömmelstroet B.W., Hensgens C.M., Keltjens J.T., van der Drift C., Vogels G.D. Purification and properties of 5,10-methylenetetrahydromethanopterin reductase, a coenzyme F420-dependent enzyme, from Methanobacterium thermoautotrophicum strain delta H. J. Biol. Chem. 1990;265:1852–1857. doi: 10.1016/S0021-9258(19)39907-7. [DOI] [PubMed] [Google Scholar]
- 213.Vaupel M., Thauer R.K. Coenzyme F420-dependent N5,N10-methylenetetrahydromethanopterin reductase (Mer) from Methanobacterium thermoautotrophicum strain Marburg. Cloning, sequencing, transcriptional analysis, and functional expression in Escherichia coli of the mer gene. Eur. J. Biochem. 1995;231:773–778. doi: 10.1111/j.1432-1033.1995.0773d.x. [DOI] [PubMed] [Google Scholar]
- 214.Shima S., Warkentin E., Grabarse W., Sordel M., Wicke M., Thauer R.K., Ermler U. Structure of coenzyme F(420) dependent methylenetetrahydromethanopterin reductase from two methanogenic archaea. J. Mol. Biol. 2000;300:935–950. doi: 10.1006/jmbi.2000.3909. [DOI] [PubMed] [Google Scholar]
- 215.Takao M., Kobayashi T., Oikawa A., Yasui A. Tandem arrangement of photolyase and superoxide dismutase genes in Halobacterium halobium. J. Bacteriol. 1989;171:6323–6329. doi: 10.1128/jb.171.11.6323-6329.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.McCready S., Marcello L. Repair of UV damage in Halobacterium salinarum. Biochem. Soc. Trans. 2003;31:694–698. doi: 10.1042/bst0310694. [DOI] [PubMed] [Google Scholar]
- 217.Brudler R., Hitomi K., Daiyasu H., Toh H., Kucho K.-i., Ishiura M., Kanehisa M., Roberts V.A., Todo T., Tainer J.A., et al. Identification of a new cryptochrome class. Mol. Cell. 2003;11:59–67. doi: 10.1016/S1097-2765(03)00008-X. [DOI] [PubMed] [Google Scholar]
- 218.Kleine T., Lockhart P., Batschauer A. An Arabidopsis protein closely related to Synechocystis cryptochrome is targeted to organelles. Plant J. 2003;35:93–103. doi: 10.1046/j.1365-313X.2003.01787.x. [DOI] [PubMed] [Google Scholar]
- 219.Selby C.P., Sancar A. A cryptochrome/photolyase class of enzymes with single-stranded DNA-specific photolyase activity. Proc. Natl. Acad. Sci. USA. 2006;103:17696–17700. doi: 10.1073/pnas.0607993103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Zhang F., Scheerer P., Oberpichler I., Lamparter T., Krauss N. Crystal structure of a prokaryotic (6-4) photolyase with an Fe-S cluster and a 6,7-dimethyl-8-ribityllumazine antenna chromophore. Proc. Natl. Acad. Sci. USA. 2013;110:7217–7222. doi: 10.1073/pnas.1302377110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.White R.H. Analysis and characterization of the folates in the nonmethanogenic archaebacteria. J. Bacteriol. 1988;170:4608–4612. doi: 10.1128/jb.170.10.4608-4612.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Maden B.E. Tetrahydrofolate and tetrahydromethanopterin compared: Functionally distinct carriers in C1 metabolism. Biochem. J. 2000;350(Pt 3):609–629. doi: 10.1042/bj3500609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.de Crecy-Lagard V. Variations in metabolic pathways create challenges for automated metabolic reconstructions: Examples from the tetrahydrofolate synthesis pathway. Comput. Struct. Biotechnol. J. 2014;10:41–50. doi: 10.1016/j.csbj.2014.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.Pfeiffer F., Schuster S.C., Broicher A., Falb M., Palm P., Rodewald K., Ruepp A., Soppa J., Tittor J., Oesterhelt D. Evolution in the laboratory: The genome of Halobacterium salinarum strain R1 compared to that of strain NRC-1. Genomics. 2008;91:335–346. doi: 10.1016/j.ygeno.2008.01.001. [DOI] [PubMed] [Google Scholar]
- 225.Sato S., Nakada Y., Kanaya S., Tanaka T. Molecular cloning and nucleotide sequence of Thermus thermophilus HB8 trpE and trpG. Biochim. Biophys. Acta. 1988;950:303–312. doi: 10.1016/0167-4781(88)90126-1. [DOI] [PubMed] [Google Scholar]
- 226.Slock J., Stahly D.P., Han C.Y., Six E.W., Crawford I.P. An apparent Bacillus subtilis folic acid biosynthetic operon containing pab, an amphibolic trpG gene, a third gene required for synthesis of para-aminobenzoic acid, and the dihydropteroate synthase gene. J. Bacteriol. 1990;172:7211–7226. doi: 10.1128/jb.172.12.7211-7226.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Schadt H.S., Schadt S., Oldach F., Sussmuth R.D. 2-Amino-2-deoxyisochorismate is a key intermediate in Bacillus subtilis p-aminobenzoic acid biosynthesis. J. Am. Chem. Soc. 2009;131:3481–3483. doi: 10.1021/ja809283u. [DOI] [PubMed] [Google Scholar]
