Abstract
Background
Vancomycin-resistant Enterococcus faecium (VREfm) strains are one of the most important pathogens causing nosocomial infections in Germany. Due to limited treatment options and an increased risk for acquisition in immunocompromised children, surveillance to monitor occurrence of VREfm in paediatric clinical facilities is of critical importance. Following an unusual accumulation of VREfm positive patients between April 2019 and August 2020 at Dr. von Hauner Children’s Hospital in Munich, Germany, our study aimed to identify dynamics and routes of transmission, and analyse the affected population in view of previously described host risk factors for VREfm colonisation or infection.
Methods
The hospital database was used to collect epidemiological and clinical data of VREfm cases. Descriptive statistical analyses were conducted to outline patient characteristics and depict possible differences between VREfm-colonised and -infected children. An outbreak investigation determining genetic relatedness among VREfm isolates was performed by core genome multilocus sequence typing (cgMLST). To examine potential transmission pathways, results of genome analysis were compared with epidemiological and clinical data of VREfm positive patients.
Results
VREfm acquisition was documented in a total of 33 children (< 18 years). Seven VREfm-colonised patients (21.2%), especially those with a haemato-oncological disease (4/7; p = 0.011), showed signs of clinical infection. cgMLST analysis revealed seven distinct clusters, demonstrating a possible connection within each clonal lineage. Additional eight singletons were identified. Comparison with epidemiological and clinical data provided strong evidence for a link between several VREfm positive patients within the hospital.
Conclusions
A nosocomial spread—at least in part—was the most likely reason for the unusual accumulation of VREfm cases. The study highlights that there is a constant need to increase efforts in hygiene measures, infection control and antibiotic stewardship to combat VREfm transmission events within German paediatric hospitals. Continuous monitoring of adherence to respective policies might reduce the occurrence of clustered cases and prevent future outbreaks.
Keywords: Vancomycin-resistant Enterococcus faecium, Paediatrics, Epidemiology, Outbreak investigation, Nosocomial cluster, Germany
Background
Enterococci are Gram-positive, catalase-negative, facultative anaerobic bacteria that commonly inhabit the gastrointestinal tract of humans and animals. Of all Enterococcus species known to date, Enterococcus faecalis and E. faecium are the most common commensal organisms in humans [1]. Both are characterised by great tenacity to hostile environmental conditions, including high NaCl concentration, bile salts, pH (4.5–10.0) and extreme temperature (5–65 °C), enabling them to persist, grow and spread under a range of stresses [1, 2]. In addition to their role as an essential part of the microflora, E. faecium and E. faecalis are of great medical significance. They are important nosocomial pathogens causing a variety of infections, such as urinary tract and surgical site infections, peritonitis, bacteraemia and in severe cases bloodstream infections and endocarditis [3–6]. A major challenge is the occurrence of intrinsic and acquired antibiotic resistance, which significantly reduces possibilities for therapy. In particular, the glycopeptide resistance genotypes vanA and vanB in vancomycin-resistant E. faecium (VREfm) isolates cause fundamental therapeutic problems [4, 6–8].
Considering the increased risk for persistence and transmission in hospitals, VREfm infections and their treatment play an important role in clinical practice [1, 2, 9–11]. Since the beginning of the twenty-first century, an increased spread of vanA- and vanB-positive E. faecium strains has been detected in German hospitals [5, 12, 13]. This resulted in major outbreaks of VREfm infections and colonisations, which led to a continuous expansion of resistance rates in subsequent years [5, 12–14]. According to a recent analysis of data on E. faecium isolates from the Antibiotic Resistance Surveillance (ARS) of the Robert Koch Institute (RKI), the proportion of existing vancomycin resistance in German hospitals increased significantly from 11.2% in 2014 to 26.1% in 2017 [15]. Due to proven evidence on prolonged hospital stay, higher costs and excess mortality amongst VREfm-colonised and -infected patients, the World Health Organization (WHO) assigned VREfm as a high priority pathogen on its global priority list of antibiotic-resistant bacteria [16–18].
Despite the fact that paediatric facilities are currently not considered classic risk areas for VREfm occurrence, hospitalised children and neonates especially those with severe comorbidities are highly susceptible for VREfm acquisition, colonisation and subsequent infection after contact with these bacteria [19–25]. Possessing immunological naivety and requiring intensive care, they present a fundamentally vulnerable patient group, for whom infections remain an important cause of death [22, 23, 26]. Therefore, it is of great interest to investigate frequent occurrence of VREfm in neonatal, interdisciplinary paediatric and paediatric surgical facilities, examine the affected population and identify spread dynamics. Potential VREfm clusters can thus be detected and current measures for prevention and control of healthcare-associated infections can be reviewed, adapted and improved.
Between April 2019 and August 2020, an unusual accumulation of VREfm cases was observed at Dr. von Hauner Children’s Hospital in Munich, Germany. The aim of the study was to identify or exclude a clonal spread, determine possible nosocomial transmission routes, analyse the affected population in view of previously described host risk factors for VREfm colonisation or infection, give suggestions to improve prevention measures and thereby reduce the rate of future VREfm-colonised and -infected patients at Dr. von Hauner Children’s Hospital.
Methods
Study design and study population
This study was designed as a monocentric, descriptive retrospective analysis investigating data of children aged < 18 years with acquired VREfm isolates between April 2019 and August 2020. The analysis focused on both colonised and infected patients at Dr. von Hauner Children’s Hospital, a 180-bed paediatric tertiary teaching hospital in Munich, Germany. As a part of the Ludwig-Maximilians-University (LMU) Klinikum, it combines general paediatrics and paediatric surgery, provides outpatient care and treats about 7500 inpatient cases every year [27]. Following local proximity and a high number of patient referrals, affected newborns on the neonatal intensive care unit (NICU) in the Polyclinic for Gynecology and Obstetrics (LMU Klinikum Campus Inner City) were included as well. During the study period, the bacteriological laboratory of Dr. von Hauner Children’s Hospital isolated VREfm from 33 patients in total. Cases of VREfm (colonisation or infection) were identified either by a microbiological analysis of rectal swabs examined due to screening for multidrug-resistant pathogens or any other clinical specimen tested for presumed bacterial infection. Routine screening using rectal swabs was performed on NICU and PICU (paediatric intensive care unit) on a weekly basis and on any patient newly admitted to these wards (starting August 2020). Rectal swabs were directly applied to VRE selection agar plates (VRE Select, reference number 63751, Bio-Rad, 85622 Feldkirchen, Germany).
VREfm isolates and cgMLST
VREfm isolates detected at the Dr. von Hauner Children’s Hospital were sent to the German National Reference Centre for Staphylococci and Enterococci at the Robert Koch Institute for further analysis. Antibiotic susceptibility testing was performed by broth microdilution and subsequent determination of the minimum inhibitory concentrations according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines and breakpoints (v10) [28]. Species identification and detection of resistance genes (vanA, vanB) were conducted using standard polymerase chain reaction (PCR) assays. An outbreak investigation determining possible clonal relatedness among the isolates was initiated by whole-genome sequencing (WGS) and typing. For this purpose, DNA derived from pure bacterial culture was isolated using the DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany). Sequencing libraries were generated with the Nextera XT DNA Library Preparation Kit (Illumina, San Diego, CA, United States) and paired-end sequencing was performed using a NextSeq instrument with a read length of 150 bp (Illumina, San Diego, CA, United States). The quality of the raw sequence data was checked using FastQC v0.11.5 [29]. Additionally, Kraken v0.10.6 was used to verify taxonomic read classification [30]. Subsequently, SPAdes v3.12.0 was used in the assembly mode ‘careful’ with default parameters for de novo assembly of high-quality sequencing reads [31]. Multilocus sequence typing (MLST) and core genome MLST (cgMLST) were performed using contigs and Ridom SeqSphere + v6.0.0 (Ridom; Münster, Germany) and established typing schemes [32, 33]. Sequence types (ST) and complex types (CT) were derived from MLST and cgMLST, respectively. Based on cgMLST (including 1423 core genes), a Minimum Spanning Tree (MST) was inferred by ignoring pairwise missing values. VREfm isolates, assigned to the same van-genotype and differing in less than 15 core genes were considered as (closely) related [34].
