Abstract
Background
Healthcare-associated infections pose a potentially fatal threat to patients worldwide and Staphylococcus aureus is one of the most common causes of healthcare-associated infections. S. aureus is a common commensal pathogen and a frequent cause of bacteremia, with studies demonstrating that nasal and blood isolates from single patients match more than 80% of the time. Here we report on a contemporary collection of colonizing isolates from those with methicillin-resistant S. aureus (MRSA) bloodstream infections to evaluate the diversity within hosts, and detail the clinical features associated with concomitant nasal colonization.
Methods
Swabs of the bilateral anterior nares were obtained from patients diagnosed with MRSA bacteremia. A single colony culture from the blood and an average of 6 colonies from the nares were evaluated for MRSA growth. For the nares cultures, we typed multiple isolates for staphylococcal protein A (spa) and derived the clonal complexes. Demographic and clinical data were obtained retrospectively from the electronic medical record system and analysed using univariate and multivariable regression models.
Results
Over an 11-month period, 68 patients were diagnosed with MRSA bloodstream infection, 53 were swabbed, and 37 (70%) were colonized with MRSA in the anterior nares. We performed molecular typing on 213 nasal colonies. Spa types and clonal complexes found in the blood were also detected in the nares in 95% of the cases. We also found that 11% of patients carried more than one clone of MRSA in the nares. Male sex and history of prior hospitalization within the past 90 days increased odds for MRSA colonization.
Conclusion
The molecular epidemiological landscape of colonization in the setting of invasive disease is diverse and defining the interplay between colonization and invasive disease is critical to combating invasive MRSA disease.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12879-022-07371-w.
Keywords: Methicillin-resistant Staphylococcus aureus, Bloodstream infections, Colonization, Molecular epidemiology
Background
Staphylococcus aureus (S. aureus) is one of the most common causes of healthcare-associated infections (HAI), associated with high mortality rates in invasive disease [1, 2]. S. aureus is also a common commensal pathogen and approximately 20–36% of the human population may be intermittently or persistently colonized, often asymptomatically [3–5]. Although S. aureus can colonize all skin surfaces, the most frequently assessed site of colonization is the anterior nares [6]. Nasal carriage plays a key role in progression onward to invasive disease [7], and persistent carriers have higher relative risk to develop bacteremia compared to non-carriers [8].
Within-host diversity of nasal carriage with S. aureus has been estimated to be 6.6% [9], and people commonly become infected with one of their colonizing isolates [10]. In a multicenter study, 82% of patients with MRSA bacteremia had identical clones in the anterior nares, and 86% of patients with MRSA colonization in anterior nares had clonally identical isolates later obtained from blood [10]. Colonization is also key to subtle transmission events and has been a central feature of outbreaks [11, 12]. In the United States, most S. aureus infections are caused by two dominant clonal complexes (CC), CC5 and CC8 [2]. CC5 has historically encompassed hospital-associated genotypes, and CC8 community genotypes. Our work [13] and that of others [14, 15] have found clonal clinical correlations are merging, with classic community genotypes becoming a common cause of hospital-associated infections. Here, we tested the associations between infection with dominant MRSA genotypes and the salient clinical features for the study population.
Given the importance of S. aureus colonization in invasive disease, we investigated the molecular epidemiological landscape of patients with MRSA bloodstream infections (BSI) with concomitant nasal colonization. Our aim was to collect a contemporary set of colonizing isolates from those with MRSA BSI, evaluate the frequency of diversity in the colonizing isolates, and define clinical features associated with the presence of colonization. We hypothesized that for patients with active MRSA BSI, there would be clinical differences between those concomitantly colonized and those who were not, as the presence of colonization itself is an important marker of disease status. We thus examined the clinical features and outcomes of patients with active MRSA BSI both with and without concomitant nasal colonization. Further definition of the link between colonization and infection will enable future avenues for mitigation of this infection.
Results
Molecular diversity of nasal colonization in those with MRSA BSI
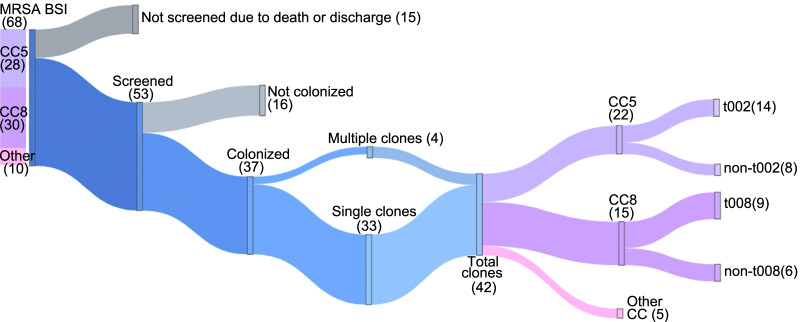
Over an 11-month period (July 2018 to May 2019), 68 unique patients were diagnosed with MRSA BSI. Sequencing of single colonies from the blood cultures revealed that 30 (44%) were caused by CC8, 28 (41%) by CC5 and 10 (15%) by other clonal complexes (Fig. 1). From the 68 patients with MRSA BSI, 53 were successfully swabbed in the nares and screened for MRSA colonization. Fifteen patients were excluded due to discharge or death at the time of identification. From the 53 individuals who underwent nasal swabbing, 37 (70%) were colonized with MRSA. The average number of colonies picked from the nasal growth plates was 6 (range 2–12), resulting in a total of 213 which underwent spa typing (Additional file 1: Table S1). It was found that 95% of patients had at least one matching spa type between the blood isolate and nasal isolate. Four of the 37 colonized patients (11%) carried multiple spa types in the anterior nares, resulting in a total of 42 distinct nasal clones. Among the individuals who had swabs positive for MRSA, 22 of the nasal spa types corresponded to CC5 and 15 corresponded to CC8, and 5 with other CCs (Fig. 1). Within the CC5 samples, t002 represented the most common spa type, present in 14 of the 22 samples (64%), while t008 presented as the most common spa type amongst the CC8 samples, in 9 of the 15 samples (60%) (Additional file 1: Table S1). Four MRSA blood isolates were also identified as novel sequence types (ST); ST6964, ST6983, ST7090, ST7094.