- 228.Isupov M.N., Boyko K.M., Sutter J.M., James P., Sayer C., Schmidt M., Schonheit P., Nikolaeva A.Y., Stekhanova T.N., Mardanov A.V., et al. Thermostable branched-chain amino acid transaminases from the archaea Geoglobus acetivorans and Archaeoglobus fulgidus: Biochemical and structural characterization. Front. Bioeng. Biotechnol. 2019;7:7. doi: 10.3389/fbioe.2019.00007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.El Yacoubi B., Phillips G., Blaby I.K., Haas C.E., Cruz Y., Greenberg J., de Crecy-Lagard V. A Gateway platform for functional genomics in Haloferax volcanii: Deletion of three tRNA modification genes. Archaea. 2009;2:211–219. doi: 10.1155/2009/428489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Grochowski L.L., Xu H., Leung K., White R.H. Characterization of an Fe(2+)-dependent archaeal-specific GTP cyclohydrolase, MptA, from Methanocaldococcus jannaschii. Biochemistry. 2007;46:6658–6667. doi: 10.1021/bi700052a. [DOI] [PubMed] [Google Scholar]
- 231.Mashhadi Z., Xu H., White R.H. An Fe2+-dependent cyclic phosphodiesterase catalyzes the hydrolysis of 7,8-dihydro-D-neopterin 2’,3’-cyclic phosphate in methanopterin biosynthesis. Biochemistry. 2009;48:9384–9392. doi: 10.1021/bi9010336. [DOI] [PubMed] [Google Scholar]
- 232.Scott J.W., Rasche M.E. Purification, overproduction, and partial characterization of beta-RFAP synthase, a key enzyme in the methanopterin biosynthesis pathway. J. Bacteriol. 2002;184:4442–4448. doi: 10.1128/JB.184.16.4442-4448.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Dumitru R.V., Ragsdale S.W. Mechanism of 4-(beta-D-ribofuranosyl)aminobenzene 5’-phosphate synthase, a key enzyme in the methanopterin biosynthetic pathway. J. Biol. Chem. 2004;279:39389–39395. doi: 10.1074/jbc.M406442200. [DOI] [PubMed] [Google Scholar]
- 234.Vaupel M., Vorholt J.A., Thauer R.K. Overproduction and one-step purification of the N5,N10-methenyltetrahydromethanopterin cyclohydrolase (Mch) from the hyperthermophilic Methanopyrus kandleri. Extremophiles. 1998;2:15–22. doi: 10.1007/s007920050038. [DOI] [PubMed] [Google Scholar]
- 235.Zerulla K., Chimileski S., Nather D., Gophna U., Papke R.T., Soppa J. DNA as a phosphate storage polymer and the alternative advantages of polyploidy for growth or survival. PLoS ONE. 2014;9:e94819. doi: 10.1371/journal.pone.0094819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236.Rodionova I.A., Vetting M.W., Li X., Almo S.C., Osterman A.L., Rodionov D.A. A novel bifunctional transcriptional regulator of riboflavin metabolism in Archaea. Nucleic Acids Res. 2017;45:3785–3799. doi: 10.1093/nar/gkw1331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 237.Kawai S., Mori S., Mukai T., Suzuki S., Yamada T., Hashimoto W., Murata K. Inorganic Polyphosphate/ATP-NAD kinase of Micrococcus flavus and Mycobacterium tuberculosis H37Rv. Biochem. Biophys. Res. Commun. 2000;276:57–63. doi: 10.1006/bbrc.2000.3433. [DOI] [PubMed] [Google Scholar]
- 238.Raffaelli N., Pisani F.M., Lorenzi T., Emanuelli M., Amici A., Ruggieri S., Magni G. Characterization of nicotinamide mononucleotide adenylyltransferase from thermophilic archaea. J. Bacteriol. 1997;179:7718–7723. doi: 10.1128/jb.179.24.7718-7723.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Raffaelli N., Emanuelli M., Pisani F.M., Amici A., Lorenzi T., Ruggieri S., Magni G. Identification of the archaeal NMN adenylyltransferase gene. Mol. Cell. Biochem. 1999;193:99–102. doi: 10.1023/A:1006968328186. [DOI] [PubMed] [Google Scholar]
- 240.Eustaquio A.S., Harle J., Noel J.P., Moore B.S. S-Adenosyl-L-methionine hydrolase (adenosine-forming), a conserved bacterial and archaeal protein related to SAM-dependent halogenases. ChemBioChem. 2008;9:2215–2219. doi: 10.1002/cbic.200800341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Deng H., Botting C.H., Hamilton J.T., Russell R.J., O’Hagan D. S-adenosyl-L-methionine:hydroxide adenosyltransferase: A SAM enzyme. Angew. Chem. Int. Ed. Engl. 2008;47:5357–5361. doi: 10.1002/anie.200800794. [DOI] [PubMed] [Google Scholar]
- 242.Fischer M., Romisch W., Schiffmann S., Kelly M., Oschkinat H., Steinbacher S., Huber R., Eisenreich W., Richter G., Bacher A. Biosynthesis of riboflavin in archaea studies on the mechanism of 3,4-dihydroxy-2-butanone-4-phosphate synthase of Methanococcus jannaschii. J. Biol. Chem. 2002;277:41410–41416. doi: 10.1074/jbc.M206863200. [DOI] [PubMed] [Google Scholar]
- 243.Haase I., Mortl S., Kohler P., Bacher A., Fischer M. Biosynthesis of riboflavin in archaea. 6,7-dimethyl-8-ribityllumazine synthase of Methanococcus jannaschii. Eur. J. Biochem. 2003;270:1025–1032. doi: 10.1046/j.1432-1033.2003.03478.x. [DOI] [PubMed] [Google Scholar]