Data collection and variables
Epidemiological and clinical data of the study population were extracted from electronic and paper-based medical records provided by the hospital database. Variables collected were age at initial detection of VREfm, sex, hospital wards patients were present during the study period and information about whether the patient had a presumed clinical infection or was colonised. Predisposition to known risk factors was identified by literature search and included in the analysis. In addition to multiple patient-related factors such as preterm birth (including young gestational age and low gestational weight), underlying immunosuppressive comorbidity (e.g. malignancy), performed surgical procedures, use of invasive devices (e.g. catheters and feeding or breathing tubes) and invasive treatments (ventilation, chemotherapy), the exposition to antibiotics was recorded [19, 20, 23, 24, 35–38]. Antibiotics prescribed within six months before initial VREfm detection were analysed and categorised into antibiotic classes and Access, Watch, or Reserve groups according to the WHO AWaRe classification of antibiotics [39]. In case of a suspected infection, the respective antibiotic used for treatment was included in the analysis.
Statistical analysis
Statistical analysis was performed using R version 4.0.5 [40]. Distribution of categorical variables in the study population was described in absolute numbers and percentages, continuous variables were illustrated with measures (median, range). Patient characteristics were further analysed regarding suspected VREfm infection (VREfm-I) and VREfm colonisation (VREfm-C). Parameters were compared with Fisher’s exact test for categorical variables and Mann–Whitney U test for continuous variables, as all quantitative data were not normally distributed. The significance level was set at 5%. Missing values were excluded for analysis. A timeline was generated to combine results from cgMLST with epidemiological and clinical data of the study population and thereby investigate potential transmission pathways within the hospital.
Results
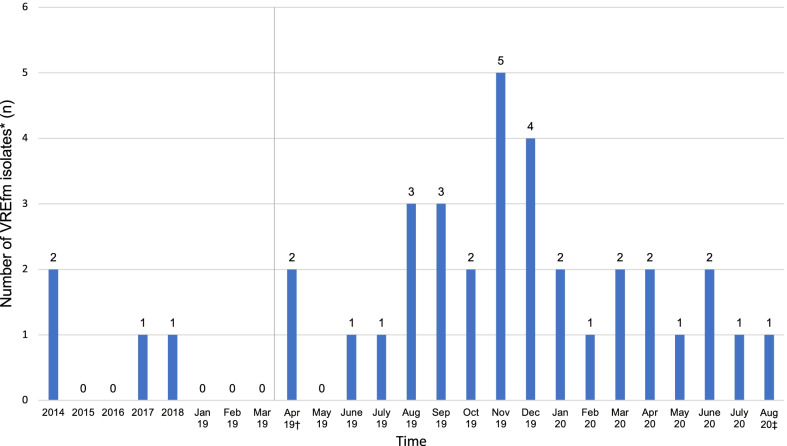
Between April 2019 and August 2020, a total of 693 children were screened for VREfm (154 (22.2%) children in non-ICU settings and 539 (77.8%) while being treated on intensive care units (NICU, PICU)). 33/693 patients were found to be colonised, accounting for a prevalence of 4.8%. Compared to previous years, the number of detected VREfm isolates showed a notable increase during the study period (see Fig. 1). Detailed baseline characteristics of children found to be VREfm positive are outlined in Table 1. Cases predominantly originated from infants with a median age of 6 months (range 0–16 years) and a male/female ratio of 1.54. At the time of VREfm detection, 26 children were treated on neonatal/paediatric intensive care units, four were identified on surgical wards, one on the bone marrow transplantation unit and two were patients cared for on general wards. Antibiotics prescribed within six months prior to detection of VREfm are shown in Table 2.
Fig. 1.
Time trend of detected vancomycin-resistant E. faecium isolates from 2014 to August 2020. †beginning of the study period at Dr. von Hauner Children’s Hospital, ‡end of the study period at Dr. von Hauner Children’s Hospital
Table 1.
Demographic and clinical data of the study population (patients < 18 years)
| Missing n (%) |
All patients (n = 33) |
VREfm-colonised patients (n = 26) |
VREfm-colonised patients with suspected infection (n = 7) |
p value* | |
|---|---|---|---|---|---|
| Sex, n (%) | 0 (0.0) | ||||
| Male | 20 (60.6) | 17 (65.4) | 3 (42.9) | 0.393 | |
| Female | 13 (39.4) | 9 (34.6) | 4 (57.1) | ||
| Age (in months) | 0 (0.0) | ||||
| Median (range) | 6 (0–198) | 5.5 (0–198) | 13 (1–184) | 0.143 | |
| Prematurity, n (%) | 4 (12.1) | ||||
| Preterm born | 16 (48.5) | 14 (53.8) | 2 (28.6) | 0.632 | |
| Mature born | 13 (39.4) | 10 (38.5) | 3 (42.9) | ||
| Gestational age (in completed weeks of gestation) | 4 (12.1) | ||||
| Median (range) | 34 (23–41) | 33.5 (23–41) | 34 (24–40) | 0.885 | |
| Gestational weight (in grams) | 8 (24.2) | ||||
|
Median (range) |
2150 (465–4430) |
2150 (465–4430) |
1665 (730–2600) |
0.96 | |
| Twin, n (%) | 0 (0.0) | ||||
| Yes | 4 (12.1) | 4 (15.4) | 0 (0.0) | 0.555 | |
| Invasive devices†, n (%) | 0 (0.0) | ||||
| NG/NJ tube | 16 (48.5) | 13 (50.0) | 3 (42.9) | 1.000 | |
| PEG/PEJ tube | 7 (21.2) | 6 (23.1) | 1 (14.3) | 1.000 | |
| Peripheral venous catheter | 21 (63.6) | 15 (57.7) | 6 (85.7) | 0.223 | |
| Central venous catheter | 17 (51.5) | 13 (50.0) | 4 (57.1) | 1.000 | |
| Arterial line | 9 (27.3) | 6 (23.1) | 3 (42.9) | 0.358 | |
| Port | 1 (3.0) | 1 (3.8) | 0 (0.0) | 1.000 | |
| Hickman catheter | 5 (15.2) | 2 (7.7) | 3 (42.9) | 0.052 | |
| Urinary catheter | 10 (30.3) | 6 (23.1) | 4 (57.1) | 0.161 | |
| Ventricular drain | 3 (9.1) | 3 (11.5) | 0 (0.0) | 1.000 | |
| Tracheostomy tube | 2 (6.1) | 2 (7.7) | 0 (0.0) | 1.000 | |
| Chest drain | 2 (6.1) | 2 (7.7) | 0 (0.0) | 1.000 | |
| Artificial stoma | 3 (9.1) | 0 (0.0) | 3 (42.9) | 0.006 | |
| Surgical procedures†, n (%) | 0 (0.0) | 18 (54.5) | 11 (42.3) | 7 (100.0) | 0.009 |
| Endoscopic procedure | 8 (24.2) | 7 (26.9) | 1 (14.3) | 0.652 | |
| Cardiothoracic surgery | 4 (12.1) | 3 (11.5) | 1 (14.3) | 1.000 | |
| Intrabdominal surgery | 5 (15.2) | 0 (0.0) | 5 (71.4) | < 0.0001 | |
| Brain surgery | 2 (6.1) | 2 (7.7) | 0 (0.0) | 1.000 | |
| Biopsy | 2 (6.1) | 1 (3.8) | 1 (14.3) | 0.385 | |
| Bone marrow aspiration | 1 (3.0) | 0 (0.0) | 1 (14.3) | 0.212 | |
| Tumour resection | 3 (9.1) | 0 (0.0) | 3 (42.9) | 0.006 | |
| Other surgical procedures‡ | 2 (6.1) | 1 (3.8) | 1 (14.3) | 0.385 | |
| Underlying diseases, n (%) | 0 (0.0) | ||||
| Haemato-oncological diseases | 6 (18.2) | 2 (7.7) | 4 (57.1) | 0.011 | |
| Cardiovascular diseases | 8 (24.2) | 6 (23.1) | 2 (28.6) | 1.000 | |
| Diseases of the respiratory system | 13 (39.4) | 13 (50.0) | 0 (0.0) | 0.027 | |
| Endocrine diseases | 3 (9.1) | 3 (11.5) | 0 (0.0) | 1.000 | |
| Gastrointestinal diseases | 15 (45.5) | 11 (42.3) | 4 (57.1) | 0.674 | |
| Genitourinary diseases | 3 (9.1) | 1 (3.8) | 2 (28.6) | 0.107 | |
| Neurological diseases | 9 (27.3) | 8 (30.8) | 1 (14.3) | 0.642 | |
| Malformation syndromes affecting multiple systems | 3 (9.1) | 2 (7.7) | 1 (14.3) | 0.524 | |
| Chromosomal abnormalities | 3 (9.1) | 3 (11.5) | 0 (0.0) | 1.000 | |
| Ventilation†, n (%) | 0 (0.0) | ||||
| Invasive ventilation | 14 (42.4) | 10 (38.5) | 4 (57.1) | 0.422 | |
| Non-invasive ventilation | 17 (51.5) | 13 (50.0) | 4 (57.1) | 1.000 | |
| Chemotherapy⁓, n (%) | 0 (0.0) | ||||
| Yes | 4 (12.1) | 1 (3.8) | 3 (42.9) | 0.023 | |
| Reanimation⁓, n (%) | 0 (0.0) | ||||
| Yes | 5 (15.2) | 3 (11.5) | 2 (28.6) | 0.282 | |
| Overall hospitalisation before initial detection of VREfm° | 0 (0.0) | ||||
| Length of stay (in days), median (range) | 38 (0–276) | 38 (0–276) | 28 (0–106) | 0.659 | |
| Number of hospital admissions, median (range) | 1 (1–34) | 1 (1–34) | 1 (1–15) | 0.484 | |
| Glycopeptide resistance genotype, n (%) | 0 (0.0) | ||||
| vanA | 15 (45.5) | 14 (53.8) | 1 (14.3) | 0.095 | |
| vanB | 18 (54.5) | 12 (46.2) | 6 (85.7) |
Signficant values (marked in bold) were defined as p < 0.05
VREfm, Vancomycin-resistant Enterococcus faecium; NG/NJ tube, nasogastric/nasojejunal tube; PEG/PEJ tube, percutaneous endoscopic gastrostomy/jejunostomy tube
*Fisher’s exact test or Mann–Whitney U test (p 0.05 was considered significant)
†Within four weeks prior to detection of VREfm
‡Including one surgery of the anus, rectum and colon and one ovarian surgery for fertility preservation
⁓Within six months prior to detection of VREfm
°Including hospitalisation at Dr. von Hauner Children’s Hospital, NICU Clinic and Polyclinic for Gynecology and Obstetrics and LMU Klinikum Großhadern
Table 2.
