Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-caused COVID-19 pandemic has transmitted to humans in practically all parts of the world, producing socio-economic turmoil. There is an urgent need for precise, fast, and affordable diagnostic testing to be widely available for detecting SARS-CoV-2 and its mutations in various phases of the disease. Early diagnosis with great precision has been achieved using real-time polymerase chain reaction (RT-PCR) and similar other molecular methods, but theseapproaches are costly and involve rigorous processes that are not easily obtainable. Conversely, immunoassays that detect a small number of antibodies have been employed for quick, low-cost tests, but their efficiency in diagnosing infected people has been restricted. The use of biosensors in the detection of SARS-CoV-2 is vital for the COVID-19 pandemic’s control. This review gives an overview of COVID-19 diagnostic approaches that are currently being developed as well as nanomaterial-based biosensor technologies, to aid future technological advancement and innovation. These approaches can be integrated into point-of-care (POC) devices to quickly identify a large number of infected patients and asymptomatic carriers. The ongoing research endeavors and developments in complementary technologies will play a significant role in curbing the spread of the COVID-19 pandemic and fill the knowledge gaps in current diagnostic accuracy and capacity.
Graphical abstract
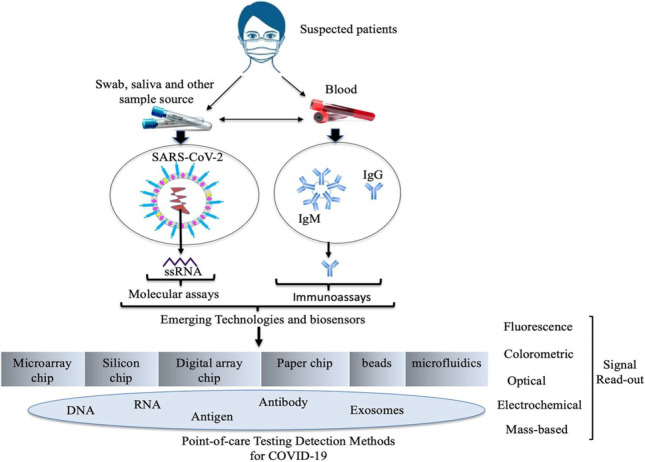
Keywords: SARS-CoV-2 infections, COVID-19, Molecularassays, Immunoassays, Biosensors, Detection
Introduction
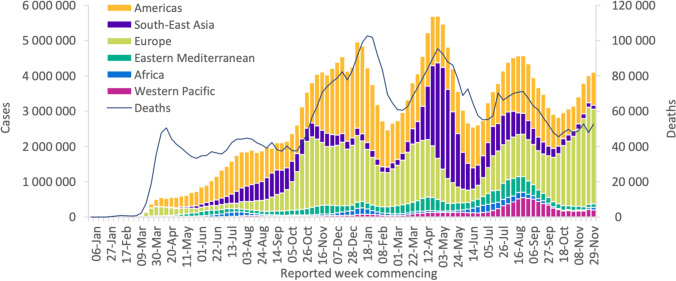
Chinese doctors first noticed a pneumonia disease caused by a new type of coronavirus in Wuhan city, China, in December 2019 [1]. The causative agent, later recognized as severe acute respiratory syndrome-related coronavirus (SARS-CoV-2), and the disease it causes is now designated as COVID-19 [2]. According to World Health Organization [3], more than 281 million COVID-19 cases have been registered worldwide since its occurrence till December 29, 2021, with more than 5.4 million deaths. The USA has the world’s top number of registered incidences and deaths. After the USA, India, Brazil, the UK, and Russia have the most cases. After the USA, the biggest numbers of deaths are in Brazil, India, Mexico, and Russia [Figure 1].
Fig. 1.
WHO reports weekly COVID-19 cases and deaths by region [4]
After the initial incidence of COVID-19, most countries were completely unprepared for its spread. Various forms of social separation and shutdowns in many locations have drastically affected the living standards of billions of people around the world. As a result of business closures and travel restrictions, the global economy has been significantly affected. Thus, many healthcare systems faced significant transmission before adequate testing was in place to allow isolation and tracking [4]. The ease of global access of precise and quick testing methodologies is critical for deciphering the complicated patterns of SARS-CoV-2 infection and immunity. To that purpose, academic institutions, research centers, and businesses globally have been working diligently to advance and manufacture vital diagnostic kits. Vaccines to prevent COVID-19 infection are currently available, and research is underway to develop therapeutics to treat and manage future epidemics of SARS-CoV-2 infection [5]. However, the difficulty of containing the COVID-19 pandemic has been exacerbated by the introduction of many SARS-CoV-2 variations, including B.1.1.7 (Alpha), B.1.351 (Beta), P1 (Gamma), and B.1.617.2 (Delta), and recently omicron SARS-CoV-2 (B.1.1.529), a new COVID-19 variant, which have enhanced spread and resistance to vaccines and treatments posing concerns to the society [6]. Therefore, diagnostics and biosecurity play a significant role in limiting the COVID-19 infection. Centralized laboratories provide specific and sensitive assays, for example, virus isolation, high-throughput polymerase chain reaction (PCR), and enzyme-linked immunosorbent assays (ELISAs); but they are expensive and subject to complex instrumentation that require well-trained operators. For COVID-19 asymptomatic infections, there are no detectable clinical manifestations, for example, pyrexia [7]. At this point, screening of individuals in crowded locations and in-time isolation of infected people cannot be performed with the utilization of infrared thermography. The reverse transcription polymerase chain reaction (RT-PCR) assay targeting the RNA of SARS-CoV-2 is presently the gold standard diagnostic approach for COVID-19 detection [8, 9]. Loop-mediated isothermal amplification (LAMP) techniques, for example, reverse transcription LAMP (RT-LAMP), are also increasing in popularity [10]. However, given the failure of many nations to implement extensive testing during the COVID-19 pandemic, further efforts and resource implementations in disease diagnostics research and innovative technology development are needed for rapid and detection of SARS-C0V-2 and its mutations [11]. Here, we review various molecular, immunological, and biosensor approaches for detection of SARS-CoV-2 infection with some commercially developed POC tests. These assays and tests will assist researchers to develop accurate, simple, fast, and inexpensive diagnostic techniques for monitoring of COVID-19. It is anticipated that this will allow researchers to track present and future infectious diseases, putting humanity in a better position to tackle disease outbreaks in the future.
The sources of SARS-CoV-2
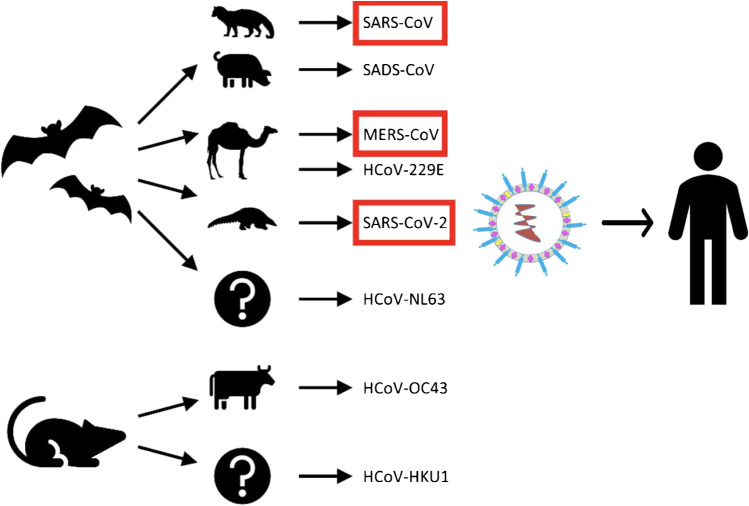
In the 1960s, human coronaviruses (HCoVs) isolated from upper respiratory infections were described as HCoV-229E and HCoV-OC43 [12]. Next, in the early 2000s, HCoV-NL63 and HCoV-HKU1 and in 2002, a SARS-CoV syndrome emerged in southern China [13]. A Middle East respiratory syndrome-related coronavirus (MERS-CoV) syndrome was observed in Saudi Arabia in 2012 [14], followed by COVID-19. The detail about emergence of coronaviruses was described by other authors [15, 16]. In the past, all human epidemics of coronaviruses and betacoronaviruses were caused by direct contact with bats, rodents, or mice and indirect transmission by human host such as birds and animals located adjacent to humans. Human coronaviruses have animal origins, as shown in Figure 2 COVID-19 source. It was rumored at first that SARS-CoV-2 originated as a result of human modification of an original coronavirus. However, to a considerable extent, our understanding of the animal source of SARS-CoV-2 is presently insufficient [17]. The virus’s reservoir carriers have yet to be identified. It is unclear whether SARS-CoV-2 was transmitted to humans via an intermediary host and, if so, which animals might have served as that intermediate host. The presence of coronavirus sequence strains RaTG13, RmYN02, and pangolin coronaviruses in animals suggests that a variety of coronaviruses comparable to SARS-CoV-2 are spreading [17]. Furthermore, whereas mutations have been implicated in the evolution of various sarbecoviruses, including SARS-CoV, it cannot be ruled out that viral RNA recombination among related coronaviruses played a role in the creation of SARS-CoV-2. However, according to the authors, it is not possible to confirm the origins of SARS-CoV-2 or refute the other theories at this point in time [18].
Fig. 2.
The origins of human coronaviruses in animals.
Adapted from Rabi et al., licensed CC BY 4.0 (2020) [15].
Targets for molecular detection
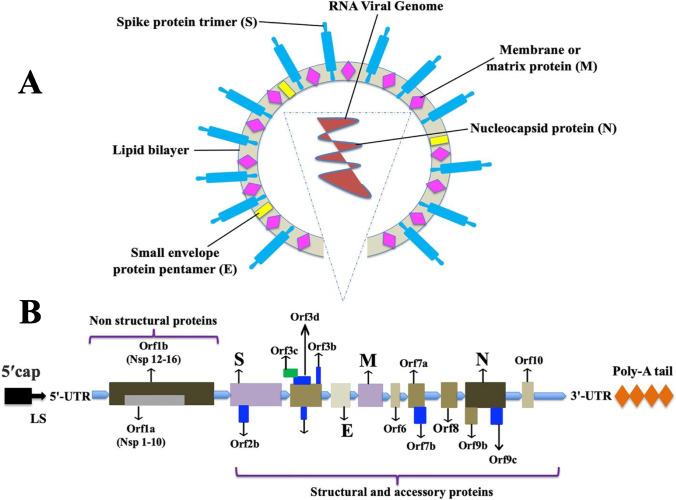
SARS-CoV-2 is a positive-sense, single-stranded RNA enveloped virus and the size of its genomic RNA is ~30 kb in length [19, 20]. The surface of virion is lipid bilayer hijacked from the host cell and the virus is mostly spherical in shape, approximately 50–200 nm in diameter [21]. The SARS-CoV-2 genome includes 5′ and 3′ untranslated regions (UTRs), six open reading frames (ORFs), including ORF1a/b, which encompasses 16 non-structural proteins (nsp) associated to the replication-transcription complex, four structural proteins, spike (S), envelope (E), membrane (M), and nucleocapsid (N), and several other non-structural, special structural, and/or ancillary ORFs (ORF3a/b, 6, 7a, 7b, 8, and 10) [22] [Figure 3 SARS-CoV-2 structure and genome]. The genes of both structural proteins and species-specific ancillary genes of non-structural proteins such as ORF1a and ORF1b, RNA-dependent RNA polymerase (RdRp), and hemagglutinin-esterase (HE) needed for viral replication are used as molecular targets. For instance, the CDC in the USA proposes two nucleocapsid protein targets (N1 and N2) [23] whereas the WHO advises an E gene assay and subsequently by a validation assay utilizing RdRp gene as a preferred diagnostic test [24]. On the other hand, China CDC targeted ORF1ab, N, and nCoC_IP2 for RNA detection. In addition different countries (Germany, France, Japan, Hong Kong, Thailand) recommended different ways of selecting targeted genes of interests for detecting SARS-CoV-2 (Table 1 Primer and probe [25–27]). A few of these molecular targets have been employed in nucleic acid diagnostics by researchers from various nations. Minimally, two molecular targets for SARS-CoV-2 diagnostics should be added to eliminate any cross-reaction with other common coronaviruses and possible variations in genes of SARS-CoV-2. Thus far, there has been no hint that any of the sequence areas included in clinical diagnostic test gives a distinct benefit. Nonetheless, to limit the consequences of changes in the gene sequences, the optimal concept would incorporate a minimal conserved area and one unique area with a positive and a negative control, particularly when the virus spreads among fresh regions.
Fig. 3.
The overall structure of SARS-CoV-2 is seen in this diagram. A The viral surface proteins such as spike protein (S), small envelope protein (E), and membrane protein (M) are embedded in a lipid bilayer envelope generated from the host cell. Inside the viral envelope is single-stranded positive-sense viral RNA coupled with the nucleocapsid protein (N) (above). B The RNA genome includes 5′ and 3′ untranslated regions (UTRs), a 5′ methylated cap (ME), and a 3′ poly-A tail. The genes that code for non-structural proteins (Nsp) and spike (S), membrane (M), envelope (E), and nucleocapsid (NC) proteins are shown in the diagram (below).
Table 1.
| Country | Target gene(s) | Forward primer 5′–3′ | Reverse primer 5′–3′ | Probe 5′–3′ | Amplicon size | References |
|---|---|---|---|---|---|---|
| USA (CDC) | N1 | GACCCCAAAATCAGCGAAAT | TCTGGTTACTGCCAGTT GAATCTG | FAM-ACCCCGCATTACGTTTGGTG GACC-BHQ1 | 71 bp | [25] |
| N2 | TTACAAACATTGGCCG CAAA | GCGCGACATTCCGAAGAA | FAM-ACAATTTGCCCCCAGCGCTT CAG-BHQ1 | 67 bp | [25] | |
|
N3 (Removed on 03/15/20) |
GGGAGCCTTGAATACACCAAAA | TGTAGCACGATTGCAGCATTG | FAM-AYCACATTGGCACCCGCAAT CCTG-BHQ1 | 72 bp | [25] | |
| China (CDC) | N | GGGGAACTTCTCCTGCTAGAAT | CAGACATTTTGCTCTCAAGCTG | FAM-TTGCTGCTGCTTGACAGATT -TAMRA | 99 bp | [25, 26, 27] |
| Orf1ab | CCCTGTGGGTTTTACA CTTAA | ACGATTGTGCATCAGCT GA | FAM-CCGTCTGCGGTATGTGGAA AGGTTATGG-BHQ1 | 119 bp | [25, 27] | |
| RdRp/nCoV_IP2 | ATGAGCTTAGTCCTGT TG | CTCCCTTTGTTGTGTTG T | Hex-AGATGTCTTGTGCTGCCGG TA-BHQ1 | 108 bp | [25, 26] | |
| E | TTCTTGCTTTCGTGGTATTC | CACGTTAACAATATTGCAGC | FAM-GTTACACTAGCCATCCTTACTGCGCTTCGA-BHQ1 | - | [26] | |
| France (Institut Pasteur) | RdRp gene /nCoV_IP4 | GGTAACTGGTATGATT TCG | CTGGTCAAGGTTAATATAGG | FAM-TCATACAAACCACGCCAGG-BHQ1 | 107 bp | [25] |
| E gene/E_Sa rbeco | ACAGGTACGTTAATAGTTAATAGCGT | ATATTGCAGCAGTACGCACACA | FAMACACTAGCCATCCTTACTGCGCTTCG-BHQ1 | 125 bp | [25] | |
| RdRp gene /nCoV_IP2 | ATGAGCTTAGTCCTGTTG | CTCCCTTTGTTGTGTTG T | Hex- AGATGTCTTGTGCTGCCGG TA-BHQ1 | 108 | [25] | |
| Hong Kong | N | TAATCAGACAAGGAACTGATTA | CGAAGGTGTGACTTCCATG | FAM-GCAAATTGTGCAATTTGCGG-TAMRA | 110 | [43] |
| Orf1bnsp14 | TGGGGYTTTACRGGTAACCT | AACRCGCTTAACAAAGCACTC | FAM-TAGTTGTGATGCWATCATG ACTAG-TAMRA | 132 bp | [43] | |
| Japan (NIID) | N | AAATTTTGGGGACCAGGAAC | TGGCAGCTGTGTAGGTCAAC | FAM-ATGTCGCGCATTGGCATGGA-BHQ | 155 bp | [43] |
| Thailand | N | CGTTTGGTGGACCCTCAGAT | CCCCACTGCGTTCTCCATT | FAM-CAACTGGCAGTAACCA-BQH1 | 57 bp | [43] |
| Germany (Charité) | RdRp | GTGARATGGTCATGTGTGGCGG | CARATGTTAAASACACTATTAGCATA |
P1: FAM-CCAGGTGGWACRTCATCMGGTGATGC-BBQ, P2: FAM-CAGGTGGAACCTCATCAGGAGATGC-BBQ |
100 bp | [43, 45-] |
| E | ACAGGTACGTTAATAGTTAATAGCGT | ATATTGCAGCAGTACGCACACA | FAM-ACACTAGCCATCCTTACTGC GCTTCG-BBQ | 113 bp | [43] | |
| N | CACATTGGCACCCGCAATC | GAGGAACGAGAAGAGGCTTG | FAM-ACTTCCTCAAGGAACAACATTGCCA-BBQ | 128 bp | [43] |
Targets for protein detection
The process of coronavirus infection and assembly is critical for selecting the most appropriate detection methods and laboratory testing. The structural S protein performs a crucial role in virus assembly, fusion, invasion, and replication [28]. For viral replication, the M and E proteins are required. The M protein is in charge of nutrition transmembrane transport and envelope construction, whereas the N and E proteins, as well as a number of auxiliary proteins, impede the host immune response or serve other unknown activities. The S and N proteins are among the most efficient antigen markers for COVID-19 disease diagnostics [16]. Furthermore, resemblances between SARS-CoV, MERS-CoV, and SARS-CoV-2 allow for the potential of use or modification of current diagnostic approaches and effective therapeutic for COVID-19. SARS-CoV-2, conversely, has a varied gene location and the potential for further diversity in the genomic region owing to the virus’s pandemic-range transmission [16].
The S gene determines the S proteins, which are split into two functional subunits: aminoterminal (S1) and carboxyterminal (S2) [21]. At the S1-S2 barrier, there is a beneficial furin cleavage that can promote infection of the host cell. The S1 domain comprises the six-amino-acid spike protein receptor binding domain (RBD), while the S2 domain is responsible for fusion [21]. Human angiotensin-converting enzyme 2 (ACE2) is present on respiratory, renal, and gastrointestinal cells, and SARS-CoV and SARS-CoV-2 bind to it for the onset of infection [21]. This ACE2, the SARS-CoV-2 receptor, can be matched with commercially available antibodies to detect SARS-CoV-2 S1 protein [29]. Furthermore, Serine 477 (S477) in the S protein has been the most often exchanged amino acid residue in the RBDs of SARS-CoV-2 mutants. As a result, S477 and its two most common mutations (S477G and S477N) are important to investigate and detection [30].
