Abstract
The Gal3, Gal80, and Gal4 proteins of Saccharomyces cerevisiae comprise a signal transducer that governs the galactose-inducible Gal4p-mediated transcription activation of GAL regulon genes. In the absence of galactose, Gal80p binds to Gal4p and prohibits Gal4p from activating transcription, whereas in the presence of galactose, Gal3p binds to Gal80p and relieves its inhibition of Gal4p. We have found that immunoprecipitation of full-length Gal4p from yeast extracts coprecipitates less Gal80p in the presence than in the absence of Gal3p, galactose, and ATP. We have also found that retention of Gal80p by GSTG4AD (amino acids [aa] 768 to 881) is markedly reduced in the presence compared to the absence of Gal3p, galactose, and ATP. Consistent with these in vitro results, an in vivo two-hybrid genetic interaction between Gal80p and Gal4p (aa 768 to 881) was shown to be weaker in the presence than in the absence of Gal3p and galactose. These compiled results indicate that the binding of Gal3p to Gal80p results in destabilization of a Gal80p-Gal4p complex. The destabilization was markedly higher for complexes consisting of G4AD (aa 768 to 881) than for full-length Gal4p, suggesting that Gal80p relocated to a second site on full-length Gal4p. Congruent with the idea of a second site, we discovered a two-hybrid genetic interaction involving Gal80p and the region of Gal4p encompassing aa 225 to 797, a region of Gal4p linearly remote from the previously recognized Gal80p binding peptide within Gal4p aa 768 to 881.
Signaling systems controlling the activation and repression states of genes often include interactions among multiple proteins. One well-studied example is the galactose-responsive system that governs the expression state of the galactose pathway genes (GAL genes) of the yeast Saccharomyces cerevisiae and the closely related milk yeast Kluyveromyces lactis (16, 52, 57, 62, 63, 74, 78, 79, 81). Galactose signaling operates through a three-component switch consisting of the Gal3 (or Gal1), Gal80, and Gal4 proteins. In the presence of galactose, the GAL genetic switch operates to elicit activation of the GAL genes, and in the absence of galactose, the switch operates to inhibit activation of the GAL genes (32, 40, 56).
The Gal4 protein component of the GAL genetic switch exhibits both site-specific DNA binding and transcription activation, and these activities reside in separable protein domains (11, 36, 43). Amino acids 1 to 65 of S. cerevisiae Gal4p specify binding to a 17-bp DNA site (UASGAL) upstream of the GAL genes (10, 27, 45). Amino acids within two other distinct regions, amino acids (aa) 148 to 238 (region I), and aa 768 to 881 (region II), specify weak and strong transcription activation functions, respectively (33, 43). Various mutant derivatives of activation region II exhibit a striking correlation between their relative transcription activation potentials and their relative affinities for yTFIIB and yTBP proteins (75), giving credence to the view that Gal4p activates transcription through interactions which facilitate the recruitment of RNA polymerase II (4, 37).
Within transcription activation region II of Gal4p is a shorter region, defined by aa 850 to 874, that binds to the Gal80 protein, an inhibitor of Gal4p-mediated transcription activation (35, 41, 42, 51, 69, 77). The determinants of Gal4p for Gal80p binding and inhibition are highly interspersed with the determinants for the region II transcriptional activation function, and yet they are mutationally resolvable (3, 39, 64). Consistent with this interspersion, Gal80p binding prohibits subsequent interaction of Gal4p with TATA binding protein (TBP) (3). Thus, Gal80p binding to a rather short peptide sequence within Gal4p plays a key role in the mechanism by which Gal80p inhibits the interactions of Gal4p essential to the recruitment of RNA polymerase II.
Addition of galactose to wild-type yeast cells overcomes the Gal80p inhibition of Gal4p, allowing Gal4p to activate transcription. This galactose-induced state requires either the Gal3p or the highly homologous Gal1p (6, 7, 12, 19, 47, 49, 70). It is quite well established that the presence of galactose and ATP promotes a complex of Gal3p (or Gal1p) and Gal80p in vitro (68, 76, 79), and the capacity of Gal3p to bind to Gal80p is clearly linked to Gal4p-mediated GAL gene activation in vivo (8) and in vitro (54). How Gal3p binding to Gal80p triggers galactose-induced Gal4p-mediated transcription activation has not been determined. Transformation of the Gal80p-Gal4p complex in response to galactose had been proposed earlier by Leuther and Johnston on the basis of the persistence of a Gal80p-Gal4p two-hybrid genetic interaction following galactose addition to yeast cells (38). Platt and Reece recently reported the galactose-responsive in vitro formation of a ternary complex comprising Gal4p, Gal80p and Gal3pC-322, a mutant form of Gal3p (54). Thus, the evidence to date supports a model wherein a conformational change in a Gal80p-Gal4p complex arises in response to galactose by way of Gal3p-Gal80p interaction. However, no evidence has yet been established for any type of physical change in a Gal80p-Gal4p complex in response to galactose and Gal3p.
Our efforts to understand the Gal3p-Gal80p-Gal4p transcription switch mechanism have led us to examine more closely how the Gal3p-Gal80p interaction might alter the Gal80p-Gal4p interaction. The effect of Gal3p on the Gal80p-Gal4p interaction was assessed by three independent experimental approaches. Based on these results, we propose that Gal3p binding to Gal80p promotes the dissociation of Gal80p from its site of inhibition within the principal activation region of Gal4p. The association of Gal80p with another site within the Gal4p sequence from aa 225 to 797 is suggested on the basis of two-hybrid genetic interaction experiments. These results have implications for both induction and deinduction.
MATERIALS AND METHODS
Yeast strains, media, culture conditions, and genetic techniques.
The yeast strains used in this study are listed in Table 1. Yeasts were transformed either by the one-step procedure of Chen et al. (15) or by the high-efficiency method of Gietz et al. (25). All strains were generated by standard genetic techniques. Sc338 was derived from Sc252 (34) by curing of the 2μm plasmid (21, 22). Sc800, Sc817, and Sc818 were derived from Y187 (29) as follows. The GAL3 gene in Y187 was replaced with a gal3Δ::Kanr cassette by a one-step gene disruption method (28). The gal3Δ::Kanr cassette was made by PCR with oligonucleotide primers ALOK45 and ALOK46 (see Table 3) and pUG6 DNA containing the Kanr cassette (28). The resulting strain, Sc798, was subjected to 5-fluoroorotic acid selection (9) to yield the ura3 derivative, Sc800. The GAL1 gene of Sc800 was replaced with a gal1Δ-1740::URA3 (8) disruption cassette to yield Sc818. Sc804 is a lacZ mutant of Sc818. Strain Sc817 is a 5-fluoroorotic acid-selected ura3 derivative of Sc804. Sc813 was derived from yeast strain L40 (5) by replacement of the chromosomal GAL4 gene of L40 with gal4Δ::Kanr, a cassette constructed by PCR with SAM01 and SAM02 oligonucleotides (see Table 3) and pUG6 DNA. Sc830 was derived from YT6::6lexOP (67) by replacement of the chromosomal GAL3 with the gal3Δ::Kanr cassette as described above.
TABLE 1.
Yeast strains used in this study
| Strain | Genotype | Reference or source |
|---|---|---|
| Sc338 | MATa leu2-3,112 ura3-52 ade1 ile MEL1 cir-0 | This study |
| Y187 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met URA3::GAL1-lacZ | 29 |
| Sc798 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met URA3::GAL1-lacZ gal3Δ::loxP-KanMX-loxP | This study |
| Sc800 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met ura3::GAL1-lacZ gal3Δ::loxP-KanMX-loxP | This study |
| Sc804 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met ura3::GAL1-lacZ gal3Δ::loxP-KanMX-loxP gal1Δ::URA3 | This study |
| Sc817 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met gal3Δ::loxP-KanMX-loxP gal1Δ::ura3 | This study |
| Sc818 | MATα gal4 gal80 his3 trp1-901 ade2-101 ura3-52 leu2-3,112 met ura3::GAL1-lacZ gal3Δ::loxP-KanMX-loxP gal1Δ::ura3 | This study |
| Sc813 | MATa his3 Δ200 trp1-901 leu2-3,112 ade2 LYS2::(lexAop)4-HIS3 URA3::(lexAop)8-lacZ gal4Δ::loxP-KanMX-loxP | This study |
| Sc830 | MATα gal4 gal80 his3 trp1 ade ara1 ura3 leu2 met URA3::(lexAop)6-lacZ gal3Δ::loxP-KanMX-loxP | This study |
TABLE 3.
Sequences of the oligonucleotides used in this study
| Oligonucleotide | Sequence |
|---|---|
| G3 | 5′-CGGATCCGCATGGACTACAACAAGAGATCT-3′ |
| G4 | 5′-CGGATCCGCTTAACTATAATGCGAGAT-3′ |
| ALOK45 | 5′-GAATACAAACGTTCCAATATTCAGTTCTCCGGTCAGACAGGTCGACAACCCTTAAT-3′ |
| ALOK46 | 5′-AGTACCCAAGGCAGGCTTCGAAACTATAATTGCGTCGTGGATCTGATATCACCTA-3′ |
| ALOK49 | 5′-CCCCGTTGTTGTCTCACCATAT-3′ |
| ALOK52 | 5′-ACCATTACAATGGATGACGTCTATAACTATCTATTCG-3′ |
| ALOK53 | 5′-ATCCATTGTAATGGTATTAAACATCCC-3′ |
| BCO2 | 5′-GACAAGCCGACAACCTTGATTGGAG-3′ |
| SAM01 | 5′-CAGATTTTCAGCTTCATCTCCAGATTGTGTCTACCAGGTCGACAACCCTTAAT-3′ |
| SAM02 | 5′-ACTTGCGGGGTTTTTCAGTATCTACGATTCATTTTAGTGGATCTGATATCACCTA-3′ |
Solid and liquid growth media for yeast were prepared essentially as described by Rose et al. (58). Yeast cells were shake-cultured in YEP (0.5% yeast extract, 1.0% Bacto Peptone) or synthetic complete (SC) medium lacking specific supplements as required for plasmid selection. The carbon sources used were glucose (2.0 or 0.05% [wt/vol]), galactose (2.0% [wt/vol]), glycerol (3.0% [vol/vol]), and lactic acid (2.0% [vol/vol]). The pH of the lactic acid (Fisher, A162-1) was adjusted to 5.7 with KOH.
Yeasts used as a source of extract for in vitro protein interaction assays were cultured as follows. Sc800 carrying either pMEGA3-Δ4 or pMEGA3-Δ4Δ3, Sc817 carrying pMEGA3-Δ4Δ80, and Sc338 carrying either pMEGA3 or pMEGA3-Δ3 were first grown to saturation in 5 ml of SC medium lacking uracil and containing 2% glucose (see below and Table 2 for plasmid descriptions). A 1-ml aliquot was then transferred to 1 liter of SC medium lacking leucine and containing 2% glycerol, 2% lactic acid, and 0.05% glucose, and the cells were shake-cultured at 30°C to a density corresponding to an optical density at 600 nm (OD600) of 1.0 to 1.2. Sc800 with pYGSTG4AD was cultured to an OD600 of 0.8 to 1.0 in 1 liter of SC medium lacking leucine and containing 2% glucose.
TABLE 2.
