Abstract
Periodontitis is a multi-etiologic infection characterized clinically by pathologic loss of periodontal ligament and alveolar bone. Herpesviruses and specific bacterial species are major periodontal pathogens which cooperate synergistically in producing severe periodontitis. Cellular immunity against herpesviruses and humoral immunity against bacteria are key periodontal host defenses. Genetic, epigenetic and environmental factors are modifiers of periodontal disease severity. MicroRNAs (miRNAs) are a class of noncoding, gene expression-based, post-transcriptional regulatory RNAs of great importance for maintaining tissue homeostasis. Aberrant expression of miRNAs has been associated with several medical diseases. Periodontal bacteria and herpesviruses elaborate several miRNAs which are of current research interest. This review attempts to conceptualize the role of periodontal miRNAs in the pathogenesis of periodontitis. The diagnostic potential of salivary miRNAs is also addressed. Employment of miRNA technology in periodontics represents an interesting new preventive and therapeutic possibility.
1. Introduction
Periodontitis is a multi-etiologic disease involving bacteria and herpesviruses, protective and destructive host defenses, and disease modifying factors such as genetic, epigenetic and environmental determinants.1–4 Periodontitis affects approximately 50% and severe/progressive forms 11% of the global population.5 Periodontitis exerts broad pathogenic effects and is a major public health problem. Severe periodontitis can lead to tooth loss and increased risk for cardiovascular disease, stroke, cancer, Alzheimer’s disease, diabetes, rheumatoid arthritis, adverse pregnancy outcome and 50 other systemic diseases.6,7 Although periodontitis is not a fatal disease per se, dissemination (via the gingival and systemic circulation) of infectious agents present in periodontitis to various sites in the body may result in the development of life-threatening diseases. Periodontal herpesvirus virions seem to comprise the greatest risk for transmission and colonization of nonoral sites. The 8 human herpesviruses can infect epithelial cells, endothelial cells, hematopoietic cells, connective tissue cells, neural ganglia, parenchymal cells, and other cell types, and can cause disease in virtually every organ system in the human body, including the kidneys, liver, pancreas, heart, brain, lungs, stomach, adrenal glands, genitalia, eye, blood vessels, intestines, joints, skin, and salivary glands.8,9
The interplay between microbial virulence, genetics, epigenetic factors, and host defenses determines disease susceptibility and eventually disease outcome. Epigenetic modifications alter the physical structure of DNA and the biological response to environmental stimuli. The shift in DNA morphology causes a change in gene expression which, despite being not encoded in the DNA sequence, participates in chromatin remodeling (DNA methylation and histone modification) and gene activation or inactivation.10,11 The epigenetic noncoding RNAs (ncRNAs) are divided, based on the length of nucleotides (nts), into short ncRNAs (<30 nts; miRNAs), short interfering RNAs (siRNAs), piwi-interacting RNAs (piRNAs) and long RNAs (lncRNAs; >200 nts; mediating gene silencing, DNA methylation, heterochromatin formation and histone modification).12–15 The human genome encodes ~2300 true miRNAs and binds approximately 60% of the genes.16 Guided by 2–8 nucleotides of their sequence, miRNAs complementarily bind to the 3′ untranslated regions (3′UTRs) of the target messenger RNA (mRNA). The result may be mRNA cleavage or translational suppression. Messenger RNA degradation comprises one of the main final outcomes of miRNA targeted transcripts.12,13 Due to imperfect complementary binding by miRNAs, a single miRNA molecule can bind to >100 different mRNAs and become a key negative regulator of gene expression.
Noncoding RNA (ncRNA)-mediated transcriptional and post-transcriptional modifications have biological significance. miRNAs can interfere with the messenger RNA (mRNAi) regulation of post-transcriptional gene expression.17,18 The miRNA synchronization of gene expression is an important mechanism governing genetic regulations of multiple physiological and pathophysiologic processes. Changes in miRNA expression, identified by microarrays, RT-PCR and RNA sequencing, may occur in physiological and pathologic conditions, including cancer, inflammatory disease, autoimmune and endocrine disorders, and serve as a biomarker or molecular therapeutic target.19,20 Cancer onset, progression, angiogenesis, and metastasis have been related to under-expression of miRNAs, termed oncomiRNAs and metastamiRNAs, and to tumor suppressor miRNAs.20,21 miRNAs may play a distinct role also in epithelial stem cell differentiation with tooth development. For example, specific sets of miRNAs exist in incisor vs. molar teeth and in epithelium vs. mesenchymal tissue,22 pointing to roles in determining size and shape of teeth. Conditional deletion of the RNase III enzyme Dicer1 in the dental epithelium and the mesenchyme hampers differentiation of progenitor cells, leading to irregularity in miRNA expression and to malformed incisors and molars.23
Periodontal tissues are strongly trophic to various viruses, including herpesviruses, papillomaviruses, Torque Teno viruses, SARS-CoV-2 and HIV. An interesting feature of some human viruses is their ability to encode for miRNAs, despite lacking proteins involved in the generation of mature miRNAs. Several viral miRNAs have been identified in animal viruses as well. Members of the herpesvirus family, which contains large DNA viruses with 100–250 kilobases genomes, encode multiple functional miRNAs that possibly contribute to herpesvirus pathosis in periodontitis, periimplantitis and endodontic pathosis.9,24–26 However, little is known concerning herpesvirus-encoded miRNAs in periodontal disease and their biological mechanisms.
Periodontal cellular and herpesvirus ncRNAs conduct epigenetic regulation, which has been linked to periodontal inflammation and resolution, including myeloid cell differentiation and migration, bacterial phagocytosis, and Toll-like receptor signaling. That differentially expressed ncRNAs occur in untreated and treated periodontitis lesions suggest a role in tissue homeostasis, wound healing and host defense. Human cellular and viral encoded miRNAs may in fact reveal a new facet of the host-pathogen interactions in periodontal disease. This article focuses on cellular and viral miRNAs in periodontopathogenesis and on the mechanisms regulated by disease-associated miRNAs. The article also discusses miRNAs as a promising diagnostic and prognostic tool for periodontal disease management. Noncoding RNAs governing genetic regulation may help to monitor periodontal treatment, and miRNAs in saliva and gingival crevicular fluid may serve as valuable non-invasive resources for research.
2. miRNA biogenesis
The first step in miRNA biogenesis is the generation of primary miRNA transcripts in the nucleus (pri-miRNA) (Figure 1). miRNA gene is usually transcribed by RNA polymerase II with the characteristic 5′ cap and 3′ poly-A tail. Pri-miRNAs can vary in lengths ranging from hundreds to thousands of nucleotides.12–15, The pri-miRNA undergoes post-transcriptionally several maturation steps. RNase III endoribonuclease and Drosha recognize the localized secondary structures on the pri-miRNAs and generates the stem loop 70- to 100-nucleotide-long hairpin shaped RNA termed precursor miRNA (pre-miRNAs). These pre-miRNAs are then translocated by exportin-5 to the cytoplasm, where they are acted upon by the RNase III enzyme Dicer.12–15 Dicer-mediated cleavage of the pre-miRNA at their terminal loop yields a 19- to 25-nucleotide-long double-stranded RNA structure. After exclusion of one strand of the duplex, the other strand is incorporated into the miRNA-induced silencing complex (mi-RISC) as the mature miRNA.12–14 The hallmark of miRNA mediated gene dysregulation is a reduction in protein expression, with or without reduced mRNA expression.12–14 This aberration can lead to derangements in the immune system, autoimmune disorders, lymphocyte proliferation, and cancer.19,20,27 The differential expression of miRNAs in embryogenesis and differentiation of various cell types and tissues, point to the importance of miRNAs in various developmental processes. The mature miRNA-guided protein machinery associated with mRNAs may ultimately affect targets’ structural stability or interfere with translation.12–15
Figure 1: Model of host and viral-encoded miRNA biogenesis.

Human primary miRNAs (pri-miRNA) are generally transcribed RNA polymerase II (Pol II), while viral miRNAs are transcribed by RNA pol II and in some instances RNA pol III or tRNase Z. Subsequent to pri-miRNAs, both human and viral miRNA biogenesis is largely similar. Pri-miRNA are processed to precursor- miRNAs (pre-miRNA) in the nucleus by the “microprocessor complex” composed of Drosha and DGCR8. Pre-miRNA are exported to cytoplasm by Exportin 5 and Ran-GTPase complex. In the cytoplasm, pre-miRNAs are processed to duplex miRNA by RNA endonuclease Dicer and its co-factor TRBP. Dicer and TRBP relay the short dsRNA to an Argonaute homolog (Ago1-4 isoforms in mammals) and GW182 (TNRC6A). The mature miRNA guided protein machinery targets respective mRNAs that eventually affects target’s structural stability or interferes with translation.
DGCR8- DiGeorge Syndrome Critical Region Gene 8; TRBP: Transactivation response element RNA-Binding Protein; TNRC6A- Trinucleotide Repeat Containing Adaptor 6A.
Human herpesviruses encode the largest number of mature miRNA species of all known virus families. Herpesviruses depend on the host miRNA processing and effector machinery to generate functionally mature v-miRNAs. To date, no herpesvirus protein has been identified that participates in the v-miRs biogenesis. In the absence of herpesvirus proteins, an expression vector with cloned viral precursor miRNAs was needed to form mature Epstein-Barr virus miRNAs.28–30 However, herpesvirus proteins can potentially impaire host miRNAs function and biogenesis, thereby preventing the host miRNA-mediated v-miRs biogenesis.28,31,32
Similar to human miRNAs, herpesvirus-encoded miRNA biogenesis commences with the transcription of primary miRNA transcripts (pri-miRNAs) mediated predominantly by RNA polymerase II (Pol II). Pri-miRNAs contains one or several hairpin loop structures in which the mature miRNA sequence is embedded (Figure 1).28,32 Besides RNA pol II, some herpesviruses utilize RNA III pol, tRNase Z or other non-canonical pathways to generate mature v-miRs.33–35 For simplicity, this article will focus on the predominant canonical miRNA biogenesis pathway for v-miR biogenesis.
