Abstract
The mouse H19 gene is expressed from the maternal chromosome exclusively. A 2-kb region at 2 to 4 kb upstream of H19 is paternally methylated throughout development, and these sequences are necessary for the imprinted expression of both H19 and the 5′-neighboring Igf2 gene. In particular, on the maternal chromosome this element appears to insulate the Igf2 gene from enhancers located downstream of H19. We analyzed the chromatin organization of this element by assaying its sensitivity to nucleases in nuclei. Six DNase I hypersensitive sites (HS sites) were detected on the unmethylated maternal chromosome exclusively, the two most prominent of which mapped 2.25 and 2.75 kb 5′ to the H19 transcription initiation site. Five of the maternal HS sites were present in expressing and nonexpressing tissues and in embryonic stem (ES) cells. They seem, therefore, to reflect the maternal origin of the chromosome rather than the expression of H19. A sixth maternal HS site, at 3.45 kb upstream of H19, was detected in ES cells only. The nucleosomal organization of this element was analyzed in tissues and ES cells by micrococcal nuclease digestion. Specifically on the maternal chromosome, an unusual and strong banding pattern was obtained, suggestive of a nonnucleosomal organization. From our studies, it appears that the unusual chromatin organization with the presence of HS sites (maternal chromosome) and DNA methylation (paternal chromosome) in this element are mutually exclusive and reflect alternate epigenetic states. In addition, our data suggest that nonhistone proteins are associated with the maternal chromosome and that these might be involved in its boundary function.
In mammals, the maternal and paternal genomes are both required for normal development (8, 37, 48). Their functional nonequivalence is mediated by genomic imprinting, an epigenetic mechanism that gives rise to differential expression of the maternal and paternal alleles of certain genes. To date, 28 imprinted genes have been identified in the mouse (5), and many of these genes are expressed in a parental allele-specific manner in humans as well (41). The precise epigenetic features that allow mammalian cells to distinguish the parental alleles of imprinted genes are still poorly understood (41, 47). CpG methylation (40), allelic timing of replication (17), and differential allelic chromatin organization (17) are among the epigenetic features that have been correlated with imprinting. CpG methylation is involved in at least the somatic maintenance of imprinted gene expression (33), and all imprinted genes so far analyzed have sequences that are allele-specifically methylated (40, 41). There have been few reports that indicate a correlation of differential chromatin organization with imprinting. The promoter of the maternally expressed mouse H19 gene was shown to display differential sensitivity to restriction endonucleases (4, 18) and to DNase I (50), and we and others have detected parental chromosome-specific chromatin conformation and paternal DNase I hypersensitivity in the imprinted U2afl-rsl gene in the mouse (16, 45). To further address the question whether differential chromatin organization is a common feature of regions involved in the regulation of imprinting, we set out to analyze nuclease sensitivity in an imprinting control element upstream of the mouse H19 gene.
The imprinted H19 gene on distal mouse chromosome 7 is expressed from the maternal allele and encodes RNA with no apparent protein-coding capacity (3, 39). During prenatal and early-postnatal development, H19 is expressed in tissues of mesodermal and endodermal origin, in which the gene and several kilobases of upstream sequences are hypermethylated on the inactive paternal allele (4, 7, 14, 18, 44). Detailed methylation studies have pinpointed a core region, localized at approximately 2 to 4 kb upstream of the gene, in which CpG dinucleotides are paternally methylated throughout development (38, 52, 53, 57). Furthermore, a study with transgenic mice has suggested that the H19 upstream region is necessary for the imprinted expression of H19 (13). A targeted deletion of the endogenous H19 gene that included 10 kb of upstream sequences resulted in biallelic expression of the 5′ neighboring insulin-like growth factor 2 (Igf2) and insulin 2 (Ins2) genes. This clearly demonstrated that sequences upstream and/or within H19 are also necessary for the paternal expression of these two neighboring imprinted genes (31). Two subsequent targeted deletions of H19 did not include upstream sequences and had only minor effects on Igf2 expression (27, 42). Recently, a targeted deletion was performed in which a 1.6-kb element corresponding to the core region of paternal methylation upstream of H19 was deleted. When paternally transmitted, this deletion led to activation of the paternal H19 gene whereas maternal transmission resulted in activation of the maternal Igf2 gene (51). Based on these targeting experiments, it has been proposed that one of the functions of this upstream element is to insulate the maternal Igf2 gene from enhancers located downstream of H19 (27, 51). This postulate is supported by the finding that when endoderm-specific enhancers downstream of the H19 gene were moved to a position between Igf2 and H19 (at a location upstream of the proposed boundary element), the Igf2 gene became derepressed (58).
Whatever the exact functions of this “boundary–imprinting-control element” on maternal versus paternal chromosomes, these must be determined by its epigenetic status. Mice deficient in the main methyltransferase, DNMT1, show very little methylation in this upstream element (56) and display an increased level of H19 and decreased Igf2 expression (33). Conversely, biallelic methylation of the H19 upstream element is associated with absence of H19 expression and with biallelic Igf2 expression (11). However, from recent work on transgenic mice it follows that hypomethylation of the H19 upstream element is not sufficient to repress Igf2 in cis. Hence, unmethylated YAC transgene constructs containing the Igf2-H19 region, when transfected into differentiated cells, showed expression of both H19 and Igf2 (59). This, together with the finding (34) that the H19 upstream element functions as a silencer in Drosophila (an organism in which methylation is absent [54]) and as a boundary element on the unmethylated maternal chromosome in the mouse, suggests involvement of epigenetic features other than DNA methylation.
For these reasons, we studied the chromatin organization throughout the entire H19 upstream element and detected distinct differences in nuclease sensitivity between the parental chromosomes. Within the core region of differential methylation, multiple DNase I hypersensitive sites (HS sites) and an unusual chromatin organization were identified on the unmethylated maternal chromosome. The pronounced differences in chromatin between the parental chromosomes were present in all expressing and nonexpressing tissues analyzed and therefore reflect the parental origin of the chromosomes rather than expression of H19.
MATERIALS AND METHODS
Mice and ES cells.
F1 mice bearing Mus musculus and M. spretus alleles were produced by crossing C57BL/6 females to M. spretus males. All M. musculus mice analyzed in this study were of the C57BL/6 genotype. Mice with a 13-kb H19-targeted deletion (ΔH19 [31]) were kindly provided by S. M. Tilghman and were mated with C57BL/6 males or females to produce offspring hemizygous for this deletion which encompasses 10 kb of upstream sequences. The embryonic stem (ES) cell lines analyzed were androgenetic lines AG-A and AK; parthenogenetic lines PR3, PR8, and PR18, and (M. musculus × M. spretus)F1 cell lines SF1-G, SF1-1, and SF1-3 (2, 11, 15, 16). For the assays on nuclei, ES cells were cultured in ES medium (2) containing 103 Units of recombinant leukemia inhibitory factor (ESGRO; BRL) per ml in the absence of feeder cells. Assays were performed on semiconfluent early-passage ES cells (passages 4 to 6 for the androgenetic and parthenogenetic lines; passages 10 to 14 for other lines) which showed <10% of morphologically visible differentiation.
Nuclease sensitivity assays and Southern blotting.
