Abstract
The Helicobacter pylori iceA gene was recently identified as a genetic marker for the development of peptic ulcer in a Western population. To assess the significance of iceA subtypes of H. pylori in relation to peptic ulcer, 140 Japanese clinical isolates (88 from Fukui and 52 from Okinawa) were characterized. Sequence analysis of the iceA1 gene from 25 representative Japanese strains was also carried out to identify the differences in iceA between the ulcer group and the gastritis group. The iceA1 genotype was not correlated with the presence of peptic ulceration in either area. In addition, sequence analysis led to identification of five deletions and five point mutations (a nonsense mutation or a 1-bp insertion) within the iceA1 open reading frame corresponding to previously published sequences. These mutations were identified in both clinical groups (ulcer and gastritis groups) in each area. Local DNA sequence analysis revealed that the endpoints of all five deletions coincided with direct repeats. We also found four strains that carried longer iceA1 open reading frames compared with that for strain 60190. In conclusion, carriage of an iceA1 strain does not seem to be a risk factor for peptic ulcer in Japanese subjects. The critical mutations in the iceA1 gene in some isolates from patients with peptic ulcers suggested that IceA does not participate in the pathogenesis of peptic ulcer in Japan. We also found deletion hot spots that were associated with direct repeats in iceA1 and that favored a small-deletion model of slipped mispairing events during replication. We showed that iceA1 sequence variations may be useful tools for analysis of the population genetics of H. pylori.
Helicobacter pylori infection is now recognized as a significant risk factor for both gastric and duodenal ulcers, gastric adenocarcinoma, and MALT lymphoma (11, 16, 17, 20). However, only a minority of infected patients develop such severe diseases, and most infected individuals are asymptomatic throughout life. Such variations in clinical outcome may be due to the considerable genetic diversity of the H. pylori strains that cause infection, although host factors may also be important for the development of disease.
Strains that possess two virulence factors, vacuolating cytotoxin encoded by vacA and a 40-kb DNA segment named the cag pathogenicity island (4), are associated with severe diseases in the West (3, 6, 7, 9, 21–23). The vacA genotype (3) and the presence of cagA (which is at one end of cag pathogenicity island) are mainly used as simple genetic markers for these virulence traits. However, close associations between these virulence factors and clinical presentation have not been confirmed in East Asian populations, and most isolates in this region were cagA positive and of the vacA s1 type, regardless of clinical manifestations (12, 13, 15, 26). Some sequence studies have shown that these two virulence-associated genes (cagA and vacA) are distinct between strains from East Asia and Europe (12, 25). Recent studies of the population genetics of H. pylori strains from diverse geographic locations demonstrated that Asian strains were assigned to the same “Asian” clonal group, probably reflecting descent from distinct common ancestors, and cagA and vacA are more diverse than other housekeeping genes among strains (1). Therefore, different combinations of bacterial and/or human factors may be critical determinants of disease in East Asian population. Moreover, sequence analysis of virulence genes may provide useful makers for study of the population genetics of H. pylori.
Recently, Peek et al. (18) reported on a novel H. pylori gene, iceA (induced by contact with epithelium). There are two different alleles of this gene (iceA1 and iceA2), and iceA1 is replaced by iceA2 at the same locus in many strains. iceA2 is a gene that is completely unrelated to iceA1 or other known proteins. Although iceA1 encodes a homolog of a putative restriction endonuclease (nlaIIIR) of Neisseria lactamica (14), the role of the iceA gene product during human infection remains unknown and the translational start site of iceA1 has not yet been confirmed (23). Interestingly, carriage of iceA1 was shown to be weakly but significantly associated with peptic ulcer in studies of H. pylori strains from Europe (Holland) and the United States (18, 23).
In the study described here, we assessed the linkage between iceA genotype and peptic ulcer in two different areas of Japan. Moreover, we sequenced and analyzed the full-length iceA gene from isolates obtained from patients with peptic ulcer or chronic gastritis and also examined whether there are different sequence motifs or segments that can be used to distinguish between Japanese and Western strains.
MATERIALS AND METHODS
Patients.
Two different areas of Japan were selected as sources of strains for iceA analysis. Fukui is a typical rural prefecture located in the central Japanese mainland (Honshu), while Okinawa consists of islands in the southwestern part of Japan and has a history and food culture different from those for other parts of Japan. Clinical isolates of H. pylori were obtained from 140 Japanese patients (88 patients from Fukui and 52 patients from Okinawa) during gastroduodenal endoscopy at the Second Department of Internal Medicine, Fukui Medical University, Fukui, and Division of Internal Medicine, Okinawa Chubu Hospital, Okinawa, respectively. The 88 patients in Fukui consisted of 49 with peptic ulcer (mean age, 53.7 years) and 39 with chronic gastritis (mean age, 58.3 years). The 52 Okinawan patients consisted of 19 with peptic ulcer (mean age, 58.3 years) and 33 with chronic gastritis (mean age, 59.6 years). Patients underwent endoscopy because of upper abdominal complaints or as part of an annual health check. Ulcer was defined as sharply delineated mucosal defects of at least 5 mm in one dimension, with depth. Patients with ulcer scars showing retraction with converging folds at endoscopy and the presence of past medical or endoscopic records of ulcer were included in the ulcer group. Patients who had received nonsteroidal anti-inflammatory drugs or antacids were excluded from this study, and none of the patients had recently been prescribed antibiotics.