- 244.Phillips G., Grochowski L.L., Bonnett S., Xu H., Bailly M., Blaby-Haas C., El Yacoubi B., Iwata-Reuyl D., White R.H., de Crecy-Lagard V. Functional promiscuity of the COG0720 family. ACS Chem. Biol. 2012;7:197–209. doi: 10.1021/cb200329f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Graham D.E., Xu H., White R.H. A member of a new class of GTP cyclohydrolases produces formylaminopyrimidine nucleotide monophosphates. Biochemistry. 2002;41:15074–15084. doi: 10.1021/bi0268798. [DOI] [PubMed] [Google Scholar]
- 246.Graupner M., Xu H., White R.H. The pyrimidine nucleotide reductase step in riboflavin and F(420) biosynthesis in archaea proceeds by the eukaryotic route to riboflavin. J. Bacteriol. 2002;184:1952–1957. doi: 10.1128/JB.184.7.1952-1957.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Romisch-Margl W., Eisenreich W., Haase I., Bacher A., Fischer M. 2,5-diamino-6-ribitylamino-4(3H)-pyrimidinone 5’-phosphate synthases of fungi and archaea. FEBS J. 2008;275:4403–4414. doi: 10.1111/j.1742-4658.2008.06586.x. [DOI] [PubMed] [Google Scholar]
- 248.Ammelburg M., Hartmann M.D., Djuranovic S., Alva V., Koretke K.K., Martin J., Sauer G., Truffault V., Zeth K., Lupas A.N., et al. A CTP-dependent archaeal riboflavin kinase forms a bridge in the evolution of cradle-loop barrels. Structure. 2007;15:1577–1590. doi: 10.1016/j.str.2007.09.027. [DOI] [PubMed] [Google Scholar]
- 249.Mashhadi Z., Xu H., Grochowski L.L., White R.H. Archaeal RibL: A new FAD synthetase that is air sensitive. Biochemistry. 2010;49:8748–8755. doi: 10.1021/bi100817q. [DOI] [PubMed] [Google Scholar]
- 250.Caforio A., Driessen A.J.M. Archaeal phospholipids: Structural properties and biosynthesis. Biochim. Biophys. Acta Mol. Cell Biol. Lipids. 2017;1862:1325–1339. doi: 10.1016/j.bbalip.2016.12.006. [DOI] [PubMed] [Google Scholar]
- 251.Vannice J.C., Skaff D.A., Keightley A., Addo J.K., Wyckoff G.J., Miziorko H.M. Identification in Haloferax volcanii of phosphomevalonate decarboxylase and isopentenyl phosphate kinase as catalysts of the terminal enzyme reactions in an archaeal alternate mevalonate pathway. J. Bacteriol. 2014;196:1055–1063. doi: 10.1128/JB.01230-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 252.De Rosa M., Gambacorta A. The lipids of archaebacteria. Prog. Lipid Res. 1988;27:153–175. doi: 10.1016/0163-7827(88)90011-2. [DOI] [PubMed] [Google Scholar]
- 253.Dawson K.S., Freeman K.H., Macalady J.L. Molecular characterization of core lipids from halophilic archaea grown under different salinity conditions. Org. Geochem. 2012;48:1–8. doi: 10.1016/j.orggeochem.2012.04.003. [DOI] [Google Scholar]
- 254.Oren A., Ventosa A., Grant W.D. Proposed minimal standards for description of new taxa in the order Halobacteriales. Int. J. Syst. Bacteriol. 1997;47:233–238. doi: 10.1099/00207713-47-1-233. [DOI] [Google Scholar]
- 255.Oren A. Molecular ecology of extremely halophilic Archaea and Bacteria. FEMS Microbiol. Ecol. 2002;39:1–7. doi: 10.1111/j.1574-6941.2002.tb00900.x. [DOI] [PubMed] [Google Scholar]
- 256.Kushwaha S.C., Kramer J.K.G., Kates M. Isolation and characterization of C50-carotenoid pigments and other polar isoprenoids from Halobacterium cutirubrum. Biochim. Biophys. Acta. 1975;398:303–314. doi: 10.1016/0005-2760(75)90146-0. [DOI] [PubMed] [Google Scholar]
- 257.Yang Y., Yatsunami R., Ando A., Miyoko N., Fukui T., Takaichi S., Nakamura S. Complete biosynthetic pathway of the C50 carotenoid bacterioruberin from lycopene in the extremely halophilic archaeon Haloarcula japonica. J. Bacteriol. 2015;197:1614–1623. doi: 10.1128/JB.02523-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Giani M., Miralles-Robledillo J.M., Peiro G., Pire C., Martinez-Espinosa R.M. Deciphering pathways for carotenogenesis in haloarchaea. Molecules. 2020;25:1197. doi: 10.3390/molecules25051197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Dummer A.M., Bonsall J.C., Cihla J.B., Lawry S.M., Johnson G.C., Peck R.F. Bacterioopsin-mediated regulation of bacterioruberin biosynthesis in Halobacterium salinarum. J. Bacteriol. 2011;193:5658–5667. doi: 10.1128/JB.05376-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Tachibana A. A novel prenyltransferase, farnesylgeranyl diphosphate synthase, from the haloalkaliphilic archaeon, Natronobacterium pharaonis. FEBS Lett. 1994;341:291–294. doi: 10.1016/0014-5793(94)80475-3. [DOI] [PubMed] [Google Scholar]