Plasmids used in this study
| Plasmid | Descriptiona | Reference or source |
|---|---|---|
| pRS415 | CEN6 ARS LEU2 | 65 |
| pRS414 | CEN6 ARS TRP1 | 65 |
| pVP16 | 2μm LEU2 ADH1pro-VP16 (codons 413–490) | 5 |
| pBDG80 | 2μm HIS3 ADH1 pro-GAL4 DNA binding domain (codons 1–147)-GAL80 | This study |
| pBDG80S2 | 2μm HIS3 ADH1 pro-GAL4 DNA binding domain (codons 1–147)-GAL80S-2 | This study |
| PAKS68 | CEN ARS1 LEU2 ADH1 pro-GAL4AD (codons 768–881) | This study |
| pG4ADVP16 | CEN ARS1 LEU2 ADH1 pro-GAL4AD (codons 768–881)-VP16 (codons 413–490) | This study |
| pG4AD*VP16 | CEN ARS1 LEU2 ADH1 pro-GAL4ADThr859-Ile-VP16 (codons 413–490) | This study |
| pG3VP16 | CEN ARS1 TRP1 GAL3-VP16 (codons 413–490) | This study |
| pTEB16 | CEN ARS1 TRP1 GAL3 | 8 |
| pMPW66 | CEN ARS1 TRP1 ADH2 pro-GAL3 | 8 |
| pYGSTG4AD | 2μm LEU2 ADH1 pro-GAL4 (codons 768–881)-GST | This study |
| pEGSTG4AD | GST-GAL4AD (codons 768–881) | This study |
| pMEGA3 | 2μm URA3 leu2-d GAL4901 GAL80 GAL3 | This study |
| pMEGA3-Δ4 | 2μm URA3 leu2-d GAL80 GAL3 | This study |
| pMEGA3-Δ4Δ3 | 2μm URA3 leu2-d GAL80 | This study |
| pMEGA3-Δ4Δ80 | 2μm URA3 leu2-d GAL3 | This study |
| pMEGA3-Δ3 | 2μm URA3 leu2-d GAL4901 GAL80 | This study |
| pVP16G80 | 2μm LEU2 ADH1 pro-VP16 (codons 413–490)-GAL80 | This study |
| pAKS87 | CEN ARS1 TRP1 ADH1 pro-VP16 (codons 413–490)-GAL80 | This study |
| pLexAG4-225-797 | 2μm HIS3 ADH1 pro-lexA (codons 1–202)-GAL4 (codons 225–797) | This study |
| pLexAG4-534-797 | 2μm HIS3 ADH1 pro-lexA (codons 1–202)-GAL4 (codons 534–797) | This study |
| pLexAG4-225-547 | HIS3 ADH1 pro-lexA (codons 1–202)-GAL4 (codons 225–534) | This study |
| pLEUG4 | CEN6 ARS LEU2 GAL4 | This study |
| pSB32-Gal4mCla | CEN ARS LEU2 GAL4-Δ codons 148–728 | 18 |
| pSB32-miniGal4-2 | CEN ARS LEU2 GAL4-Δ codons 210–716 | 18 |
pro indicates promoter; unless it is indicated, all promoters are native.
For two-hybrid analyses, Sc830 and Sc818, each carrying three different plasmids, and Y187, Sc798, and Sc813, each carrying two different plasmids, were cultured at 30°C in 5 ml of selective SC medium containing 2% glucose. At a culture OD600 of 1.0, a 200-μl aliquot was transferred to the appropriate SC medium containing either 2% glycerol, 2% lactic acid, and 0.05% glucose (noninduced) or 2% glycerol, 2% lactic acid, 2% galactose, and 0.05% glucose (induced). The cells were shake-cultured at 30°C to a density corresponding to an OD600 of 0.35 to 0.45.
Yeast cell extracts.
Yeast whole-cell extracts for the two-hybrid β-galactosidase assay were prepared by vortexing with glass beads (0.45 mm in diameter). The breakage buffer consisted of 100 mM Tris (pH 8.0), 0.2 mM dithiothreitol (DTT) and 10 mM phenylmethylsulfonyl fluoride. Extracts were stored at −80°C until use. The protein concentrations of the extracts were determined by a Bio-Rad assay.
Yeast cell extracts for the in vitro protein interaction assays in Fig. 3 were prepared by vortexing with glass beads. Prior to the addition of glass beads, the cell pellet from 100 ml of culture was resuspended in 425 μl of buffer A (10 mM HEPES [pH 7.5], 200 mM NaCl, 0.5% Triton X-100, 10 mM NaF, 0.2 mM Na3VO4, 0.5 mM EDTA, 2 mM DTT, 1 μg of leupeptin per ml, 1 μg of pepstatin per ml, 20 μg of aprotinin per ml). Yeast cell extracts for the in vitro protein interaction assays in Fig. 4 to 7 were prepared by a French pressure cell procedure. A 15-ml volume of buffer A containing the resuspended cell pellet from a 6-liter culture was loaded into the pressure cell, frozen for 30 min in a mixture of dry ice and ethanol, and subjected to 1,500 lb/in2 as described by Verner and Weber (72). Extracts were stored at −80°C. Protein concentrations were estimated by the procedure of Peterson (53).
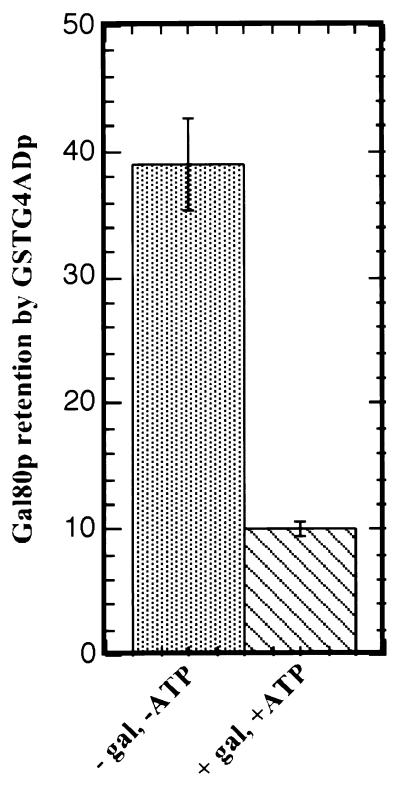
FIG. 3.
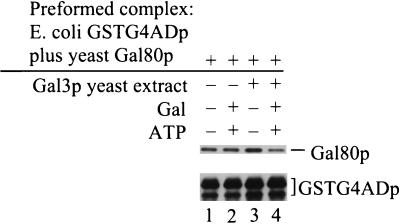
The presence of Gal3p, galactose, and ATP reduces Gal80p retention by GSTG4ADp. (A) Extracts were prepared from Gly-Lac-grown cultures of yeast strain Sc800 (gal4Δ gal80Δ gal3Δ) carrying pYGSTG4AD, pMEGA3-Δ4Δ3 (Gal80p), or pMEGA3-Δ4 (Gal80p and Gal3p). The GT-Sepharose-bound GSTG4ADp was incubated for 2 h at 4°C with an aliquot of yeast extract (1,000 μg of protein) containing either Gal80p (lanes 1 and 2) or Gal80p plus Gal3p (lanes 3 and 4) in the absence (−) or presence (+) of galactose and ATP. GT-Sepharose-bound proteins were subjected to Western immunoblotting. Blots were probed with rabbit polyclonal anti-Gal80p and anti-GST. (B) E. coli-expressed GSTG4ADp instead of yeast-expressed GSTG4ADp was used together with the same yeast extracts used in panel A. A 70-μl volume of the E. coli extract in combination with 1,000 μg of yeast extract (2×) or 35 μl of the E. coli extract in combination with 500 μg of yeast extract (1×) were mixed with the binding buffer. The binding reactions were carried out in the presence (+) or absence (−) of galactose and ATP. GT-Sepharose-bound proteins were analyzed as above.
FIG. 4.
Incubation of GSTG4ADp-Gal80p complex with Gal3p, galactose, and ATP results in loss of bound Gal80p. The GSTG4ADp-Gal80p complex was first formed on GT-Sepharose beads. This GT-Sepharose-bound GSTG4ADp-Gal80p complex was incubated for 2 h at 4°C with an aliquot of a gal4Δ gal80Δ yeast extract either lacking Gal3p (Sc817) (lanes 1 and 2) or containing Gal3p (Sc817 bearing pMEGA3-Δ4Δ80) (lanes 3 and 4) in the absence (−) or presence (+) of galactose and ATP. GT-Sepharose-bound proteins were analyzed by Western immunoblotting with rabbit polyclonal anti-Gal80p and anti-GST. The amount of GSTG4ADp loaded per lane in lanes 3 and 4 was more than that used per lane in lanes 1 and 2.
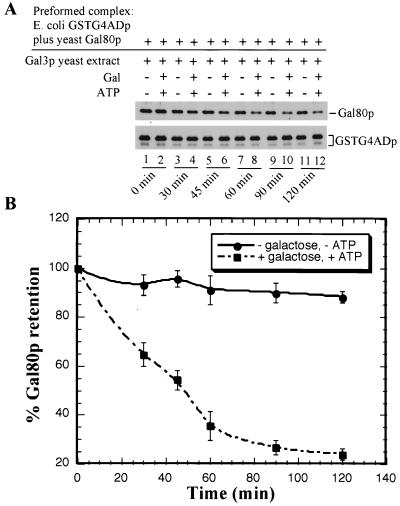
FIG. 7.
Kinetics of loss of Gal80p from GT-Sepharose bound GSTG4ADp-Gal80p complex in the presence of Gal3p, galactose, and ATP. (A) First, GSTG4ADp-Gal80p complex was formed on glutathione-Sepharose beads. The GT-Sepharose-bound GSTG4ADp-Gal80p complex was mixed with binding buffer and gal4Δ gal80Δ yeast extract containing Gal3p (Sc817 bearing pMEGA3-Δ4Δ80) and incubated in the absence (lanes 1, 3, 5, 7, 9, and 11) or presence (lanes 2, 4, 6, 8, 10, and 12) of galactose and ATP. At the times indicated, the Sepharose beads were pelleted and washed, and the bound proteins were fractionated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (8% polyacrylamide) and immunoblotted with anti-Gal80p (top immunoblot) or anti-GST (bottom immunoblot). (B) Densitometric scans were performed to quantify the Gal80p and GSTG4ADp signals on the immunoblot. The OD600 corresponding to the amount of Gal80p remaining bound to GSTG4ADp relative to the amount bound at time zero (100%), per unit GSTG4ADp, is plotted as a function of time. Each data point plotted represents the average of four independent GSTG4AD pull-down experiments. A smooth curve fit for the data points is shown as lines.
Yeast whole-cell extracts used for the immunoprecipitation experiments in Fig. 1 were prepared by the French pressure cell method as above, except that the breakage buffer consisted of 50 mM NaPO4 (pH 7.2), 5 mM EDTA, 50 mM NaF, 1 μg of leupeptin per ml, 1 μg of pepstatin per ml, 20 μg of aprotinin per ml, 0.2 mM Na3VO4, and 1 mM phenylmethylsulfonyl fluoride.
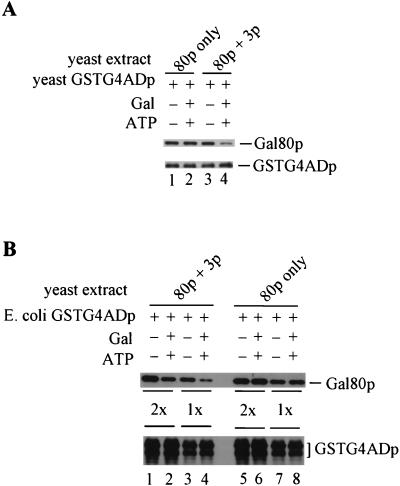
FIG. 1.
Immunoprecipitates of full-length Gal4p show reduced levels of Gal80p in the presence of Gal3p, galactose, and ATP. Protein extracts were prepared from Gly-Lac-grown cultures of yeast strain Sc338 carrying either plasmid pMEGA3 (GAL4901 GAL80 GAL3) or plasmid pMEGA3-Δ3 (GAL4901 GAL80). The anti-901 monoclonal antibody was used to immunoprecipitate 901-tagged Gal4p from yeast extracts comprising 3 mg of proteins. Immunoprecipitations were carried out in the absence (lane 1) and presence (lane 2) of galactose and ATP. Precipitated proteins were analyzed by Western immunoblotting. Gal4p and Gal80p were detected on the same blot by using a combination of rabbit polyclonal anti-Gal4p and anti-Gal80p. (A) Immunoprecipitations were performed on extracts of Sc338 carrying pMEGA3. Due to markedly different intensities of immunologically detected Gal4p901 and Gal80p bands, two different chemiluminescence film exposures (A1 and A2) of the same immunoblot are shown. wt, wild type. (B) Immunoprecipitations were performed on extracts of Sc338 carrying pMEGA3-Δ3.
Bacterial strains, culture conditions, and extracts.
Escherichia coli DH5α was used as bacterial host during plasmid construction. For the production of GSTGal4ADp in E. coli, strain BL21(DE3) (Novagen) was transformed with plasmid pEGSTG4AD and shake-cultured at 37°C in 1 liter of Luria broth containing ampicillin (50 μg/ml) to an OD600 of 0.6. IPTG (isopropyl-β-d-thiogalactopyranoside) was then added to a final concentration of 1 mM, and the cells were shake-cultured for an additional 3 h at 37°C. The cells were harvested by centrifugation at 4,000 × g for 30 min and washed once with buffer (20 mM HEPES [pH 7.5], 200 mM NaCl). The washed cell pellet from 500 ml of culture was resuspended in 50 ml of lysis buffer consisting of 20 mM HEPES (pH 7.5), 200 mM NaCl, 10 mM β-mercaptoethanol, and 20 μg of aprotinin per ml. Cell lysates were prepared by sonication and centrifuged at 10,000 × g for 15 min. The supernatant, containing GSTG4ADp, was frozen on dry ice and stored at −85°C.
Plasmid descriptions and constructions.
