Summary
Bispecific antibodies (Bispecifics) demonstrate exceptional clinical potential to address some of the most complex diseases. However, Bispecific production in a single cell often requires the correct pairing of multiple polypeptide chains for desired assembly. This is a considerable hurdle that hinders the development of many immunoglobulin G (IgG)-like bispecific formats. Our approach focuses on the rational engineering of charged residues to facilitate the chain pairing of distinct heavy chains (HC). Here, we deploy structure-guided protein design to engineer charge pair mutations (CPMs) placed in the CH3-CH3′ interface of the fragment crystallizable (Fc) region of an antibody (Ab) to correctly steer heavy chain pairing. When used in combination with our stable effector functionless 2 (SEFL2.2) technology, we observed high pairing efficiency without significant losses in expression yields. Furthermore, we investigate the relationship between CPMs and the sequence diversity in the parental antibodies, proposing a rational strategy to deploy these engineering technologies.
Subject areas: Biochemistry, Bioengineering, Biomolecular engineering, Structural biology
Graphical abstract

Highlights
-
•
Crystal structures unveil molecular basis of SEFL2.2 and CPM technologies
-
•
Next gen structure-guided design of CPMs to steer HC-HC pairing
-
•
Top CPMs show high pairing efficiency and optimal expression and stability
-
•
Balancing CPM charge distribution minimizes impact of sequence diversity
Biochemistry; Bioengineering; Biomolecular engineering; Structural biology
Introduction
Designed to recognize two distinct epitopes in the same or different targets, Bispecifics represent a new generation of large molecule therapeutics (Labrijn et al., 2019; Wang et al., 2019). Bispecifics have been gaining traction as a way to confer new therapeutic functionalities, such as the simultaneous engagement of T cells and tumor cells seen in Bispecific T cell Engagers (BiTEs) (Brinkmann and Kontermann, 2017). However, these therapeutics are more complex than conventional monoclonal antibodies (mAbs) and present additional challenges at every stage of development (Brinkmann and Kontermann, 2017; Klein et al., 2012).
IgG is one of the most common scaffolds used to develop Bispecifics (Labrijn et al., 2019). IgG molecules are composed of 2 identical HCs, each pairing with identical light chains (LCs) that fold together into a symmetrical “Y” shape. The HC/HC interactions within a wild type (WT) IgG include the flexible hinge region via disulfide bonds, the CH2/CH2 interface via the N-linked glycans, and in the CH3/CH3 interface via direct protein interactions (Merchant et al., 1998). However, the expression of two non-identical HCs in a single cell will typically result in a bispecific heterodimer and the two corresponding homodimers. Such impurities will affect productivity yields and cost of goods (Carter, 2001; Suresh et al., 1986). In addition, the presence of undesired homodimer species can have toxic effects on the patient. Therefore, to drive the pairing of 2 distinct HCs required for the assembly of hetero-Fc containing Bispecifics, a specific heterodimerization interface must be engineered. As a result, a number of engineering approaches have been developed to increase the ratio of heterodimer to homodimer, including but not limited to knobs-into-holes (Atwell et al., 1997; Merchant et al., 1998; Ridgway et al., 1996), strand-exchange engineered domain body (Davis et al., 2010), electrostatic steering (Gunasekaran et al., 2010; Leaver-Fay et al., 2016; Liu et al., 2015) and other additional approaches (Moore et al., 2011; Von Kreudenstein et al., 2013). However, those technologies require further improvements if Bispecific yields are to be comparable with those of mAbs (Gong et al., 2021).
The electrostatic steering approach, enabled by the incorporation of CPMs, is one of the preferred technologies with many Bispecifics currently in clinical development (Gunasekaran et al., 2010; Labrijn et al., 2019). By introducing negative charges on one chain and positive charges on the other, attractive electrostatic forces drive heterodimerization while repulsive forces prevent homodimer formation. A well-known case study is the CPMv1, where charged residues were rationally manipulated to drive the desired chain pairing in the stable CH3/CH3′ interface (Gunasekaran et al., 2010). However, such approaches, when applied as a platform, often yield suboptimal molecules and may need to be combined with additional engineering technologies. For example, in scenarios where little to no effector function is desired, multiple mutations can abrogate FcγR binding, such as the SEFL2.2 IgG1 scaffold (Jacobsen et al., 2017). However, combining protein engineering solutions developed independently can often have unanticipated results.
Here, we sought to unveil the molecular basis for SEFL2.2 and CPMv1 technologies by determining their crystal structures, which we then used to guide the development of next generation CPMs. Furthermore, we have investigated the optimal deployment of CPMs across the CH3/CH3′ domain interface and explored the impact on stability properties. Altogether, these findings provide valuable insights into further refinement of the engineering of Bispecifics.
Results
Molecular structure of the SEFL2.2 Fc
The SEFL2.2 IgG1 scaffold was originally developed to replace the IgG2 therapeutic scaffold in clinical settings where little to no effector function is desired. This was accomplished through the substitution of N297 with a glycine, resulting in deglycosylation of the Fc and subsequent reduction of FcγR binding (Jacobsen et al., 2017). However, the deglycosylation destabilized the Fc, and thus a covalent disulfide bond was engineered between the R292 and V302 (Jacobsen et al., 2017).
To investigate the molecular basis and the impact of the SEFL2.2 design on the Fc, we sought to determine its crystal structure. Co-crystals of SEFL2.2 Fc were generated in complex with a minimized Z-domain (Mini-Z) peptide. This engineered two-helix bundle of 34 amino acids binds tightly to the CH2-CH3 hinge region in the Fc and serves to optimize Fc intramolecular contacts, improving the conditions for crystal formation (Braisted and Wells, 1996; Starovasnik et al., 1997). Before crystallization, SEFL2.2 Fc was mixed with Mini-Z peptide at 1:2 molar ratio to attain suitable crystals diffracting to 1.95 Å. The asymmetric unit contains a single complex composed of SEFL2.2 Fc (2 chains) interacting with two copies of the Mini-Z peptide at the Fc between CH2 and CH3 (PDB: 7LUR; Figure S1A and Table S1).
To investigate potential conformational changes to the Fc structure upon removal of N297 glycan and the additional disulfide bond modification to create SEFL2.2, we compared this structure with a 2.3 Å crystal structure of WT IgG1 Fc also bound to mini-Z domain (PDB: 1OQO) (Figures 1A and 1B). Overall alignment of SEFL2.2 Fc with WT Fc showed a root-mean-square deviation (RMSD) of 1.2 Å, indicating that the structures were highly similar (Figure S1B). The structural alignment for the CH2 domains alone (chain A SEFL2.2 CH2 superimposed on top of chain A WT CH2) and for the CH3/CH3′ hetero-domain (SEFL2.2 CH3 dimer with WT CH3 dimer) were even more similar with an RMSD of 0.38 Å and 0.31 Å, respectively (Figures 1C and 1D). Despite the high degree of structural conservation observed for these individual domains, major quaternary rearrangements were noted at the CH2/CH2 interface. Indeed, the N-terminus of the SEFL2.2 Fc CH2 domain is 21 Å apart from its counterpart, instead of the 15.5 Å for WT Fc (Figure 1B). This 5.5 Å difference may indicate SEFL2.2 is more dynamic than the WT Fc. This is in agreement with total buried surface area (BSA) calculations (Krissinel and Henrick, 2007), which revealed the chains of SEFL2.2 Fc are slightly more exposed than the WT Fc (2035.7 Å2 vs 2197.9 Å2, respectively). In the WT Fc, the CH2/CH2 interface is solely mediated by sugar-sugar interactions between the 2 N297-linked glycans (Man3NAG2) (Figures 1D and 1E), and the removal of these glycan interactions in the SEFL2.2 can likely be attributed to this more dynamic behavior. As reported previously, the deletion of N297 glycan reduced the thermostability of SEFL2.2 scaffold, which can further indicate a more dynamic protein (Jacobsen et al., 2017). A careful look into the CH2 interface reveals 10 residues that lost their sugar-protein interactions (Figure 1E). Among them, residues with aromatic side chains (F and V) became solvent exposed and others with long and flexible side chains (R and K) noted a possible increase in rotamer conformations.
Figure 1.
Structural analysis of SEFL2.2 Fc
(A) Schematic representation of WT Fc (PDB: 1OQO) and SEFL2.2 Fc (PDB: 7LUR).
(B) Superimposition of WT Fc (gray) with SEFL2.2 Fc (green), shown in cartoon representation with N297 glycan shown in sticks and C292-C302 in yellow spheres. Distances between the CH2 N-termini were measured using the Cα of G237 of each chain. Distances between the CH3 C-termini were measured using Cα of L443 of each chain.