- 261.Bale N.J., Sorokin D.Y., Hopmans E.C., Koenen M., Rijpstra W.I.C., Villanueva L., Wienk H., Sinninghe Damste J.S. New insights into the polar lipid composition of extremely halo(alkali)philic euryarchaea from hypersaline lakes. Front. Microbiol. 2019;10:377. doi: 10.3389/fmicb.2019.00377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262.Kates M. Biology of halophilic bacteria, Part II. Membrane lipids of extreme halophiles: Biosynthesis, function and evolutionary significance. Experientia. 1993;49:1027–1036. doi: 10.1007/BF01929909. [DOI] [PubMed] [Google Scholar]
- 263.Kates M., Moldoveanu N., Stewart L.C. On the revised structure of the major phospholipid of Halobacterium salinarium. Biochim. Biophys. Acta. 1993;1169:46–53. doi: 10.1016/0005-2760(93)90080-S. [DOI] [PubMed] [Google Scholar]
- 264.Kellermann M.Y., Yoshinaga M.Y., Valentine R.C., Wormer L., Valentine D.L. Important roles for membrane lipids in haloarchaeal bioenergetics. Biochim. Biophys. Acta. 2016;1858:2940–2956. doi: 10.1016/j.bbamem.2016.08.010. [DOI] [PubMed] [Google Scholar]
- 265.Elling F.J., Becker K.W., Konneke M., Schroder J.M., Kellermann M.Y., Thomm M., Hinrichs K.U. Respiratory quinones in Archaea: Phylogenetic distribution and application as biomarkers in the marine environment. Environ. Microbiol. 2016;18:692–707. doi: 10.1111/1462-2920.13086. [DOI] [PubMed] [Google Scholar]
- 266.Petrova T.E., Boyko K.M., Nikolaeva A.Y., Stekhanova T.N., Gruzdev E.V., Mardanov A.V., Stroilov V.S., Littlechild J.A., Popov V.O., Bezsudnova E.Y. Structural characterization of geranylgeranyl pyrophosphate synthase GACE1337 from the hyperthermophilic archaeon Geoglobus acetivorans. Extremophiles. 2018;22:877–888. doi: 10.1007/s00792-018-1044-5. [DOI] [PubMed] [Google Scholar]
- 267.Tachibana A., Yano Y., Otani S., Nomura N., Sako Y., Taniguchi M. Novel prenyltransferase gene encoding farnesylgeranyl diphosphate synthase from a hyperthermophilic archaeon, Aeropyrum pernix—Molecular evolution with alteration in product specificity. Eur. J. Biochem. 2000;267:321–328. doi: 10.1046/j.1432-1327.2000.00967.x. [DOI] [PubMed] [Google Scholar]
- 268.Jain S., Caforio A., Fodran P., Lolkema J.S., Minnaard A.J., Driessen A.J.M. Identification of CDP-archaeol synthase, a missing link of ether lipid biosynthesis in Archaea. Chem. Biol. 2014;21:1392–1401. doi: 10.1016/j.chembiol.2014.07.022. [DOI] [PubMed] [Google Scholar]
- 269.Morii H., Koga Y. CDP-2,3-Di-O-geranylgeranyl-sn-glycerol:L-serine O-archaetidyltransferase (archaetidylserine synthase) in the methanogenic archaeon Methanothermobacter thermautotrophicus. J. Bacteriol. 2003;185:1181–1189. doi: 10.1128/JB.185.4.1181-1189.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 270.Morii H., Kiyonari S., Ishino Y., Koga Y. A novel biosynthetic pathway of archaetidyl-myo-inositol via archaetidyl-myo-inositol phosphate from CDP-archaeol and D-glucose 6-phosphate in methanoarchaeon Methanothermobacter thermautotrophicus cells. J. Biol. Chem. 2009;284:30766–30774. doi: 10.1074/jbc.M109.034652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 271.Vences-Guzman M.A., Geiger O., Sohlenkamp C. Sinorhizobium meliloti mutants deficient in phosphatidylserine decarboxylase accumulate phosphatidylserine and are strongly affected during symbiosis with alfalfa. J. Bacteriol. 2008;190:6846–6856. doi: 10.1128/JB.00610-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 272.Conover R.K., Doolittle W.F. Characterization of a gene involved in histidine biosynthesis in Halobacterium (Haloferax) volcanii: Isolation and rapid mapping by transformation of an auxotroph with cosmid DNA. J. Bacteriol. 1990;172:3244–3249. doi: 10.1128/jb.172.6.3244-3249.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Grisolia V., Carlomagno M.S., Nappo A.G., Bruni C.B. Cloning, structure, and expression of the Escherichia coli K-12 hisC gene. J. Bacteriol. 1985;164:1317–1323. doi: 10.1128/jb.164.3.1317-1323.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 274.Loc’h J., Blaud M., Rety S., Lebaron S., Deschamps P., Bareille J., Jombart J., Robert-Paganin J., Delbos L., Chardon F., et al. RNA mimicry by the fap7 adenylate kinase in ribosome biogenesis. PLoS Biol. 2014;12:e1001860. doi: 10.1371/journal.pbio.1001860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 275.Ren H., Wang L., Bennett M., Liang Y., Zheng X., Lu F., Li L., Nan J., Luo M., Eriksson S., et al. The crystal structure of human adenylate kinase 6: An adenylate kinase localized to the cell nucleus. Proc. Natl. Acad. Sci. USA. 2005;102:303–308. doi: 10.1073/pnas.0407459102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 276.Moon S., Kim J., Koo J., Bae E. Structural and mutational analyses of psychrophilic and mesophilic adenylate kinases highlight the role of hydrophobic interactions in protein thermal stability. Struct. Dyn. 2019;6:024702. doi: 10.1063/1.5089707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 277.Chen L., Zhou C., Yang H., Roberts M.F. Inositol-1-phosphate synthase from Archaeoglobus fulgidus is a class II aldolase. Biochemistry. 2000;39:12415–12423. doi: 10.1021/bi001517q. [DOI] [PubMed] [Google Scholar]