The plasmids used in this study are listed in Table 2. The oligonucleotides used in the construction of the plasmids are listed in Table 3. In most cases, plasmid construction comprised multiple steps, and the details are available upon request.
pMEGA3 consists of the GAL4, GAL80, GAL3, and URA3 genes contained within a BamHI fragment inserted at the BamHI site of pC1/1. pC1/1 consists of pBR322, the entire 2μm plasmid of yeast, and leu2-d, the promoter-defective allele of LEU2 (31). Cells bearing pMEGA3 overproduce the Gal4, Gal80, and Gal3 proteins and show normal regulation of GAL genes (65a). pMEGA3-Δ3 (pAKS101) was constructed by deletion of the GAL3 promoter and two-thirds of the GAL3 open reading frame (ORF) from GAL3 in pMEGA3. Cells bearing pMEGA3-Δ3 overproduce Gal4p and Gal480p. pMEGA3-Δ4 (pMEGA4) is essentially pMEGA3 lacking the GAL4 gene. Cells bearing the pMEGA3-Δ4 overproduce Gal80p and Gal3p. pMEGA3-Δ4Δ3 (pMEGA5) is essentially pMEGA3 lacking the GAL4 and GAL3 genes. Cells bearing the pMEGA3-Δ4Δ3 overproduce Gal80p. pMEGA3-Δ4Δ80 (pMEGA11) is essentially pMEGA3 lacking the GAL4 and GAL80 genes. Cells bearing the pMEGA3-Δ4Δ80 overproduce Gal3p.
Plasmid pYGSTG4AD (pAKS71), carrying a GST-GAL4AD fusion expressed from the yeast ADH1 promoter, was constructed from the 750-bp SmaI-BamHI fragment of GST from pMPW60 (8) and a SmaI-BglII-gapped pGAD424 (Clontech). Plasmid pEGSTG4AD (pPXAKSGSTG4AD), used for expression of GSTG4AD in E. coli, was derived as an in-frame insertion of GAL4AD (codons 768 to 881) from GAL4 in pYCp50 (59) into SmaI-EcoRI-gapped pGEX2TK (Pharmacia Biotech). Plasmid pBDG80 (pLWBDG80) was constructed by inserting the GAL80 ORF as a PCR-generated BamHI fragment (using oligonucleotides G3 and G4) at the BamHI site of pMA424 (44). Plasmid pVP16G80 (pLWG80VP16) and pAKS87 encode a fusion of VP16 aa 413 to 490 with Gal80p. pVP16G80 was constructed by inserting the GAL80 ORF, as a PCR-generated BamHI fragment (using oligonucleotides G3 and G4) at the BamHI site of pVP16 (5). pAKS87 was constructed by inserting a 3.0-kb ClaI-NotI fragment obtained from pVP16G80 into NarI-NotI-digested pRS414. Plasmid pG4ADVP16 (pAKS70), consisting of ADH1pro-GAL4AD (codons 768 to 881)-VP16 (codons 413 to 490) on a CEN ARS LEU2 backbone, was constructed from pCRF-2 (61, 71a), pGAD424 (Clontech), and pYCplac111 (26). Plasmid pG4AD*VP16 (pAKS77) is a mutant derivative of pG4ADVP16 bearing Ile instead of Thr at aa 859 in the Gal4 activation domain. pG4AD*VP16 was constructed by fusion PCR mutagenesis (2), employing two PCR steps. In the first step, pG4ADVP16 was used as the template in two separate PCRs. Oligonucleotides ALOK49 (outside primer) and ALOK53 (mutagenic primer) were used for one reaction, and oligonucleotides ALOK52 (mutagenic primer) and BC02 (outside primer) were used for the other reaction (the oligonucleotides are given in Table 3). These reactions produced a 5′ product of 730 bp and a 3′ product of 770 bp. In the second step, a single PCR was carried out with 1 μl of each of the purified PCR products from the above reactions as a template together with oligonucleotides ALOK49 and BC02 as primers. The resulting full-length PCR product was digested with MluI and MscI and used to replace the corresponding wild-type region of GAL4AD-VP16 in pG4ADVP16 to yield pG4AD*VP16. pBDG80S2 (pCLAKS40) is a mutant derivative of pBDG80 containing the GAL80S-2 allele (encoding a product in which Lys at aa 351 replaces Glu) (20, 50, 51) in place of the wild-type GAL80. pG3VP16 (pAKS33) is a pTEB16 (Table 2) derivative that encodes a Gal3 (aa 1 to 524)-VP16 (aa 413 to 490) fusion protein. Plasmid LexAG4-225-797 (pAKS44), which encodes a LexA (aa 1 to 202)-Gal4 (aa 225 to 797) fusion protein, was constructed by fusing GAL4 codons 225 to 797 to LEXA codons 1 to 202 in pEG202 (from R. Brent). Plasmid pLexAG4-225-547 (pAKS35), which encodes a LexA (aa 1 to 202)-Gal4 (aa 225 to 547) fusion protein, was constructed by fusing the region of GAL4 comprising codons 225 to 547 to LEXA codons 1 to 202 in pEG202. Plasmid pLexAG4-534-797 (pAKS1), which encodes a fusion protein of LexA (aa 1 to 202)-Gal4 (aa 534 to 797), was constructed by fusing GAL4 codons 534 to 797 to LEXA codons 1 to 202 in pEG202.
Plasmid pLEUG4 (pAKS91), which encodes the wild-type Gal4p, was constructed by inserting a 3.6-kb BamHI-HindIII fragment containing the entire GAL4 gene obtained from pBM292 (33) into BamHI-HindIII-digested pRS415.
G4ADp-Gal80p interaction assays.
For the interaction assays Fig. 3A, a 100-μl aliquot of yeast extract comprising 700 μg of protein and containing GSTG4ADp was dispensed into a 1.5-ml Eppendorf tube and the volume was adjusted to 500 μl with buffer A. A 25-μl aliquot of GT-Sepharose bead suspension (Pharmacia Biotech) was added, and the tube was rotated for 90 min at 4°C. The beads were pelleted, and the pellet was washed twice with 500 μl of buffer A. A 100-μl aliquot of yeast extract comprising 1,000 μg of protein (100 μl) containing either Gal80p (Sc800 bearing pMEGA3-Δ4Δ3) or Gal80p plus Gal3p (Sc800 bearing pMEGA3-Δ4) was added to the beads, and the volume was adjusted to 250 μl with buffer A. To this 250-μl volume, an additional 250 μl of buffer A supplemented either with 10 mM MgCl2 or with 10 mM MgCl2–4 mM ATP–4% (wt/vol) galactose was added, and the sample was rotated for 2 h at 4°C. The beads were pelleted by centrifugation and subsequently washed three times with buffer A supplemented with either 5 mM MgCl2 or 5 mM MgCl2–2 mM ATP–2% galactose. A 70-μl volume of 1× electrophoresis loading buffer (48) was added to the bead pellet, and the sample was heated at 100°C for 8 min. For the interaction assay in Fig. 3B, the above procedure was used except that 70 μl of an E. coli extract (GSTG4ADp) together with 100 μl of yeast extract comprising 1,000 μg of protein (2×) or 35 μl of E. coli extract (GSTG4ADp) together with 50 μl of yeast extract comprising 500 μg of protein (1×) were used. For the in vitro protein interaction assays in Fig. 4 to 7, an 80 μl-aliquot of the E. coli extract (GSTG4ADp) was added to 25 μl of the GT-Sepharose bead suspension. The volume was then adjusted to 500 μl with buffer A, and the sample was rotated for 90 min at 4°C to allow binding of GSTG4ADp to the GT-Sepharose beads. The beads were pelleted and washed twice with 500 μl of buffer A. A 75-μl aliquot of yeast extract comprising 1,600 μg of protein from Sc800 bearing pMEGA3-Δ4Δ3 (the source of Gal80p) was added to the beads, and the sample was rotated for 2 h at 4°C to allow saturation of the bound GSTG4ADp with Gal80p. The beads were pelleted and washed twice with buffer A. A volume of yeast extract comprising 2,300 μg of protein prepared from Sc817 carrying either no plasmid or pMEGA3-Δ4Δ80 (GAL3) was added to the washed bead pellet. The volume was adjusted to 250 μl with buffer A. An additional 250 μl of buffer A supplemented with either 10 mM MgCl2, 10 mM MgCl2–4.0% galactose (wt/vol), 10 mM MgCl2–4.0 mM ATP, or 10 mM MgCl2–4% (wt/vol) galactose–4.0 mM ATP was immediately added. The sample was then rotated for 2 h at 4°C or, for the experiment in Fig. 7, rotated for 0, 30, 45, 60, 90, or 120 min as indicated. The beads were pelleted and washed three times with 500 μl of the indicated buffer. The proteins retained on the beads were eluted by heating at 100°C with 100 μl of the 1× electrophoresis loading buffer (48) for 8 min.
Immunoprecipitation.
Frozen yeast lysates were thawed on ice and immediately clarified by centrifugation at 50,000 rpm for 30 min in a TL-100.3 rotor in a Beckman TL-100 ultracentrifuge. Immunoprecipitation was carried out as previously described by Fujiki and Verner (23) with slight modifications. An aliquot of the clarified yeast lysate containing 3 mg of protein was adjusted to 800 μl with IP buffer (10 mM HEPES [pH 7.5], 200 mM NaCl, 1.0% Triton X-100, 10 mM NaF, 0.2 mM Na3VO4, 2 mM DTT, 2 mM EDTA, 20 μg of aprotinin per ml) supplemented with either 2.0% nonfat dry milk or 2.0% nonfat dry milk–2.0 mM ATP–2.0% (wt/vol) galactose. Anti-GD monoclonal antibody (a gift from S. S. Tevethia [14]) served as a nonspecific control. Anti-GD antibody was added to each sample, and the samples were rocked at 4°C for 60 min. Next, 50 μl of protein A-Sepharose beads (50.0% suspension) (Pharmacia Biotech) was added to each sample, and the samples were rocked for an additional 30 min at 4°C. The protein A-Sepharose beads were then pelleted, and the supernatant was transferred to a fresh tube. A 50-μl aliquot of the anti-901 monoclonal antibody (14) was then added to each sample, and the samples were rocked for 60 min at 4°C. Next, a 50-μl aliquot of protein A-Sepharose beads (50% suspension) were added, and the samples were rocked for 50 minutes at 4°C. The beads were pelleted and washed three times with IP buffer. A 60-μl aliquot of the 1× electrophoresis loading buffer was added to the Sepharose bead pellet, and the sample was heated at 100°C for 8 min.
Immunoblotting.
For the immunoblots in Fig. 3 to 5, an 18-μl aliquot (for Gal80p detection) and a 9-μl aliquot (for GSTG4ADp detection) of the GT-Sepharose eluate was applied to a sodium dodecyl sulfate (SDS)–8.0% polyacrylamide gel (acrylamide/bisacrylamide ratio, 30:0.8). Proteins were electrotransferred to nitrocellulose membrane, and the membrane was incubated (2 h at room temperature) with either anti-Gal80p (1:500 dilution) serum or anti-glutathione S-transferase (GST) (1:4,000 dilution) serum (provided by Andrew Waskiewicz and Jon Cooper). After being washed, the blots were incubated (1 h at room temperature) with anti-rabbit horseradish peroxidase-linked secondary antibody (Amersham Life Science). The proteins were detected by using a chemiluminescence reagent (Renaissance; NEN Life Science Products).
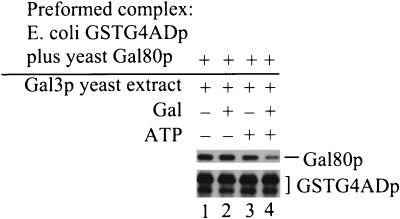
FIG. 5.
Both galactose and ATP are required for the Gal3p-mediated loss of Gal80p from GSTG4ADp-Gal80p complex. First, GSTG4ADp-Gal80p complex was formed on GT-Sepharose beads, as for Fig. 4. This GT-Sepharose-bound GSTG4ADp-Gal80p complex was incubated for 2 h at 4°C with the gal4Δ gal80Δ yeast extract containing Gal3p (Sc817 bearing pMEGA3-Δ4Δ80). Either no galactose or ATP (lane 1), galactose only (lane 2), ATP only (lane 3) or galactose and ATP (lane 4) were included in the binding buffer. GT-Sepharose-bound proteins were analyzed by Western immunoblotting with rabbit polyclonal anti-Gal80p and anti-GST.
For analyses of the results of the immunoprecipitation experiments in Fig. 1, SDS–7.5% polyacrylamide gels were used to fractionate proteins. Immunodetection of the Gal4p and Gal80p was carried out with anti-Gal4p (1:1,000 dilution) serum and anti-Gal80p (1:500 dilution) serum as above.