3.1. miRNAs and Periodontal Disease
Gingival epithelial and macrophages/Langerhans immune cells are the first responders to microbes and microbial by-products (lipopolysaccharide, lipoteichoic acid, DNA) via specific Toll-like receptors (Figure 2).36 miRNAs partly regulate Immune cell functions, and an altered miRNA expression can exert profound impact on immune reactions.37 Proinflammatory stimulators including lipopolysaccharide activate miRNAs, which may play a modulatory role in myeloid cells of the periodontal host response.38,39 miRNAs participate also in regulation of gene stressor patterns in various cellular and molecular pathways that are associated with inflammation and periodontal disease.40–42
Figure 2. Regulation of innate immune responses by cellular and viral miRNAs.
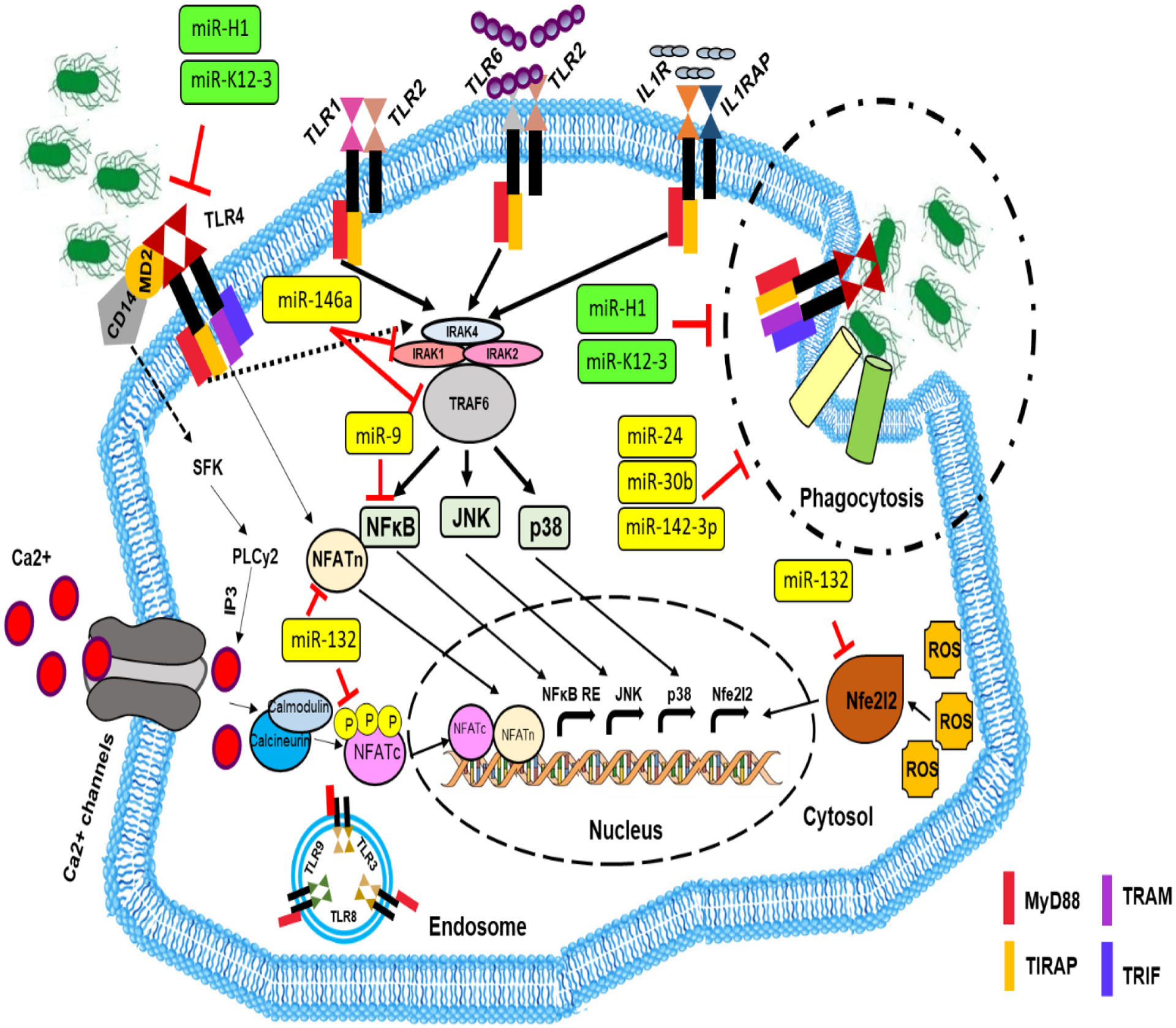
Tissue homeostasis at the gingival sulcus is disrupted due to the dysbiotic colonization of “keystone” periodontal pathogens elevating the pathogenicity of the whole community. Microbial products (PAMPs/DAMPs) induce innate immune signaling once bound to their ligand. TLRs (except TLR3) recruit the adaptor protein MyD88, thereby activating activate NFκB and MAP kinases by acting on their upstream activators IRAK, TRAF6 and TAK1. Activated NFκB and MAPK translocate to the nucleus and bind their respective motifs in the promoters of target genes to induce transcription. This includes early response and inflammatory genes like TNFα, IL1β, IL6, IL12, IL23, causing increased cytokine/chemokine production. Various cellular miRNAs including miR-9, miR-132 and miR-146a are shown to regulate innate immunity in response to periodontopathogenic challenge. Viral miRNAs also impair innate immunity by suppressing bacterial phagocytosis and cytokine secretion that may create favorable microenvironment for bacteria. HSV-1 miR-H1 and KSHV miR-K12-3 are two viral miRNAs that are enriched in inflamed gingiva and exhibit immunomodulatory functions.
TLR- Toll-like Receptor; PAMP- Pathogen Associated Molecular Pattern; DAMP- Damage Associated Molecular Pattern; MyD88- Myeloid Differentiation Primary Response Gene 88; NFκB- Nuclear Factor Kappa-light-chain-enhancer of activated B cells; MAPK- Mitogen-Activated Protein Kinase; IRAK- Interleukin 1 Receptor Associated Kinase; TRAF6- TNF Receptor Associated Factor 6; TAK1- Transforming growth factor-β-Activated Kinase 1; TNFα- Tumor Necrosis Factor α; IL- Interleukin; PLCγ- Phospholipase γ; JNK- c-Jun N-terminal kinases; HSV-1- Herpes Simplex Virus-1; KSHV- Kaposi Sarcoma-Associated Herpesvirus; TIRAP: TIR Domain Containing Adaptor Protein; TRAF- TNF receptor Associated Factor; TRIF- TIR-domain-containing adapter-inducing interferon-β; ROS- Reactive Oxygen Species; NFATn- Nuclear Factor of Activated T-Cells n; IP3- Inositol 1,4,5-tris Phosphate
Table 1 summarizes miRNAs which show differential expression in periodontal disease as compared to healthy gingiva.26,43–46 miRNA expression differs between gingival biopsies obtained from obese/non-obese subjects with or without periodontal disease. A wide-range upregulation of miR-15a, miR-18a, miR-22, miR-30d, miR-30e, miR-103, miR-106b, miR-130a, miR-142-3p, miR-185 and miR-210 has been reported periodontal disease.27,47 A significant upregulation occurs of periodontal miRNA species that share inflammatory and metabolic targets.44 Co-regulation between miRNAs and protein coding genes may take place in innate and adaptive immune responses and in tissue healing events. The pathophysiology of periodontitis may also involve miRNAs and regulatory frameworks governing immune cell differentiation and function.43 Dissecting the downstream pathways of miRNAs, however, is a daunting task but of crucial importance for identifying miRNAs that are part of the host defense, contribute to pre-inflammatory or pro-resolution cascades, are responsive to pattern recognition receptor signaling, and triggers of Toll-like receptor signaling, immune cell activation and macrophage polarization.43–47 Detailed miRNA profiles of periodontal health, disease and disease resolution may also provide a robust focus measure on the suitability of miRNAs to serve diagnostic purposes.
Table 1.
List of studies that reported differential expression of miRNAs in periodontal disease. Only a few differentially expressed miRNAs are cited as example.
| Differential expression of Selected miRNAs in periodontal disease | miRNA Detection Technique | Study |
|---|---|---|
| Upregulated miRNAs (11): miR-15a, miR-18a, miR-22, miR-30d, miR-30e, miR-103, miR-106b, miR-130a, miR-142-3p, miR-185, and miR-210 | miRNA PCR array | Perri et al., 2012 |
|
Upregulated miRNAs (91): hsa-miR-451, hsa-miR-223, hsa-miR-486-5p, hsa-miR-3917 Downregulated miRNAs (68): hsa-miR-1246, hsa-miR-1260, hsa-miR-141, hsa-miR-1260b, hsa-miR-203, hsa-miR-210, hsa-miR-205* |
miRNA microarray | Stoecklin-Wasmer et al., 2012 |
|
Upregulated miRNAs (17): hsa-miR-150, hsa-miR-223, and hsa-miR-200b Downregulated miRNAs (22): hsa-miR-379, hsa-miR-199a-5p, and hsa-miR-214. |
miRNA microarray | Ogata et al., 2014 |
|
Upregulated miRNAs (39): miR-21-5p, let-7f-5p, miR-29b/c, miR-24-3p, miR-27a-3p Downregulated miRNAs (2): miR-423-5p, miR-196b-5p |
miRNA PCR array | Naqvi et al., 2019 |
|
Upregulated miRNA (1): miR-143–3p Downregulated miRNA (1): miR-222-3p |
miRNA PCR array | Pettiette et al., 2019 |
|
Upregulated miRNAs: hsa-miR-29c Downregulated miRNAs: hsa-miR-27b-5p, hsa-miR-25-5p, hsa-miR-26a-5p, hsa-miR-26b-5p |
miRNA PCR array | Uttamani et al (unpublished) |
refers to passenger microRNA strand.