Isolation and culture of H. pylori.
Two gastric biopsy specimens were sequentially taken from the gastric body and antrum with a sterilized endoscope. Biopsy samples were immediately put into Cary-Blair-N transport medium (Nissui Seiyaku Co., Tokyo, Japan) and were cultured on Trypticase Soy Agar-II plates containing 5% sheep blood (Nippon Becton Dickinson, Tokyo, Japan) for 5 days at 37°C under microaerobic conditions (5% O2, 15% CO2, 80% N2). The specimens obtained from Okinawan patients were sent by air in the same transport medium in an icebox and were cultured within 24 h after biopsy. A single colony from each patient was picked from the culture plate and was inoculated onto another fresh culture plate. A few colonies from the second culture plate were inoculated into 20 ml of brucella broth containing 10% fetal calf serum and were cultured for 3 days under the same conditions described above. H. pylori cells were harvested from the bacterial suspensions by centrifugation at 1,300 × g for 10 min. Chromosomal DNA was extracted from the pellets by the protease and phenol-chloroform method, suspended in 300 μl of TE buffer (10 mM Tris HCl, 1 mM EDTA), and stored at 4°C until PCR amplification. The type strains NCTC 11916 and NCTC 11637 were also analyzed for their iceA genotypes.
Typing of iceA gene.
On the basis of information from a previous study (25), two primer sets, iceA1F (5′-GTGTTTTTAACCAAAGTATC-3′) and iceA1R (5′-CTATAGCCASTYTCTTTGCA-3′) for iceA1 and iceA2F GTTGGGTATATCACAATTTAT 3′) and iceA2R (5′-TTRCCCTATTTTCTAGTAGGT-3′) for iceA2, were used to determine the iceA genotype (23). PCR was performed in 50-μl reaction mixtures containing 1 μl of genomic DNA (50 to 100 ng), 250 nM (each) primer, 1× reaction buffer, 1.5 mM MgCl2, 1 U of AmpliTaq DNA polymerase, and distilled, sterilized water in a GeneAmp 2400 PCR system (Perkin-Elmer Japan, Chiba, Japan). After boiling at 94°C for 5 min, amplification was carried out for 25 cycles of 94°C for 30 s, 50°C for 30 s, and 72°C for 30 s. The mixture was then held at 72°C for 7 min to complete the elongation step and was finally stored at 4°C. PCR products were separated by 2% agarose gel electrophoresis and were examined under UV illumination.
cagA status and vacA genotype were determined previously by PCR-based typing or DNA sequencing for vacA and by hybridization for cagA, respectively (12; Y. Ito, T. Azuma, S. Ito, H. Suto, H. Miyaji, Y. Yamazaki, M. Kuriyama, Y. Kohli, and Y. Keida, Gastroenterology 116, abstr. A196, 1999).
DNA sequencing of entire iceA gene.
Fourteen isolates from Fukui (7 from patients with peptic ulcer and 7 from patients with chronic gastritis only) and 11 isolates from Okinawa (5 from patients with peptic ulcer and 6 from patients with chronic gastritis only) were randomly selected for sequence analysis of iceA1. The region comprising the entire iceA region of each isolate was amplified with the iceA-spanning primer set IAS-F and IAS-R (Table 1). PCR conditions were as follows: heating at 94°C for 5 min, followed by 25 cycles consisting of 94°C for 30 s, 55°C for 30 s, and 72°C for 30 s. The tubes were held at 72°C for 7 min, before storage at 4°C. The PCR products were then purified with Centricon-100 Concentrator columns (Amicon, Beverly, Mass.). DNA sequencing was performed by the dideoxynucleotide chain termination method with a BigDye Terminator Cycle Sequencing Ready Reaction Mix (Perkin-Elmer Japan) in an ABI PRISM 310 Genetic Analyzer (Perkin-Elmer Japan). According to the manufacturer's protocol, reagent mixtures containing 5 μl of purified PCR product, 3.2 pmol of primer, 8 μl of Terminator Cycle Sequencing Ready Reaction Mix, and 5 μl of sterilized distilled water were prepared. Reaction tubes were placed in the thermal cycler, and thermal cycling was started under the following conditions: 96°C for 10 s, 50°C for 5 s, and 60°C for 4 min, which was repeated for 25 cycles. Cycle sequencing reactions were performed for both DNA strands. Nucleotide sequences were aligned and analyzed with GENTYX-MAC, version 8.0 (Software Development Co., Tokyo, Japan).