For the immunoblots in Fig. 6 and 7, a 5-μl aliquot (for Gal80p) and an 1-μl aliquot (for GSTG4ADp) of the GT-Sepharose eluate were subjected to SDS-polyacrylamide gel electrophoresis (8.0% acrylamide). Immunodetection of the GSTG4ADp and Gal80p was done with anti-GST (1:30,000 dilution) serum and anti-Gal80 (1:1000 dilution) serum as above.
FIG. 6.
In the presence of Gal3p, GSTG4ADp-Gal80p complexes are destabilized fourfold by galactose and ATP. First, GSTG4ADp-Gal80p complex was formed on GT-Sepharose beads, as for Fig. 4 and 5. This GT-Sepharose-bound GSTG4ADp-Gal80p complex was incubated for 2 h at 4°C with the gal4Δ gal80Δ yeast extract containing Gal3p (Sc817 bearing pMEGA3-Δ4Δ80). GT-Sepharose-bound proteins were analyzed by Western immunoblotting with rabbit polyclonal anti-Gal80p and anti-GST. Densitometric scans were performed to quantify the Gal80p and GSTG4ADp bands on the immunoblot. The densitometric units of Gal80p retained per unit of GSTG4ADp for each of six independent experiments were averaged to obtain the values shown. Error bars indicate the standard deviations.
Densitometric analysis.
Quantification of the Gal80p and GSTG4ADp bands detected by the chemiluminescence immunoblots in Fig. 6 and 7 was performed by scanning the immunoblots with a Molecular Dynamics 100A scanning densitometer running Soft Ware Quantity One from PDI. To establish that the band intensities produced for Gal80p and GSTG4ADp were within the linear response range for the chemiluminescence method, we performed pilot experiments in which we loaded different amounts of the GT-Sepharose eluate per lane and used several different film exposure times. By this procedure, we established that 5- and 1-μl aliquots of the GT-Sepharose eluate for Gal80p and GSTG4ADp, respectively, provided results meeting the requirement of linearity (data not shown).
β-Galactosidase assay.
β-Galactosidase assays were performed as described by Adams et al. (1), except that 50 μl of 10% SDS and 270 μl of β-mercaptoethanol were added to 100 ml of Z-buffer prior to use. The β-galactosidase filter assay was done as described by Vojtek and Hollenberg (73).
RESULTS
Retention of Gal80p in immunoprecipitates of Gal4p is reduced in the presence of galactose, ATP, and Gal3p.
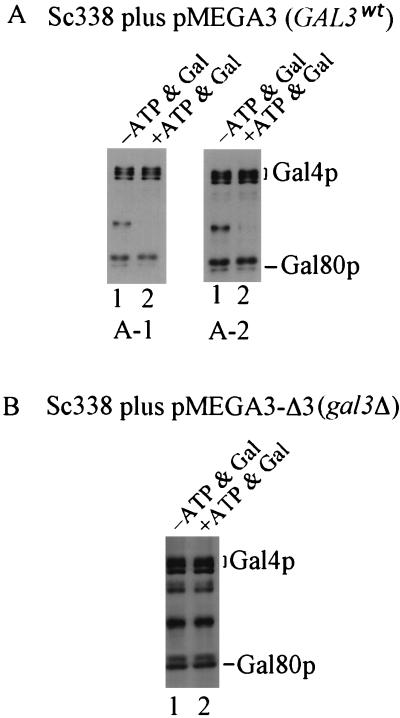
Yeast two-hybrid genetic data suggested a model wherein Gal80p does not dissociate from Gal4p upon galactose induction (38). To test this model by a more direct method, we carried out coimmunoprecipitation experiments. Anti-901 antibody directed against 901 epitope (14)-tagged Gal4p was used to coimmunoprecipitate Gal80p from extracts of Gly-Lac-grown yeast cells Sc338 carrying pMEGA3 or pMEGA3-Δ3. Coimmunoprecipitation was carried out in the presence or absence of galactose and ATP. The 901 epitope-tagged Gal4p exhibited fully wild-type behavior in its transcription activation, inhibition by Gal80p, and carbon-responsive phosphorylation (74a). Immunoprecipitates were analyzed by Western immunoblotting for the presence of Gal4p and Gal80p by using anti-Gal4p and anti-Gal80p sera, respectively. Three independent coimmunoprecipitations were performed for each cell extract in the presence or absence of galactose and ATP. The amount of Gal80p relative to the amount of Gal4p in the immunoprecipitate was compared. When GAL3 was present on the plasmid (pMEGA3) in yeast strain Sc338, we consistently observed a lower yield of Gal80p in the presence than in the absence of galactose and ATP (Fig. 1A, lanes 1 and 2). However, when GAL3 was absent from the plasmid (pMEGA3-Δ3) in Sc338, similar amounts of Gal80p were coimmunoprecipitated in the presence and absence of galactose and ATP (Fig. 1B). The chromosomal GAL3 gene of Sc338 is expressed at a low level in Gly-Lac, a level 40-fold lower than when GAL3 is carried on pMEGA3 (data not shown). Thus, the effect of Gal3p in these experiments is essentially due to Gal3p encoded by pMEGA3. Since galactose and ATP promote in vitro association of Gal3p and Gal80p, we reasoned that the reduction in the amount of Gal80p coimmunoprecipitated when Gal3p, galactose, and ATP were present might be relevant to how galactose triggers the Gal3p-mediated relief of Gal80p inhibition of Gal4p activity in vivo. Accordingly, a more detailed in vivo investigation of the effect of Gal3p on Gal4p-Gal80p interaction was undertaken by using a yeast two-hybrid approach.
The presence of Gal3p and galactose reduces lacZ reporter expression from a BDGal80p-Gal4ADVP16p two-hybrid pair.
We determined the effect of Gal3p and galactose on a two-hybrid genetic interaction between Gal80p and a Gal4 peptide comprising aa 768 to 881. This Gal4 peptide exhibits Gal80p binding (35, 42), TBP binding (3, 46, 75), and transcription activation activities (43). The wild-type Gal80p or a superrepressor mutant Gal80S-2p fused to the Gal4 peptide comprising aa 1 to 147 was used as bait. This Gal4 peptide comprises DNA-binding (10, 27, 45), nuclear localization (66), and dimerization (13, 24, 45) determinants. The wild-type Gal4p aa 768 to 881 (G4AD) or mutant Gal4p* aa 768 to 881 (G4AD*) fused to the COOH-terminal 78 aa (aa 413 to 490) of the herpesvirus transcriptional activator VP16 was used as prey (G4ADVP16p or G4AD*VP16p respectively). It was necessary to use the G4ADVP16 fusion protein instead of the G4AD protein alone since Gal80p represses the transcriptional activation function of Gal4p by physical association with a region within Gal4p aa 768 to 881. Bait and prey fusion constructs were expressed from the yeast ADH1 promoter. Bait plasmid, pBDG80, and prey plasmid, pG4ADVP16, and the corresponding controls, pRS414 (TRP1 vector alone) and pRS415 (LEU2 vector alone), were transformed into yeast strain Sc818 (gal4Δ gal80Δ gal3Δ gal1Δ). Expression of the UASGAL-lacZ fusion carried as a chromosomal copy in Sc818 provided a report of two-hybrid interaction as determined by β-galactosidase activity assays on extracts of Gly-Lac- and Gal-Gly-Lac-grown cells. The two-hybrid assay results are shown in Table 4. The combination of pBDG80 and pG4ADVP16 produced a high level of lacZ expression compared to pBDG80 alone, and the lacZ expression levels were similar in the presence and absence of galactose. These results indicated that there was a two-hybrid interaction between BDG80p and G4ADVP16p, which was not affected by galactose in the absence of Gal3p.
TABLE 4.
Effect of Gal3p on the two-hybrid interactions of BDG80 and G4ADVP16 alleles
| Plasmids | Proteins | β-Galactosidase activitya in:
|
Ratiob | |
|---|---|---|---|---|
| Gly-Lac | Gal-Gly-Lac | |||
| pBDG80 + pRS415 + pRS414 | BDG80 | 11.27 | 11.64 | 1.0 |
| pBDG80 + pG4ADVP16 + pRS414 | BDG80 + G4ADVP16 | 509.82 | 571.64 | 0.9 |
| pBDG80 + pG4ADVP16 + pTEB16 | BDG80 + G4ADVP16 + Gal3 | 494.55 | 222.91 | 2.2 |
| pBDG80 + pG4A*DVP16 + pTEB16 | BDG80 + G4AD*VP16 + Gal3 | 9.82 | 9.46 | 1.0 |
| pBDG80 + pAKS68 + pTEB16 | BDG80 + G4AD + Gal3 | 11.27 | 11.64 | 1.0 |
| pBDG80S2 + pRS415 + pRS414 | BDG80S2 | 6.18 | 5.82 | 1.1 |
| pBDG80S2 + pG4ADVP16 + pRS414 | BDG80S2 + G4ADVP16 | 542.18 | 620.36 | 0.9 |
| pBDG80S2 + pG4ADVP16 + pTEB16 | BDG80S2 + G4ADVP16 + Gal3 | 552.73 | 617.82 | 0.9 |
β-Galactosidase activity in yeast protein extracts is expressed in arbitrary units (1 Unit = 1,000OD420 per min per mg of protein). The values are averages of 4 to 12 independent transformants. Standard errors were 10 to 15%. The yeast host strain was Sc818 (gal4Δ gal80Δ gal3Δ gal1Δ ura3::GAL1-lacZ).
Ratio of β-galactosidase activity in Gly-Lac to that in Gal-Gly-Lac.
The effect of Gal3p on the BDG80p-G4ADVP16p two-hybrid interaction was then determined in the presence and absence of galactose. When plasmids pBDG80, pG4ADVP16, and pTEB16 (CEN vector carrying GAL3 expressed from its native promoter) were present in the absence of galactose, lacZ expression levels were similar to those attained in the absence of GAL3. However, when galactose was present, the cells carrying the GAL3 plasmid (pTEB16) exhibited an approximately twofold lower level of lacZ expression compared to when galactose was absent.
Since Gal1p, like Gal3p, interacts with Gal80p in a galactose-dependent manner and promotes GAL gene expression in response to galactose, we would expect that the presence of chromosomal GAL1 would cause reduced BDG80p-G4ADVP16p interaction in response to galactose. To test this, we repeated the BDG80p-G4ADVP16p two-hybrid assay in host strain Sc798. Sc798 (gal4Δ gal80Δ gal3Δ GAL1) is isogenic to Sc818 (gal4Δ gal80Δ gal3Δ gal1Δ) except for the GAL1 gene. In Sc798 (GAL1), the BDG80p-G4ADVP16p-driven reporter expression in galactose was half that observed in Gly-Lac (Table 5). Such a reduction was not observed in Sc818 (gal1Δ) (Table 4). This indicates that Gal1p, produced by BDG80p-G4ADVP16p interaction at the GAL1 gene promoter of Sc798 (Table 5), is responsible for the reduced lacZ reporter expression. Thus, both Gal3p and Gal1p, under conditions known to favor their binding to Gal80p (i.e., galactose), caused a reduction of lacZ reporter expression from the Gal80p-G4ADp two-hybrid interaction. The relative potencies of Gal3p and Gal1p in reducing BDG80p-G4ADVP16p interaction in response to galactose cannot be determined from these experiments, since the relative levels of Gal3p and Gal1p present were not determined.
TABLE 5.
Effect of Gal1p on the two-hybrid interaction of BDG80p and G4ADVP16p
| Proteins | Relevant genotype of host strain | β-Galactosidase activitya in:
|
Ratiob | |
|---|---|---|---|---|
| Gly-Lac | Gal-Gly-Lac | |||
| BDG80 | gal4Δ gal80Δ gal3Δ GAL1 | 5.09 | 4.00 | 1.3 |
| BDG80 + G4ADVP16 | gal4Δ gal80Δ gal3Δ GAL1 | 427.27 | 218.55 | 2.0 |
β-Galactosidase activity in yeast protein extracts is expressed in arbitrary units (1 unit = 1,000OD420 per min per mg of protein). The values are averages of four independent transformants. Standard errors were 3 to 5%. The yeast host strain was Sc798 (gal4Δ gal80Δ gal3Δ).
Ratio of β-galactosidase activity in Gly-Lac to that in Gal-Gly-Lac.