- 278.Neelon K., Roberts M.F., Stec B. Crystal structure of a trapped catalytic intermediate suggests that forced atomic proximity drives the catalysis of mIPS. Biophys. J. 2011;101:2816–2824. doi: 10.1016/j.bpj.2011.10.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 279.Maurer S., Ludt K., Soppa J. Characterization of copy number control of two Haloferax volcanii replication origins using deletion mutants and haloarchaeal artificial chromosomes. J. Bacteriol. 2018;200:e00517-17. doi: 10.1128/JB.00517-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 280.Chamovitz D., Misawa N., Sandmann G., Hirschberg J. Molecular cloning and expression in Escherichia coli of a cyanobacterial gene coding for phytoene synthase, a carotenoid biosynthesis enzyme. FEBS Lett. 1992;296:305–310. doi: 10.1016/0014-5793(92)80310-D. [DOI] [PubMed] [Google Scholar]
- 281.Serino L., Reimmann C., Baur H., Beyeler M., Visca P., Haas D. Structural genes for salicylate biosynthesis from chorismate in Pseudomonas aeruginosa. Mol. Gen. Genet. 1995;249:217–228. doi: 10.1007/BF00290369. [DOI] [PubMed] [Google Scholar]
- 282.Dawson A., Chen M., Fyfe P.K., Guo Z., Hunter W.N. Structure and reactivity of Bacillus subtilis MenD catalyzing the first committed step in menaquinone biosynthesis. J. Mol. Biol. 2010;401:253–264. doi: 10.1016/j.jmb.2010.06.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 283.Schmidt D.M., Hubbard B.K., Gerlt J.A. Evolution of enzymatic activities in the enolase superfamily: Functional assignment of unknown proteins in Bacillus subtilis and Escherichia coli as L-Ala-D/L-Glu epimerases. Biochemistry. 2001;40:15707–15715. doi: 10.1021/bi011640x. [DOI] [PubMed] [Google Scholar]
- 284.Palmer D.R., Garrett J.B., Sharma V., Meganathan R., Babbitt P.C., Gerlt J.A. Unexpected divergence of enzyme function and sequence: "N-acylamino acid racemase" is o-succinylbenzoate synthase. Biochemistry. 1999;38:4252–4258. doi: 10.1021/bi990140p. [DOI] [PubMed] [Google Scholar]
- 285.Chen Y., Jiang Y., Guo Z. Mechanistic insights from the crystal structure of Bacillus subtilis o-succinylbenzoyl-CoA synthetase complexed with the adenylate intermediate. Biochemistry. 2016;55:6685–6695. doi: 10.1021/acs.biochem.6b00889. [DOI] [PubMed] [Google Scholar]
- 286.Jiang M., Chen M., Guo Z.F., Guo Z. A bicarbonate cofactor modulates 1,4-dihydroxy-2-naphthoyl-coenzyme a synthase in menaquinone biosynthesis of Escherichia coli. J. Biol. Chem. 2010;285:30159–30169. doi: 10.1074/jbc.M110.147702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 287.Suvarna K., Stevenson D., Meganathan R., Hudspeth M.E. Menaquinone (vitamin K2) biosynthesis: Localization and characterization of the menA gene from Escherichia coli. J. Bacteriol. 1998;180:2782–2787. doi: 10.1128/JB.180.10.2782-2787.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 288.Cheng Z., Sattler S., Maeda H., Sakuragi Y., Bryant D.A., DellaPenna D. Highly divergent methyltransferases catalyze a conserved reaction in tocopherol and plastoquinone synthesis in cyanobacteria and photosynthetic eukaryotes. Plant Cell. 2003;15:2343–2356. doi: 10.1105/tpc.013656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 289.Koike-Takeshita A., Koyama T., Ogura K. Identification of a novel gene cluster participating in menaquinone (vitamin K2) biosynthesis. Cloning and sequence determination of the 2-heptaprenyl-1,4-naphthoquinone methyltransferase gene of Bacillus stearothermophilus. J. Biol. Chem. 1997;272:12380–12383. doi: 10.1074/jbc.272.19.12380. [DOI] [PubMed] [Google Scholar]
- 290.Leffers H., Gropp F., Lottspeich F., Zillig W., Garrett R.A. Sequence, organization, transcription and evolution of RNA polymerase subunit genes from the archaebacterial extreme halophiles Halobacterium halobium and Halococcus morrhuae. J. Mol. Biol. 1989;206:1–17. doi: 10.1016/0022-2836(89)90519-6. [DOI] [PubMed] [Google Scholar]
- 291.Madon J., Zillig W. A form of the DNA-dependent RNA polymerase of Halobacterium halobium, containing an additional component, is able to transcribe native DNA. Eur. J. Biochem. 1983;133:471–474. doi: 10.1111/j.1432-1033.1983.tb07487.x. [DOI] [PubMed] [Google Scholar]