TABLE 1.
Primers used for sequence analysis of iceA
| Procedure and primer (direction) | Nucleotide sequencea | Position |
|---|---|---|
| iceA1 sequencing | ||
| IAS-F (+) | 5′-CGTGGGCGATGATGTGAAGATTG-3′ | 426–446b |
| IAS-R (−) | 5′-CGTCCCAGCGAACAGATCACAA-3′ | 1403–1424b |
| IAS-2F (+) | 5′-GGCAACTCTGAAAACACTC-3′ | 934–952b |
| IAS-2R (+) | 5′-TCCTGTATTGTATGGGGTC-3′ | 1192–1210b |
| Mutation screening | ||
| iceA1 1033C/T (+) | 5′-CAGACTCTTGATGATTTCT-3′ | 489–511c |
| IAS-R (−) | 5′-CGTCCCAGCGAACAGATCACAA-3′ | 1403–1424b |
| 1172 del5 (+) | 5′-TGAATATGACGGTTGCT-3′ | 661–677d |
| IAS-R (−) | 5′-CGTCCCAGCGAACAGATCACAA-3′ | 1403–1424b |
Statistics.
The association between iceA genotype and clinical outcome in both areas were analyzed by Fisher's exact probability test. A P value of less than 0.05 was considered to indicate statistical significance.
Nucleotide sequence accession numbers.
The DNA sequences of the iceA1 genes of each strain characterized here were deposited in the GenBank database (accession nos. AF157527 to AF157551).
RESULTS
Typing of iceA.
A total of 140 Japanese isolates and 2 type strains (strains NCTC 11916 and NCTC 11637) were investigated for determination of their iceA genotypes. The distributions of iceA genotypes in two areas of Japan (Fukui and Okinawa) are shown in Table 2. One hundred thirty-six Japanese isolates (97.1%) could be typed by this PCR-based method, and no isolates containing both iceA1 and iceA2 were found. For four Fukui strains, no PCR products were obtained with the allele-specific primers. DNA sequencing of full-length iceA determined the genotypes of these untypeable strains. Although these isolates were of the iceA1 type, deletion of nucleotides corresponding to the iceA1F primer region prevented typing (data not shown).
TABLE 2.
Distribution of iceA genotype in each area (Fukui and Okinawa)
| Genotype | No. (%) of strains
|
|||
|---|---|---|---|---|
| Fukuia
|
Okinawab
|
|||
| Peptic ulcer | Chronic gastritis | Peptic ulcer | Chronic gastritis | |
| iceA1 | 33 (67.3) | 26 (66.7) | 15 (78.9) | 25 (75.8) |
| iceA2 | 16 (32.7) | 13 (33.3) | 4 (21.1) | 8 (24.2) |
| Total | 49 | 39 | 19 | 33 |
P = 0.617.
P = 0.538.
On amplification of iceA1, the sizes of the PCR products were slightly different among isolates, suggesting the existence of a deletion in this region. van Doorn et al. (23) reported that there are two iceA2 type strains; one showed an amplicon of 229 bp with the iceA2-specific primer set, and the other yielded a 334-bp amplicon containing a 105-bp in-frame duplication. All Japanese iceA2 type strains yielded an amplicon of 229 bp. In contrast, NCTC 11916 and NCTC 11637 were found to contain an iceA2 gene yielding a 334-bp amplicon. As shown in Table 2, there was no significant association between iceA genotype and peptic ulcer disease in each area (Fukui, P = 0.617; Okinawa, P = 0.538). Isolates possessing iceA1 were found more frequently in Okinawa (76.9%) than in Fukui (67.0%), but the difference was not significant (P = 0.147). As shown in our previous study, all of the Japanese strains in this study were cagA positive. Moreover, almost all Fukui strains and more than 80% of Okinawan strains were of the s1/m1 vacA type. No significant associations were found between these two virulence-associated gene subtypes and the iceA genotype.
Sequence analysis of iceA1.
Primer set IAS-F and IAS-R successfully amplified the entire iceA1 regions of all 25 strains. The sequence data for the iceA1 genes from 25 Japanese strains and 5 Western strains deposited in the GenBank database are summarized in Table 3. Alignment of the iceA1 gene between Japanese strains and 60190 (GenBank accession no. U43917) revealed that the iceA1 gene in many Japanese strains contained in-frame deletions compared with that of strain 60190. We found small deletions of six nucleotides, TAATTT at nucleotide 780, AATTTG at nucleotide 781, or TTTGGA at nucleotide 783, of the iceA open reading frame of strain 60190. These three deletions were considered to be caused by same deletion mechanism (at the same direct repeats), and the deletion of six nucleotides at nucleotide 780 (780del6) was found more frequently than the other deletions. Therefore, these three deletions are arbitrarily designated 780del6. Another deletion of 24 nucleotides at nucleotide 1246 was designated 1246del24. Three deletions, a five-nucleotide deletion at nucleotide 809 (809del5), a seven-nucleotide deletion at nucleotide 914 (914del7), and a five-nucleotide deletion at nucleotide 1172 (1172del5), caused frameshifts that resulted in early translational termination of iceA1. These in-frame deletions were frequently identified among Japanese strains (Tables 3 and 4). For example, 809del5 was found in 14 (56.0%) of 25 Japanese isolates and 914del7 was identified in 11 isolates (44.0%). Furthermore, three different deletions (780del6, 809del5, and 914del7) were simultaneously observed in nine Japanese isolates. The local DNA sequences surrounding these five deletions revealed that these deletion endpoints coincided with direct repeats of between 5 and 10 nucleotides (Table 4).