Fig. 1.
Sankey flow chart diagram of 68 patients with MRSA BSI detailing nasal colonization status with breakdown by spa and clonal complex (CC)
Epidemiological and clinical characteristics associated with bacteremia and colonization
The 68 patients with MRSA BSI were predominantly male (n = 41, 60%), between the age of 55 and 69 (n = 28; 41%). The patients were 34% (n = 23) non-Hispanic white, 34% (n = 23) non-Hispanic black, 28% (n = 19) Hispanic, and 4% (n = 3) belonged to other races or ethnicity groups (Table 1). Patients were more frequently admitted from home (n = 46; 68%), but more than half of the patients had a history of previous hospitalization within the prior 90 days (n = 42; 62%). Community-onset (CO) infections, or infections which present within the first 3 days of hospitalization, were more common than hospital-onset (HO) (n = 40; 59% vs n = 28; 41%) infections, however 75% (n = 51) of patients had an invasive device in place at the time of infection. A prior personal history of colonization, as defined as any prior positive culture for MRSA from any site, or a documented history of prior MRSA infection or colonization, was present 28% of the time. The sources of infection were predominantly deemed to be unknown/other (32%), vascular access (22%), and skin and soft tissue (21%), followed by peripheral IV (10%), and pneumonia (7%) (Table 1).
Table 1.
Demographics and clinical characteristics of patients with MRSA BSI
| Factor, n (%) | (n = 68) |
|---|---|
| Sex | |
| Male | 41 (60) |
| Female | 27 (40) |
| Race/ethnicity | |
| Non-hispanic white | 23 (34) |
| Non-hispanic black | 23 (34) |
| Hispanic/latinos | 19 (28) |
| Asian | 2 (3) |
| Other/not reported | 1 (1) |
| Age at time of infection | |
| 18–54 years | 18 (26) |
| 55–69 years | 28 (41) |
| ≥ 70 years | 22 (32) |
| History of IV drug use | 4 (6) |
| HIV | 7 (10) |
| Admission source | |
| Homea | 46 (68) |
| NH/Rehab/LTACH | 15 (22) |
| Outside hospital | 7 (10) |
| Prior hospital admission (90 days) | 42 (62) |
| Length of hospital stay prior to BSI | |
| CO-MRSA | 40 (59) |
| HO-MRSA | 28 (41) |
| Clonal complex | |
| CC5 | 28 (41) |
| CC8 | 30 (44) |
| Other | 10 (15) |
| Frequent healthcare interaction | |
| Hemodialysis | 14 (21) |
| Infusion centerb | 14 (21) |
| None | 40 (59) |
| Presence of invasive devicec | 51 (75) |
| Invasive procedures (missing = 1)d | 25 (37) |
| Wound presente | 35 (51) |
| Comorbiditiesf | |
| Myocardial infarction | 11 (16) |
| Congestive heart failure | 18 (26) |
| Peripheral vascular disease | 11 (16) |
| Cerebrovascular disease | 11 (16) |
| Dementia | 8 (12) |
| Chronic pulmonary disease | 15 (22) |
| Connective tissue disease | 3 (4) |
| Peptic ulcer disease | 1 (1) |
| Mild liver disease | 4 (6) |
| Diabetes (no complications) | 18 (26) |
| Diabetes with organ damage | 11 (16) |
| Hemi or paraplegia | 7 (10) |
| Moderate/severe renal disease | 14 (21) |
| Solid tumor | 2 (3) |
| Leukemia | 1 (1) |
| Lymphoma/multiple myeloma | 10 (15) |
| Moderate/severe liver disease | 5 (7) |
| Metastatic solid tumor | 11 (16) |
| Charlson comorbidity index (CCI) | |
| 0–3 | 13 (19) |
| 4–5 | 20 (29) |
| 6–8 | 22 (32) |
| > 8 | 13 (19) |
| History of transplantg | 8 (12) |
| History of MRSA colonizationh | 19 (28) |
| Presumed source of MRSA BSI | |
| Peripheral IV | 7 (10) |
| Skin and soft tissue infection | 14 (21) |
| Pneumonia | 5 (7) |
| Diabetic foot infection | 1 (1) |
| Vascular access | 15 (22) |
| Sacral wound | 4 (6) |
| Other/unknown sourcei | 22 (32) |
IV intravenous, NH nursing home, Rehab rehabilitation facility, LTACH long-term acute care hospital, CO community-onset, HO hospital-onset, MRSA methicillin-resistant Staphylococcus aureus, BSI bloodstream infection, HIV human immunodeficiency virus
a “Admission from home”: included nonmedical residences such as home, group homes, assisted living facilities, and homeless shelters
b“Infusion center”: outpatient centers for chemotherapy, intravenous fluids, intravenous immunomodulators, and blood products
c“Presence of invasive device”: included pacemaker, implantable cardioverter defibrillator (ICD), left ventricular assist device (LVAD), vascular access (excluding peripheral intravenous catheters), orthopedic hardware, nephrostomy, suprapubic catheter, ileal conduit, foley catheter, arteriovenous graft placement (AVG), percutaneous endoscopic gastrostomy (PEG) tube, or ostomy