Porphyromonas gingivalis, a major periodontal pathogen, stimulates Toll-like receptors 2 and 4 signaling,48 and Toll-like receptor 2 activation also initiates neutrophil recognition of P. gingivalis.49 Toll-like receptors 2,4 and 7/8, which activate tumor necrosis factor-α and interleukin-1β via lipopolysaccharides, other bacterial cell wall components or in-vitro treatment, can upregulate miR-9 in human polymorphonuclear neutrophils and monocytes in MyD88-dependent and NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells)-dependent manners.50 miR-9 overexpression negatively regulates the expression of NF-κB1, a key member of the NF-κB family, which may subdue inflammation. Lee et al (2011) found ~2.5-fold higher expression of miR-9 in inflamed vs. healthy gingiva.51 Periodontal bacteria may induce moderate level of anti-inflammatory miRNAs to subdue exuberant host immune responses, possibly by suppressing NF-κB, tumor necrosis factor-α or interleukin-1β.52 Whether miR-9 overexpression triggers an anti-inflammatory loop in periodontal disease remains to be established.
The role of miRNAs in periodontal disease is complex. Jiang et al showed, in human periodontal ligament cells, that P. gingivalis lipopysaccharide reduced expression of miR-146a, Toll-like receptor 2 and 4, NF-κB, p65 nuclear activity and proinflammatory cytokines, but activated Toll-like receptor 2 and 4 and NF-κB-induced proinflammatory genes (Figure 2).53 He et al. reported that miR-146a, by virtue of targeting interleukin-1 receptor-associated kinase 1 (IRAK1) and tumor necrosis factor receptor-associated factor 6 (TRAF6) at their 3’UTRs, exert both pro- and anti-inflammatory attributes and suggested a positive regulation of innate and adaptive immunity via NF-κB signaling pathway.54 In addition, P. gingivalis challenge of THP-1 macrophages (an immortalized monocyte-like cell line) increased the expression level of miR-132 via Toll-like receptor 2 and 4 and via NF-κB signaling, while a decreased level of miR-132 strongly suppressed tumor necrosis factor-α secretion (Figure 2).55 Upregulation of miR-29b triggered osteoclast differentiation in tumor necrosis factor-α/RANKL-treated cells, and similar miR-29b-mediated mechanism of bone breakdown may contribute to periodontal bone loss.56 Franceschetti et al. demonstrated that upregulation of all three members of miR-29 family were key to a robust osteoclastogenesis, while miR-29 inhibition stymied bone breakdown.57
Phagocytosis is an attribute of monocytes, macrophages and dendritic cells.58,59 miR-24, miR-30b and miR-142-3p regulate negatively phagocytoses in macrophages and dendritic cells, as overexpression of these miRNAs significantly attenuates the phagocytosis of Escherichia coli and Staphylococcus aureus and suppresses proinflammatory cytokine secretion (tumor necrosis factor-α and interleukin-6, 8 and 12p40) from phagocytic cells (Figure 2).60–62 Interestingly, miR-24, miR-30b and miR-142-3p were down-regulated in gingivitis compared to healthy gingiva. Suppression of these pro-resolving miRNA networks might be a host response against bacterial-induced exuberant inflammation; however, dysregulated suppression of such miRNAs may generate overt inflammation and contribute to periodontal pathology.
To sum up, different miRNAs exhibit unique and varying expression patterns that may contribute to hyperinflammation or resolution of periodontal disease.
3.2. miRNA and periodontal inflammatory cells
Figure 3 illustrates unique subset of miRNAs that regulate immune cell-specific functions. It is notably that some miRNAs exhibit overlapping yet mutually exclusive functions in different immune cells. miRNAs also play an active role in regulation of the overall immune response in periodontitis development, and miRNA expression is markedly different in gingival tissue of periodontitis and control animals.41 miRNA expression in periodontal immune cells is the topic of this section.
Figure 3. microRNA regulates unique function and phenotype of various immune cell relevant to periodontal disease pathogenesis.

A diverse set of immune cells contribute to immunity against periodontopathogens. To elicit adept immune response in a dynamic manner, rapid changes in transcriptional profiles can occur. miRNAs can play important role in swift polarization of immune cells necessary to mount robust immune activation. A few miRNAs have been characterized that may be critical for the acquisition of various immune cell and osteoclasts relevant to periodontal disease.
Th- T helper; Treg- T regulatory
Neutrophils
Neutrophils are the predominant mammalian cell type in the periodontal pocket where they exert anti-bacterial function by phagocytosis and by enzymatic activity and reactive oxygen species.63–65 The tumor necrosis factor-α enriched environment of periodontal disease leads to induction of endothelial adhesion molecules (E-selectin and intercellular adhesion molecule-1) and a concomitant downregulation of their miR-31 and miR-17-3p targeting miRNAs.66 A downregulation of these miRNAs occcurs both in human and mouse periodontitis models.41 Downregulation of miR-17-3p also supports the expression of its target interleukin-8, which is essential for the recruitment of neutrophils. miR-451 is also downregulated in periodontal disease versus healthy controls.41 Conversely, a systemic administration of miR-451 negatively affected the infiltration of neutrophils in an air-pouch model of local inflammation.67 Evidently, miRNAs can support periodontal infiltration of neutrophils during disease development.
Macrophages
Monocytes and macrophages are key cellular components of innate and adaptive immunity and play vital functions in maintaining tissue homeostasis. miRNA profiles can significantly modulate and alter the attributes of macrophages. Macrophage exposure in vitro to lipopolysaccharide of Aggregatibacter actinomycetemcomitans and P. gingivalis cause a downregulation of miR-29b and let-7f. Validated targets of miR-29b are interleukin-6 receptor α and interferon-γ-inducible protein 30 and of flet-7f, a suppressor of cytokine signaling 4 and thrombospondin-1.39 Macrophage activation and function in gingivitis are also guided by differential regulation of miR-146 and miR-155 via NF-κB-dependent and Toll-like receptor pathways in macrophages.44,53,68–70 Interestingly, miR-146 acts as a negative feedback regulator in innate immunity by targeting the transcription factor RelB and miR-155 which exert dual effects on macrophage activation and function by upregulating both pro- and anti-inflammatory pathways in context dependent manners.71
An abundance of M1/M2 macrophages in periodontal disease is strongly associated with expression of miR-155 and miR-125b, with most upregulation occurring in M1 macrophages in agreement with a polarization of M1 macrophages in periodontitis. miR-155 promotes the M1 macrophage phenotype via suppression of the CCAAT/enhancer-binding protein-β signaling cascade and via degradation of SHIP-1, IL13Rα1 or SMAD2/3 during macrophage programming and activation.69,72–75 Similarly, overexpression of miR-125b enhances macrophage activation via targeting IRF4.75,76 miR-24b also serves as a negative regulator of macrophage classical activation and may promote an alternate pathway of macrophage activation. Downregulation of miR-24 will likely activate pro-inflammatory gene networks and may participate in host defense responses.26,60,77,78
Macrophages are promptly recruited to the sites of periodontal infection and encounter antigens from invading organisms.79 Several miRNAs are involved in leukocyte differentiation, innate signaling and macrophage activation.19,60,80,81 Key miRNA players such as miR-155, miR-146a, miR-127and miR-125b may modulate macrophage activation state.81–83 Overexpression of miR-155 drives macrophages to M1 phenotype, by regulating Akt-dependent M1/M2 MΦ polarization, and suppresses M2 phenotype by downregulating the expression of interleukin-13Rα1. Overexpression of miR-24, miR-30b, and miR-142-3p in lipopolysaccharide-stimulated monocytes, macrophages, and dendritic cells can also inhibit myeloid cell functionality including reduction of pro-inflammatory cytokine secretion.60 Over-expression of miR-146a can significantly decrease the production of tumor necrosis factor-α in M1 macrophages and increase the production of M2 marker genes such as Arg1 by targeting the Notch signaling pathway.84 Divergent miRNA expression occurring in periodontal disease can influence monocyte-to-macrophage differentiation and affect the macrophage functionality in disease pathogenesis (Figure 4).
Figure 4. microRNA regulate inflammatory and pro-resolving macrophage phenotype.

A unique subset of pro- (red) and anti-inflammatory (blue) miRNAs have been characterized in the context of periodontal disease or periodontopathogen challenge. Dysregulation in the expression of specific miRNAs could be an important strategy by perioddontopathogens to alter the functional phenotype of macrophages (Mφ), a critical microbicidal innate immune cell. For instance, in periodontopathogen challenged myeloid cells miR-24 favors M2 over M1 phenotype by inhibiting proinflammatory cytokine production and inducing M2-associated surface markers
A challenge of miR-24 with either lipopolysaccharide of P.gingivalis, A. actinomycetemcomitans or Escherichia coli resulted in downregulation of miR-24 expression, similar to the downregulation of miR-24 observed in diseased gingival biopsies after IgG stimulation.26,60,77,78 Also, miR-24 in lipopolysaccharide-challenged macrophages exhibited reduced secretion of the proinflammatory cytokines tumor necrosis factor-α, interleukin-12p40 and interleukin-6.60 Overexpression of miR-24 confers immunosuppressive phenotypes in macrophages by inducing CD206 in alternatively activated (M2 phenotype) macrophages but not in classically activated (M1 phenotype) macrophages.77 Together, these findings suggest an importaant role of miRNAs in shaping the immune response against periodontopathogens.