- 292.Rivlin A.A., Chan Y.L., Wool I.G. The contribution of a zinc finger motif to the function of yeast ribosomal protein YL37a. J. Mol. Biol. 1999;294:909–919. doi: 10.1006/jmbi.1999.3309. [DOI] [PubMed] [Google Scholar]
- 293.de Crecy-Lagard V., Forouhar F., Brochier-Armanet C., Tong L., Hunt J.F. Comparative genomic analysis of the DUF71/COG2102 family predicts roles in diphthamide biosynthesis and B12 salvage. Biol. Direct. 2012;7:32. doi: 10.1186/1745-6150-7-32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 294.Ng S.Y., Chaban B., VanDyke D.J., Jarrell K.F. Archaeal signal peptidases. Microbiology (Reading) 2007;153:305–314. doi: 10.1099/mic.0.2006/003087-0. [DOI] [PubMed] [Google Scholar]
- 295.Raut P., Glass J.B., Lieberman R.L. Archaeal roots of intramembrane aspartyl protease siblings signal peptide peptidase and presenilin. Proteins. 2021;89:232–241. doi: 10.1002/prot.26009. [DOI] [PubMed] [Google Scholar]
- 296.Arndt E., Scholzen T., Kromer W., Hatakeyama T., Kimura M. Primary structures of ribosomal proteins from the archaebacterium Halobacterium marismortui and the eubacterium Bacillus stearothermophilus. Biochimie. 1991;73:657–668. doi: 10.1016/0300-9084(91)90045-3. [DOI] [PubMed] [Google Scholar]
- 297.Scholzen T., Arndt E. Organization and nucleotide sequence of ten ribosomal protein genes from the region equivalent to the spectinomycin operon in the archaebacterium Halobacterium marismortui. Mol. Genet. Genomics. 1991;228:70–80. doi: 10.1007/BF00282450. [DOI] [PubMed] [Google Scholar]
- 298.Otaka E., Higo K., Itoh T. Yeast ribosomal proteins. VIII. Isolation of two proteins and sequence characterization of twenty-four proteins from cytoplasmic ribosomes. Mol. Gen. Genet. 1984;195:544–546. doi: 10.1007/BF00341461. [DOI] [PubMed] [Google Scholar]
- 299.Ban N., Nissen P., Hansen J., Moore P.B., Steitz T.A. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science. 2000;289:905–920. doi: 10.1126/science.289.5481.905. [DOI] [PubMed] [Google Scholar]
- 300.Dresios J., Chan Y.L., Wool I.G. The role of the zinc finger motif and of the residues at the amino terminus in the function of yeast ribosomal protein YL37a. J. Mol. Biol. 2002;316:475–488. doi: 10.1006/jmbi.2001.5374. [DOI] [PubMed] [Google Scholar]
- 301.Zhu X., Dzikovski B., Su X., Torelli A.T., Zhang Y., Ealick S.E., Freed J.H., Lin H. Mechanistic understanding of Pyrococcus horikoshii Dph2, a [4Fe-4S] enzyme required for diphthamide biosynthesis. Mol. Biosyst. 2011;7:74–81. doi: 10.1039/C0MB00076K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 302.Zhu X., Kim J., Su X., Lin H. Reconstitution of diphthine synthase activity in vitro. Biochemistry. 2010;49:9649–9657. doi: 10.1021/bi100812h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 303.Su X., Lin Z., Chen W., Jiang H., Zhang S., Lin H. Chemogenomic approach identified yeast YLR143W as diphthamide synthetase. Proc. Natl. Acad. Sci. USA. 2012;109:19983–19987. doi: 10.1073/pnas.1214346109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 304.Uthman S., Bar C., Scheidt V., Liu S., ten Have S., Giorgini F., Stark M.J., Schaffrath R. The amidation step of diphthamide biosynthesis in yeast requires DPH6, a gene identified through mining the DPH1-DPH5 interaction network. PLoS Genet. 2013;9:e1003334. doi: 10.1371/journal.pgen.1003334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 305.Bolhuis A., Matzen A., Hyyrylainen H.L., Kontinen V.P., Meima R., Chapuis J., Venema G., Bron S., Freudl R., van Dijl J.M. Signal peptide peptidase- and ClpP-like proteins of Bacillus subtilis required for efficient translocation and processing of secretory proteins. J. Biol. Chem. 1999;274:24585–24592. doi: 10.1074/jbc.274.35.24585. [DOI] [PubMed] [Google Scholar]
- 306.Nam S.E., Kim A.C., Paetzel M. Crystal structure of Bacillus subtilis signal peptide peptidase A. J. Mol. Biol. 2012;419:347–358. doi: 10.1016/j.jmb.2012.03.020. [DOI] [PubMed] [Google Scholar]