Nuclei were purified as described in detail previously (15). Briefly, tissue samples (most frequently from two animals) were homogenized, filtered through cheesecloth, and centrifuged at 4°C. Pelleted cells were resuspended in a 0.3 M sucrose buffer containing 0.2% Nonidet P-40 and were lysed on ice. After lysis, aliquots were layered onto a 1.2 M sucrose buffer and centrifuged at 4°C. Nuclear pellets thus obtained were resuspended in either DNase I or micrococcal nuclease (MNase) buffer at 0.5 × 107 to 1 × 107 nuclei/ml, and nuclease sensitivity assays were performed immediately thereafter. DNase I assays were conducted in 0.3 M sucrose–60 mM KCl–15 mM NaCl–5 mM MgCl2–0.1 mM EGTA–15 mM Tris-HCl (pH 7.5)–0.5 mM dithiothreitol by adding different quantities of DNase I (Boehringer Mannheim) to 200-μl aliquots and incubating the mixture for 10 min at 25°C. MNase (Pharmacia) digestions were performed in 500-μl aliquots (at 300 U/ml) at 37°C in 15 mM Tris-HCl (pH 7.5)–15 mM NaCl–60 mM KCl–0.15 mM β-mercaptoethanol–0.15 mM spermine–0.5 mM spermidine–0.34 M sucrose, 10 mM NaHSO3–1 mM CaCl2. DNase I and MNase reactions were terminated by the addition of an equal volume of 20 mM EDTA (pH 8.0)–1% sodium dodecyl sulfate. Proteinase K was then added to a final concentration of 200 μg/ml, and digestion was performed overnight at 37°C. Genomic DNA was extracted twice with phenol-chloroform, ethanol precipitated, and dissolved in water. After endonuclease digestion, DNA was electrophoresed on 1.0% agarose gels in 1× Tris-borate-EDTA (TBE) buffer, alkali blotted onto Hybond-N+ membrane (Amersham), and UV cross-linked (Stratalinker; Stratagene). Hybridization with radiolabelled fragments and washing of membranes, were performed as described previously (10). Control hybridizations for the MNase assays were performed with a 250-bp HindIII-PstI fragment from the 5′ extremity of the mouse Gapdh gene as the probe. Densitometric measurements on X-ray films were performed with a Bio-Rad GelDoc-1000 apparatus.
RESULTS
Constitutive maternal DNase I HS sites in the H19 upstream region.
We analyzed the DNase I sensitivity of chromatin in the H19 upstream region in adult liver, brain, and kidney tissue dissected from interspecific F1 hybrids between M. musculus females and M. spretus males [(C57BL/6 × M. spretus)F1]. To distinguish between the parental chromosomes, we used restriction fragment length polymorphisms (RFLPs) between the two mouse species. Figure 1 shows the chromosomal region analyzed and indicates the BstXI and SacI RFLPs detected in the H19 upstream region, as well as the genomic DNA probes that were used in this study.
FIG. 1.
H19 upstream region. BstXI (B) and SacI (S) restriction sites in the H19 gene (solid box) and 8-kb of upstream sequences were derived from published sequence information (26; accession no. Af049091) or were mapped in genomic DNA. HhaI restriction sites (vertical bar below the line) located upstream of H19 are also indicated. Fragments H19-6 (a 1.1-kb SacI-BstXI fragment), H19-7 (a 1-kb BamHI-SacI fragment), H19-1 (a 1-kb BamHI fragment [30]), and H19-11 (a 0.4-kb SacI-BamHI fragment) were used as probes to compare M. musculus (M) and M. spretus (S) genomic DNAs. Fragments corresponding to the SacI and BstXI RFLPs identified between the two mouse species are indicated above the sequence.
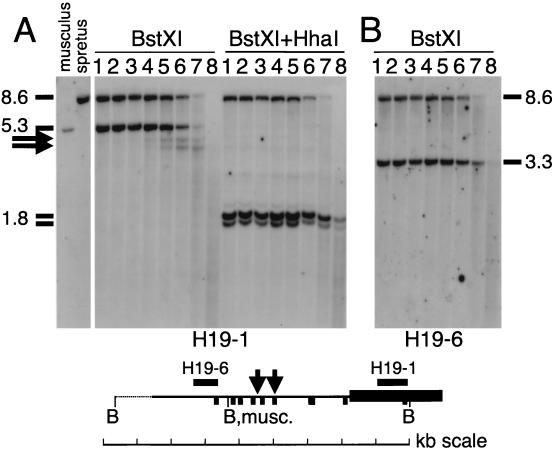
We used the BstXI RFLP to first analyze the DNase I sensitivity of the entire differentially methylated upstream region and the 5′ part of the gene. By hybridizing with a probe (H19-1) from the 5′ portion of the H19 gene, we found that in adult liver (Fig. 2) and adult brain (results not shown) the parental chromosomes displayed comparable generalized sensitivity to DNase I. In nuclei from both tissues, the 5.3-kb maternal fragment showed a similar overall DNase I sensitivity to that of the paternal fragment of 8.6 kb. In addition, the autoradiogram revealed two distinct HS sites at approximately 2.2 kb (digestion product of about 3.7 kb) and 2.7 kb (product of about 4.2 kb) upstream of the H19 gene. Within the promoter region of the H19 gene, no DNase I HS sites were present (Fig. 2A) but promoter DNase I hypersensitivity was detected in neonatal liver in which the H19 gene is expressed at high levels (data not shown). To determine on which parental chromosome the two DNase I HS sites were present, we digested the same adult liver DNase I series with both BstXI and the methylation-sensitive restriction enzyme HhaI. In the H19 upstream region, HhaI digestion leaves the fully methylated paternal chromosomes intact and completely digests the unmethylated maternal allele (52). We thus were able to analyze the DNase I digestion pattern specifically on the paternal allele. No hypersensitivity was seen in the H19 upstream region on the paternal allele (Fig. 2A), thus establishing that the two HS sites in the DNase I series digested with BstXI alone were on the maternal chromosome. That there is no DNA hypersensitivity on the paternal chromosome was confirmed by hybridizing the same DNase I series with a further-upstream fragment (probe H19-6; Fig. 2B). This probe detects the M. spretus-specific 8.6-kb BstXI fragment and a 3.3-kb M. musculus-specific fragment (Fig. 1). No hypersensitivity products deriving from the 8.6-kb paternal fragment were detected.
FIG. 2.
Allele-specific DNase I sensitivity in the H19 upstream region. (A) The BstXI polymorphism between M. musculus and M. spretus was used to analyze adult liver from (C57BL/6 × M. spretus)F1 mice. After incubation of nuclei with increasing concentrations of DNase I (lanes 1 to 8 correspond to 0, 20, 50, 100, 150, 200, 300, and 500 U/ml, respectively), DNA was extracted and digested with either BstXI (series to the left) or BstXI plus HhaI (series to the right). After electrophoresis, hybridization was carried out with probe H19-1. DNase I digestion products are indicated by horizontal arrows. For size determination of fragments (indicated in kilobases), 0.05 μg of size marker DNA (1-kb ladder; Promega) migrated together with 10 μg of BstXI-digested mouse DNA (bands were visualized by hybridization with a radiolabeled size marker [data not shown]). (B) The (C57BL/6 × M. spretus)F1 BstXI-digested liver DNase I series from panel A, probed with H19-6. The map shows the position of BstXI (B) sites relative to H19 (thick black bar). HhaI sites are shown as vertical bars, and genomic probes are shown as horizontal bars. The polymorphic, M. musculus-specific BstXI site is indicated as B, musc. Vertical arrows indicate the approximate positions of the two DNase I HS sites within the 5.3-kb M spretus-specific (maternal) BstXI fragment. Restriction sites derive from published sequence information or were mapped in genomic DNA. The interrupted line indicates a region for which no sequence information was available.