Prior investigations on numerous SARS-CoV-2 genome sequences showed about 4% genomic mutation of the entire 220 strains tested [31], implying that distinct strains overlap, which could pose a difficult role for many diagnostic approaches. The coronavirus exhibits S protein immunogens to detect the COVID-19 causal virus [32]. The immune system’s production of immune globulins improves the detection method. As a result, immune globulins are critical for detecting COVID-19 and maybe treating it [33]. Antibody detection is based on the proteins immunoglobulin M (IgM) and (IgG) and (IgA) [34]. Apart from antibodies, there might be other proteins and cellular markers in infected patients’ respiratory, blood, and fecal samples [35] that might be confused with other illnesses. Therefore, various approaches must be employed to detect the impacts of SARS-CoV-2, initially to establish the diagnosis, followed by detecting the viral nucleic acid, particular viral proteins, or virions of the SARS-CoV-2 virus [16]. In-depth understanding of the morphology and biochemical properties of SARS-CoV-2 infection will facilitate the development of novel diagnostics for detection of COVID-19.
Diagnosis of SARS-CoV-2
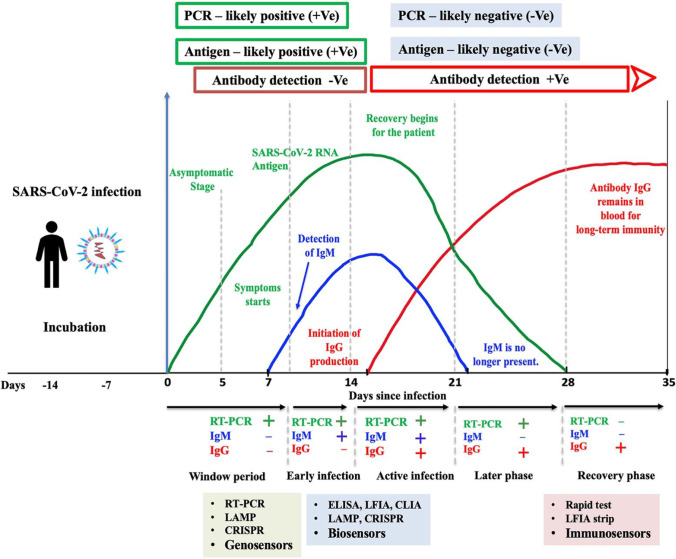
SARS-CoV-2 presents a distinctive biological feature that poses a number of obstacles to world healthcare structures and has resulted in a weak reaction to the infectious disease’s containment. Numerous innovative and improved diagnostic techniques have been produced making them accessible at a range of settings due to the research and advancement attempts of clinical laboratories and university researchers. SARS-CoV-2 can now be detected using a number of different diagnostic procedures [36]. Despite the fact that these approaches are robust and sensitive, their sensitivities have been modified by the selected diagnostic method, burden of the virus, and person-specific immune reaction. Because of the virus’s extended incubation time, one of the most difficult aspects of diagnosing COVID-19 is the necessity to use several detection strategies. It is important to distinguish between certain time points for SARS-CoV-2 detection, performing genetic components any time point after infection, and antibodies must be detected using serological methods at distinct time points after infection. Along the course of illness, Figure 4 depicts a graphical time frame regarding the suitable diagnostic approaches.
Fig. 4.
The most studied detection approaches along the trajectory of infection for SARS-CoV-2. The figure depicts the dynamic range of SARS-CoV-2 infectivity, viral RNA, and host immunoglobulins (IgM and IgG), as well as COVID-19, time kinetics. Upon hitting a measurable level in blood, antibodies undergo seroconversion. Note: The amounts of each antibody shown in this chart are for illustration purposes only and do not represent actual values. This is an exemplary design, and it should be noted that there are variations in the literature, particularly for the slope tails. We chose to preserve the excellence and quality until more data was gathered and a consensus is reached on the time courses.
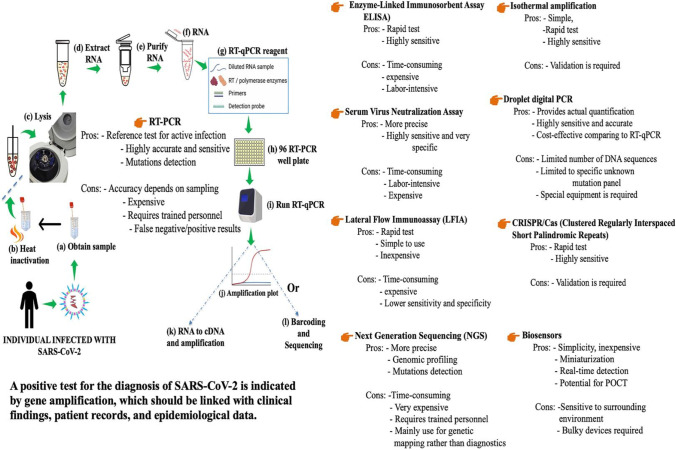
SARS-CoV-2 has been detected using a range of analytical approaches [36]. Figure 5 summarizes some key diagnostic approaches for detecting SARS-CoV-2. Nucleic acid is quantified using molecular methods and genosensors, and infected patients’ immunoresponse is quantified through protein detection (antigen and antibody) using immunoassays and immunosensors [36]. Immunoassays are price-efficient, sensitive, fast, and selective. However, they need time-consuming washing processes that are difficult to automate [37]. Contrasting to immunological tests, genetic content-related approaches usually display higher sensitivity and selectivity [37]. Nucleic acids detection, on the other hand, frequently necessitates rigorous procedures and technically talented workers [37], making consequently bulk testing in a pandemic event impractical. As a result, issues must be resolved in both methods of detection in order to develop an efficient diagnosis, as this will be described more in this review study.
Fig. 5.
An overview of the most important detection approaches with their major characteristics
Molecular detection methods
Nucleic acid detection methods have developed rapidly and become a revolutionary technology for molecular diagnostics. The identification of RNA from the SARS-CoV-2 virus, the pandemic’s causative infectious agent, is used to provide a molecular diagnosis of COVID-19. The initial genome sequence of SARS-CoV-2 was determined with metagenomic RNA sequencing, an impartial and high-throughput technique for sequencing numerous genomes [38, 39]. Subsequently, the sequence was added to the GenBank sequence repository portal that allows scientists to design primers and probes for nucleic acid-based SARS-CoV-2 diagnostics [38].
Specimen collection handling and storage
Improper specimen collection, storage, and transport are likely to produce false test results for COVID-19. For SARS-CoV-2, the Centers for Disease Control and Prevention (CDC, USA) suggests testing lower respiratory tract specimens, if accessible (https://www.cdc.gov/coronavirus/2019-ncov/hcp/infection-control-recommendations.html). In addition, sputum, oropharyngeal, or nasopharyngeal swab/wash can be collected using sets that are available commercially that varies from one supplier to the next [40]. Nasopharyngeal specimens may miss early infection; a deeper specimen may need to be obtained by bronchoscopy [41]. These procedures require legitimate biosafety precautions, well-trained staff, and may not be accessible in many parts of the world. Saliva and serum are potential sources of biomarkers for testing, monitoring, and prevention control in SARS-CoV-2 infections [42, 43]. However, only 15% infected individuals hospitalized with pneumonia showed detectable RNA in serum. Personal protective equipment, such as N95 masks or face masks, eye protection, gloves, and a gown are recommended during specimen collection. Specimens should be stored at 2–8 °C for up to 72 h or at −70 °C or below for delayed use (https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html).
Extraction and purification
Performance of the COVID-19 molecular diagnostic assays is dependent upon the amount and quality of template RNA purified from specimens. Several commercial vendors offer RNA extraction kits and procedures that have been qualified and validated for recovery and purity of RNA for use [44].
Standard PCR assay
The PCR test for COVID-19 is a molecular one that looks for genetic information of SARS-CoV-2, the virus that causes COVID-19, in the upper respiratory tract. Researchers employ PCR technology to convert small amounts of RNA from specimens into DNA, which is then repeated until SARS-CoV-2 is detected if it exists. Due to its enormous versatility, high sensitivity, and high sequence specificity, the PCR assay has become a routine practice for identifying coronaviruses [45]. Usually, coronavirus RNA is converted into cDNA by reverse transcription. Next, PCR is performed followed by amplicon analysis through conventional detection methods or instruments such as gel electrophoresis and sequencing methods for detecting coronaviruses [46]. Since it was approved for use in February 2020, the PCR test has been the reliable standard for diagnosing COVID-19 (https://www.eurofinsus.com/safer-work/resources/blogs/blog-what-makes-covid-19-pcr-testing-the-gold-standard/). However, these methods are tedious and expensive to use in clinical settings.
Real-time PCR (qPCR) as a reference test
Nucleic acid-based real-time reverse transcriptase-PCR (RT-PCR) assay is advantageous for early diagnosis of coronavirus infection [47]. In general, the assay design process requires two principal steps: (a) sequence alignment, primer, and probe design, and (b) assay optimization and testing. In January 2020, the China CDC disclosed publicly the genetic sequence of the SARS-CoV-2 [38]. This empowered countries to design primers for RT-PCR tests that have become the reference test for the diagnosis of COVID-19. A number of SARS-related viral genome sequences were aligned and analyzed to design a set of primers and probes [48]. The RT-PCR assay can be structured as a two-target framework, where one universal primer set can detect various coronaviruses including SARS-CoV-2 and a second primer set only recognizes SARS-CoV-2. One of the existing problems of the qRT-PCR process is the use of a fluorescent tag attaching to the primary signals generated by amplified DNA. This system not only adds to the equipment’s cost but also adds to its interpretations. Developing countries and outside laboratory settings with little resources are less interested in this technology. The efficacy of commercialized RT-PCR kits has not been rigorously controlled. For clinical sample analysis, the presence of safe and stable outside positive controls could be avoided in real-time RT-PCR assays [49]. For now, the quickly mutating nature of SARS-CoV-2 necessitates an accurate detection of genetically diverse coronaviruses [50]. Multiplex real-time RT-PCR assays can be used to detect coronavirus accurately with great sensitivity avoiding the risk of false-negative results [47]. For assay optimization, controls need to be carefully chosen to guarantee the quality of the test and to distinguish experimental errors. Real-time RT-PCR cycle threshold (Ct) values represent the number of amplification cycles required for the target gene to exceed a threshold level [51]. The Ct values are inversely related to viral load and can provide an indirect method for measuring the copy number of viral RNA in the sample; however, the utilization of Ct values as an intermediary of viral load is impacted by the test itself and components inside the sample matrix that can influence amplification competency [52]. It has recently been proposed that the viral load of SARS-CoV-2 might be a significant factor in deciding both disease intensity and probability of transmission [53, 54]. There are some constraints to RT-qPCR, for instance, sample quality must be adequate, and qualified personnel are necessary. Furthermore, in the primary phases of the disease, RT-PCR tests may miss detecting SARS-CoV-2. False negatives have been reported in participants for up to 2 weeks, according to some publications [55]. The variability in viral infectivity, and the volume of virus in patient populations, could be the cause. Differences can arise as a result of the disease’s progression. It is also likely that the sample site is not appropriate, the virus-containing specimens are not collected, or that the detection procedure is not consistent. These flaws restrict the utility of RT-PCR tests in a variety of situations. Despite these drawbacks, RT-PCR is still the gold standard for verifying COVID-19 diagnosis. Since the outbreak of the pandemic, numerous efforts have been made to produce handheld PCR devices [56, 57]. CovidNudge, a lab-on-a-chip device, can conduct sample preparation and real-time RT-PCR in remote locations. The Cove Nudge test, which is now being used in the UK, costs roughly 10 pounds for single test (equal to USD 13.80), which is almost ten times less than the average cost of a traditional RT-PCR test in the country (around 100 pounds). The potential issues presented by existing RT-PCR devices can be minimized with the refining and optimization of PCR chip and biosensor construction. In another study, microchip-based RT-qPCR was developed for detection of SARS-CoV-2 from nasopharyngeal swab RNA extracts and the limit of detection was 1 viral copy per reaction [58]. The microchip kit reduces reaction volumes by tenfold, resulting in decreased reagent use and shorter assay times while meeting the gold standard in sensitivity of relatively high procedures. However, the lack of acceptable high-quality DNA extraction procedures that can allow users to avoid the need for laboratory equipment is an obstacle in field deployable real-time PCR systems. These constraints, taken together, hinder microchip PCR methods from becoming a realistic portable system of choice for COVID-19 diagnosis in the field.
Other nucleic acid-based diagnostic techniques for SARS-CoV-2
Digital PCR
The digital PCR (dPCR) method can be used to quantify a minimal amount of viral load, track the virus in the environment, and assess the efficacy of anti-SARS-CoV-2 medications [59] and detect viral mutations [60]. Digital PCR empowers the total measurement of target nucleic acids. Digital droplet PCR (ddPCR) divides the sample into several droplets, each of which acts as a separate PCR reactor, and includes a couple of no target sequences [61]. Amplification-positive segments are measured and quantified after PCR. ddPCR is the most commonly utilized partitioning process, accessible to the commercial units. The partitioning process includes microwell plates, capillaries, oil emulsion, and miniaturized channels [62]. SARS-CoV-2 virus loads can be quantified more accurately and precisely using ddPCR [63]. The sensitivity of ddPCR (around 10−2 copy/mL) is greater than that of traditional PCR, allowing it to detect a very low amount of virus. For instance, when comparing pharyngeal swab specimens from COVID-19 recovering patients, ddPCR identified viral RNA (Chinese CDC sequences) in 9 of 14 (64.2%) RT-PCR negative specimens [63]. Vasudevan and colleagues developed ddPCR to quantify SARS-CoV-2 viral load from crude lysate without nucleic acid purification. The limit of detection was 10 copies per reaction [64]. The viral loads of samples taken from various parts of the same patient were also contrasted: the load was maximum in pharyngeal specimens, and lowest in fecal samples and in serum samples [65].
The main benefit of employing dPCR is its high sensitivity and high-throughput screening, both of which are essential for COVID-19 detection. However, there are a few issues that must be addressed before dPCR can be employed in regular diagnostics. The dPCR, like other PCR testing, necessitates the use of costly equipment, reagents, and expert knowledge to run the process. The manufacture of dPCR chips is a time-consuming process with many phases. Furthermore, like with other POC testing, certain regulations and procedures must be followed to ensure the quality of dPCR system results.
Isothermal amplification
Nucleic acid-based assays using isothermal amplification are being adapted as an alternative to instrumentation-based laboratory-dependent assays for POC SARS-CoV-2 detection [66]. Isothermal amplification methods use a single temperature and do not require thermal cycling. These techniques include LAMP [67], recombinase polymerase amplification (RPA) [68], transcription-mediated amplification (TMA) or nucleic acid sequence-based amplification (NASBA) [69, 70], and rolling circle amplification (RCA) [71] and have been used for pathogens detection. The RT-NASBA assay was developed for detection of SARS-CoV [72] and the sensitivity was equivalent to real-time RT-PCR assay. Recently, Jones and colleagues developed an RT-LAMP assay for SARS-CoV-2 and SARS-CoV-2 variant RNA amplification detection directly from nasopharyngeal swab samples taken from patients. The advantage of this method was there was no requirement to isolate the RNA from the specimen [73]. The RCA assay was used to detect SARS-CoV both in liquid and solid phases [74]. The assay was rapid, more sensitive, and inexpensive than conventional PCR assay. These isothermal amplification assays could be integrated with inexpensive portable devices for detection of COVID-19. These technologies run at a constant temperature throughout the operation, avoiding the need for thermocyclers. These approaches also allow for real-time amplification measurements, and amplicons can be recognized by monitoring fluorescence or inspecting for color change by naked eye. Gel electrophoresis is no longer necessary because of this capacity. Because of their minimal power needs and ease, isothermal amplification technologies could be more utilized than standard PCR. This enables them to be incorporated into easy, inexpensive portable platforms. Isothermal amplification techniques range in terms of total quantity of primers and enzymes employed, the temperature at which the amplification takes place, and the sorts of templates used. Table 2 [75–87] summarizes the several isothermal amplification platforms established for detecting SARS-CoV-2 in both laboratory and clinical samples.
Table 2.
Summary of the several isothermal amplification platforms established for SARS-CoV-2 diagnostics in both laboratory and clinical samples [Refs. 75-87]
| Procedure | Samples | Target gene region of the primers | Analytical sensitivity (LOD) | Pretreatment or need for RNA extraction from the sample | Validation with clinical samples | Ref. |
|---|---|---|---|---|---|---|
|
Triplex RT-LAMP Detection: visual colorimetric coupled to RT-qPCR for fluorescence monitoring |
Clinical samples from human saliva | ORF1ab, S, and ORF7a, N2 | 250 copies/reaction | NO | Yes | [75] |
|
One-tube RT-LAMP Detection: visual colorimetric coupled to RT-qPCR for fluorescence monitoring with SYBR Green |
Clinical samples from human nasopharyngeal swabs | ORF1ab | 200 copies/mL | Yes | Yes | [76] |
|
RT-LAMP coupled with a disposable cartridge Detection: colorimetric |
Clinical samples from human oropharyngeal and nasopharyngeal swabs | N, E | 300 copies/reaction | Yes | Yes | [77] |
|
RT-LAMP coupled with an aluminum block embedded with a cartridge heater Detection: fluorescence |
SARS-CoV-2/Australia/QLD02/2020 strain grown in the laboratory | RdRp | 100 copies/reaction | Yes | No | [78] |
|
RT-LAMP on microfluidic chips Detection: real-time monitoring fluorescence using smartphone camera |
Clinical samples and synthetic nasal fluid spiked with virus | ORF1a, S, ORF8, and N | 50 copies/μL | No | Yes | [79] |
|
Saliva-Dry-RT-LAMP incorporated with Biobox Detection: visual fluorometric coupled to RT-qPCR platform |
Clinical samples from human saliva and nasopharyngeal swabs | S spike, RdRp | 1 copy/μL | Yes | Yes | [80] |
|
RT-LAMP platform coupled with CRISPR-Cas12-based DETECTR with lateral flow strip Detection: visual fluorescence coupled to RT-qPCR platform |
Clinical samples from human nasopharyngeal and oropharyngeal swabs | N, E | 500 copies/μL | Yes | Yes | [81] |
|
RT-LAMP CRISPR on electric-field microfluidic chips Detection: fluorescence |
Contrived and clinical samples from human nasopharyngeal swabs | N, E | 10 copies/μL | NO | Yes | [82] |
|
RT-LAMP on paper microfluidic chips (portable COVIDISC) Detection: fluorescence produced by an intercalating dye (SYTO-82) coupled to qRT-PCR |
Clinical samples from human nasopharyngeal swabs | RdRp | 1 genome copies/μL | NO | Yes | [83] |
|
One-tube sandwich RCA electrochemical biosensor Detection: pulse voltammetry coupled to validate in RT-qPCR platform |
Clinical samples from human nasopharyngeal swabs | N, S | 1 copy/μL | Yes | Yes | [84] |
|
RCA on multi-microelectrode array microchips with electrochemical biosensor Detection: pulse voltammetry |
Plasmids containing SARS-CoV-2 template | N and RdRp | 0.972 fg/μL (RdRp gene) and 3.925 fg/μL (N gene) | NO | NO | [85] |
|
RT-RPA- and CRISPR-based one-tube OR-DETECTR technology integrated with lateral flow strip Detection: fluorescence |
-Artificial samples -Human pharyngeal samples |
N and RdRp | 2.5 copies/μL | Yes | Yes | [86] |
|
NASBA-based PHANTOM technology Detection: color, luminescence |
-Human nasopharyngeal samples | ORF1ab (Nsp13) | 100 copies/reaction | Yes | Yes | [87] |
CRISPR/Cas system
There are other nucleic acid-based tests that could be utilized for SARS-CoV-2 detection. The CRISPR/Cas method is an efficient genome editing platform that was recently applied to pathogen detection [88, 89]. The CRISPR-associated enzyme Cas13a has recently been adapted for developing SHERLOCK detection technology for RNA sensing [90]. Virus-derived RNA targets are reversely transcribed to complementary DNA (cDNA) followed by isothermal amplification in which the amplicons are transcribed back into RNA. Next, the CRISPR/Cas13a complex containing a designed guide RNA binds with the isothermally amplified RNA product and Cas13a is activated [91]. Cas13a subsequently cleaves surrounding fluorophore-quencher dual-labelled probes producing a fluorescent signal. The SHERLOCK technology based on CRISPR/Cas13a integrated with isothermal amplification for SARS-CoV-2 detection has been reported [92], evaluated in clinical settings [93], and authorized by the US FDA for urgent COVID-19 applications [94]. Subsequently, this strategy was validated into a simplified STOPCovid test using nasopharyngeal swabs from COVID-19 patients [94]. This test is equivalent to RT-qPCR-based SARS-CoV-2 tests and has a limit of detection of 100 copies of viral genome. Recently, a CRISPR-based diagnostic assay was developed for SARS-CoV-2 mutation detection that can be used in analyzing nasopharyngeal specimens without RNA purification. The authors further reported that an engineered AsCas12a enzyme makes it possible for detection of wildtype and mutated SARS-CoV-2 that can be amplified using LAMP assay [95]. An inexpensive, modular heater could be developed that is fit and suitable for a disposable, single-use test to be used in the POC settings [94].