d“Invasive procedures”: included any invasive procedures or surgery occurring within 1 month before first positive blood culture for MRSA, excluding electroencephalogram (EEG), electrocardiogram (EKG), and transthoracic echocardiogram (TTE)
e“Wound present”: presence of a chronic skin wound overlying the sacrum, limb, abdomen, or other body part
f“Comorbidities”: as defined by the Charlson Comorbidity Index (CCI); refer to standard definitions for CCI
g“History of transplant”: included solid organ and bone marrow transplant
h“History of MRSA colonization”: any positive culture from urine, sputum, tissue, or nares with MRSA prior to the positive MRSA blood culture or a documented history of prior MRSA infection or colonization
i“Other/Unknown source”: MRSA infection from urinary source, osteomyelitis, surgical site infection, spinal infection, septic arthritis or cardiac device infection, or an unknown/not reported source
Multivariable analysis found that those with concomitant MRSA colonization were more likely to be male, comprising 73% (n = 27) of the colonized patients compared to 44% (n = 7) in the non-colonized group (OR, 5.06; 95% CI, 1.64–22.01; p = 0.03) (Table 2). Patients were more likely to have a prior hospital admission within 90 days (n = 26; 70%) compared to the non-colonized (n = 6; 38%) (OR, 3.96; 95% CI, 1.01–15.56; p = 0.05). Patients with a history of renal disease including need for hemodialysis were more likely to be colonized (OR, 15.52; 95% CI, 1.13–213.37; p = 0.04), while dementia was negatively associated with colonization (OR, 0.10; 95% CI, 0.01–1.02; p = 0.05) (Additional file 2: Table S2). There were no significant differences in outcomes between the two groups (Additional file 3: Table S3). All patients who underwent swabbing were treated with standard of care anti-MRSA therapy (intravenous vancomycin), and there was no significant difference between the time of bacteremia and the time of nasal swab acquisition between colonized and non-colonized patients (Wilcoxon rank sum t-test p = 0.1738). Patients were not actively decolonized with anti-MRSA antibiotics, such as intranasal mupirocin.
Table 2.
Demographics and clinical characteristics of patients with MRSA BSI with and without nasal colonization
| Factor | Colonized N = 37 (%) |
Not colonized N = 16 (%) |
Univariate analysis OR (95% CI) |
p value | Multivariable analysis OR (95% CI) |
p value |
|---|---|---|---|---|---|---|
| Sex | ||||||
| Male | 27 (73) | 7 (44) | 3.47 (1.02–11.82) | 0.05 | 5.06 (1.64–22.01) | 0.03 |
| Female | 10 (27) | 9 (56) | Reference | Reference | ||
| Race/ethnicity | ||||||
| Non-hispanic white | 13 (35) | 7 (44) | Reference | |||
| Non-hispanic black | 11 (30) | 4 (25) | 1.48 (0.34–6.43) | 0.60 | ||
| Hispanic/latino | 11 (30) | 5 (31) | 1.19 (0.29–4.81) | 0.81 | ||
| Asian | 1 (3) | 0 (0) | – | – | ||
| Other/not reported | 1 (3) | 0 (0) | – | – | ||
| Age at time of infection | ||||||
| 18–54 years | 12 (32) | 2 (13) | Reference | |||
| 55–69 years | 16 (43) | 6 (38) | 0.44 (0.08–2.60) | 0.37 | ||
| ≥ 70 years | 9 (24) | 8 (50) | 0.19 (0.03–1.11) | 0.06 | ||
| History of IV drug use | ||||||
| Yes | 2 (5) | 1 (6) | 0.86 (0.07–10.19) | 0.90 | ||
| No | 35 (95) | 15 (94) | Reference | |||
| HIV diagnosis | ||||||
| Yes | 6 (16) | 0 (0) | – | – | ||
| No | 31 (84) | 16 (100) | – | – | ||
| Admission source | ||||||
| Home | 22 (59) | 13 (81) | Reference | |||
| NH/Rehab/LTACH | 10 (27) | 2 (13) | 2.96 (0.56–15.63) | 0.20 | ||
| Other hospital | 5 (14) | 1 (6) | 2.96 (0.31–28.14) | 0.35 | ||
| Prior hospital admission (90 days) | ||||||
| Yes | 26 (70) | 6 (38) | 3.94 (1.15–13.52) | 0.03 | 3.96 (1.01–15.56) | 0.05 |
| No | 11 (30) | 10 (63) | Reference | Reference | ||
| Length of hospital stay prior to BSI | ||||||
| CO-MRSA | 20 (54) | 10 (63) | Reference | |||
| HO-MRSA | 17 (46) | 6 (38) | 1.42 (0.43–4.71) | 0.57 | ||
| Clonal complex | ||||||
| CC8 | 13 (35) | 9 (56) | Reference | |||
| CC5 | 19 (51) | 5 (31) | 2.63 (0.72–9.66) | 0.15 | ||
| Other | 5 (14) | 2 (13) | 1.73 (0.27–10.97) | 0.56 | ||
| Frequent healthcare interaction | ||||||
| Hemodialysis | 12 (32) | 1 (6) | 8.00 (0.92–69.84) | 0.06 | ||
| Infusion Center | 7 (19) | 3 (19) | 1.56 (0.33–7.24) | 0.57 | ||
| None | 18 (49) | 12 (75) | Reference | |||
| Presence of invasive device | ||||||
| Yes | 30 (81) | 10 (63) | 2.57 (0.70–9.48) | 0.16 | ||
| No | 7 (19) | 6 (38) | Reference | |||
| Invasive procedures | ||||||
| Yes | 13 (35) | 4 (25) | 1.63 (0.44–6.07) | 0.47 | ||
