Abstract
E2F is a family of transcription factors that regulates the cell cycle. It is widely accepted that E2F-mediated transactivation of a set of genes is the critical activity that governs cellular progression through G1 into S phase. In contrast to this hypothesis, we demonstrate that E2F actually suppresses the onset of S phase in two cell types when the cells are arrested by gamma irradiation. Our findings indicate that in these cells, the critical event triggering progression from G0/G1 arrest into S phase is the release of E2F-mediated transrepression of cell cycle genes, not transactivation by E2F. Furthermore, our data suggest that E2F-mediated transactivation is not necessary for the G1/S-phase transition in these cells.
The E2F proteins are transactivating factors that interact with the promoters of several genes whose expression is necessary for cell cycle progression, and it has been thought that E2F transactivation of a subset of these genes is necessary to drive the cell through G1 into S phase. E2F family members form complexes with the retinoblastoma protein (pRb), p107, and p130 (pocket proteins) during specific periods of the cell cycle (25). The transactivation function of E2F is inhibited when E2F is bound by pRb or one of the other pocket proteins. Since it is thought that transactivation by E2F is necessary for the transition from G1 to S phase, it has been accepted that inactivation of E2F-mediated transactivation by pocket proteins in this manner would be sufficient to inhibit cellular proliferation (45). Accordingly, complexes in which E2F is bound by pocket proteins were initially assumed to be transcriptionally inactive. However, it was subsequently found that these complexes are not inactive: they are now known to have transcriptional repressor activity. Thus, whereas it was thought that E2F-pocket protein complexes are impotent bystanders in the regulation of cell cycle gene expression, it is now clear that they have the potential to actively inhibit the expression of genes that contain binding sites for E2F in their promoters. The role of this repressor activity in cell cycle control is not fully understood.
The hypothesis that E2F transactivation is essential to drive cellular proliferation was initially derived from several studies that concluded that E2F-binding sites within promoters function primarily as enhancers. Many early studies, however, were performed either with minimal promoters (22, 23) or in the presence of DNA tumor virus proteins that affect E2F activity (e.g., adenovirus E1a or human papillomavirus E7) (3, 14, 22, 39, 43). It is now thought, however, that in the context of some promoters, E2F sites have no enhancer activity whatsoever; instead, in these promoters E2F sites are negative regulatory elements. Indeed, in the absence of DNA tumor virus proteins, E2F sites have been found to act as repressive elements in a large number of E2F-regulated cellular promoters (see Table 1). Furthermore, it has frequently been reported that the E2F sites in the dihydrofolate reductase (DHFR) promoter are enhancers; however, their activity was initially studied in HeLa cells, which are transformed by the DNA tumor virus oncoprotein E7 (3). Therefore, it is notable that the group that reported that the E2F sites in the DHFR promoter function as enhancers in HeLa cells subsequently reported that the same sites function solely as repressive elements in nontransformed fibroblasts (17). In light of the increasing recognition that E2F sites can function solely as repressive elements, it not surprising that Dyson recently asked, “Should we think of E2F-binding sites as activators of gene expression in S phase, or as elements that confer cell cycle regulated repression in G0/G1?” (8).
TABLE 1.
Studies in which E2F sites have been shown to act as repressive elements in cellular promoters
| Gene | Reference(s) |
|---|---|
| cdc-2 | 5, 40 |
| b-myb | 21, 53 |
| c-myc | N. Hays, personal communication |
| cdc-6 | 50 |
| DHFR | 17, 34, 35 |
| E2F-1 | 15, 18, 20, 24 |
| E2F-2 | 32 |
| E2F-3 | J. Nevins, personal communication |
| Cyclin A | 31 |
| Cyclin E | 27 |
| Insulin-like growth factor 1 | 29 |
| Orc1 | 26 |
| n-myc | 13 |
| Thymidine kinase | 7 |
| Retinoblastoma protein | 51 |
| p107 | 38 |
It has previously been shown that E2F overexpression is sufficient to drive rat fibroblasts that are arrested in G0/G1 by serum starvation into S phase and that this activity is dependent upon the transactivation function of E2F (19, 33). Thus, it was concluded that transactivation by E2F is necessary for progression into S phase. However, even overexpression of E2F fails to fully upregulate several S-phase genes in serum-starved cells (6). This suggests that serum starvation inhibits proliferation by targeting other cell-cycle-regulatory pathways in addition to the E2F pathway. Hence, the finding that E2F transactivation is necessary for the onset of S phase in serum-starved cells may be misleading in regard to conclusions about the role of E2F-mediated transactivation in the normal cell cycle; i.e., it is possible that E2F-mediated transactivation is required to compensate for the downregulation or inhibitory effect of another cell cycle-regulatory pathway in serum-starved cells. To better define the role of E2F in S-phase entrance, we sought a method of arresting cells that is more specific for the pocket protein-E2F pathway than is serum starvation.
Many types of cells arrest when they are treated with gamma irradiation during G1. However, Rb−/Rb− cells are defective in their response to irradiation in that they continue to progress into S phase after treatment with gamma irradiation during early G1 (4, 12, 36). Consequently, it has been suggested that Rb is an essential component of the arrest pathway (12). It has been postulated that the cell-cycle-regulatory activity of pRb is primarily mediated by its binding to E2F (45), so it seemed likely that the signal for gamma-irradiation-induced G1 arrest is specifically channeled through the Rb-E2F pathway. Therefore, we reasoned that gamma irradiation might be a more selective tool than serum starvation for studying the role of E2F in the regulation of progression from G0/G1 into S phase. Furthermore, the use of gamma irradiation arrests cells under conditions in which all cellular proteins are at physiological concentrations; thus, artifactual findings that can potentially occur when cell cycle arrest is brought about by overexpression of proteins such as pRb or cyclin-dependent kinase inhibitors are avoided.
To delineate the role of E2F in the progression from G0/G1 to S phase, we used a transfected-competitor strategy to block the effects of the endogenous E2F in gamma-irradiated cells. Plasmids and oligonucleotides that contain binding sites for specific transcription factors have been used by several groups in transfection assays to block transcription factor activity in vivo. In fact, competitor plasmids containing enhancer sequences were used by Schöler and Gruss to first demonstrate the existence of enhancer-binding transcription factors in vivo (30). They found that the promoter activity of transfected reporter constructs was decreased by cotransfection of competitor plasmids that contain the same enhancer sequences as the promoters in their reporter constructs. They hypothesized that this occurred because the competitor plasmids bound and sequestered trans-acting “cellular factors,” i.e., transcription factors, that were necessary for the activity of the enhancer regions of the promoters in the reporter plasmids. More recently, competitor oligonucleotides containing NF-κB binding sites were used by Nabel and colleagues to demonstrate that NF-κB has an essential role in phorbol ester-induced cellular adhesion (9). Transfection of these competitors blocked the integrin production and cellular adhesion that normally occur when HL-60 cells are treated with phorbol esters. This was thought to occur because the competitors sequestered and thereby functionally inactivated cellular NF-κB. The Nabel group has also found that transfection of octamer-binding sites inhibits interleukin-2 secretion by Jurkat T leukemia cells to a degree similar to that observed when the octamer site in the interleukin-2 enhancer is mutated (2). Similarly, transfection of oligonucleotides containing mef-1 binding sites has been found to block myogenesis and transfection of oligonucleotides containing PU.1 sites inhibits hematopoiesis (28, 42). Because many groups have been successful in using transcription factor-binding competitors to disrupt disparate cellular processes, we constructed plasmids that contain up to 96 E2F binding sites to use as competitors for sequestering and inactivating cellular E2F to elucidate the role of E2F in gamma-irradiation-induced G0/G1 arrest.