TABLE 3.
Sequence analysis of iceA1 gene from H. pylori strains in Japana
| Location, disease, and strain | cagA | vacA | Cytotoxin activityf | iceA ORF (bp)c | In-frame deletionb
|
Point mutation | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| 780del6 | 809del5 | 914del7 | 1172del5 | 1246del24 | ||||||
| Fukui | ||||||||||
| CG (n = 7) | ||||||||||
| F13 | + | s1/m1 | 384 | + | + | − | − | − | ||
| F15 | + | s1/m1 | 1 | 384 | + | + | − | − | − | |
| F16 | + | s1/m1 | 2 | 117 | + | + | + | − | + | 1033C/T |
| F36 | + | s1/m1 | 2 | 534 | − | − | − | − | − | |
| F37 | + | s1/m1 | 1 | 519 | − | − | − | − | − | |
| F38 | + | s1/m1 | 6 | 684 | − | − | − | − | − | |
| F43 | + | s1/m1 | 7 | 696 | − | − | − | − | − | |
| PUD (n = 7) | ||||||||||
| F70 | + | s1/m1 | 0 | 360 | + | + | + | − | + | |
| F72 | + | s1/m1 | 4 | 684 | − | − | − | − | − | |
| F73 | + | s1/m1 | 3 | 384 | − | + | + | − | − | |
| F79 | + | s1/m1 | 7 | 117 | + | + | + | − | + | 1033C/T |
| F82 | + | s1/m1 | 4 | 360 | + | + | + | − | + | |
| F83 | + | s1/m1 | 5 | 360 | + | + | + | − | + | |
| F84 | + | s1/m1 | 7 | 117 | + | + | + | − | + | 1033C/T |
| Total no. pos/total no. | 8/14 | 9/14 | 7/14 | 0/14 | 6/14 | |||||
| Okinawa | ||||||||||
| CG (n = 6) | ||||||||||
| OK104 | + | s1/m1 | 1 | 267 | + | + | − | + | − | |
| OK106 | + | s1/m1 | 1 | 384 | + | + | + | − | − | 1059insT, 1090insA |
| OK107 | + | s1/m2 | 0 | 141 | − | − | − | + | − | 1059insT |
| OK111 | + | s1/m1 | 2 | 141 | + | − | − | − | − | |
| OK115 | + | s1/m1 | 2 | 510 | − | − | − | − | + | 786G/A |
| OK129 | + | s1/m2 | 0 | 357 | − | − | − | − | + | |
| PUD (n = 5) | ||||||||||
| OK99 | + | s1/m1 | 4 | 384 | + | + | + | − | − | |
| OK102 | + | s1/m1 | 2 | 384 | + | + | + | − | − | |
| OK108 | + | s1/m1 | 0 | 684 | − | − | − | − | − | |
| OK113 | + | s1/m1 | 0 | 357 | − | − | − | − | + | 904C/T |
| OK134 | + | s1/m1 | 1 | 384 | + | + | + | − | − | |
| Total no. pos/total no. | 6/11 | 5/11 | 4/11 | 2/11 | 2/11 | |||||
| Alaskad | ||||||||||
| 209 | Unknown | Unknown | Unknown | 684 | − | − | − | − | − | |
| 214 | Unknown | Unknown | Unknown | 459 | − | − | − | − | − | 1059delT |
| 218 | Unknown | Unknown | Unknown | 660 | − | − | − | − | + | |
| Total no. pos/total no. | 0/3 | 0/3 | 0/3 | 0/3 | 1/3 | |||||
| United Kingdom | ||||||||||
| 60190 | + | s1/m1 | Pos | 584 | − | − | − | − | − | |
| 26695e | + | s1/m1 | Pos | 516 | − | − | − | − | − | |
| Total | 0/2 | 0/2 | 0/2 | 0/2 | 0/2 | |||||
Abbreviations: ORF, open reading frame; CG, chronic gastritis; PUD, peptic ulcer disease; del, deletion; ins, insertion; pos, positive.
+, presence of deletion; −, absence of deletion.
Nucleotide corresponding to published sequence of H. pylori 60190 including the iceA open reading frame and upstream sequence (GenBank accession no. U43917).