| No | 24 (65) | 12 (75) | Reference | |||
| Wound present | ||||||
| Yes | 21 (57) | 6 (38) | 2.19 (0.66–7.29) | 0.20 | ||
| No | 16 (43) | 10 (63) | Reference | |||
| Charlson comorbidity index (CCI) | ||||||
| 0–3 | 4 (11) | 5 (31) | Reference | Reference | ||
| 4–5 | 12 (32) | 4 (25) | 3.75 (0.66–21.25) | 0.14 | 7.44 (0.95–58.45) | 0.06 |
| 6–8 | 15 (41) | 4 (25) | 4.69 (0.84–26.08) | 0.08 | 7.84 (1.03–59.84) | 0.05 |
| > 8 | 6 (16) | 3 (19) | 2.50 (0.37–16.89) | 0.35 | 3.68 (0.44–30.88) | 0.23 |
| History of transplant | ||||||
| Yes | 6 (16) | 1 (6) | 2.90 (0.32–26.33) | 0.34 | ||
| No | 31 (84) | 15 (94) | Reference | |||
| History of MRSA colonization | ||||||
| Yes | 14 (38) | 2 (13) | 4.26 (0.84–21.60) | 0.08 | ||
| No | 23 (62) | 14 (88) | Reference | |||
| Presumed source of MRSA BSI | ||||||
| Peripheral IV | 5 (14) | 2 (13) | 1.09 (0.19–6.33) | 0.92 | ||
| Skin and soft tissue infection | 7 (19) | 4 (25) | 0.70 (0.17–2.84) | 0.62 | ||
| Pneumonia | 3 (8) | 1 (6) | 1.32 (0.13–13.78) | 0.81 | ||
| Diabetic foot infection | 1 (3) | 0 (0) | – | – | ||
| Vascular access | 11 (30) | 3 (19) | 1.83 (0.43–7.74) | 0.41 | ||
| Sacral wound | 2 (5) | 2 (13) | 0.40 (0.05–3.13) | 0.38 | ||
| Other/unknown | 8 (22) | 4 (25) | 0.83 (0.21–3.28) | 0.79 |
See Table 1 for definitions
Bold = significant at ≤ 0.05
IV intravenous, NH nursing home, Rehab rehabilitation facility, LTACH long-term acute care hospital, CO community-onset, HO hospital-onset, MRSA methicillin-resistant Staphylococcus aureus, BSI bloodstream infection, HIV human immunodeficiency virus
When comparing clinical characteristics associated with two predominant clonal complexes (CC5 and CC8) in patients bacteremic during the entire study period, admission from a rehabilitation facility was significantly more common in those infected with CC5 (n = 11; 39%) compared to CC8 (n = 4; 13%) (OR, 3.78; CI 1.02–14.06; p = 0.05) (Table 3). Furthermore, patients in the CC5 group had significantly higher rates of hospitalizations within the prior 90 days (n = 22; 79%) compared to those with CC8 (n = 15; 50%) (OR, 3.67; CI 1.16–11.60; p = 0.03). In the multivariable analysis, the presence of an invasive device was 93% (n = 26) in the CC5 group and 60% (n = 18) in the CC8 group (OR, 9.75; CI 1.87–50.85; p = 0.007). Although not approaching significance, the primary source of BSI due to CC8 vs. CC5 was 13% vs 5% in skin derived infections (due to either peripheral IV or skin and soft tissue infection). While the 30-day mortality rate for all bacteremic patients was 26%, there were no differences in mortality and outcomes between those infected with either CC8 or CC5.
Table 3.
Demographics and clinical characteristics of patients with MRSA BSI due to CC5 vs. CC8
| Factor | CC5 N = 28 (%) |
CC8 N = 30 (%) |
Univariate analysis OR (95% CI) |
p value | Multivariable analysis OR (95% CI) |
p value |
|---|---|---|---|---|---|---|
| Sex | ||||||
| Male | 20 (71) | 16 (53) | 2.19 (0.74–6.50) | 0.16 | 2.63 (0.80–8.61) | 0.11 |
| Female | 8 (29) | 14 (47) | Reference | Reference | ||
| Race/ethnicity | ||||||
| Non-hispanic white | 10 (36) | 11 (37) | Reference | |||
| Non-hispanic black | 11 (39) | 8 (27) | 1.51 (0.43–5.28) | 0.52 | ||
| Hispanic/latino | 7 (25) | 9 (30) | 0.86 (0.23–3.16) | 0.82 | ||
| Asian | 0 (0) | 1 (3) | – | – | ||
| Other/not reported | 0 (0) | 1 (3) | – | – | ||
| Age at time of infection | ||||||
| 18–54 years | 7 (25) | 8 (27) | Reference | |||
| 55–69 years | 9 (32) | 15 (50) | 0.69 (0.19–2.54) | 0.57 | ||
| ≥ 70 years | 12 (43) | 7 (23) | 1.96 (0.49–7.77) | 0.34 | ||
| History of IV drug use | ||||||
| Yes | 0 (0) | 3 (10) | – | – | ||
| No | 28 (100) | 27 (90) | – | – | ||
| HIV diagnosis | ||||||
| Yes | 3 (11) | 3 (10) | 1.08 (0.20–5.86) | 0.93 | ||
| No | 25 (89) | 27 (90) | Reference | |||
| Admission source | ||||||
| Home | 16 (57) | 22 (73) | Reference | |||
| NH/Rehab/LTACH | 11 (39) | 4 (13) | 3.78 (1.02–14.06) | 0.05 | ||
| Other hospital | 1 (4) | 4 (13) | 0.34 (0.04–3.37) | 0.36 | ||
| Prior hospital admission (90 days) | ||||||
| Yes | 22 (79) | 15 (50) | 3.67 (1.16–11.60) | 0.03 | ||
| No | 6 (21) | 15 (50) | Reference | |||
| Length of hospital stay prior to BSI | ||||||
| CO-MRSA | 17 (61) | 19 (63) | Reference | |||
| HO-MRSA | 11 (39) | 11 (37) | 1.12 (0.39–3.23) | 0.84 | ||
| Frequent healthcare interaction | ||||||
| Hemodialysis | 9 (32) | 4 (13) | 3.00 (0.77–11.62) | 0.11 | ||
| Infusion center | 4 (14) | 6 (20) | 0.89 (0.21–3.72) | 0.87 | ||