Dendritic cells
Dendritic cells are phagocytic antigen-presenting cells that are present in periodontal, mucosal, lymphoid and skin tissues and serve as a link between innate and adaptive immunity.85,86 In mouse and human periodontitis models, significant upregulation takes place in dendritic cell-related miRNAs, including miR-24, miR-30, miR-126, miR-142, miR-146, miR-155 and let-7i. These miRNAs are involved in maturation and functions of dendritic cells (and macrophages), suggesting another miRNAs participation in the pathogenesis of periodontal disease.41 Studies show that miR-155 increases the pathogen-binding ability of dendritic cells and promotes antigen-specific T cell activation via targeting downregulation of PU.1.87,88 Furthermore, both miR-155 and let-7i enhances interleukin-12 production and induction of proinflammatory cytokines via targeting SOCS-1 expression in mature dendritic cells.89–91
T cells
T cells predominate in cell-mediated (macrophage/lymphocyte) immunity and are imperative for polyclonal B cell activation. Mature T cells are categorized as CD4 T and CD8 T cells based on two major surface receptor molecules. The T cell receptor signal activation initiates several downstream pathways that result in cytokine production and T helper cell differentiation,92 and in significant changes in the miRNA profile.93 T cell activation of CD4 and CD8 causes upregulation of the miR-181 family, miR-17–92 clusters, miR-214, miR-146a, miR-155 and let-7 and downregulation of miR-29, miR-125 and miR-216.94 Upregulation of the miR-181 family miRNAs negatively targets multiple phosphatases, including SH2 domain-containing protein tyrosine phosphatase 2, protein tyrosine phosphatase, non-receptor type 22 and dual-specificity protein phosphatase 5 and 6, and negatively regulate the activation of T cells.95–97 The miR-181a expression results in enhanced phosphorylation of immunoreceptor tyrosine-based activation motifs on the cytosolic side of the T-cell receptor/CD3 complex.98,99 Upregulation of the miR-214 and miR-17–92 cluster during T cell activation targets the tensin homologue and the negative phosphatase regulator in the PI3K/Ak strain transforming pathway.100,101 miR-146 functions as a feedback regulator of NF-κB signalling and modulates T cell division, division and growth and productive immune response.102 miR-146 also targets tumor necrosis factor receptor associated factor 6 and interleukin 1 receptor associated kinase 1 in activated T cells for feedback control of the intensity and duration of NF-κB signaling.98,102
Th1-type phenotype is known as proinflammatory and regulated by miR-29 and the miR-17–92 cluster. miR-29 inhibits Th1 cell differentiation and interferon-γ production in CD4 T cells by targeting T-box transcription factor Tbx21 and eomesodermin (Eomes; Th1 specific transcription factors) and interferon-γ mRNA expression.103 miR-17–92 cluster negatively effects Th1 differentiation and interferon-γ production via targeting transforming growth factor-β receptor type 2 and cAMP-responsive element-binding protein-1. These factors are induced in cluster targets and are involved in regulatory T cell differentiation.104 Th2 cells are associated with humoral immunity and show anti-inflammatory properties and are regulated by miR-155, let-7 family members and miR-126.105,106 MiR-155 inhibits Th2 differentiation107 and the let-7 family negatively regulates Th2 differentiation by targeting interleukin-13.108 Knockdown of miR-126 after T cell activation exhibits increased interleukin 5 and 13 expression and favors a Th2 differentiation.109
Dysregulation between T regulatory cells and Th17 cells may contribute to periodontitis.110 T regulatory cells possess a differential miRNA expression pattern characterized by high levels of miR-155 and miR-146 and low levels of miR-24, miR-31 and miR-125.111,112 miR-155 directly targets SOCS-1 (an inhibiter of STAT5 signaling) during T regulatory cell differentiation and is essential for maintaining stability and function of the T regulatory cell lineage.113 Both miR-24 and miR-31 target forkhead box P3, a lineage specification factor of T regulatory cells, and a downregulation of these miRNAs results in increased forkhead box P3 expression and enhanced T regulatory cell differentiation and function.114,115
Th17 cells recruit neutrophils and macrophages to control periodontal disease.116–118 miR-17–92 cluster and miR-19b favor Th17 cell differentiation,119,120 and miR-17 promotes Th17 polarization by inhibiting its target, the ikaros family zinc-finger 4.119 miR-17 and miR-19b suppress inducible regulatory T cell differentiation.104 Another miRNA that favors Th17 cell function is miR-155, which post-transcriptionally suppresses the inhibitory effects of Jumonji and AT-rich interaction domain-containing 2.121 On the other hand, miR-301 suppresses the expression of the E3 SUMO-protein ligase PIAS3, a negative regulator of STAT3 pathway and Th17 differentiation.122
miRNA expression studies have revealed an upregulated expression of miR-17, miR-19, miR-155 and miR-301 in inflamed gingiva of periodontitis patients and animals compared to healthy controls.41 These miRNAs promote the development of Th17 cells, which help initiate and maintain the inflammatory response in periodontal disease. Inhibition of these miRNAs may induce T regulatory cell formation and shift the immune response to a Th2 profile to attenuate, if warranted,, the severity of periodontal disease.
B cells
B cells comprised a critical proportion of the inflammatory infiltrate in diseased periodontal tissues and occur at elevated levels in advanced stages of periodontitis.123,124 Inflammation-related miRNAs (miR-125, miR-148, miR-155, miR-181 and miR-217) are upregulated in severe periodontitis where they regulate B cell terminal differentiation by targeting a network of transcription factors. MiR-125b targets transcription factors (B lymphocyte-induced maturation protein-1 and interferon regulatory factor 4) and perturb B cell terminal differentiation.125 miR-148 targets BTB domain and CNC homologue 2 expression and promotes plasma cell lineage commitment plus fosters germinal centre B cell survival via targeting pro-apoptotic factors PTEN and Bim.126,127 miR-217 stabilizes the expression of the Bcl-6 transcriptional repressor in germinal centre and facilitates generation of class-switched antibodies.128 Both miR-155 and miR-181 negatively regulate the expression of activation-induced cytidine deaminase (AID) to reinforce and repress AID expression.129,130
3.3. Herpesvirus miRNAs and periodontal disease
Herpesvirus miRNA expression profile can provide valuable information regarding viral presence, state of latency/reactivation and pathogenic role in periodontal disease. Herpesviruses encode various proteins to subvert host cellular pathways and the discovery of virus-encoded miRNAs highlights yet another multifaceted molecule by which herpesviruses can evade the host immune response. Since the first report of herpesvirus-derived miRNAs (v-miRs) in 2004,33 a large number (~250) of v-miRs sequences have been deposited in the miRBase (http://www.mirbase.org/cgi-bin/browse.pl).
Herpesviruses express miRNAs during the latent and lytic phases.32,131,132 Numerous reports demonstrate a direct regulation of host transcriptome by v-miRs.26,29,32,131,133 All herpesviruses except varicella-zoster virus and herpesvirus-6A encode multiple miRNAs. Mature miRNAs are reported in Epstein-Barr virus (n=44), human cytomegalovirus (n=26), herpesvirus-6B (n=8), herpes simplex virus-1 (n=27), and Kaposi sarcoma-associated herpesvirus (n=25).32,131
Figure 5 shows the genomic organization of v-miRs of the 5 most prominent herpesviruses in oral diseases. Because herpesviruses lack proteins and rely on the host protein machinery to generate viral miRNA, they are under positive selection pressure to maintain the sequence of miRNAs precursors. Stem-loop structure recognized by specific host enzymes ensures generation of mature miRNAs.12,13 Mutations in miRNA precursor can impair sequence complementarity and alter the molecular secondary structure required for interaction with RNA-binding endonucleases.
Figure 5. Genome organization of viral microRNAs in 5 herpesviruses associated with oral diseases.

EBV- Epstein-Barr Virus; HHV6B- Human Herpesvirus 6B; HSV-1- Herpes Simplex Virus-1; KSHV- Kaposi Sarcoma-Associated Herpesvirus; HCMV- Human Cytomegalovirus
Specific herpesvirus miRNAs accumulate with higher expression level and prevalence in periodontal disease than in periodontal health. Herpes simplex virus-1 miR-H1–5p and ICP0 transcript were present in inflamed gingiva but not detectable in most healthy gingival biopsies (Figure 6A, D).25 Cytomegalovirus miR-US4 and Kaposi sarcoma virus-miR-K12-3 also showed higher expression levels and prevalence and in inflamed than in healthy gingiva. High level of v-miR expression is suggestive of an active herpesvirus state (Figure 6B,C). Higher v-miRs expression was also detected in obese human subjects with periodontal disease than with healthy periodontium.26
Figure 6. Herpesvirus miRNA and mRNA are enriched in inflamed gingival biopsies.

Total RNA isolated from periodontally healthy and disease gingiva was used for viral miRNA and mRNA quantification by reverse transcription PCR (RT-qPCR). Dot plots show mean Ct values of (A) hsv1-miR-H1, (B) hcmv-miR-US4 and (C) kshv-miR-K12-3, and (D) ICP0 mRNA of HSV-1 in gingiva of healthy and periodontitis subjects (n=6/group). Ct value ≥35 was considered as not detected. Lower Ct values indicate higher expression. Numbers of positive samples are mentioned for each group. Student’s t-test was used to calculate p-values. **p<0.01; ***p<0.001.