Antibiotic use within six months prior to detection of VREfm in the study population
| All patients (n = 33) |
VREfm-colonised patients (n = 26) |
VREfm-colonised patients with suspected infection (n = 7) |
p value* | |
|---|---|---|---|---|
| Antibiotics total, median (range) | 5 (0–14) | 5 (0–14) | 7 (3–14) | 0.150 |
| Antibiotic classes, n (%) | ||||
| Aminoglycosides | 5 (15.2) | 5 (19.2) | 0 (0.0) | 0.559 |
| Beta-lactam/beta-lactamase inhibitor | 22 (66.7) | 16 (61.5) | 6 (85.7) | 0.378 |
| Carbapenems | 16 (48.5) | 10 (38.5) | 6 (85.7) | 0.039 |
| First-generation cephalosporins | 3 (9.1) | 3 (11.5) | 0 (0.0) | 1.000 |
| Fluoroquinolones | 1 (3.0) | 0 (0.0) | 1 (14.3) | 0.212 |
| Glycopeptides | 11 (33.3) | 9 (34.6) | 2 (28.6) | 1.000 |
| Imidazoles | 4 (12.1) | 3 (11.5) | 1 (14.3) | 1.000 |
| Macrolides | 6 (18.2) | 6 (23.1) | 0 (0.0) | 0.301 |
| Penicillins | 12 (36.4) | 11 (42.3) | 1 (14.3) | 0.223 |
| Phosphonics | 1 (3.0) | 0 (0.0) | 1 (14.3) | 0.212 |
| Second-generation cephalosporins | 8 (24.2) | 7 (26.9) | 1 (14.3) | 0.652 |
| Third-generation cephalosporins | 14 (42.4) | 13 (50.0) | 1 (14.3) | 0.195 |
| Trimethoprim/sulfonamide combinations | 9 (27.3) | 5 (19.2) | 4 (57.1) | 0.068 |
| Unknown antibiotic class | 2 (6.1) | 2 (7.7) | 0 (0.0) | 1.000 |
| Antibiotic groups AWaRe classification† | ||||
| Access Antibiotics, median (range) | 1 (0–6) | 1 (0–6) | 1 (0–3) | 0.629 |
| Watch Antibiotics, median (range) | 4 (0–13) | 3 (0–13) | 4 (2–12) | 0.120 |
Signficant values (marked in bold) were defined as p < 0.05
VREfm, Vancomycin-resistant Enterococcus faecium
*Fisher’s exact test or Mann–Whitney U test (p < 0.05 was considered significant)
†excluding two unknown antibiotics
A clinical VREfm infection was presumed in seven patients (21.2%), all of whom were previously colonised with the respective VREfm strain. Central-line associated infection was detected in three patients. Two patients suffered a surgical wound infection, one patient was diagnosed with a urinary tract infection and another one showed positive blood cultures, suggesting an invasive systemic infection. Demographic and clinical findings from patients with suspected VREfm-I and VREfm-C are presented in Table 1. Our data demonstrate that children who developed signs of a bacterial infection were significantly more likely to have had a temporary artificial stoma (p = 0.006), to have undergone a recent surgical procedure (p = 0.009) or to have received carbapenems (p = 0.039) or chemotherapy (p = 0.023) before initial VREfm detection. Similarly, the presence of an underlying haemato-oncological diagnosis (p = 0.011) was significantly more likely in VREfm-infected compared to VREfm-colonised children. An underlying respiratory disease (p = 0.027) was significantly less likely in patients with a presumed VREfm-I than in those without clinical symptoms.
All suspected VREfm infections were treated with Reserve antibiotics. Six patients received linezolid, one patient received daptomycin and another child was treated with a linezolid/daptomycin combination.
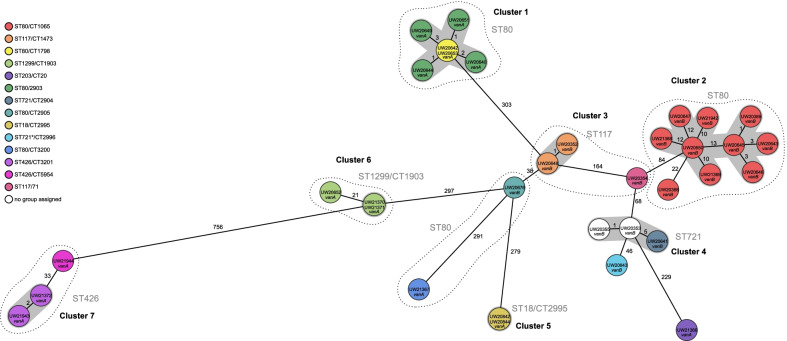
Genome analysis (MLST) showed that VREfm isolates belonged to seven different STs. The most commonly detected STs were ST80 (n = 18), ST721 (n = 4), ST117 (n = 3), ST424 (n = 3) and ST1299 (n = 3). Regarding detection of resistance genes, PCR analysis identified vanB cluster in 54.5% (n = 18) and vanA cluster in 45.5% of isolates (n = 15). More detailed cgMLST analysis revealed genetic links, divided into seven distinct clusters (allele difference ≤ 15, Fig. 2). The clonal lineage ST80/CT1065 vanB represented the largest group, containing almost one third (n = 10) of all VREfm isolates. The combination of cgMLST with epidemiological and clinical data of the study population is shown in Fig. 3.
Fig. 2.