We report here that E2F becomes a potent transcriptional repressor in cells that are arrested in G0/G1 by gamma irradiation, and we examine the effect of sequestering E2F repressor complexes in these cells with E2F-binding competitor plasmids. We found that sequestration of the E2F complexes allows cells to bypass the gamma-irradiation-induced cell cycle block. These results indicate that E2F repressor activity is necessary for the gamma-irradiation-induced G0/G1 block. Another group using a different approach to disrupt E2F activity and different methods of arresting the cell cycle has recently found that E2F has an essential role in cell cycle arrest induced in cells lines other than those used in the present study (52). Thus, when considered together, these complementary studies indicate that E2F-mediated repression has a critical role in cell cycle arrest of a broad range of cells. Beyond this conclusion, the findings presented in our study strongly suggest that activation by E2F is not necessary for the G1-to-S-phase transition in certain cell lines.
MATERIALS AND METHODS
Plasmids.
To construct p24-E2F-COMPETITOR and p24-E2Fm-CONTROL, pGEM-3 (Promega) was altered to remove several potential E2F-binding sites. The sequence TTTGCCGG (nucleotides 850 to 857) was changed to TAAAACGG and the sequence TTTGCGGC was changed to TTTGCAGC (resulting in a silent mutation in the β-lactamase gene) by site-directed mutagenesis, and the fragment between the BsrBI site at nucleotide 2015 and the HindIII site (which contains several potential E2F-binding sites) was removed by restriction endonuclease digestion and subsequent blunt-end ligation. Twelve copies of the oligonucleotide containing two E2F-binding sites from the adenovirus E2A promoter or two mutant E2F sites (46) were cloned into sites in the multiple cloning sequence of the resulting vector. To construct pCOMP-4-E-E2F, pCOMP-12-E-E2F, pCOMP-24-E-E2F, pCOMP-48-E-E2F, and pCOMP-96-E-E2F, the oligonucleotide containing two E2F-binding sites was excised from pSKE2F (46) by restriction endonuclease digestion and the appropriate numbers of copies of this oligonucleotide were cloned into sites in the multiple cloning sequence of pGEM-3. pCOMP-12-E-E2Fm was constructed by cloning six copies of an oligonucleotide containing two mutant E2F-binding sites (46) into sites in the multiple cloning sequence of pGEM-3. The DHFR E2F site competitors were constructed by inserting the appropriate number of copies of the oligonucleotide CTGCAGTCTAGAGGTACCACAATTTCGCGCCAAACTTGACAATTTCG CGCCAAATTGGGTACCTCTAGACTGCAG into the multiple cloning sequence of pGEM-3. E2F sites are in boldface type, and sequences used for cloning are underlined. To construct p3E2F-CAT, the oligonucleotide AGCTTT TCGCGCTTAAATTTGAGAAAGTTTTCGCGCTTAAATTTGAGAAAGTT TTCGCGCTTAAATTGAGATCTATATATAG was cloned between the HindIII site and BamHI site (replacing the original sequence) of pE1bCAT (a gift from M. R. Green). E2F sites are in boldface type, and sequences used for cloning are underlined. p12sE1a and pRSVCAT have been described previously (46).
Tissue culture, transfections, reporter assays, and immunoblotting.
WS1 (ATCC CRL-1502), IMR90 (CCL-186), NRK-52E (ATCC CRL-1571), and Mv 1 Lu (ATCC CCL-64) cells were all maintained in Dulbecco modified Eagle medium with 10% fetal bovine serum. For luciferase assays, NRK-52E cells on 35-mm plates were transfected with 0.3 μg of the reporter plasmids and 5 μg of the competitor or control plasmids, as indicated, using Fugene 6 (Boehringer Mannheim) as directed by the manufacturer. Cells were harvested after 36 h, and luciferase assays were performed according to the manufacturer's directions (Promega). For chloramphenicol acyteltransferase (CAT) assays, Mv 1 Lu cells (ATCC) on 60-mm plates were transfected with 0.8 μg of the reporter plasmid, 0.8 μg of 12s E1a expression vector, and 15 μg of pCOMP-12-E-E2F or pCOMP-12-E-E2Fm, as indicated, by the calcium phosphate method and then harvested for CAT assays as described previously (45). For immunoblotting, cells were transfected as for CAT assays. Thirty-six hours after transfection, cells were harvested and immunoblotting was performed with anti-E1a polyclonal serum (Santa Cruz) as described previously (47).
EMSA.
Whole-cell extracts were prepared by passing cells in hypotonic lysis buffer (10 mM HEPES [pH 7.95], 400 mM KCl, 1 mM EDTA, 0.5 mM dithiothreitol [DTT], 5% glycerol, COMPLETE protease inhibitor [Boehringer Mannheim] and 10 μM NaVO4) through a hypodermic needle. Ten micrograms of total protein extract was incubated at room temperature with salmon sperm DNA and plasmid or oligonucleotide competitors, as indicated, in DNA-protein binding buffer (20 mM HEPES [pH 7.3], 50 mM KCl, 2.5 mM MgCl2, 0.5 mM EDTA, 0.5 mM DTT, 10% glycerol). After 5 min, 2 ng of a radiolabeled probe was added to the reaction mixture, followed by incubation for 20 min. Samples were electrophoresed through a 4% polyacrylamide gel (0.5× Tris-borate-EDTA) at 120 V for 1.5 h. The following oligonucleotides were annealed to complimentary oligonucleotides and used as probes and competitors: E2A, 5′-GGGATTTAAGTTTCGCGCCCTTTCTCAA-3′; Sp1, 5′-ATTCGATCGGGG CGGGGCGAGC-3′; NF-κB, 5′-AGTTGAGGGGACTTTCCCAGGC-3′; OCT1, 5′-TGTCGAATGCAAATCACTAGAA-3′; ATF, 5′-AGAGATTGCCTGACGTCAGAGAGCTAG-3′; E-box, 5′-CAGTATCACGTGTCATAGG-3′. Antibodies used in electrophoretic mobility shift assays (EMSA) all from Santa Cruz, were as follows: pRb, C-15; p107, C-18; p130, C-20; E2F-1, KH95; E2F-4, C-108. Two micrograms of each antibody was used per lane, where indicated (6 μg of antibody [total] for pocket protein cocktail), except that 6 μg of the E2F-1 antibody was used.
RT-PCR.
Reverse transcriptase (RT) reactions were performed with equal amounts of RNA isolated from IMR90 cells or gamma-irradiated IMR90 cells by using Retroscript (Ambion) as directed by manufacturer. One microliter from the RT reactions was added to each PCR. To confirm that the actin RT-PCR was in the linear range, PCR was performed with β-actin primers with both 0.1 and 1.0 μl of RT. All reactions were cycled 23 times, and the products were evaluated on ethidium bromide-stained agarose gels. All PCR products were of the expected size: 200, 530, 781, and 510 bp for b-myb, DHFR, thymidine kinase (TK), and β-actin, respectively. The following oligonucleotides were used as primers for RT-PCR: b-myb sense, 5′-GATGTGCCGGAGCAGAGGGATAG-3′; b-myb antisense, 5′-GTCCATGGCCCCTTGACAAGGTC-3′; DHFR sense, 5′-ATGGTTGGTTCGCTAAACTGCATC-3′; DHFR antisense, 5′-GAGAGAACACCTGGGTATTCTGGC-3′; TK sense, 5′-CGGGGGCAGATCCAGGTGATTC-3′; TK antisense, 5′-CCCAGAAGGCCAAGGTGTGG-3′; β-actin sense, 5′-GTGATGGTGGGCATGGGTCA-3′; β-actin antisense, 5′-TTAATGTCACGCACGATTTCCC-3′.