Immunoassay-based diagnostic methods
An immunoassay is a diagnostic test that can measure current or previous viral infection utilizing antigen-antibody interactions, either by utilizing monoclonal antibodies to detect viral antigens in clinical samples or by utilizing cloned viral antigens to recognize patient antibodies directed against the virus. These antibodies are present in patients who have recovered from SARS-CoV-2 infection and exist in blood, tissues, and other biomarkers throughout the body [96]. A COVID-19 disease biomarker can be measured in different kinds of immunoassays [96]. Although the outcome is similar, the method and technology behind the test may differ entirely. One of the most significant differences is that although most immunoassays employ labels, others do not. A number of labels such as enzymes, radioactive isotopes, fluorogenic reporters, DNA reporters, chemical probes, and nanomaterials are used in labelled immunoassays to modify or detect the antibodies and antigen analytes. The two very common enzyme probes used in immunoassays are horseradish peroxidase (HRP) and alkaline phosphatase (AP) or glucose oxidase (GO) and are referred to as enzyme immunoassays (EIAs) [96–98], ELISAs [96–98], and enzyme-multiplied immunoassay technique (EMIT) [96–98]. Radioimmunoassays (RIAs) [96, 97, 99] use radioactive isotopes to label antigen or antibody. Fluoroimmunoassays (FIAs) [100] are similar to RIAs, except instead of a radioisotope, the label is a fluorophore (e.g., Rhodamin and phycoerythrin). Protein microarrays are a type of immunoassay in which fluorogenic reporters are frequently used [101]. A relatively innovative method to immunoassays encompasses integrating real-time quantitative polymerase chain reaction (RT-qPCR) with conventional immunoassay methods named as real-time immunoquantitative PCR (iqPCR) [102, 103]. DNA reporters are used as label in these assays. Other label-based immunoassays are particle counting immunoassays (PACIAs) [104], liposome immunoassays (LIAs) [97, 105], flow-injecting immunoassays (FIIs) [106], chemiluminescence immunoassays (CLIAs) [107], and lateral flow or immunochromatographic immunoassays [108]. Label-free immunoassays use detection methods that do not rely on labeling or modification. The main features of these different immunoassays are summarized in Table 3 immunoassays. Each approach has its own set of benefits and downsides, so we may pick and choose the most suitable method to optimize our test for COVID-19 diagnostics.
Table 3.
A summary table of different immunoassays with their main features.
| Types | Immunoassays | Labels/features | Detection read-out | Sensitivity | Pros | Cons | References |
|---|---|---|---|---|---|---|---|
| Labelled immunoassays | EIA, ELISA, or EMIT | Enzymes (HRP, AP, GO) | Color | High |
-Sensitivity: high -Speed: rapid -Automation: excellent -Usability: wide -Shelf-life: long -High-throughput |
-Personnel required: trained -Health hazard: substrate carcinogenic -Time consuming: 16–20 h -Sample volume: μL–mL -Per sample: expensive -May cause false positive due to poor specificity and color transition |
[97, 98] |
| Radioimmunoassays (RIAs) | Radioactive isotopes | Radiation | Ultrahigh |
-Specific and sensitive -Minimal analyte detection -Automation: excellent |
-Personnel required: skilled and licensed -Health hazard: YES -Expensive radioactive elements -Expensive instrumentations -Isotopes must be used within few weeks -Shelf-life: short -Time consuming: 24–48 h -Sample volume: μL–mL -Contamination may cause false positive |
[97, 99] | |
| Fluoroimmunoassays (FIAs) | Fluorogenic reporters (phycoerythrin, Rhodamin) | Fluorescence | High |
-Sensitive, specific, and safe -Rapid -Fluorescence background: low -Automation: simple and accessible |
-Fluorescent equipment dependent -Bleaching of photographs -Necessity to improve assay parameters |
[100] | |
|
Real-time immunoquantitative PCR (iqPCR) |
DNA reporters | Fluorescence | Ultrahigh |
-More sensitive and reliable than ELISA -Background signal: low -Applicability: diverse -Could be developed multicomponent immunoassays |
Limitations: non-specific binding of the assay components to the solid phase | [102, 103] | |
| Particle counting immunoassays (PACIAs) | Polystyrene beads | Counting | High | -Simple, sensitive, homogeneous, rapid | -Quantitative results are variable | [104] | |
| Liposome immunoassays (LIAs) | Liposomal-encapsulated labels |
-Fluorescence -Electrical signal for analytical results |
Ultrasensitive |
-Personnel required: minimally trained -Health hazard: NONE -Sample volume: μL -Automation: ideal |
-Commercially available: very few -Speed (h): 2 |
[97, 105] | |
| Flow-injecting immunoassays (FIIs) | Enzyme: (HRP, GO), chemiluminescent | Fluorescence | High |
-Speed: 10 min or less -Avoidance of using direct labeling like dyes -Chemiluminescent label in solid phase -Selectivity: high -Easiness |
-Carryover of high samples -Sample throughput -Revival result in a drop of sensitivity |
[106] | |
| Chemiluminescence immunoassays (CLIAs) | Chemical probes | Visible light | Ultrahigh |
-Sensitivity: excellent -Reagent: steady -High-throughput -Automation: easy and can be incorporated in high-throughput |
-External equipment dependent to capture the light signal -Substantial reagent usage -Contaminating reagents cause poor selectivity and reproducibility |
[107] | |
| Lateral flow or immunochromatographic immunoassays | Colloidal gold nanoparticle | Visible color | Low |
-Rapid and easy to use -Inexpensive and safe -No need external equipment to visualize the signal -Can be used as POC test -Shelf-life: long -Utilization of nanoparticle markers could improve precision |
-Sensitivity and specificity: low because of sample pattern -Color perception: biased to individual |
[108] | |
| Label-free immunoassays | Immunosensors | Label-free and setup on transducers |
-Optical (color, fluorescence) -Electrochemical -Mass-based (piezoelectric, magnetoelastic) |
High |
-Rapid -Selectivity & sensitivity: high -Automation: YES -High-throughput -Potential to use for onsite detection |
-Labels are costly -Electrode: difficult to sustain -Buffer preparation: challenging -Personnel required: highly trained |
[36] |
It is also important to emphasize that there is a drive to advance label-free immunoassays right now. Physical principles such as beneficial and harmful light intervention, resonance circumstances, and how variations in efficient reflection coefficient affect these parameters underpin these ingenious technologies. In label-free assays, antigen-antibody interaction can be detected with no requirement of a light-emitting label. As a result, the sensitivity of the test can be enhanced while the operational time is reduced.
Test importance based on antigen detection
Rapid antigen testing identifies fragments of virus outer surface proteins as a marker for detection of an active COVID-19 infection. This test could allow additional information before or at the time of sampling for molecular testing [109]. When a HCoV clinical samples contain an adequate amount of antigen, it will bind to specific antibodies fixed to a device that will allow visualization. The antigens distinguished are expressed only when the viral infection is effectively replicating; thus, such tests are best used to detect acute or early infection. The infection starting time, the quality of specimen collection, and processing, elevated viral load in a sample, may increase the diagnostic sensitivity of rapid antigen tests [110]. Present methods for influenza and respiratory syncytial virus experience the effect of imperfect sensitivity to preclude the diagnosis of infection [111]. The same difficulty would presumably persist for SARS-CoV-2, and the tests would require execution with clear guidance and comprehensive oversight. There are eleven commercially available SARS-CoV-2 antigen quick diagnostic assays were developed for analytical sensitivity testing in order to identify the SARS-CoV-2 variants of concern alpha, beta, gamma, and zeta [112]. Although analytical testing with cultured virus can be used as a surrogate for clinical validity, it is not a substitute for clinical examinations. Despite minor variations in sensitivity, the authors [112] demonstrated that Ag-RDTs are still successful in detecting variants of concern.
Even though the sensitivity of these assays is lower than that of RT-PCR, they can reliably detect elevated viral loads correlated with the accumulation of infectious viral particles, rendering them valuable public health measures [113]. Monoclonal antibodies against the nucleocapsid protein of SARS-CoV-2 have been created, which may provide the foundation for a fast POC antigen detection test [114].
Test importance based on antibody detection
Antibody testing detects antibodies against the virus in subjects who have been infected with the virus in the past. Serological tests, utilizing ELISA, detect the presence of antibodies (such as immunoglobulins IgA, IgM, and IgG) from clinical specimens [115]. Antibodies are generated over days to weeks after infection exposure and allow the virus to be readily detectable [116]. A recent study reported that both IgM and IgG antibodies were recognized 5 days after incidence of SARS-CoV-2 [117]. An antibody test can be performed to facilitate the diagnosis of SARS-CoV-2 infections, where molecular assays were not satisfactory, and specimens were collected improperly. In the case of some patients, antibody response was developed only in the second week after exhibiting symptoms [118]. Certain factors such as age, nutritional status, onset and severity of infection, some medications, and immunosuppression may affect the production of antibodies [118]. Cross-reactivity of SARS-CoV-2 antibodies to other non–SARS-CoV-2 HCoV proteins may generate false-positive results [118]. Plasma samples of COVID-19 patients were tested against the (S) protein of SARS-CoV-2 and SARS-CoV and showed a high frequency of cross-reactivity among them [119]. An antibody test does not need to be rapid for patients already hospitalized with complications, since they would not need to use a screening test. However, an antibody test could be suitable as supplementary diagnostic tools for patients under medical care with late complication stages of the infection, when RT-PCR may be negative since viral shedding decreases 10 days after onset of symptoms [120]. Currently, serological tests for detection of SARS-CoV-2 are under development [35, 117]. Recently, IgG and IgM from human serum of COVID-19 patients were detected using an ELISA assay [117] using the Rp3 nucleocapsid antigen protein from SARS-CoV-2. The Rp3 has a 90% amino acid sequence similarity to other SARS-related viruses. Testing against SARS-CoV-2 through ELISA could be feasible, as more data on the infection becomes available. A POC serology test could reveal the presence or absence of patient antibodies against the viral infections in patients’ blood. This information allows medical staff to take necessary steps for proper treatment and mitigation of COVID-19 SARS-CoV-2 infections.
Technology advancements for SARS-CoV-2 diagnostics
In the next section, we will review recent technology developments for the diagnosis of SARS-CoV-2 infections.
Microarray methods
The microarray technology measures the nucleic acid expression level of a large number of genes simultaneously using hybridization probes [121, 122]. The technique could be adapted to use for detection of SARS-CoV-2. A PCR assay integrated with microarray technology was developed for detection of SARS coronavirus [123]. However, SARS-CoV can rapidly mutate, leading to the development of a microarray to detect single nucleotide polymorphisms (SNPs) characteristic of the (S) gene of SARS-CoV and generating 100% accuracy in sample detection [124]. A non-fluorescence oligonucleotide array was developed for detection of coronavirus at the genus level that produced equal sensitivity with that of individual RT-PCR assay [125]. The quantitative sensitivity and specificity of RT-qPCR tests for COVID-19 are higher under optimal conditions. When the sampling pool is expanded using asymptomatic people, the sensitivity drops and false negatives occur. Furthermore, RT-qPCR has a longer turnaround time because the majority of the time is spent extracting RNA from swab samples [126]. Damin and colleagues [126] developed a microarray-based assay coupled with hybridization probes for SARS-CoV-2 RNA detection with high sensitivity from nasopharyngeal swab samples. This technology also includes a RNAGEM that offers a viable alternative to commercially available RNA extraction kits. This microarray technology has the potential to be used in the field deployable platform with multiplexing capability for detection of other viral and bacterial diseases. A portable and POC diagnostic platform based on the microarray chip, the Mobile Analysis Platform (MAP), was developed for detection of influenza (flu) A/B, RSV, and MERS coronavirus [127]. The total turnaround time of the MAP device including automated sample processing, PCR amplification, and detection was 110 min. The clinical outcomes of MAP were 97% for flu A, 100% for flu B, and 100% for respiratory syncytial infection (RSV). The approximate limit of detection (LOD) of the MAP was 30 copies/assay for RSV and 1500 copies/assay for MERS coronavirus. A recent study reported that SARS-CoV-2 infection mutated into at least 30 different strains [128]. These mutations are capable of changing its pathogenicity. The microarray technology could help to detect SARS-CoV intra- and inter-species transmission with the novel emergence of SARS-CoV-2 strains.
Microfluidic methods
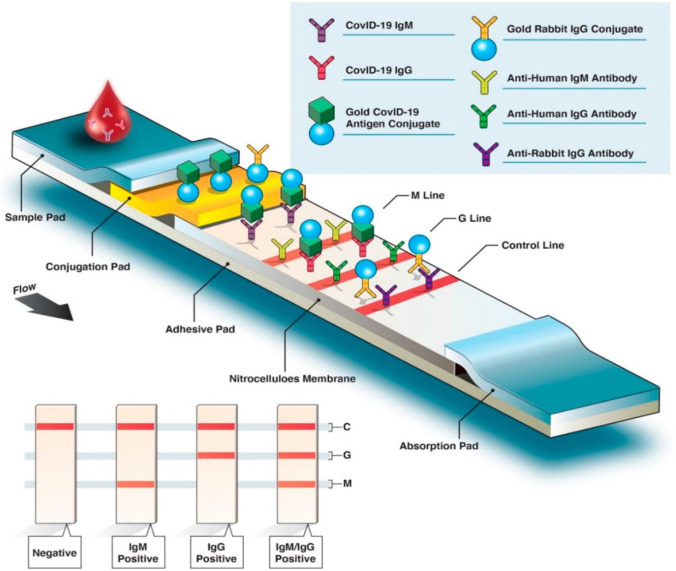
The microfluidic devices concerned with the precise control and manipulation of fluids within microscopic channels [129]. These devices comprise a small-sized chip with micrometer-sized channels and reaction chambers. The scalability and quality of microfluidics largely depends on the fabrication process of the device. These chips can be fabricated with inorganic, organic polymer, and paper. Inorganic materials include silicon, glass, and ceramic [121]. Polymers include polydimethylsiloxane (PDMS), poly (methyl methacrylate) (PMMA), polypropylene (PP), polyimide (PI), hydrogel, polystyrene (PS), polyethylene terephthalate (PET), and polycarbonate (PC) [130]. Paper is a flexible cellulose and polyethersulfone membrane material that can detect several biotargets [131]. The key advantages of utilizing paper microfluidics incorporate scaling down, small sample volume, fast detection times, and portability [132]. A lateral flow device (LFD) is a paper-based biosensor developing lateral flow immunoassay (LFIA) tests and lateral flow nucleic acid tests, respectively, that produce a colorimetric signal output [133]. A commercial lateral flow assay incorporated with a paper-like membrane strip is coated with two borderlines, one with gold nanoparticle-antibody conjugates and one that captures antibodies in the other [133]. The clinical sample is transferred to the membrane and proteins are passed across the strip by capillary force. Next, the antigens attach to the gold nanoparticle functionalized antibody and the complex moves along through the membrane. When they get to the other borderline, the complex is immobilized by the capture antibodies, and a colored line appears [134]. The LFIA test is simple and rapid and potentially be used as a POC test. In most cases, the LFIA test strips determine the appearance of patient antibodies (IgG, IgM, or IgA) generated by SARS-CoV-2 antigens using human serum, whole blood, or a drop of blood from finger prick (Figure 5 LIFA antibody detection) (Figure 6).
Fig. 6.
A quick look at the fast diagnostic serological test. Colorimetric lateral flow immunoassay (LFIA).
Reproduced with permission from Ghaffari, A. et al. Copyright MDPI (2020), Diagnostics [108]
Nucleic acid-based testing can be integrated into lateral flow methods for POC detection of SARS-CoV-2. Microchip, digital array chip, and paper-based microfluidic biosensors for fast diagnosis of COVID-19 could be advanced as a result of developments in POC testing to identify antibodies, antigens, or nucleic acids in raw samples which include saliva, sputum, and blood [135]. They have a number of benefits, including being cost-effective, sensitive, specific, easy to use, fast and efficient, instrument-free, and accessible to target consumers (ASSURED) [135]. The findings can be retrieved quickly and easily, allowing for quick judgment and reducing the chances of human-to-human disease exposure. With the ongoing COVID-19 pandemic, a better overview of the existing microfluidic-based detection techniques can aid physicians and scientists in incorporating innovative, fast, and suitable clinical detection methods that mitigate societal expenditures, speed the examination process, improve diagnostic performance, and ultimately halt the pandemic globally [136].