Minimum spanning tree of 34 vancomycin-resistant E. faecium isolates from 33 children and one mother. The number of varying alleles is shown next to the black lines. Colouring was based on cgMLST (analysis of 1423 genes of the nuclear genome). Grey areas connect isolates that belong to a cluster based on the definitions of SeqSphere + (maximum allele difference = 15 alleles). Dashed lines include isolates that belong to one sequence type (ST). ST = sequence type (multilocus sequence typing (MLST), analysis of 7 housekeeping genes), CT = complex type (core genome MLST (cgMLST)). SeqSphere option: pairwise ignoring missing values. Task templates: E. faecium cgMLST v1.1; E. faecium MLST v1.0. *adk: Novel allele, ST may indicate nearest ST. MLST-Finder: http://www.genomicepidemiolog
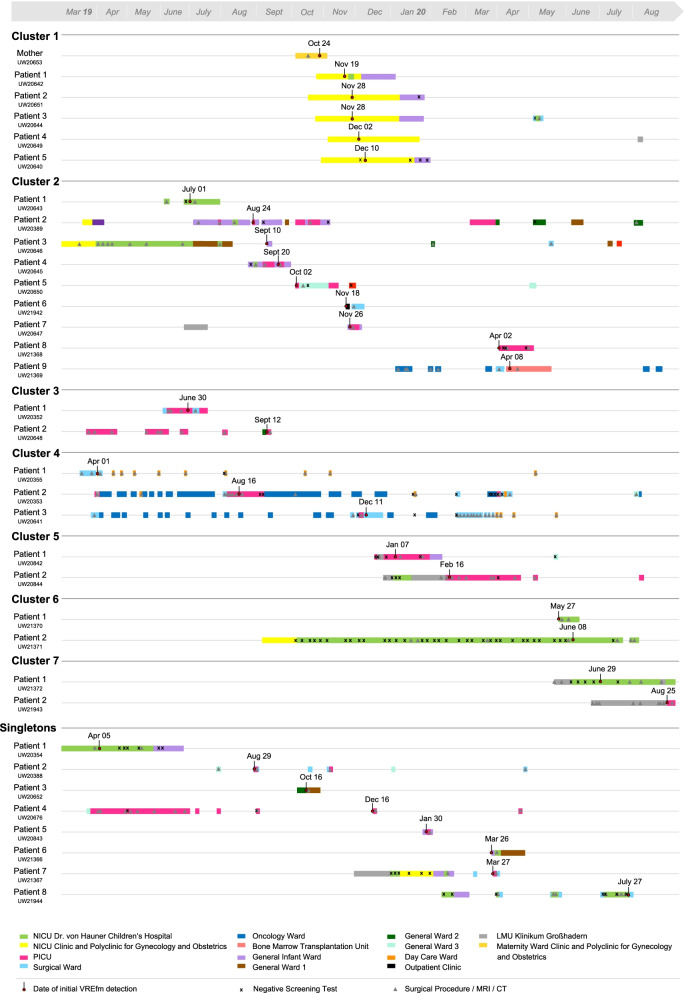
Fig. 3.
Timeline for each patient combining results from cgMLST with epidemiological and clinical data
Cluster 1 was found at the Polyclinic for Gynecology and Obstetrics. It contained five premature babies and a mother, who had been hospitalised due to complications with twin pregnancy. The mother (corresponding VREfm isolate ID UW20653, see Fig. 2 and 3), screened positive for VREfm on 24 October 2019, representing the initial case of this cluster. The first newborn (UW20642) was found to be colonised on 19 November, followed by both twins of the initial case (UW20651, UW20644) on 28 November. Subsequently, two additional patients (UW20649, UW20640) were screened positive beginning of December. All newborns were inpatients during the same time.
Cluster 2 contained nine different isolates, detected within a period of ten months. Regarding the clinical history of affected patients, four children (patients 1, 4, 5, 8; UW20643, UW20645, UW20650, UW21368) had already been tested positive for VREfm during a previous external hospitalisation. After admission to Dr. von Hauner Children’s Hospital, patient 1 was an inpatient on NICU during the same time (June 2019) as patient 3 (UW20646), patient 2 (UW20389) was an inpatient on the general infant ward during the same time (August/September 2019) as patient 4 and patients 2 and 3 both had a gastroscopy on the same day (29 July 2019) performed by the same surgical team. Apart from two negative test results from patients 1 and 4, no screening tests were performed before initial VREfm detection.
Cluster 3 occurred on PICU. The first colonised patient (UW20352) was a child with a malignant solid tumour, who was transferred to Munich for elective surgical treatment in June 2019. Seventy-four days after the primal case a second patient (UW20648) was screened positive for VREfm. In both cases, no screening tests were performed before initial detection of the respective bacterial strain.
Another three isolates, identified in 2019 within a period of nine months, built Cluster 4. All three affected children had an underlying oncological disease but were present on different wards when they were first screened VREfm positive. Except for one negative test result (patient 3; UW20641), no screening tests were performed before initial detection. Patients 1 (UW20355) and 3 were hospitalised at the same surgical ward end of March/beginning of April and patient 2 (UW20353) was admitted several times to the oncology ward for chemotherapy during the same period as patient 3. In addition, patients 1 and 2 had a surgical procedure one day apart (01 April and 31 March) performed by the same surgeon.
Cluster 5 was detected on PICU in January/February 2020. Two patients (UW20842, UW20844) were screened positive for VREfm within 40 days. Before initial detection the patients were screened negative several times, however, patient 2 was temporarily hospitalised at another site of the LMU Klinikum, where no data for potential screening tests were available.
Further two isolates formed Cluster 6. The initial case (UW21370) in this cluster was an infant screened positive for VREfm on first day of admission (27 May 2020) to the NICU of Dr. von Hauner Children’s Hospital. A second patient (UW21371) was found to be colonised on 08 June, located at the same hospital ward during the same time as the initial case. Before detection of VREfm, patient 2 was screened negative several times.
Cluster 7 included two isolates. The first patient (UW21372) with detected VREfm on 29 June 2020, was a child suffering from cardiac defect and severe postischemic brain injury who was transferred from another site of the LMU Klinikum to the NICU of Dr. von Hauner Children’s Hospital for further treatment. Before initial detection, the patient was tested negative several times. Patient 2 (UW21943) was screened positive on first day of admission to PICU on 25 August 2020. Regarding clinical history, patient 2 had evidence for VREfm colonisation during a previous hospital stay (10 August 2020) at another site of the LMU Klinikum, where patient 1 had been present two months before. In addition, patient 1 had a temporal overlap regarding hospital ward with both patients from Cluster 6.
Eight remaining isolates were considered singletons (a single clone that has no close relatives in the cgMLST). However, clinical-epidemiological data revealed evidence for a possible link between several cases (Fig. 2). The hospitalisation of patient 1 (UW20354) was congruent with the hospitalisation of patient 3 (UW20646) from Cluster 2. Patient 4 (UW20676) was present on PICU during the same time as both patients (UW20352, UW20648) from Cluster 3 and patients 7 (UW21367) and 8 (UW21944) showed an epidemiological connection in terms of hospital wards with children (UW20844, UW21370, UW21371, UW21372) from Cluster 5–7. A temporal and personnel concordance in several performed surgical procedures presented another potential connection for some patients with no cluster assignment.
Discussion
In this retrospective study, we aimed to investigate the accumulation of VREfm isolates during April 2019 and August 2020 at Dr. von Hauner Children’s Hospital. The main objective was to investigate a clonal spread, determine possible nosocomial transmission routes and analyse the affected population while in parallel evaluating previously described host risk factors for VREfm carriage or infection. In total, we found 33 children to be VREfm positive during the study period. CgMLST of all isolates revealed a polyclonal structure with a suspected spread demonstrated within seven distinct clusters and eight singletons. In combination with epidemiological and clinical data, our observations provided compelling evidence for transmission of VREfm between patients within the hospital.
Cluster 1, 3, 5, 6 and 7 consisted of isolates from children, who were present on identical hospital wards either during the same time or during a period following shortly thereafter (maximum seven weeks). These findings suggest a likely nosocomial transmission—a frequent and relevant issue that has been described in particular for VREfm compared to other multidrug-resistant microorganisms [41]. This fits well with the current state of research, where direct transmission between colonised or infected patients as well as indirect transmission via the patient’s surroundings are discussed as the most probable routes of spread [2, 10, 11, 42, 43]. Regarding the duration of VREfm-C in paediatric patients, periods ranging from several weeks to over six months are described [44–47]. Due to their extensive resilience, enterococci are known to survive even longer (up to several years) in hospital environments [1, 2]. Drees et al. found that patients admitted to rooms previously occupied by VRE carriers had a significantly higher risk of VRE acquisition [48]. Contaminated drip stands, ventilation tubes, incubators, thermometers, and other VRE positive surfaces were confirmed to play an important role in transmission pathways and multiple cleaning practices were shown to be inefficient for their decontamination [49–53]. In addition to transmission dynamics via the environment, these findings could be the result of close and constant interaction between patients and healthcare staff. Especially hands or gloves have been shown to act as a potential reservoir and transmission vehicle for nosocomial bacteria [9, 54, 55]. Moreover, we have shown that Cluster 4 included VREfm positive patients, detected at different hospital wards. However, all three children had an underlying haemato-oncological disease and two patients had undergone a surgical procedure with the same surgical team one day apart from each other. These results indicate potential transmission via nursing staff or attending physicians in oncology or contamination of the hospital environment in general. Cluster 2, containing isolates from nine patients, showed a poor clinical-epidemiological linkage. Multiple children had been tested positive for VREfm during a previous external hospitalisation or had a positive test result on their first day of admission, assuming that they had already been colonised before initial detection at Dr. von Hauner Children’s Hospital. Nevertheless, we found some temporal overlap in terms of hospital wards or performed surgical procedures, again suggesting nosocomial transmission of VREfm.