HSV-1- Herpes Simplex Virus 1; HCMV- Human Cytomegalovirus; KSHV- Kaposi Sarcoma-Associated Herpesvirus; ICP0- Infected Cell Protein 0
v-miR expression may impair host innate and cellular functions. Evidence relates an overexpression of herpes simplex virus-1 (miR-H1), Kaposi sarcoma virus (miR-K12-3) and cytomegalovirus (miR-UL70) miRNAs to marked changes in the host transcriptome miRnome in gingival epithelial cells and macrophages.26,133 High v-miR level is also related to functional modulation of cytokine/chemokine secretion, host transcriptome (including mRNA and miRNA) changes, and impaired bacterial phagocytosis.26,133 Attenuation of bacterial phagocytosis is of particular interest because ex vivo data correlate v-miRs accumulation with increased bacterial growth in gingival biopsies.134,135 A transcriptome-wide impact of vmiRs on various host cells suggests a pathogenic role by perturbing cellular pathways. High gingival expression of v-miRs may contribute to periodontal disease by impairing host immune responses and cellular functions.
Epstein-Barr virus and Kaposi sarcoma virus are oncogenic herpesviruses involved in oro-nasal carcinomas and other cancers. Herpesvirus miRs may contribute to carcinogenesis, metastasis, and resistance to treatment, and v-miRs profiles may be helpful in judging cancer diagnosis, disease activity, prognosis and response to treatment. Herpesvirus spread and persistence may be facilitated by suppression of immune recognition by v-miRs.
Various vmiRs are potential candidates for carcinoma assessment tests. Zhang et al. (2015) found plasma levels of miR-BART7 and miR-BART13 decreases after radiotherapy, suggesting the usefulness of vmiR expression to monitor therapeutic response.136 Plasma levels of miR-BART7 can separate patients with nasopharyngeal carcinoma from healthy subjects, suggesting circulating vmiRs are potential diagnostic markers.137 High levels of miR-BART7 confer resistance to nasopharyngeal carcinoma likely due to dysregulation of cancer-related cellular gene networks. These findings point to a role of vmiRs in cancer development and diagnostics. However, the reason specific sets of v-miRs are enriched in disease and participate in disease pathogenesis are largely unknown. Only limited data exist regarding herpesvirus transcript profiles in diseased and healthy oral tissues. Herpes simplex virus-1 ICP0 mRNA occurs with high expression level in inflamed gingival biopsies (Figure 6D).
4. Saliva for oral herpesvirus identification
Detection of miRNAs in saliva, serum and plasma supports their clinical application as potential biomarkers. Recent studies found that salivary miRNA expression can distinguish periodontitis and healthy controls.138–140 Saliva testing represents an interesting alternative to nasopharyngeal swabs and serum sampling due to lower costs, increased patient comfort, and reduced transmission risk to healthcare personnel from swab-induced coughing and sneezing and from blood-borne pathogens. Furthermore, patients may collect their own salivary samples at home, causing a reduction in demand for healthcare personnel and testing supplies. Also, compared to conventional gene testing, miRNAs exhibit exclusive biogenesis, can modulate entire gene programs, and may minimize off-target test results. A single miRNA molecule that target multiple genes can potentially influence simultaneously the expression of several proteins.
Initial research points to the feasibility of employing a saliva-based test for rapid detection of herpesvirus mRNA and miRNA. Herpes simplex virus-1 mRNA and miRNA have shown even higher expression and detection frequency in saliva than in gingival crevicular fluid in subjects with periodontal disease (Figure 7); similar difference was not observed for the RNU6B endogenous gene which served as control. These initial results support the notion of detecting salivary herpesvirus vmiRs to assess periodontal disease severity.
Figure 7. Salivary detection of viral transcript for periodontal health monitoring.
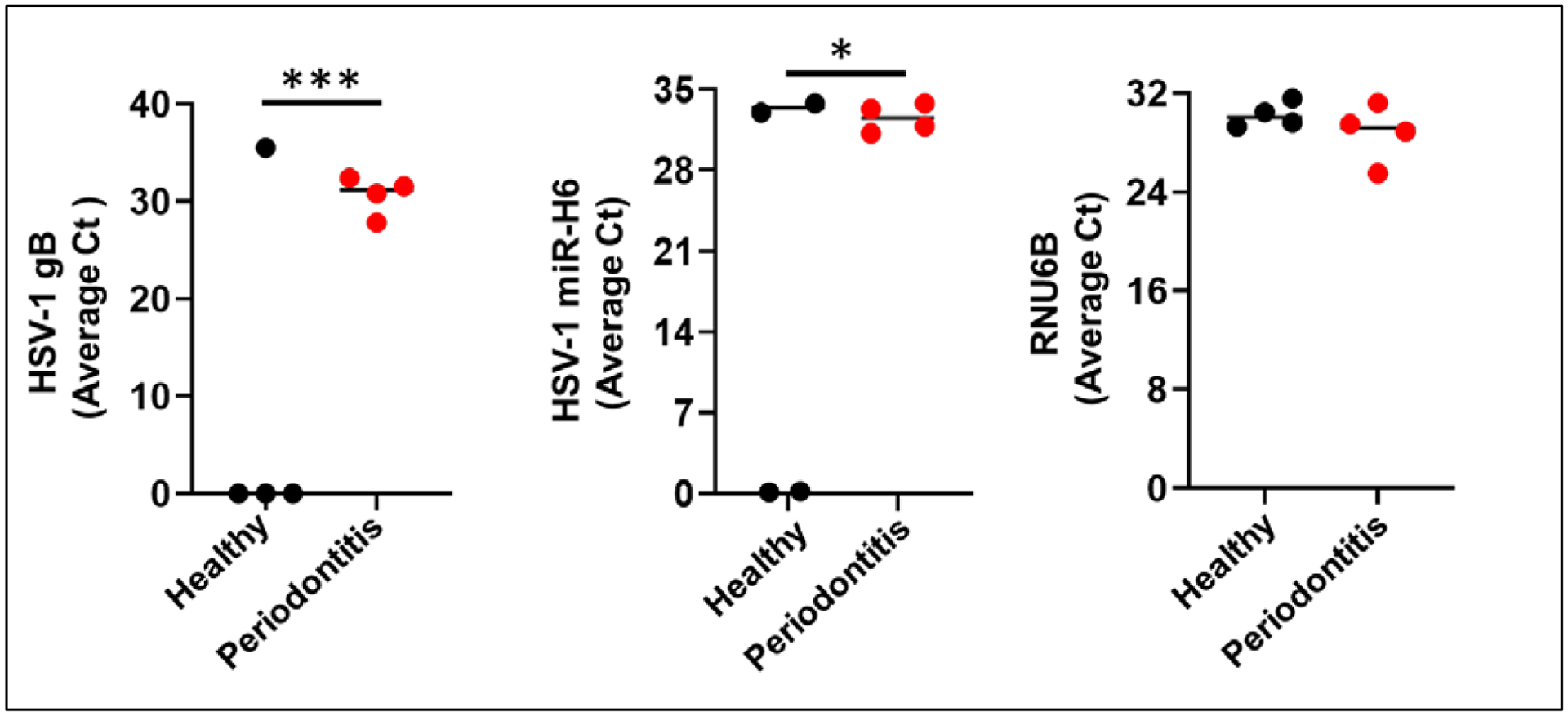
Total RNA was isolated from saliva of periodontally healthy and diseased subjects (n=4/group) and the expression of HSV-1 mRNA (gB) and miRNA (miR-H6) was examined by quantitative reverse transcription PCR (RT-qPCR). As a control, RNA, U6B Small Nuclear (RNU6B) was used. Student’s t-test was used to calculate p-values. *p<0.05; ***p<0.001.
5. Future Directions
The long-term research goal is to translate the miRNA diagnostic and therapeutic possibilities into patient management, but several challenges remain. High throughput sequencing at single-cell resolution will be critical to identify the periodontal cellular sites of miRNA production. Studies on the role of periodontal miRNA in cytokines/chemokine production may be needed to distinguish protective and adverse immune reactions.
Herpesviruses are at the center of the host-pathogen interaction in severe periodontal disease, and miRNAs perform potentially dual functions by targeting multiple herpesvirus and host transcripts that regulate herpesvirus replication and persistence. Certain miRNAs are associated with either the herpesvirus lytic or latent stage which therefore make them candidates to gauge the herpesvirus lifecycle status. As lytic herpesviruses are triggers of progressive periodontitis, identification of vmiRs that support productive herpesvirus infections may help identify mechanisms of periodontal disease progression and assist in periodontal diagnosis, treatment planning and treatment outcome assessment. Synthetically derived siRNAs or anti-miRNA oligonucleotides may also be used therapeutically to target mRNA for translational repression.27,141,142 Future dental practice will most likely employ saliva-based biomarkers, including miRNAs, to diagnose oral diseases and guide more precision-based treatment. However, the application of miRNAs in dental practice will depend on design of novel assays and technologies that can be accessed by dental practitioners. Further research into the biology of periodontal miRNAs is warranted and encouraged.
Acknowledgements:
Research in ARN lab is funded by the NIH/NIDCR grants R01DE027980 and R03DE027147. We are thankful to Dr. Juhi Uttamani and Dr. Raza Ali Naqvi for their valuable inputs in manuscript drafting and Dr. Araceli Valverde for help with the preparation of illustrations.