(C) Superimposition of CH3/CH3 dimers.
(D) Superimposition of CH2/CH2 dimers and glycan interface.
(E) Conformational shift of glycan binding residues (in sticks) in the CH2.
(F) Non-canonical disulfide bond representation.
(G) 2Fo-Fc electron density map for non-canonical disulfide bond countered at 1.0ϭ.
(H) Rosetta energy score of WT Fc and 49 variants with different candidate disulfides.
Analysis of the engineered disulfide demonstrated successful cross-linking between β-strands D and E (Figure 1F) (Jacobsen et al., 2017). Although the R292C and V302C mutations enable a correct disulfide bond as demonstrated by the electron density (Figure 1G), we sought to investigate the insertion of other disulfide bonds as well. After identifying alternative residue pairs where the Cβ distance <6.5 Å, we modeled and ranked a total of 49 candidate disulfides using Rosetta (Figures 1H and S2). Of note, most designs resulted in a substantial increase in Rosetta energy scores relative to the parental WT structure (ΔREU>0; Figure 1H), indicating suboptimal mutations. Although Rosetta energy units are arbitrary and relative, higher scores could translate to unfavorable properties such as decreased Fc stability or even lower protein expression for each candidate design. From this in-silico analysis, the top-ranking disulfide was the same R292C and V302C mutation pair selected for stabilizing the CH2 domain, suggesting that this disulfide is a quality cross-linking solution as evidenced by the crystal structure and supported by the observed increase in CH2 melting temperature (Tm) up to ∼80°C (Jacobsen et al., 2017).
Mechanism of action for SEFL2.2 Fc
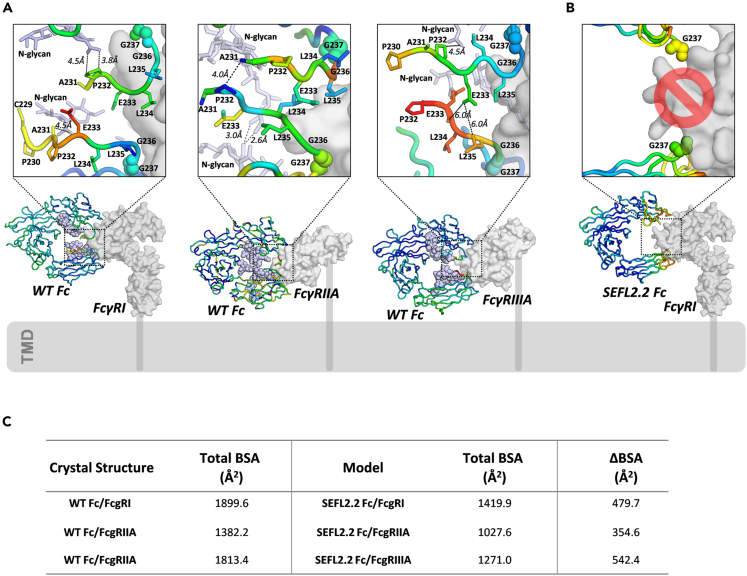
Although SEFL2.2 is an efficient strategy to reduce Fc binding to FcγRI, FcγRIIA and to FcgRIIIA receptors (Jacobsen et al., 2017; Liu et al., 2017), little is known about its mechanism of action. To unveil the molecular basis that mediates the loss of Fcγ receptor activity for SEFL2.2, we compared this structure with the co-crystal structures of 3 Fcγ receptors [FcγRI (PDB: 4X4M), FcγRIIA (PDB: 3RY6) and FcγRIIIA (PDB: 5XJE)], each determined in complex with WT Fc (Figure 2A) (Lu et al., 2015; Ramsland et al., 2011; Sakae et al., 2017). Focusing the structural analysis on the CH2/CH2 interface and the lower hinge regions that promote Fcγ interactions (Figure 2A), we observed that IgG1 residues from A231 to L235 in the lower hinge region all contribute to a network of protein-protein or protein-sugar interactions across the CH2/CH2 interface (Figure 2A). Notably, some of those interactions also take place across the sugar moieties from both N297 linked glycans (Man3NAG2) (Figure 2A), providing a rationale for the reduction of FcγR binding in the deglycosylated SEFL2.2 design. To provide further structural evidence, we superimposed the crystal structure of SEFL2.2 Fc with the crystal structure of WT Fc in complex with FcγRI (Figure 2B). Indeed, deletion of the CH2 glycan in the SEFL2.2 destabilized the CH2/CH2 interface and thus, in an allosteric manner, disrupted the A231-L235 motif (not visible in the crystal structure likely because of excessive flexibility). This potentially unstructured hinge is likely to interfere with FcγR binding rather than promote interface and complex formation (Figure 2B). This is in alignment with other effector less technologies focused on mutating P232E, L234A/L235A, or L235E residues (Chappel et al., 1991; Schlothauer et al., 2016; Xu et al., 2000) all located in the highly unstable A231-L235 motif as mapped on a WT Fc/FcγRI cartoon representation with relevant side chain mutations depicted in sticks (Figure S3). To better understand how much the SEFL2.2/FcγR interface was indeed reduced, we calculated the BSA of SEFL2.2 Fc modeled with 3 FcγRs. As shown in Figure 2C, SEFL2.2 resulted in a reduction of 479.7, 354.6, and 524,4 Å2 BSA for Fc complexes FcγRI, FcγRIIA and FcγRIIIA, respectively, most likely because of the structural rearrangements in the lower hinge.
Figure 2.
The mechanism for loss of Fcγ receptor binding by SEFL2.2 Fc
(A) Complex structures of WT IgG1 Fc binding to Fcγ receptors modeled onto the surface of transmembrane domain (TMD).
(B) Model of SEFL2.2 Fc interface with FcγRI using the WT Fc (not depicted) as a reference.
(C) The surface areas (Å2) buried on the FcγRI, FcγRII, and FcγRIII by WT Fc and SEFL2.2 Fc.
Molecular structure of the CPMv1 Fc
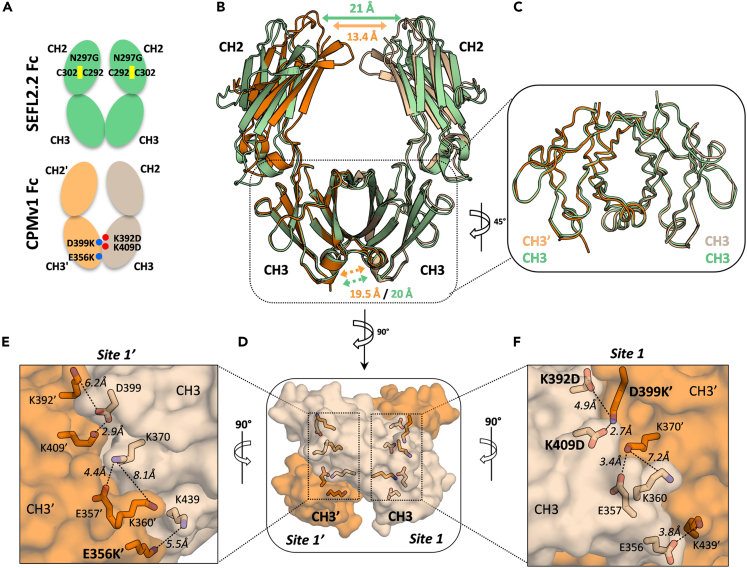
One strategy to enhance antibody Fc heterodimer formation is the use of engineered CPMs (Gunasekaran et al., 2010). To investigate whether the incorporation of CPMs in the CH3 domains could alter the CH3/CH3′ interface, we sought to determine the crystal structure of an early CPM version known as CPMv1 (Gunasekaran et al., 2010). This CPMv1 Fc contained two distinct chains carrying compatible CPMs (but not the SEFL2.2 technology described above), and was solved to 2.45 Å, with the asymmetric unit containing a single copy of the heterodimeric Fc (PDB: 7LUS; Figure S4A). As with the SEFL2.2 Fc structure described above, we found that CPMv1 did not trigger major changes in the Fc quaternary structure (Figure S4B). Furthermore, analysis of a similar CPM structure previously published also showed a good structural conservation upon the introduction of CPMs (Leaver-Fay et al., 2016). Of note, the distance between the CH2/CH2′ interface in the CPMv1 Fc structure is only ∼13.4 Å between the two N-terminal points, highlighting the influence deglycosylated SEFL2.2 technology has on these domains observed above (Figures 1B, 3A, and 3B).
Figure 3.