COVID-19 detection with nanomaterial-enhanced biosensors: new horizons
Nanomaterials and nanotechnology are used to make ultrasensitive biosensors for detecting nucleic acids, and proteins. A biosensor has many features that produce detection very efficient. These characteristics include measurable approach, distinctive functionality, good selectivity, rapid response times, portability, miniaturization, and precision. Various nanomaterial-advanced biosensors are devices wherein the transducer [137] has been transformed to encapsulate the intended bioreceptor components of pathogens (including viruses) derived from biological samples [138]. They transform the biological reaction into transducers, such as optical-, electrochemical-, and mass-based micromechanical (piezoelectric, magnetic, magneto-optical, and magnetoelastic) signals read-out, and detect it rapidly and correctly [139]. Nucleic acids (DNA, RNA, and PNA molecules), DNA probes, enzymes, organelles, proteins, aptamers, exosomes, entire cells, biomimetics, microorganisms, phages, and tissues are examples of bioreceptors [138]. In the current COVID-19 pandemic that has compromised the health of the world’s people and economies, nanomaterial-enhanced biosensors could be an effective device for speedy, efficient, portable, mass manufacturing compatibility and a more promising diagnostic alternative to conventional diagnostic methods [140]. Electrochemical and optical biosensors have sparked a lot of interest among diagnostic users because of their ease of detection, downsizing, sensor size flexibility, and sensitive parameter controllability [140].
Electrochemical biosensors
An electrochemical biosensor incorporates the sensitivity of electroanalytical strategies and the specificity of biorecognition components to detect chemical and biological analytes [141]. In 2019, Layqah and Eissa used carbon array electrodes coated with gold nanoparticles to develop voltammetric immunosensors for detection of the MERS-CoV [142]. These sensors, on the other hand, were technical in character, necessitating the use of high-quality analytical reagents and components. Moreover, in order to achieve superior sensitivity, extra labeling and signal amplification were needed, which could result in increased expenditures during future downsizing. Alafeef and coworkers (2020) advanced a fast, inexpensive, simple, and quantitative paper-assisted electrochemical sensor chip to allow the digital detection of SARS-CoV-2 RNA targeting N gene. The authors use gold nanoparticles (AuNPs), coated with extremely specific antisense probes (ssDNA) attached on a paper-based graphene electrochemical framework producing an electronic impulse amplification. This system has the ability to create a simple and quick handheld device [143]. Carbon electrode arrays modified with nanostructured gold nanoparticles were developed for multiplexed detection of coronavirus targeting spike protein S1 [142]. The LOD was 0.4 and 1.0 pg/mL for HCoV and MERS-CoV, respectively. The method was single step, sensitive, and accurate. Using a universal electrochemical biosensing, a multilevel analyte (DNA, RNA, protein) could be detected in a single platform [144]. This technology could be refined and implemented for rapid use in identifying SARS-CoV-2. Biosensors with two-dimensional nanomaterials like graphene or black phosphorus are being developed for POC diagnostics and could be utilized for COVID-19 testing [145]. For instance, in a recent work, a smart antibody sensor based on gold nanoarchitecture-aided laser-scribed graphene biosensor electrodes for SARS-CoV-2 detection with an estimated LOD of 2.9 ng/mL was recently implemented [146]. Screen-printing is used to make graphene-assisted electrodes because of being inexpensive, easy fabrication, and increased efficiency. In addition to graphene, biosensors based on black phosphorus (BP) or phosphorene could be investigated for COVID-19 diagnosis [147]. Black phosphorus has outstanding electrochemical capabilities that improve test sensitivity and selectivity because of its intrinsic redox characteristics.
Optical biosensors
Novel photonic technologies have become promising for a broad array of applications in numerous disciplines for disease detection in recent decades. Various factors can be employed in diagnostics, such as energy, polarization, absorption, fluorescence, light scattering, amplitude, decay time, and/or phase. The surface plasmon resonance, localized surface plasmon resonance, and fiber optics in biosensing applications have gotten a lot of interest because of their great sensitivity, and enforcement, as well as rapid reaction [148, 149]. These approaches and criteria could be combined with nanomaterials to determine if fluorescence, luminescence, or colorimetric detection is the best option for SARS-COV-2 virus detection. A nanophotonic LIGHT technology was developed for fluorescence-based rapid detection of E. coli from urine samples [150]. This technology works in three ways: (1) selective enrichment filtering of E. coli from urine samples, (2) photothermal cell lysis, and (3) ultrafast photonic PCR on a chip. The end-point detection was 103 CFU/mL in 10 min. A lens to focus the excitation light LEDs requires minimal power consumption and is extremely low in cost compared with laser sources, making them an ideal PCR heating source for isothermal amplification developing POC testing. The refinement of LIGHT technology integrated with an inexpensive paper method could be an effective solution for the prevention and identification of emergencies related to health and biosafety as well as the clinical management of community-acquired SARS-CoV-2 infections. A quantum dot-based lateral flow immunochromatographic strips (QD-LFICSs) immunoassay integrated with a portable 3D-printed optical read-out platform was developed for detection of monoclonal antibodies of avian influenza virus (AIV) H7N9 [151]. The LOD was 0.0268 hemagglutination units (HAU) that was compared with a real-time PCR assay and was more sensitive than conventional colloidal gold ICSs. The LFA provides cost-effective POC tests that may be more practical for primary care physicians than a lot of the other methods. Chen and colleagues devised a multiplex reverse transcription loop-mediated isothermal amplification coupled with a nanoparticle-based lateral flow biosensor targeting RdRp and N genes [152]. This could be developed for widespread use for other infectious disease pathogens. Contrasted with serology tests that distinguish the presence of target antibodies, antigen-assisted lateral flow assays (LFAs) are less sensitive than RT-PCR yet may move toward the clinical sensitivity of RT-PCR with additional development and advancement. For instance, Standard Diagnostics as of lately commercialized an antigen-detecting LFA (list number 09COV30D) with a self-announced by and large sensitivity of 84% and specificity of 100% contrasted with RT-PCR [153]. While this is an amazing beginning, further exploration is expected to enhance this sensitivity and advancement of reasonable fast antigen LFA. A rapid, novel automated chemiluminescent immunoassay (CLIA)-based commercial product MAGLUMI™ 2000 Plus was developed to study the kinetics of IgM and IgG antibodies when COVID-19 patients produce low viral load in the late stage [154]. Both IgM and IgG rapidly increased after the onset of fever. IgG requires at least 12 days to attain 100% sensitivity, while the highest positive rate achieved for IgM was 88% throughout the study period. However, chemiluminescent immunoassay tests for IgM and IgG antibodies detect an immune response too late to do any good in minimizing transmission. An SPR-based biosensor integrated with a gold substrate was developed for detection of SARS-CoV surface antigen [155]. The LOD of 200 ng/mL was detected within 10 min. The SARS-CoV-2 can be detected using CANNARY biosensor technology. This technique uses a cell-based immunosensor that integrates capture of the SARS-COV-2 with signal amplification to produce a result in 3–5 min (www.pathsensors.com/psi-sars-cov-2-biosensor/). Subsequently, a quick, efficient, and cost-effective interferometric optical biosensor test capable of detecting immunoglobulins in serum and saliva samples was developed [156]. We expect that developing real-time, inexpensive, label-free nanophotonic biosensors will open up new possibilities for detecting COVID-19 in a timely and effective manner [157].
Other biosensors
The combined electromechanical properties of piezoelectric materials make them ideal for use as sensors and actuators in smart buildings and devices for COVID-19 diagnostics [158]. Piezoelectric materials have also emerged as key nanomaterials that may be integrated with living tissue employed in diagnostics that are compact bioelectronic and biochemical devices. Magnetic and magnetostrictive materials, on the other hand, may transform magnetic power into mechanical power or the other way around, and they are utilized to make sensors and actuators [158]. Additional material-based biosensors, namely textile-based, film-based, or carbon-based biosensors, have also been proposed for COVID-19 application [159]. They are being created to enhance the efficiency and detection sensitivity of current biosensors, allowing the users additional viable options. Simple manufacturing procedures and enhanced assay effectiveness have been established for textile-based biosensors including thread-based, fabric-based, or cloth-based biosensors. Polysiloxanes with variable hydrophobicity were used into thread-based biosensors in one study to defer fluid flow in lateral flow assays and increase detection sensitivity [160, 161]. Apart from textile-based biosensors, film-based biosensors have been created to detect infectious microorganisms from raw samples. For example, a thin polyester film base, a sample compartment, a lid, a reaction compartment, and a waste chamber have all been incorporated in a film-based biosensor [162].
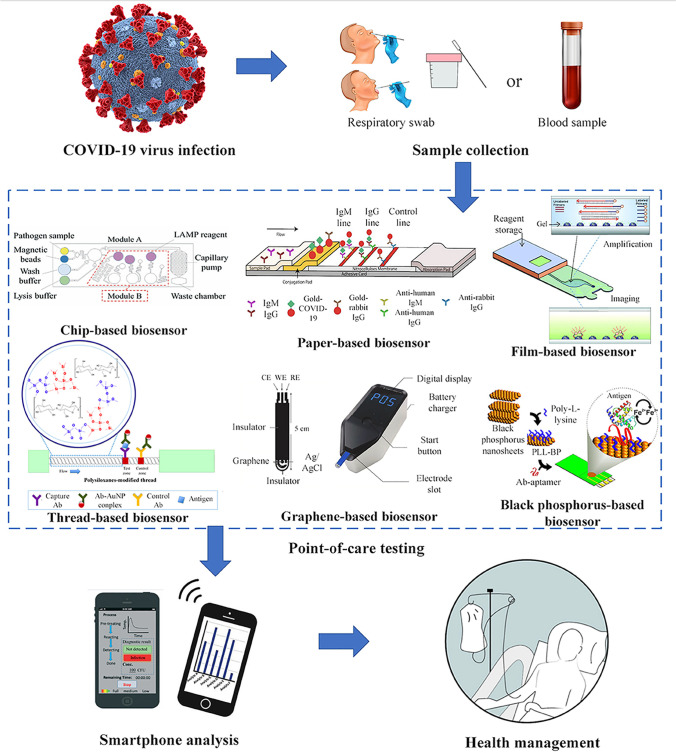
Biosensors with nanomaterial enhancements can open up new possibilities, such as more effective, convenient, and safe uses. Costs, environmental sensitivity, the need for large devices, and regulatory difficulties should all be addressed before the product is released to the market [163]. Table 4 shows the methodology, their principles, the samples required, the cost, the benefits, and drawbacks of biosensor technologies over traditional methods for detecting SARS-COV-2. Table 5 summarizes various nanomaterial-enhanced biosensors and their features for SARS-CoV-2 virus detection. Although numerous approaches for detecting SARS-CoV-2 virus can be used, the pandemic transmission of COVID-19 necessitates the development of POC [Figure 7].
Table 4.
A summary table with all of the procedures, their concepts, the samples needed, the cost, and the benefits and drawbacks of biosensor technology against traditional methods for detecting SARS-COV-2.
| Molecular assays | Immunological assays | Nanomaterials-based biosensors | ||||||
|---|---|---|---|---|---|---|---|---|
| 1. Tests | RT-PCR | Isothermal PCR |
Nucleic acid hybridization microarray | Antigen | ELISA | Lateral flow | Chemiluminescence | Electrochemical Optical Mass-based |
| 2. Specimens | Upper respiratory | Upper respiratory | Upper respiratory | Blood serum or plasma | Blood serum or plasma | Blood serum or plasma | Blood serum or plasma | Upper respiratory, blood, or urine |
| 2. Target | RNA | RNA | RNA | Viral protein | Antibody against virus in patient’s blood | Antibody against virus in patient’s blood | Antibody against virus in patient’s blood | RNA, antigen, antibody |
| 3. Diagnosis of asymptomatic cases | Negative or anticipated inadequate value | Negative or anticipated inadequate value | Negative or anticipated inadequate value | Negative or anticipated inadequate value | Likely false negative in early infection | Likely false negative in early infection | Likely false negative in early infection | The principle varies depending on the approach |
| 4. Diagnosis of symptomatic cases | Current reference test | Likely correlate with reference test | Likely correlate with reference test | Under development or in regulatory phase | Diagnosing acute infections is likely very limited around the time of symptom onset | Diagnosing acute infections is likely very limited around the time of symptom onset | Diagnosing acute infections is likely very limited around the time of symptom onset | The principle varies depending on the approach |
| 5. Screening of viral shedding in recovery phase | Negative or anticipated inadequate value | Negative or anticipated inadequate value | Negative or anticipated inadequate value | Probably negative or anticipated inadequate value | Generally, do not reflect active virus but detect patient’s antibody | Generally, do not reflect active virus but detect patient’s antibody | Generally, do not reflect active virus but detect patient’s antibody | The principle varies depending on the approach |
| Advantages |
-Current reference test, -Very sensitive and specific, if performed accurately -Rapid RT-PCR can be used at POCT |
-Does not require thermocycler, -Likely very sensitive and specific, if performed accurately -Cost: very cost-effective -Likely to be used a POCT |
-Simple and -Fast -Could be used a POCT |
-Fast -More accurate than lateral flow assays -Cost: not very costly -Give quantitative information |
-Cost: inexpensive -Increase specificity -Fast -User-friendly -Could be used a POCT |
-Sensitive -Rapid |
-Sensitive -Fast -User-friendly |
|
| Disadvantages |
-Expensive -Laborious and -Time-consuming -Tests still need to be performed in the centralized laboratory. -High risk of false negative due to bad sampling -Likely shortage of reagents |
-Likely shortage of reagents | Cost: expensive |
-Complex to develop -Cost: variable |
-Expensive -Laborious and -Time-consuming -Tests still need to be performed in the centralized laboratory -Likely shortage of reagents Possible false-negative results if performed too early onset of infection -Possible false-positive results if mixed with other diseases |
-Provides only qualitative information (presence or absence of antibodies) -Likely shortage of reagents (blocking buffer, wash buffer, coating buffer) -Possible false-negative results if performed too early onset of infection -Possible false-positive results if mixed with other diseases |
-Cost: expensive | -Cost: some forms are expensive (for, e.g., magnetic and plasmonic methods require costly instrumentations) |
Table 5.
Different biosensor approaches based on nanomaterials and their features for SARS-CoV-2 detection
| SARS-CoV-2 | |||||
|---|---|---|---|---|---|
| Samples | Nanomaterials | Transducer | Bioreceptor | LOD | Ref. |
| Human nasopharyngeal swabs | Two-electrode screen-printed carbon electrode | Electrochemistry-pulse voltammetry | DNA/cDNA/RNA | 1 copy/μL | [84] |
| Human nasal swab or saliva | Graphene-ssDNA-AuNP | Digital electrochemical–Ramon spectrum | RNA | 6.9 copies/μL | [143] |
| Upper respiratory | Plasmonic chip | Optical fluorescence-LSPR | cDNA/nucleic acid | 0.220 pM | [164] |
| Human nasopharyngeal swab, culture | Graphene sheet | Electrochemical-FET | S1 spike protein antigen | LOD: 1.6 × 101 pfu/mL in culture medium 2.42 × 102 copies/mL in clinical samples | [165] |
| Culture | Membrane-engineered vero cells (vero/anti-S1) | Bioelectric recognition assay (BERA) | SARS-CoV-2 S1 spike protein antigen | 1 fg/mL | [166] |
| Human serum/nasopharyngeal swab | Gold nanorods | SPR | Antibody | 111.11 deg/RIU | [167] |
| Human oropharyngeal and nasopharyngeal swab | Magnetic γ Fe2O3 nanoparticles | Colorimetric-3,3′,5,5′-tetramethylbenzidine (TMB) | S protein of SARS-CoV-2 | 4.98 ng·mL−1 | [168] |
| Human saliva | Screen-printed gold electrode | Electrochemical-colorimetric | Spike antigen of SARS-CoV-2 | 1 pg/mL | [169] |
Fig. 7.
Point-of-care (POC) for COVID-19. Reprinted with permission from Choi, J. et al. Development of point-of-care biosensors for COVID-19. Front Chem 8: 517. Copyright (2019) Frontiers in Chemistry [159].
Benefits and limitations of nucleic acids, antigens, and antibody tests POC devices
Because of its great specificity and capacity to swiftly create myriad of copies of a single RNA or DNA sample, RT-PCR is the preferred technique for detecting the SARS-CoV-2 virus. However, because of the range and precision in temperatures required for the amplification of nucleic acids in the sample, it is challenging to convert this technique to a portable device. Similarly, the typical RT-PCR procedure uses kits available in the market today for RNA extraction and purification. The kit extracts and filters RNA from the viral sample, and based on the amount of fluid utilized after purification, it may also concentrate it, which helps to improve assay sensitivity.
To speed up the process of diagnostics in vitro accessing the global commercial market, the USA used an Emergency Use Authorization (EUA) to allow urgent use of such devices for detection of SARS-CoV-2 or diagnosis of COVID-19 [170]. Some instances of EUA-approved RT-PCR-based POC devices that have been successful include Accula SARS-CoV-2 Test (Mesa Biotech Inc.), cobas SARS-CoV-2 and Influenza A/B Nucleic Acid Test (Roche Molecular Systems, Inc.), The Xpert Xpress SARS-CoV-2, Xpert Xpress SARS-CoV-2/Flu/RSV, and Xpert Xpress SARS-CoV-2 DoD (all from Cepheid), Visby Medical COVID-19 Point-of-Care Test (Visby Medical, Inc.), and BioFire Respiratory Panel 2.1-EZ (BioFire Diagnostics, LLC) [171]. These EUA-authorized tests can be used at the POC (i.e., in patient care environments operating under a CLIA Certificate of Waiver, Certificate of Compliance, or Certificate of Accreditation).
The elimination of the viral purification phase and the reduction in instrumentation complexity are the two main advantages of using isothermal amplification-based techniques for SARS-CoV-2 virus detection [79]. Abbott Diagnostics Scarborough, Inc. released the ID NOW COVID-19 test, which uses RT-LAMP, shortly after the pandemic began. This was the first time an isothermal method had been approved by the EUA for COVID-19 testing [79]. Another EUA-authorized test (Cue COVID-19 Test, Cue Health) uses isothermal amplification in a single-use cartridge to detect the virus from direct nasal swabs in 20 min using an electrochemical detection approach with a limit of detection of 20 genome copies per sample [172]. Cue COVID-19 molecular diagnostic test was approved in March 2021 for nonprescription, over-the-counter usage, and healthcare practitioners without CLIA certification [172]. Lucira COVID-19 All-in-One single-use Test Kit [173] features a handheld battery-powered instrument that performs an RT-LAMP assay for COVID-19 detection. This device was first approved by the FDA (USA) for COVID-19 testing at home using self-collected nasal swabs.