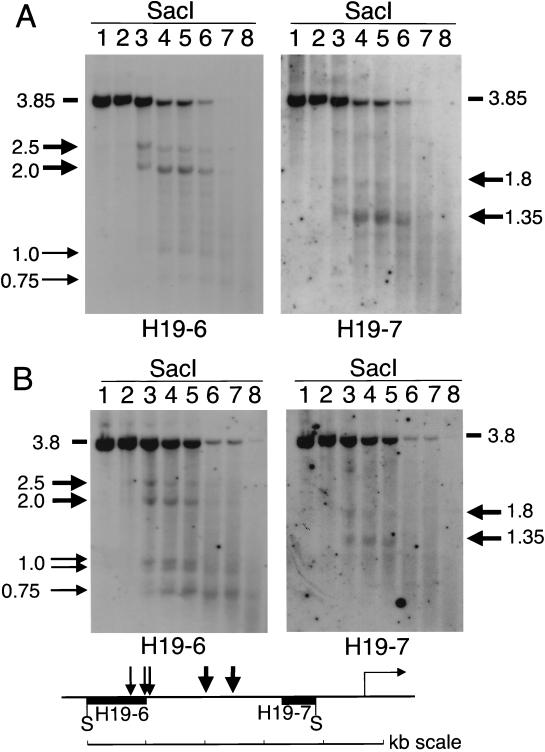
We mapped the two prominent maternal DNase I HS sites more accurately within a 3.85-kb SacI fragment which comprises the core region of the paternally methylated upstream sequences (see Fig. 6). M. musculus liver nuclei were incubated at increasing concentrations of DNase I, and after extraction, genomic DNA samples were digested with the restriction endonuclease SacI. Using DNA fragments from both ends of the SacI restriction fragment as probes, we determined that the two maternal DNase I HS sites were located at 2.75 and 2.25 kb upstream of the transcription initiation site (Fig. 3A). These mapping positions place the two prominent HS sites within the 2-kb core of paternal methylation, at 0.5 and 1 kb 5′ to a 0.46-kb G-rich region that contains multiple copies of a 9-bp repeat sequence (52). Additional, less prominent DNase I HS sites, that could not be detected in the BstXI-digested DNase I series in Fig. 2, were detected further upstream in the 5′ part of the SacI fragment. These less prominent sites also mapped within the core region of paternal methylation, at 3.65 and 4 kb upstream of the transcription initiation site (Fig. 3A; also see Fig. 7). By performing a double digestion of the DNase I series with SacI and HhaI we established that all the DNase I HS sites in liver were present on maternal chromosomes only (results not shown).
FIG. 6.
Allele-specific MNase digestion in ES cells. (A) Nuclei were extracted from early-passage parthenogenetic (line PR8) and androgenetic (line AG-A) cells. Incubation with MNase, gel electrophoresis, and hybridization with probe H19-6 were as in the experiment in Fig. 5B. Lanes 1 to 5 correspond to 0, 30, 45, 60, and 120 s of digestion, respectively, with MNase (Maternal indicates parthenogenetic cells; Paternal indicates androgenetic cells). Maternal MNase digestion products are numbered as in Fig. 5B. Subsequent hybridization of the blot with a Gapdh probe showed that MNase digestion was the same in the parthenogenetic and androgenetic cells (results not shown). (B) Comparison of MNase and DNase I digestion profiles in parthenogenetic cells. The left-hand lane shows the same MNase digestion as shown in lane 3 of panel A. The right-hand lane shows the DNase I digestion on the same nuclei shown (Fig. 4A, lane 6).
FIG. 3.
Maternal DNase I HS sites in liver (A) and brain (B). (A) C57BL/6 adult liver nuclei were incubated with increasing concentrations of DNase I (lanes 1 to 8 correspond to 0, 20, 50, 100, 150, 200, 300, and 500 U/ml, respectively). DNA samples were digested with SacI, and Southern hybridization was performed with H19-6 and H19-7. (B) Analysis of C57BL/6 adult brain was performed as for adult liver in panel A. Fragment sizes are indicated in kilobases (size estimations were performed as in the experiment in Fig. 2). DNase I products are indicated by horizontal arrows. Not indicated is a faint DNase I product of 2.25 kb that was more readily detectable in ES cells (Fig. 4). The map shows the maternal DNase I HS sites in liver and brain, the H19-6 and H19-7 probes, the SacI (S) sites, and the H19 transcription initiation site (horizontal arrow).
FIG. 7.
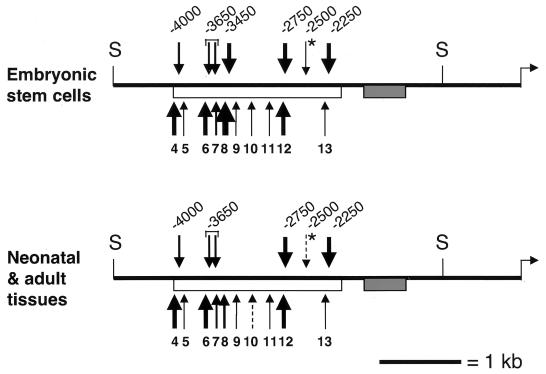
Summary maps of maternal MNase and DNase I sensitivity in ES cells and tissues (liver, brain, and kidney). Vertical arrows above the lines indicate maternal DNase I HS sites identified in this study; the thickness reflects their prominence in the material analyzed. For each DNase I HS site, the nucleotide position relative to the H19 transcription initiation site (horizontal arrow) is indicated. All HS sites map within the core region of paternal DNA methylation (open bar [53, 57]). The HS site marked with an asterisk was visible in ES cells but barely detectable in adult tissues. Vertical arrows below the line indicate the maternal chromosome-specific MNase digestion sites within the paternally methylated region (numbering is as in Fig. 5 and 6). SacI sites (S) are indicated; the shaded bar indicates a 461-bp G-rich region with 32 copies of a 9-bp repeat sequence (26, 52).
We next analyzed DNase I sensitivity in the brain, an ectodermal tissue in which no H19 and Igf2 expression is observed during development or in the adult animal (except for the leptomeningal and choroid plexus epithelia [14, 49]). As in the liver, two prominent DNase I HS sites were present at 2.25 and 2.75 kb upstream of the gene, and the HS sites at 3.65 and 4 kb upstream of the H19 gene were also detected in the brain (Fig. 3B). These latter sites appeared to be stronger in adult brain than in adult liver, and this revealed that the DNase I hypersensitivity at 3.65 kb upstream of the H19 gene consists of two closely linked HS sites (Fig. 3B). A double digestion of the DNase I series with SacI and HhaI was performed to established that in brain, all five DNase I HS sites were present on maternal chromosomes only (results not shown, but see Fig. 5C). We also analyzed nuclei from adult kidney, and this revealed the same maternal DNase I HS sites as were detected in the liver and brain (results not shown).
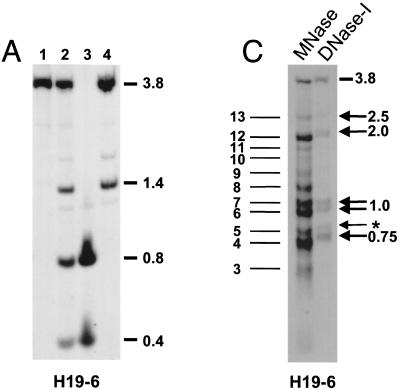
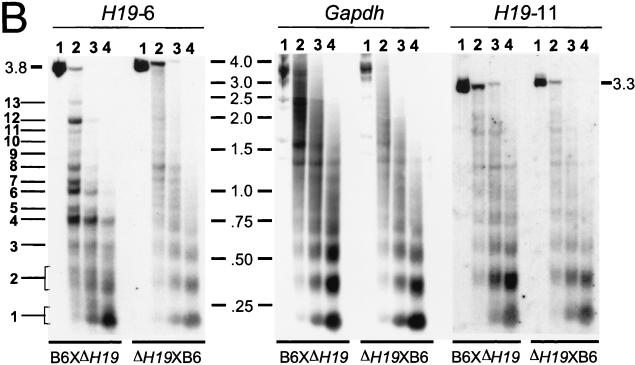
FIG. 5.