To determine whether carbon-responsive changes in the levels of BDG80p and G4ADVP16p might influence the two-hybrid results, we determined the levels of these proteins in extracts of cells grown in the presence or absence of galactose under the conditions of our two-hybrid experiments. Western immunoblotting and densitometry showed that both proteins were present at higher levels in cells cultured in the presence of galactose in either the presence or absence of Gal3p (data not shown). This result was expected, since BDG80p and G4ADVP16p were expressed from the yeast ADH1 promoter, a promoter known to be more active in fermentable sugar media than in nonfermentable media (17, 71). Therefore, based on promoter activity driving the expression of bait and prey, one would expect higher two-hybrid lacZ reporter expression in the presence than in the absence of galactose. Thus, the twofold reduction in lacZ reporter activity we found in response to galactose in the presence of Gal3p (or Gal1p) (Tables 4 and 5) is likely to be an underestimate of the effect of Gal3p and galactose on the Gal80p-Gal4p (aa 768 to 881) interaction.
To ascertain that our bait and prey molecules exhibited behavior consistent with known properties of Gal80p and Gal4p we performed further genetic experiments, taking advantage of mutant proteins with known properties. As one test, we repeated the two-hybrid assay in the presence of Gal3p but used BDG80p paired with G4AD*VP16p instead of G4ADVP16p. G4AD*VP16p bears the Thr859-to-Ile mutation within G4AD. This mutation in the context of full-length Gal4p causes an in vivo constitutive phenotype, indicating an inability to interact with repressor Gal80p (39, 64). G4AD*VP16p did not give rise to a two-hybrid interaction with BDG80p (Table 4), suggesting that the BDG80p-G4ADVP16p two-hybrid interaction was physiologically relevant.
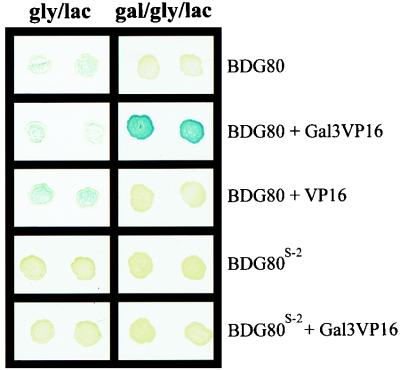
To determine whether the two-hybrid interaction between BDG80p and G4ADVP16p responds to Gal3p in a physiologically relevant manner, we took advantage of the GAL80S-2 allele. The GAL80S-2 allele blocks galactose-triggered, Gal3p-mediated Gal4p activation in vivo, and its encoded Gal80S-2 protein, in contrast to the wild-type Gal80 protein, does not interact with Gal3p in vitro (76). We performed two-hybrid interaction assays with the pBDG80-pG3VP16 and pBDG80S2-pG3VP16 pairs. As expected, BDGal80p, but not BDGal80S-2p, showed galactose-dependent interaction with Gal3VP16p (Fig. 2). We then performed a two-hybrid interaction assay pairing the bait BDGal80S-2p with G4ADVP16p. The BDGal80S-2p-G4ADVP16p interaction, in contrast to the BDGal80p-G4ADVP16p interaction, was not reduced in the presence compared to the absence of Gal3p and galactose (Table 4). The slightly increased lacZ expression in the presence of galactose was probably due to the increased expression levels observed from the ADH1 promoter in galactose compared to glycerol-lactic acid (see above). Thus, the observed action of Gal3p together with galactose in reducing the BDGal80p-G4ADVP16p-mediated lacZ expression occurred only under conditions allowing Gal3p-Gal80p interaction.
FIG. 2.
Two-hybrid interactions between wild-type or mutant BDG80p and Gal3VP16p. Yeast transformants were spotted onto a nitrocellulose membrane (Optitran; Schleicher & Schuell) placed on the surface of SC selective agar plates containing either Gly-Lac or Gal-Gly-Lac as the carbon source and allowed to grow for 2 days at 30°C. β-Galactosidase activity was assayed on these membranes. The carbon source is indicated at the top. The two-hybrid bait and prey molecules are indicated on the right.
Gal80p retention by GSTG4AD (aa 768 to 881) is reduced in the presence of Gal3p, galactose, and ATP.
The lower lacZ expression observed for the BDG80p-G4ADVP16p pair in the presence compared to the absence of Gal3p and galactose could arise from a reduced BDG80p-G4ADp interaction caused by Gal3p-Gal80p interaction. Alternatively, lower lacZ expression could arise without any change in the BDG80p-G4ADVP16p interaction if Gal3p binding to Gal80p caused reduced access of VP16 to its target(s) through steric hindrance. To determine directly whether Gal3p could reduce an interaction between Gal80p and the Gal4 peptide from aa 768 to 881, we performed in vitro pull-down assays. A GSTG4AD (GAL4 codons 768 to 881) fusion construct carried on plasmid pYGSTG4AD was expressed in yeast from the yeast ADH1 promoter. GSTG4ADp from yeast extract was bound to GT-Sepharose beads, and the GT-Sepharose-bound GSTG4ADp was washed and subsequently incubated with yeast extracts prepared from Gly-Lac-grown Sc800 (gal4Δ gal80Δ gal3Δ) carrying either pMEGA3-Δ4Δ3 (GAL80 only) or pMEGA3-Δ4 (GAL80 and GAL3). Since both galactose and ATP are required for the in vitro Gal3p-Gal80p interaction (68, 76, 79), incubations were carried out in either the presence or absence of galactose and ATP. Retention of Gal80p by GSTG4ADp was determined by Western immunoblotting with anti-Gal80p and anti-GST.
The amount of Gal80p retained was similar in both the absence and presence of galactose and ATP when no Gal3p was present (Fig. 3A, lanes 1 and 2). In contrast, when Gal3p was present, less Gal80p was retained in the presence than compared in the absence of galactose and ATP (lanes 3 and 4). We conclude that in the presence of galactose and ATP, Gal3p reduces the retention of Gal80p by GSTG4ADp in vitro.
To facilitate the production and isolation of GSTG4ADp for further analyses, we constructed GSTG4AD in an E. coli expression vector. Plasmid pEGSTG4AD was propagated in E. coli to produce GSTG4ADp for a GSTG4ADp-Gal80p binding assay. The format of this GSTG4ADp-Gal80p binding experiment was essentially that of the above experiment which utilized the yeast expressed GSTG4ADp. The E. coli-expressed GSTG4ADp retained similar amounts of Gal80p in both the presence and absence of galactose and ATP when Gal3p was absent (Fig. 3B, lanes 5 to 8). In contrast, when Gal3p was present, the amount of Gal80p retained by GSTG4ADp was markedly smaller in the presence than in the absence of galactose and ATP (lanes 1 to 4). These results confirmed the results with yeast GSTG4ADp and validated the use of E. coli-expressed GSTG4ADp in our subsequent experiments.
To determine whether the above results with GSTG4ADp arose due to the GST moiety, we performed Gal80p-G4ADp binding experiments in the presence and absence of Gal3p, galactose, and ATP by using a His6-tagged G4ADp fusion instead of the GSTG4ADp fusion. Our results were essentially the same as above (data not shown). Thus, the results observed for GSTG4ADp were not an artifact due to the presence of GST but, rather, reflected the properties of the Gal4ADp-Gal80p interaction under the various conditions tested. The observed reduction in Gal80p retention in the presence of Gal3p, galactose, and ATP could be due to a destabilization of GSTG4ADp-Gal80p complex or to slower formation of the GSTG4ADp-Gal80p complex in presence of Gal3p, galactose, and ATP, or both.
Preformed GSTG4ADp-Gal80p complex is destabilized by Gal3p in the presence of galactose and ATP.
We performed a two-step experiment to determine whether Gal3p reduced the stability of a preformed GSTG4ADp-Gal80p complex in the presence of galactose and ATP. First, a predetermined amount of E. coli GSTG4ADp bound to GT-Sepharose was saturated with Gal80p from a whole-cell extract of Gly-Lac-grown yeast strain Sc800 (gal4Δ gal80Δ gal3Δ) carrying pMEGA3-Δ4Δ3 (GAL80). The resulting GSTG4ADp-Gal80p complex was washed and subsequently incubated in the presence or absence of galactose and ATP at 4°C for 2 h with whole-cell extract of Gly-Lac-grown cells of Sc817 (gal4Δ gal80Δ gal3Δ) or Sc817 carrying pMEGA3-Δ4Δ80 (GAL3). In the absence of Gal3p, no difference in Gal80p retention was observed in the presence or absence of galactose and ATP (Fig. 4, lanes 1 and 2). When Gal3p was present, a marked decrease in the retention of Gal80p was observed in the presence compared to the absence of galactose and ATP (lanes 3 and 4). We conclude that in vitro, Gal3p in the presence of galactose and ATP causes destabilization of preformed Gal4ADp-Gal80p complex.
Both ATP and galactose are required for maximum Gal3p-mediated destabilization of the preformed E. coli GSTG4ADp-Gal80p complex.
Both galactose and ATP are required for the Gal80p-Gal3p (Gal1p) interaction (8, 68, 76, 79). To determine whether both galactose and ATP are necessary for the observed Gal3p-mediated destabilization of G4ADp-Gal80p complex, we performed the following experiment. GSTG4ADp-Gal80p complexes bound to glutathione-Sepharose beads were incubated with yeast whole-cell extract from Gly-Lac-grown strain Sc817 carrying pMEGA3-Δ4Δ80 (GAL3) in the presence of either galactose or ATP alone, in the presence of both galactose and ATP, or in the absence of both galactose and ATP. The amount of Gal80p retained by GSTG4ADp was determined by Western immunoblotting, as above. The addition of either galactose or ATP alone had little or no effect on the amount of Gal80p retained by GSTG4ADp (Fig. 5, lanes 1 to 3). In contrast, when both galactose and ATP were added, less Gal80p was retained by GSTG4ADp compared to when both galactose and ATP were absent (compare lanes 1 and 4). We conclude that destabilization of an in vitro GSTG4ADp-Gal80p complex by Gal3p requires galactose and ATP, both of which are required for Gal3p-Gal80p complex formation.
All of the experiments above indicated that in the presence of galactose (and ATP) Gal3p destabilizes a complex between Gal80p and Gal4p. Galactose (and ATP) have no effect in the absence of Gal3p. The GSTG4ADp pull-down experiments emphasize that this effect of Gal3p is on the interaction of Gal80p with the Gal4p activation domain, aa 768 to 881. To quantify the effect of galactose plus ATP on the Gal80p-GSTG4ADp interaction in the presence of Gal3p, we performed six independent GSTG4ADp-Gal80p pull-down experiments similar to the experiments illustrated in Fig. 5, except that we did not test the individual effects of galactose and ATP. For these experiments, we established that the GSTG4ADp and Gal80p band intensities detected by chemiluminescence were within the linear range for lane load versus response (see Materials and Methods). The amount of Gal80p retained by GSTG4ADp was 3.5- to 4.2-fold smaller in the presence than in the absence of galactose and ATP (Fig. 6). Two additional experiments were performed in an identical fashion to the above, except that the alkaline phosphatase detection method was used, as described previously (48), for Western immunoblot band detection. By the alkaline phosphatase detection method, we observed a fourfold-lower retention of Gal80p by GSTG4AD in the presence than in the absence of galactose and ATP (data not shown). Thus, both the chemiluminescence and alkaline phosphatase detection methods provided essentially identical results.
Kinetics of loss of retention of Gal80p from the GSTG4ADp-Gal80p complex in response to Gal3p, galactose, and ATP.
To determine the relative rates of loss of Gal80p from the GSTG4ADp-Gal80p complex in the presence of Gal3p and in the presence or absence of galactose and ATP, we performed four independent time course experiments. Preformed GT-Sepharose-GSTG4ADp-Gal80p complexes were incubated in the absence or presence of galactose and ATP at 4°C with whole-cell extract of Gly-Lac-grown Sc817 cells carrying pMEGA3-Δ4Δ80(GAL3). At various times over the course of 2 h, the retention of Gal80p by GSTG4ADp was determined by Western immunoblotting and densitometry. For these experiments, we established that the GSTG4ADp and Gal80p band intensities detected by chemiluminescence were within the linear range for lane load versus response (see Materials and Methods). The retention of Gal80p decreased over time markedly in the presence of galactose and ATP but only slightly in the absence of galactose and ATP (Fig. 7). Figure 7A illustrates the immunoblot of one of the four experiments. Plots in Fig. 7B represent the average values obtained for all four independent pull-down experiments. The ratio of the slopes of the time courses in Fig. 7B taken over the first 60 min indicates a sevenfold-higher rate of loss of Gal80p from GSTG4ADp in the presence than in the absence of galactose and ATP.
Gal80p exhibits a two-hybrid interaction with Gal4p aa 225 to 797.