The N protein is the target of the majority of antigen tests now available. The utilization of the S protein, on the other hand, may be more particular because it shares less sequence homology with prior SARS-CoV and MERS viruses [174]. Antigens that have worked well in POC devices that have been authorized by the EUA include the CareStart COVID-19 Antigen test (Access Bio, Inc.), LumiraDx SARS-CoV-2 Ag Test (LumiraDx UK Ltd.), BinaxNOW COVID-19 Ag Card marketed by Abbott Diagnostics Scarborough, Inc., Becton, Dickinson and Company, LLC marketed BD Veritor System for Rapid Detection of SARS-CoV-2, Clip COVID Rapid Antigen Test (Luminostics, Inc.), QuickVue SARS Antigen Test, Sofia 2 SARS Antigen FIA, Sofia 2 Flu + SARS Antigen FIA (all three from Quidel Corporation), and Status COVID-19/Flu (Princeton BioMeditech Corp.) [171]. The N protein antigen from SARS-CoV-2 can be detected qualitatively using any of these approaches. As a result, they all have an extraction buffer to rupture the viral proteins in the sample and release the viral nucleoproteins inside [171]. These technologies are allowed to be used at the POC and hands-on efforts are required.
The Ellume COVID-19 Home Test (Ellume Limited) was approved as the first antigen test to be licensed for nonprescription home testing by the EUA in December 2021 [175]. The EUA-authorized BinaxNOW COVID-19 Ag Card Home Test is another antigen test for home use marketed by Abbott Diagnostics Scarborough, Inc. that required prescription for use [176]. However, the BinaxNOW group also achieved approval for self-testing nonprescription home use for different aged people [177, 178]. Two additional home use antigen-testing devices have been added to the move [179, 180]. It is presently unclear if patients who cured from SARS-CoV-2 infection gained antibodies that can prevent them from re-infection, as some recognized and suspicious cases have been reported to have been exposed to the virus again [181]. The time of SARS-CoV-2 infection exposure, as well as the nature of the specimen taken for antibody testing from people who have recently been exposed to SARS-CoV-2 infection, could result in antibody positive to spread the infection [182].
Several serology tests, including EUA submissions pending, have recently been approved by EUA. However, only a few have been approved as POC devices, such as lateral flow immunoassay-based tests were developed for IgG/IgM antibody screening for COVID-19 and was marketed by Fa-StepTM, Assure Tech. (Hangzhou, China), RightSignTM, Biotest Biotech (Hangzhou, China), RapCoVTM, Advaite, Inc. (Malvern, PA, USA), MidaSpotTM Nirmidas Biotech, Inc. (PALO ALTO, CA, USA), and SiennaTM-Clarity COVIBLOCK™ (Örninkatu, Salo, Finland) [171].
Surveillance and epidemiology of SARS-CoV-2
Here, we describe some emerging diagnostic technologies that can be adapted for detecting COVID-19. However, the containment of SARS-CoV-2 likely involves many complex challenges as the asymptomatic individuals are still capable of transmitting the virus [183]. Therefore, curbing SARS-CoV-2 transmission needs extensive surveillance, sharing epidemiological data, and patient monitoring. Cell phones can be utilized for this purpose as they possess connectivity, hardware to expedite electronic reporting, epidemiological databasing, and POC tests [184]. One possibility is voluntary digital tracing of COVID-19 by protecting personal privacy and developing an application programming interface (APIs) [185]. Flexible and wearable microfluidic sensor devices could be used to monitor health and online pathogen detection [186]. This wearable technology could be developed for detection and monitoring of SARS-CoV. Using cell phone photography, the test results could be sent to a physician for immediate precision telemedicine care. A strategy relating international genomic surveillance and prompt evaluation of the genetic changes of new variants is needed to get ready for the pandemic’s succeeding stage, which will facilitate the growth and upgrading of diagnostics, vaccines, drug discovery, and non-pharmaceutical interference [187].
Artificial intelligence-based COVID-19 disease management
In the battle against COVID-19, artificial intelligence (AI) can be incredibly useful. In contrast to the SARS-CoV-1 outbreak in 2003, AI has contributed a more significant job including predicting and tracking of vulnerable areas of COVID-19 outbreak, contact tracing, advance diagnosis, protein structure prediction, development of therapeutics, and vaccines [188]. Man-made intelligence-empowered advances, like the AI robot, smart toilet, and cell phone, could offer important help for following, diagnostics, and therapeutics. As a nonsurgical system that can capture and analyze urine and stool samples, the smart toilet could be an effective technique for diagnosing COVID-19 where the nasopharyngeal test yielded a negative result [189]. For health tracking, scanners, force and movement detectors, and urine test strips have been used [190]. Diagnostics and service robots with AI centers can be utilized for collecting clinical samples and gathering tests from COVID-19 patients in jam-packed public spots, like customs, air terminals, ports, rail route stations, and so on [191]. The robot can work in hazardous conditions, consequently decreasing the pressure from clinical experts and medical care staff. With the steady progress of AI innovations, more intelligent and more impressive automated frameworks under control with the capability to sanitize, convey prescription drugs and food, monitor temperature and fundamental signs, diagnose, and help dissemination control will become tools to battle against potential viral epidemic outbreaks [191].
Outlook
Despite massive attempts and significant progress made by people from around the world to combat the pandemic, COVID-19 has become unrestricted and poses a risk to human societal structure. Both from human and viral perspectives, the causes are complex. However, two components are likely to have a larger role in the scenario: virus mutations and the presence of symptomless individuals.
Detecting emerging SARS-CoV-2 mutated variants
The SARS-CoV-2 virus has been constantly evolving during the recent worldwide pandemic. However, certain newly discovered variations have acquired much more mutations in a short period of time, generating alarm all across the world [192]. Mutations in SARS-CoV-2 present a profound influence on the need to swiftly pinpoint alterations in the vial genomic sequences that cause virus spread, posing a considerable difficulty for virus detection. Numerous parameters determine the effect of variants on a test’s effectiveness, along with the variation’s sequence, fundamental design distinctions in every test, and the incidence of the mutants in the individuals of many geographic locations [193]. The molecular qRT-PCR tests with a point mutation can result in false-negative results in detection [194]. As a result, mutated variants can limit the amplification sensitivity. Due to the possibility of selected mutations in SARS-CoV-2, specific locus of targeted genes must be avoided when designing oligos. Therefore, generating degenerate oligos is a viable solution to the SARS-CoV-2 mutation issue. In accordance with WHO’s revised guidelines on variant identifiers, dynamic strains can be documented as B.1.1.7 (Alpha), B1.351 (Beta), P.1 (Gamma), B.1.617.2 (Delta), B.1.617.1 (Kappa), P.2 (Zeta), and B.1.1.529 (Omicron) influencing viral spread, disease intensity, diagnostics, and vaccination effectiveness posing threat to the society [195]. Screening GISAID information revealed an upsurge of S protein amino acid mutations in various geographic locations [196]. Certain major mutations in the spike protein of B.1.1.7 (e.g., deletion 69–70, 144 and substitution K417N, K417T, E484K, N501Y, A570D, D614G, P681H, T716I, S982A, D1118H, and many others) have gotten a lot of attention in the scientific community and variant of concerns (VOC) are closely monitored [31, 197]. The analysis of sequencing materials sent to GenBank and GISAID revealed that mutations in the ORF1ab gene were most common in Germany and China [198, 199]. Another report revealed that the COVID-19 diagnostic primers suggested by the US CDC have showed mutations in all targets, while RT-PCR setups for the N gene included in Japan, Thailand, and China demonstrated numerous mutations in multiple groups, indicating the N gene may not be a consistent site. Therefore, the RT-PCR assay setup should be improved routinely to detect new alpha, beta, gamma, and delta variants [200, 201]. Over 30 mutations were found in the S gene of the most diverse SARS-CoV-2 omicron variant, raising threats about disease control and prevention [193]. Therefore, more accurate, simple-to-use, and uncomplicated detection procedures for SARS-CoV-2 mutations are urgently needed. Existing methods for determining possible mutations of SARS-CoV-2 include next-generation sequencing (NGS), metagenomic NGS, RT-qPCR, CRISPR/Cas, and RT-LAMP. Conventional detection techniques, on the other hand, are frequently difficult, rigorous, and have low sensitivity. SARS-CoV-2 and its mutation can be detected with effective biosensors such as surface plasmon resonance (SPR), local surface plasmon resonance (LSPR), and surface-enhanced Raman spectroscopy (SERS) using electrochemistry, piezoelectricity, fluorescence, and electrochemiluminescence signal reading.
Mutations in the SARS-CoV-2 virus can cause a shift in viral proteins, which can affect the results of an antigen or serology test. Antibody screening techniques frequently use antibodies (IgM and IgG) that uniquely attach to both the S and N proteins. Using thermally deactivate omicron samples, Abbott BinaxNOW and Quidel QuickVue antigen tests showed the omicron variant with similar accuracy as other variations [193]. These findings are not a substitute for ongoing clinical trials employing specimens infected with live virus, and they do not preclude the chance that the omicron variation has an effect on test efficiency. Although a study found that thus far discovered SARS-CoV-2 spike mutations have no effect on antibody testing [202], we choose to use kits that have antibody directed for the less often mutant N protein if at all possible. To address new variations and mutations, a number of public health organizations, university researchers, research institutions, and private enterprises are collaborating. To follow this dangerous infection more extensively, more genetic monitoring investigations employing focused genetic tests or whole viral genome sequencing approaches should be conducted. Different detection methods for identification of SARS-CoV-2 mutated variants are listed in Table 6.
Table 6.
Identification of SARS-CoV-2 variants using detection methods
| Methods | Properties | Ref. |
|---|---|---|
| Whole genome sequencing (WGS) |
-Shotgun sequencing or amplicon-based -The entire genome can be sequenced -Unbiased approach -Requirement of equipments and bioinformatic analysis -Takes several days -Costly |
[203, 204] |
| Sanger sequencing |
-Partial next-generation sequencing -Amplicon based -Targeted for whole or partial S gene -Requirement of equipments and bioinformatic analysis -Takes several days -Costly |
[205] |
| Multiplex RT-PCR |
-Targeted for deletion or target failure (drop out) (for instance, S gene target failure variants but not all) -S gene target failure can be integrated with E, N, ORF1 RT-PCR assay -The significance of the findings should not be compromised. -Further co-relation assessment and validation is required |
[193, 206, 208] |
| Single nucleotide polymorphisms (SNPs) assays |
-RT-PCR assay could be used for detecting SNPs (e.g., spike N501Y and HV69-70del mutations, present in B.1.1.7/501Y.V1 VOC) -Positive controls should be used in the assay -Some commercial melting curve analysis is used to detect SNPs, e.g., HV69-70del, K417N, N439K, Y453F, E484K, N501Y, A570D, D614G, P681H, or V1176F. -Further validation is required using sequencing |
[207] |
| Isothermal amplifications (RT-LAMP, TMA) |
-Rapid test -Need less resources -Highly sensitive and specific -Some protocols are limited to differentiate between variants -Proper clinical validation is required |
[209] |
| CRISPR |
-Turnaround time 20–50 min -Low level instrumentation is required -Higher sensitivity and/or specificity -Possibility for medium mutation detection |
[210] |
| Rapid antigen detection test |
-Shorter turnaround time -Inexpensive -Sensitivity is generally lower that RT-PCR -Targeting N protein, new variant B.1.1.7/501Y.V1 was detected -Further validation test is required |
[211] |
| Neutralization assays and antigenic characterization |
-Assess variants risk assessment, determining if VOI is a VOC -Requires biosafety level (BSL)-3 -Labor intensive |
[212] |
| Antibody test |
-Turnaround time is < 1 h -Low level instrumentation is required -Due to immunodeficiency and cross-activity sensitivity and/or specificity is medium -Possibility for lower mutation detection |
[193] |
Detecting infected people who are symptomless
SARS-CoV-2 identification is complicated by asymptomatic cases. Sometimes asymptomatic cases may still have virus particles in the upper respiratory system that are beneath the point of confinement of nucleic acid detection, and particular anti-SARS-CoV-2 IgG and IgM antibodies that are not detectable. Therefore, “false negative” findings are prone to be found in asymptomatic cases. As a result, developing effective strong testing for detecting asymptomatic case is more critical. Furthermore, efficient assessment services are primarily dependent on screening intensity and response rapidity, with improved test sensitivity coming in subsequently [213]. Even if the sensitivity of RT-qPCR is poor, it is possible that it will still create a pivotal impact. Another suggestion is to use an integration of RT-qPCR tests that are correctly scheduled with several repeats. For instance, once weekly (days 0 and 7) and on day 14th during the isolation phase and serological antibody testing for anti-SARS-CoV-2 IgM and IgG could be performed for the same time frame mentioned above. If individuals who have been discharged from quarantine are still releasing live virus, it is possible that other people can be infected from them [214]. As a result, in some situations, self-isolation is suggested for up to 1 month. Alternative RT-qPCR and serological antibody testing tests should be strongly advised.
Conclusion
An unsettling trajectory marks the spread of SARS-CoV-2. Advancement and implementation of viable technologies for SARS-CoV-2 molecular-, immunologic-, and biosensor-based tests for precise diagnosis of individuals infected with the SARS-CoV-2 is crucial to control the pandemic of COVID-19. Even vaccinated people may harbor and spread live viral infection without showing signs and symptoms. Therefore, molecular, immunological, and biosensor tests will continue to be important and rapid POC devices dramatically enhance COVID-19 care and minimize the expenses after vaccination phases and emergence of new variants [215]. The existing RT-qPCR-based diagnostic tests have some limitations, as they miss some cases of infection if performed inaccurately. Furthermore, they can only be executed in central laboratories by exceptionally trained operators. Thus, there is a basic need to develop rapid, reliable, inexpensive, and simple POC tests in order to detect and isolate patients who are at risk and need prompt care. Individuals may spread the infection on to numerous others before diagnosis, adding to the continued worldwide transmission of COVID-19. The fast, lateral flow immunoassay, colloidal gold immunochromatography, and automated chemiluminescent immunoassay tests for IgM and IgG could supplement the current COVID-19 testing by RT-PCR. However, there is a need to rigidly develop and assess the clinical performance of rapid tests of SARS-CoV-2. The microarray chip, silicon chip, paper chip, digital array chip, microfluidic, and beads-based assays could be refined and optimized for developing novel portable devices for POC SARS-CoV-2 detection. The next generation of nanomaterial-enhanced portable biosensors could be developed for real-time SARS-CoV-2 and its mutations detection at the remote locations with electrochemical-, optical-, and mass-based signal read-out [216]. Isothermal amplification assays (for instance, RT-LAMP, RT-RPA, RT-RCA, and NASBA), CRISPR/Cas, and immunoassays (such as lateral flow) could be integrated with lateral flow paper strip, microfluidic lab-on-a-chip devices, and inexpensive biosensor devices for visual detection of SARS-CoV-2 and its emerging mutated variants. The discovery of novel biomarkers (such as DNA, RNA, microRNA, proteins, exosomes, cells, and metabolites) is also the key in identifying ones that are associated with the early stages of COVID-19 infection. It is vital to find ways to lower the total price of sample collecting, testing, and assessment. In this case, using saliva as a sample could be a viable option providing the test sensitivity is not compromised. Saliva has been shown to be a viable option for detecting SARS-CoV-2 in the upper respiratory tract [79]. When it comes to mass screening, saliva has a numerous benefit over nasopharyngeal swabs, including the ability to be self-administered. We feel that using either nose swabs from the anterior nostrils or saliva to scale tests will be more efficient. Some commercially available POC tests are under development before they are utilized for the diagnosis of SARS-CoV-2. Recently, certain firms have been awarded multi-million-dollar contracts to manufacture antigen and molecular tests to be utilized at POC or at home in order to increase testing capacity [217–220]. The introduction of omicron SARS-CoV-2 (B.1.1.529), a new COVID-19 variant, is placing burden on diagnostic companies who determine the existence of SARS-CoV-2 to verify that their findings are unaffected. The ongoing endeavors will additionally lead to develop robust, reliable, fast, and simple to execute in vitro diagnostic assays and smart diagnostics for COVID-19. The integration of portable devices and assays could be developed for analysis of multiplexed results of pathogens. Digital biosensor technologies could be employed in the future to advance digital barcode fragments and a computer simulation framework for multiplex analyte measurement [216]. The integration of diagnostics and cell phones should give higher correspondence in population surveillance and epidemiology of COVID-19. For POC diagnosis, real-time monitoring via a cellular telephone is critical. It specializes in the derivatization of tangible handheld barcode components and possesses antibodies that are critical for biomarker adhesion to the extracellular environment. This can be used to detect biomarkers for SARS-CoV-2. There are as yet noteworthy knowledge and information gaps in health research that are vital for an effective public health response in the case of pandemics caused by COVID-19.
Acknowledgements
The author is grateful to Professor Stephen M. Boyle and Professor Tanya LeRoith (Virginia Tech) for critical reading of the manuscript.
Data availability
Not applicable for this submission.
Declarations
Ethics approval
This research did not involve any human and/or animals.
Conflict of interest
The author declares no competing interests.
Source of biological material
This research did not involve any human and/or animals.
Statement on animal welfare
This research did not involve any human and/or animals.
Footnotes
Author’s affiliation changed in the course of preparing the manuscript.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Note: Information on SARS-CoV-2 are advancing rapidly. A portion of this review may change as more studies and clinical performances of POC diagnostic devices become accessible. Some featured studies also identified shortcomings with their experimental study and design.