| None | 15 (54) | 20 (67) | Reference | |||
| Presence of invasive device | ||||||
| Yes | 26 (93) | 18 (60) | 8.67 (1.73–43.49) | 0.009 | 9.75 (1.87–50.85) | 0.007 |
| No | 2 (7) | 12 (40) | Reference | |||
| Invasive procedures (missing = 1) | ||||||
| Yes | 10 (36) | 11 (37) | 0.91 (0.31–2.67) | 0.86 | ||
| No | 18 (64) | 18 (60) | Reference | |||
| Wound present | ||||||
| Yes | 18 (64) | 12 (40) | 2.70 (0.93–7.82) | 0.07 | ||
| No | 10 (36) | 18 (60) | Reference | |||
| Charlson comorbidity index (CCI) | ||||||
| 0–3 | 5 (18) | 7 (23) | Reference | |||
| 4–5 | 9 (32) | 8 (27) | 1.58 (0.35–7.00) | 0.55 | ||
| 6–8 | 11 (39) | 8 (27) | 1.93 (0.45–8.33) | 0.38 | ||
| > 8 | 3 (11) | 7 (23) | 0.60 (0.10–3.54) | 0.57 | ||
| History of transplant | ||||||
| Yes | 4 (14) | 1 (3) | 4.83 (0.51–46.18) | 0.17 | ||
| No | 24 (86) | 29 (97) | Reference | |||
| History of MRSA colonization | ||||||
| Yes | 9 (32) | 5 (17) | 2.37 (0.68–8.23) | 0.17 | ||
| No | 19 (68) | 25 (83) | Reference | |||
| Presumed source of MRSA BSI | ||||||
| Peripheral IV | 1 (4) | 5 (17) | 0.19 (0.02–1.70) | 0.14 | ||
| Skin and soft tissue infection | 4 (14) | 8 (27) | 0.46 (0.12–1.74) | 0.25 | ||
| Pneumonia | 2 (7) | 1 (3) | 2.23 (0.19–26.06) | 0.52 | ||
| Diabetic foot infection | 1 (4) | 0 (0) | – | – | ||
| Vascular access | 8 (29) | 6 (20) | 1.60 (0.48–5.38) | 0.45 | ||
| Sacral wound | 4 (14) | 0 (0) | – | – | ||
| other/unknown | 8 (29) | 10 (33) | 0.80 (0.26–2.45) | 0.70 |
See Table 1 for definitions
Bold = significant at ≤ 0.05
IV intravenous, NH nursing home, Rehab rehabilitation facility, LTACH long-term acute care hospital, CO community-onset, HO hospital-onset, MRSA methicillin-resistant Staphylococcus aureus, BSI bloodstream infection, HIV human immunodeficiency virus
Methods
Study setting, patient identification, and isolate selection
The Mount Sinai Hospital (MSH) is a quaternary care facility. Under the approval of the MSH institutional review board (HS # 18-00662; 13-00981), adults (≥ 18 years old) with MRSA BSI were identified by the MSH Clinical Microbiology Laboratory as part of standard clinical care between July 2018 to May 2019. Identification and susceptibility of MRSA was performed by the Clinical Microbiology Laboratory using VITEK®2 (bioMerieux). Per hospital policy, all patients with MRSA BSI are placed on contact isolation with staff donning gown and gloves for interactions. Swabbers washed their hands prior to donning gown and gloves, and additionally wore surgical masks during sample collection. Bilateral anterior nares were sampled using a single, pre-labeled, sterile swab (Puritan® PurFlock® Ultra Sterile Flocked Collection Devices) moisturized with sterile 0.9% NaCl saline solution inserted up to 2.5 cm from the edge of the nares and rolled 5 times in each nostril. Swabs were plated directly on CHROMID® MRSA plates. After 24 h of incubation at 37 °C, the plate was evaluated for positive MRSA growth. If positive for MRSA, the isolate lawns were collected and used to inoculate Tryptic Soy Broth (TSB) with 15% glycerol cryovials. If negative for growth, the plates were re-evaluated at 48 h. When needed, Staphaurex™ Latex Agglutination Test was used to confirm coagulase positivity. The cryovial contents were homogenized using a Vortex-Genie 2 and then stored in a − 80 °C freezer.
For the molecular work up, the frozen stocks from the nasal swab stocks were plated on BD BBL™ Trypticase™ Soy Agar (TSA II™) with Sheep Blood and incubated at 37 °C. Selection of colonies was performed using a previously described replica plating technique [16] in which nasal swab stocks were diluted in 1X PBS and spread onto TSA II plates before incubation for 16 h at 37 °C to facilitate an even spread of distinct colonies. From these base plates we replica plated onto new TSA II Blood plate, BD BBL™ Prepared Plated Media: Mannitol Salt Agar, CHROMID® MRSA plates, and BD BBL™ Oxacillin Screen Agar (Mueller Hinton Agar w/6 µg/mL Oxacillin, 4% NaCl) Prepared Media plates, respectively. These replica plates were grown at 37 °C for 24 h before examination for phenotypic differences in colony morphology. The aim was to select approximately 6 (ideally morphologically diverse) colonies (Additional file 1: Table S1) for every swab sample, and bacterial stocks were prepared using TSB with 15% glycerol. All colonies were analysed using a Bruker MALDI Biotyper® System to confirm the growth of S. aureus.