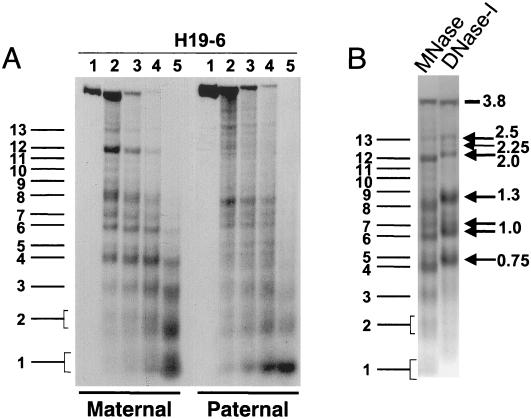
Allele-specific MNase digestion in the brain. (A) Unaltered DNA methylation in brain samples from 1-week-old mice that were hemizygous for a 13-kb deletion comprising the H19 gene and 10 kb of upstream sequences (ΔH19 [31]). Lanes: 1, SacI-digested brain DNA of a C57BL/6 mouse; 2, SacI-HhaI-digested C57BL/6 brain DNA; 3, SacI-HhaI-digested (C57BL/6 × ΔH19)F1 brain DNA; 4, SacI-HhaI-digested (ΔH19 × C57BL/6)F1 brain DNA. Hybridization was with probe H19-6. The 0.4- and 0.8-kb bands are indicative of complete HhaI digestion (absence of DNA methylation); the 1.4-kb band indicates an HhaI restriction site that is partially methylated on the paternal chromosome. (B) MNase digestions on nuclei extracted from brains of 1-week-old mice that were hemizygous for ΔH19. B6XΔH19 corresponds to (C57BL/6 × H19)F1, and ΔH19XB6 corresponds to (ΔH19 × C57BL/6)F1. Lanes 1 to 4 correspond to 0, 30, 60, 120 s of incubation with MNase, respectively. DNA was digested with SacI, and hybridization was carried out with H19-6, H19-11, and a Gapdh probe. Fragment sizes were determined as explained in the legend to Fig. 2. Maternal MNase bands are numbered 1 to 13 starting from the mononucleosomal band. The estimated sizes of bands 4 to 13 are 0.7, 0.8, 0.95, 1.05, 1.25, 1.4, 1.6, 1.8, 2.0, and 2.5 kb, respectively. (C) Comparison of MNase and DNase I digestion profiles. The left-hand lane shows the same MNase digestion as shown in panel B (lane 2, B6XΔH19); the right-hand lane shows a DNase I digestion on the same nuclei. Arrows indicate DNase I HS sites with their size in kilobases to the right. The weak DNase I site indicated with an asterisk was clearly detectable only in brain tissue. No DNase I HS sites were detected in tissues from (ΔH19 × C57BL/6)F1 mice (results not shown).
Maternal hypersensitivity is present in ES cells.
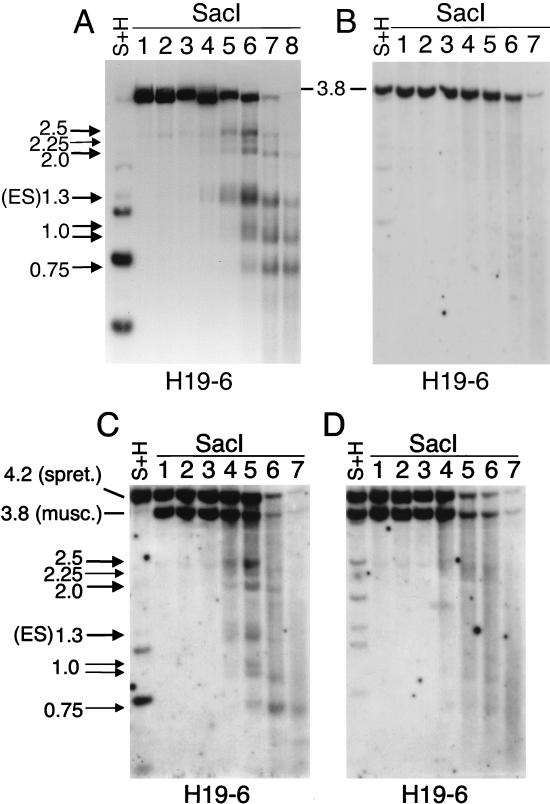
From our tissue-specific analysis, it followed that the five DNase I HS sites which we identified in the H19 upstream element are present in tissues of different embryonic origin and that their presence is not associated with the expression of the H19 gene. To determine whether these five maternal HS sites are present before implantation and differentiation into the different embryonic lineages and whether both parental chromosomes are required for their establishment and maintenance, we performed DNase I studies on monoparental ES cell lines. Both parthenogenetic (with maternal chromosomes only) and androgenetic (with paternal chromosomes only) cell lines were studied. Three parthenogenetic ES cell lines were analyzed that were at an early passage and showed virtually no DNA methylation in the 3.85-kb SacI fragment (Fig. 4A). All five DNase I HS sites which we had detected in liver, brain, and kidney tissue, at 2.25, 2.65, 3.65 (doublet), and 4 kb upstream of the transcription initiation site, were also present in the three parthenogenetic cell lines. An additional HS site was identified at approximately 3.45 kb upstream of the H19 gene. From the thickness of the corresponding band (at ∼1.3 kb), it appears that this additional, strong HS site corresponds to a somewhat broader region of hypersensitivity. A seventh, less prominent DNase I HS site was identified at approximately 2.5 kb upstream of the transcription initiation site (the 2.25-kb band). In two early-passage androgenetic ES cell lines which we analyzed, we did not detect DNase I hypersensitivity and the DNA in the upstream region was completely methylated (Fig. 4B). Our analysis of monoparental ES cells suggested that only the chromosome of maternal origin is required for the maintenance of the chromatin organization characterized by DNase I hypersensitivity and absence of DNA methylation. Conversely, for maintenance of the paternal epigenotype (DNA methylation), only the chromosome of paternal origin is necessary.
FIG. 4.
Maternal DNase I HS sites in ES cells. (A) Parthenogenetic ES cells (line PR8). The left-hand lane shows SacI-HhaI (S+H)-digested genomic DNA, in which the two lower bands correspond to complete HhaI digestion products. In lanes 1 to 8, nuclei were incubated with increasing concentrations of DNase I (corresponding to 0, 50, 100, 200, 300, 400, 500, and 750 U/ml, respectively). DNA was digested with SacI and, after electrophoresis, hybridized with H19-6. Comparable results were obtained with two other parthenogenetic ES cell lines (results not shown). The HS site marked ES was detected in ES cells only. The 2.25-kb DNase I digestion product is not ES cell specific; although fainter, it was also visible in liver, brain (Fig. 3), and kidney tissue (results not shown). (B) Androgenetic ES cells (line AG-A). Lanes 1 to 7 contain 0, 50, 100, 200, 400, 600, and 800 U of DNase I, per ml, respectively. Southern blotting was as in panel A. Comparable results were obtained with another androgenetic line (results not shown). (C) Biparental (C57BL/6 × M. spretus)F1 ES cells (line SF1-G [11]). The DNase I assay were performed as in panel B. This series was also digested with SacI plus HhaI (S+H) and hybridized with H19-6 (results not shown). This yielded no DNase I hypersensitivity products, demonstrating that in this ES cell line the HS sites are present on the unmethylated maternal chromosome. (D) Biparental (C57BL/6 × M. spretus)F1 ES cells (line SF1-1 [11]). The DNase I assay and Southern blot hybridization were performed as in panel B. Another ES cell line with biallelic methylation in the H19 upstream element (SF1-3 [11]) also showed no DNase I HS sites (results not shown).
We next set out to analyze chromatin in biparental ES cells, in an (M. musculus × M. spretus)F1 cell line (SF1-G) which we have shown previously to have unaltered maternal H19 and paternal Igf2 expression in the ES cells themselves and in completely ES-derived fetuses (11). In this ES line, the nine HhaI sites in the SacI restriction fragment were methylated on the paternal chromosome and fully digested on the maternal chromosome (Fig. 4C). As in the parthenogenetic ES cell lines, seven distinct DNase I HS sites were present on the maternal chromosomes exclusively, five of which corresponded to the HS sites detected in adult tissues. The other two sites mapped at approximately 2.5 and 3.45 kb upstream of H19 (as in the parthenogenetic ES cells). Our analysis of monoparental and biparental ES cell lines implies that already at the blastocyst stage, at least in cells of the inner cell mass, the maternal chromosome-specific DNase I HS sites are established in the H19 upstream element.
We have recently shown that methylation in the H19 upstream element (and in the differentially methylated regions within other imprinting genes) can become altered on derivation and culture of ES cells, and in three ES cell lines we observed almost complete methylation on the maternal (in addition to the paternal) allele. This gain of maternal methylation was consistently associated with absence of H19 expression (on Northern analysis) and biallelic Igf2 expression in the ES cell lines (22) and completely ES cell-derived fetuses (11). To determine whether this gain of methylation was associated with alterations in DNase I sensitivity, we performed DNase I assays on these three ES cell lines. As seen in Fig. 4D, the maternal DNase I HS sites were absent or strongly reduced in intensity. Therefore, although chromatin and methylation changes had occurred on derivation and culturing of these ES cell lines, DNA methylation and the presence of DNase I hypersensitivity appeared to be mutually exclusive, as in all other cell lines and tissues analyzed.