Accession no. AF001537-9.
Accession no. AE000626.
The relative activity of vacuolating cytotoxin was defined as the index number at the maximum dilution, as described previously (12).
TABLE 4.
Local DNA sequences surrounding deletions in iceA1 gene
| Deletion | Strainb | Sequencea | Repeated element | No. of isolates positive/total no. of isolates (%) |
|---|---|---|---|---|
| 780del6 | 60190 | AGAATTTAATTTGGAGTTT | AGAATTT or AATTT | 14/25 (56.0) |
| F38 | AGAATTTGAATTAGAATTT | |||
| F82 | AGAATT AGAATTT | |||
| 809del5 | 60190 | AAACTCTAGGAAATTCTA | AAAYTCTA | 14/25 (56.0) |
| F38 | AAACTCTAGGAAATTCTA | |||
| F82 | AAACTC AATTCTA | |||
| 914del7 | 60190 | GTGTGGTGTGCGTGGC | GTGCG or GTGTGS | 11/25 (44.0) |
| F38 | GTGCGGTGTGCGTGGC | |||
| F82 | GTGCG TGGC | |||
| 1172del5 | 60190 | GGTTGTGTGGGCTGTTAT | GGTTGT | 2/25 (8.0) |
| F38 | GGTTGTGTGGGTTGTTAT | |||
| OK104 | GGTTGC TTGTTAT | |||
| 1246del24 | 60190 | GGGTATCAAAAAGGCTATG | GGGTATCAAA | 9/25 (36.0) |
| F38 | GGGTATCAAAAAGGTTATT | |||
| F82 | GGGTATCAAA | |||
| 1246del24 | 60190 | GTGATGGGTATCAAATTGG | ||
| F38 | ATGAGGGGTATCAAATTGG | |||
| F82 | TTGG |
Nucleotides corresponding to the iceA open reading frame and its upstream sequence of strain 60190 (GenBank accession no. U43917). del, deletion.
Strains 60190 and F38 are representative Western and Japanese strains, respectively. F82 and OK104 are representative strains carrying each mutation. Underlined nucleotides indicate direct repeats (overlapping in 914del7). Y is T or C, and S is G or C.
In addition to deletions, five different point mutations that caused early translational termination of iceA1 were identified in seven strains (Table 3). Due to combinations of these mutations (frameshift and point mutations), the length of the predicted iceA1 open reading frame was different among strains (range, 117 to 696 bp). Of interest, 4 Japanese strains (strains F38, F72, OK108 and F43) possessed a longer iceA1 open reading frame (684 or 696 bp) than that of strain 60190 (534 bp). A putative ribosomal binding site (AGGA) was identified 8 bp upstream of the ATG translational initiation codon. The lengths of the predicted amino acid sequences of iceA1 from these four Japanese isolates were similar to that of NlaIIIR of N. lactamica (228 or 232 versus 230 amino acids). The amino acid identities of the iceA1 open reading frames from these four isolates (isolates F38, F72, OK108, and F43) and NlaIIIR were 56.2, 53.4, 57.1, and 53.4%, respectively.
Detection of specific mutation of iceA1 gene.
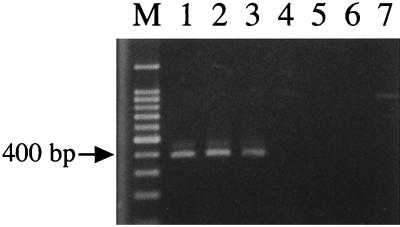
Strains with two mutations, 1033C/T (C-to-T nonsense mutation at nucleotide 1033) and 1072del5, were selected for further analysis because carriage of these mutations causes early translational termination of iceA1. We screened 99 Japanese isolates with the iceA1 genotype for these mutations by PCR amplification using the specific primer sets listed in Table 1. We evaluated the performances of these two primer sets for detection of each mutation by using 25 strains that were sequenced as described above and confirmed the usefulness of these primer sets (Fig. 1). A nonsense mutation (1033C/T) was identified in 5 (8.5%) of 59 iceA1 type strains from Fukui, while this mutation was not found in any of 40 Okinawan strains examined. In contrast, 1072del5 was identified in 7 (17.5%) of 40 Okinawan iceA1 strains but in none of 59 Fukui strains.
FIG. 1.
Detection of 1033C/T nonsense mutation by PCR. Lanes 1 to 3, strains carrying the 1033C/T mutation (strains F16, F79 and F84, respectively); lanes 4 to 7, strains without 1033C/T (strains F13, F15, F82, and F83, respectively); lane M, size marker (100-bp ladder).