Molecular typing
Single colonies from the clinical MRSA blood cultures underwent DNA extraction and PacBio whole genome sequencing as part of a surveillance program, as described previously [11]. From the resulting genomes a custom script (https://github.com/mjsull/spa_typing) was run to identify the spa type. The raw sequence data for the blood genomes have been deposited in the National Center for Biotechnology Information SRA database under Bioproject PRJNA470993.
DNA extraction of the nasal cultures was performed on a ThermoFisher KingFisher Flex 96 using Applied BioSystems MagMAX™ DNA Multi-Sample Ultra 2.0 Kits. Typing of the spa gene was performed by first using PCR to amplify the spa sequence through use of previously described methods [17], and then Sanger sequenced by Psomagen Inc. The Lasergene software package (version 17, DNAstar) was used to generate consensus sequences for spa and ST typing. Assignment of new STs was performed by submission of the DNA sequences of seven housekeeping genes (arcC, aroE, glpF, gmk, pta, tpi, and yqiL) to the online MLST database https://pubmlst.org/ [18]
Patient data collection and statistical analyses
Under approval obtained from the Mount Sinai Hospital Institutional Review Board (HS# 17-00825), demographic and clinical data were obtained retrospectively from the electronic medical record system, including admission data, presumed source of the BSI based on Infectious Diseases (ID) consults, comorbidities, and prior outpatient health care exposures. Chart abstraction and data entries were placed into the REDCap [19] system and verified by two different ID trained individuals. All patients diagnosed with MRSA BSI received an ID consultation at the time of diagnosis, as per standard practice at our institution. Surveillance definitions included hospital-onset MRSA (HO-MRSA), defined as positive cultures on or after the fourth day after hospital admission, and community-onset MRSA (CO-MRSA), defined as BSI presenting within the 72-h hospital admission interval [20]. We also evaluated in-hospital outcomes and death, especially those considered to be related to the MRSA BSI. We categorized all non–normally distributed continuous variables into discrete categorical groups. The univariate logistic regression model was performed initially, and variables with p values ≤ 0.2 were placed in the multivariable logistic regression model. All variables with p ≤ 0.05 were considered statistically significant. All analyses were performed in SAS (version 9.4) [21].
Discussion
Colonization is an important feature of invasive MRSA disease. In our study we found that 70% of patients with MRSA BSI were colonized with MRSA in the anterior nares. In 95% of these cases, the MRSA clones present in the blood were also found in the nares, suggesting infection from an endogenous source.
As nasal colonization was associated with history of a recent hospitalization within the prior 90 days, presumably these individuals may become colonized with exposure to the healthcare setting. While baseline colonization is not assessed on admission at our hospital, nasal colonization with MRSA has been independently associated with healthcare exposure in prior studies [22]. The relationship between recent hospitalization and colonization was implicated through the higher rates of CC5 amongst those with prior healthcare exposures and higher overall rates of colonization with CC5. This may signal differences in the avenues in which BSI due to CC5 and CC8 are acquired given the association of CC5 and the healthcare setting. Additionally, a higher proportion of male patients with MRSA BSI were found to be colonized when compared to women. This finding is consistent with prior studies and may be attributed to differences in the microbiome of the skin such as sweat, sebum, and hormone production along with differences in behavior and hand-hygiene practices [23, 24].
The overall genetic diversity found among the nasal clones was 11%. However, within-patient nasal cultures exhibited a high degree of relatedness with one another, with most clustering into the same genetic clades. A prior study focused on nasal carriage clonality estimated a 6.6% diversity [9], while one pediatric study found 30% diversity over longitudinal time points [25], suggesting that carriage of discordant S. aureus clones in individuals with nasal colonization occur regularly and may lead to horizontal genetic exchange among clones. Nonetheless, there is a paucity of data regarding S. aureus diversity in nasal colonization. With evaluations limited to a small number of colonies, the actual diversity is likely greater [26].
There were few overall differences in the clinical characteristics of patients infected with either CC5 or CC8 MRSA, upholding prior findings that indicate that historical hospital and community associations of clones are indeed blurred in endemic regions. Skin and soft tissue and peripheral intravenous catheter (PIV) infections tended to correlate more frequently to the CC8 MRSA. While CC8 is associated with community skin derived infections [27], we have previously observed that CC8 was also associated with PIV infections [13] highlighting the importance of community clones causing hospital-onset infections. Colonization was associated with patients with prior history of renal disease, which is a well-established clinical association given the frequent interaction with healthcare facilities and likelihood of requiring vascular access [28].
There are some limitations in this study. The patient population at our institution is complex and might not represent the same population in smaller community hospitals. Regional and global differences in the prevalence of MRSA colonization and clonal types may limit generalizability. The retrospective chart review portion may be subject to errors in chart abstraction. Although we expect that the numerically dominant clones are most likely to be selected, particularly in the setting of bacteremia, we are likely underrepresenting the full diversity [29]. While it is understood that people become infected with their colonizing isolate, we are unable to directly address directionality, as there may have been a single exposure that led to acquisition at both sites. Additionally, exposure to intravenous anti-MRSA antibiotics and/or choice of anti-MRSA antibiotics was not assessed and may have differed between colonized and non-colonized groups. Finally, we focused on MRSA-BSI due to the focus of our molecular surveillance program although we recognize that methicillin-susceptible S. aureus (MSSA) also causes significant disease [30].
Conclusions
Colonization is an essential component of invasive MRSA disease, and we found high rates of colonization, and colonization with more than one clone at a frequency of 11%. This contemporary sample collection adds to the paucity of literature on the frequency and the diversity of colonization in the presence of active MRSA BSI. We found colonization to be associated with male sex and those with a recent hospitalization, highlighting potential high-risk groups. Combined molecular and clinical analyses will assist in defining the intra-host and inter-host dynamics of MRSA and enable the development of targeted approaches to curtail disease.