Structural analysis of CPMv1 Fc
(A) Schematic representation of SEFL2.2 Fc (PDB: 7LUR) and CPMv1 Fc (PDB: 7LUS).
(B) Superimposition of CPMv1 Fc (orange and wheat) with SEFL2.2 Fc (green), shown in cartoon representation. Distances between the CH2 N-termini were measured using the Cα of G237 of each chain. Distances between the CH3 C-termini were measured using Cα of L443 of each chain.
(C) Superimposition of CH3/CH3′ dimers.
(D–F) Surface representation of CH3/CH3′ dimer interface with key residues shown in sticks. Dotted lines represent the distances between side-chains.
In regards to the structural influence of the CPMs, the C-terminal points in the CH3 domains display a virtually identical distance to that seen in SEFL2.2 Fc (19.5 and 20 Å, respectively) and the CH3 domains from these 2 Fc structures are also very conserved (RMSD = 0.449 Å) (Figures 3B and 3C). Moreover, the total BSA between the two chains of CPMv1 Fc is 2291.1 Å2, which is greater by 255.4 Å2 than SEFL2.2 Fc (2035.7 Å2) and 93.2 Å2 greater than WT Fc (2197.9 Å2), potentially because of the CPMs engineered in the CH3/CH3′ interface. Further analysis revealed the structural mechanisms underpinning the CPMs in the heterodimeric CH3/CH3′ protein interface (Figure 3D). The engineered CPMs span two sites throughout the CH3 interface, with a single E356K′ mutation (designed to repulse K439 in CH3′) in site 1′ and a mutation ‘triad’ in site 1 (K392D and K409D in the CH3 and D399K′ in the CH3′) (Figures 3D–3F, S4C, and S4D). Interestingly, the triad engineered in site 1 shows a clear reposition of these side chains as result of CPMv1 design. Here, the side chains from D399K′ with K409D (2.7 Å) and D399K′ with K392D (4.9 Å), in site 1, move closer to their counterpart than observed for the native residues in site 1′, likely translating to a stronger binding energy (Figures 3E and 3F). Notably, in site 1′, the single E356K′ shows an opposite effect by repelling the native K439 (Figure 3E), suggesting this mutation does not favor heterodimerization and could be reengineered for further improvement. Altogether, the structural data strongly suggests that CPMv1 can be deployed without disrupting the CH3 folding and the CH3/CH3′ interface, and it seems to be compatible with the SEFL2.2 technology.
Screening for next generation of CPMs
To determine the optimal CPM design and to improve upon CPMv1 in the SEFL2.2 scaffold, we deployed a structure-guided approach to design a panel of 43 CPM variants across the CH3/CH3′ interface. These variants comprise 1 to 3 charge pairs that encompass two of the sites described in the CPMv1 design in addition to a third site described here (Figure 4A, Table S2). To facilitate the differentiation of correct paired species (heterodimer) from mispaired (homodimer), the 43 CPM variants and the SEFL2.2 (negative control) were tested on a monovalent single-chain variable fragment (scFv)-Fc format (Figure 4B).
Figure 4.
Screening of CPM variants in the CH3/CH3′ interface
(A) Diagram representation of the three positions (reference Edelman Unit) available for the design of CPMs.
(B) Schematic representation of format used for CPMs screening.
(C) Expression and analytical assessment of the 43 CPM variants and SEFL2.2 control.
(D) Trends of expression and analytical assessment per number of charge pairs.
All molecules were transiently expressed in 293-6E cells and purified by protein A for further analytics. Interestingly, all 43 CPM variants displayed expression levels lower than the SEFL2.2 control (Figure 4C, Table S2), suggesting that many of the CPMs explored here are unfavorable overall. Indeed, the loss in expression was even higher in those variants with charged mutations inserted in all three positions (1+ 2+3), whereas some combinations with mutations at only one (1) – v107 or two positions (1+3 and 1+ 2) – v1, v103, v104, and v118 were able to reach yields approaching that of the control SEFL2.2 homodimer.
To assess the ability of these new CPM variants in driving the HC/HC pairing, we determined the % main peak (MP) by micro capillary electrophoresis (MCE) of correct species (monovalent scFv-Fc) for these 43 variants and the SEFL2.2 control (Figures 4C and S5, Table S2). All variants showed a ∼ 2-fold increase overall in percent correct species as determined by % MP, a significant improvement compared to the 47.6% observed in the negative SEFL2.2 control. Here, variants carrying mutations in all three positions (1+2+3) showed high percentages of correct pairing in particular (with many >95%), highlighting that these mutations are efficient at preventing homodimer formation despite the lower expression levels.
To better compare these variants, we proceeded to correct the expression titer by the MP% determined by MCE (correct species = Titer × MP%) (Figure 4C). This analysis revealed that despite the lower expression levels, over 50% (24/43) of the engineered Fc variants resulted in greater heterodimerization formation compared to SEFL2.2 control (Figure 4C). Of note, variants CPMv103 and CPMv104 were top candidates. These two designs in particular are variants of CPMv1, but introduce either K439D or K439E to form a productive interaction with E356K in the heterodimer (unlike the repulsive interaction observed with E356K in CPMv1) (Table S2 and Figure 3E).
Although these 43 CPM variants have incorporated a variety of charged residues (K, E and D), we observed correlations between the number of charge pairs and protein expression and correct pairing. In this case, while an increase in the mutations selected here appear to correlate with a decrease in protein expression, the designs with more mutations tend to enhance the desired repulsion/attraction mechanism (Figure 4D). Thus, the optimal variants appear to strike a balance between expression and correct pairing, often consisting of two charge-pairs (Figure 4D).
CPMs are pH sensitive
As charge pair mutations operate based on electrostatic principles, we evaluated top CPM designs at different pHs to further distinguish design limitations. To this end we selected 6 CPM variants: CPMv1 and CPMv103 (high performing variants with mutations at positions (1 + 3)); CPMv106 and CPMv131 (high performing variants with modifications at positions (1 + 2)); CPMv112 and CPMv126 (variants with high % correct pairing with modifications at positions (1 + 2+3)). To avoid the complexity from HC/LC pairing, we deployed an altered SEFL2.2 IgG format (DEVD-IgG) mimicking the quaternary structure of a Hetero-IgG format (Figure 5A). In addition to CPMs in Fc region, DEVD-IgG carries a DEVD (a protease cleavage site) insertion between CH1 domain and the hinge region of CPM (−) HC to increase the molecular weight (MW) differentiation of correct and mispaired species in liquid chromatography–mass spectrometry (LC-MS) analytics. As expected, the expression levels for CPMv1 and CPMv103 variants are also very similar in this format (Figure 5B), although the performance of CPMv112 and CPMv126 was somewhat improved (Figures 4C and 5B). When using CEX to purify these molecules, the profile was mostly similar across this panel with the elution of a single main peak at a similar percentage of NaCl gradient buffer at pH 5.6 (Figure 5C). However, when the purification buffer was changed from pH 5.6 down to 5.0, CPMv106, CPMv131, CPMv112, and CPMv126 displayed multiple peaks (up to 3) with the third peak consisting of protein aggregate, eluting when the salt in the buffer was raised to 1 M NaCl (Figure 5C). In contrast, CPMv1 and CPMv103 retained a single CEX peak with the desired species and similar retention time, suggesting that these two variants are more resistant to pH variation. In some cases, a single mutation was seen to affect the CEX profile. For example, the addition of K360E to CPMv106 to create CPMv131 displayed an improved CEX profile at pH 5.0 in which the third peak of protein aggregate was eliminated. These results demonstrate that although K360E has not been recognized as playing a direct role across the CH3/CH3′ interface (Figures 3E and 3F), it may improve the CH3 electrostatic surface and create an environment where the entire molecule is less susceptible to acidic conditions. This decrease in aggregate species was also observed between CPMv112 and CPMv126, two variants closely related to CPMv126 but also carrying the K360E.
Figure 5.
Purification profiles for selected CPM variants
(A) Schematic representation of format used for CPM screening with the CPMs represented in red and blue and a DEVD site inserted into the upper hinge of one HC.
(B) Expression levels of CPM variants after harvest.
(C) CEX profiles for CPM variants assessed with buffers at pH 5.0 and pH 5.6. Arrows highlight main peaks.
CPM variants show different stability profiles
In addition to the pH sensitivity observed above, we also explored key stability metrics common to drug development (Jain et al., 2017). Thus, we sought to characterize the aggregation propensity, viscosity, and Tm for the CPM variants described above in an IgG1 SEFL2.2 scaffold (CPMv1, CPMv103, CPMv106, CPMv131, CPMv112, and CPMv126).