Unusual chromatin organization on the maternal chromosome.
To analyze the status of chromatin around the DNase I HS sites in the H19 upstream region, we analyzed its nucleosomal organization on the maternal and the paternal chromosomes by digestion with MNase. MNase digests preferentially the linker DNA between nucleosomes and therefore permits the determination of the positioning of nucleosomes (which, for most of the genome, are found once every ∼200 bp). Our aim was to determine whether, in the H19 upstream element, nucleosomes are uniformly distributed in a canonical array or whether there are discontinuities in the nucleosomal array indicative of nonhistone protein binding. Brains dissected from neonatal mice that were hemizygous for a 13-kb targeted deletion comprising the H19 upstream region (ΔH19 [31]) were analyzed by MNase digestion. For these brain samples, we first verified whether their methylation status had not become altered in the H19 upstream element due to the absence of either the maternal [in (ΔH19 × C57BL/6)F1 animals] or the paternal [in (C57BL/6 × ΔH19)F1 animals] allele. In brain samples with only the maternal copy of this region, all the HhaI sites within the 3.85-kb SacI fragment were unmethylated. Samples with only the paternal copy of this region, in contrast, showed complete methylation of these HhaI sites (Fig. 5A). Hence, methylation levels in the H19 upstream element had not become altered due to hemizygosity in this region.
Brain nuclei from the reciprocal types of hemizygous mice were incubated with MNase. DNA extracted from these series was digested with SacI, Southern blotted, and hybridized with different H19 probes (Fig. 5B). When probed with fragment H19-6, the patterns of hybridization were strikingly different between the reciprocal heterozygotes. Brains from hemizygous animals with only the maternal copy of the H19 region displayed several prominent bands, which were missing or very weak in hemizygotes with only paternal copies. As a control for MNase digestion, the same blot was hybridized with a Gapdh probe. Identical (nucleosomal) digestion profiles were obtained, and this established that, overall, MNase digestion was the same in the reciprocal hemizygotes. Comparison of the Gapdh and H19-6 hybridization profiles revealed a striking feature: hybridizing bands on the maternal chromosomes (i.e., those numbered 4 to 13 in Fig. 5B) did not fall into a canonical nucleosomal array with regularly spaced nucleosomes once every ∼200 bp. For example, bands 4 and 5 were separated by, at most, 100 bp and bands 12 and 13 were separated by about 500 bp. These unusual size differences between MNase digestion products cannot be accounted for by nucleosomes. In addition, some of the maternal chromosome bands (bands 4, 6, 7, 8, and 12) had very high intensity and already appeared at the lowest concentration of MNase. As a second control experiment, the same blot was hybridized with a (PCR-amplified) 294-bp probe from the 5′-most extremity of the SacI restriction fragment (a smaller fragment of H19-6 [results not shown]). This yielded the same maternal MNase digestion products as observed on hybridization with the larger H19-6 fragment (albeit slightly weaker in intensity) and confirmed that the maternal chromosome-specific MNase bands did constitute a single digestion profile (i.e., all the cells had the same nucleosomal organization in this region).
On the paternal chromosomes [in (ΔH19 × C57BL/6)F1 animals], in contrast, the MNase digestion profile was smeary and no strong bands were apparent (on H19-6 hybridization) in the region corresponding to maternal chromosome-specific digestion products 4 to 13, except perhaps for band 8 (Fig. 5A). In addition, the MNase digestion profile for the paternal chromosome was very similar to that of Gapdh. We also hybridized the blot with probe H19-7 (located at the 3′ end of the 3.85-kb SacI fragment). This probe also revealed distinct bands on the maternal chromosome and a rather smeary MNase digestion profile on the paternal chromosome (results not shown). Finally, hybridization with a probe from the H19 gene itself (fragment H19-11) demonstrated that within an SacI fragment containing the 3′ part of the H19 gene, the nucleosomal organization is the same on both parental alleles (Fig. 5B).
Since both the DNase I HS sites and the noncanonical MNase digestion profile suggested association of nonhistone proteins on the maternal chromosome, we were interested in directly comparing the digestion patterns obtained with these two nucleases. SacI-digested genomic DNAs of the brain MNase series (corresponding to sample 2 in Fig. 5B) and of a DNase I digestion on the same nuclei were run next to each other and, after transfer onto nylon membrane, hybridized with probe H19-6. The DNase I HS sites were at the same position as or very close to MNase sites that were clearly indicative of a nonnucleosomal organization (Fig. 5C; also see Fig. 7). Hence, the DNase I HS site at 4 kb upstream of the H19 transcription initiation site mapped between MNase sites 4 and 5, which are separated by <100 bp. The DNase I HS site doublet, at 3.65 kb upstream of H19, mapped between the closely linked MNase sites 6 and 7. The two 3′-most HS sites (at 2.75 and 2.25 kb upstream of H19) mapped close to MNase sites 12 and 13, which are separated by ∼500 bp.
MNase assays were also performed on livers from hemizygous ΔH19 animals, and corresponding SacI-digested DNAs were hybridized with H19-6. This gave the same unusual profile on the maternal chromosome as in the brain; in this tissue, the paternal chromosomes had a nondistinct, normal MNase digestion profile (results not shown).
Since the maternal DNase I HS sites are fully established in ES cells, we wished to determine whether these early embryonic cells also displayed the unusual nonnucleosomal MNase digestion profile on the maternal chromosome. MNase digestions were performed on nuclei from parthenogenetic and androgenetic ES cells (Fig. 6). The maternal chromosomes in parthenogenetic ES cells displayed the same unusual MNase digestion profile in the H19 upstream region as they did in the liver and brain. However, MNase digestion product 8 was more prominent and broader than in these tissues, and this band colocalized with the prominent DNase I HS site at 3.45 kb upstream of H19, which we observed in ES cells only. The paternal chromosomes in androgenetic ES cells, in contrast, yielded a rather smeary MNase digestion profile, and no strong bands were apparent in the region corresponding to maternal chromosome-specific digestion products 4 to 13; this was similar to the digestion profile of the paternal chromosomes in the liver and brain.
DISCUSSION
The unusual chromatin organization on the maternal chromosome is constitutive and suggests association of nonhistone proteins.
In this paper, we describe parental allele-specific DNase I hypersensitivity in a 2-kb region, 2 to 4 kb upstream of the mouse H19 gene. All five constitutive DNase I HS sites which we identified in this boundary–imprinting-control element were present on the unmethylated maternal chromosome only and were detected in tissue samples dissected from animals of different genetic origins. The five sites mapped to 2.25, 2.5, 3.65 (doublet), and 4 kb upstream of the transcription initiation site (Fig. 7). An additional, strong maternal DNase I site was detected in ES cells only, at 3.45 kb upstream of H19. In a recent independent study, Hark and Tilghman identified two regions of maternal DNase I hypersensitivity (at approximately 2.4 and 3.8 kb upstream of H19 [24]) in adult liver. Our DNase I study included both expressing (liver and kidney) and nonexpressing (brain) neonatal and adult tissues, as well as monoparental and biparental ES cell lines. The constitutive presence of the five HS sites seems to reflect the maternal origin of the chromosome rather than the transcriptional activity of H19.
It has been shown that DNase I HS sites correspond frequently to small regions where nucleosomes have been either disrupted or displaced by binding of nonhistone proteins (6, 12, 23). That there is indeed, on the maternal chromosome, a nonnucleosomal organization in the H19 upstream element follows from our MNase assays on brain, liver, and ES cells. In contrast to the methylated paternal chromosomes, which displayed a rather undefined digestion profile suggestive of a normal nucleosomal organization, a distinct and noncanonical banding pattern was apparent on the maternal chromosome in the region of DNase I hypersensitivity (Fig. 7). Taken together, our DNase I and MNase assays demonstrate that the H19 upstream element has a nonnucleosomal chromatin organization on the maternal chromosome with likely association of nonhistone proteins, and this complements recent chromatin studies by others (24, 50).