S-phase analysis.
WS1 or NRK52E cells on 12-well plates were transfected with the indicated amount of either competitor or control plasmid using Effectene (Qiagen) as directed by the manufacturer and then serum starved for 48 h. One hour after refeeding with medium containing 10% fetal bovine serum, cells were either sham treated or exposed to 500 rads of gamma irradiation. Fourteen hours after refeeding, BrdU (Sigma) was added to the medium. Eighteen hours after refeeding, the cells were fixed and stained for BrdU. The cells in randomly chosen fields were scored for BrdU until a total of 2,000 cells were evaluated for each transfection. To assess transfection efficiency in these experiments, 0.2 μg of competitor and control plasmids were cotransfected with 0.1 μg of pEGFP-C2 (Clontech) using Effectene (Qiagen) as directed by the manufacturer on a 24-well plate. After 24 h, representative fields were photographed.
RESULTS
E2F sites in the DHFR promoter function as repressors in quiescent cells.
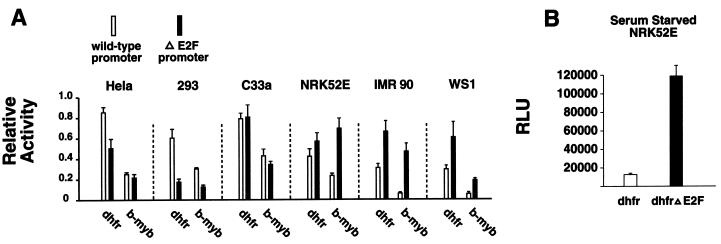
It has been hypothesized that E2F sites function as enhancers in some promoters and repressors in others; i.e., the E2F sites in the DHFR promoter are frequently described as enhancers and those in the b-myb promoter are thought to be repressors (8, 37). However, the composition of E2F complexes varies between cell types (1); thus, it is likely that E2F activity also varies between different types of cells. Indeed, we have previously demonstrated that E2F sites in synthetic promoters can act as either positive or negative elements in a cell type-dependent fashion (47). Additionally, whereas Azizkhan and colleagues did find that the E2F sites in the DHFR promoter are primarily activators in HeLa cells, they subsequently found that the E2F sites in the DHFR promoter function solely as repressors in fibroblasts (3, 17). To confirm and extend these studies, we examined the activity of the E2F sites in the DHFR and b-myb promoters in several cell types (Fig. 1A). We found that the net activity of the DHFR promoter E2F sites in proliferating cells depends upon the cell type in which they are examined. Most importantly, whereas the E2F sites in the DHFR promoter have been thought to be primarily transcriptional activators, we found that in nonimmortalized human fibroblasts, IMR90 and WS1, the net effect of the E2F sites in the DHFR promoter is repressive. Furthermore, whereas the E2F sites in the b-myb promoter are widely thought to function solely as repressors, we found that in some cells they are primarily activators. Thus it is clear that E2F site activity varies with cell type, and it is therefore possible that the precise role of E2F in the regulation of cellular proliferation varies with cell type. We also examined the activity of the E2F sites in the DHFR promoter in G0 cells. In proliferating NRK52E cells, the E2F sites in the DHFR promoter have little effect on the promoter's activity (Fig 1A); however, when these cells are made quiescent by serum starvation, the E2F sites in the DHFR promoter become transcriptional repressors (Fig. 1B). Thus, E2F site activity is dependent upon the cellular environment. In this context, we contend that E2F sites have the potential to act as repressors in all of the cellular promoters in which their activity has been studied to date (Table 1). This further underscores the importance of determining the role of E2F-mediated repression in control of the cell cycle.
FIG. 1.
The E2F sites in the DHFR promoter can act as transcriptional repressors. (A) A comparison of the activity of reporter constructs driven by wild-type DHFR and b-myb promoters to reporters driven by DHFR and b-myb promoters from which the E2F sites are deleted demonstrates that E2F site activity varies with cell type. Luciferase values were normalized between cell lines to facilitate presentation. (B) E2F sites are transcriptional repressors in serum-starved NRK52E cells. Twelve hours after transfection, the medium was replaced with serum-free medium; 48 h later, the cells were harvested for luciferase assays. Results are reported in relative light units (RLU).
Plasmids containing E2F-binding sites efficiently and specifically compete for E2F complexes.
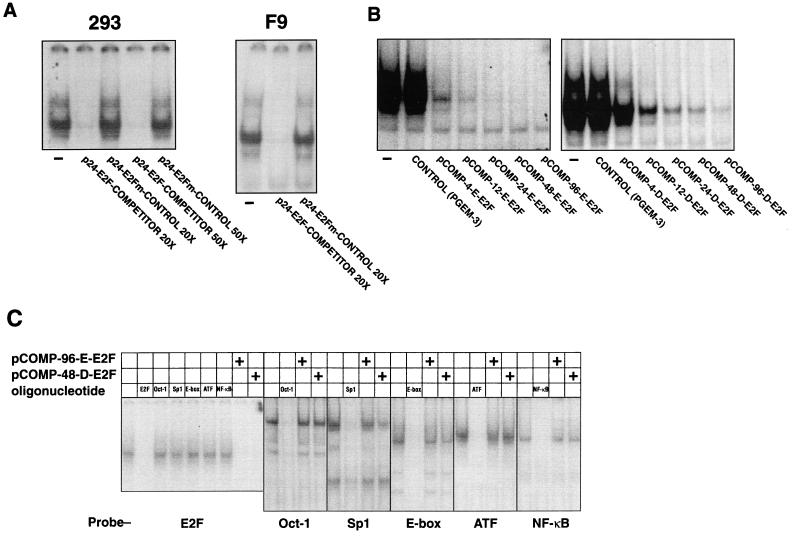
As outlined in the introduction, we have used a competitor plasmid strategy to block E2F activity in the cell. We first performed several assays to examine the effectiveness and specificity of such plasmids. We initially constructed two plasmids, p-24-E2F-COMPETITOR and p-24-E2Fm-CONTROL, that are exactly the same except that they contain 24 E2F binding sites and 24 mutant E2F binding sites, respectively (see Materials and Methods). We assessed the ability of each plasmid to sequester the E2F-binding activity from 293 and F9 cells by EMSA analysis. We chose these cell lines because they have a particularly large amount of the “free,” or transactivating, form of E2F; thus, they allowed us to test the ability of our competitor plasmids to bind the activating form of E2F. Whereas p-24-E2F-COMPETITOR efficiently competed for 293 and F9 cell E2F binding activity, p-24-E2Fm-CONTROL was completely ineffective (Fig. 2A). We then reasoned that the ability of an E2F competitor plasmid to sequester E2F should depend on the number of E2F sites contained in the plasmid. To test this hypothesis we made a series of constructs, using pGEM-3 (Promega) as a backbone, that contain 4, 8, 12, 24, 48, and 96 E2F sites, each with the sequence of those found in the E2a promoter (see Materials and Methods). These plasmids were designated pCOMP-4-E-E2F, pCOMP-8-E-E2F, pCOMP-12-E-E2F, pCOMP-24-E-E2F, pCOMP-48-E-E2F, and pCOMP-96-E-E2F (“E-E2F” indicates that the E2F-binding sites were derived from the E2a promoter), respectively. We also constructed a series of analogous plasmids that contain E2F sites with the sequence of those found in the human DHFR promoter; those plasmids are pCOMP-4-D-E2F, pCOMP-8-D-E2F, pCOMP-12-D-E2F, pCOMP-24-D-E2F, pCOMP-48-D-E2F, and pCOMP-96-D-E2F (“D-E2F” indicates that the E2F-binding sites were derived from the DHFR promoter). Both series of constructs were effective in competing for E2F-binding activity from 293 cells, and the efficiency with which each competed was dependent upon the number of E2F sites contained (Fig. 2B). We note that the E2F sites from the E2a promoter were consistently more effective in binding E2F than were the E2F sites derived from the DHFR promoter; however, the DHFR-E2F competitors can block all E2F-binding activity (Fig. 2C). The activity of both series of competitor plasmids in these assays was highly specific since representative plasmids from each series, pCOMP-96-E-E2F and pCOMP-48-D-E2F, did not compete effectively for any of several other transcription factors under the same conditions in which they completely sequestered E2F (Fig. 2C). Thus, a variety of competitor plasmids that contain E2F sites can sequester E2F efficiently and specifically.