To investigate the aggregation propensity, we conducted stress tests of 2- and 4-week incubation at 40°C, 4-week at 25°C, and pH jump, followed by size exclusion chromatography (SEC) to separate the correct species (MP) and aggregates (high molecular weight, HMW) (Figures 6A and 6B). After a 2-week incubation at 40°C, a common protocol used to stress biological molecules, all six CPM variants maintained over 95% correct species as determined by SEC %MP (Figure 6A and Table S3). Although the percentage of low molecular weight (LMW) species was similar among all CPMs (Table S3), there was a minor, yet noticeable, increase in HMW species at 2-week, and 4-week samples (Figure 6B). After 4 weeks, the HMW percentage ranged from 0.6% to 1.1%, with CPMv1 having the highest level of HMW species, whereas CPMv103, CPMv131, and CPMv126 displayed the lowest levels at ∼0.6%. Interestingly, in the pH-jump stability experiment (consisting of a shift from pH 5.2 to pH 7 to simulate injection conditions), CPMv1 also showed a higher percentage of HMW relative to the other 5 variants (Figure 6B and Table S3).
Figure 6.
Stability assessment of selected CPM variants.
(A and B) Determination of protein species by analytical SEC after stress tests.
(C) DSC thermogram for CPM variants. On top, schematic representation of Tm1 and Tm2 as result of SEFL2.2 and CPM engineering.
Viscosity is another important measurement often used to guide the development of biologics aimed at subcutaneous delivery at high concentration (Sharma et al., 2014). In this experiment, the six purified CPM variants at 150 mg/mL were measured with a cone and plate viscometer. Results showed that all CPM variants were within an acceptable viscosity range, with CPMv1 having the highest viscosity at 8.2 cP and other variants ranging from 6.6 cP to 7.4 cP (Table S3).
Tm is a good measure for the overall folding stability of the molecule and thus of its long-term stability (Jain et al., 2017). With the combination of SEFL2.2 and CPM, the transition order of CH2 and CH3 was reversed relative to unmodified Fc domains. The SEFL2.2 disulfide bond increases the Tm of CH2 to ∼80°C (Jacobsen et al., 2017), whereas the CPMs decrease the Tm of CH3 (Figure 6C). Here, purified protein samples were measured by differential scanning calorimetry (DSC). Each of the variants showed two melting peaks with the Tm1 associated with the variation in unfolding of CH3 domain that contains CPMs and the Tm2 associated with the conserved disulfide-stabilized SEFL2.2 CH2 and the fragment antigen-binding region (Fab) (Figure 6C, Table S3) (Jacobsen et al., 2017). Interestingly, the Tm of CPMv1 CH3 domain (71.3°C) was well below the reported Tm for IgG1 CH3 domains (∼82°C) (Garber and Demarest, 2007), highlighting how these mutations are less favorable than WT residues even in the heterodimer (Gunasekaran et al., 2010; Jacobsen et al., 2017; Leaver-Fay et al., 2016). However, some of the improved designs were able to overcome this unfavorability, with CPMv103 (77.8°C) approaching that of the WT CH3 Tm. Thus, CPMv103 represents an ideal engineering solution for HC-HC pairing, with high expression levels, efficient pairing, and a good stability profile.
To further isolate the unfolding process of the engineered CH3 domains, we performed differential hydrogen deuterium exchange mass spec (HDX-MS). For these experiments, we explored the variants CPMv1 and CPMv103 in a SEFL2.2 hetero-Fc format (Figure 7A) so as to avoid any possible contributions/noise from the Fab region. As expected, there were no differences in deuterium incorporation observed in the CH2 domain between these two CPMs (Figure S6). In contrast, compared to CPMv1, two peptides in the CH3 domain (349–364 and 433–444) of CPMv103 were found to exhibit less deuterium incorporation (Figures 7B and 7C), indicating that CH3 domain of CPMv103 is more stable and less solvent accessible. Upon closer inspection, the peptide 349–364 (YTLPPSRKEMTKNQVS) contains the E356K mutation in both CPM variants, while the peptide 433–444 contains the native K439 in CPMv1 (HNHYTQKSLSLS) and the K439D mutation in CPMv103 (HNHYTQDSLSLS). This suggests that K439D mutation in CPMv103 likely establishes a productive interaction with the positively charged E356K mutation bringing the CH3/CH3′ interface tighter together and consequently more resistant to deuterium incorporation. Thus, the HDX-MS data pinpoints that the increased stability for CPMv103 comes from the enhanced interactions by a newly formed charge pair in the CH3 domain. Moreover, the HDX-MS data also showed that the CPM engineering in the CH3 domain does not impact the local conformation of the SEFL2.2 CH2 domain.
Figure 7.
Characterization of CPMv1 and v103 by HDX-MS
(A) Schematic representation of SEFL2.2 Fc with CPMv1 and CPMv103.
(B) Structure of CPMv1 CH3/CH3′ interface with selected residues shown in spheres.
(C) HDX plots showing the level of deuterium incorporation in both CPMv1 and CPMv103 variants.
Deployment of CPM is impacted by the Fv region
Although the surrogate formats used above to develop the Fc CPMs allowed for thorough characterization, they lack any therapeutic benefit or bispecific functionality. Thus, we took our top performing CPM design, v103, and evaluated it in 4 Hetero-IgG SEFL2.2 molecules, consisting of Fv regions A through H (Figure 8 and Table S4). As each Hetero-IgG contained two distinct Fv regions, this allowed us to determine the impact of sequence diversity. Although we could clearly demonstrate efficient pairing in our surrogate molecules, the introduction of additional sequences could skew this relationship (such as overexpression or underexpression of one polypeptide chain). Moreover, to also understand the impact of the CPM distribution across the CH3/CH3′ interface on protein expression and the percentage of species of interests versus mispaired species, we have assessed 2 different scenarios: i) the swapping of the binary positive/negative distribution and ii) where the charge distribution from the CPMv103 in the CH3/CH3′ interface is mixed (D399K and K439D on HC_1 (in blue); K409D, K392D, and E356K on HC_2 (in green)), creating a balanced charge distribution (BCD), followed by consequent swapping. Surprisingly, the expression data (captured by the Protein A column) showed that in case of scenario i) one orientation, negatively charged mutations K409D K392D K439D on the HC_1 (in blue) and positively charged mutations D399K E356K on the HC_2 (in green), increased the expression of 3/4 molecules up to ∼2 fold (Figure 8A). This random deployment for v103 also appeared to impact protein folding and impurities. Indeed, Hetero-IgG A×B went from displaying over 13.3% of ½-Ab (∼75 kDa) to near 0% upon CPMs swapping (Figure 8B) as determined by the analytical SEC. This ∼75 kDa impurity is likely the result of an over expression of one HC over the other and consequent inability to form homodimers because of the repulsive charges generated by these CPMs. In contrast, C×D, E×F and G×H molecules all saw a rise of this impurity (Figure 8B). This appears to suggest a relationship between the protein sequence in the complementarity-determining regions (CDRs) and the 5 charged mutations deployed in the CH3/CH3′ dimer. In striking contrast, the swapping in scenario ii, the swapping of CPMv103BCD did not impact the expression of this 4-molecule panel with the median values almost identical (Figure 8A). Regarding the ½-Ab species, and apart from the A×B molecule in CPMv103BCD swap, the BCD design appeared to greatly reduce this ∼75 kDa impurity, which will translate to higher final recovery of the desired species (Figure 8B).
Figure 8.
Rational deployment of CPMv103 in Hetero-IgG molecules
(A) Protein A yields of CPMv103 and CPMv103BCD each in both versions, swapping and not-swapping, applied to 4 Hetero-IgG molecules.
(B) Characterization of impurity levels (½-Ab species) measured by SEC and LC-MS. The ½-Ab species with over 3% detected (dash line) were characterized by LC-MS and highlighted with schematics.
Discussion
The ability to drive new types of protein assembly has the potential to create a promising generation of biologic therapeutics. Antibody-like proteins composed of distinct polypeptide chains can enable the binding of 2 or more therapeutic targets. However, the understanding of the structure/protein expression relationship has been insufficient, limiting the development of multispecific Abs.
In this study, we sought to unveil the molecular basis of two key technologies, SEFL2.2 and CPMv1, and whether each of them can be implemented simultaneously without observing detrimental effects. Moreover, we pursued the development of the next generation of CPMs by identifying the CPMv103 while shedding light toward building a rational deployment of CPMs.