DISCUSSION
In this study, we showed that there were no significant differences in the proportions of strains with the iceA1 genotype between peptic ulcer and chronic ulcer groups. We also showed that the iceA1 genotype did not appear to be a reliable marker of peptic ulcer disease among Japanese subjects in two areas separated by more than 1,300 km. Our data were inconsistent with those of recent reports in the West (18, 23) suggesting a significant association between iceA1 genotype and peptic ulcer. It is possible that asymptomatic patients infected with iceA1 type strains develop peptic ulcer disease later in life. However, this is unlikely because the difference in mean age among the two clinical groups was not significant. We also could not exclude the possibility that patients with peptic ulcer may be simultaneously infected with iceA1 and iceA2 strains because we used a single representative strain to assess the iceA genotype in each patient. However, concurrent infection with more than two different strains assessed by randomly amplified polymorphic DNA fingerprinting in Japanese patients (Fukui) occurs at a frequency of only about 5% (unpublished data). In addition, iceA1 type strains were more frequently found in Okinawa (76.9%) than in Fukui (67.0%), although the number of patients with peptic ulcers in the Okinawan group (36.5%) was smaller than the number in the Fukui group (55.7%). Therefore, our results could not support previous studies in which iceA1 type strains were shown to have a virulence potential.
Sequence analysis of the iceA1 genes from 25 strains was also carried out to obtain new insight into the genetic differences between the peptic ulcer group and the gastritis group. Interestingly, we found five deletions within the iceA1 open reading frame of Japanese strains compared with previously published sequences, and these deletions were frequently identified in Japanese isolates, suggesting deletion hot spots in iceA1. It is well known that local DNA sequences containing repeat sequences (direct repeats or inverted repeats) may cause deletion by misalignment during DNA replication or recombination (8, 19). In this study, we found that deletion endpoints coincided with direct repeats. Therefore, these direct repeats may be responsible for each deletion identified within the iceA1 genes of Japanese strains. However, we found three deletions simultaneously among Japanese isolates. Moreover, Japanese strains have highly homologous vacA and cagA genes (12). Therefore, a founder effect may also participate in the high prevalence of in-frame deletions in Japanese strains as well as mutational hot spots.
In this study, we also found five different point mutations that caused premature termination of iceA1 translation. Recently, we reported that various vacA mutations are responsible for a deficiency of cytotoxin activity among Japanese strains (13). Therefore, it is likely that iceA1 containing such mutations has no function, although we could not exclude the possibility that the truncated protein remains functional. Recent reports suggested that point mutations in specific H. pylori genes were shown to be associated with disease (5) or antibiotic resistance (10, 24). Therefore, we expected that truncating mutations of iceA1 occurred only in strains from patients with gastritis. However, these truncating mutations were identified in iceA1 genes from not only the gastritis group but also the peptic ulcer group, and the shortest open reading frame (117 bp) in iceA1 was identified in isolates from both clinical groups. These results suggested that the iceA1 gene product does not contribute critically to the pathogenesis of peptic ulcer disease in the Japanese population.
In this study, we found four Japanese isolates that possess an iceA open reading frame longer than that of strain 60190. The predicted amino acid sequences of iceA1 gene products from these four isolates were similar to that of the NlaIIIR protein. Further studies should be carried out to determine whether 228- or 232-amino-acid IceA1 proteins are functional and the truncated IceA1 is not. Recently, Raudonikiene and Berg (A. Raudonikiene and D. E. Berg, GenBank accession nos. AF001537 to AF001539) reported on three iceA1-type Alaska strains with a different translational start point from that of strain 60190, as shown in this study, and two of these three strains contained mutations, 1246del24 and 1059insT, respectively.
We also showed geographical variation in iceA1 mutations. Four deletions (780del6, 809del5, 914del7, and 1246del24) were distributed uniformly in both areas, suggesting a very early origin. In contrast, a nonsense mutation at nucleotide 1033 (1033C/T) was identified only among the Fukui strains, while 5-bp deletion at nucleotides 1172 to 1176 (1172del5) was found only among Okinawan strains. Therefore, the profile of iceA1 mutations may represent the nature of H. pylori lineages. Most other H. pylori genes studied to date were highly polymorphic only for point mutations and not for insertion or deletion differences. This study indicated that various alleles of iceA1 may be very useful as indicators of evolutionary divergence in H. pylori gene pools from different regions, for the tracing of strain types, and for the differentiation of single versus multiple infections.
In conclusion, we could not confirm the significance of iceA genotyping as a predictor of peptic ulcer disease, and we found that many Japanese isolates contained iceA1 mutations, regardless of the clinical outcome. These results indicated that the iceA genotype is not a useful marker of virulence in this population and that the progression from gastritis to peptic ulcer must require some other genes or factors including the genetic susceptibility of the host. However, we showed that iceA1 sequence variations may provide useful markers that indicate the divergence of H. pylori strains from different geographic regions.
ACKNOWLEDGMENTS
We thank Douglas E. Berg for critical reading of the manuscript and valuable comments, Manabu Inuzuka for stimulating discussion, and Syuko Murayama and Akiyo Yamakawa for technical assistance.