FIG. 2.
Competitor plasmids containing E2F sites bind E2F efficiently and specifically as assessed by EMSA. (A) A plasmid that contains 24 E2F-binding sites efficiently binds the E2F in lysates from 293 and F9 cells, whereas a plasmid that is exactly the same except that it contains 24 mutant E2F sites is ineffective in binding E2F. The plasmid concentrations were adjusted so that the molar concentration of plasmid E2F sites was either 20:1 or 50:1 (as indicated) compared with the molar concentration of the radiolabeled E2F probe. (B) The efficiency with which a plasmid competes for E2F is dependent on the number of E2F sites it contains. One hundred nanograms of each of the competitor plasmids was used in the assays. Two nanograms of a radiolabeled oligonucleotide containing binding sites for E2F was used as a probe. (C) Under the same conditions that E2F competitor plasmids sequester E2F, they fail to compete for other transcription factors. Radiolabeled oligonucleotides containing the indicated transcription factor-binding sites were used as probes. Either 100 ng of E2F competitor plasmid or 50 ng of unlabeled oligonucleotide competitor was used, as indicated.
Competitor plasmids containing E2F sites readily block E2F transactivation activity in vivo.
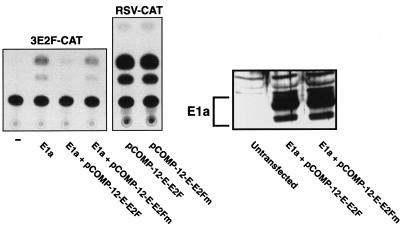
To determine if such competitor plasmids effectively block E2F function in vivo, we first examined their effect on a reporter plasmid that is driven by three E2F-binding sites (p3E2F-CAT) in the presence of E1a, an adenovirus protein that releases E2F from pocket proteins converting cellular E2F to its free, transactivating form (1). As expected, E1a expression activated p3E2F-CAT (Fig. 3). When an E2F competitor plasmid with 12 E2F-binding sites was cotransfected, however, activation was blocked. In contrast, cotransfection of a plasmid with 12 mutant E2F sites with the E1a expression vector was without effect and, as we found in our in vitro assays, the ability of the competitor plasmid to bind E2F in vivo is specific, since transfection of the E2F competitor plasmid had no effect on the activity of a promoter that does not contain E2F-binding sites (Fig. 3). Thus, E2F competitor plasmids efficiently bind, sequester, and thereby inactivate the transactivating forms of E2F in vivo, even when endogenous E2F activity is maximally activated by E1a.
FIG. 3.
E2F competitor plasmids block E2F activity in vivo. A minimal reporter construct containing three E2F sites and a TATA box (p3E2F-CAT) is activated by E1a. A competitor plasmid containing 12 E2F-binding sites blocks E2F-mediated transactivation, whereas a plasmid containing 12 mutant E2F sites does not. The effect of the competitor plasmid is specific for E2F since it has no effect on RSV-CAT, a reporter plasmid that lacks E2F sites. An immunoblot for E1a indicates that the competitor plasmid does not affect the level of E1a expression.
E2F sites are efficient silencers in gamma-irradiated cells.
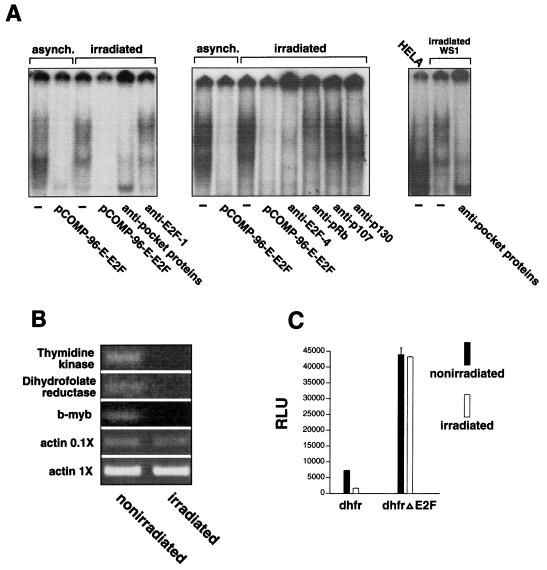
As outlined in the introduction, we used gamma-irradiated cells as tools to study the role of E2F in cellular progression from G0/G1 to S phase. We first examined the change in E2F-binding activity that occurs when cells are treated with gamma irradiation. Gamma irradiation of G1 fibroblasts resulted in a marked alteration of E2F-binding activity when assessed by EMSA. In general, the E2F complexes from gamma-irradiated cells migrated more slowly than those from asynchronously growing cells (Fig. 4A, left and middle panels), suggesting that gamma irradiation induces formation of new pocket protein-E2F complexes or changes in preexisting pocket protein-E2F complexes. Indeed, the E2F was almost completely bound by pocket proteins in the gamma-irradiated cells as evidenced by the finding that the migration of all E2F-binding activity was altered by a cocktail of antibodies to pRb, p107, and p130, whereas the same amount of an E2F-1 antibody, used as a control, had little effect on complex migration (Fig. 4A, left panel). Only E2F that is not bound by pocket proteins can function as a transactivator; hence, our data suggest that there is little if any of the transactivating form of E2F in cells arrested in G0/G1 by gamma irradiation. In fact, we found that most of the E2F activity in irradiated cells was E2F-4 (Fig. 4A, middle panel), an E2F family member that has been implicated in transcriptional repression, not activation (25). To determine which of the pocket proteins bind to E2F in gamma-irradiated cells, we added antibodies to pRb, p107, and p130 individually to the cell lysates from gamma-irradiated cells and then performed EMSA. These assays indicated that the E2F in gamma-irradiated cells is primarily bound by pRb and p130, since the pRb antibody altered the faster-migrating complexes and the p130 antibody altered the slower-migrating complexes, whereas the p107 antibody had little if any effect on complex migration (Fig. 4A, middle panel). Finally, we compared the migration of the E2F from HeLa cells with the migration of E2F complexes from gamma-irradiated fibroblasts. Because the oncoprotein E7 is expressed in HeLa cells, almost all of the E2F in these cells is in the free, transactivating form. A comparison of the E2F from HeLa cells with the E2F from gamma-irradiated fibroblasts again indicates that there is little free E2F in gamma-irradiated cells (Fig. 4A, right panel). Thus, most, if not all, of the E2F in gamma-irradiated cells is found in complexes that have the potential to act as transrepressors.
FIG. 4.