The crystal structure of SEFL2.2 Fc at 1.95 Å enabled us to understand the molecular basis for the loss of function toward Fcγ receptor binding. The simple removal of the N297-linked glycan in the CH2 region induces an opening of the CH2/CH2′ interface critical for Fcγ receptor binding. This elegant approach could, however, reduce the overall thermostability of the IgG1-SEFL2.2 molecule. To mitigate any potential biophysical liability, we deployed a structure-guided design coupled with Rosetta aimed at strengthening the CH2 domain stability by inserting a non-canonical disulfide bond. This rationally guided and comprehensive design confirmed that the R292C and V302C mutations initially reported (Jacobsen et al., 2017), form indeed a strong covalent bond rescuing the thermostability profile.
The crystal structure of CPMv1 at 2.45 Å not only confirmed the rationale behind the concept of attraction/repulsion applied to 2 polypeptide chains mediated by CPMs (Leaver-Fay et al., 2016), but most importantly, it informed the design of further improved CPMs. This extensive effort yielded several CPM designs with improved characteristics over the initial CPMv1 such as protein expression, purification profiles and biophysical properties, all key attributes for a therapeutic molecule. From those, we have selected the CPMv103 which shows similar expression levels and purification to the CPMv1 while showing a remarkable increase in thermostability because of the insertion of the K439D mutation. This mutation, absent in the CPMv1, may enable productive contacts with the E356K′ residue in the opposite CH3 domain (Figure 3E) and therefore, overall strengthening the CH3/CH3′ interface in the bispecific heterodimer.
To create fast workflows, the deployment of technologies like CPMs is often executed as a rigid platform, where building blocks (i.e., Abs) are forced into those engineered scaffolds. However, the disregard of the possible impact of the sequence diversity often leads to suboptimal results (Gong et al., 2021), prompting us to investigate the relationship between CPMs and the sequence diversity present in the CDRs. Although it remains unclear the mechanism by which the CPMs in the CH3/CH3′ interface is affected by the Fv, the deployment of such charged mutations can have a detrimental effect on the expression of the desired heterodimer. However, by balancing the charge distribution of those 5 charged residues in the CPMv103 across the HC/HC interface, we have succeeded in minimizing the impact of sequence diversity. Therefore, the CPMv103BCD design shows more promise when deployed as a technology platform.
The continuous perfection of protein scaffolds coupled with the rational deployment of those same tools will enable us to predict success and ultimately, control the outcome of the development of biologics as therapeutics.
Limitations of the study
Among the numerous published approaches to knocking out effector function and driving bispecific IgG heavy chain pairing, our studies were limited to building upon our prior art of SEFL2.2 aglycosylated IgG CH2 domains and electrostatic steering with modifications to naturally charged residues at the CH3′ CH3 interface.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Chemicals, peptides, and recombinant proteins | ||
| Esp3I (BsmBI) | Thermo Scientific | Cat# FD0454 |
| T4 DNA ligase | Thermo Scientific | Cat# EL0011 |
| QIAGEN Plasmid Maxi Kit | QIAGEN | Cat# 12165 |
| PEImax | Polysciences | Cat# 24765-2 |
| FreeStyle F-17 medium | Gibco | Cat# A1383502 |
| Tryptone-N1 | Organotechnie | Cat# 19553 |
| Glucose | Thermo Fisher | Cat# A2494001 |
| Sodium valproate | MP Biomedicals | Cat# 0215206480 |
| Opti-MEM medium | Thermo Fisher | Cat# 11058021 |
| Lipofectamine LTX | Thermo Fisher | Cat# 15338500 |
| FabRICATOR enzyme | Genovis AB | Cat# A0-FR1-250 |
| Deposited data | ||
| SEFL2.2 Fc | This paper | PDB:7LUR |
| CPMv1 Fc | This paper | PDB:7LUS |
| Experimental models: Cell lines | ||
| HEK 293-6E | National Research Council of Canada | N/A |
| CHO-K1 | Amgen | N/A |
| Recombinant DNA | ||
| pTT5 | National Research Council of Canada | N/A |
| Synthesized DNA fragments | Twist Bioscience | N/A |
| Software and algorithms | ||
| XDS | Kabsch, 2010 | https://xds.mr.mpg.de/ |
| CCP4 program suite | Emsley et al., 2010; Winn et al., 2011 | http://www.ccp4.ac.uk/ |
| Scalepack | Otwinowski and Minor, 1997 | https://hkl-xray.com |
| Phaser | Adams et al., 2010 | http://phenix-online.org |
| COOT | Emsley et al., 2010 | http://www2.mrc-lmb.cam.ac.uk |
| Qt-PISA | Krissinel and Henrick, 2007 | https://www.ebi.ac.uk |
| PyMOL molecular graphics system | Schrodinger | https://pymol.org/2/ |
| RosettaScripts | Fleishman et al., 2011 | https://www.rosettacommons.org/software |
| Chromeleon software | Thermofisher | https://www.thermofisher.com/ |
| MassAnalyzer | Zhang et al., 2012 | https://pubs.acs.org/doi/10.1021/ac901193n |
| Origin software v7.0 | OriginLab | https://www.originlab.com/ |
| Graphpad Prism v6.02 | GraphPad Software | https://www.graphpad.com/ |
| Others | ||
| Protein A MabSelect SuRe column | Cytiva | Cat# 11003493 |
| HiTrap SP HP CEX column | GE Life Sciences | Cat# GE29-0513-24 |
| HiTrap MabSelect SuRE column | GE Life Sciences | Cat# GE11-0034-93 |
| HiTrap Desalting column | GE Life Sciences | Cat# GE17-1408-01 |
| HiLoad Superdex 200pg | Cytiva | Cat# 28989335 |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Fernando Garces (fgarces@amgen.com).
Materials availability
There are restrictions to the availability of plasmids, cell lines, and proteins generated in this study due to the lack of an external centralized repository for its distribution and our need to maintain the stock.
Experimental model and subject details
Cell lines
The human embryonic kidney 293-6E (HEK 293-6E) suspension cell line, an immortalized cell line derived from an aborted female fetus, was initially developed by the National Research Council of Canada (Durocher et al., 2002). Cells were cultured in FreeStyle F-17 medium (ThermoFisher) at 37°C and 5% CO2, on a shake platform set to 120 RPM.
The CHO-K1 suspension cell line, an epithelial cell line derived from the ovary of the Chinese hamster, was cultured in growth medium containing 50% of EX-CELL 302 Serum-Free Medium (Sigma), 50% of Amgen in-house developed CHO growth medium, and 2mM L-glutamine (Gibco). Cells were incubated at 37°C and 5% CO2, on a shake platform set to 120 RPM.
Method details
Crystallography and model design
SEFL2.2 Fc (initial concentration 7.75 mg/mL) was mixed with Mini-Z domain of Protein A at a 1:2 molar ratio and incubated at 4°C for 1 hour. Complex crystals were grown in 0.2 M ammonium tartrate, 20% w/v PEG3350 and 10% NDSB-221 detergent using sitting drop vapor diffusion at 20°C. Crystals were frozen with Mitegen LV Cryo Oil. SEFL2.2 Fc diffraction data were collected at the Advanced Light Source, Berkeley, California, Beamline 5.0.2 at a temperature of 100 K and a wavelength of 1.0 Å. Data sets were processed with XDS (Kabsch, 2010) and scaled with the CCP4 program suite (Winn et al., 2011). The structure was solved to 1.95 Å via molecular replacement using Molrep (Emsley et al., 2010) with an in-house effector functionless Fc crystal structure and the structure of Mini-Z from the structure of a G0 version of Fc bound to a minimized version of Protein A (PDB:1OQO).
CPMv1 Fc crystals were grown in 12.5% w/v PEG550MME, 12.5% w/v PEG20K, 0.1 M Tris-HCl (pH 8.5), 20% 2,3-butanediol, 3% ethanol and frozen directly from the crystal drop. CPMv1 Fc diffraction data were collected at the Advanced Photon Source, Lemont, Illinois, Beamline SER-CAT 22-ID at a temperature of 100 K and a wavelength of 1.0 Å. Data sets were processed with HKL2000 and scaled with Scalepack (Otwinowski and Minor, 1997). The structure was solved to 2.45 Å via molecular replacement using Phaser (Adams et al., 2010) and a crystal structure of a Fc-fragment of human IgG2 antibody (PDB:4HAF).
For both SEFL2.2 Fc and CPMv1 Fc structures, iterative refinement cycles were performed with the CCP4 program suite (Winn et al., 2011) and COOT (Emsley et al., 2010). Buried surface area was calculated using Qt-PISA (Krissinel and Henrick, 2007). Figure illustrations were created using PyMOL molecular graphics system, version 1.8 (Schrödinger).