References
- 1.Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020 doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Gorbalenya AE, Baker SC, Baric R, Groot RJ, Drosten C, Gulyaeva AA, Haagmans BL, Lauber C, Leontovich AM, Neuman BW, Penzar D. Severe acute respiratory syndrome-related coronavirus: the species and its viruses—a statement of the Coronavirus Study Group. Nat Microbiol. 2020;5(4):536–544. doi: 10.1038/s41564-020-0695-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.WHO https://covid19.who.int/. Accessed 23 Dec 2021
- 4.Clark E, Chiao EY, Amirian ES. Why contact tracing efforts have failed to curb coronavirus disease 2019 (COVID-19) transmission in much of the United States. Clin Infect Dis. 2021;72(9):e415–9. doi: 10.1093/cid/ciaa1155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhang N, Li C, Hu Y, Li K, Liang J, Wang L, Du L, Jiang S. Current development of COVID-19 diagnostics, vaccines and therapeutics. Microbes Infect. 2020;22(6–7):231–5. doi: 10.1016/j.micinf.2020.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Raman R, Patel KJ, Ranjan K. COVID-19: unmasking emerging SARS-CoV-2 variants, vaccines and therapeutic strategies. Biomolecules 2021;11(7):993. 10.3390/biom11070993 [DOI] [PMC free article] [PubMed]
- 7.Shalhoub S, Farahat F, Al-Jiffri, , et al. IFN-α2a or IFN-β1a in combination with ribavirin to treat Middle East respiratory syndrome coronavirus pneumonia: a retrospective study. J Antimicrob Chemother. 2015;70:2129–2132. doi: 10.1093/jac/dkv085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Centers for Disease Control and Prevention. Interim guidance for antigen testing for SARS-CoV-2. https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antigen-tests-guidelines.html. Accessed 23 Dec 2021
- 9.Binnicker MJ. Challenges and controversies to testing for COVID-19. J Clin Microbiol. 2020;58(11):e01695–20. doi: 10.1128/JCM.01695-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wang R, Qian C, Pang Y, Li M, Yang Y, Ma H, Zhao M, Qian F, Yu H, Liu Z, Ni T. opvCRISPR: one-pot visual RT-LAMP-CRISPR platform for SARS-cov-2 detection. Biosens Bioelectron. 2021;15(172):112766. doi: 10.1016/j.bios.2020.112766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Vandenberg O, Martiny D, Rochas O, van Belkum A, Kozlakidis Z. Considerations for diagnostic COVID-19 tests. Nat Rev Microbiol. 2021;19(3):171–83. doi: 10.1038/s41579-020-00461-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sheahan TP, Sims AC, Leist SR, Schäfer A, Won J, Brown AJ, Montgomery SA, Hogg A, Babusis D, Clarke MO, Spahn JE. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat Commun. 2020;11(1):1–4. doi: 10.1038/s41467-019-13940-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Peiris Js Ls, Poon LL, Guan Y, Yam LY, Lim W, Nicholls J, Yee WK, Yan WW, Cheung MT, Cheng VC, Chan KH. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet. 2003;361(9366):1319–25. 10.1016/s0140-6736(03)13077-2 [DOI] [PMC free article] [PubMed]
- 14.Assiri A, Al-Tawfiq JA, Al-Rabeeah AA, et al. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: a descriptive study. Lancet Infect Dis. 2013;13:752–761. doi: 10.1016/S1473-3099(13)70204-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Rabi FA, Al Zoubi MS, Kasasbeh GA, Salameh DM, Al-Nasser AD. SARS-CoV-2 and coronavirus disease 2019: what we know so far. Pathogens. 2020;9(3):231. doi: 10.3390/pathogens9030231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Alhalaili B, Popescu IN, Kamoun O, Alzubi F, Alawadhia S, Vidu R. Nanobiosensors for the detection of novel coronavirus 2019-nCoV and other pandemic/epidemic respiratory viruses: a review. Sensors (Basel). 2020;20(22):6591. doi: 10.3390/s20226591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Delahay RJ, de la Fuente J, Smith GC, Sharun K, Snary EL, Girón LF, Nziza J, Fooks AR, Brookes SM, Lean FZ, Breed AC. Assessing the risks of SARS-CoV-2 in wildlife. One Health Outlook. 2021;3(1):1–4. doi: 10.1186/s42522-021-00039-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26(4):450–2. doi: 10.1038/s41591-020-0820-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–273. doi: 10.1038/s41586-020-2951-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wu A, Peng Y, Huang B, et al. Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe. 2020;27:325–328. doi: 10.1016/j.chom.2020.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Xu X, Chen P, Wang J, et al. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci. 2020;63:457–460. doi: 10.1007/s11427-020-1637-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Brant AC, Tian W, Majerciak V, Yang W, Zheng ZM. SARS-CoV-2: from its discovery to genome structure, transcription, and replication. Cell Biosci. 2021;11(1):1–7. doi: 10.1186/s13578-021-00643-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A, Diaz G, Cohn A, Fox L, Patel A, Gerber SI, Kim L, Tong S, Lu X, Lindstrom S, Pallansch MA, Weldon WC, Biggs HM, Uyeki TM, Pillai SK, Washington State 2019-nCoV Case Investigation Team First case of 2019 novel coronavirus in the United States. N Engl J Med. 2020;382:929–936. doi: 10.1056/NEJMoa2001191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DKW, Bleicker T, Brunink S, Schneider J, Schmidt ML, Mulders D, Haagmans BL, van der Veer B, van den Brink S, Wijsman L, Goderski G, Romette JL, Ellis J, Zambon M, Peiris M, Goossens H, Reusken C, Koopmans MPG, Drosten C. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25:2000045. doi: 10.2807/1560-7917.ES.2020.25.3.2000045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Center for Health Security, Johns Hopkins Bloomberg School of Public Health. Comparison of national RT-PCR primers, probes, and protocols for SARS-CoV-2 diagnostics. 2020. Accessed at https://www.centerforhealthsecurity.org/resources/COVID-19/COVID-19-fact-sheets/200410-RT-PCR.pdf. Accessed 25 Dec 2021
- 26.Niu P, Lu R, Zhao L, Wang H, Huang B, Ye F, Wang W, Tan W. Three novel real-time RT-PCR assays for detection of COVID-19 virus. China CDC Wkly. 2020;2(25):453. 10.46234/ccdcw2020.116 [DOI] [PMC free article] [PubMed]
- 27.Etievant S, Bal A, Escuret V, et al. Performance assessment of SARS-CoV-2 PCR assays developed by WHO referral laboratories. J Clin Med. 2020;9:1871. doi: 10.3390/jcm9061871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jackson CB, Farzan M, Chen B, Choe H. Mechanisms of SARS-CoV-2 entry into cells. Nat Rev Mol Cell Biol. 2022;23(1):3–20. doi: 10.1038/s41580-021-00418-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lee JH, Choi M, Jung Y, Lee SK, Lee CS, Kim J, Kim J, Kim NH, Kim BT, Kim HG. A novel rapid detection for SARS-CoV-2 spike 1 antigens using human angiotensin converting enzyme 2 (ACE2) Biosens Bioelectron. 2021;1(171):112715. doi: 10.1016/j.bios.2020.112715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Singh A, Steinkellner G, Köchl K, Gruber K, Gruber CC. Serine 477 plays a crucial role in the interaction of the SARS-CoV-2 spike protein with the human receptor ACE2. Sci Rep. 2021;11(1):1–1. doi: 10.1038/s41598-021-83761-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Harvey WT, Carabelli AM, Jackson B, Gupta RK, Thomson EC, Harrison EM, Ludden C, Reeve R, Rambaut A, Peacock SJ, Robertson DL. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol. 2021;19(7):409–24. doi: 10.1038/s41579-021-00573-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Grifoni A, Sidney J, Zhang Y, Scheuermann RH, Peters B, Sette A. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe. 2020;27(4):671–80. doi: 10.1016/j.chom.2020.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chen L, Xiong J, Bao L, Shi Y. Convalescent plasma as a potential therapy for COVID-19. Lancet Infect Dis. 2020;20(4):398–400. doi: 10.1016/S1473-3099(20)30141-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zervou FN, Louie P, Stachel A, Zacharioudakis IM, Ortiz-Mendez Y, Thomas K, Aguero-Rosenfeld ME. SARS-CoV-2 antibodies: IgA correlates with severity of disease in early COVID-19 infection. J Med Virol. 2021 doi: 10.1002/jmv.27058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Xiang J, Yan M, Li H, Liu T, Lin C, Huang S, Shen C. Evaluation of enzyme-linked immunoassay and colloidal gold-immunochromatographic assay kit for detection of novel coronavirus (SARS-Cov-2) causing an outbreak of pneumonia (COVID-19). MedRxiv. 2020;ppmedrxiv-20028787.
- 36.Mattioli IA, Hassan A, Oliveira ON, Jr, Crespilho FN. On the challenges for the diagnosis of SARS-CoV-2 based on a review of current methodologies. ACS Sens. 2020;5(12):3655–77. doi: 10.1021/acssensors.0c01382. [DOI] [PubMed] [Google Scholar]
- 37.Alpdagtas S, Ilhan E, Uysal E, Sengor M, Ustundag CB, Gunduz O. Evaluation of current diagnostic methods for COVID-19. APL Bioeng. 2020;4(4):041506. doi: 10.1063/5.0021554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wu F, Zhao S, Yu B, et al. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579:265–269. doi: 10.1038/s41586-020-2008-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Miller S, Chiu C, Rodino KG, et al. Point-counterpoint: should we be performing metagenomic next-generation sequencing for infectious disease diagnosis in the clinical laboratory? J Clin Microbiol. 2020;58:e01739–19. doi: 10.1128/JCM.01739-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Nairz M, Bellmann-Weiler R, Ladstätter M, Schüllner F, Zimmermann M, Koller AM, Blunder S, Naschberger H, Klotz W, Herold M, Kerndler S. Overcoming limitations in the availability of swabs systems used for SARS-CoV-2 laboratory diagnostics. Sci Rep. 2021;11(1):1–9. doi: 10.1038/s41598-021-81782-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Loeffelholz MJ, Tang YW. Laboratory diagnosis of emerging human coronavirus infections—the state of the art. Emerg Microbes Infect. 2020;9(1):747–56. doi: 10.1080/22221751.2020.1745095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sapkota D, Søland TM, Galtung HK, Sand LP, Giannecchini S, To KK, Mendes-Correa MC, Giglio D, Hasséus B, Braz-Silva PH. COVID-19 salivary signature: diagnostic and research opportunities. J Clin Pathol. 2021;74(6):344–9. doi: 10.1136/jclinpath-2020-206834. [DOI] [PubMed] [Google Scholar]
- 43.Ceron JJ, Lamy E, Martinez-Subiela S, Lopez-Jornet P, Capela-Silva F, Eckersall PD, Tvarijonaviciute A. Use of saliva for diagnosis and monitoring the SARS-CoV-2: a general perspective. J Clin Med. 2020;9(5):1491. doi: 10.3390/jcm9051491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ambrosi C, Prezioso C, Checconi P, Scribano D, Sarshar M, Capannari M, Tomino C, Fini M, Garaci E, Palamara AT, De Chiara G. SARS-CoV-2: comparative analysis of different RNA extraction methods. J Virol Methods. 2021;1(287):114008. doi: 10.1016/j.jviromet.2020.114008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Uhlenhaut C, Cohen JI, Pavletic S, et al. Use of a novel virus detection assay to identify coronavirus HKU1 in the lungs of a hematopoietic stem cell transplant recipient with fatal pneumonia Transpl Infect Dis. 2012;14:79-85. 10.1111/j.1399-3062.2011.00657.x [DOI] [PMC free article] [PubMed]
- 46.Setianingsih TY, Wiyatno A, Hartono TS, et al. Detection of multiple viral sequences in the respiratory tract samples of suspected Middle East respiratory syndrome coronavirus patients in Jakarta, Indonesia 2015–2016. Int J Infect Dis. 2019;86:102–107. doi: 10.1016/j.ijid.2019.06.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tombuloglu H, Sabit H, Al-Suhaimi E, Al Jindan R, Alkharsah KR. Development of multiplex real-time RT-PCR assay for the detection of SARS-CoV-2. PLoS ONE. 2021;16(4):e0250942. doi: 10.1371/journal.pone.0250942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ai T, Yang Z, Hou H, et al. Correlation of chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: a report of 1014 cases. Radiology. 2020;20064210.1148/radiol.2020200642 [DOI] [PMC free article] [PubMed]
- 49.Yu XF, Pan JC, Ye R, Xiang HQ, Kou Y, Huang ZC. Preparation of armored RNA as a control for multiplex real-time reverse transcription-PCR detection of influenza virus and severe acute respiratory syndrome coronavirus. Clin Microbiol. 2008;46(3):837–41. doi: 10.1128/JCM.01904-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lauring AS, Hodcroft EB. Genetic variants of SARS-CoV-2—what do they mean? Jama. 2021;325(6):529–31. doi: 10.1001/jama.2020.27124. [DOI] [PubMed] [Google Scholar]
- 51.Rao SN, Manissero D, Steele VR, Pareja J. A narrative systematic review of the clinical utility of cycle threshold values in the context of COVID-19. Infect Dis Ther. 2020;9(3):573–586. doi: 10.1007/s40121-020-00324-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bustin SA, Mueller R. Real-time reverse transcription PCR (qRT-PCR) and its potential use in clinical diagnosis. Clin Sci (Lond) 2005;109(4):365–79. doi: 10.1042/CS20050086. [DOI] [PubMed] [Google Scholar]
- 53.Joynt GM, Wu WK. Understanding COVID-19: what does viral RNA load really mean? Lancet Infect Dis. 2020;20:635–636. doi: 10.1016/S1473-3099(20)30237-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Greddes L. Puzzle over viral load. New Sci. 2020;245:8. 10.1016/S0262-4079(20)30658-8 [DOI] [PMC free article] [PubMed]
- 55.Kanji JN, Zelyas N, MacDonald C, Pabbaraju K, Khan MN, Prasad A, Hu J, Diggle M, Berenger BM, Tipples G. False negative rate of COVID-19 PCR testing: a discordant testing analysis. Virol J. 2021;18(1):1–6. doi: 10.1186/s12985-021-01489-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zowawi HM, Alenazi TH, AlOmaim WS, Wazzan A, Alsufayan A, Hasanain RA, Aldibasi OS, Althawadi S, Altamimi SA, Mutabagani M, Alamri M. Portable RT-PCR system: a rapid and scalable diagnostic tool for COVID-19 testing. J Clin Microbiol. 2021;59(5):e03004–20. doi: 10.1128/JCM.03004-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Renzoni A, Perez F, Ngo Nsoga MT, Yerly S, Boehm E, Gayet-Ageron A, Kaiser L, Schibler M. Analytical evaluation of Visby medical RT-PCR portable device for rapid detection of SARS-CoV-2. Diagnostics (Basel). 2021;11(5):813. doi: 10.3390/diagnostics11050813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Cojocaru R, Yaseen I, Unrau PJ, Lowe CF, Ritchie G, Romney MG, Sin DD, Gill S, Slyadnev M. Microchip RT-PCR detection of nasopharyngeal SARS-CoV-2 samples. J Mol Diagn. 2021;23(6):683–90. doi: 10.1016/j.jmoldx.2021.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tan C, Fan D, Wang N, Wang F, Wang B, Zhu L, Guo Y. Applications of digital PCR in COVID-19 pandemic. View (Beijing). 2021;2(2):20200082. doi: 10.1002/VIW.20200082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Wong YC, Lau SY, Wang To KK, Mok BW, Li X, Wang P, Deng S, Woo KF, Du Z, Li C, Zhou J. Natural transmission of bat-like severe acute respiratory syndrome coronavirus 2 without proline-arginine-arginine-alanine variants in coronavirus disease 2019 patients. Clin Infect Dis. 2021;73(2):e437–44. doi: 10.1093/cid/ciaa953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Baker M. Digital PCR hits its stride. Nat Methods. 2012;9(6):541–4. doi: 10.1038/nmeth.2027. [DOI] [Google Scholar]
- 62.Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, Vessella RL, Tewari M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 2013;10(10):1003–5. doi: 10.1038/nmeth.2633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Dong L, Zhou J, Niu C, Wang Q, Pan Y, Sheng S, Wang X, Zhang Y, Yang J, Liu M, Zhao Y. Highly accurate and sensitive diagnostic detection of SARS-CoV-2 by digital PCR. Talanta. 2021;1(224):121726. doi: 10.1016/j.talanta.2020.121726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Vasudevan HN, Xu P, Servellita V, Miller S, Liu L, Gopez A, Chiu CY, Abate AR. Digital droplet PCR accurately quantifies SARS-CoV-2 viral load from crude lysate without nucleic acid purification. Sci Rep. 2021;11(1):1–9. doi: 10.1038/s41598-020-80715-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lu R, Wang J, Li M, Wang Y, Dong J, Cai W. SARS-CoV-2 detection using digital PCR for COVID-19 diagnosis, treatment monitoring and criteria for discharge. MedRxiv. 2020. 10.1101/2020.03.24.20042689.
- 66.Rabe BA, Cepko C. SARS-CoV-2 detection using isothermal amplification and a rapid, inexpensive protocol for sample inactivation and purification. Proc Natl Acad Sci U S A. 2020;117(39):24450–8. doi: 10.1073/pnas.2011221117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Notomi T, Okayama H, Masubuchi H, et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000;28:E63. doi: 10.1093/nar/28.12.e63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Piepenburg O, Williams CH, Stemple DL, et al. DNA detection using recombination proteins. PLoS Biol. 2006;4:e204. doi: 10.1371/journal.pbio.0040204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Compton J. Nucleic acid sequence-based amplification. Nature. 1991;350:91–2. doi: 10.1038/350091a0. [DOI] [PubMed] [Google Scholar]
- 70.Dhar BC, Reed AJ, Mitra S, Sanchez PR, Nedorezova DD, Connelly RP, Rohde KH, Gerasimova YV. Cascade of deoxyribozymes for the colorimetric analysis of drug resistance in Mycobacterium tuberculosis. Biosens Bioelectron. 2020;1(165):112385. doi: 10.1016/j.bios.2020.112385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Dean FB, Nelson JR, Giesler TL, et al. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 2001;11:1095–1099. doi: 10.1101/gr.180501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Keightley MC, Sillekens P, Schippers W, et al. Real-time NASBA detection of SARS-associated coronavirus and comparison with real-time reverse transcription-PCR. J Med Virol. 2005;77:602–8. doi: 10.1002/jmv.20498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Jones L, Bakre A, Naikare H, Kolhe R, Sanchez S, Mosley YY, Tripp RA. Isothermal amplification and fluorescent detection of SARS-CoV-2 and SARS-CoV-2 variant virus in nasopharyngeal swabs. PLoS ONE. 2021;16(9):e0257563. doi: 10.1371/journal.pone.0257563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wang WK, Fang CT, Chen HL, et al. Detection of severe acute respiratory syndrome coronavirus RNA in plasma during the course of infection. J Clin Microbiol. 2005;43:962–965. doi: 10.1128/JCM.43.2.962-965.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Yamazaki W, Matsumura Y, Thongchankaew-Seo U, Yamazaki Y, Nagao M. Development of a point-of-care test to detect SARS-CoV-2 from saliva which combines a simple RNA extraction method with colorimetric reverse transcription loop-mediated isothermal amplification detection. J Clin Virol. 2021;1(136):104760. doi: 10.1016/j.jcv.2021.104760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.He Y, Xie T, Tong Y. Rapid and highly sensitive one-tube colorimetric RT-LAMP assay for visual detection of SARS-CoV-2 RNA. Biosens Bioelectron. 2021;1(187):113330. doi: 10.1016/j.bios.2021.113330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Deng H, Jayawardena A, Chan J, Tan SM, Alan T, Kwan P. An ultra-portable, self-contained point-of-care nucleic acid amplification test for diagnosis of active COVID-19 infection. Sci Rep. 2021;11(1):1–2. doi: 10.1038/s41598-021-94652-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Sreejith KR, Umer M, Dirr L, Bailly B, Guillon P, von Itzstein M, Soda N, Kasetsirikul S, Shiddiky MJ, Nguyen NT. A portable device for LAMP based detection of SARS-CoV-2. Micromachines (Basel). 2021;12(10):1151. doi: 10.3390/mi12101151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Ganguli A, Mostafa A, Berger J, Aydin MY, Sun F, de Ramirez SA, Valera E, Cunningham BT, King WP, Bashir R. Rapid isothermal amplification and portable detection system for SARS-CoV-2. Proc Natl Acad Sci U S A. 2020;117(37):22727–35. doi: 10.1073/pnas.2014739117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Toppings NB, Mohon AN, Lee Y, Kumar H, Lee D, Kapoor R, Singh G, Oberding L, Abdullah O, Kim K, Berenger BM. A rapid near-patient detection system for SARS-CoV-2 using saliva. Sci Rep. 2021;11(1):1–9. doi: 10.1038/s41598-021-92677-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Broughton JP, Deng X, Yu G, Fasching CL, Servellita V, Singh J, Miao X, Streithorst JA, Granados A, Sotomayor-Gonzalez A, Zorn K. CRISPR—Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020;38(7):870–4. doi: 10.1038/s41587-020-0513-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Ramachandran A, Huyke DA, Sharma E, Sahoo MK, Huang C, Banaei N, Pinsky BA, Santiago JG. Electric field-driven microfluidics for rapid CRISPR-based diagnostics and its application to detection of SARS-CoV-2. Proc Natl Acad Sci U S A. 2020;117(47):29518–25. doi: 10.1073/pnas.2010254117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Garneret P, Coz E, Martin E, Manuguerra JC, Brient-Litzler E, Enouf V, González Obando DF, Olivo-Marin JC, Monti F, van der Werf S, Vanhomwegen J. Performing point-of-care molecular testing for SARS-CoV-2 with RNA extraction and isothermal amplification. PLoS ONE. 2021;16(1):e0243712. doi: 10.1371/journal.pone.0243712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Chaibun T, Puenpa J, Ngamdee T, Boonapatcharoen N, Athamanolap P, O’Mullane AP, Vongpunsawad S, Poovorawan Y, Lee SY, Lertanantawong B. Rapid electrochemical detection of coronavirus SARS-CoV-2. Nat Commun. 2021;12(1):1. doi: 10.1038/s41467-021-21121-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Kim HE, Schuck A, Lee SH, Lee Y, Kang M, Kim YS. Sensitive electrochemical biosensor combined with isothermal amplification for point-of-care COVID-19 tests. Biosens Bioelectron. 2021;15(182):113168. doi: 10.1016/j.bios.2021.113168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sun Y, Yu L, Liu C, Ye S, Chen W, Li D, Huang W. One-tube SARS-CoV-2 detection platform based on RT-RPA and CRISPR/Cas12a. J Transl Med. 2021;19(1):1. doi: 10.1186/s12967-021-02741-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Chakravarthy A, Anirudh KN, George G, Ranganathan S, Shettigar N, Suchitta U, Palakodeti D, Gulyani A, Ramesh A. Ultrasensitive RNA biosensors for SARS-CoV-2 detection in a simple color and luminescence assay. medRxiv. 2021;4(12):e202101213. 10.26508/lsa.202101213. [DOI] [PMC free article] [PubMed]
- 88.Berber B, Aydin C, Kocabas F, Guney-Esken G, Yilancioglu K, Karadag-Alpaslan M, Caliseki M, Yuce M, Demir S, Tastan C. Gene editing and RNAi approaches for COVID-19 diagnostics and therapeutics. Gene Ther. 2021;28(6):290–305. doi: 10.1038/s41434-020-00209-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Dhar BC, Steimberg N, Mazzoleni G. Point-of-care pathogen detection with CRISPR-based programmable nucleic acid binding proteins. ChemMedChem. 2021;16(10):1566–75. doi: 10.1002/cmdc.202000782. [DOI] [PubMed] [Google Scholar]
- 90.Kellner MJ, Koob JG, Gootenberg JS, et al. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc. 2019;14:2986–3012. doi: 10.1038/s41596-019-0210-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.O’Connell MR. Molecular mechanisms of RNA targeting by Cas13-containing type VI CRISPR-Cas systems. J Mol Biol. 2019;431:66–87. doi: 10.1016/j.jmb.2018.06.029. [DOI] [PubMed] [Google Scholar]
- 92.Zhang F, Abudayyeh OO, Gootenberg JS. A protocol for detection of COVID-19 using CRISPR diagnostics. A protocol for detection of COVID-19 using CRISPR diagnostics. 2020;8.