The E2F in cells treated with gamma irradiation is a potent transcriptional repressor. (A) There is little free E2F in irradiated WS1 cells. A cocktail containing antibodies to the three pocket proteins alters the migration of all of the E2F complexes in an EMSA using lysates from cells treated with 1,000 rads of gamma irradiation, whereas an equal amount of E2F-1 antibody used as a control has only a minimal effect (left panel). Almost all of the E2F in irradiated cells is E2F-4; pRb and p130 are the predominant pocket proteins found in E2F complexes in irradiated cells (middle panel). A comparison of the E2F from irradiated WS1 cells with the E2F from HeLa cells confirms that there is little if any free E2F in the irradiated cells (right panel). (B) The mRNAs from three genes that are thought to be regulated by E2F, the thymidine kinase, DHFR, and b-myb genes, are markedly reduced in irradiated IMR 90 cells as assessed by RT-PCR. RT-PCR of β-actin was used as a control for RNA loading. The PCR for β-actin was performed with two different concentrations of the RT products to confirm that the PCR was in the linear range. The contrast and magnification of the images of the thymidine kinase, DHFR, and b-myb bands was increased to facilitate visualization (the same changes were applied to the entire image). (C) The activity of a DHFR reporter construct is severely repressed in irradiated NRK52E cells compared to the activity of the same construct in cells that are not irradiated, whereas a DHFR promoter construct from which the E2F sites are deleted is unaffected by radiation.
Finally, to determine if E2F functions as a transcriptional repressor in gamma-irradiated cells, we examined the activity of E2F-promoter binding sites in cells that were gamma-irradiated. We first examined the expression of three genes that are thought to be regulated by E2F, the DHFR, thymidine kinase, and b-myb genes, in irradiated cells. Gamma irradiation resulted in a sharp downregulation of expression of these genes in human fibroblasts (Fig. 4B). Next, we transfected cells with reporter constructs driven by the DHFR promoter and a mutant DHFR promoter from which the E2F sites had been deleted. These cells were first synchronized and then treated with gamma irradiation during early G1. The activity of the wild-type DHFR promoter was markedly repressed in irradiated cells compared with its activity in cells that were not irradiated (Fig. 4C). The inhibition of promoter activity in the irradiated cells was mediated by the E2F sites because the mutant DHFR promoter construct that lacked E2F sites was unaffected by gamma irradiation. Most importantly, the E2F in gamma-irradiated cells is a potent transcriptional repressor as evidenced by the finding that the wild-type promoter was much less active than the mutant DHFR promoter that lacked E2F sites (Fig 4C). Thus, the E2F in gamma-irradiated cells is bound by pRb and p130 and functions as a transcriptional repressor.
E2F is required for the G0/G1 block induced by gamma irradiation.
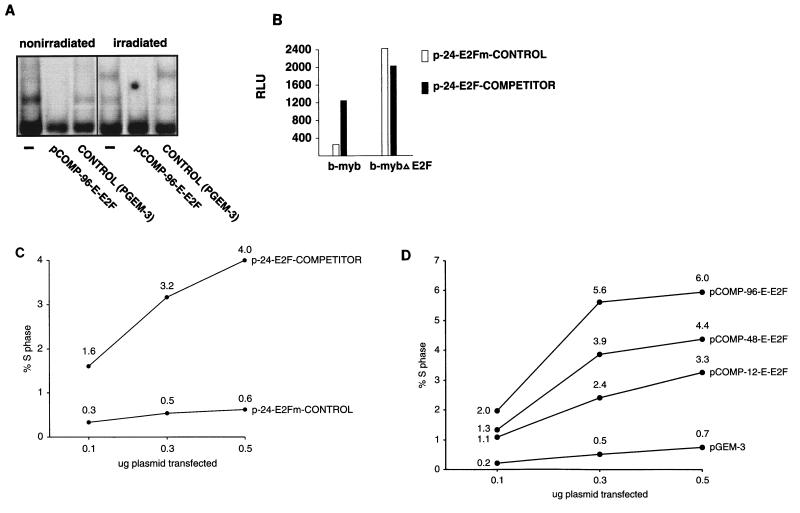
pCOMP-96-E-E2F binds all E2F complexes in gamma-irradiated fibroblasts (Fig. 5A). We show here and have previously demonstrated that transfected E2F competitor plasmids can relieve E2F-mediated transcriptional repression in vivo (Fig. 5B and reference 46, Fig. 1). Therefore, we used E2F competitor plasmids to assess the role of gamma-irradiation-induced E2F-repressor complexes in the G0/G1 block of gamma-irradiated cells. We first used WS1 cells, a human diploid skin fibroblast cell line, because they are derived from normal tissue, they are nonimmortalized, and they undergo arrest for a prolonged period in response to gamma irradiation, and we have found that they transfect at high efficiency (15 to 30% of WS1 cells are transfected when assessed by transfection of a green fluorescent protein (GFP) expression vector [data not shown]). When WS1 cells synchronized by serum starvation were released into G1 by serum refeeding, 20 to 40% of the cells entered S phase within 18 h (data not shown; see analogous data for NRK52E cells in Fig. 5E). As expected, WS-1 cells that were transfected with a control plasmid (p-24-E2Fm-CONTROL) and then irradiated in G1 did not enter S phase (Fig. 5C). Strikingly, however, transfection of the competitor plasmid p-24-E2F-COMPETITOR prior to irradiation relieved the block, as evidenced by the finding that a significant number of cells entered S phase and the number of cells entering S phase correlated with the amount of p-24-E2F-COMPETITOR transfected (Fig. 5C). It is important to note that these are transient transfections; thus, the majority of cells will remain arrested because they are not transfected. However, we have now repeated this experiment in WS1 cells more than 15 times with several different competitor and control plasmid preparations, and the results are highly reproducible and completely unambiguous. Indeed, we consistently find that approximately 10-fold-more WS1 cells transfected with pCOMP-96-E-E2F enter S phase than cells transfected with control plasmids (Fig. 5D). Furthermore, the efficiency with which the competitor plasmids relieve the G0/G1 block correlates with the number of E2F sites in the competitor plasmid (Fig. 5D). This is consistent with our finding that the efficiency with which competitor plasmids compete for E2F binding is dependent upon the number of E2F sites they contain (Fig 2B). We have also used several different transfection reagents (data not shown) and obtained similar results. The competitor plasmids also had the same effect in a second cell line (Fig. 5E and below). Finally, transfection of competitor plasmids containing the E2F sites from the DHFR promoter (instead of those from the E2a promoter) also release WS1 cells from the gamma-irradiation-induced G0/G1 block (data not shown; see data for NRK52E cells in Fig. 5E and below). These results indicate that the G0/G1-to-S-phase block in gamma-irradiated cells is subverted when E2F is sequestered by the competitor. Furthermore, since sequestration of E2F functionally inactivates it (e.g., see Fig. 3 and 5B and reference 46, Fig. 1) and the E2F in gamma-irradiated cells functions as a transcriptional repressor, these results indicate that the G0/G1-to-S-phase block in gamma-irradiated cells must be dependent upon E2F-mediated transcriptional repression, not simple inactivation of E2F. Therefore, E2F is required to prevent exit from G0/G1 in gamma-irradiated cells.
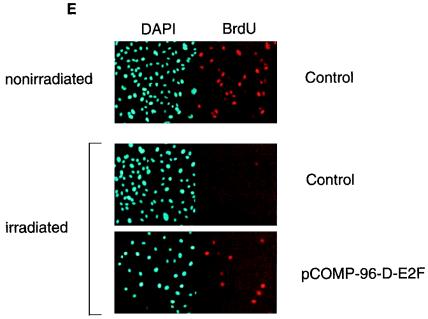
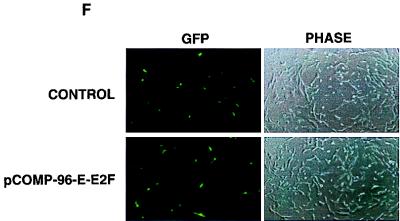
FIG. 5.