Although the functional significance of the unusual maternal chromatin organization is unknown, this element upstream of the mouse H19 gene can act as a silencer in transgenic flies (34) and insulates the Igf2 gene from enhancers that are downstream of the H19 gene in the mouse (21, 27, 51, 58). The boundary-insulator property of this element is observed on the maternal allele in the mouse, which is unmethylated in this region (53), as is the Drosophila genome (54). This suggests that epigenetic features other than methylation are responsible for its role. We propose that the unusual, somatically stable chromatin configuration of this element induces a higher-order conformation on the maternal chromosome such that the Igf2 gene becomes insulated from enhancers downstream of the H19 gene (and thereby becomes repressed [32]).
The DNase I HS sites were observed in parthenogenetic but not in androgenetic ES cell lines. For technical reasons, we were unable to determine whether they are present in oocytes. Their presence in parthenogenetic ES cells suggests, however, that this might well be the case, since parthenogenetic embryos are derived from activated eggs. In addition, our analysis of ES cells (and neonatal and adult tissues) confirms that this boundary–imprinting-control element is constitutively kept methylated on the paternal allele and that the maternal allele is constitutively organized in an unusual chromatin configuration. The question arises whether factors associated with the HS sites are important in keeping the maternal chromosome unmethylated at critical stages of development, for example during the genome-wide de novo methylation that occurs during gastrulation. In relation to this distinct possibility, it is interesting that a prominent maternal DNase I HS site was detected in ES cells only, at 3.45 kb upstream of the H19 transcription initiation site. Also from our in vitro differentiation studies (22), it appears that this site is ES cell specific.
Nuclease hypersensitivity and DNA methylation are mutually exclusive and appear to reflect alternate epigenetic states.
In all cell and tissue samples analyzed, it was only on the unmethylated chromosome that DNase I HS sites were observed, and this was also the case in ES cell lines in which the parental epigenotypes had become altered as a consequence of in vitro manipulation. This suggests a mechanism by which DNA methylation and the unusual chromatin organization with DNase I hypersensitivity are mutually exclusive. Could this imply that methylation precludes the unusual chromatin organization, or that the unusual chromatin organization of this element does not favor its methylation? Alternatively, are these epigenotypes interdependent and regulating each other? To mimic the maternal chromosome on which H19 is active and Igf2 is repressed, Webber and Tilghman (59) transfected yeast artificial chromosomes containing the Igf2 and H19 genes into differentiated cell lines and found that both genes were expressed at high levels. In comparison with an earlier transgenic study by Ainscough et al. (1) in which large yeast artificial chromosomes containing both Igf2 and H19 showed imprinted expression after germ line transmission, this suggests that during gametogenesis the decision is made between methylation (paternal inheritance) and the unusual chromatin organization (maternal inheritance). When factors required to methylate this region are available (during spermatogenesis), the upstream element becomes methylated, and this would not allow formation of the unusual chromatin organization to occur. In contrast, nonhistone factors that promote an unusual chromatin organization might be available during oogenesis, and once such organization is achieved, this would prevent methylation. It should be noted that the differential DNase I sensitivity in the H19 upstream element is very similar to the situation in the imprinted U2afl-rsl gene. In this paternally expressed mouse gene, two prominent DNase I HS sites are constitutively present in the 5′ untranslated region of the unmethylated paternal gene only (16, 45). Like in H19, these HS sites map precisely to the sequence element that corresponds to the core of parental allele-specific DNA methylation (46). In addition, we observed that loss of methylation in U2afl-rsl (in ES cells) is associated with gain of DNase I hypersensitivity (22). This suggests that in this imprinted mouse gene also, DNA methylation and unusual chromatin organization are mutually exclusive and reflect alternate epigenetic states. However, the validity of such a model would require testing in germ cells or in primordial germ cell lines.
The similarities between the H19 upstream region and the U2afl-rsl gene lead to the question whether differentially methylated regions within other imprinted genes (41) have a similar parental allele-specific chromatin configuration, characterized by constitutive DNase I hypersensitivity on the unmethylated allele. It might be that this holds true only for imprinting regulatory elements, since we did not detect any structural differences in the H19 gene itself. In addition, in all the regions that have been studied in the neighboring Igf2 gene, both parental alleles were equally sensitive to DNase I (15, 43).
Chromatin regulatory function of the H19 upstream element.
It is not known which nonhistone proteins are associated with the H19 upstream element, and our assays do not permit us to pinpoint the precise DNA sequences to which factors are bound. It can be expected, however, that some of these factors are ubiquitously expressed, since five of the maternal DNase I HS sites were detected in all fetal and adult tissues analyzed. In addition, some of the nonhistone proteins associated with this region could be evolutionarily conserved, since this imprinting element can silence adjacent reporter genes in transgenic flies. In particular, it has been demonstrated that a 1.2-kb sequence 5′ to the G-rich repeat region is responsible for this silencing effect (34). Interestingly, precisely within these sequences we mapped the two most prominent constitutive DNase I HS sites (at 2.25 and 2.75 kb upstream of the H19 transcription initiation site), in close vicinity to two MNase sensitive sites separated by some 500 bp (Fig. 7).
Little is known about mammalian boundary elements-insulators, and these regulatory elements have been characterized in greater detail in nonmammalian species. It has been shown that they are associated with constitutive HS sites and establish higher-order domains of chromatin that affect the interaction between enhancers and promoters (9, 20). For example, the “specialized chromatin structures” at the boundaries of the Drosophila hsp70 genes, which are associated with multiple DNase I HS sites, can insulate the white gene from both positive and negative chromosomal position effects and can insulate the hsp70 promoter from activation by enhancers (29, 55). Specific peptides (BEAF-32A and BEAF-32B) are constitutively bound to these elements, and immunostaining localizes these factors to interband regions and puffs of polytene chromosomes, suggesting a role in the organization of chromosomal domains (25, 60). Another well-characterized insulator is the Drosophila retrotransposon element gypsy, which inhibits the interaction of distal enhancers with the transcription complex but does not affect the action of enhancers positioned proximally (19). The “suppressor of Hairy-wing” and “modifier of mdg4” proteins have been characterized as components of the gypsy insulator; the latter confers on the insulator the ability to unidirectionally affect enhancer function (19, 35). Although these fly insulators are not homologous to the H19 upstream element, sequence comparison does not exclude mechanistic similarities or common structural features, such as an involvement of short repeat sequences (found both in the gypsy insulator [35] and in the H19 upstream element [52, 53]).
Apart from their possible role in insulating the Igf2 gene from enhancers downstream of the H19 gene, a key question that remains is why the constitutive HS sites upstream of H19 (and in the 5′ untranslated region of U2afl-rsl) are present only on the unmethylated maternal allele. Do associated factors not bind to methylated DNA (28), or are HS sites established in female germ cells and stably maintained throughout development (6)? The consistent inverse correlation between methylation and DNase I hypersensitivity brings about the additional possibility that factors bound to the HS sites are involved in keeping this allele unmethylated throughout development. These are intriguing questions, and future research should address the likely interrelationship between the unusual, maternal chromatin organization characterized by DNase I hypersensitivity and paternal DNA methylation in this boundary–imprinting-control element.
ACKNOWLEDGMENTS
We thank W. Reik, G. Kelsey, and M. Constância for helpful discussions and careful reading of the article, S. M. Tilghman for providing the H19 deletion mice, M. S. Bartolomei for providing the H19 upstream SacI fragment and for communicating results prior to publication, and M. A. Surani for providing the H19 gene fragment.
This work was supported by the Ministry of Agriculture, Fisheries and Food (to R.F. and W. Reik), the Royal Society (to R.F.), and The Babraham Institute (R.F. is a Babraham Research Fellow).
S.K. and A.A. contributed equally to this work.