References
- 1.Kinane DF, Stathopoulou PG, Papapanou PN. Periodontal diseases. Nat Rev Dis Primers. 2017;3:17038. [DOI] [PubMed] [Google Scholar]
- 2.Loos BG, Van Dyke TE. The role of inflammation and genetics in periodontal disease. Periodontol 2000. 2020;83(1):26–39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Slots J Primer on etiology and treatment of progressive/severe periodontitis: A systemic health perspective. Periodontol 2000. 2020;83(1):272–276. [DOI] [PubMed] [Google Scholar]
- 4.Kulkarni V, Bhatavadekar NB, Uttamani JR. The effect of nutrition on periodontal disease: a systematic review. J Calif Dent Assoc. 2014;42(5):302–311. [PubMed] [Google Scholar]
- 5.Herrera D, Meyle J, Renvert S, Jin L. (2018) White Paper on Prevention and Management of Periodontal Diseases for Oral Health and General Health. Geneva, FDI World Dental Federation, 2018. [Google Scholar]
- 6.Genco RJ, Sanz M. Clinical and public health implications of periodontal and systemic diseases: an overview. Periodontol 2000. 2020;83(1):7–13. [DOI] [PubMed] [Google Scholar]
- 7.Bui FQ, Almeida-da-Silva CLC, Huynh B, Trinh A, Liu J, Woodward J, Asadi H, Ojcius DM. Association between periodontal pathogens and systemic disease. Biomed J. 2019;42(1):27–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Slots J Focal infection of periodontal origin. Periodontol 2000. 2019;79(1): 233–235. [DOI] [PubMed] [Google Scholar]
- 9.Slots J Periodontal herpesviruses: prevalence, pathogenicity, systemic risk. Periodontol 2000. 2015;69(1):28–45. [DOI] [PubMed] [Google Scholar]
- 10.Reik W Stability and flexibility of epigenetic gene regulation in mammalian development. Nature. 2007;447:425–432. [DOI] [PubMed] [Google Scholar]
- 11.Wilson AG. Epigenetic regulation of gene expression in the inflammatory response and relevance to common diseases. J Periodontol. 2008;79(8 Suppl):1514–1519. [DOI] [PubMed] [Google Scholar]
- 12.Bartel DP. Metazoan MicroRNAs. Cell. 2018;173(1):20–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ha M, Kim VN. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol. 2014;15(8):509–524. [DOI] [PubMed] [Google Scholar]
- 14.Cech TR, Steitz JA. The noncoding RNA revolution-trashing old rules to forge new ones. Cell. 2014;157(1):77–94. [DOI] [PubMed] [Google Scholar]
- 15.Naqvi AR, Islam MN, Choudhury NR, Haq QM. The fascinating world of RNA interference. Int J Biol Sci. 2009;5(2):97–117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Alles J, Fehlmann T, Fischer U, et al. An estimate of the total number of true human miRNAs. Nucleic Acids Res. 2019;47(7):3353–3364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Guil S, Esteller M. DNA methylomes, histone codes and miRNAs: tying it all together. Int J Biochem Cell Biol. 2009;41(1):87–95. [DOI] [PubMed] [Google Scholar]
- 18.O’Connell RM, Rao DS, Chaudhuri AA. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol. 2010;10:111–122. [DOI] [PubMed] [Google Scholar]
- 19.Mendell JT, Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012;148(6):1172–1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014;9:287–314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Scapoli L, Palmieri A, Lo Muzio L, et al. MicroRNA expression profiling of oral carcinoma identifies new markers of tumor progression. Int J Immunopathol Pharmacol. 23(4):1229–1234. [DOI] [PubMed] [Google Scholar]
- 22.Klein OD, Lyons DB, Balooch G, et al. An FGF signaling loop sustains the generation of differentiated progeny from stem cells in mouse incisors. Development. 2008;135(2):377–385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cao H, Wang J, Li X, et al. MicroRNAs play a critical role in tooth development. J Dent Res. 2010;89(8):779–784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhong S, Naqvi A, Bair E, Nares S, Khan AA. Viral microRNAs identified in human dental pulp. J Endod. 2017;43(1):84–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Naqvi AR, Seal A, Shango J, et al. Herpesvirus-encoded microRNAs detected in human gingiva alter host cell transcriptome and regulate viral infection. Biochim Biophys Acta Gene Regul Mech 2018;1861(5):497–508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Naqvi AR, Brambila MF, Martínez G, Chapa G, Nares S. Dysregulation of human miRNAs and increased prevalence of HHV miRNAs in obese periodontitis subjects. J Clin Periodontol. 2019;46(1):51–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kulkarni V, Uttamani JR, Naqvi AR, Nares S. microRNAs: Emerging players in oral cancers and inflammatory disorders. Tumour Biol. 2017;39(5):1010428317698379. [DOI] [PubMed] [Google Scholar]
- 28.Grundhoff A, Sullivan CS. Virus-encoded microRNAs. Virology. 2011;411(2):325–343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Skalsky RL, Corcoran DL, Gottwein E, et al. The viral and cellular microRNA targetome in lymphoblastoid cell lines. PLoS Pathog. 2012;8(1):e1002484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Umbach JL, Cullen BR. In-depth analysis of Kaposi’s sarcoma-associated herpesvirus microRNA expression provides insights into the mammalian microRNA-processing machinery. J Virol. 2010;84(2):695–703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Happel C, Ramalingam D, Ziegelbauer JM. Virus-Mediated Alterations in miRNA Factors and Degradation of Viral miRNAs by MCPIP1. PLoS Biol. 2016;14(11):e2000998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cullen BR. How do viruses avoid inhibition by endogenous cellular microRNAs? PLoS Pathogens. 2013;9(11):e1003694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Pfeffer, Zavolan FA, Grasser M, et al. Identification of virus-encoded microRNAs, Science. 2004;304(5671):734–736. [DOI] [PubMed] [Google Scholar]
- 34.Diebel KW, Smith AL, van Dyk LF. Mature and functional viral miRNAs transcribed from novel RNA polymerase III promoters. RNA. 2010;16(1):170–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bogerd HP, Karnowski HW, Cai X, Shin J, Pohlers M, Cullen BR. A mammalian herpesvirus uses noncanonical expression and processing mechanisms to generate viral MicroRNAs. Mol Cell. 2010;37(1):135–142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Huang CB, Alimova Y, Ebersole JL. Macrophage polarization in response to oral commensals and pathogens. Pathog Dis. 2016;74(3):ftw011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chen S, Constantinides C, Kebschull M, Papapanou P. MicroRNAs Regulate Cytokine Responses in Gingival Epithelial Cells. Infect Immun. 2016;84(12):3282–3289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Naqvi AR, Zhong S, Dang H, Fordham JB, Nares S, Khan A. Expression profiling of LPS responsive miRNA in primary human macrophages. J Microb Biochem Technol. 2016;8(2):136–143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Naqvi AR, Fordham JB, Khan A, Nares S. microRNAs responsive to A. actinomycetemcomitans and P. gingivalis LPS modulate expression of genes regulating innate immunity in human macrophages. Inn Immun. 2014;20(5):540–551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kebschull M, Papapanou PN. Mini but mighty: microRNAs in the pathobiology of periodontal disease. Periodontol 2000. 2015;69(1):201–220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Luan X, Zhou X, Naqvi A, et al. MicroRNAs and immunity in periodontal health and disease. Int J Oral Sci. 2018;10(3):24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Naqvi RA, Gupta M, George A, Naqvi AR. MicroRNAs in shaping the resolution phase of inflammation. Semin Cell Dev Biol. 2021:S1084–9521(21)00066–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Stoecklin-Wasmer C, Guarnieri P, Celenti R, Demmer RT, Kebschull M, Papapanou PN. MicroRNAs and their target genes in gingival tissues. J Dent Res. 2012;91(10):934–940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Perri R, Nares S, Zhang S, Barros SP, Offenbacher S. MicroRNA modulation in obesity and periodontitis. J Dent Res. 2012;91(1):33–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ogata Y, Matsui S, Kato A, et al. MicroRNA expression in inflamed and noninflamed gingival tissues from Japanese patients. J Oral Sci. 2014;56(4):253–260. [DOI] [PubMed] [Google Scholar]
- 46.Pettiette MT, Zhang S, Moretti AJ, Kim SJ, Naqvi AR, Nares S. MicroRNA Expression Profiles in External Cervical Resorption. J Endod. 2019;45(9):1106–1113.e2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Naqvi AR, Shango J, Seal A, Shukla D, Nares S. Herpesvirus and MicroRNAs: New Pathogenesis Factors in Oral Infection and Disease? Front Immunol. 2018;9:2099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lu W, Gu JY, Zhang YY, Gong DJ, Zhu YM, Sun Y. Tolerance induced by Porphyromonas gingivalis may occur independently of TLR2 and TLR4. PLoS One. 2018;13(7):e0200946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Sochalska M, Potempa J. Manipulation of Neutrophils by Porphyromonas gingivalis in the Development of Periodontitis. Front Cell Infect Microbiol. 2017;7:197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bazzoni F, Rossato M, Fabbri M, et al. Induction and regulatory function of miR-9 in human monocytes and neutrophils exposed to proinflammatory signals. Proc Natl Acad Sci USA. 2009;106(13):5282–5287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lee YH, Na HS, Jeong SY, Jeong SH, Park HR, Chung J. Comparison of inflammatory microRNA expression in healthy and periodontitis tissues. Biocell. 2011;35(2):43–49. [PubMed] [Google Scholar]
- 52.Liu W, Wang X, Zheng Y, et al. Electroacupuncture inhibits inflammatory injury by targeting the miR-9-mediated NF-κB signaling pathway following ischemic stroke. Mol Med Rep. 2016:13:1618–1626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jiang SY, Xue D, Xie YF, Zhu DW, Dong YY, Wei CC, Deng JY. The negative feedback regulation of microRNA-146a in human periodontal ligament cells after Porphyromonas gingivalis lipopolysaccharide stimulation. Inflamm Res. 2015;64(6):441–451. [DOI] [PubMed] [Google Scholar]