E2F-mediated transcriptional repressor activity is necessary for gamma-irradiation-induced G0/G1 block. (A) An E2F competitor plasmid effectively sequesters the E2F from both nonirradiated and irradiated WS1 cells as assessed by EMSA. (B) An E2F competitor plasmid relieves repression of an E2F-regulated reporter gene. The activity of a b-myb luciferase construct increased approximately fourfold when cotransfected with p-24-E2F-COMPETITOR compared with its activity when cotransfected with p-24-E2Fm-CONTROL. p-24-E2F-COMPETITOR had no effect on the activity of a b-myb luciferase construct from which the E2F sites had been deleted. (C) Transfection of plasmids that bind E2F release gamma-irradiated cells from a G0/G1 block. Cells were transfected as indicated, serum starved for 48 h, refed, irradiated, and then assessed for S-phase entry. Values for each data point are shown on the graph. This experiment has now been repeated more than 15 times with WS1 cells using several different preparations of competitor plasmids and controls. The results are completely reproducible. (D) The efficiency with which a competitor plasmid releases cells from the gamma-irradiation-induced G0/G1 block depends upon both the number of E2F sites the competitor plasmid contains and the amount of competitor plasmid transfected. This experiment was performed with WS1 cells, as in panel C. (E) Competitor plasmids containing the E2F sites from the DHFR promoter release NRK52E cells from the gamma-irradiation-induced G0/G1 block. The experiment was performed as in panel C except that NRK52E cells were transfected with p-COMP-96-D-E2F (contains E2F sites from the DHFR promoter) and pGEM-3 (control), as indicated. The views shown are typical fields. This experiment has now been repeated more than eight times with NRK52E cells with several different preparations of competitor plasmids and controls, including three times with p-24-E2Fm-CONTROL and p-24-E2F-COMPETITOR. The results are completely reproducible. (F) pCOMP-96-E-E2F and pGEM-3 transfect with equal efficiency. An excess of each of these plasmids was cotransfected with a GFP expression vector, and representative fields were photographed after 24 h. p24-E2F-COMPETITOR and p24-E2Fm-CONTROL were assessed in the same manner and were found to transfect with the same efficiency (data not shown).
We next examined the effect of sequestering cellular E2F using pCOMP-96-D-E2F on the response to gamma irradiation of NRK52E cells. NRK52E is a nontransformed epithelioid cell line that is derived from rat kidney (16). We found that transfection of pCOMP-96-D-E2F released NRK52E cells from the G0/G1 block induced by gamma irradiation, whereas a control plasmid (pGEM-3) had no effect (Fig. 5E). The same results were obtained whether we used pCOMP-96-D-E2F or pCOMP-96-E-E2F (data not shown). Importantly, we have also performed the same experiments in NRK52E cells using p-24-E2F-COMPETITOR and p-24-E2Fm-CONTROL three times and obtained results similar (data not shown) to those shown for WS1 cells in Fig. 5C. Therefore, we have found that a variety of E2F competitor plasmids are effective in releasing cells from the gamma-irradiation induced G0/G1 block.
Finally, to demonstrate that the competitor plasmids and control plasmids we used transfect with equal efficiency, we transfected an excess of each along with a GFP expression vector. The transfection efficiency is the same for both the competitor and control plasmids (Fig. 5F). Therefore, we have clearly demonstrated in two unrelated mammalian cell types that E2F-repressor complexes have a critical role in maintaining the gamma-irradiation-induced G0/G1 cell cycle block.
DISCUSSION
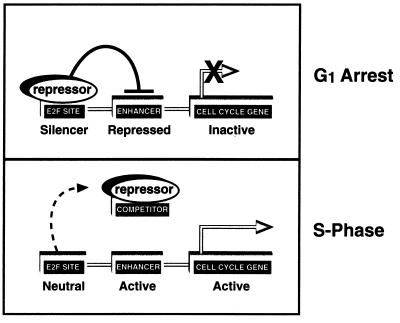
In contrast to the accepted model of cell cycle regulation in which transactivation by E2F is thought to regulate cell cycle progression, our data suggest that at least in some cells E2F may function as an “off switch,” limiting proliferation by repressing transcription of growth-promoting genes. In this model, E2F site-mediated repression regulates cell cycle progression by inhibiting promoter activity that would otherwise function to drive the cell into S phase (Fig. 6). If this model proves correct, it would at least in part resolve the phenotypic conundrum presented by the E2F-1 knockout mouse (11, 49). These mice exhibit hyperplasia, neoplasia, and a decreased level of apoptosis. Clearly, these characteristics are evidence against a model of cell cycle control in which E2F drives proliferation.
FIG. 6.
Model for the mechanism by which a transfected competitor plasmid releases cells from the gamma-irradiation-induced G0/G1 block. The upper panel is supported by our finding that the E2F in gamma-irradiated cells is a potent transcriptional repressor, and the lower panel is supported by our finding that competitor plasmids containing E2F-binding sites bind and sequester E2F complexes in vitro and in vivo.
Expression of E2F-1, E2F-2, and E2F-3 is upregulated as cells progress through G1, and it has been proposed that transactivation mediated by these proteins is necessary for the onset of S phase (25). It is important to note that our experiments do not formally exclude the possibility that the competitor plasmids fail to completely inhibit the transactivating activity associated with E2F-1, E2F-2, and E2F-3 and that the residual transactivating activity drives gene expression that is necessary for the G1-to-S-phase transition. We find this scenario unlikely, however, for the following reasons. (i) Even a competitor plasmid with only 12 E2F sites blocks maximal levels of E2F transactivation, as evidenced by the finding that it blocks E1a activation of an E2F-driven reporter construct. It is likely that the amount of free E2F in E1a-expressing cells far exceeds that found in cells progressing normally through G1, so it is doubtful that the transactivating E2F in irradiated cells exceeds the E2F-binding capacity of pCOMP-96-E-E2F. (ii) The competitors readily bind free E2F, as assessed by EMSA of 293 and F9 cell lysates (Fig. 2A). (iii) Most of the E2F found in quiescent and proliferating cells is E2F-4, and the level of E2F-4 does not vary significantly during the cell cycle (41). It is highly unlikely that the competitor plasmids titrate out enough E2F-4 to release the cells from a G0 block without also binding the relatively small amount of additional E2F activity that is expressed during G1. (iv) There is a positive correlation between the number of E2F sites in the transfected competitor and the number of cells that enter S phase after gamma irradiation (Fig. 5D). If E2F-mediated transactivation was necessary for entry into S phase, it would be expected that beyond a certain point the number of cells entering S phase would fall off as the number of E2F sites in the competitors increased. We have not found this to be so. Considering these arguments, our data strongly suggests that E2F-mediated transactivation is not necessary for the G1-to-S-phase transition under the conditions of our experiments.
We also note that there is convincing evidence that E2F-mediated transactivation plays a role in cell cycle progression. For example, the proportion of SAOS-2 cells in G1 increases when E2F activity is inhibited (10, 48). SAOS-2 cells, however, are transformed cells that do not express pRb. The loss of pRb is likely to increase the overall level of E2F transactivation activity in these cells above the level found in normal, pRb-expressing cells, and it is probable that the transformed phenotype in part results from and is dependent upon this increase. Indeed, we have shown that E2F sites are more effective enhancers in cells that do not express wild-type pRb, such as SAOS-2 cells (46). Thus, it may be that the proportion of SAOS-2 cells in G1 increases when E2F-mediated transactivation is inhibited because they lose some of the features characteristic of transformed cells, such as an accelerated growth rate. Indeed, it has been pointed out that in these and similar experiments, S-phase entry is not completely eliminated (8). It may be that inhibition of E2F-mediated transactivation just slows progression through the cell cycle. In this context, it is tempting to speculate that a certain threshold of E2F-regulated gene expression must be exceeded for progression through G1 and into S phase and that further increases of expression serve to increase the rate of progression through G1. This is supported by the recent finding that even though E2F-1 is not necessary for the G1-to-S-phase progression of cycling cells, it does contribute to the rate of progression from G0 to S phase (44). It is certainly conceivable that submaximal levels of E2F-regulated gene expression are sufficient to allow S-phase entry and that these levels are readily achieved by release of E2F-mediated repression.