Overall, our findings are in line with other reports, confirming VREfm transmission within and between wards by WGS/cgMLST and epidemiological data [10, 33, 56–58]. An inter-hospital spread can be assumed since the predominant clonal lineage in Bavaria ST80/CT1065 vanB represented the most commonly detected group (n = 10) at Dr. von Hauner Children’s Hospital [59]. Especially isolates of Cluster 2 harboured the ST80/CT1065 vanB group, which may explain the poor clinical-epidemiological linkage and thus indicate a cross-contamination event with external hospitals. Frequent patient referrals between hospitals and specialities, in particular within different sites of the LMU Klinikum, may demonstrate a reason for the dissemination.
Interestingly, some of the genetically unrelated VREfm isolates showed a relevant connection in terms of epidemiological and clinical data of affected patients (patients of Cluster 6 and Cluster 7; patients of the singleton group and Cluster 2, 3, 5, 6 or 7). In each case, VREfm isolates harboured van-genotypes, which were identical with genotypes of possible connected clusters (Fig. 2). This could refer to genetic mobility of vanA and vanB variants, allowing resistance to spread among different clonal lineages by horizontal gene transfer (HGT) [8, 60, 61]. Arredondo-Alonso et al. as well as Pinholt et al. previously identified high frequencies in HGT, especially of the vanA transposon Tn1546 and corresponding vanA plasmids among unrelated E. faecium isolates as an alternative route of vancomycin resistance transmission in hospitals [62, 63].
To improve prevailing measures for the prevention of nosocomial VREfm spread at Dr. von Hauner Children’s Hospital, we evaluated affected patients, especially those with a presumed clinical infection requiring antibiotic therapy. Thereby, we have shown that more than half of all VREfm positive children were premature babies with young gestational age and low gestational weight. Regarding possible differences between VREfm-colonised and VREfm-infected patients, the presence of a temporary artificial stoma, a recent surgical procedure, previous treatment with carbapenems, preceding chemotherapy and underlying haemato-oncological disease were significantly associated with the development of clinical symptoms. These findings are in line with current studies, identifying preterm babies as well as immunosuppressed paediatric intensive care patients—in particular children with a haematological/oncological diagnosis—as a high-risk population for VREfm-C and VREfm-I [19, 20, 35, 64–67]. High exposure to antibiotics, especially third-generation cephalosporins, seems to further increase the risk [19, 35, 68].
Some important limitations need to be taken into account when interpreting our results. First, our study lacked negative test results in some patients prior to initial VREfm detection making it difficult to determine the exact time point of VREfm acquisition. This temporal imprecision may have influenced the accuracy of epidemiological data. Furthermore, we neither had information on patients’ room numbers or bays on NICU/PICU (to identify direct roommates) nor were samples of the patients’ environment available. Therefore, we were only able to make statements on ward level. Transmission via environmental contamination or healthcare staff (hands/gloves) could only be assumed, not confirmed. Second, it must be noted that clonal lineages such as ST80/CT1065 seem to have a very stable core genome (low allele differences), which often leads to the formation of clusters in cgMLST with no epidemiological link (neither temporally nor spatially) [59]. Third, focusing on cgMLST may have led to miss the confirmation of potential epidemiological links as a result of an overestimation of non-related isolates by excluding the analysis of possible HGT [62]. A polyclonal VREfm colonisation (also as a result of HGT) is conceivable, as usually only one colony (1 clone) per patient and time point is microbiologically processed, which does not necessarily reflect the totality of possible colonisation.
However, our findings regarding VREfm spread are still relevant and valuable. Future efforts should and will aim on exploring new and better ways to reduce nosocomial transmission events at Dr. von Hauner Children’s Hospital. In essence, the majority of existing international recommendations on the prevention of VREfm-C and VREfm-I in hospital call for improved hygiene measures, educational activities and screening as key interventions in suspected outbreak scenarios in clinical settings [69–74]. Following this study’s outbreak investigation it is essential to continuously review adherence to basic (hand) hygiene measures, in particular to the implementation of the “Five Moments for Hand Hygiene” [69–73]. Furthermore, we will aim to establish a new action plan consisting of a prevention bundle tailored to our affected population. Considering our findings we conclude that the bundle should include an active screening of rectal swabs for high-risk patients namely children and infants on intensive care units, surgical wards, the oncology ward and the bone marrow transplantation unit as well as a passive screening of every specimen taken for clinical indication [69–72, 74]. Routine surveillance of women with a high-risk pregnancy and high prenatal antibiotic use hospitalised at the Clinic and Polyclinic for Gynecology and Obstetrics should be considered. To achieve a reduction in clonal spread and horizontal transmission events, screening should be performed at regular intervals beginning on the first day of hospital admission [74, 75]. A subsequent isolation strategy for every VRE carrier including individual sanitary facilities for older children and mothers as well as enhanced barrier measures (gowns/gloves) for all contact persons constitute meaningful measures to reduce nosocomial VREfm dissemination [69–72, 75–77]. Improved cleaning and disinfection methods during and after hospitalisation of VREfm carriers and the involvement of affected patients and accompanying persons in hygiene measures can magnify the effect [50, 70–72, 74, 78]. Complementary to our current standard hygiene measures (disinfectant: Kohrsolin FF 0.5% or Terralin protect 0.5%) we consider to use ultraviolet-C (UV-C) light as an additional method to enhance terminal disinfection of patient rooms [79]. UV-C radiation has been used after detection of Cluster 1 at the Clinic and Polyclinic for Gynecology and Obstetrics. In general, it has been shown that efforts to reduce the use of unnecessary antibiotics are key to avoid selection of multidrug-resistant pathogens [80]. Despite no consistent scientific consensus about the impact of antibiotic stewardship programs (ASP) on VRE acquisition in general, an implementation in paediatric patients seems to be promising [69, 81, 82]. Further studies should regularly monitor the effectiveness of infection control measures and adherence to respective policies using defined suitable target variables. Finally, refinements to the examination of genomic data by a new approach that also includes the analyses of HGT mobilisation and polyclonal colonisation to effectively confirm potential epidemiological links may provide more accurate results for surveillance [62, 83].
Conclusions
In conclusion, the Dr. von Hauner Children’s Hospital witnessed a substantial increase in the detection of VREfm isolates between April 2019 and August 2020, a dynamic that can be—at least in part—attributed to suggested nosocomial transmission events. Our study highlights the importance of protecting intensive care patients, who were mainly affected by the outbreak. In view of the monocentric character of this study, results may not entirely be generalisable to other clinical settings. However, findings and conclusions can serve as an example for comparable paediatric tertiary teaching hospitals. To achieve a reduction of transmission it is critical to further investigate VREfm genetic profiles and epidemiological links between colonised/infected patients, hospital environment and healthcare staff. Additional prospective studies are needed to continuously improve preventive efforts in hygiene measures, infection control and ASP to combat the spread of VREfm between hospitalised children and infants in Germany.
Acknowledgements
We thank the microbiological laboratory team at Dr. von Hauner Children’s Hospital for technical assistance and all doctors and nurses providing clinical care to the patients included in this retrospective study.
Abbreviations
- ASP
Antibiotic stewardship program
- AWaRe
Access Watch Reserve
- cgMLST
Core genome multilocus sequence typing
- CT
Complex type
- E. faecalis
Enterococcus faecalis
- E. faecium
Enterococcus faecium
- EUCAST
European Committee on Antimicrobial Susceptibility Testing
- HGT
Horizontal gene transfer
- LMU
Ludwig-Maximilians-University
- MLST
Multilocus sequence typing
- MST
Minimum Spanning Tree
- NICU
Neonatal intensive care unit
- PCR
Polymerase chain reaction
- PICU
Paediatric intensive care unit
- RKI
Robert Koch Institute
- ST
Sequence type
- UV-C
Ultraviolet-C
- VRE
Vancomycin-resistant enterococci
- VREfm
Vancomycin-resistant Enterococcus faecium
- VREfm-C
VREfm colonisation
- VREfm-I
VREfm infection
- WGS
Whole-genome sequencing
- WHO
World Health Organization
Authors' contributions
IT, LK, MM, JH and UvB contributed to the conception and design of the study. GW and RW performed the genome analysis of VREfm isolates. IT and MM were responsible for data acquisition. IT, LK and UvB analysed and interpreted the results. IT, LK and UvB drafted and/or revised the manuscript. VH and SS were involved in clinical care of patients and provided additional clinical input. All authors read and approved the final manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL. There has been no specific funding for this study.