- 307.Rangaswamy V., Altekar W. Ketohexokinase (ATP:D-fructose 1-phosphotransferase) from a halophilic archaebacterium, Haloarcula vallismortis: Purification and properties. J. Bacteriol. 1994;176:5505–5512. doi: 10.1128/jb.176.17.5505-5512.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 308.Anderson I., Scheuner C., Goker M., Mavromatis K., Hooper S.D., Porat I., Klenk H.P., Ivanova N., Kyrpides N. Novel insights into the diversity of catabolic metabolism from ten haloarchaeal genomes. PLoS ONE. 2011;6:e20237. doi: 10.1371/journal.pone.0020237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 309.Williams T.J., Allen M.A., Liao Y., Raftery M.J., Cavicchioli R. Sucrose metabolism in haloarchaea: Reassessment using genomics, proteomics, and metagenomics. Appl. Environ. Microbiol. 2019;85:e02935-18. doi: 10.1128/AEM.02935-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 310.Chaga G., Porath J., Illeni T. Isolation and purification of amyloglucosidase from Halobacterium sodomense. Biomed. Chromatogr. 1993;7:256–261. doi: 10.1002/bmc.1130070504. [DOI] [PubMed] [Google Scholar]
- 311.Rudolph B., Hansen T., Schonheit P. Glucose-6-phosphate isomerase from the hyperthermophilic archaeon Methanococcus jannaschii: Characterization of the first archaeal member of the phosphoglucose isomerase superfamily. Arch. Microbiol. 2004;181:82–87. doi: 10.1007/s00203-003-0626-4. [DOI] [PubMed] [Google Scholar]
- 312.Aulkemeyer P., Ebner R., Heilenmann G., Jahreis K., Schmid K., Wrieden S., Lengeler J.W. Molecular analysis of two fructokinases involved in sucrose metabolism of enteric bacteria. Mol. Microbiol. 1991;5:2913–2922. doi: 10.1111/j.1365-2958.1991.tb01851.x. [DOI] [PubMed] [Google Scholar]
- 313.Qu Q., Lee S.J., Boos W. Molecular and biochemical characterization of a fructose-6-phosphate-forming and ATP-dependent fructokinase of the hyperthermophilic archaeon Thermococcus litoralis. Extremophiles. 2004;8:301–308. doi: 10.1007/s00792-004-0392-5. [DOI] [PubMed] [Google Scholar]
- 314.Ohnishi H., Kitamura H., Minowa T., Sakai H., Ohta T. Molecular cloning of a glucoamylase gene from a thermophilic Clostridium and kinetics of the cloned enzyme. Eur. J. Biochem. 1992;207:413–418. doi: 10.1111/j.1432-1033.1992.tb17064.x. [DOI] [PubMed] [Google Scholar]
- 315.Ahmed H., Ettema T.J., Tjaden B., Geerling A.C., van der Oost J., Siebers B. The semi-phosphorylative Entner-Doudoroff pathway in hyperthermophilic archaea: A re-evaluation. Biochem. J. 2005;390:529–540. doi: 10.1042/BJ20041711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 316.Chai Y., Kolter R., Losick R. A widely conserved gene cluster required for lactate utilization in Bacillus subtilis and its involvement in biofilm formation. J. Bacteriol. 2009;191:2423–2430. doi: 10.1128/JB.01464-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 317.Gao C., Wang Y., Zhang Y., Lv M., Dou P., Xu P., Ma C. NAD-independent L-lactate dehydrogenase required for L-lactate utilization in Pseudomonas stutzeri A1501. J. Bacteriol. 2015;197:2239–2247. doi: 10.1128/JB.00017-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 318.Pfrimer P., de Moraes L.M., Galdino A.S., Salles L.P., Reis V.C., De Marco J.L., Prates M.V., Bloch C., Jr., Torres F.A. Cloning, purification, and partial characterization of Bacillus subtilis urate oxidase expressed in Escherichia coli. J. Biomed. Biotechnol. 2010;2010:674908. doi: 10.1155/2010/674908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 319.Lee Y., Lee D.H., Kho C.W., Lee A.Y., Jang M., Cho S., Lee C.H., Lee J.S., Myung P.K., Park B.C., et al. Transthyretin-related proteins function to facilitate the hydrolysis of 5-hydroxyisourate, the end product of the uricase reaction. FEBS Lett. 2005;579:4769–4774. doi: 10.1016/j.febslet.2005.07.056. [DOI] [PubMed] [Google Scholar]
- 320.Kim K., Park J., Rhee S. Structural and functional basis for (S)-allantoin formation in the ureide pathway. J. Biol. Chem. 2007;282:23457–23464. doi: 10.1074/jbc.M703211200. [DOI] [PubMed] [Google Scholar]
- 321.Xu Z., Jiang W.H., Jiao R.S., Yang Y.L. [Cloning, sequencing and high expression in Escherichia coli of D-hydantoinase gene from Burkholderia pickettii] Sheng Wu Gong Cheng Xue Bao. 2002;18:149–154. [PubMed] [Google Scholar]
- 322.Ho Y.Y., Huang Y.H., Huang C.Y. Chemical rescue of the post-translationally carboxylated lysine mutant of allantoinase and dihydroorotase by metal ions and short-chain carboxylic acids. Amino Acids. 2013;44:1181–1191. doi: 10.1007/s00726-012-1451-3. [DOI] [PubMed] [Google Scholar]
- 323.Schultz A.C., Nygaard P., Saxild H.H. Functional analysis of 14 genes that constitute the purine catabolic pathway in Bacillus subtilis and evidence for a novel regulon controlled by the PucR transcription activator. J. Bacteriol. 2001;183:3293–3302. doi: 10.1128/JB.183.11.3293-3302.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 324.Martinez-Rodriguez S., Garcia-Pino A., Las Heras-Vazquez F.J., Clemente-Jimenez J.M., Rodriguez-Vico F., Garcia-Ruiz J.M., Loris R., Gavira J.A. Mutational and structural analysis of L-N-carbamoylase reveals new insights into a peptidase M20/M25/M40 family member. J. Bacteriol. 2012;194:5759–5768. doi: 10.1128/JB.01056-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 325.Werner A.K., Romeis T., Witte C.P. Ureide catabolism in Arabidopsis thaliana and Escherichia coli. Nat. Chem. Biol. 2010;6:19–21. doi: 10.1038/nchembio.265. [DOI] [PubMed] [Google Scholar]