Structural modeling was performed using RosettaScripts and the Talaris2014 score function (Fleishman et al., 2011). Using the “fastrelax” protocol, five cycles of backbone minimization and rotamer optimization brought the template structures to a local energy minimum. For each potential disulfide, Cysteines were computationally introduced for both residues and forced into a disulfide. This was followed by 50 Monte Carlo based simulated annealing steps for the peptide backbone and surrounding residues. The final models were ranked relative to each other using the Talaris2014 score and the unweighted disulfide potential term.
Molecular biology
DNA fragments of variable and constant domains of intended molecules (HC, LC, scFv-Fc) were synthesized separately by Twist Bioscience, and then cloned into a mammalian transient expression vector using a Golden Gate Assembly method (Engler et al., 2008). HCs were constructed in SEFL2.2 IgG1 with desired CPMs (Table S2). In the case of Hetero-Fc constructs, Fc sequences with CPMs were separately cloned into two different mammalian expression vectors carrying a Puromycin or Hygromycin resistant gene. After sequence confirmation, transfection-grade DNA was prepared using a Maxi plasmid purification kits (Qiagen, catalog # 12165). Plasmids were mixed at a mass based ratio of 1:1 for Hetero-Fc and monovalent scFv-Fc molecules, 1:1:1:1 (LC1:HC1:LC2:HC2) for classic Hetero-IgGs and 2:1:1 (LC:HC1:HC2) for DEVD-IgGs.
Mammalian expression
All monovalent scFv-Fc, DEVD-IgG and Hetero-IgG molecules were expressed using a transient HEK 293-6E expression system, initially developed by the National Research Council of Canada (Durocher et al., 2002). For monovalent scFv-Fc and DEVD-IgG molecules, a slightly modified transfection process was used (Estes et al., 2015). Briefly, after cells reaching 1.5×106 viable cells/mL, transfections were done by mixing 0.5 μg DNA with 2.0 μL PEImax (Polysciences, catalog # 24765-2) in 100 μL FreeStyle F-17 medium (Gibco, catalog # A1383502) and incubating for 10 min before complex was added to 900 μL of cell culture. In the case of the Hetero-IgG molecules, Sodium Valproate was also added after transfection (Backliwal et al., 2008; Jäger et al., 2013). In this case, cells were grown to 2×106 viable cells/mL for transfection. Transfections were done by mixing 0.5 μg DNA with 1.5 μL PEImax in 100 μL FreeStyle F-17 medium for 10 min before complex was added to 900 μL of cell culture. Twenty-four hours after transfection, fresh media was added to double the volume of the culture with the addition of Tryptone-N1 (Organotechnie, catalog # 19553) and glucose (Thermo Fisher, catalog # A2494001) to achieve final concentrations of 2.5 g/L and 4.5 g/L, respectively. Four days after transfection, sodium valproate (MP Biomedicals, catalog # 0215206480) was added to a final concentration of 3.75 mM. Six days after transfection, conditioned medium was harvested by centrifugation followed by vacuum filtration through a 0.22 μm filter.
In the case of Hetero-Fc constructs, expression was done using an internal stable CHO-K1 process with the incorporation of transposase technology. For transfection, cells were centrifuged and resuspended at 2×106 viable cells/mL in Opti-MEM medium (Thermo Fisher, catalog # 11058021). Transfections were done by mixing 2 μg Hetero-Fc constructs DNA with 2 μg transposase DNA and 10 μL Lipofectamine LTX (Thermo Fisher, catalog #15338500) in 1 mL of Opti-MEM, incubating for 15-20 minutes, and adding complex to 1 mL of resuspended cells. Cultures were incubated at 37°C and 5% CO2, on a shake platform set to 120 RPM. Five hours after transfection, 2 mL of our internally developed growth media was added. Seventy-two hours after transfection, cells were resuspended in growth media with the addition of Puromycin at 10 μg/mL and Hygromycin at 600 μg/mL and passaged by dilution until growth and viability reached pre-transfection levels. Cultures were inoculated for production by media exchange at 1.5×106 cells/mL in an internally developed production media. Seven days after inoculation conditioned medium was harvested.
Protein purification
Purification of CPMv1 Fc and SEFL2.2 Fc constructs was performed for crystallography. CPMv1 Fc IgG was purified as previously described (Jacobsen et al., 2017). Briefly, IgGs were recovered from the clarified CHO-K1 cell condition media using a two-columns purification process. First, proteins were affinity captured using MabSelect SuRe (Cytiva, catalog # 11003493), followed by a wash with PBS buffer and elution using 100 mM sodium acetate pH 3.6. The eluted samples were then equilibrated to pH 5.0, loaded onto an SP-HP column (GE Life Sciences, catalog # GE29-0513-24), followed by washing with 20mM acetic acid, pH 5.0 and elution with 0-600 mM NaCl gradient pH 5.0. Peak fractions were pooled and further dialyzed into sodium acetate buffer (10 mM acetic acid, 9% sucrose, pH 5.2). CPMv1 Fc was subsequently cleaved with FabRICATOR enzyme (Genovis AB, catalog # A0-FR1-250). The Fc was re-purified on a Protein A MabSelect SuRe column (Cytiva, catalog # 11003493), followed by elution with 100 mM sodium acetate pH 3.5. The pH was adjusted to pH 5.0 using Trizma base, then purified by SEC in 25 mM Hepes, 150 mM NaCl, pH7.5. In case of SEFL2.2 Fc, 1.6 L of conditioned medium was loaded onto a Protein A MabSelect SuRe column and washed with TBS buffer (25 mM Tris HCl pH 7.4 and 100 mM NaCl), then eluted with 25 mM Tris, 500 mM arginine, pH 7.5. The sample was loaded onto a HiTrap SP HP CEX column (GE Life Sciences, catalog # GE29-0513-24), washed with 50 mM sodium acetate, pH 5.0 and eluted with 30 CV 0-400 mM NaCl gradient in sodium acetate, pH 6.0. The pooled peak fractions were buffer exchanged with 25 mM Hepes, 150 mM NaCl, pH7.5.
Purification of monovalent scFv-Fc molecules was performed utilizing an ÄKTA purifier connected to an autosampler and using a tandem 2-dimensional purification method as previously described (Yoo et al., 2014). Briefly, 35 mL of conditioned medium was loaded onto a 1 mL HiTrap MabSelect SuRE column (GE Life Sciences, catalog # GE11-0034-93), washed with TBS buffer and eluted with 2 CV of 100 mM sodium acetate, pH 3.6. The eluate was directly loaded onto a 5 mL HiTrap Desalting column (GE Life Sciences, catalog # GE17-1408-01) and buffer exchanged into formulation buffer of 10 mM KPO4, 75 mM Lysine, 4% Trehalose, pH 6.0.
To purify DEVD-IgG molecules, 5 L of conditioned medium were loaded onto 200 mL MabSelect SuRe column and washed with PBS. The bound protein was eluted with 0.1M sodium acetate, pH 3.6. Three separate aliquots of ∼150 mg of purified Protein A sample were further purified by SEC using HiLoad Superdex 200pg (Cytiva, catalog # 28989335) equilibrated in 50mM MES, 100mM NaCl, pH 6.0. Small-scale CEX was performed to determine optimal buffer conditions for the molecules at two different pH. 2 mg of SEC pooled samples were diluted up to 5 mL in 50 mM sodium acetate at either pH5.0 or pH 5.6. Each sample was loaded onto a 1mL HiTrap SP HP CEX column and eluted with 30 CV 0-400 mM NaCl gradient in each pH buffer. Peak fractions were analyzed by MCE, UPLC and LC-MS for final samples. Large-scale CEX was next performed on the SEC protein samples at the desired buffer pH 5.6. The SEC pool was loaded onto a 10 mL SP HP column equilibrated in 50 mM sodium acetate, pH 5.6 and eluted with 30 CV 0-400 mM NaCl gradient. Peak fractions were pooled and dialyzed in 10 mM sodium acetate, 9% sucrose, pH5.2.
The Hetero-IgG CPM molecules were purified over Protein A affinity resin, followed by SP HP CEX as described in DEVD-IgG protocol; however, omitting the SEC column.
For all molecules the final protein samples were analyzed for purity by MCE using a Caliper instrument (Perkin Elmer), and by UPLC (Waters) with a BEH C-200, 4.6×150 mm column in 100 mM sodium phosphate, 50 mM NaCl, 7.5 % ethanol, pH 6.9. LC-MS was also performed to determine the correct mass of each molecule.