Availability of data and materials
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
This study was approved by the Ludwig-Maximilians-University ethics committee under project number 21–0334. Due to the retrospective character of the study, no informed consent was required. The processing of personal data was anonymised and in full accordance with local data protection laws.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Fisher K, Phillips C. The ecology, epidemiology and virulence of Enterococcus. Microbiology. 2009;155:1749–1757. doi: 10.1099/mic.0.026385-0. [DOI] [PubMed] [Google Scholar]
- 2.Suleyman G, Alangaden G, Bardossy AC. The role of environmental contamination in the transmission of nosocomial pathogens and healthcare-associated infections. Curr Infect Dis Rep. 2018;20:12. doi: 10.1007/s11908-018-0620-2. [DOI] [PubMed] [Google Scholar]
- 3.Hidron AI, Edwards JR, Patel J, Horan TC, Sievert DM, Pollock DA, et al. NHSN annual update: antimicrobial-resistant pathogens associated with healthcare-associated infections: annual summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2006–2007. Infect Control Hosp Epidemiol. 2008;29:996–1011. doi: 10.1086/591861. [DOI] [PubMed] [Google Scholar]
- 4.Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10:266–278. doi: 10.1038/nrmicro2761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Remschmidt C, Schröder C, Behnke M, Gastmeier P, Geffers C, Kramer TS. Continuous increase of vancomycin resistance in enterococci causing nosocomial infections in Germany—10 years of surveillance. Antimicrob Resist Infect Control. 2018;7:54. doi: 10.1186/s13756-018-0353-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.O'Driscoll T, Crank CW. Vancomycin-resistant enterococcal infections: epidemiology, clinical manifestations, and optimal management. Infect Drug Resist. 2015;8:217–230. doi: 10.2147/IDR.S54125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hollenbeck BL, Rice LB. Intrinsic and acquired resistance mechanisms in enterococcus. Virulence. 2012;3:421–433. doi: 10.4161/viru.21282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Werner G, Coque TM, Hammerum AM, Hope R, Hryniewicz W, Johnson A, et al. Emergence and spread of vancomycin resistance among enterococci in Europe. Eurosurveillance. 2008 doi: 10.2807/ese.13.47.19046-en. [DOI] [PubMed] [Google Scholar]
- 9.Hayden MK, Blom DW, Lyle EA, Moore CG, Weinstein RA. Risk of hand or glove contamination after contact with patients colonized with vancomycin-resistant enterococcus or the colonized patients' environment. Infect Control Hosp Epidemiol. 2008;29:149–154. doi: 10.1086/524331. [DOI] [PubMed] [Google Scholar]
- 10.Correa-Martinez CL, Tönnies H, Froböse NJ, Mellmann A, Kampmeier S. Transmission of vancomycin-resistant enterococci in the hospital setting: uncovering the patient-environment interplay. Microorganisms. 2020 doi: 10.3390/microorganisms8020203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhou X, Willems RJL, Friedrich AW, Rossen JWA, Bathoorn E. Enterococcus faecium: from microbiological insights to practical recommendations for infection control and diagnostics. Antimicrob Resist Infect Control. 2020;9:130. doi: 10.1186/s13756-020-00770-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gastmeier P, Schröder C, Behnke M, Meyer E, Geffers C. Dramatic increase in vancomycin-resistant enterococci in Germany. J Antimicrob Chemother. 2014;69:1660–1664. doi: 10.1093/jac/dku035. [DOI] [PubMed] [Google Scholar]
- 13.Klare I, Witte W, Wendt C, Werner G. Vancomycin-resistant enterococci (VRE) Bundesgesundheitsbl. 2012 doi: 10.1007/s00103-012-1564-6. [DOI] [PubMed] [Google Scholar]
- 14.Borgmann S, Schulte B, Wolz C, Gruber H, Werner G, Goerke C, et al. Discrimination between epidemic and non-epidemic glycopeptide-resistant E. faecium in a post-outbreak situation. J Hosp Infect. 2007;67:49–55. doi: 10.1016/j.jhin.2007.06.002. [DOI] [PubMed] [Google Scholar]
- 15.Markwart R, Willrich N, Haller S, Noll I, Koppe U, Werner G, et al. The rise in vancomycin-resistant Enterococcus faecium in Germany: data from the German Antimicrobial Resistance Surveillance (ARS) Antimicrob Resist Infect Control. 2019;8:147. doi: 10.1186/s13756-019-0594-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jung E, Byun S, Lee H, Moon SY, Lee H. Vancomycin-resistant Enterococcus colonization in the intensive care unit: clinical outcomes and attributable costs of hospitalization. Am J Infect Control. 2014;42:1062–1066. doi: 10.1016/j.ajic.2014.06.024. [DOI] [PubMed] [Google Scholar]
- 17.Chiang H-Y, Perencevich EN, Nair R, Nelson RE, Samore M, Khader K, et al. Incidence and outcomes associated with infections caused by vancomycin-resistant enterococci in the united states: systematic literature review and meta-analysis. Infect Control Hosp Epidemiol. 2017;38:203–215. doi: 10.1017/ice.2016.254. [DOI] [PubMed] [Google Scholar]
- 18.Tacconelli E, Carrara E, Savoldi A, Harbarth S, Mendelson M, Monnet DL, et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis. 2018;18:318–327. doi: 10.1016/S1473-3099(17)30753-3. [DOI] [PubMed] [Google Scholar]
- 19.Flokas ME, Karageorgos SA, Detsis M, Alevizakos M, Mylonakis E. Vancomycin-resistant enterococci colonisation, risk factors and risk for infection among hospitalised paediatric patients: a systematic review and meta-analysis. Int J Antimicrob Agents. 2017;49:565–572. doi: 10.1016/j.ijantimicag.2017.01.008. [DOI] [PubMed] [Google Scholar]
- 20.Andersson P, Beckingham W, Gorrie CL, Kennedy K, Daveson K, Ballard SA, et al. Vancomycin-resistant Enterococcus (VRE) outbreak in a neonatal intensive care unit and special care nursery at a tertiary-care hospital in Australia—a retrospective case-control study. Infect Control Hosp Epidemiol. 2019;40:551–558. doi: 10.1017/ice.2019.41. [DOI] [PubMed] [Google Scholar]
- 21.Iosifidis E, Evdoridou I, Agakidou E, Chochliourou E, Protonotariou E, Karakoula K, et al. Vancomycin-resistant Enterococcus outbreak in a neonatal intensive care unit: epidemiology, molecular analysis and risk factors. Am J Infect Control. 2013;41:857–861. doi: 10.1016/j.ajic.2013.02.005. [DOI] [PubMed] [Google Scholar]
- 22.Posfay-Barbe KM, Zerr DM, Pittet D. Infection control in paediatrics. Lancet Infect Dis. 2008;8:19–31. doi: 10.1016/S1473-3099(07)70310-9. [DOI] [PubMed] [Google Scholar]
- 23.Simon A, Gröger N, Engelhart S, Molitor G, Exner M, Bode U, et al. Vancomycin resistant Enterococci (VRE)—importance, prevention and management in pediatrics—a review. Hyg Med. 2004;29:259–275. [Google Scholar]
- 24.Vydra J, Shanley RM, George I, Ustun C, Smith AR, Weisdorf DJ, et al. Enterococcal bacteremia is associated with increased risk of mortality in recipients of allogeneic hematopoietic stem cell transplantation. Clin Infect Dis. 2012;55:764–770. doi: 10.1093/cid/cis550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Adams DJ, Eberly MD, Goudie A, Nylund CM. Rising vancomycin-resistant enterococcus infections in hospitalized children in the United States. Hosp Pediatr. 2016;6:404–411. doi: 10.1542/hpeds.2015-0196. [DOI] [PubMed] [Google Scholar]
- 26.Khan AM, Morris SK, Bhutta ZA. Neonatal and perinatal infections. Pediatr Clin North Am. 2017;64:785–798. doi: 10.1016/j.pcl.2017.03.008. [DOI] [PubMed] [Google Scholar]
- 27.Vorstand des Klinikums der Universität München (LMU Klinikum). Leistungsbericht 2019. 2019. https://cdn0.scrvt.com/4d3e519fe5939342b95c7312343779ef/2b5753a974394f8e/2a0bea82de85/Leistungsbericht-2019_PRINT_korr.pdf. Accessed 23 June 2021.
- 28.European Committee on Antimicrobial Susceptability Testing (EUCAST). Antimicrobial susceptibility testing. 2021. https://eucast.org/ast_of_bacteria/. Accessed 19 May 2021.