- 93.Hou T, Zeng W, Yang M, Chen W, Ren L, Ai J, Wu J, Liao Y, Gou X, Li Y, Wang X. Development and evaluation of a rapid CRISPR-based diagnostic for COVID-19. PLoS Pathog. 2020;16(8):e1008705. doi: 10.1371/journal.ppat.1008705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Joung J, Ladha A, Saito M, Segel M, Bruneau R, Huang ML, Kim NG, Yu X, Li J, Walker BD, Greninger AL. Point-of-care testing for COVID-19 using SHERLOCK diagnostics. medRxiv. 2020. 10.1101/2020.05.04.20091231.
- 95.Ooi KH, Liu MM, Tay JW, Teo SY, Kaewsapsak P, Jin S, Lee CK, Hou J, Maurer-Stroh S, Lin W, Yan B. An engineered CRISPR-Cas12a variant and DNA-RNA hybrid guides enable robust and rapid COVID-19 testing. Nat Commun. 2021;12(1):1–23. doi: 10.1038/s41467-021-21996-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Burrell CJ, Howard CR, Murphy FA. Laboratory diagnosis of virus diseases. Fenner White’s Med Virol. 2017:135.
- 97.Monroe D. Novel liposome immunoassays for detecting antigens, antibodies, and haptens. J Liposome Res. 1989;1(3):339–77. doi: 10.3109/08982108909036001. [DOI] [Google Scholar]
- 98.Mohit E, Rostami Z, Vahidi H. A comparative review of immunoassays for COVID-19 detection. Expert Rev Clin Immunol. 2021;17(6):573–599. doi: 10.1080/1744666X.2021.1908886. [DOI] [PubMed] [Google Scholar]
- 99.Laureano AF, Riboldi M. The different tests for the diagnosis of COVID-19—a review in Brazil so far. JBRA Assist Reprod. 2020;24(3):340. doi: 10.5935/1518-0557.20200046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hierholzer JC, Halonen PE, Bingham PG, Coombs RA, Stone YO. Antigen detection in human respiratory coronavirus infections by monoclonal time-resolved fluoroimmunoassay. Clin Diagn Virol. 1994;2(3):165–79. doi: 10.1016/0928-0197(94)90020-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Chatterjee DK, Sitaraman K, Baptista C, Hartley J, Hill TM, Munroe DJ. Protein microarray on-demand: a novel protein microarray system. PLoS ONE. 2008;3(9):e3265. doi: 10.1371/journal.pone.0003265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Masson PL. Particle counting immunoassay—an overview. J Pharm Biomed Anal. 1987;5(2):113–7. doi: 10.1016/0731-7085(87)80013-4. [DOI] [PubMed] [Google Scholar]
- 103.McKie A, Samuel D, Cohen B, Saunders NA. Development of a quantitative immuno-PCR assay and its use to detect mumps-specific IgG in serum. J Immunol Methods. 2002;261(1–2):167–75. doi: 10.1016/s0022-1759(02)00003-0. [DOI] [PubMed] [Google Scholar]
- 104.Gofflot S, El Moualij B, Zorzi D, Melen L, Roels S, Quatpers D, Grassi J, Vanopdenbosch E, Heinen E, Zorzi W. Immuno-quantitative polymerase chain reaction for detection and quantitation of prion protein. J Immunoassay Immunochem. 2004;25(3):241–58. doi: 10.1081/ias-200028044. [DOI] [PubMed] [Google Scholar]
- 105.Ning B, Huang Z, Youngquist BM, Scott JW, Niu A, Bojanowski CM, Zwezdaryk KJ, Saba NS, Fan J, Yin XM, Cao J. Liposome-mediated detection of SARS-CoV-2 RNA-positive extracellular vesicles in plasma. Nat Nanotechnol. 2021;16(9):1039–44. 10.1038/s41565-021-00939-8 [DOI] [PMC free article] [PubMed]
- 106.Gübitz G, Shellum C. Flow-injection immunoassays. Anal Chim Acta. 1993;283(1):421–8. doi: 10.1016/0003-2670(93)85253-G. [DOI] [Google Scholar]
- 107.Infantino M, Grossi V, Lari B, Bambi R, Perri A, Manneschi M, Terenzi G, Liotti I, Ciotta G, Taddei C, Benucci M. Diagnostic accuracy of an automated chemiluminescent immunoassay for anti-SARS-CoV-2 IgM and IgG antibodies: an Italian experience. J Med Virol. 2020;92(9):1671–5. doi: 10.1002/jmv.25932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Ghaffari A, Meurant R, Ardakani A. COVID-19 serological tests: how well do they actually perform? Diagnostics. 2020;10(7):453. doi: 10.3390/diagnostics10070453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Khan S, Nakajima R, Jain A, et al. Analysis of serologic cross-reactivity between common human coronaviruses and SARS-CoV-2 using coronavirus antigen microarray. bioRxiv. 2020;24:006544. 10.1101/2020.03.24.006544
- 110.Bruning AHL, Aatola H, Toivola H, et al. Rapid detection and monitoring of human coronavirus infections. New Microbes New Infect. 2018;9:52–55. doi: 10.1016/j.nmni.2018.04.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Merckx J, Wali R, Schiller I, et al. Diagnostic accuracy of novel and traditional rapid tests for influenza infection compared with reverse transcriptase polymerase chain reaction: a systematic review and meta-analysis. Ann Intern Med. 2017;167:394–409. doi: 10.7326/M17-0848. [DOI] [PubMed] [Google Scholar]
- 112.Bekliz M, Adea K, Essaidi-Laziosi M, Sacks JA, Escadafal C, Kaiser L, Eckerle I. SARS-CoV-2 antigen-detecting rapid tests for the delta variant. Lancet Microbe. 2021 doi: 10.1016/S2666-5247(21)00302-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Mina MJ, Parker R, Larremore DB. Rethinking Covid-19 test sensitivity—a strategy for containment. N Engl J Med. 2020;383(22):e120. doi: 10.1056/NEJMp2025631. [DOI] [PubMed] [Google Scholar]
- 114.Sheridan C. Fast, portable tests come online to curb coronavirus pandemic. Nat Biotechnol. 2020;38:515–518. doi: 10.1038/d41587-020-00010-2. [DOI] [PubMed] [Google Scholar]
- 115.Xiao SY, Wu Y, Liu H. Evolving status of the 2019 novel coronavirus infection: proposal of conventional serologic assays for disease diagnosis and infection monitoring. J Med Virol. 2020;92:464–467. doi: 10.1002/jmv.25702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Guo L, Ren L, Yang S, et al. Profiling early humoral response to diagnose novel coronavirus disease (COVID-19) Clin Infect Dis. 2020 doi: 10.1093/cid/ciaa310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Zhang W, Du RH, Li B, et al. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg Microbes Infect. 2020;9:386–389. doi: 10.1080/22221751.2020.1729071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Okba NMA, Muller MA, Li W, et al. SARS-COV-2 specific antibody responses in COVID-19 patients. Accessed at https://www.medrxiv.org/content/10.1101/2020.03.18.20038059v1 on 15 may 2020.
- 119.Lv H, Wu NC, Tsang OT, et al. Cross-reactive antibody response between SARS-CoV-2 and SARS-CoV infections. Cell Rep. 2020;31:107725. doi: 10.1016/j.celrep.2020.107725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.To KK, Tsang OT, Leung WS, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20:565–574. doi: 10.1016/S1473-3099(20)30196-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Kricka LJ. Microchips, microarrays, biochips and nanochips—personal laboratories for the 21st century. EJIFCC. 2000;12:105–108. doi: 10.1016/s0009-8981(01)00451-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Dhar BC, Cimarelli L, Singh KS, et al. Molecular detection of a potentially toxic diatom species. Int J Environ Res Public Health. 2015;12:4921–4941. doi: 10.3390/ijerph120504921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Shi R, Ma W, Wu Q, et al. Design and application of 60mer oligonucleotide microarray in SARS coronavirus detection. Chin Sci Bull. 2003;48:1165–1169. doi: 10.1007/BF03183928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Guo X, Geng P, Wang Q, et al. Development of a single nucleotide polymorphism DNA microarray for the detection and genotyping of the SARS coronavirus. J Microbiol Biotechnol. 2014;24:1445–1454. doi: 10.4014/jmb.1404.04024. [DOI] [PubMed] [Google Scholar]
- 125.Luna LKD, Heiser Regamey N, et al. Generic detection of coronaviruses and differentiation at the prototype strain level by reverse transcription-PCR and nonfluorescent low-density microarray. J Clin Microbiol. 2007;45:1049–1052. doi: 10.1128/JCM.02426-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Damin F, Galbiati S, Gagliardi S, Cereda C, Dragoni F, Fenizia C, Savasi V, Sola L, Chiari M. CovidArray: a microarray-based assay with high sensitivity for the detection of SARS-CoV-2 in nasopharyngeal swabs. Sensors. 2021;21(7):2490. doi: 10.3390/s21072490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Hardick J, Metzgar D, Risen L, et al. Initial performance evaluation of a spotted array Mobile Analysis Platform (MAP) for the detection of influenza A/B, RSV, and MERS coronavirus Diagn. Microbiol Infect Dis. 2018;91:245–247. doi: 10.1016/j.diagmicrobio.2018.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Yao H, Lu X, Chen Q, et al. Patient-derived mutations impact pathogenicity of SARS-CoV-2. Accessed at https://www.medrxiv.org/content/10.1101/2020.04.14.20060160v2 on 25 May 2020.
- 129.Whitesides GM. The origins and the future of microfluidics. Nature. 2006;442:368–73. doi: 10.1038/nature05058. [DOI] [PubMed] [Google Scholar]
- 130.Wang S, Chinnasamy T, Lifson MA, et al. Flexible substrate-based devices for point-of-care diagnostics. Trends Biotechnol. 2016;34:909–921. 10.1016/j.tibtech.2016.05.009 [DOI] [PMC free article] [PubMed]
- 131.Shafiee H, Asghar W, Inci F, et al. Paper and flexible substrates as materials for biosensing platforms to detect multiple biotargets. Sci Rep. 2015;5:8719. doi: 10.1038/srep08719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Dhar BC, Lee NY. Lab-on-a-chip technology for environmental monitoring of microorganisms. BioChip J. 2018;12:173–183. doi: 10.1007/s13206-018-2301-5. [DOI] [Google Scholar]
- 133.Sachdeva S, Davis RW, Saha AK. Microfluidic point-of-care testing: commercial landscape and future directions. Front Bioeng Biotechnol. 2021;15(8):1537. doi: 10.3389/fbioe.2020.602659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Alsohaimi IH. Analytical detection methods for diagnosis of COVID-19: developed methods and their performance. Biotechnol Biotechnol Equip. 2021;35(1):196–207. doi: 10.1080/13102818.2020.1865838. [DOI] [Google Scholar]
- 135.Antiochia R. Based biosensors: frontiers in point-of-care detection of COVID-19 disease. Biosensors. 2021;11(4):110. doi: 10.3390/bios11040110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Mu HY, Lu YL, Hsiao TH, Huang JH. Microfluidic-based approaches for COVID-19 diagnosis. Biomicrofluidics. 2020;14(6):061504. doi: 10.1063/5.0031406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Grieshaber D, Grieshaber D, MacKenzie R, Vörös J, Reimhult E. Electrochemical biosensors-sensor principles and architectures. Sensors. 2008;8(3):1400–58. doi: 10.3390/s80314000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Kaya SI, Karadurmus L, Ozcelikay G, Bakirhan NK, Ozkan SA. Electrochemical virus detections with nanobiosensors. In Nanosensors for smart cities. 2020 (pp. 303-326). Elsevier. 10.1016/B978-0-12-819870-4.00017-7.
- 139.Saylan Y, Yilmaz F, Özgür E, Derazshamshir A, Yavuz H, Denizli A. Molecular imprinting of macromolecules for sensor applications. Sensors. 2017;17(4):898. doi: 10.3390/s17040898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Kamat S, Kumari M, Jayabaskaran C. Nano-engineered tools in the diagnosis, therapeutics, prevention, and mitigation of SARS-CoV-2. J Control Release. 2021;10(338):813–36. doi: 10.1016/j.jconrel.2021.08.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Cheng MS, Toh C, et al. Novel biosensing methodologies for ultrasensitive detection of viruses. Analyst. 2013;138:6219–6229. doi: 10.1039/c3an01394d. [DOI] [PubMed] [Google Scholar]
- 142.Layqah LA, Eissa S. An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Microchim Acta. 2019;186(4):1. doi: 10.1007/s00604-019-3345-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Alafeef M, Dighe K, Moitra P, Pan D. Rapid, ultrasensitive, and quantitative detection of SARS-CoV-2 using antisense oligonucleotides directed electrochemical biosensor chip. ACS Nano. 2020;14(12):17028–45. doi: 10.1021/acsnano.0c06392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Lin M, Song P, Zhou G, et al. Electrochemical detection of nucleic acids, proteins, small molecules and cells using a DNA-nanostructure-based universal biosensing platform. Nat Protoc. 2016;11:1244–1263. doi: 10.1038/nprot.2016.071. [DOI] [PubMed] [Google Scholar]
- 145.Lim WY, Lan BL, Ramakrishnan N. Emerging biosensors to detect severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2): a review. Biosensors. 2021;11(11):434. doi: 10.3390/bios11110434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Beduk T, Beduk D, de Oliveira Filho JI, Zihnioglu F, Cicek C, Sertoz R, Arda B, Goksel T, Turhan K, Salama KN, Timur S. Rapid point-of-care COVID-19 diagnosis with a gold-nanoarchitecture-assisted laser-scribed graphene biosensor. Anal Chem. 2021;93(24):8585–8594. doi: 10.1021/acs.analchem.1c01444. [DOI] [PubMed] [Google Scholar]
- 147.Laghrib F, Saqrane S, El Bouabi Y, Farahi A, Bakasse M, Lahrich S, El Mhammedi MA. Current progress on COVID-19 related to biosensing technologies: new opportunity for detection and monitoring of viruses. Microchem J. 2021;160:105606. doi: 10.1016/j.microc.2020.105606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Halas NJ, Lal S, Chang W, et al. Plasmons in strongly coupled metallic nanostructures. Chem Rev. 2011;111:3913–3961. doi: 10.1021/cr200061k. [DOI] [PubMed] [Google Scholar]
- 149.Vo-Dinh T, Wang H, Scaffidi J. Plasmonic nanoprobes for SERS biosensing and bioimaging. J. Biophotonics. 2010;3:89–102. doi: 10.1002/jbio.200910015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Cho B, Lee SH, Song J, Bhattacharjee S, Feng J, Hong S, Song M, Kim W, Lee J, Bang D, Wang B. Nanophotonic cell lysis and polymerase chain reaction with gravity-driven cell enrichment for rapid detection of pathogens. ACS Nano. 2019;13(12):13866–74. doi: 10.1021/acsnano.9b04685. [DOI] [PubMed] [Google Scholar]
- 151.Xiao M, Huang L, Dong X, et al. Integration of a 3D-printed read-out platform with a quantum dot-based immunoassay for detection of the avian influenza A (H7N9) virus. Analyst. 2019;144:2594–2603. doi: 10.1039/c8an02336k. [DOI] [PubMed] [Google Scholar]
- 152.Chen X, Zhou Q, Li S, Yan H, Chang B, Wang Y, Dong S. Rapid and visual detection of SARS-CoV-2 using multiplex reverse transcription loop-mediated isothermal amplification linked with gold nanoparticle-based lateral flow biosensor. Front Cell Infect Microbiol. 2021;11:581239. 10.3389/fcimb.2021.581239. [DOI] [PMC free article] [PubMed]
- 153.Products—standard Q COVID-19 Ag http://sdbiosensor.com/xe/product/7672 (accessed May 3, 2020).