This work was supported by a grant-in-aid for Scientific Research (grant 11670555) from the Ministry of Education, Science, Sports and Culture of Japan.
REFERENCES
- 1.Achtman M, Azuma T, Berg D E, Ito Y, Morelli G, Pan Z-J, Suerbaum S, Thompson S A, van der Ende A, van Doorn L-J. Recombination and clonal groupings within Helicobacter pylori from different geographical regions. Mol Microbiol. 1999;32:459–470. doi: 10.1046/j.1365-2958.1999.01382.x. [DOI] [PubMed] [Google Scholar]
- 2.Akopyants N S, Clifton S W, Kersulyte D, Crabtree J E, Youree B E, Reece C A, Bukanov N O, Drazek E S, Roe B A, Berg D E. Analysis of the cag pathogenicity island of Helicobacter pylori. Mol Microbiol. 1998;28:37–53. doi: 10.1046/j.1365-2958.1998.00770.x. [DOI] [PubMed] [Google Scholar]
- 3.Atherton J C, Cao P, Peek R M, Tummuru M K R, Blaser M J, Cover T L. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. J Biol Chem. 1995;270:17771–17777. doi: 10.1074/jbc.270.30.17771. [DOI] [PubMed] [Google Scholar]
- 4.Censini S, Lange C, Xiang Z, Crabtree J E, Ghiara P, Borodovsky M, Rappuoli R, Covacci A. cag, a pathogenicity island of Helicobacter pylori, encodes type I-specific and disease-associated virulence factors. Proc Natl Acad Sci USA. 1996;93:14648–14653. doi: 10.1073/pnas.93.25.14648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chang C-S, Chen L-T, Yang J-C, Lin J-T, Chang K-C, Wang J-T. Isolation of a Helicobacter pylori protein, FldA, associated with mucosa-associated lymphoid tissue lymphoma of the stomach. Gastroenterology. 1999;117:82–88. doi: 10.1016/s0016-5085(99)70553-6. [DOI] [PubMed] [Google Scholar]
- 6.Cover T L, Dooley C P, Blaser M J. Characterization of and human serologic response to proteins in Helicobacter pylori broth culture supernatants with vacuolizing cytotoxin activity. Infect Immun. 1990;58:603–610. doi: 10.1128/iai.58.3.603-610.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Crabtree J E, Taylor J D, Wyatt J I, Heatley R V, Shallcross T M, Tompkins D S, Rathbone B J. Mucosal IgA recognition of Helicobacter pylori 120 kDa protein, peptic ulceration, and gastric pathology. Lancet. 1991;338:332–335. doi: 10.1016/0140-6736(91)90477-7. [DOI] [PubMed] [Google Scholar]
- 8.Farabaugh P J, Schmeissner U, Hofer M, Miller J H. Genetic studies on the lac repressor. VII. On the molecular nature of spontaneous hotspots in the lacI gene of Escherichia coli. J Mol Biol. 1978;126:847–863. doi: 10.1016/0022-2836(78)90023-2. [DOI] [PubMed] [Google Scholar]
- 9.Figura N, Guglielmetti P, Rossolini A, Barberi A, Cusi G, Musmanno R A, Russi M, Quaranta S. Cytotoxin production by Campylobacter pylori strains isolated from patients with peptic ulcers and from patients with chronic gastritis only. J Clin Microbiol. 1989;27:225–226. doi: 10.1128/jcm.27.1.225-226.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Goodwin A, Kersulyte D, Sisson G, van Zanten S J O V, Berg D E, Hoffman P S. Metronidazole resistance in Helicobacter pylori is due to null mutations in a gene (rdxA) that encodes an oxygen-insensitive NADPH nitroreductase. Mol Microbiol. 1998;28:383–393. doi: 10.1046/j.1365-2958.1998.00806.x. [DOI] [PubMed] [Google Scholar]
- 11.Hentschel E, Brandstatter G, Dragosics B, Hirschl A M, Nemec H, Schutze K, Taufer M, Wurzer H. Effect of ranitidine and amoxicillin plus metronidazole on the eradication of Helicobacter pylori and the recurrence of duodenal ulcer. N Engl J Med. 1993;328:308–312. doi: 10.1056/NEJM199302043280503. [DOI] [PubMed] [Google Scholar]
- 12.Ito Y, Azuma T, Ito S, Miyaji H, Hirai M, Yamazaki Y, Sato F, Kato T, Kohli Y, Kuriyama M. Analysis and typing of the vacA gene from cagA-positive strains of Helicobacter pylori isolated in Japan. J Clin Microbiol. 1997;35:1710–1714. doi: 10.1128/jcm.35.7.1710-1714.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ito Y, Azuma T, Ito S, Suto H, Miyaji H, Yamazaki Y, Kohli Y, Kuriyama M. Full-length sequence analysis of the vacA gene from cytotoxic and noncytotoxic Helicobacter pylori. J Infect Dis. 1998;178:1391–1398. doi: 10.1086/314435. [DOI] [PubMed] [Google Scholar]