One interpretation consistent with all of our results is that in the presence of galactose (and ATP), Gal3p binds to Gal80p complexed with Gal4p within the region of Gal4p aa 768 to 881 and destabilizes the complex. Leuther and Johnston (38), using a two-hybrid approach, found evidence consistent with persistence of a Gal80p-Gal4p complex in the presence of galactose. Our results could be reconciled with those of Leuther and Johnston (38) if there were a second Gal80p binding site present on Gal4p outside the G4AD (aa 768 to 881) region.
We used a two-hybrid approach to test for a possible second Gal80p binding site on Gal4p. Three different segments of Gal4p fused to the LexA DNA binding domain (aa 1 to 202) were used individually as baits paired with VP16Gal80p as prey. The LexA-Gal4p aa 225 to 797 bait, paired with VP16Gal80p, yielded levels of lacZ expression markedly above background, whereas each of the other two Gal4 segment baits (Gal4p aa 225 to 547 and aa 534 to 797) did not (Table 6). These results suggest that in addition to the classical Gal80p binding site located within Gal4p aa 768 to 881, there is a second site, within Gal4p aa 225 to 797, with which Gal80p associates.
TABLE 6.
Two-hybrid interaction of LexA-Gal4p aa 225 to 797 and VP16Gal80p
| Plasmids | Proteins | β-Galactosidase activitya |
|---|---|---|
| pEG202 + pVP16G80 | LexA(aa1–202) + VP16Gal80 | 9.86 ± 0.96 |
| pLexAG4-225–547 + pVP16G80 | LexA(aa1–202)-Gal4(aa225–547) + VP16Gal80 | 11.52 ± 1.73 |
| pLexAG4-534–797 + pVP16G80 | LexA(aa1–202)-Gal4(aa534–797) + VP16Gal80 | 10.41 ± 1.07 |
| pLexAG4-225–797 + pVP16G80 | LexA(aa1–202)-Gal4(aa225–797) + VP16Gal80 | 330.28 ± 6.01 |
| pLexAG4-225–797 + pVP16 | LexA(aa1–202)-Gal4(aa225–797) + VP16 | 16.8 ± 1.2 |
β-Galactosidase activity in yeast protein extracts is expressed in arbitrary units (1 unit = 1,000OD420 per min per mg of protein. The yeast host strain used was Sc813 [gal4Δ gal80Δ URA3::(lexA)8-lacZ].
To verify the two-hybrid interaction of VP16Gal80p with the middle region of Gal4p, we performed two-hybrid assays with yeast strain Y187 (29), in which VP16Gal80p was partnered with either the wild-type Gal4p or Gal4p variants lacking the middle region. We used two such Gal4 variants, Gal4mCla (lacking aa 148 to 728) and miniGal4-2 (lacking aa 210 to 716) (18). The results shown in Table 7 illustrate that for wild-type Gal4p partnered with VP16Gal80p, lacZ expression increases in response to galactose. This result is in agreement with earlier results of Leuther and Johnston (38). In contrast, VP16Gal80p partnered with either of the two Gal4ps lacking the middle region, shows reduced lacZ expression in response to galactose. These results are expected if Gal80p is able to associate with the middle region of Gal4p in the presence of galactose. On the basis of these results, we suggest that in addition to the classical Gal80p binding site located within Gal4p aa 768 to 881 there is a second site, within Gal4p aa 225 to 797, with which Gal80p associates.
TABLE 7.
Compared to wild-type Gal4p, mutant Gal4ps lacking the middle region show reduced two-hybrid interaction with VP16Gal80p in the presence of galactose
| Plasmids | Proteins | β-Galactosidase activitya in:
|
|
|---|---|---|---|
| Gly-Lac | Gal | ||
| pLEUG4 + pAKS87 | Wild-type Gal4 + VP16Gal80 | 4,167 | 6,157 |
| pSB32-miniGal4-2 + pAKS87 | Mini-Gal4-2 + VP16Gal80 | 5,830 | 3,495 |
| pSB32-Gal4mCla + pAKS87 | Gal4mCla + VP16Gal80 | 5,925 | 3,840 |
β-Galactosidase activity in yeast protein extracts is expressed in arbitrary units (1 unit = 1,000OD420 per min per mg of protein). The values are averages of four independent transformants. Standard errors were 5 to 10%. The yeast host strain was Y187 (gal4Δ gal80Δ URA3::GAL1-lacZ).
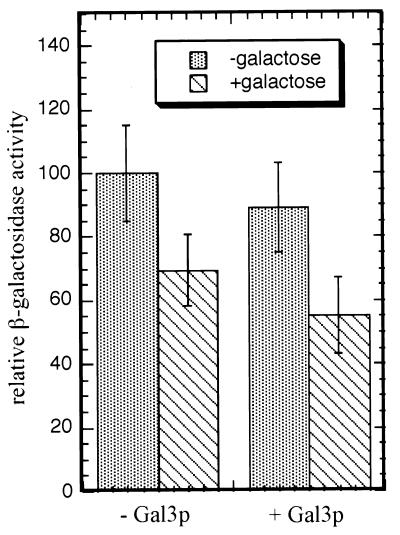
To determine the effect of Gal3p and galactose on the two-hybrid interaction between VP16Gal80p and the Gal4p middle region, we carried out two-hybrid assays with yeast strain Sc830 (gal4Δ gal80Δ gal3Δ) carrying plasmids pLexAG4-225–797 and pVP16G80 together with either pMPW66 (CEN ARS TRP1 plasmid with GAL3 expressed from the ADH2 promoter) or pRS414 (control vector) (Table 2). The β-galactosidase activities in whole-cell extracts are shown Fig. 8. These results indicate that Gal3p has no effect on the interaction of VP16Gal80p with the middle region of Gal4p and that galactose has a slight negative effect.
FIG. 8.
Gal3p does not have an effect on the two-hybrid interaction between VP16Gal80p and the middle region of Gal4p (aa 225 to 797). Yeast strain Sc830 was cotransformed with pLexAG4-225-797 and pVP16G80 together with either pMPW66 (GAL3 expressed from ADH2 promoter) or pRS414 (empty vector). Extracts were prepared from transformants grown in the presence or absence of galactose and assayed for β-galactosidase activity. The activity obtained from the interaction between LexAGal4-225-797p and VP16Gal80p in the absence of both Gal3p and galactose was set to 100. All the results were normalized accordingly. The values are the averages of eight independent transformants. Error bars show standard deviations.
DISCUSSION
Essential features of signal transduction from galactose to Gal4p transcriptional activation are beginning to emerge. Gal3p is able to complex with Gal80p in vitro in the presence of galactose and ATP (68, 76, 79). Gal3C mutant proteins capable of binding Gal80p in vitro in the absence of galactose confer a constitutive phenotype, linking galactose-promoted Gal3p-Gal80p complex formation to Gal4p-mediated transcription activation in vivo (8). A genetic two-hybrid interaction between Gal80p and Gal4p persists in galactose-grown yeast, suggesting nondissociation of Gal80p and Gal4p upon induction (38). Galactose- and ATP-enhanced formation of a ternary protein complex composed of Gal80p, a Gal4p variant (aa 1 to 94 and 768 to 881), and Gal3pC-322 has been established in vitro (54). Finally, Gal80p binding to a region within the principal activation region (aa 768 to 881) of Gal4p prohibits TBP binding (3). Taken together, these observations suggest a model wherein galactose activates Gal3p binding to Gal80p, causing a conformational change in the Gal4p-Gal80p complex and thereby freeing the Gal4p activation region from Gal80p and allowing it to bind transcription complex proteins.
By performing three distinct types of experiments, we have provided evidence for the dissociation of Gal80p from Gal4p in specific response to Gal3p and galactose (and ATP). First, we observed that immunoprecipitates of epitope-tagged full-length Gal4p from yeast extracts consistently contain less coimmunoprecipitated Gal80p in the presence than in the absence of Gal3p, galactose, and ATP. Second, reporter gene expression from a two-hybrid genetic interaction between Gal80p and G4ADp was reduced in response to the presence of Gal3p and galactose. Third, GT-Sepharose pull-down assays of yeast extract-derived Gal80p with GSTG4ADp consistently showed reduced levels of Gal80p retained by GSTG4ADp in the presence compared to the absence of Gal3p, galactose, and ATP. The presence of galactose and ATP in the presence of Gal3p caused a 3.5- to 4.2-fold reduction in the amount of Gal80p retained by GSTG4ADp over the course of 2 h (Fig. 6).
Our time course experiments revealed that preformed GSTG4ADp-Gal80p complexes decay slowly in the presence of Gal3p, even when galactose and ATP are absent (Fig. 7). We attribute this slow decay to Gal3p and not to some other effect of the incubation conditions, since under identical conditions, we observed that in the absence of Gal3p the GSTG4ADp-Gal80p complex was stable even in the presence of galactose and ATP (Fig. 4). Galactose-independent Gal3p-mediated decay of the Gal80p-Gal4p complex is not surprising, since overproduction of Gal3p (and Gal1p) causes galactose-independent Gal4p mediated transcription activation (6). Moreover, low levels of galactose-independent binding of Gal3p to Gal80p have been observed previously (74a). The capacity of Gal3p to bind to Gal80p and cause decay of the Gal80p-Gal4p complex in the absence of galactose may well contribute to the low basal Gal4p-dependent transcription activation characteristic of some of the GAL gene promoters (30, 55).
The relative effect of the presence of galactose and ATP on the ability of Gal3p to dissociate preformed GSTG4ADp-Gal80p complexes is represented by the difference in the rate of loss of Gal80p from GSTG4ADp in our time course experiments. Over the course of the first 60 min, as averaged over four independent experiments, the apparent decay of GSTG4ADp-Gal80p complexes in response to Gal3p was on the order of sevenfold more rapid in the presence than in the absence of galactose and ATP (Fig. 7B). The results of these time course experiments and the results of the other experiments reported here support a model wherein a galactose-dependent Gal3p-Gal80p interaction weakens the association of Gal80p with Gal4p.
Dissociation of Gal80p and Gal4p was not evident in the experiments of Platt and Reece (54). Using purified components in a DNA electrophoretic mobility shift assay, they showed that galactose and ATP promotes the binding of Gal3pC-322 to a Gal80p-Gal4p-DNA complex in vitro to yield a ternary protein complex. They employed Gal4p (aa 1 to 94 and 768 to 881) consisting of Gal4p aa 1 to 94 fused to the same Gal4p activation domain (aa 768 to 881) we used here in fusion with GST. However, in their experiments the binding-reaction mixtures may not have been incubated for a sufficient time prior to gel loading to ensure that equilibrium had been reached. Thus, the ternary complexes captured upon loading the gel at 20 min of incubation under the conditions of their experiments may have presented only an early phase, dominated by association of Gal3p with the Gal80p-Gal4p-DNA complex. Moreover, the experiments of Platt and Reece used different ion concentration, pH, and buffer conditions than did the work presented herein. Additionally, the experiments of Platt and Reece used a Gal3pC-322 constitutive mutant protein rather than wild-type Gal3p as we did here. Such differences might explain why our experiments revealed dissociation and their experiments did not.
We hypothesize that the decreased stability of the Gal80p-Gal4p complex in response to Gal3p and galactose (and ATP) shown herein represents a significant mechanistic event subsequent to the Gal3p-Gal80p binding step. Ansari et al. (3) have shown that interaction of Gal80p with Gal4p aa 840 to 881 (G4AD) prohibits the binding of TBP. Thus, reduced stability of the Gal80p-G4ADp complex would increase the probability of G4AD participation in forming complexes with TBP and perhaps other general transcription factor proteins (TFIIB, etc.) (3, 18, 46, 75).
An additional finding of potential mechanistic significance is the two-hybrid genetic interaction of Gal80p with a segment of Gal4p (aa 225 to 797) that lies linearly distant from the classical Gal80p binding site (Gal4p aa 850 to 874) (Table 6). Neither of the two approximate halves (aa 225 to 547 and aa 534 to 797) of this region tested positive for a two-hybrid interaction with Gal80p. Although the interaction of VP16Gal80p with the Gal4p middle region (aa 225 to 797) appears slightly reduced in the presence compared to the absence of galactose, the presence or absence of Gal3p has no effect (Fig. 8). Thus, the galactose-triggered Gal3p-interaction with Gal80p does not play a role in the interaction of Gal80p with the middle region of Gal4p, unlike in the interaction of Gal80p with the carboxy-terminal domain of Gal4p. The slight negative effect of galactose, independent of Gal3p, might be due to a direct effect of galactose on Gal80p, but no evidence for galactose binding to Gal80p has been reported.