ACKNOWLEDGMENTS
We thank Roger Watson and Peggy Farnham for gifts of the b-myb and DHFR plasmids and L. Carayannopoulos and N. Levy for review of the manuscript.
S.J.W. is supported in part by the NIH and the American Lung Association, and U.W. received support from the Deutscher Akademischer Austauschdienst.
S.H., B.L.C., B.E.D., and U.W. contributed equally to this work.
REFERENCES
- 1.Bagchi S, Raychaudhuri P, Nevins J R. Adenovirus E1A proteins can dissociate heteromeric complexes involving the E2F transcription factor: a novel mechanism for E1A transactivation. Cell. 1990;62:659–669. doi: 10.1016/0092-8674(90)90112-r. [DOI] [PubMed] [Google Scholar]
- 2.Bielinska A, Shivdasani R A, Zhang L Q, Nabel G J. Regulation of gene expression with double-stranded phosphorothioate oligonucleotides. Science. 1990;250:997–1000. doi: 10.1126/science.2237444. [DOI] [PubMed] [Google Scholar]
- 3.Blake M C, Azizkhan J C. Transcription factor E2F is required for efficient expression of the hamster dihydrofolate reductase gene in vitro and in vivo. Mol Cell Biol. 1989;9:4994–5002. doi: 10.1128/mcb.9.11.4994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brugarolas J, Moberg K, Boyd S D, Taya Y, Jacks T, Lees J A. Inhibition of cyclin-dependent kinase 2 by p21 is necessary for retinoblastoma protein-mediated G1 arrest after gamma-irradiation. Proc Natl Acad Sci USA. 1999;96:1002–1007. doi: 10.1073/pnas.96.3.1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dalton S. Cell cycle regulation of the human cdc2 gene. EMBO J. 1992;11:1797–1804. doi: 10.1002/j.1460-2075.1992.tb05231.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.DeGregori J, Kowalik T, Nevins J R. Cellular targets for activation by the E2F1 transcription factor include DNA synthesis- and G1/S-regulatory genes. Mol Cell Biol. 1995;15:4215–4224. doi: 10.1128/mcb.15.8.4215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dou Q P, Zhao S, Levin A H, Wang J, Helin K, Pardee A B. G1/S-regulated E2F-containing protein complexes bind to the mouse thymidine kinase gene promoter. J Biol Chem. 1994;269:1306–1313. [PubMed] [Google Scholar]
- 8.Dyson N. The regulation of E2F by pRB-family proteins. Genes Dev. 1998;12:2245–2262. doi: 10.1101/gad.12.15.2245. [DOI] [PubMed] [Google Scholar]
- 9.Eck S L, Perkins N D, Carr D P, Nabel G J. Inhibition of phorbol ester-induced cellular adhesion by competitive binding of NF-κB in vivo. Mol Cell Biol. 1993;13:6530–6536. doi: 10.1128/mcb.13.10.6530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Fan J G, Bertino J R. Functional roles of E2F in cell cycle regulation. Oncogene. 1997;14:1191–1200. doi: 10.1038/sj.onc.1200940. [DOI] [PubMed] [Google Scholar]
- 11.Field S J, Tsai F Y, Kuo F, Zubiaga A M, Kaelin W G, Jr, Livingston D M, Orkin S H, Greenberg M E. E2F-1 functions in mice to promote apoptosis and suppress proliferation. Cell. 1996;85:549–561. doi: 10.1016/s0092-8674(00)81255-6. [DOI] [PubMed] [Google Scholar]
- 12.Harrington E A, Bruce J L, Harlow E, Dyson N. pRB plays an essential role in cell cycle arrest induced by DNA damage. Proc Natl Acad Sci USA. 1998;95:11945–11950. doi: 10.1073/pnas.95.20.11945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hiebert S W, Blake M, Azizkhan J, Nevins J R. Role of E2F transcription factor in E1a-mediated transactivation of cellular genes. J Virol. 1991;65:3547–3552. doi: 10.1128/jvi.65.7.3547-3552.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hiebert S W, Lipp M, Nevins J R. E1A-dependent transactivation of the human MYC promoter is mediated by the E2F factor. Proc Natl Acad Sci USA. 1989;86:3594–3598. doi: 10.1073/pnas.86.10.3594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hsiao K M, McMahon S L, Farnham P J. Multiple DNA elements are required for the growth regulation of the mouse E2F1 promoter. Genes Dev. 1994;8:1526–1537. doi: 10.1101/gad.8.13.1526. [DOI] [PubMed] [Google Scholar]
- 16.Huu D-N, Rosenblum E N, Zeigel R F. Persistent infection of a rat kidney cell line with Rauscher murine leukemia virus. J Bacteriol. 1966;92:1133–1140. doi: 10.1128/jb.92.4.1133-1140.1966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jensen D E, Black A R, Swick A G, Azizkhan J C. Distinct roles for Sp1 and E2F sites in the growth/cell cycle regulation of the dhfr promoter. J Cell Biochem. 1997;67:24–31. doi: 10.1002/(sici)1097-4644(19971001)67:1<24::aid-jcb3>3.0.co;2-y. [DOI] [PubMed] [Google Scholar]
- 18.Johnson D G. Regulation of E2F-1 gene expression by p130 (Rb2) and D-type cyclin kinase activity. Oncogene. 1995;11:1685–1692. [PubMed] [Google Scholar]
- 19.Johnson D G, Schwarz J K, Cress W D, Nevins J R. Expression of transcription factor E2F1 induces quiescent cells to enter S phase. Nature. 1993;365:349–352. doi: 10.1038/365349a0. [DOI] [PubMed] [Google Scholar]
- 20.Johnson D G, Ohtani K, Nevins J R. Autoregulatory control of E2F1 expression in response to positive and negative regulators of cell cycle progression. Genes Dev. 1994;8:1514–1525. doi: 10.1101/gad.8.13.1514. [DOI] [PubMed] [Google Scholar]
- 21.Lam E W, Watson R J. An E2F-binding site mediates cell-cycle regulated repression of mouse b-myb transcription. EMBO J. 1993;12:2705–2713. doi: 10.1002/j.1460-2075.1993.tb05932.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Moberg K H, Logan T J, Tyndall W A, Hall D J. Three distinct elements within the murine c-myc promoter are required for transcription. Oncogene. 1992;7:411–421. [PubMed] [Google Scholar]
- 23.Mudryj M, Hiebert S W, Nevins J R. A role for the adenovirus inducible E2F transcription factor in a proliferation dependent signal transduction pathway. EMBO J. 1990;9:2179–2184. doi: 10.1002/j.1460-2075.1990.tb07387.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Neuman E, Flemington E K, Sellers W R, Kaelin W G., Jr Transcription of the E2F-1 gene is rendered cell cycle dependent by E2F DNA-binding sites within its promoter. Mol Cell Biol. 1994;14:6607–6615. doi: 10.1128/mcb.14.10.6607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Nevins J R. Toward an understanding of the functional complexity of the E2F and retinoblastoma families. Cell Growth Differ. 1998;9:585–593. [PubMed] [Google Scholar]