- 154.Padoan A, Cosma C, Sciacovelli L, et al. Analytical performances of a chemiluminescence immunoassay for SARS-CoV-2 IgM/IgG and antibody kinetics. Clin Chem Lab Med. 2020;58:1081–1088. doi: 10.1515/cclm-2020-0443. [DOI] [PubMed] [Google Scholar]
- 155.Park TJ, Hyun MS, Lee, Lee HJ, et al. A self-assembled fusion protein-based surface plasmon resonance biosensor for rapid diagnosis of severe acute respiratory syndrome. Talanta. 2009;79:295–301. 10.1016/j.talanta.2009.03.051 [DOI] [PMC free article] [PubMed]
- 156.Murillo AM, Tomé-Amat J, Ramírez Y, Garrido-Arandia M, Valle LG, Hernández-Ramírez G, Tramarin L, Herreros P, Santamaría B, Díaz-Perales A, Holgado M. Developing an optical interferometric detection method based biosensor for detecting specific SARS-CoV-2 immunoglobulins in serum and saliva, and their corresponding ELISA correlation. Sensors Actuators B Chem. 2021;345:130394. doi: 10.1016/j.snb.2021.130394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Asghari A, Wang C, Yoo KM, Rostamian A, Xu X, Shin JD, Dalir H, Chen RT. Fast, accurate, point-of-care COVID-19 pandemic diagnosis enabled through advanced lab-on-chip optical biosensors: opportunities and challenges. Appl Phys Rev. 2021;8(3):031313. doi: 10.1063/5.0022211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Narita F, Wang Z, Kurita H, Li Z, Shi Y, Jia Y, Soutis C. A review of piezoelectric and magnetostrictive biosensor materials for detection of COVID-19 and other viruses. Adv Mater. 2021;33(1):2005448. doi: 10.1002/adma.202005448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Choi JR. Development of point-of-care biosensors for COVID-19. Front chem. 2020;27(8):517. doi: 10.3389/fchem.2020.00517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Choi JR, Nilghaz A, Chen L, Chou KC, Lu X. Modification of thread-based microfluidic device with polysiloxanes for the development of a sensitive and selective immunoassay. Sensors Actuators B Chem. 2018;1(260):1043–51. doi: 10.1016/j.snb.2018.01.102. [DOI] [Google Scholar]
- 161.Rasmi Y, Li X, Khan J, Ozer T, Choi JR. Emerging point-of-care biosensors for rapid diagnosis of COVID-19: current progress, challenges, and future prospects. Anal Bioanal Chem. 2021;18:1–23. doi: 10.1007/s00216-021-03377-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Kumar V, Brent JR, Shorie M, Kaur H, Chadha G, Thomas AG, Lewis EA, Rooney AP, Nguyen L, Zhong XL, Burke MG. Nanostructured aptamer-functionalized black phosphorus sensing platform for label-free detection of myoglobin, a cardiovascular disease biomarker. ACS Appl Mater Interfaces. 2016;8(35):22860–8. doi: 10.1021/acsami.6b06488. [DOI] [PubMed] [Google Scholar]
- 163.Campos EV, Pereira AE, De Oliveira JL, Carvalho LB, Guilger-Casagrande M, De Lima R, Fraceto LF. How can nanotechnology help to combat COVID-19? Opportunities and urgent need. J Nanobiotechnol. 2020;18(1):1–23. doi: 10.1186/s12951-020-00685-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Qiu G, Gai Z, Tao Y, Schmitt J, Kullak-Ublick GA, Wang J. Dual-functional plasmonic photothermal biosensors for highly accurate severe acute respiratory syndrome coronavirus 2 detection. ACS Nano. 2020;14(5):5268–77. doi: 10.1021/acsnano.0c02439. [DOI] [PubMed] [Google Scholar]
- 165.Seo G, Lee G, Kim MJ, Baek SH, Choi M, Ku KB, Lee CS, Jun S, Park D, Kim HG, Kim SJ. Rapid detection of COVID-19 causative virus (SARS-CoV-2) in human nasopharyngeal swab specimens using field-effect transistor-based biosensor. ACS Nano. 2020;14(4):5135–42. doi: 10.1021/acsnano.0c02823. [DOI] [PubMed] [Google Scholar]
- 166.Mavrikou S, Moschopoulou G, Tsekouras V, Kintzios S. Development of a portable, ultra-rapid and ultra-sensitive cell-based biosensor for the direct detection of the SARS-CoV-2 S1 spike protein antigen. Sensors. 2020;20(11):3121. doi: 10.3390/s20113121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Das CM, Guo Y, Yang G, Kang L, Xu G, Ho HP, Yong KT. Gold nanorod assisted enhanced plasmonic detection scheme of COVID-19 SARS-CoV-2 spike protein. Adv Theory Simul. 2020;3(11):2000185. doi: 10.1002/adts.202000185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Büyüksünetçi YT, Çitil BE, Tapan U, Anık Ü. Development and application of a SARS-CoV-2 colorimetric biosensor based on the peroxidase-mimic activity of γ-Fe2O3 nanoparticles. Microchim Acta. 2021;188(10):1–9. doi: 10.1007/s00604-021-04989-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Karakuş E, Erdemir E, Demirbilek N, Liv L. Colorimetric and electrochemical detection of SARS-CoV-2 spike antigen with a gold nanoparticle-based biosensor. Anal Chim Acta. 2021;1182:338939. doi: 10.1016/j.aca.2021.338939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.U.S. Food & Drug Administration. Emergency use authorization of medical products and related authorities. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/emergency-use-authorization-medical-products-and-related-authorities (accessed 2021-01-09)
- 171.Genomeweb. Coronavirus test tracker: commercially available COVID-19 diagnostic tests. https://www.360dx.com/coronavirus-test-tracker-launched-covid-19-tests (accessed 2020-12-19)
- 172.Cue Health leading the fight against COVID-19. https://www.cuehealth.com/what-is-cue/how-cue-detects-covid-19 (accessed 2020-12-19).
- 173.Lucira check it. https://www.lucirahealth.com/ (accessed April 6, 2021).
- 174.Mojsoska B, Larsen S, Olsen DA, Madsen JS, Brandslund I, Alatraktchi FA. Rapid SARS-CoV-2 detection using electrochemical immunosensor. Sensors. 2021;21(2):390. doi: 10.3390/s21020390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.National Institutes of Health. NIH-funded COVID-19 home test is first to receive over-the-counter authorization from FDA. https://www.nih.gov/news-events/news-releases/nih-funded-covid-19-home-test-first-receive-over-counter-authorization-fda (accessed 2021-11-03).
- 176.Abbott. Abbott’s BinaxNOW COVID-19 rapid test receives FDA emergency use authorization for first virtually guided, at-home rapid test using Emed’s digital health platform. https://abbott.mediaroom.com/2020-12-16-Abbotts-BinaxNOW-COVID-19-Rapid-Test-Receives-FDA-Emergency-Use-Authorization-for-First-Virtually-Guided-At-Home-Rapid-Test-Using-eMeds-Digital-Health-Platform (accessed 2021-11-02).
- 177.U.S. Food & Drug Administration Emergency Use Authorization. BinaxNOW COVID-19 antigen self-test. https://www.fda.gov/media/147251/download (accessed 2021-04-05).
- 178.U.S. Food & Drug Administration Emergency Use Authorization. BinaxNOW COVID-19 Ag Card 2 Home Test. https://www.fda.gov/media/147256/download (accessed 2021-04-05).
- 179.U.S. Food & Drug Administration Emergency Use Authorization. QuickVue At-Home OTC COVID-19 Test. https://www.fda.gov/media/147247/download (accessed 2021-11-05).
- 180.U.S. Food & Drug Administration Emergency Use Authorization. QuickVue At-Home COVID-19 Test. https://www.fda.gov/media/146309/download (Accessed 2021-11-05).
- 181.Centers for Disease Control and Prevention. Using antibody tests for COVID-19. https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antibody-tests.html. (accessed 2021-01-29).
- 182.Theel ES, Slev P, Wheeler S, Couturier MR, Wong SJ, Kadkhoda K. The role of antibody testing for SARS-CoV-2: is there one? J Clin Microbiol. 2020;58(8):e00797–20. doi: 10.1128/JCM.00797-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.He X, Lau EHY, Wu P, et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med. 2020;26:672–675. doi: 10.1038/s41591-020-0869-5. [DOI] [PubMed] [Google Scholar]
- 184.Maghdid HS, Ghafoor KZ, Sadiq AS, et al. A novel AI-enabled framework to diagnose coronavirus COVID 19 using smartphone embedded sensors. Accessed at arXiv:2003.07434 on 26 May 2020.
- 185.Apple and Google partner on COVID-19 contact tracing technology. Accessed at https://www.apple.com/newsroom/2020/04/apple-and-google-partner-on-covid-19-contact-tracing-technology/ on 26 May 2020.
- 186.Yang B, Kong J, Fang X. Bandage-like wearable flexible microfluidic recombinase polymerase amplification sensor for the rapid visual detection of nucleic acids. Talanta. 2019;204:685–692. doi: 10.1016/j.talanta.2019.06.031. [DOI] [PubMed] [Google Scholar]
- 187.Oude Munnink BB, Worp N, Nieuwenhuijse DF, Sikkema RS, Haagmans B, Fouchier RA, Koopmans M. The next phase of SARS-CoV-2 surveillance: real-time molecular epidemiology. Nat Med. 2021;27(9):1518–24. doi: 10.1038/s41591-021-01472-w. [DOI] [PubMed] [Google Scholar]
- 188.Arora N, Banerjee AK, Narasu ML. The role of artificial intelligence in tackling COVID-19. Future Virol. 2020;15(11):717–24. doi: 10.2217/fvl-2020-0130. [DOI] [Google Scholar]
- 189.Kipkorir V, Cheruiyot I, Ngure B, Misiani M, Munguti J. Prolonged SARS-CoV-2 RNA detection in anal/rectal swabs and stool specimens in COVID-19 patients after negative conversion in nasopharyngeal RT-PCR test. J Med Virol. 2020;92(11):2328–2331. doi: 10.1002/jmv.26007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Park SM, Won DD, Lee BJ, Escobedo D, Esteva A, Aalipour A, Ge TJ, Kim JH, Suh S, Choi EH, Lozano AX. A mountable toilet system for personalized health monitoring via the analysis of excreta. Nat Biomed Eng. 2020;4(6):624–35. doi: 10.1038/s41551-020-0534-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Zhao Z, Ma Y, Mushtaq A, Rajper AM, Shehab M, Heybourne A, Song W, Ren H. Applications of robotics, artificial intelligence, and digital technologies during COVID-19: A Review. Disaster Med Public Health Prep. 2021;1-11. 10.1017/dmp.2021.9. [DOI] [PMC free article] [PubMed]
- 192.Brief TA. Rapid increase of a SARS-CoV-2 variant with multiple spike protein mutations observed in the United Kingdom. Epidemiology. 2020;7:1–3. [Google Scholar]
- 193.https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/sars-cov-2-viral-mutations-impact-covid-19-tests. Accessed 25 Dec 2021
- 194.Bru D, Martin-Laurent F, Philippot L. Quantification of the detrimental effect of a single primer-template mismatch by real-time PCR using the 16S rRNA gene as an example. Appl Environ Microbiol. 2008;74(5):1660–3. doi: 10.1128/AEM.02403-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.World Health Organization. https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/; May, 2021.
- 196.GISAID [Internet]; https://www.gisaid.org/. Accessed 25 Dec 2021
- 197.Salvatori G, Luberto L, Maffei M, Aurisicchio L, Roscilli G, Palombo F, Marra E. SARS-CoV-2 SPIKE PROTEIN: an optimal immunological target for vaccines. J Transl Med. 2020;18:1–3. doi: 10.1186/s12967-020-02392-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Farkas C, Fuentes-Villalobos F, Garrido JL, Haigh J, Barría MI. Insights on early mutational events in SARS-CoV-2 virus reveal founder effects across geographical regions. PeerJ. 2020;8:e9255. doi: 10.7717/peerj.9255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Koyama T, Platt D, Parida L. Variant analysis of SARS-CoV-2 genomes. Bull World Health Organ. 2020;98(7):495. doi: 10.2471/BLT.20.253591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Ziegler K, Steininger P, Ziegler R, Steinmann J, Korn K, Ensser A. SARS-CoV-2 samples may escape detection because of a single point mutation in the N gene. Eurosurveillance. 2020;25(39):2001650. doi: 10.2807/1560-7917.ES.2020.25.39.2001650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Kaden R. Early phylogenetic diversification of SARS-CoV-2: determination of variants and the effect on epidemiology, immunology, and diagnostics. J Clin Med. 2020;9(6):1615. doi: 10.3390/jcm9061615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Barnes CO, West AP, Jr, Huey-Tubman KE, Hoffmann MA, Sharaf NG, Hoffman PR, Koranda N, Gristick HB, Gaebler C, Muecksch F, Lorenzi JC. Structures of human antibodies bound to SARS-CoV-2 spike reveal common epitopes and recurrent features of antibodies. Cell. 2020;182(4):828–42. doi: 10.1016/j.cell.2020.06.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.European Centre for Disease Prevention and Control. Sequencing of SARS-CoV-2: first update 2021. Available from: https://www.ecdc.europa.eu/sites/default/files/documents/Sequencing-of-SARS-CoV-2-first-update.pdf. Accessed 25 Dec 2021
- 204.World Health Organization. Genomic sequencing of SARS-CoV-2: a guide to implementation for maximum impact on public health 2021. Available from: https://apps.who.int/iris/handle/10665/338480. Accessed 26 Dec 2021
- 205.Université de Genève and Hôpitaux Universitaires de Genève. Protocol for specific RT-PCRs for marker regions of the Spike region indicative of the UK SARS-CoV-2 variant B.1.1.7 and the South African variant 501Y.V2 2021. Available from: https://www.hug.ch/sites/interhug/files/structures/laboratoire_de_virologie/protocol_amplification_voc_20 201 201_uk_geneva.pdf
- 206.Volz E MS, Chand M, Barrett JC, Johnson R, Geidelberg L, et al. Transmission of SARS-CoV-2 lineage B. 1.1. 7 in England: insights from linking epidemiological and genetic data. 2021 [cited 4 January 2021]. Available from: https://www.medrxiv.org/content/10.1101/2020.12.30.20249034v1.full
- 207.Gulay Korukluoglu, Kolukirik M, Bayrakdar F, Ozgumus GG, Altas AB, Cosgun Y, et al. 40 minutes RT-qPCR assay for screening Spike N501Y and HV69-70del mutations 2021. Available from: https://www.biorxiv.org/content/10.1101/2021.01.26.428302v1.full.pdf
- 208.Vogels CB, Breban M, Alpert T, Petrone ME, Watkins AE, Hodcroft E, et al. PCR assay to enhance global surveillance for SARS-CoV-2 variants of concern 2021. 2021.01.28.21250486]. Available from: https://www.medrxiv.org/content/medrxiv/early/2021/02/01/2021.01.28.21250486.full.pdf. Accessed 26 Dec 2021
- 209.Dao Thi VL, Herbst K, Boerner K, Meurer M, Kremer LP, Kirrmaier D, et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Science Translational Medicine. 2020;12(556):eabc7075. [DOI] [PMC free article] [PubMed]
- 210.Liang Y, Lin H, Zou L, Zhao J, Li B, Wang H, Lu J, Sun J, Yang X, Deng X, Tang S. CRISPR-Cas12a-based detection for the major SARS-CoV-2 variants of concern. Microbiol Spectr. 2021;9(3):e01017–21. doi: 10.1128/Spectrum.01017-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.European Centre for Disease Prevention and Control. Options for the use of rapid antigen tests for COVID-19 in the EU/EEA and the UK 2020. Available from: https://www.ecdc.europa.eu/en/publicationsdata/options-use-rapid-antigen-tests-covid-19-eueea-and-uk. Accessed 26 Dec 2021
- 212.Riepler L, Rössler A, Falch A, Volland A, Borena W, von Laer D, Kimpel J. Comparison of four SARS-CoV-2 neutralization assays. Vaccines. 2021;9(1):13. doi: 10.3390/vaccines9010013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Larremore DB, Wilder B, Lester E, Shehata S, Burke JM, Hay JA, Tambe M, Mina MJ, Parker R. Test sensitivity is secondary to frequency and turnaround time for COVID-19 screening. Science advances. 2021;7(1):eabd5393. [DOI] [PMC free article] [PubMed]
- 214.Zhang W, Du RH, Li B, Zheng XS, Yang XL, Hu B, Wang YY, Xiao GF, Yan B, Shi ZL, Zhou P. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg Microbes Infect. 2020;9(1):386–9. doi: 10.1080/22221751.2020.1729071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Bleier BS, Ramanathan M, Jr, Lane AP. COVID-19 vaccines may not prevent nasal SARS-CoV-2 infection and asymptomatic transmission. Otolaryngol Head Neck Surg. 2021;164(2):305–7. doi: 10.1177/0194599820982633. [DOI] [PubMed] [Google Scholar]
- 216.Xi H, Jiang H, Juhas M, Zhang Y. Multiplex biosensing for simultaneous detection of mutations in SARS-CoV-2. ACS Omega. 2021;6(40):25846–59. doi: 10.1021/acsomega.1c04024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Biden Jr JR. National Strategy for the COVID-19 Response and Pandemic Preparedness: January 2021. Simon and Schuster. 2021. https://www.whitehouse.gov/wp-content/uploads/2021/01/National-Strategy-forthe-COVID-19-Response-and-Pandemic-Preparedness.pdf.
- 218.Ellume awarded $231.8M from US DoD, HHS for OTC, at-home COVID-19 test. https://www.360dx.com/immunoassays/ellume-awarded-2318m-us-dod-hhs-otc-home-covid-19-test#.YB2n6HdkgRU%20 (accessed 2021-25-05).
- 219.National Institutes of Health. NIH mobilizes national innovation initiative for COVID-19 diagnostics. https://www.nih.gov/news-events/news-releases/nih-mobilizes-national-innovation-initiative-covid-19-diagnostics (accessed 2021-25-06).
- 220.NIH RADx program awards $29.3M to increase SARS-CoV-2 testing capacity. https://www.360dx.com/covid-19/nih-radx-program-awards-293m-increase-sars-cov-2-testing-capacity?utm_source=Sailthru&utm_medium=email&utm_campaign=360DN%20Tues%202021-04-06&utm_term=360Dx%20Daily%20News#.YGx5umRKjeo (accessed 2021-04-06).
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable for this submission.