Supplementary Information
Additional file 1: Table S1. MRSA isolate typing.
Additional file 2: Table S2. Comorbidities of patients with MRSA BSI with and without nasal colonization.
Additional file 3: Table S3. Outcomes of patients with MRSA BSI with and without nasal colonization.
Abbreviations
- HAI
Healthcare-associated infections
- MRSA
Methicillin-resistant S. aureus
- BSI
Bloodstream infections
- spa
Staphylococcal protein A
- S. aureus
Staphylococcus aureus
- CC
Clonal complex
- CO
Community-onset
- HO
Hospital-onset
- MSH
Mount Sinai Hospital
- TSB
Tryptic Soy Broth
- TSA II™
Trypticase™ Soy Agar
- ID
Infectious diseases
- HO-MRSA
Hospital-onset MRSA
- CO-MRSA
Community-onset MRSA
- ST
Sequence type
- MSSA
Methicillin-susceptible S. aureus
Authors contributions
ERS, AvdG, ACD, MC, DRA wrote the manuscript. ERS, AvdG, MC, DRA, MG, RS, HvB revised the manuscript. ERS, ACD, ABC, DN, LF, AM, RS, GP, DRA performed the clinical evaluations. AvdG, FS, JP, MG, MC, DRA, BA, HvB performed analysis of clinical samples. ACD, DRA performed statistical analyses. AO, KC, HvB assisted with data collection. AvdG, IO, MJS, MLS, BA, MC, HvB performed isolate sequencing. All authors read and approved the final manuscript.
Funding
This research was supported in part by R01 AI119145 (H.v.B.). Dr. Altman has received institutional funding and funding from participation in ContraFect corporation clinical trials. The funders had no role in the study design, data collection and interpretation, or the decision to submit the work for publication.
Availability of data and materials
The raw sequence data for the blood genomes that support the findings of this study have been deposited in the National Center for Biotechnology Information SRA database under Bioproject PRJNA470993; other data used and/or analysed during the current study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
The study protocols were reviewed and approved by the Icahn School of Medicine at Mount Sinai Institutional Review Board. Use and disclosure of protected health information (PHI) was approved under a waiver of authorization and a waiver of informed consent was obtained. Collection and processing of bacterial samples were covered with protocols HS # 18-00662 and # 13-00981, and the clinical data collection with the protocol HS # 17-00825. This study was performed in accordance with the Helsinki Declaration of 1964, and its later amendments.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Erika Reategui Schwarz and Adriana van de Guchte contributed equally to this work.
References
- 1.Lowy FD. Staphylococcus aureus infections. N Engl J Med. 1998;339(8):520–532. doi: 10.1056/NEJM199808203390806. [DOI] [PubMed] [Google Scholar]
- 2.Chambers HF, Deleo FR. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat Rev Microbiol. 2009;7(9):629–641. doi: 10.1038/nrmicro2200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wertheim HF, Melles DC, Vos MC, van Leeuwen W, van Belkum A, Verbrugh HA, et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect Dis. 2005;5(12):751–762. doi: 10.1016/S1473-3099(05)70295-4. [DOI] [PubMed] [Google Scholar]
- 4.Muthukrishnan G, Lamers RP, Ellis A, Paramanandam V, Persaud AB, Tafur S, et al. Longitudinal genetic analyses of Staphylococcus aureus nasal carriage dynamics in a diverse population. BMC Infect Dis. 2013;13:221. doi: 10.1186/1471-2334-13-221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.VandenBergh MF, Yzerman EP, van Belkum A, Boelens HA, Sijmons M, Verbrugh HA. Follow-up of Staphylococcus aureus nasal carriage after 8 years: redefining the persistent carrier state. J Clin Microbiol. 1999;37(10):3133–3140. doi: 10.1128/JCM.37.10.3133-3140.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kluytmans J, van Belkum A, Verbrugh H. Nasal carriage of Staphylococcus aureus: epidemiology, underlying mechanisms, and associated risks. Clin Microbiol Rev. 1997;10(3):505–520. doi: 10.1128/CMR.10.3.505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Davis KA, Stewart JJ, Crouch HK, Florez CE, Hospenthal DR. Methicillin-resistant Staphylococcus aureus (MRSA) nares colonization at hospital admission and its effect on subsequent MRSA infection. Clin Infect Dis. 2004;39(6):776–782. doi: 10.1086/422997. [DOI] [PubMed] [Google Scholar]
- 8.Wertheim HF, Vos MC, Ott A, van Belkum A, Voss A, Kluytmans JA, et al. Risk and outcome of nosocomial Staphylococcus aureus bacteraemia in nasal carriers versus non-carriers. Lancet. 2004;364(9435):703–705. doi: 10.1016/S0140-6736(04)16897-9. [DOI] [PubMed] [Google Scholar]
- 9.Cespedes C, Said-Salim B, Miller M, Lo SH, Kreiswirth BN, Gordon RJ, et al. The clonality of Staphylococcus aureus nasal carriage. J Infect Dis. 2005;191(3):444–452. doi: 10.1086/427240. [DOI] [PubMed] [Google Scholar]
- 10.von Eiff C, Becker K, Machka K, Stammer H, Peters G. Nasal carriage as a source of Staphylococcus aureus bacteremia. Study Group. N Engl J Med. 2001;344(1):11–16. doi: 10.1056/NEJM200101043440102. [DOI] [PubMed] [Google Scholar]