- 26.Ohtani K, DeGregori J, Leone G, Herendeen D R, Kelly T J, Nevins J R. Expression of the HsOrc1 gene, a human ORC1 homolog, is regulated by cell proliferation via the E2F transcription factor. Mol Cell Biol. 1996;16:6977–6984. doi: 10.1128/mcb.16.12.6977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ohtani K, DeGregori J, Nevins J R. Regulation of the cyclin E gene by transcription factor E2F1. Proc Natl Acad Sci USA. 1995;92:12146–12150. doi: 10.1073/pnas.92.26.12146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Parekh R, Bag J. Inhibition of myogenesis in mouse C2 cells by double-stranded phosphorothioate oligodeoxynucleotides containing mef-1 sequence. J Biol Chem. 1996;271:17372–17376. doi: 10.1074/jbc.271.29.17372. [DOI] [PubMed] [Google Scholar]
- 29.Porcu P, Grana X, Li S, Swantek J, De Luca A, Giordano A, Baserga R. An E2F binding sequence negatively regulates the response of the insulin-like growth factor I (IGF-I) promoter to simian virus 40 T antigen and to serum. Oncogene. 1994;9:2125–2134. [PubMed] [Google Scholar]
- 30.Schöler H R, Gruss P. Specific interaction between enhancer-containing molecules and cellular components. Cell. 1984;36:403–411. doi: 10.1016/0092-8674(84)90233-2. [DOI] [PubMed] [Google Scholar]
- 31.Schulze A, Zerfass K, Spitkovsky D, Middendorp S, Berges J, Helin K, Jansen-Durr P, Henglein B. Cell cycle regulation of the cyclin A gene promoter is mediated by a variant E2F site. Proc Natl Acad Sci USA. 1995;92:11264–11268. doi: 10.1073/pnas.92.24.11264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sears R, Ohtani K, Nevins J R. Identification of positively and negatively acting elements regulating expression of the E2F2 gene in response to cell growth signals. Mol Cell Biol. 1997;17:5227–5235. doi: 10.1128/mcb.17.9.5227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shan B, Lee W H. Deregulated expression of E2F-1 induces S-phase entry and leads to apoptosis. Mol Cell Biol. 1994;14:8166–8173. doi: 10.1128/mcb.14.12.8166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Slansky J E, Farnham P J. Transcriptional regulation of the dihydrofolate reductase gene. Bioessays. 1996;18:55–62. doi: 10.1002/bies.950180111. [DOI] [PubMed] [Google Scholar]
- 35.Slansky J E, Farnham P J. Introduction to the E2F family: protein structure and gene regulation. Curr Top Microbiol Immunol. 1996;208:1–30. doi: 10.1007/978-3-642-79910-5_1. [DOI] [PubMed] [Google Scholar]
- 36.Slebos R J C, Lee M H, Plunkett B S, Kessis T D, Williams B O, Jacks T, Hedrick L, Kastan M B, Cho K R. p53-dependent G1 arrest involves pRb-related proteins and is disrupted by the human papillomavirus 16 E7 protein. Proc Natl Acad Sci USA. 1994;91:5320–5324. doi: 10.1073/pnas.91.12.5320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Smith E J, Leone G, DeGregori J, Jakoi L, Nevins J R. The accumulation of an E2F-p130 transcriptional repressor distinguishes a G0 cell state from a G1 cell state. Mol Cell Biol. 1996;16:6965–6976. doi: 10.1128/mcb.16.12.6965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Smith E J, Leone G, Nevins J R. Distinct mechanisms control the accumulation of the Rb-related p107 and p130 proteins during cell growth. Cell Growth Differ. 1998;9:297–303. [PubMed] [Google Scholar]
- 39.Thalmeier K, Synovzik H, Mertz R, Winnacker E L, Lipp M. Nuclear factor E2F mediates basic transcription and transactivation by E1a of the human MYC promoter. Genes Dev. 1989;3:527–536. doi: 10.1101/gad.3.4.527. [DOI] [PubMed] [Google Scholar]
- 40.Tommasi S, Pfeifer G P. In vivo structure of the human cdc2 promoter: release of a p130–E2F-4 complex from sequences immediately upstream of the transcription initiation site coincides with induction of cdc2 expression. Mol Cell Biol. 1995;15:6901–6913. doi: 10.1128/mcb.15.12.6901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Verona R, Moberg K, Estes S, Starz M, Vernon J P, Lees J A. E2F activity is regulated by cell cycle-dependent changes in subcellular localization. Mol Cell Biol. 1997;17:7268–7282. doi: 10.1128/mcb.17.12.7268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Voso M T, Burn T C, Wulf G, Lim B, Leone G, Tenen D G. Inhibition of hematopoiesis by competitive binding of transcription factor PU.1. Proc Natl Acad Sci USA. 1994;91:7932–7936. doi: 10.1073/pnas.91.17.7932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wade M, Kowalik T F, Mudryj M, Huang E-S, Azizkhan J C. E2F mediates dihydrofolate reductase promoter activation and multiprotein complex formation in human cytomegalovirus infection. Mol Cell Biol. 1992;12:4364–4374. doi: 10.1128/mcb.12.10.4364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wang Z M, Yang H, Livingston D M. Endogenous E2F-1 promotes timely G0 exit of resting mouse embryo fibroblasts. Proc Natl Acad Sci USA. 1998;95:15583–15586. doi: 10.1073/pnas.95.26.15583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Weinberg R A. The retinoblastoma protein and cell cycle control. Cell. 1995;81:323–330. doi: 10.1016/0092-8674(95)90385-2. [DOI] [PubMed] [Google Scholar]
- 46.Weintraub S J, Prater C A, Dean D C. Retinoblastoma protein switches the E2F site from positive to negative element. Nature. 1992;358:259–261. doi: 10.1038/358259a0. [DOI] [PubMed] [Google Scholar]
- 47.Weintraub S J, Chow K N B, Luo R X, Zhang S H, He S, Dean D C. Mechanism of active transcriptional repression by the retinoblastoma protein. Nature. 1995;375:812–815. doi: 10.1038/375812a0. [DOI] [PubMed] [Google Scholar]
- 48.Wu C L, Classon M, Dyson N, Harlow E. Expression of dominant-negative mutant DP-1 blocks cell cycle progression in G1. Mol Cell Biol. 1996;16:3698–3706. doi: 10.1128/mcb.16.7.3698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yamasaki L, Jacks T, Bronson R, Goillot E, Harlow E, Dyson N J. Tumor induction and tissue atrophy in mice lacking E2F-1. Cell. 1996;85:537–548. doi: 10.1016/s0092-8674(00)81254-4. [DOI] [PubMed] [Google Scholar]
- 50.Yan Z, DeGregori J, Shohet R, Leone G, Stillman B, Nevins J R, Williams R S. Cdc6 is regulated by E2F and is essential for DNA replication in mammalian cells. Proc Natl Acad Sci USA. 1998;95:3603–3608. doi: 10.1073/pnas.95.7.3603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zacksenhaus E, Gill R M, Phillips R A, Gallie B L. Molecular cloning and characterization of the mouse RB1 promoter. Oncogene. 1993;8:2343–2351. [PubMed] [Google Scholar]
- 52.Zhang H S, Postigo A A, Dean D C. Active transcriptional repression by the Rb-E2F complex mediates G1 arrest triggered by p16INK4a, TGFbeta, and contact inhibition. Cell. 1999;97:53–61. doi: 10.1016/s0092-8674(00)80714-x. [DOI] [PubMed] [Google Scholar]
- 53.Zwicker J, Liu N, Engeland K, Lucibello F C, Muller R. Cell cycle regulation of E2F site occupation in vivo. Science. 1996;271:1595–1597. doi: 10.1126/science.271.5255.1595. [DOI] [PubMed] [Google Scholar]