Additional evidence for the interaction of VP16Gal80p with the middle region of Gal4p came from experiments with well-characterized mini-Gal4ps lacking the middle region. If dissociation of Gal80p from the Gal4AD region (aa 768 to 881) is favored in galactose and if Gal80p can associate with the middle region of Gal4p, reporter expression will be driven by both the Gal4p activation domain (aa 768 to 881) and the activation domain of VP16 (VP16Gal80p, associated with the Gal4p middle region). Thus, increased reporter expression in response to galactose is expected if full-length Gal4p is used. Leuther and Johnston observed a galactose-responsive increase in reporter gene expression in their experiments with Gal80VP16 and full-length Gal4p (38). However, the use of Gal4ps lacking the middle region should fail to yield an increase in reporter expression in response to galactose. Indeed, we observed this expected result (Table 7). These data are consistent with the notion that the middle region is required for retention of VP16Gal80p in our experiments. Based on these results, we hypothesize that two sites for Gal80p binding exist on Gal4p, one being the classic site comprising Gal4p aa 850 to 874 and the other being a site comprising part(s) of Gal4p aa 225 to 797 (Fig. 9).
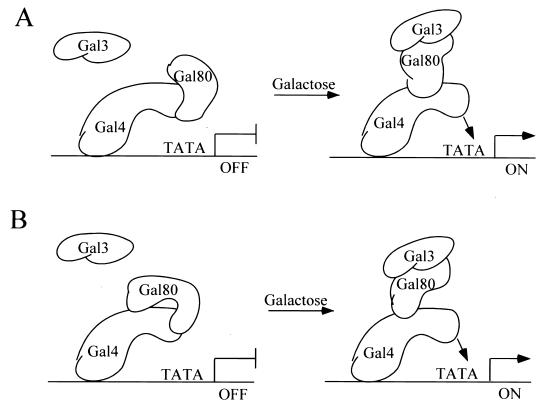
FIG. 9.
Two-site model for conformational change in the Gal80p-Gal4p complex in response to galactose-induced Gal3p-Gal80p complex formation. (A) Association of Gal80p with Gal4p at its classical inhibition target site within the Gal4p activation domain. In response to galactose, Gal3p binding to Gal80p destabilizes the association of Gal80p with the classical inhibition site. Gal80p now associates with another site within Gal4p aa 225 to 797, thereby freeing Gal4p to recruit a polymerase II complex for transcription initiation. (B) Model in which Gal80p is bound to both sites simultaneously in the absence of galactose.
Consistent with a two-site model for Gal80p-Gal4p interaction, we found that the loss of retention of Gal80p in response to Gal3p, galactose, and ATP was markedly higher for GSTG4ADp than for Gal4p901. We observed a small but reproducible reduction of Gal80p retained by full-length coimmunoprecipitated Gal4p901 compared with a 3.5- to 4.2-fold reduction of Gal80p retained by GSTG4ADp. Stabilization of the Gal4p901-Gal80p complex by anti-901 might account for this difference. However, the transfer of Gal80p to a second location on Gal4p901 which is absent in GSTG4ADp (e.g., Gal4 aa 225 to 797) could also explain the difference. The development of a physical assay for Gal80p binding to Gal4p aa 225 to 797 and the use of methods such as protein cross-linking may help distinguish between these possibilities and test the two-site model.
Association of Gal80p with the middle region of Gal4p in the induced state, as suggested by a two-site model of Gal80p-Gal4p interaction, could reduce the search time needed for Gal80p to find its inhibitory site on Gal4p as galactose levels fall. This would be a possible mechanism by which Gal80p inhibition of Gal4p could be rapidly responsive to declining galactose levels.
The precise nature of the change in Gal80p leading to its dissociation from the principal activation domain of Gal4p is another of several facets of the Gal3p-Gal80p-Gal4p switch mechanism that warrants further investigation. Gal80p phosphorylation state changes in response to carbon source have been reported (80), and such changes could, if further documented, provide potential insight into the mechanism of this switch. Also, phosphorylation of Gal4p at serine 699 has been reported to alter the relationship of Gal4p to Gal80p inhibition (60). It is conceivable that such alterations in Gal4p will provide clues to the nature of Gal3p-triggered alteration of Gal80p. For example, changes in Gal4p phosphorylation state might influence which binding site is favored by Gal80p. These issues and potentialities will provide some of the focus for future work on the GAL switch. The assays we have developed here should contribute to a widening arsenal of technical capabilities for exploring the workings of the GAL switch.
ACKNOWLEDGMENTS
We thank J. Cooper, S. J. Elledge, R. D. Gietz, J. H. Hegemann, P. Heiter, S. A. Johnston, S. S. Tevethia, S. J. Triezenberg, A. Waskieawicz, and L. W. Yuan for providing the plasmids, strains, and antisera used in this work. We thank M. Fried, I. Ropson, and R. Shiman for useful discussions. We thank R. Blank, A. Hopper, G. Peng, S. Sarkar, and D. Spector for critical evaluation of the manuscript.
This work is supported by Public Health Service grant GM27975 to James E. Hopper.
REFERENCES
- 1.Adams A, Gottschling D E, Kaiser C A, Stearns T. Methods in yeast genetics: a laboratory course manual. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1997. [Google Scholar]
- 2.Amberg D C, Botstein D, Beasley E M. Precise gene disruption in Saccharomyces cerevisiae by double fusion polymerase chain reaction. Yeast. 1995;11:1275–1280. doi: 10.1002/yea.320111307. [DOI] [PubMed] [Google Scholar]
- 3.Ansari A Z, Reece R J, Ptashne M. A transcriptional activating region with two contrasting modes of protein interaction. Proc Natl Acad Sci USA. 1998;95:13543–13548. doi: 10.1073/pnas.95.23.13543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Barberis A, Pearlberg J, Simkovich N, Farrell S, Reinagel P, Bamdad C, Sigal G, Ptashne M. Contact with a component of the polymerase II holoenzyme suffices for gene activation. Cell. 1995;81:359–368. doi: 10.1016/0092-8674(95)90389-5. [DOI] [PubMed] [Google Scholar]
- 5.Bartel P L, Fields S. Analyzing protein-protein interactions using two-hybrid system. Methods Enzymol. 1995;254:241–263. doi: 10.1016/0076-6879(95)54018-0. [DOI] [PubMed] [Google Scholar]
- 6.Bhat P J, Hopper J E. Overproduction of the GAL1 or GAL3 protein causes galactose-independent activation of the GAL4 protein: evidence for a new model of induction for the yeast GAL/MEL regulon. Mol Cell Biol. 1992;12:2701–2707. doi: 10.1128/mcb.12.6.2701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bhat P J, Oh D, Hopper J E. Analysis of the GAL3 signal transduction pathway activating GAL4 protein-dependent transcription in Saccharomyces cerevisiae. Genetics. 1990;125:281–291. doi: 10.1093/genetics/125.2.281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Blank T E, Woods M P, Lebo C M, Xin P, Hopper J E. Novel Gal3 proteins showing altered Gal80p binding cause constitutive transcription of Gal4p-activated genes in Saccharomyces cerevisiae. Mol Cell Biol. 1997;17:2566–2575. doi: 10.1128/mcb.17.5.2566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Boeke J D, LaCroute F, Fink G R. A positive selection for mutants lacking orotidine-5′-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197:345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- 10.Bram R J, Kornberg R D. Specific protein binding to far upstream activating sequences in polymerase II promoters. Proc Natl Acad Sci USA. 1985;82:43–47. doi: 10.1073/pnas.82.1.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Brent R, Ptashne M. A eukaryotic transcriptional activator bearing the DNA specificity of a prokaryotic repressor. Cell. 1985;43:729–736. doi: 10.1016/0092-8674(85)90246-6. [DOI] [PubMed] [Google Scholar]
- 12.Broach J R. Galactose regulation in Saccharomyces cerevisiae. The enzymes encoded by the GAL7, 10, 1 cluster are co-ordinately controlled and separately translated. J Mol Biol. 1979;131:41–53. doi: 10.1016/0022-2836(79)90300-0. [DOI] [PubMed] [Google Scholar]
- 13.Carey M, Kakidani H, Leatherwood J, Mostashari F, Ptashne M. An amino-terminal fragment of GAL4 binds DNA as a dimer. J Mol Biol. 1989;209:423–432. doi: 10.1016/0022-2836(89)90007-7. [DOI] [PubMed] [Google Scholar]
- 14.Cavender J F, Conn A, Epler M, Lacko H, Tevethia M J. Simian virus 40 large T antigen contains two independent activities that cooperate with a ras oncogene to transform rat embryo fibroblasts. J Virol. 1995;69:923–934. doi: 10.1128/jvi.69.2.923-934.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chen D C, Yang B C, Kuo T T. One-step transformation of yeast in stationary phase. Curr Genet. 1992;21:83–84. doi: 10.1007/BF00318659. [DOI] [PubMed] [Google Scholar]
- 16.Czyz M, Nagiec M M, Dickson R C. Autoregulation of GAL4 transcription is essential for rapid growth of Kluyveromyces lactis on lactose and galactose. Nucleic Acids Res. 1993;21:4378–4382. doi: 10.1093/nar/21.18.4378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Denis C L, Ferguson J, Young E T. mRNA levels for the fermentative alcohol dehydrogenase of Saccharomyces cerevisiae decrease upon growth on a nonfermentable carbon source. J Biol Chem. 1983;258:1165–1171. [PubMed] [Google Scholar]
- 18.Ding W V, Johnston S A. The DNA binding and activation domains of Gal4p are sufficient for conveying its regulatory signals. Mol Cell Biol. 1997;17:2538–2549. doi: 10.1128/mcb.17.5.2538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Douglas H, Pelroy G. A gene controlling inducibility of the galactose pathway enzymes in Saccharomyces. Biochim Biophys Acta. 1963;68:155–156. [Google Scholar]
- 20.Douglas H C, Hawthorne C D. Uninducible mutants in the gal i locus of Saccharomyces cerevisiae. J Bacteriol. 1972;109:1139–1143. doi: 10.1128/jb.109.3.1139-1143.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Erhart E, Hollenberg C P. Curing of Saccharomyces cerevisiae 2-μm DNA by transformation. Curr Genet. 1981;3:83–89. doi: 10.1007/BF00365710. [DOI] [PubMed] [Google Scholar]
- 22.Erhart E, Hollenberg C P. The presence of a defective LEU2 gene on 2 mu DNA recombinant plasmids of Saccharomyces cerevisiae is responsible for curing and high copy number. J Bacteriol. 1983;156:625–635. doi: 10.1128/jb.156.2.625-635.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Fujiki M, Verner K. Coupling of cytosolic protein synthesis and mitochondrial protein import in yeast. Evidence for cotranslational import in vivo. J Biol Chem. 1993;268:1914–1920. [PubMed] [Google Scholar]
- 24.Gadhavi P L. Structural dissection of the DNA-binding domain of the yeast transcriptional activator GAL4 reveals an alpha-helical region responsible for dimerization. J Protein Chem. 1998;17:591–598. doi: 10.1007/BF02780960. [DOI] [PubMed] [Google Scholar]
- 25.Gietz D, St. Jean A, Woods R A, Schiestl R H. Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res. 1992;20:1425. doi: 10.1093/nar/20.6.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gietz R D, Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base-pair restriction sites. Gene. 1988;74:527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- 27.Giniger E, Varnum S M, Ptashne M. Specific DNA binding of GAL4, a positive regulatory protein of yeast. Cell. 1985;40:767–774. doi: 10.1016/0092-8674(85)90336-8. [DOI] [PubMed] [Google Scholar]
- 28.Guldener U, Heck S, Fielder T, Beinhauer J, Hegemann J H. A new efficient gene disruption cassette for repeated use in budding yeast. Nucleic Acids Res. 1996;24:2519–2524. doi: 10.1093/nar/24.13.2519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Harper J W, Adami G R, Wei N, Keyomarsi K, Elledge S J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell. 1993;75:805–816. doi: 10.1016/0092-8674(93)90499-g. [DOI] [PubMed] [Google Scholar]
- 30.Igarashi M, Segawa T, Nogi Y, Suzuki Y, Fukasawa T. Autogenous regulation of the Saccharomyces cerevisiae regulatory gene GAL80. Mol Gen Genet. 1987;207:273–279. doi: 10.1007/BF00331589. [DOI] [PubMed] [Google Scholar]
- 31.Irani M, Taylor W E, Young E T. Transcription of the ADH2 gene in Saccharomyces cerevisiae is limited by positive factors that bind competitively to its intact promoter region on multicopy plasmids. Mol Cell Biol. 1987;7:1233–1241. doi: 10.1128/mcb.7.3.1233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Johnston M, Carlson M. Regulation of carbon and phosphate utilization. II. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1992. [Google Scholar]