- 326.Werner A.K., Medina-Escobar N., Zulawski M., Sparkes I.A., Cao F.Q., Witte C.P. The ureide-degrading reactions of purine ring catabolism employ three amidohydrolases and one aminohydrolase in Arabidopsis, soybean, and rice. Plant Physiol. 2013;163:672–681. doi: 10.1104/pp.113.224261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 327.Kardinahl S., Schmidt C.L., Hansen T., Anemuller S., Petersen A., Schafer G. The strict molybdate-dependence of glucose-degradation by the thermoacidophile Sulfolobus acidocaldarius reveals the first crenarchaeotic molybdenum containing enzyme—An aldehyde oxidoreductase. Eur. J. Biochem. 1999;260:540–548. doi: 10.1046/j.1432-1327.1999.00201.x. [DOI] [PubMed] [Google Scholar]
- 328.Kang B.S., Kim Y.M. Cloning and molecular characterization of the genes for carbon monoxide dehydrogenase and localization of molybdopterin, flavin adenine dinucleotide, and iron-sulfur centers in the enzyme of Hydrogenophaga pseudoflava. J. Bacteriol. 1999;181:5581–5590. doi: 10.1128/JB.181.18.5581-5590.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 329.Xi H., Schneider B.L., Reitzer L. Purine catabolism in Escherichia coli and function of xanthine dehydrogenase in purine salvage. J. Bacteriol. 2000;182:5332–5341. doi: 10.1128/JB.182.19.5332-5341.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 330.Karatza P., Frillingos S. Cloning and functional characterization of two bacterial members of the NAT/NCS2 family in Escherichia coli. Mol. Membr. Biol. 2005;22:251–261. doi: 10.1080/09687860500092927. [DOI] [PubMed] [Google Scholar]
- 331.Desguin B., Soumillion P., Hols P., Hausinger R.P. Nickel-pincer cofactor biosynthesis involves LarB-catalyzed pyridinium carboxylation and LarE-dependent sacrificial sulfur insertion. Proc. Natl. Acad. Sci. USA. 2016;113:5598–5603. doi: 10.1073/pnas.1600486113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 332.He J., Yin W., Galperin M.Y., Chou S.H. Cyclic di-AMP, a second messenger of primary importance: Tertiary structures and binding mechanisms. Nucleic Acids Res. 2020;48:2807–2829. doi: 10.1093/nar/gkaa112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 333.Fischer S., John von Freyend S., Sabag-Daigle A., Daniels C.J., Allers T., Marchfelder A. Assigning a function to a conserved archaeal metallo-beta-lactamase from Haloferax volcanii. Extremophiles. 2012;16:333–343. doi: 10.1007/s00792-012-0433-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 334.Spath B., Schubert S., Lieberoth A., Settele F., Schutz S., Fischer S., Marchfelder A. Two archaeal tRNase Z enzymes: Similar but different. Arch. Microbiol. 2008;190:301–308. doi: 10.1007/s00203-008-0368-4. [DOI] [PubMed] [Google Scholar]
- 335.Desmarais J.J., Flamholz A.I., Blikstad C., Dugan E.J., Laughlin T.G., Oltrogge L.M., Chen A.W., Wetmore K., Diamond S., Wang J.Y., et al. DABs are inorganic carbon pumps found throughout prokaryotic phyla. Nat. Microbiol. 2019;4:2204–2215. doi: 10.1038/s41564-019-0520-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 336.Corrigan R.M., Grundling A. Cyclic di-AMP: Another second messenger enters the fray. Nat. Rev. Microbiol. 2013;11:513–524. doi: 10.1038/nrmicro3069. [DOI] [PubMed] [Google Scholar]
- 337.Gundlach J., Mehne F.M., Herzberg C., Kampf J., Valerius O., Kaever V., Stulke J. An essential poison: Synthesis and degradation of cyclic di-AMP in Bacillus subtilis. J. Bacteriol. 2015;197:3265–3274. doi: 10.1128/JB.00564-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 338.Commichau F.M., Heidemann J.L., Ficner R., Stulke J. Making and breaking of an essential poison: The cyclases and phosphodiesterases that produce and degrade the essential second messenger cyclic di-AMP in bacteria. J. Bacteriol. 2019;201:e00462-18. doi: 10.1128/JB.00462-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 339.Yin W., Cai X., Ma H., Zhu L., Zhang Y., Chou S.H., Galperin M.Y., He J. A decade of research on the second messenger c-di-AMP. FEMS Microbiol. Rev. 2020;44:701–724. doi: 10.1093/femsre/fuaa019. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Not applicable.