- 11.Berbel Caban A, Pak TR, Obla A, Dupper AC, Chacko KI, Fox L, et al. PathoSPOT genomic epidemiology reveals under-the-radar nosocomial outbreaks. Genome Med. 2020;12(1):96. doi: 10.1186/s13073-020-00798-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Senn L, Clerc O, Zanetti G, Basset P, Prod'hom G, Gordon NC, et al. The stealthy superbug: the role of asymptomatic enteric carriage in maintaining a long-term hospital outbreak of ST228 methicillin-resistant Staphylococcus aureus. MBio. 2016;7(1):e02039–e2115. doi: 10.1128/mBio.02039-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dupper AC, Sullivan MJ, Chacko KI, Mishkin A, Ciferri B, Kumaresh A, et al. Blurred molecular epidemiological lines between the two dominant methicillin-resistant Staphylococcus aureus clones. Open Forum Infect Dis. 2019;6(9):ofz302. doi: 10.1093/ofid/ofz302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Seybold U, Kourbatova EV, Johnson JG, Halvosa SJ, Wang YF, King MD, et al. Emergence of community-associated methicillin-resistant Staphylococcus aureus USA300 genotype as a major cause of health care-associated blood stream infections. Clin Infect Dis. 2006;42(5):647–656. doi: 10.1086/499815. [DOI] [PubMed] [Google Scholar]
- 15.Popovich KJ, Weinstein RA, Hota B. Are community-associated methicillin-resistant Staphylococcus aureus (MRSA) strains replacing traditional nosocomial MRSA strains? Clin Infect Dis. 2008;46(6):787–794. doi: 10.1086/528716. [DOI] [PubMed] [Google Scholar]
- 16.Lederberg J, Lederberg EM. Replica plating and indirect selection of bacterial mutants. J Bacteriol. 1952;63(3):399–406. doi: 10.1128/jb.63.3.399-406.1952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pardos de la Gandara M, Raygoza Garay JA, Mwangi M, Tobin JN, Tsang A, Khalida C, et al. Molecular types of methicillin-resistant Staphylococcus aureus and methicillin-sensitive S. aureus strains causing skin and soft tissue infections and nasal colonization, identified in Community Health Centers in New York City. J Clin Microbiol. 2015;53(8):2648–2658. doi: 10.1128/JCM.00591-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jolley KA, Bray JE, Maiden MCJ. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018;3:124. doi: 10.12688/wellcomeopenres.14826.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. Research electronic data capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42(2):377–381. doi: 10.1016/j.jbi.2008.08.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.CDC. Multidrug-Resistant Organisms & Clostrioides difficile Infection (MDRO/CDI) Module. 2018.
- 21.Base SAS 9.4 Procedures Guide: Statistical Procedures: SAS Institute Inc. SAS Institute; 2013.
- 22.Gorwitz RJ, Kruszon-Moran D, McAllister SK, McQuillan G, McDougal LK, Fosheim GE, et al. Changes in the prevalence of nasal colonization with Staphylococcus aureus in the United States, 2001–2004. J Infect Dis. 2008;197(9):1226–1234. doi: 10.1086/533494. [DOI] [PubMed] [Google Scholar]
- 23.Humphreys H, Fitzpatick F, Harvey BJ. Gender differences in rates of carriage and bloodstream infection caused by methicillin-resistant Staphylococcus aureus: are they real, do they matter and why? Clin Infect Dis. 2015;61(11):1708–1714. doi: 10.1093/cid/civ576. [DOI] [PubMed] [Google Scholar]
- 24.Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011;9(4):244–253. doi: 10.1038/nrmicro2537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mongkolrattanothai K, Gray BM, Mankin P, Stanfill AB, Pearl RH, Wallace LJ, et al. Simultaneous carriage of multiple genotypes of Staphylococcus aureus in children. J Med Microbiol. 2011;60(Pt 3):317–322. doi: 10.1099/jmm.0.025841-0. [DOI] [PubMed] [Google Scholar]
- 26.Paterson GK, Harrison EM, Murray GGR, Welch JJ, Warland JH, Holden MTG, et al. Capturing the cloud of diversity reveals complexity and heterogeneity of MRSA carriage, infection and transmission. Nat Commun. 2015;6:6560. doi: 10.1038/ncomms7560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.DeLeo FR, Otto M, Kreiswirth BN, Chambers HF. Community-associated meticillin-resistant Staphylococcus aureus. Lancet. 2010;375(9725):1557–1568. doi: 10.1016/S0140-6736(09)61999-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zacharioudakis IM, Zervou FN, Ziakas PD, Mylonakis E. Meta-analysis of methicillin-resistant Staphylococcus aureus colonization and risk of infection in dialysis patients. J Am Soc Nephrol. 2014;25(9):2131–2141. doi: 10.1681/ASN.2013091028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Didelot X, Walker AS, Peto TE, Crook DW, Wilson DJ. Within-host evolution of bacterial pathogens. Nat Rev Microbiol. 2016;14(3):150–162. doi: 10.1038/nrmicro.2015.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kourtis AP, Hatfield K, Baggs J, Mu Y, See I, Epson E, et al. Vital signs: epidemiology and recent trends in methicillin-resistant and in methicillin-susceptible Staphylococcus aureus bloodstream infections—United States. MMWR Morb Mortal Wkly Rep. 2019;68(9):214–219. doi: 10.15585/mmwr.mm6809e1. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Table S1. MRSA isolate typing.
Additional file 2: Table S2. Comorbidities of patients with MRSA BSI with and without nasal colonization.
Additional file 3: Table S3. Outcomes of patients with MRSA BSI with and without nasal colonization.
Data Availability Statement
The raw sequence data for the blood genomes that support the findings of this study have been deposited in the National Center for Biotechnology Information SRA database under Bioproject PRJNA470993; other data used and/or analysed during the current study are available from the corresponding author on reasonable request.