- 54.He L, Wang Z, Zhou R, Xiong W, Yang Y, Song N, Qian J. Dexmedetomidine exerts cardioprotective effect through miR-146a-3p targeting IRAK1 and TRAF6 via inhibition of the NF-κB pathway. Biomed Pharmacother. 2021;133:110993. [DOI] [PubMed] [Google Scholar]
- 55.Park MH, Park E, Kim H-J, et al. Porphyromonas gingivalis-induced miR-132 regulates TNFα expression in THP-1 derived macrophages. Springer Plus. 2016;5:761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kagiya T, Nakamura S. Expression profiling of miRNAs in RAW264.7 cells treated with a combination of tumor necrosis factor alpha and RANKL during osteoclast differentiation. J Periodontal Res. 2013;48(3):373–85. [DOI] [PubMed] [Google Scholar]
- 57.Franceschetti T, Kessler CB, Lee SK, Delany AM. miR-29 promotes murine osteoclastogenesis by regulating osteoclast commitment and migration. J Biol Chem. 2013;288(46):33347–33360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kubach J, Becker C, Schmitt E, Steinbrink K, Huter E, Tuettenberg A, Jonuleit H. Dendritic cells: sentinels of immunity and tolerance. Int J Hematol. 2005;81(3):197–203. [DOI] [PubMed] [Google Scholar]
- 59.Mosser DM, Edwards JP. Exploring the full spectrum of macrophage activation. Nat Rev Immunol. 2008;8(12):958–969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Naqvi AR, Fordham JB, Nares S. miR-24, miR-30b, and miR-142-3p regulate phagocytosis in myeloid inflammatory cells. J Immunol. 2015;194(4):1916–1927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Fordham JB, Naqvi AR, Nares S. Regulation of miR-24, miR-30b, and miR-142-3p during macrophage and dendritic cell differentiation potentiates innate immunity. J Leukoc Biol. 2015;98(2):195–207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Naqvi AR, Fordham JB, Ganesh B, Nares S. miR-24, miR-30b and miR-142-3p interfere with antigen processing and presentation by primary macrophages and dendritic cells. Sci Rep. 2016;6:32925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Andersen E, Cimasoni G. A rapid and simple method for counting crevicular polymorphonuclear leucocytes. J Clin Periodontol. 1993;20(9):651–655. [DOI] [PubMed] [Google Scholar]
- 64.Bender JS, Thang H, Glogauer M. Novel rinse assay for the quantification of oral neutrophils and the monitoring of chronic periodontal disease. J Periodontal Res. 2006;41(3):214–220. [DOI] [PubMed] [Google Scholar]
- 65.Liu RK, Cao CF, Meng HX, Gao Y. Polymorphonuclear neutrophils and their mediators in gingival tissues from generalized aggressive periodontitis. J Periodontol. 2001;72(11):1545–1553. [DOI] [PubMed] [Google Scholar]
- 66.Suárez Y, Wang C, Manes TD, Pober JS. Cutting edge: TNF-induced microRNAs regulate TNF-induced expression of E-selectin and intercellular adhesion molecule-1 on human endothelial cells: feedback control of inflammation. J Immunol. 2010;184(1):21–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Murata K, Yoshitomi H, Furu M, Ishikawa M, Shibuya H, Ito H, Matsuda S. MicroRNA-451 down-regulates neutrophil chemotaxis via p38 MAPK. Arthritis Rheumatol 2014;66(3):549–559. [DOI] [PubMed] [Google Scholar]
- 68.Xie YF, Shu R, Jiang SY, Liu Dl, Zhang Xl. Comparison of microRNA profiles of human periodontal diseased and healthy gingival tissues. Int J Oral Sci. 2011;3:125–134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.O’Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci USA. 2007;104(5):1604–1609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Taganov KD, Boldin MP, Chang KJ, Baltimore D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci USA. 2006;103(33):12481–12486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Etzrodt M, Cortez-Retamozo V, Newton A, et al. Regulation of monocyte functional heterogeneity by miR-146a and Relb. Cell Rep. 2012;1(4):317–324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Cai X, Yin Y, Li N, Zhu D, Zhang J, Zhang CY, Zen K. Re-polarization of tumor-associated macrophages to pro-inflammatory M1 macrophages by microRNA-155. J Mol Cell Biol. 2012;4(5):341–343. [DOI] [PubMed] [Google Scholar]
- 73.Martinez-Nunez RT, Louafi F, Sanchez-Elsner T. The interleukin 13 (IL-13) pathway in human macrophages is modulated by microRNA-155 via direct targeting of interleukin 13 receptor alpha1 (IL13Ralpha1). J Biol Chem. 2011;286(3):1786–1794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Louafi F, Martinez-Nunez RT, Sanchez-Elsner T. MicroRNA-155 targets SMAD2 and modulates the response of macrophages to transforming growth factor-β. J Biol Chem. 2010;285(53):41328–41336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Sierra-Filardi E, Puig-Kröger A, Blanco FJ, et al. Activin A skews macrophage polarization by promoting a proinflammatory phenotype and inhibiting the acquisition of anti-inflammatory macrophage markers. Blood. 2011;117(9):5092–5101. [DOI] [PubMed] [Google Scholar]
- 76.Squadrito ML, Etzrodt M, De Palma M, Pittet MJ. MicroRNA-mediated control of macrophages and its implications for cancer. Trends Immunol. 2013;34(7):350–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Fordham JB, Naqvi AR, Nares S. miR-24 regulates macrophage polarization and plasticity. J. Clin. Cell. Immunol 2015;6(5):362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Naqvi AR, Fordham JB, Nares S. MicroRNA target Fc receptors to regulate Ab-dependent Ag uptake in primary macrophages and dendritic cells. Inn Immun. 2016;22(7):510–521. [DOI] [PubMed] [Google Scholar]
- 79.Harokopakis E, Albzreh MH, Martin MH, Hajishengallis G. TLR2 transmodulates monocyte adhesion and transmigration via Rac1- and PI3K-mediated inside-out signaling in response to Porphyromonas gingivalis fimbriae. J Immunol. 2006; 176(12): 7645–7656. [DOI] [PubMed] [Google Scholar]
- 80.Kulkarni V, Naqvi AR, Uttamani JR, Nares S. MiRNA-target interaction reveals cell- specific post-transcriptional regulation in mammalian cell lines. Int J Mol Sci. 2016;17(1):72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Fordham JB, Naqvi AR, Uttamani JR, Kulkarni V, Nares S. MicroRNA: Dynamic Regulators of Macrophage Polarization and Plasticity. Frontiers Imm 2017;8:1062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Zhang Y, Zhang M, Li X, et al. Silencing MicroRNA-155 attenuates cardiac injury and dysfunction in viral myocarditis via promotion of M2 phenotype polarization of macrophages. Sci Rep 2016;6:22613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Ying H, Kang Y, Zhang H, et al. MiR-127 modulates macrophage polarization and promotes lung inflammation and injury by activating the JNK pathway. J Immunol. 2015;194(3):1239–1251. [DOI] [PubMed] [Google Scholar]
- 84.Huang C, Liu XJ, QunZhou, et al. MiR-146a modulates macrophage polarization by inhibiting Notch1 pathway in RAW264.7 macrophages. Int Immunopharmacol. 2016;32:46–54. [DOI] [PubMed] [Google Scholar]
- 85.Venkatesan G, Uppoor A, Naik DG. Redefining the role of dendritic cells in periodontics. J Indian Soc Periodontol. 2013;17:700–705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Wilensky A, Segev H, Mizraji G, Shaul Y, Capucha T, Shacham M, Hovav AH. Dendritic cells and their role in periodontal disease. Oral Dis. 2014;20(2):119–126. [DOI] [PubMed] [Google Scholar]
- 87.Martinez-Nunez RT, Louafi F, Friedmann PS, Sanchez-Elsner T. MicroRNA-155 modulates the pathogen binding ability of dendritic cells (DCs) by down-regulation of DC-specific intercellular adhesion molecule-3 grabbing non-integrin (DC-SIGN). J Biol Chem. 2009;284(24):16334–16342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Dunand-Sauthier I, Santiago-Raber ML, Capponi L, et al. Silencing of c-Fos expression by microRNA-155 is critical for dendritic cell maturation and function. Blood. 2011;117(17):4490–4500. [DOI] [PubMed] [Google Scholar]
- 89.Huffaker TB, O’Connell RM. miR-155-SOCS1 as a functional axis: satisfying the burden of proof. Immunity. 2015;43(1):3–4. [DOI] [PubMed] [Google Scholar]
- 90.Kim SJ, Gregersen PK, Diamond B. Regulation of dendritic cell activation by microRNA let-7c and BLIMP1. J Clin Invest. 2013;123(2):823–833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Mescher MF, Curtsinger JM, Agarwal P, et al. Signals required for programming effector and memory development by CD8 + T cells. Immunol. Rev 2006;211:81–92. [DOI] [PubMed] [Google Scholar]
- 92.Hwang JR, Byeon Y, Kim D, Park SG. Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development. Exp Mol Med. 2020;52(5):750–761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Gutiérrez-Vázquez C, Rodríguez-Galán A, Fernández-Alfara M. et al. miRNA profiling during antigen-dependent T cell activation: A role for miR-132–3p. Sci Rep. 2017;7:3508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Teteloshvili N, Smigielska-Czepiel K, Kroesen BJ, et al. T-cell activation induces dynamic changes in miRNA expression patterns in CD4 and CD8 T-cell subsets. MicroRNA. 2015;4:117–122. [DOI] [PubMed] [Google Scholar]
- 95.Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004;303(5654):83–86. [DOI] [PubMed] [Google Scholar]
- 96.Li QJ, Chau J, Ebert PJ, et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell. 2007;129(1):147–161. [DOI] [PubMed] [Google Scholar]