Stability characterization by SEC
For the stability studies, molecules were concentrated to 70 mg/mL at pH 5.2 before incubation at 25°C or 40°C. Samples were harvested at either 2 weeks or 4 weeks time point and analyzed by SEC. For the pH-jump study, a 20× solution of PBS pH 7.0 was added to the samples at 5% of the final volume. The samples are either analyzed without incubation (T0) or after incubation at 40°C for 24 hours.
For the SEC analysis, ∼70 μg of each sample was injected into an Acquity UPLC (Waters) equipped with a BEH200, 4.6×300 mm column. The mobile phase was 100 mM sodium phosphate and 500 mM NaCl at pH 6.8 at a flow rate of 0.3 mL/min. Data was analyzed with Chromeleon software (Thermo fisher).
Stability characterization by DSC
DSC was performed using a MicroCal VP-capillary differential scanning calorimeter (Malvern Panalytical). A scanning range of 20°C to 100°C with a scan rate of 60°C/h was used. All samples were diluted to approximately 1 mg/mL before measurements. Baseline correction was performed by using cubic function in Origin software (v7.0). Gaussian fits were performed using non-two state cursor imitation. The best fit judged by chi-squre values was obtained after evaluating multiple iterations. Final fit was plotted in Excel.
Stability characterization by viscosity
Viscosity was measured using 80 μL of each sample (concentrated to 150 mg/mL and added PS80 to 0.01%) on a cone and plate rheometer (AR-G2, TA Instruments). The rheometer was equipped with a 20 mm anodized aluminum cone geometry with a cone angle of 1.998°. Reported results represent the viscosity at a shear rate of 1000 s-1 at 23°C.
Stability characterization by HDX-MS
HDX-MS experiments were performed to pinpoint differences in protein structure between CPMv1 and CPMv103, as previously described (Zhang et al., 2010, 2018). The H/D exchange reaction was initiated by diluting protein samples with 10 mM acetate in D2O (pD 5.2) as indicated for a predetermined time at 37°C. The exchange reaction was quenched and the protein was denatured by mixing the solution in a 1:1 ratio with ice-cold 200 mM sodium phosphate, 4 M guanidine-HCl and 0.5 M tris(2-carboxyethyl)phosphine (pH 2.4). Digested peptides were separated by a reversed-phase HPLC and further analyzed by an Orbitrap mass spectrometer (Elite, Thermo Fisher). All data were processed with the software MassAnalyzer (Zhang et al., 2012) for peptide identification and deuterium level calculation. All HDX-MS data were normalized to a 100% deuterium concentration and the percent deuterium incorporation was plotted against labeling time on a log scale with Graphpad Prism v6.02 (Graphpad software).
Acknowledgments
The authors are grateful to Mark Daris, Ning Sun, Alison Castellano, Michelle Wu, Vivian Bi, and Noi Nuanmanee for assistance in cloning, expression and purification of scFv-Fc, DEVD-IgG and Hetero-IgG molecules; to Hannah Catterall, Deniz Temel and Victoria Jann for assistance in measuring Tm by DSC, viscosity by cone and plate and stability by SEC. This study was funded by Amgen Inc.
Author contributions
Project design by B.E., F.G. and Z.W.; Crystallography and structural analysis by A.S. and D.W.; Protein cloning, expression and purification by D.G., B.E., V.L., C.M. and D.L.; Protein characterization by S.S.; HDX-MS by J.Z.; Manuscript written or edited by B.E., A.S., D.G., T.P.R., F.G., and Z.W.
Declaration of interests
The authors declare no competing interests.
Published: December 17, 2021
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2021.103447.
Contributor Information
Fernando Garces, Email: fgarces@amgen.com.
Zhulun Wang, Email: zwang.t@gmail.com.
Supplemental information
Data and code availability
Atomic coordinates and structural factors for SEFL2.2 Fc and CPMv1 Fc have been deposited with the Protein Data Bank under accession codes of 7LUR and 7LUS, respectively. This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
- Adams P.D., Afonine P.V., Bunkóczi G., Chen V.B., Davis I.W., Echols N., Headd J.J., Hung L.W., Kapral G.J., Grosse-Kunstleve R.W., et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atwell S., Ridgway J.B., Wells J.A., Carter P. Stable heterodimers from remodeling the domain interface of a homodimer using a phage display library. J. Mol. Biol. 1997;270:26–35. doi: 10.1006/jmbi.1997.1116. [DOI] [PubMed] [Google Scholar]
- Backliwal G., Hildinger M., Kuettel I., Delegrange F., Hacker D.L., Wurm F.M. Valproic acid: A viable alternative to sodium butyrate for enhancing protein expression in mammalian cell cultures. Biotechnol. Bioeng. 2008;101:182–189. doi: 10.1002/bit.21882. [DOI] [PubMed] [Google Scholar]
- Braisted A.C., Wells J.A. Minimizing a binding domain from protein A. Proc. Natl. Acad. Sci. U S A. 1996;93:5688–5692. doi: 10.1073/pnas.93.12.5688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brinkmann U., Kontermann R.E. The making of bispecific antibodies. MAbs. 2017;9:182–212. doi: 10.1080/19420862.2016.1268307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carter P. Bispecific human IgG by design. J. Immunol. Methods. 2001;248:7–15. doi: 10.1016/s0022-1759(00)00339-2. [DOI] [PubMed] [Google Scholar]
- Chappel M.S., Isenman D.E., Everett M., Xu Y.Y., Dorrington K.J., Klein M.H. Identification of the Fc gamma receptor class I binding site in human IgG through the use of recombinant IgG1/IgG2 hybrid and point-mutated antibodies. Proc. Natl. Acad. Sci. U S A. 1991;88:9036–9040. doi: 10.1073/pnas.88.20.9036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis J.H., Aperlo C., Li Y., Kurosawa E., Lan Y., Lo K.M., Huston J.S. SEEDbodies: Fusion proteins based on strand-exchange engineered domain (SEED) CH3 heterodimers in an Fc analogue platform for asymmetric binders or immunofusions and bispecific antibodies. Protein Eng. Des. Sel. 2010;23:195–202. doi: 10.1093/protein/gzp094. [DOI] [PubMed] [Google Scholar]
- Durocher Y., Perret S., Kamen A. High-level and high-throughput recombinant protein production by transient transfection of suspension-growing human 293-EBNA1 cells. Nucleic Acids Res. 2002;30:E9. doi: 10.1093/nar/30.2.e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emsley P., Lohkamp B., Scott W.G., Cowtan K. Features and development of coot. Acta Crystallogr. D Biol. Crystallogr. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engler C., Kandzia R., Marillonnet S. A one pot, one step, precision cloning method with high throughput capability. PLoS One. 2008;3:e3647. doi: 10.1371/journal.pone.0003647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Estes B., Hsu Y.R., Tam L.T., Sheng J., Stevens J., Haldankar R. Uncovering methods for the prevention of protein aggregation and improvement of product quality in a transient expression system. Biotechnol. Prog. 2015;31:258–267. doi: 10.1002/btpr.2021. [DOI] [PubMed] [Google Scholar]
- Fleishman S.J., Leaver-Fay A., Corn J.E., Strauch E.M., Khare S.D., Koga N., Ashworth J., Murphy P., Richter F., Lemmon G., et al. RosettaScripts: A scripting language interface to the Rosetta macromolecular modeling suite. PLoS One. 2011;6:e20161. doi: 10.1371/journal.pone.0020161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garber E., Demarest S.J. A broad range of Fab stabilities within a host of therapeutic IgGs. Biochem. Biophys. Res. Commun. 2007;355:751–757. doi: 10.1016/j.bbrc.2007.02.042. [DOI] [PubMed] [Google Scholar]