- 29.Andrews S, Krueger F, Segonds-Pichon F, Biggins L, Krueger C, Wingett S. FastQC: a quality control tool for high throughput sequence data. Cambridge: Babraham institute; 2012. [Google Scholar]
- 30.Wood DE, Salzberg SL. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014;15:R46. doi: 10.1186/gb-2014-15-3-r46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Homan WL, Tribe D, Spark S, Li M, Hogg G, Spalburg E, et al. Multilocus sequence typing scheme for Enterococcus faecium. J Clin Microbiol. 2002;40(6):1963–1971. doi: 10.1128/JCM.40.6.1963-1971.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.de Been M, Pinholt M, Top J, Bletz S, Mellmann A, van Schaik W, et al. Core genome multilocus sequence typing scheme for high-resolution typing of Enterococcus faecium. J Clin Microbiol. 2015;53:3788–3797. doi: 10.1128/JCM.01946-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nationales Referenzzentrum für Staphylokokken und Enterokokken. FAQ zu Genomsequenzierung. 2019. https://www.rki.de/DE/Content/Infekt/NRZ/Staphylokokken/Praeanalytikhandbuch/FAQ-Genomsequenzierung.pdf?__blob=publicationFile. Accessed 19 May 2021.
- 35.Singh-Naz N, Sleemi A, Pikis A, Patel KM, Campos JM. Vancomycin-resistant Enterococcus faecium colonization in children. J Clin Microbiol. 1999;37:413–416. doi: 10.1128/JCM.37.2.413-416.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fossi Djembi L, Hodille E, Chomat-Jaboulay S, Coudrais S, De Santis N, Gardes S, et al. Factors associated with Vancomycin-resistant Enterococcus acquisition during a large outbreak. J Infect Public Health. 2017;10:185–190. doi: 10.1016/j.jiph.2016.04.010. [DOI] [PubMed] [Google Scholar]
- 37.Nourse C, Murphy H, Byrne C, O'Meara A, Breatnach F, Kaufmann M, et al. Control of a nosocomial outbreak of vancomycin resistant Enterococcus faecium in a paediatric oncology unit: risk factors for colonisation. Eur J Pediatr. 1998;157:20–27. doi: 10.1007/s004310050760. [DOI] [PubMed] [Google Scholar]
- 38.Haas EJ, Zaoutis TE, Prasad P, Li M, Coffin SE. Risk factors and outcomes for vancomycin-resistant enterococcus bloodstream infection in Children. Infect Control Hosp Epidemiol. 2010;31:1038–1042. doi: 10.1086/655464. [DOI] [PubMed] [Google Scholar]
- 39.World Health Organization. The 2019 WHO AWaRe classification of antibiotics for evaluation and monitoring of use. 2019. https://apps.who.int/iris/handle/10665/327957. Accessed 19 May 2021.
- 40.R Core Team. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. 2021. https://www.R-project.org/.
- 41.Erb S, Frei R, Dangel M, Widmer AF. Multidrug-resistant organisms detected more than 48 hours after hospital admission are not necessarily hospital-acquired. Infect Control Hosp Epidemiol. 2017;38:18–23. doi: 10.1017/ice.2016.226. [DOI] [PubMed] [Google Scholar]
- 42.Zhou Q, Moore C, Eden S, Tong A, McGeer A. Factors associated with acquisition of vancomycin-resistant enterococci (VRE) in roommate contacts of patients colonized or infected with VRE in a tertiary care hospital. Infect Control Hosp Epidemiol. 2008;29:398–403. doi: 10.1086/587187. [DOI] [PubMed] [Google Scholar]
- 43.Grundmann H, Bärwolff S, Tami A, Behnke M, Schwab F, Geffers C, et al. How many infections are caused by patient-to-patient transmission in intensive care units? Crit Care Med. 2005;33:946–951. doi: 10.1097/01.ccm.0000163223.26234.56. [DOI] [PubMed] [Google Scholar]
- 44.Lister DM, Tan K, Carse E, Stuart RL. Clearance of infant vancomycin-resistant Enterococcus faecium carriage after a neonatal inpatient outbreak. Am J Infect Control. 2016;44:1172–1173. doi: 10.1016/j.ajic.2016.03.032. [DOI] [PubMed] [Google Scholar]
- 45.Henning KJ, Delencastre H, Eagan J, Boone N, Brown A, Chung M, et al. Vancomycin-resistant Enterococcus faecium on a pediatric oncology ward: duration of stool shedding and incidence of clinical infection. Pediatr Infect Dis J. 1996;15:848–854. doi: 10.1097/00006454-199610000-00004. [DOI] [PubMed] [Google Scholar]
- 46.Schuster F, Graubner UB, Schmid I, Weiss M, Belohradsky BH. Vancomycin-resistant-enterococci—colonization of 24 patients on a pediatric oncology unit. Klin Padiatr. 1998;210:261–263. doi: 10.1055/s-2008-1043889. [DOI] [PubMed] [Google Scholar]
- 47.Nourse C, Byrne C, Murphy H, Kaufmann ME, Clarke A, Butler K. Eradication of vancomycin resistant Enterococcus faecium from a paediatric oncology unit and prevalence of colonization in hospitalized and community-based children. Epidemiol Infect. 2000;124:53–59. doi: 10.1017/S095026889900326X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Drees M, Snydman DR, Schmid CH, Barefoot L, Hansjosten K, Vue PM, et al. Prior environmental contamination increases the risk of acquisition of vancomycin-resistant enterococci. Clin Infect Dis. 2008;46:678–685. doi: 10.1086/527394. [DOI] [PubMed] [Google Scholar]
- 49.McDermott H, Skally M, O'Rourke J, Humphreys H, Fitzgerald-Hughes D. Vancomycin-resistant enterococci (VRE) in the intensive care unit in a nonoutbreak setting: identification of potential reservoirs and epidemiological associations between patient and environmental VRE. Infect Control Hosp Epidemiol. 2018;39:40–45. doi: 10.1017/ice.2017.248. [DOI] [PubMed] [Google Scholar]
- 50.Dancer SJ. Controlling hospital-acquired infection: focus on the role of the environment and new technologies for decontamination. Clin Microbiol Rev. 2014;27:665–690. doi: 10.1128/CMR.00020-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Schulz-Stübner S, Schmidt-Warnecke A, Hwang J-H. VRE transmission via the reusable breathing circuit of a transport ventilator: outbreak analysis and experimental study of surface disinfection. Intensive Care Med. 2013;39:975–976. doi: 10.1007/s00134-013-2842-y. [DOI] [PubMed] [Google Scholar]
- 52.Golan Y, Doron S, Sullivan B, Snydman DR. Transmission of vancomycin-resistant enterococcus in a neonatal intensive care unit. Pediatr Infect Dis J. 2005;24:566–567. doi: 10.1097/01.inf.0000164762.03930.0a. [DOI] [PubMed] [Google Scholar]
- 53.Porwancher R, Sheth A, Remphrey S, Taylor E, Hinkle C, Zervos M. Epidemiological study of hospital-acquired infection with vancomycin-resistant Enterococcus faecium: possible transmission by an electronic ear-probe thermometer. Infect Control Hosp Epidemiol. 1997;18:771–773. doi: 10.1086/647535. [DOI] [PubMed] [Google Scholar]
- 54.Duckro AN, Blom DW, Lyle EA, Weinstein RA, Hayden MK. Transfer of vancomycin-resistant enterococci via health care worker hands. Arch Intern Med. 2005;165:302–307. doi: 10.1001/archinte.165.3.302. [DOI] [PubMed] [Google Scholar]
- 55.Jackson SS, Harris AD, Magder LS, Stafford KA, Johnson JK, Miller LG, et al. Bacterial burden is associated with increased transmission to health care workers from patients colonized with vancomycin-resistant Enterococcus. Am J Infect Control. 2019;47:13–17. doi: 10.1016/j.ajic.2018.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Xanthopoulou K, Peter S, Tobys D, Behnke M, Dinkelacker AG, Eisenbeis S, et al. Vancomycin-resistant Enterococcus faecium colonizing patients on hospital admission in Germany: prevalence and molecular epidemiology. J Antimicrob Chemother. 2020;75:2743–2751. doi: 10.1093/jac/dkaa271. [DOI] [PubMed] [Google Scholar]
- 57.Hansen SK, Andersen L, Detlefsen M, Holm A, Roer L, Antoniadis P, et al. Using core genome multilocus sequence typing (cgMLST) for vancomycin-resistant Enterococcus faecium isolates to guide infection control interventions and end an outbreak. J Glob Antimicrobial Resist. 2021;24:418–423. doi: 10.1016/j.jgar.2021.02.007. [DOI] [PubMed] [Google Scholar]