- 97.Zimmerman EI, Dollins CM, Crawford M, et al. Lyn kinase-dependent regulation of miR181 and myeloid cell leukemia-1 expression: implications for drug resistance in myelogenous leukemia. Mol Pharmacol. 2010;78(5)811–817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Baumjohann D, Ansel KM. MicroRNA-mediated regulation of T helper cell differentiation and plasticity. Nat Rev Immunol. 2013;13(9):666–678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Fu G, Rybakin V, Brzostek J, Paster W, Acuto O, Gascoigne NR. Fine-tuning T cell receptor signaling to control T cell development. Trends Immunol. 2014;35(7):311–318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Jindra PT, Bagley J, Godwin JG, Iacomini J. Costimulation-dependent expression of microRNA-214 increases the ability of T cells to proliferate by targeting Pten. J Immunol. 2010;185(2):990–997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Liu SQ, Jiang S, Li C, Li QJ. miR-17–92 cluster targets phosphatase and tensin homology and ikaros family zinc finger 4 to promote TH17-mediated inflammation. J Biol Chem. 2014;289(18):12446–12456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Yang L, Boldin MP, Yu Y, et al. miR-146a controls the resolution of T cell responses in mice. J Exp Med. 2012;209(9):1655–1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Steiner DF, Thomas MF, Hu JK, et al. MicroRNA-29 regulates T-box transcription factors and interferon-γ production in helper T cells. Immunity. 2011;35(2):169–181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Jiang S, Li C, Olive V, et al. Molecular dissection of the miR-17–92 cluster’s critical dual roles in promoting Th1 responses and preventing inducible Treg differentiation. Blood. 2011;118(20):5487–5497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Garlet GP. Destructive and protective roles of cytokines in periodontitis: a re-appraisal from host defense and tissue destruction viewpoints. J Dent Res. 2010;89(12):1349–1363. [DOI] [PubMed] [Google Scholar]
- 106.Yücel OumlOuml, Berker E, Gariboğlu S, Otlu H. Interleukin-11, interleukin-1β, interleukin-12 and the pathogenesis of inflammatory periodontal diseases. J Clin Periodontol. 2008;35(5):365–370. [DOI] [PubMed] [Google Scholar]
- 107.Banerjee A, Schambach F, DeJong CS, Hammond SM, Reiner SL. Micro-RNA-155 inhibits IFN-γ signaling in CD4+ T cells. Eur J Immunol. 2010;40(1):225–231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Kumar M, Ahmad T, Sharma A, et al. Let-7 microRNA-mediated regulation of IL-13 and allergic airway inflammation. J Allergy Clin. Immunol 2011;128(5):1077–1085. [DOI] [PubMed] [Google Scholar]
- 109.Mattes J, Collison A, Plank M, Phipps S, Foster PS. Antagonism of microRNA-126 suppresses the effector function of TH2 cells and the development of allergic airways disease. Proc Natl Acad Sci USA. 2009;106(44):18704–18709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Dutzan N, Kajikawa T, Abusleme L, et al. A dysbiotic microbiome triggers TH17 cells to mediate oral mucosal immunopathology in mice and humans. Sci Transl Med. 2018;10(463):eaat0797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Rouas R, Fayyad-Kazan H, El Zein N, et al. Human natural Treg microRNA signature: role of microRNA-31 and microRNA-21 in FOXP3 expression. Eur J Immunol. 2009;39(6):1608–1618. [DOI] [PubMed] [Google Scholar]
- 112.Sethi A, Kulkarni N, Sonar S, Lal G. Role of miRNAs in CD4 T cell plasticity during inflammation and tolerance. Front Genet. 2013;4:8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Lu LF, Thai TH, Calado DP, et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity. 2009;30(1):80–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Lu LF, Boldin MP, Chaudhry A, et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell. 2010;142(6):914–929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Fayyad-Kazan H, Rouas R, Fayyad-Kazan M, et al. MicroRNA profile of circulating CD4-positive regulatory T cells in human adults and impact of differentially expressed microRNAs on expression of two genes essential to their function. J Biol Chem. 2012;287(13):9910–9922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Allam JP, Duan Y, Heinemann F, et al. IL-23-producing CD68+ macrophage-like cells predominate within an IL-17-polarized infiltrate in chronic periodontitis lesions. J Clin Periodontol. 2011;38(10):879–886. [DOI] [PubMed] [Google Scholar]
- 117.Zhao L, Zhou Y, Xu Y, Sun Y, Li L, Chen W. Effect of non-surgical periodontal therapy on the levels of Th17/Th1/Th2 cytokines and their transcription factors in Chinese chronic periodontitis patients. J Clin Periodontol. 2011;38(6):509–516. [DOI] [PubMed] [Google Scholar]
- 118.Laurence A, O’Shea JJ. TH-17 differentiation: of mice and men. Nat Immunol. 2007;8():903–905. [DOI] [PubMed] [Google Scholar]
- 119.Liu SQ, Jiang S, Li C, Li QJ. miR-17–92 cluster targets phosphatase and tensin homology and ikaros family zinc finger 4 to promote TH17-mediated inflammation. J Biol Chem. 2014;289(18):12446–12456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Yang L, Boldin MP, Yu Y, et al. miR-146a controls the resolution of T cell responses in mice. J Exp Med. 2012;209(9):1655–1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Escobar TM, Kanellopoulou C, Kugler DG, et al. miR-155 activates cytokine gene expression in Th17 cells by regulating the DNA-binding protein Jarid2 to relieve polycomb-mediated repression. Immunity. 2014;40(6):865–879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Mycko MP, Cichalewska M, Machlanska A, Cwiklinska H, Mariasiewicz M, Selmaj KW. microRNA-301a regulation of a T-helper 17 immune response controls autoimmune demyelination. Proc Natl Acad Sci USA. 2012;109(20):E1248–E1257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Seymour GJ, Powell RN, Cole KL, et al. Experimental gingivitis in humans. A histochemical and immunological characterization of the lymphoid cell subpopulations. J Periodontal Res. 1983;18(4):375–385. [DOI] [PubMed] [Google Scholar]
- 124.Stoufi ED, Taubman MA, Ebersole JL, Smith DJ. Preparation and characterization of human gingival cells. J Periodontal Res. 1987;22(2):144–149. [DOI] [PubMed] [Google Scholar]
- 125.Gururajan M, Haga CL, Das S, et al. MicroRNA 125b inhibition of B cell differentiation in germinal centers. Int Immunol. 2010;22(7):583–592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Porstner M, Winkelmann R, Daum P, et al. miR-148a promotes plasma cell differentiation and targets the germinal center transcription factors Mitf and Bach2. Eur J Immunol. 2015;45(4):1206–1215. [DOI] [PubMed] [Google Scholar]
- 127.Gonzalez-Martin A, Adams BD, Lai M, et al. The microRNA miR-148a functions as a critical regulator of B cell tolerance and autoimmunity. Nat Immunol. 2016;17(4):433–440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.de Yebenes VG, Bartolomé-Izquierdo N, Nogales-Cadenas R, et al. miR-217 is an oncogene that enhances the germinal center reaction. Blood. 2014;124(2):229–239. [DOI] [PubMed] [Google Scholar]
- 129.Teng G, Hakimpour P, Landgraf P, et al. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity. 2008;28(5):621–629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.de Yébenes VG, Belver L, Pisano DG, et al. miR-181b negatively regulates activation-induced cytidine deaminase in B cells. J Exp Med. 2008;205(10):2199–2206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Naqvi AR. Immunomodulatory roles of human herpesvirus-encoded microRNA in host-virus interaction. Rev Med Virol. 2020;30(1):e2081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Ahmad I, Valverde A, Siddiqui H, Schaller S, Naqvi AR. Viral MicroRNAs: Interfering the Interferon Signaling. Curr Pharm Des. 2020;26(4):446–454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Naqvi AR, Shango J, Seal A, Shukla D, Nares S. Viral miRNAs alter host cell miRNA profiles and modulate innate immune responses. Front Immunol. 2018;9:433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Naqvi A, Schaller S, Nares S, Pettiette M, Zhang S, Moretti A, Brambila M, Martinez G, Chapa G. Porphyromonas gingivalis gene expression in subjects with periodontal disease. J Dent Res. 2020;99:2075. [Google Scholar]
- 135.Siddiqui H, Nares S, Pettiette M, Zhang S, Moretti A, Brambila M, Martinez G, Chapa G, Naqvi A. Aggregatibacter actinomycetemcomitans transcript profiles in chronic periodontitis. J Dent Res. 2020;99:972. [Google Scholar]
- 136.Zhang G, Zong J, Lin S, et al. Circulating Epstein-Barr virus microRNAs miR-BART7 and miR-BART13 as biomarkers for nasopharyngeal carcinoma diagnosis and treatment. Int J Cancer. 2015;136(5):E301–E12. [DOI] [PubMed] [Google Scholar]
- 137.Chan JY, Gao W, Ho WK, Wei WI, Wong TS. Overexpression of Epstein-Barr virus-encoded microRNA-BART7 in undifferentiated nasopharyngeal carcinoma. Anticancer Res. 2012;32(8):3201–3210. [PubMed] [Google Scholar]
- 138.Fujimori K, Yoneda T, Tomofuji T, et al. Detection of salivary miRNAs reflecting chronic periodontitis: A pilot study. Molecules. 2019;24(6):1034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Al-Rawi NH, Al-Marzooq F, Al-Nuaimi AS, Hachim MY, Hamoudi R. Salivary microRNA 155, 146a/b and 203: A pilot study for potentially non-invasive diagnostic biomarkers of periodontitis and diabetes mellitus. PLoS One. 2020;15(8):e0237004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Nik Mohamed Kamal NNS, Awang RAR, Mohamad S, Shahidan WNS. Plasma- and saliva exosome profile reveals a distinct MicroRNA signature in chronic periodontitis. Front Physiol. 2020;11:587381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Kole R, Krainer AR, Altman S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat Rev Drug Discov. 2012;11(2):125–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Krongbaramee T, Zhu M, Qian Q, et al. Plasmid encoding microRNA-200c ameliorates periodontitis and systemic inflammation in obese mice. Mol Ther Nucleic Acids. 2021;23:1204–1216. [DOI] [PMC free article] [PubMed] [Google Scholar]