- Gong D., Riley T.P., Bzymek K.P., Correia A.R., Li D., Spahr C., Robinson J.H., Case R.B., Wang Z., Garces F. Rational selection of building blocks for the assembly of bispecific antibodies. MAbs. 2021;13:1870058. doi: 10.1080/19420862.2020.1870058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gunasekaran K., Pentony M., Shen M., Garrett L., Forte C., Woodward A., Ng S.B., Born T., Retter M., Manchulenko K., et al. Enhancing antibody Fc heterodimer formation through electrostatic steering effects: Applications to bispecific molecules and monovalent IgG. J. Biol. Chem. 2010;285:19637–19646. doi: 10.1074/jbc.M110.117382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobsen F.W., Stevenson R., Li C., Salimi-Moosavi H., Liu L., Wen J., Luo Q., Daris K., Buck L., Miller S., et al. Engineering an IgG scaffold lacking effector function with optimized developability. J. Biol. Chem. 2017;292:1865–1875. doi: 10.1074/jbc.M116.748525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jäger V., Büssow K., Wagner A., Weber S., Hust M., Frenzel A., Schirrmann T. High level transient production of recombinant antibodies and antibody fusion proteins in HEK293 cells. BMC Biotechnol. 2013;13:52. doi: 10.1186/1472-6750-13-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain T., Sun T., Durand S., Hall A., Houston N.R., Nett J.H., Sharkey B., Bobrowicz B., Caffry I., Yu Y., et al. Biophysical properties of the clinical-stage antibody landscape. Proc. Natl. Acad. Sci. U S A. 2017;114:944–949. doi: 10.1073/pnas.1616408114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kabsch W. XDS. Acta Crystallogr. D Biol. Crystallogr. 2010;66:125–132. doi: 10.1107/S0907444909047337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein C., Sustmann C., Thomas M., Stubenrauch K., Croasdale R., Schanzer J., Brinkmann U., Kettenberger H., Regula J.T., Schaefer W. Progress in overcoming the chain association issue in bispecific heterodimeric IgG antibodies. MAbs. 2012;4:653–663. doi: 10.4161/mabs.21379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krissinel E., Henrick K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 2007;372:774–797. doi: 10.1016/j.jmb.2007.05.022. [DOI] [PubMed] [Google Scholar]
- Labrijn A.F., Janmaat M.L., Reichert J.M., Parren P. Bispecific antibodies: A mechanistic review of the pipeline. Nat. Rev. Drug Discov. 2019;18:585–608. doi: 10.1038/s41573-019-0028-1. [DOI] [PubMed] [Google Scholar]
- Leaver-Fay A., Froning K.J., Atwell S., Aldaz H., Pustilnik A., Lu F., Huang F., Yuan R., Hassanali S., Chamberlain A.K., et al. Computationally designed bispecific antibodies using negative state repertoires. Structure. 2016;24:641–651. doi: 10.1016/j.str.2016.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L., Jacobsen F.W., Everds N., Zhuang Y., Yu Y.B., Li N., Clark D., Nguyen M.P., Fort M., Narayanan P., et al. Biological characterization of a stable effector functionless (SEFL) monoclonal antibody scaffold in vitro. J. Biol. Chem. 2017;292:1876–1883. doi: 10.1074/jbc.M116.748707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Z., Leng E.C., Gunasekaran K., Pentony M., Shen M., Howard M., Stoops J., Manchulenko K., Razinkov V., Liu H., et al. A novel antibody engineering strategy for making monovalent bispecific heterodimeric IgG antibodies by electrostatic steering mechanism. J. Biol. Chem. 2015;290:7535–7562. doi: 10.1074/jbc.M114.620260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu J., Chu J., Zou Z., Hamacher N.B., Rixon M.W., Sun P.D. Structure of FcgammaRI in complex with Fc reveals the importance of glycan recognition for high-affinity IgG binding. Proc. Natl. Acad. Sci. U S A. 2015;112:833–838. doi: 10.1073/pnas.1418812112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merchant A.M., Zhu Z., Yuan J.Q., Goddard A., Adams C.W., Presta L.G., Carter P. An efficient route to human bispecific IgG. Nat. Biotechnol. 1998;16:677–681. doi: 10.1038/nbt0798-677. [DOI] [PubMed] [Google Scholar]
- Moore G.L., Bautista C., Pong E., Nguyen D.H., Jacinto J., Eivazi A., Muchhal U.S., Karki S., Chu S.Y., Lazar G.A. A novel bispecific antibody format enables simultaneous bivalent and monovalent co-engagement of distinct target antigens. MAbs. 2011;3:546–557. doi: 10.4161/mabs.3.6.18123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otwinowski Z., Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- Ramsland P.A., Farrugia W., Bradford T.M., Sardjono C.T., Esparon S., Trist H.M., Powell M.S., Tan P.S., Cendron A.C., Wines B.D., et al. Structural basis for Fc gammaRIIa recognition of human IgG and formation of inflammatory signaling complexes. J. Immunol. 2011;187:3208–3217. doi: 10.4049/jimmunol.1101467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ridgway J.B., Presta L.G., Carter P. 'Knobs-into-holes' engineering of antibody CH3 domains for heavy chain heterodimerization. Protein Eng. 1996;9:617–621. doi: 10.1093/protein/9.7.617. [DOI] [PubMed] [Google Scholar]
- Sakae Y., Satoh T., Yagi H., Yanaka S., Yamaguchi T., Isoda Y., Iida S., Okamoto Y., Kato K. Conformational effects of N-glycan core fucosylation of immunoglobulin G Fc region on its interaction with Fcgamma receptor IIIa. Sci. Rep. 2017;7:13780. doi: 10.1038/s41598-017-13845-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlothauer T., Herter S., Koller C.F., Grau-Richards S., Steinhart V., Spick C., Kubbies M., Klein C., Umana P., Mossner E. Novel human IgG1 and IgG4 Fc-engineered antibodies with completely abolished immune effector functions. Protein Eng. Des. Sel. 2016;29:457–466. doi: 10.1093/protein/gzw040. [DOI] [PubMed] [Google Scholar]
- Sharma V.K., Patapoff T.W., Kabakoff B., Pai S., Hilario E., Zhang B., Li C., Borisov O., Kelley R.F., Chorny I., et al. In silico selection of therapeutic antibodies for development: Viscosity, clearance, and chemical stability. Proc. Natl. Acad. Sci. U S A. 2014;111:18601–18606. doi: 10.1073/pnas.1421779112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starovasnik M.A., Braisted A.C., Wells J.A. Structural mimicry of a native protein by a minimized binding domain. Proc. Natl. Acad. Sci. U S A. 1997;94:10080–10085. doi: 10.1073/pnas.94.19.10080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suresh M.R., Cuello A.C., Milstein C. Bispecific monoclonal antibodies from hybrid hybridomas. Methods Enzymol. 1986;121:210–228. doi: 10.1016/0076-6879(86)21019-8. [DOI] [PubMed] [Google Scholar]
- Von Kreudenstein T.S., Escobar-Carbrera E., Lario P.I., D'Angelo I., Brault K., Kelly J., Durocher Y., Baardsnes J., Woods R.J., Xie M.H., et al. Improving biophysical properties of a bispecific antibody scaffold to aid developability: Quality by molecular design. MAbs. 2013;5:646–654. doi: 10.4161/mabs.25632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Q., Chen Y., Park J., Liu X., Hu Y., Wang T., McFarland K., Betenbaugh M.J. Design and production of bispecific antibodies. Antibodies (Basel) 2019;8:43. doi: 10.3390/antib8030043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winn M.D., Ballard C.C., Cowtan K.D., Dodson E.J., Emsley P., Evans P.R., Keegan R.M., Krissinel E.B., Leslie A.G., McCoy A., et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 2011;67:235–242. doi: 10.1107/S0907444910045749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu D., Alegre M.L., Varga S.S., Rothermel A.L., Collins A.M., Pulito V.L., Hanna L.S., Dolan K.P., Parren P.W., Bluestone J.A., et al. In vitro characterization of five humanized OKT3 effector function variant antibodies. Cell Immunol. 2000;200:16–26. doi: 10.1006/cimm.2000.1617. [DOI] [PubMed] [Google Scholar]
- Yoo D., Provchy J., Park C., Schulz C., Walker K. Automated high-throughput protein purification using an AKTApurifier and a CETAC autosampler. J. Chromatogr. A. 2014;1344:23–30. doi: 10.1016/j.chroma.2014.04.014. [DOI] [PubMed] [Google Scholar]
- Zhang J., Chalmers M.J., Stayrook K.R., Burris L.L., Garcia-Ordonez R.D., Pascal B.D., Burris T.P., Dodge J.A., Griffin P.R. Hydrogen/deuterium exchange reveals distinct agonist/partial agonist receptor dynamics within vitamin D receptor/retinoid X receptor heterodimer. Structure. 2010;18:1332–1341. doi: 10.1016/j.str.2010.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J., Woods C., He F., Han M., Treuheit M.J., Volkin D.B. Structural changes and aggregation mechanisms of two different dimers of an IgG2 monoclonal antibody. Biochemistry. 2018;57:5466–5479. doi: 10.1021/acs.biochem.8b00575. [DOI] [PubMed] [Google Scholar]
- Zhang Z., Zhang A., Xiao G. Improved protein hydrogen/deuterium exchange mass spectrometry platform with fully automated data processing. Anal. Chem. 2012;84:4942–4949. doi: 10.1021/ac300535r. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Atomic coordinates and structural factors for SEFL2.2 Fc and CPMv1 Fc have been deposited with the Protein Data Bank under accession codes of 7LUR and 7LUS, respectively. This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.