REFERENCES
- 1.Ainscough J F, Koide T, Tada M, Barton S, Surani M A. Imprinting of Igf2 and H19 from a 130 kb YAC transgene. Development. 1997;124:3621–3632. doi: 10.1242/dev.124.18.3621. [DOI] [PubMed] [Google Scholar]
- 2.Allen N D, Barton S C, Hilton K, Norris M L, Surani M A. A functional analysis of imprinting in parthenogenetic embryonic stem cells. Development. 1994;120:1473–1482. doi: 10.1242/dev.120.6.1473. [DOI] [PubMed] [Google Scholar]
- 3.Bartolomei M S, Zemel S, Tilghman S M. Parental imprinting of the mouse H19 gene. Nature. 1991;351:153–155. doi: 10.1038/351153a0. [DOI] [PubMed] [Google Scholar]
- 4.Bartolomei M S, Webber A L, Brunkow M E, Tilghman S M. Epigenetic mechanisms underlying the imprinting of the mouse H19 gene. Genes Dev. 1993;7:1663–1673. doi: 10.1101/gad.7.9.1663. [DOI] [PubMed] [Google Scholar]
- 5.Beechey C V, Cattanach B M. Genetic and physical imprinting map of the mouse. United Kingdom: Mammalian Genetics Unit, Harwell; 1998. http://www.mgu.har.mrc.ac.uk . World Wide Web ( http://www.mgu.har.mrc.ac.uk) ) [Google Scholar]
- 6.Boyes J, Felsenfeld G. Tissue-specific factors additively increase the probability of the all-or-none formation of a hypersensitive site. EMBO J. 1996;15:2496–2507. [PMC free article] [PubMed] [Google Scholar]
- 7.Brandeis M, Kafri T, Ariel M, Chaillet J R, McCarrey J, Razin A, Cedar H. The ontogeny of allele-specific methylation associated with imprinted genes in the mouse. EMBO J. 1993;12:3669–3677. doi: 10.1002/j.1460-2075.1993.tb06041.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cattanach B M, Kirk M. Differential activity of maternally and paternal derived chromosome regions in mice. Nature. 1985;315:496–498. doi: 10.1038/315496a0. [DOI] [PubMed] [Google Scholar]
- 9.Chung J H, Whiteley M, Felsenfeld G. A 5′ element of the chicken beta-globin domain serves as an insulator in human erythroid cells and protects against position effect in Drosophila. Cell. 1993;74:505–514. doi: 10.1016/0092-8674(93)80052-g. [DOI] [PubMed] [Google Scholar]
- 10.Church G M, Gilbert W. Genomic sequencing. Proc Natl Acad Sci USA. 1984;81:1991–1995. doi: 10.1073/pnas.81.7.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dean W, Bowden L, Aitchison A, Klose J, Moore T, Meneses J J, Reik W, Feil R. Altered imprinted gene methylation and expression in completely ES cell-derived mouse fetuses: association with aberrant phenotypes. Development. 1998;125:2273–2282. doi: 10.1242/dev.125.12.2273. [DOI] [PubMed] [Google Scholar]
- 12.Elgin S C R. The formation and function of DNase-I hypersensitive sites in the process of gene activation. J Biol Chem. 1988;263:19259–19262. [PubMed] [Google Scholar]
- 13.Elson D A, Bartolomei M S. A 5′ differentially methylated sequence and the 3′-flanking region are necessary for H19 transgene imprinting. Mol Cell Biol. 1997;17:309–317. doi: 10.1128/mcb.17.1.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Feil R, Walter J, Allen N D, Reik W. Developmental control of allelic methylation in the imprinted mouse Igf2 and H19 genes. Development. 1994;120:2933–2943. doi: 10.1242/dev.120.10.2933. [DOI] [PubMed] [Google Scholar]
- 15.Feil R, Handel M A, Allen N D, Reik W. Chromatin structure and imprinting: developmental control of DNase-I sensitivity in the mouse insulin-like growth factor 2 gene. Dev Genet. 1995;17:240–252. doi: 10.1002/dvg.1020170309. [DOI] [PubMed] [Google Scholar]
- 16.Feil R, Boyano M D, Allen N D, Kelsey G. Parental chromosome-specific chromatin conformation in the imprinted U2afl-rsl gene in the mouse. J Biol Chem. 1997;272:20893–20900. doi: 10.1074/jbc.272.33.20893. [DOI] [PubMed] [Google Scholar]
- 17.Feil R, Kelsey G. Insights from model systems. Genomic imprinting: a chromatin connection. Am J Hum Genet. 1997;61:1213–1219. doi: 10.1086/301655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ferguson-Smith A C, Sasaki H, Cattanach B M, Surani M A. Parental-origin-specific epigenetic modification of the mouse H19 gene. Nature. 1993;362:751–755. doi: 10.1038/362751a0. [DOI] [PubMed] [Google Scholar]
- 19.Gdula D A, Gerasimova T I, Corces V G. Genetic and molecular analysis of the gypsy chromatin insulator of Drosophila. Proc Natl Acad Sci USA. 1996;93:9378–9383. doi: 10.1073/pnas.93.18.9378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gerasimova T I, Corces V G. Boundary and insulator elements in chromosomes. Curr Opin Genet Dev. 1996;6:185–192. doi: 10.1016/s0959-437x(96)80049-9. [DOI] [PubMed] [Google Scholar]
- 21.Greally J M, Starr D J, Hwang S, Song L, Jaarola M, Zemel S. The mouse H19 locus mediates a transition between imprinted and non-imprinted DNA replication patterns. Hum Mol Genet. 1998;7:91–96. doi: 10.1093/hmg/7.1.91. [DOI] [PubMed] [Google Scholar]
- 22.Gregory, R., and R. Feil. Unpublished data.
- 23.Gross D S, Garrard W T. Nuclease hypersensitive sites in chromatin. Annu Rev Biochem. 1988;57:159–197. doi: 10.1146/annurev.bi.57.070188.001111. [DOI] [PubMed] [Google Scholar]
- 24.Hark A T, Tilghman S M. Chromatin conformation of the H19 epigenetic mark. Hum Mol Genet. 1998;7:1979–1985. doi: 10.1093/hmg/7.12.1979. [DOI] [PubMed] [Google Scholar]
- 25.Hart C M, Zhao K, Laemmli U K. The scs′ boundary element: characterization of boundary element-associated factors. Mol Cell Biol. 1997;17:999–1009. doi: 10.1128/mcb.17.2.999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ishihara K, Kato R, Furuumi H, Zubair M, Sasaki H. Sequence of a 42-kb mouse region containing the imprinted H19 locus: identification of a novel muscle-specific transcription unit showing biallelic expression. Mamm Genome. 1998;9:775–777. doi: 10.1007/s003359900863. [DOI] [PubMed] [Google Scholar]
- 27.Jones B K, Levorse J M, Tilghman S M. Igf2 imprinting does not require its own DNA methylation or H19 RNA. Genes Dev. 1998;12:2200–2207. doi: 10.1101/gad.12.14.2200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kass S U, Goddard J P, Adams R L P. Inactive chromatin spreads from a focus of methylation. Mol Cell Biol. 1993;13:7372–7379. doi: 10.1128/mcb.13.12.7372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kellum R, Schedl P. A position effect assay for boundaries of higher order chromatin structure. Cell. 1991;64:941–950. doi: 10.1016/0092-8674(91)90318-s. [DOI] [PubMed] [Google Scholar]
- 30.Koide T, Ainscough J, Wijgerde M, Surani M A. Comparative analysis of Igf2/H19 imprinting domain: identification of a highly conserved intergenic DNase I hypersensitive region. Genomics. 1994;24:1–8. doi: 10.1006/geno.1994.1574. [DOI] [PubMed] [Google Scholar]
- 31.Leighton P A, Ingram R S, Eggenschwiler J, Efstratiadis A, Tilghman S M. Disruption of imprinting caused by deletion of the H19 gene region in mice. Nature. 1995;375:34–39. doi: 10.1038/375034a0. [DOI] [PubMed] [Google Scholar]