- 33.Johnston M, Dover J. Mutational analysis of the GAL4-encoded transcriptional activator protein of Saccharomyces cerevisiae. Genetics. 1988;120:63–74. doi: 10.1093/genetics/120.1.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Johnston S A, Hopper J E. Isolation of the yeast regulatory gene GAL4 and analysis of its dosage effects on the galactose/melibiose regulon. Proc Natl Acad Sci USA. 1982;79:6971–6975. doi: 10.1073/pnas.79.22.6971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Johnston S A, Salmeron J M, Jr, Dincher S S. Interaction of positive and negative regulatory proteins in the galactose regulon of yeast. Cell. 1987;50:143–146. doi: 10.1016/0092-8674(87)90671-4. [DOI] [PubMed] [Google Scholar]
- 36.Keegan L, Gill G, Ptashne M. Separation of DNA binding from the transcription-activating function of a eukaryotic regulatory protein. Science. 1986;231:699–704. doi: 10.1126/science.3080805. [DOI] [PubMed] [Google Scholar]
- 37.Koh S S, Ansari A Z, Ptashne M, Young R A. An activator target in the RNA polymerase II holoenzyme. Mol Cell. 1998;1:895–904. doi: 10.1016/s1097-2765(00)80088-x. [DOI] [PubMed] [Google Scholar]
- 38.Leuther K K, Johnston S A. Nondissociation of GAL4 and GAL80 in vivo after galactose induction. Science. 1992;256:1333–1335. doi: 10.1126/science.1598579. [DOI] [PubMed] [Google Scholar]
- 39.Leuther K K, Salmeron J M, Johnston S A. Genetic evidence that an activation domain of GAL4 does not require acidity and may form a beta sheet. Cell. 1993;72:575–585. doi: 10.1016/0092-8674(93)90076-3. [DOI] [PubMed] [Google Scholar]
- 40.Lohr D, Venkov P, Zlatanova J. Transcriptional regulation in the yeast GAL gene family: a complex genetic network. FASEB J. 1995;9:777–787. doi: 10.1096/fasebj.9.9.7601342. [DOI] [PubMed] [Google Scholar]
- 41.Lue N F, Chasman D I, Buchman A R, Kornberg R D. Interaction of GAL4 and GAL80 gene regulatory proteins in vitro. Mol Cell Biol. 1987;7:3446–3451. doi: 10.1128/mcb.7.10.3446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ma J, Ptashne M. The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell. 1987;50:137–142. doi: 10.1016/0092-8674(87)90670-2. [DOI] [PubMed] [Google Scholar]
- 43.Ma J, Ptashne M. Deletion analysis of GAL4 defines two transcriptional activating segments. Cell. 1987;48:847–853. doi: 10.1016/0092-8674(87)90081-x. [DOI] [PubMed] [Google Scholar]
- 44.Ma J, Ptashne M. A new class of yeast transcriptional activators. Cell. 1987;51:113–119. doi: 10.1016/0092-8674(87)90015-8. [DOI] [PubMed] [Google Scholar]
- 45.Marmorstein R, Carey M, Ptashne M, Harrison S C. DNA recognition by GAL4: structure of a protein-DNA complex. Nature. 1992;356:408–414. doi: 10.1038/356408a0. [DOI] [PubMed] [Google Scholar]
- 46.Melcher K, Johnston S A. GAL4 interacts with TATA-binding protein and coactivators. Mol Cell Biol. 1995;15:2839–2848. doi: 10.1128/mcb.15.5.2839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Meyer J, Walker-Jonah A, Hollenberg C P. Galactokinase encoded by GAL1 is a bifunctional protein required for induction of the GAL genes in Kluyveromyces lactis and is able to suppress the Gal3 phenotype in Saccharomyces cerevisiae. Mol Cell Biol. 1991;11:5454–5461. doi: 10.1128/mcb.11.11.5454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mylin L M, Hofmann K J, Schultz L D, Hopper J E. Regulated GAL4 expression cassette providing controllable and high-level output from high-copy galactose promoters in yeast. Methods Enzymol. 1990;185:297–308. doi: 10.1016/0076-6879(90)85026-k. [DOI] [PubMed] [Google Scholar]
- 49.Nogi Y. GAL3 gene product is required for maintenance of the induced state of the GAL cluster genes in Saccharomyces cerevisiae. J Bacteriol. 1986;165:101–106. doi: 10.1128/jb.165.1.101-106.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Nogi Y, Fukasawa T. Nucleotide sequence of the yeast regulatory gene GAL80. Nucleic Acids Res. 1984;12:9287–9298. doi: 10.1093/nar/12.24.9287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Nogi Y, Matsumoto K, Toh-e A, Oshima Y. Interaction of super-repressible and dominant constitutive mutations for the synthesis of galactose pathway enzymes in Saccharomyces cerevisiae. Mol Gen Genet. 1977;152:137–144. doi: 10.1007/BF00268810. [DOI] [PubMed] [Google Scholar]
- 52.Pan T, Halvorsen Y D, Dickson R C, Coleman J E. The transcription factor LAC9 from Kluyveromyces lactis-like GAL4 from Saccharomyces cerevisiae forms a Zn(II)2Cys6 binuclear cluster. J Biol Chem. 1990;265:21427–21429. [PubMed] [Google Scholar]
- 53.Peterson G L. A simplification of the protein assay method of Lowry et al. which is more generally applicable. Anal Biochem. 1977;83:346–356. doi: 10.1016/0003-2697(77)90043-4. [DOI] [PubMed] [Google Scholar]
- 54.Platt A, Reece R J. The yeast galactose genetic switch is mediated by the formation of a Gal4p-Gal80p-Gal3p complex. EMBO J. 1998;17:4086–4091. doi: 10.1093/emboj/17.14.4086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Post-Beittenmiller M A, Hamilton R W, Hopper J E. Regulation of basal and induced levels of the MEL1 transcript in Saccharomyces cerevisiae. Mol Cell Biol. 1984;4:1238–1245. doi: 10.1128/mcb.4.7.1238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Reece R J, Platt A. Signaling activation and repression of RNA polymerase II transcription in yeast. Bioessays. 1997;19:1001–1010. doi: 10.1002/bies.950191110. [DOI] [PubMed] [Google Scholar]
- 57.Riley M I, Hopper J E, Johnston S A, Dickson R C. GAL4 of Saccharomyces cerevisiae activates the lactose-galactose regulon of Kluyveromyces lactis and creates a new phenotype: glucose repression of the regulon. Mol Cell Biol. 1987;7:780–786. doi: 10.1128/mcb.7.2.780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Rose M D, Winston F, Heiter P. Methods in yeast genetics: a laboratory course manual. Plainview, N.Y: Cold Spring Harbor Laboratory Press; 1990. [Google Scholar]
- 59.Rose M D, Novick P, Thomas J H, Botstein D, Fink G R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60:237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- 60.Sadowski I, Costa C, Dhanawansa R. Phosphorylation of Gal4p at a single C-terminal residue is necessary for galactose-inducible transcription. Mol Cell Biol. 1996;16:4879–4887. doi: 10.1128/mcb.16.9.4879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sadowski I, Ma J, Triezenberg S, Ptashne M. GAL4-VP16 is an unusually potent transcriptional activator. Nature. 1988;335:563–564. doi: 10.1038/335563a0. [DOI] [PubMed] [Google Scholar]
- 62.Salmeron J M, Jr, Johnston S A. Analysis of the Kluyveromyces lactis positive regulatory gene LAC9 reveals functional homology to, but sequence divergence from the Saccharomyces cerevisiae GAL4 gene. Nucleic Acids Res. 1986;14:7767–7781. doi: 10.1093/nar/14.19.7767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Salmeron J M, Jr, Langdon S D, Johnston S A. Interaction between transcriptional activator protein LAC9 and negative regulatory protein GAL80. Mol Cell Biol. 1989;9:2950–2956. doi: 10.1128/mcb.9.7.2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Salmeron J M, Jr, Leuther K K, Johnston S A. GAL4 mutations that separate the transcriptional activation and GAL80-interactive functions of the yeast GAL4 protein. Genetics. 1990;125:21–27. doi: 10.1093/genetics/125.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sikorski R S, Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989;122:19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65a.Sil, A. K., et al. Unpublished data.
- 66.Silver P A, Keegan L P, Ptashine M. Amino terminus of the yeast GAL4 gene product is sufficient for nuclear localization. Proc Natl Acad Sci USA. 1984;81:5951–5955. doi: 10.1073/pnas.81.19.5951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Stone G, Sadowski I. GAL4 is regulated by a glucose-responsive functional domain. EMBO J. 1993;12:1375–1385. doi: 10.1002/j.1460-2075.1993.tb05782.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Suzuki-Fujimoto T, Fukuma M, Yano K I, Sakurai H, Vonika A, Johnston S A, Fukasawa T. Analysis of the galactose signal transduction pathway in Saccharomyces cerevisiae: interaction between Gal3p and Gal80p. Mol Cell Biol. 1996;16:2504–2508. doi: 10.1128/mcb.16.5.2504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Torchia T E, Hamilton R W, Cano C L, Hopper J E. Disruption of regulatory gene GAL80 in Saccharomyces cerevisiae: effects on carbon-controlled regulation of the galactose/melibiose pathway genes. Mol Cell Biol. 1984;4:1521–1527. doi: 10.1128/mcb.4.8.1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Torchia T E, Hopper J E. Genetic and molecular analysis of the GAL3 gene in the expression of the galactose/melibiose regulon of Saccharomyces cerevisiae. Genetics. 1986;113:229–246. doi: 10.1093/genetics/113.2.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Tornow J, Santangelo G M. Efficient expression of the Saccharomyces cerevisiae glycolytic gene ADH1 is dependent upon a cis-acting regulatory element (UASRPG) found initially in genes encoding ribosomal proteins. Gene. 1990;90:79–85. doi: 10.1016/0378-1119(90)90441-s. [DOI] [PubMed] [Google Scholar]
- 71a.Triezenberg, S. J. Personal communication.
- 72.Verner K, Weber M. Protein import into mitochondria in a homologous yeast in vitro system. J Biol Chem. 1989;264:3877–3879. [PubMed] [Google Scholar]
- 73.Vojtek A B, Hollenberg S M. Ras-Raf interaction: two-hybrid analysis. Methods Enzymol. 1995;255:331–342. doi: 10.1016/s0076-6879(95)55036-4. [DOI] [PubMed] [Google Scholar]
- 74.Webster T D, Dickson R C. The organization and transcription of the galactose gene cluster of Kluyveromyces lactis. Nucleic Acids Res. 1988;16:8011–8028. doi: 10.1093/nar/16.16.8011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74a.Woods, M. P., and J. E. Hopper. Unpublished data.
- 75.Wu Y, Reece R J, Ptashne M. Quantitation of putative activator-target affinities predicts transcriptional activating potentials. EMBO J. 1996;15:3951–3963. [PMC free article] [PubMed] [Google Scholar]
- 76.Yano K, Fukasawa T. Galactose-dependent reversible interaction of Gal3p with Gal80p in the induction pathway of Gal4p-activated genes of Saccharomyces cerevisiae. Proc Natl Acad Sci USA. 1997;94:1721–1726. doi: 10.1073/pnas.94.5.1721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Yocum R R, Johnston M. Molecular cloning of the GAL80 gene from Saccharomyces cerevisiae and characterization of a gal80 deletion. Gene. 1984;32:75–82. doi: 10.1016/0378-1119(84)90034-9. [DOI] [PubMed] [Google Scholar]
- 78.Zachariae W, Breunig K D. Expression of the transcriptional activator LAC9 (KlGAL4) in Kluyveromyces lactis is controlled by autoregulation. Mol Cell Biol. 1993;13:3058–3066. doi: 10.1128/mcb.13.5.3058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Zenke F T, Engles R, Vollenbroich V, Meyer J, Hollenberg C P, Breunig K D. Activation of Gal4p by galactose-dependent interaction of galactokinase and Gal80p. Science. 1996;272:1662–1665. doi: 10.1126/science.272.5268.1662. . (Erratum, 273:417, 1996.) [DOI] [PubMed] [Google Scholar]
- 80.Zenke F T, Kapp L, Breunig K D. Regulated phosphorylation of the Gal4p inhibitor Gal80p of Kluyveromyces lactis revealed by mutational analysis. Biol Chem. 1999;380:419–430. doi: 10.1515/BC.1999.056. [DOI] [PubMed] [Google Scholar]
- 81.Zenke F T, Zachariae W, Lunkes A, Breunig K D. Gal80 proteins of Kluyveromyces lactis and Saccharomyces cerevisiae are highly conserved but contribute differently to glucose repression of the galactose regulon. Mol Cell Biol. 1993;13:7566–7576. doi: 10.1128/mcb.13.12.7566. [DOI] [PMC free article] [PubMed] [Google Scholar]