- 14.Morgan R D, Camp R R, Wilson G G, Xu S-Y. Molecular cloning and expression of NlaIII restriction-modification system in E. coli. Gene. 1996;183:215–218. doi: 10.1016/s0378-1119(96)00561-6. [DOI] [PubMed] [Google Scholar]
- 15.Pan Z J, Berg D E, van der Hulst R W M, Su W-W, Raudonikiene A, Xiao S-D, Dankert J, Tytgat G N J, van der Ende A. Prevalence of vacuolating cytotoxin production and distribution of distinct vacA alleles in Helicobacter pylori from China. J Infect Dis. 1998;178:220–226. doi: 10.1086/515601. [DOI] [PubMed] [Google Scholar]
- 16.Parsonnet J, Friedman G D, Vandersteen D P, Chang Y, Vogelman J H, Orentreich N, Sibley R K. Helicobacter pylori infection and the risk of gastric carcinoma. N Engl J Med. 1991;325:1127–1131. doi: 10.1056/NEJM199110173251603. [DOI] [PubMed] [Google Scholar]
- 17.Parsonnet J, Hansen S, Rodriguez L, Gelb A B, Warnke R A, Jellum E, Orentreich N, Vogelman J H, Friedman G D. Helicobacter pylori infection and gastric lymphoma. N Engl J Med. 1994;330:1267–1271. doi: 10.1056/NEJM199405053301803. [DOI] [PubMed] [Google Scholar]
- 18.Peek R M, Thompson S A, Donahue J P, Tham K T, Atherton J C, Blaser M J, Miller G G. Adherence to gastric epithelial cells induces expression of a Helicobacter pylori gene, iceA, that is associated with clinical outcome. Proc Assoc Am Physicians. 1998;110:531–544. [PubMed] [Google Scholar]
- 19.Ripley L S. Frameshift mutation: determinants of specificity. Annu Rev Genet. 1990;24:189–213. doi: 10.1146/annurev.ge.24.120190.001201. [DOI] [PubMed] [Google Scholar]
- 20.Sung J J Y, Chung S C, Ling T K W, Yung M Y, Leung V K S, Ng E K W, Li M K K, Cheng A F B, Li A K C. Antibacterial treatment of gastric ulcers associated with Helicobacter pylori. N Engl J Med. 1995;332:139–142. doi: 10.1056/NEJM199501193320302. [DOI] [PubMed] [Google Scholar]
- 21.Tee W, Lambert J R, Dwyer B. Cytotoxin production by Helicobacter pylori from patients with upper gastrointestinal tract diseases. J Clin Microbiol. 1995;33:1203–1205. doi: 10.1128/jcm.33.5.1203-1205.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tomb J F, White O, Kerlavage A R, Clayton R A, Sutton G G, Fleischmann R D, Ketchum K A, Klenk H P, Gill S, Dougherty B A, Nelson K, Quackenbush J, Zhou L, Kirkness E F, Peterson S, Loftus B, Richardson D, Dodson R, Khalak H G, Glodek A, McKenney K, Fitzgerald L M, Lee N, Adams M D, Hickey E K, Berg D E, Gocayne J D, Utterback T R, Peterson J D, Kelly J M, Cotton M D, Weidman J M, Fujii C, Bowman C, Watthey L, Wallin E, Hayes W S, Borodovsky M, Karp P D, Smith H O, Fraser C M, Venter J C. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388:539–547. doi: 10.1038/41483. [DOI] [PubMed] [Google Scholar]
- 23.van Doorn L-J, Figueiredo C, Sanna R, Plaisier A, Schneeberger P, Boer W D, Quint W. Clinical relevance of the cagA, vacA, and iceA status of Helicobacter pylori. Gastroenterology. 1998;115:58–66. doi: 10.1016/s0016-5085(98)70365-8. [DOI] [PubMed] [Google Scholar]
- 24.Versalovic J, Shortridge D, Kibler K, Griffy M V, Beyer J, Flamm R K, Tanaka S K, Graham D Y, Go M F. Mutations in 23S rRNA are associated with clarithromycin resistance in Helicobacter pylori. Antimicrob Agents Chemother. 1996;40:477–480. doi: 10.1128/aac.40.2.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.van der Ende A, Pan Z-J, Bart A, van der Hulst R W, Feller M, Xiao S D, Tytgat G N, Dankert J. cagA-positive Helicobacter pylori populations in China and The Netherlands are distinct. Infect Immun. 1998;66:1822–1826. doi: 10.1128/iai.66.5.1822-1826.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wang H-J, Kuo C-H, Yeh A A M, Chang P C L, Wang W-C. Vacuolating toxin production in clinical isolates of Helicobacter pylori with different vacA genotypes. J Infect Dis. 1998;178:207–212. doi: 10.1086/515600. [DOI] [PubMed] [Google Scholar]