- 32.Leighton P A, Saam J R, Ingram R S, Stewart C L, Tilghman S M. An enhancer deletion affects both H19 and Igf2 expression. Genes Dev. 1995;9:2079–2089. doi: 10.1101/gad.9.17.2079. [DOI] [PubMed] [Google Scholar]
- 33.Li E, Beard C, Jaenisch R. Role for DNA methylation in genomic imprinting. Nature. 1993;366:362–365. doi: 10.1038/366362a0. [DOI] [PubMed] [Google Scholar]
- 34.Lyko F, Brenton J D, Surani M A, Paro R. An imprinting element from the mouse H19 locus functions as a silencer in Drosophila. Nat Genet. 1997;16:171–173. doi: 10.1038/ng0697-171. [DOI] [PubMed] [Google Scholar]
- 35.Mazo A M, Mizrokhi L J, Karavanov A A, Sedkov Y A, Krichevskaja A A, Ilyin Y V. Suppression in Drosophila: su(Hw) and Su(f) gene products interact with a region of gypsy (mdg4) regulating its transcriptional activity. EMBO J. 1989;8:903–911. doi: 10.1002/j.1460-2075.1989.tb03451.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.McGhee J D, Wood W I, Dolan M, Engel J D, Felsenfeld G. A 200 base pair region at the 5′ end of the chicken adult beta-globin gene is accessible to nuclease digestion. Cell. 1981;27:45–55. doi: 10.1016/0092-8674(81)90359-7. [DOI] [PubMed] [Google Scholar]
- 37.McGrath J, Solter D. Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell. 1984;37:179–183. doi: 10.1016/0092-8674(84)90313-1. [DOI] [PubMed] [Google Scholar]
- 38.Olek A, Walter J. The pre-implantation ontogeny of the H19 methylation imprint. Nat Genet. 1998;17:275–276. doi: 10.1038/ng1197-275. [DOI] [PubMed] [Google Scholar]
- 39.Pachnis V, Brannan C I, Tilghman S M. The structure and expression of a novel gene activated in early mouse embryogenesis. EMBO J. 1988;7:673–681. doi: 10.1002/j.1460-2075.1988.tb02862.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Razin A, Cedar H. DNA methylation and genomic imprinting. Cell. 1994;77:473–476. doi: 10.1016/0092-8674(94)90208-9. [DOI] [PubMed] [Google Scholar]
- 41.Reik W, Walter J. Imprinting mechanisms in mammals. Curr Opin Genet Dev. 1998;8:154–164. doi: 10.1016/s0959-437x(98)80136-6. [DOI] [PubMed] [Google Scholar]
- 42.Ripoche M-A, Kress C, Poirier F, Dandolo L. Deletion of the H19 transcription unit reveals the existence of a putative imprinting control element. Genes Dev. 1997;11:1596–1604. doi: 10.1101/gad.11.12.1596. [DOI] [PubMed] [Google Scholar]
- 43.Sasaki H, Jones P A, Chaillet J R, Ferguson-Smith A C, Barton S C, Reik W, Surani M A. Parental imprinting: potentially active chromatin of the repressed maternal allele of the mouse insulin-like growth factor II (Igf2) gene. Genes Dev. 1992;6:1843–1856. doi: 10.1101/gad.6.10.1843. [DOI] [PubMed] [Google Scholar]
- 44.Sasaki H, Ferguson-Smith A C, Shum A S W, Barton S C, Surani M A. Temporal and spatial regulation of H19 imprinting in normal and uniparental mouse embryos. Development. 1995;121:4195–4202. doi: 10.1242/dev.121.12.4195. [DOI] [PubMed] [Google Scholar]
- 45.Shibata H, Yoshino K, Sunahara S, Gondo Y, Katsuki M, Ueda T, Kamiya M, Muramatsu M, Murakami Y, Kalcheva I, Plass C, Chapman V M, Hayashizaki Y. Inactive allele-specific methylation and chromatin structure of the imprinted gene U2afl-rsl on mouse chromosome 11. Genomics. 1996;35:248–252. doi: 10.1006/geno.1996.0348. [DOI] [PubMed] [Google Scholar]
- 46.Shibata H, Ueda T, Kamiya M, Yoshiki A, Kusakabe M, Plass C, Held W A, Sunahara S, Katsuki M, Muramatsu M, Hayashizaki Y. An oocyte-specific methylation imprint center in the mouse U2afbp-rs/U2afl-rsl gene marks the establishment of allele-specific methylation during preimplantation development. Genomics. 1997;44:171–178. doi: 10.1006/geno.1997.4877. [DOI] [PubMed] [Google Scholar]
- 47.Surani M A. Imprinting and the initiation of gene silencing in the germ line. Cell. 1998;93:309–312. doi: 10.1016/s0092-8674(00)81156-3. [DOI] [PubMed] [Google Scholar]
- 48.Surani M A H, Barton S C, Norris M L. Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature. 1984;308:548–550. doi: 10.1038/308548a0. [DOI] [PubMed] [Google Scholar]
- 49.Svensson K, Walsh C, Fundele R, Ohlsson R. H19 is imprinted in the choroid plexus and the leptomeninges of the mouse foetus. Mech Dev. 1995;51:31–37. doi: 10.1016/0925-4773(94)00345-n. [DOI] [PubMed] [Google Scholar]
- 50.Szabó P E, Pfeifer G P, Mann J R. Characterization of novel parent-specific epigenetic modifications upstream of the imprinted mouse H19 gene. Mol Cell Biol. 1998;18:6767–6776. doi: 10.1128/mcb.18.11.6767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Thorvaldsen J L, Duran K L, Bartolomei M S. Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2. Genes Dev. 1998;12:3693–3702. doi: 10.1101/gad.12.23.3693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Tremblay K D, Saam J R, Ingram R S, Tilghman S M, Bartolomei M S. A paternal-specific methylation imprint marks the alleles of the mouse H19 gene. Nat Genet. 1995;9:407–413. doi: 10.1038/ng0495-407. [DOI] [PubMed] [Google Scholar]
- 53.Tremblay K D, Duran K L, Bartolomei M S. A 5′ 2-Kilobase-pair region of the imprinted mouse H19 gene exhibits paternal methylation throughout development. Mol Cell Biol. 1997;17:4322–4329. doi: 10.1128/mcb.17.8.4322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Urieli-Shoval S, Gruenbaum Y, Sedat J, Razin A. The absence of detectable methylated bases in Drosophila melanogaster DNA. FEBS Lett. 1982;146:148–152. doi: 10.1016/0014-5793(82)80723-0. [DOI] [PubMed] [Google Scholar]
- 55.Vazquez J, Schedl P. Sequences required for enhancer blocking activity are located within two nuclease-hypersensitive regions. EMBO J. 1994;13:5984–5993. doi: 10.1002/j.1460-2075.1994.tb06944.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Warnecke P M, Biniszkiewicz D, Jaenisch R, Frommer M, Clark S J. Sequence-specific methylation of the mouse H19 gene in embryonic stem cells deficient in the Dnmt-1 gene. Dev Genet. 1998;22:111–121. doi: 10.1002/(SICI)1520-6408(1998)22:2<111::AID-DVG1>3.0.CO;2-9. [DOI] [PubMed] [Google Scholar]
- 57.Warnecke P M, Mann J R, Frommer M, Clark S J. Bisulfite sequencing in preimplantation embryos: DNA methylation profile of the upstream region of the mouse imprinted H19 gene. Genomics. 1998;51:182–190. doi: 10.1006/geno.1998.5371. [DOI] [PubMed] [Google Scholar]
- 58.Webber A L, Ingram R S, Levorse J M, Tilghman S M. Location of enhancers is essential for the imprinting of H19 and Igf2 genes. Nature. 1998;391:711–715. doi: 10.1038/35655. [DOI] [PubMed] [Google Scholar]
- 59.Webber A L, Tilghman S M. The absence of enhancer competition between Igf2 and H19 following transfer into differentiated cells. Mol Cell Biol. 1998;18:1903–1910. doi: 10.1128/mcb.18.4.1903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhao K, Hart C M, Laemmli U K. Visualization of chromosomal domains with boundary element-associated factor BEAF-32. Cell. 1995;81:879–889. doi: 10.1016/0092-8674(95)90008-x. [DOI] [PubMed] [Google Scholar]