- 58.Pinholt M, Larner-Svensson H, Littauer P, Moser CE, Pedersen M, Lemming LE, et al. Multiple hospital outbreaks of vanA Enterococcus faecium in Denmark, 2012–13, investigated by WGS, MLST and PFGE J. Antimicrob Chemother. 2015;70:2474–2482. doi: 10.1093/jac/dkv142. [DOI] [PubMed] [Google Scholar]
- 59.Eisenberger D, Tuschak C, Werner M, Bogdan C, Bollinger T, Hossain H, et al. Whole-genome analysis of vancomycin-resistant Enterococcus faecium causing nosocomial outbreaks suggests the occurrence of few endemic clonal lineages in Bavaria, Germany. J Antimicrob Chemother. 2020;75:1398–1404. doi: 10.1093/jac/dkaa041. [DOI] [PubMed] [Google Scholar]
- 60.Freitas AR, Tedim AP, Francia MV, Jensen LB, Novais C, Peixe L, et al. Multilevel population genetic analysis of vanA and vanB Enterococcus faecium causing nosocomial outbreaks in 27 countries (1986–2012) J Antimicrob Chemother. 2016;71:3351–3366. doi: 10.1093/jac/dkw312. [DOI] [PubMed] [Google Scholar]
- 61.Sivertsen A, Billström H, Melefors Ö, Liljequist BO, Wisell KT, Ullberg M, et al. A multicentre hospital outbreak in Sweden caused by introduction of a vanB2 transposon into a stably maintained pRUM-plasmid in an Enterococcus faecium ST192 clone. PLoS ONE. 2014;9:e103274. doi: 10.1371/journal.pone.0103274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Arredondo-Alonso S, Top J, Corander J, Willems RJL, Schürch AC. Mode and dynamics of vanA-type vancomycin resistance dissemination in Dutch hospitals. Genome Med. 2021;13:9. doi: 10.1186/s13073-020-00825-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Pinholt M, Gumpert H, Bayliss S, Nielsen JB, Vorobieva V, Pedersen M, et al. Genomic analysis of 495 vancomycin-resistant Enterococcus faecium reveals broad dissemination of a vanA plasmid in more than 19 clones from Copenhagen, Denmark. J Antimicrob Chemother. 2017;72:40–47. doi: 10.1093/jac/dkw360. [DOI] [PubMed] [Google Scholar]
- 64.Hufnagel M, Liese C, Loescher C, Kunze M, Proempeler H, Berner R, et al. Enterococcal colonization of infants in a neonatal intensive care unit: associated predictors, risk factors and seasonal patterns. BMC Infect Dis. 2007;7:107. doi: 10.1186/1471-2334-7-107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Miedema CJ, Kerkhof M, Arends JP, Bergman KA, Kimpen JL. Risk factors for colonization with enterococci in a neonatal intensive care unit. Clin Microbiol Infect. 2000;6:53. doi: 10.1046/j.1469-0691.2000.00019.x. [DOI] [PubMed] [Google Scholar]
- 66.Malik RK, Montecalvo MA, Reale MR, Li K, Maw M, Munoz JL, et al. Epidemiology and control of vancomycin-resistant enterococci in a regional neonatal intensive care unit. Pediatr Infect Dis J. 1999;18:352–356. doi: 10.1097/00006454-199904000-00009. [DOI] [PubMed] [Google Scholar]
- 67.Aktürk H, Sütçü M, Somer A, Karaman S, Acar M, Ünüvar A, et al. Results of four-year rectal vancomycin-resistant enterococci surveillance in a pediatric hematology-oncology ward: from colonization to infection. Turk J Haematol. 2016;33:244–247. doi: 10.4274/tjh.2015.0368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Carmeli Y, Eliopoulos GM, Samore MH. Antecedent treatment with different antibiotic agents as a risk factor for vancomycin-resistant Enterococcus. Emerg Infect Dis. 2002;8:802–807. doi: 10.3201/eid0808.010418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Hygienemaßnahmen zur Prävention der Infektion durch Enterokokken mit speziellen Antibiotikaresistenzen: Empfehlung der Kommission für Krankenhaushygiene und Infektionsprävention (KRINKO) beim Robert Koch-Institut. Bundesgesundheitsbl. 2018;61:1310–61. doi:10.1007/s00103-018-2811-2. [DOI] [PubMed]
- 70.Centers for Disease Control and Prevention. Recommendations for preventing the spread of vancomycin resistance: recommendations of the Hospital Infection Control Practices Advisory Committee (HICPAC). MMWR 1995;44(No. RR-12). [PubMed]
- 71.Siegel JD, Rhinehart E, Jackson M, Chiarello L. Management of multidrug-resistant organisms in health care settings, 2006. Am J Infect Control. 2007;35:S165–S193. doi: 10.1016/j.ajic.2007.10.006. [DOI] [PubMed] [Google Scholar]
- 72.National Health and Medical Research Council. Australian Guidelines for the Prevention and Control of Infection in Healthcare. Canberra: Commonwealth of Australia; 2019.
- 73.WHO Patient Safety, World Health Organization. WHO guidelines on hand hygiene in health care. 2009. http://apps.who.int/iris/bitstream/handle/10665/44102/9789241597906_eng.pdf? sequence=1. Accessed 03 August 2021.
- 74.Muto CA, Jernigan JA, Ostrowsky BE, Richet HM, Jarvis WR, Boyce JM, et al. SHEA guideline for preventing nosocomial transmission of multidrug-resistant strains of Staphylococcus aureus and Enterococcus. Infect Control Hosp Epidemiol. 2003;24:362–386. doi: 10.1086/502213. [DOI] [PubMed] [Google Scholar]
- 75.Price CS, Paule S, Noskin GA, Peterson LR. Active surveillance reduces the incidence of vancomycin-resistant enterococcal bacteremia. Clin Infect Dis. 2003;37:921–928. doi: 10.1086/377733. [DOI] [PubMed] [Google Scholar]
- 76.Lepelletier D, Berthelot P, Lucet J-C, Fournier S, Jarlier V, Grandbastien B. French recommendations for the prevention of ‘emerging extensively drug-resistant bacteria’ (eXDR) cross-transmission. J Hosp Infect. 2015;90:186–195. doi: 10.1016/j.jhin.2015.04.002. [DOI] [PubMed] [Google Scholar]
- 77.Ostrowsky BE, Trick WE, Sohn AH, Quirk SB, Holt S, Carson LA, et al. Control of vancomycin-resistant enterococcus in health care facilities in a region. N Engl J Med. 2001;344:1427–1433. doi: 10.1056/NEJM200105103441903. [DOI] [PubMed] [Google Scholar]
- 78.Cheng VCC, Tai JWM, Chau PH, Lai CKC, Chuang VWM, So SYC, et al. Successful control of emerging vancomycin-resistant enterococci by territory-wide implementation of directly observed hand hygiene in patients in Hong Kong. Am J Infect Control. 2016;44:1168–1171. doi: 10.1016/j.ajic.2016.03.050. [DOI] [PubMed] [Google Scholar]
- 79.Anderson DJ, Gergen MF, Smathers E, Sexton DJ, Chen LF, Weber DJ, et al. Decontamination of targeted pathogens from patient rooms using an automated ultraviolet-C-emitting device. Infect Control Hosp Epidemiol. 2013;34(5):466–471. doi: 10.1086/670215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Barnes SL, Rock C, Harris AD, Cosgrove SE, Morgan DJ, Thom KA. The impact of reducing antibiotics on the transmission of multidrug-resistant organisms. Infect Control Hosp Epidemiol. 2017;38:663–669. doi: 10.1017/ice.2017.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Patel SJ, Saiman L. Principles and strategies of antimicrobial stewardship in the neonatal intensive care unit. Semin Perinatol. 2012;36:431–436. doi: 10.1053/j.semperi.2012.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kreitmeyr K, von Both U, Pecar A, Borde JP, Mikolajczyk R, Huebner J. Pediatric antibiotic stewardship: successful interventions to reduce broad-spectrum antibiotic use on general pediatric wards. Infection. 2017;45:493–504. doi: 10.1007/s15010-017-1009-0. [DOI] [PubMed] [Google Scholar]
- 83.Harris PNA, Wailan AM. Beyond the core genome: tracking plasmids in outbreaks of multidrug-resistant bacteria. Clin Infect Dis. 2021;72:421–422. doi: 10.1093/cid/ciaa052. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.