Abstract
Hypoglycemia is a common occurrence in critically ill patients and is associated with significant mortality and morbidity. We developed a machine learning model to predict hypoglycemia by using a multicenter intensive care unit (ICU) electronic health record dataset. Machine learning algorithms were trained and tested on patient data from the publicly available eICU Collaborative Research Database. Forty-four features including patient demographics, laboratory test results, medications, and vitals sign recordings were considered. The outcome of interest was the occurrence of a hypoglycemic event (blood glucose < 72 mg/dL) during a patient’s ICU stay. Machine learning models used data prior to the second hour of the ICU stay to predict hypoglycemic outcome. Data from 61,575 patients who underwent 82,479 admissions at 199 hospitals were considered in the study. The best-performing predictive model was the eXtreme gradient boosting model (XGBoost), which achieved an area under the received operating curve (AUROC) of 0.85, a sensitivity of 0.76, and a specificity of 0.76. The machine learning model developed has strong discrimination and calibration for the prediction of hypoglycemia in ICU patients. Prospective trials of these models are required to evaluate their clinical utility in averting hypoglycemia within critically ill patient populations.
Keywords: Hypoglycemia, Intensive care unit, Blood glucose, Critical care, Machine learning
1. Introduction
Hypoglycemia is common in hospitalized patients and has been linked to serious adverse events and mortality [1]. Both severe and moderate hypoglycemia are associated with increased risk of neurological impairment, cardiac arrhythmia, ischemia, stroke, seizures, and death [2–4]. Recent studies have found that hypoglycemia is common in critically ill patient populations, with an incidence between 10.1 and 45% [1, 5]. Furthermore, a retrospective analysis of US inpatient electronic health records (EHR) revealed that hypoglycemia was associated with a 66% increase in mortality risk and a 50% increase in length of hospitalization [6].
There is a clear impetus to reduce inpatient hypoglycemia. One strategy to prevent hypoglycemic episodes is to develop prediction tools that provide individualized risk scores. If a patient is deemed at high-risk for hypoglycemia, clinicians can provide appropriate interventions to reduce the likelihood of a hypoglycemic event, such as foregoing the use of insulin sliding scale among those at highest risk and increasing blood sugar thresholds for the sliding scale for those with modest risk.
In the last five years, several models which predict hypoglycemia within hospitalized patients have been developed using EHR datasets [7–11]. However, all previous studies have only predicted hypoglycemia among non-critically ill adult patients and have trained on data from a single hospital. Furthermore, previous studies used generalized linear modeling to predict hypoglycemic events and achieved only modest predictive power.
In this study, we aimed to develop and validate more complex machine learning models to predict the risk of hypoglycemia by using a large, multicenter intensive care unit (ICU) database.
2. Methods
2.1. Data set
The eICU Collaborative Research v2.0 database (eICU-CRD) was used for this study. The Philips eICU program is an integrated electronic medical record and analytics system that is deployed in hospitals across the United States. The Philips eICU Research Institute in partnership with the MIT Laboratory for Computational Physiology has previously developed and published the eICU-CRD, which is a de-identified, publicly-available multicenter critical care database [12]. It holds data for 200,859 admissions which represent 139,367 patient stays in 335 ICUs across the United States and is openly accessible at http://eicu-crd.mit.edu/. This database contains information on patient demographics, diagnoses, labs, vitals, medications administered throughout each patient’s ICU stay. The eICU-CRD was certified as de-identified and met Safe Harbor standards by an independent privacy expert (Privacert, Cambridge, Massachusetts, USA) (Health Insurance Portability and Accountability Act Certification No. 1031219-2). The eICU-CRD database has previously received ethical approval from the Institutional Review Boards of their hosting organizations. Since the database does not contain any personally identifiable or protected health information, a waiver for the requirement for informed consent was included in the IRB approval.
All patients within the eICU-CRD who had at least two blood glucose readings during their ICU stay were included in the study cohort. This inclusion criterion was applied across the entire multicenter dataset, regardless of the hospital where each patient was admitted. Patient demographics and prior diagnoses collected upon ICU admission were extracted. All laboratory values, vital sign recordings, and medication data from the first 2 h of each patient’s ICU stay were extracted from the dataset as well.
2.2. Outcome variable
The outcome of interest was the occurrence of a hypoglycemic event during a patient’s ICU stay. Hypoglycemia was defined as a blood glucose reading lower than 4 mmol/L (72.01 mg/dL), which is a standard cutoff used previously to define hypoglycemia [8, 9].
The outcome variable was binary. A label of 1 was assigned when a patient had any hypoglycemic event from 2 h after their ICU admission to the end of their ICU stay. If a patient did not have any hypoglycemic events during this period, the outcome variable was set to 0.
2.3. Predictor variables
A total of 44 candidate predictors were considered. These predictors included patient demographics (gender, age, ethnicity), diagnoses (diabetes, heart disease, kidney injury, etc.), laboratory results (sodium, albumin, creatinine, etc.), vital sign recordings (blood pressure, temperature, heart rate, etc.), and medications administered (insulin, dextrose, etc.). A detailed list of the predictor variables and their missingness is available in Table S1.
For all the laboratory results and vital sign recordings, the mean value of the variable prior to the second hour of the patient’s ICU stay was used as the predictor variable. The coefficient of variation of the blood glucose readings prior to the second hour was also used as a predictor variable. Additionally, the predictor variable associated with each medication was binary and was set to 1 if the patient was administered the medication within the first 2 h of ICU admission, otherwise it was set to 0.
Predictors which had missing entries over 40% were removed; the total serum protein, folate, triglycerides, lactate dehydrogenase, c-reactive protein, and bicarbonate were thus dropped. Missing data for all other features were determined using a 4-h fill-forward imputation or KNN-based imputation. The area under the receiver operating curve (AUROC) for the models trained on the KNN-imputed dataset and the 4-h fill-forward dataset differed by less than 1%, but the KNN method was ultimately chosen as it is considered more robust [13].
2.4. Statistical analysis and modeling
The patient cohort was randomly split into an 80% training/validation set and 20% testing set. A total of 15 different machine learning models were evaluated to predict hypoglycemia as shown in Table S2.
Each model was initially trained and evaluated using tenfold cross validation on the training/validation subset. The model with the highest AUROC was chosen for further development. The best predictive power was achieved with the eXtreme Gradient Boosting (XGBoost) algorithm [14].
A backwards stepwise feature selection algorithm was used to identify the 10 most important features for the XGBoost model (Fig. S1). An exhaustive grid search strategy was performed to identify the best-performing hyperparameters (e.g., tree depth, number of nodes, etc.). The continuous predicted output (ŷ) of each model was transformed into a binary output (0 or 1) by using a decision threshold (γ). A continuous output was transformed to 1 (patient is predicted to be hypoglycemic) if ŷ ≥ γ, otherwise it was labeled as 0 (patient is predicted to be non-hypoglycemic). In this study, the value of γ was selected to maximize the sum of sensitivity and specificity on the training dataset.
Finally, the XGBoost model was evaluated on the test dataset, and the model predictions were used to generate the AUROC plots. As the dataset is class-imbalanced, with the majority of patients being non-hypoglycemic, the sensitivity, specificity, and precision metrics were also computed to appropriately assess model performance. Model calibration was evaluated by building a reliability diagram and plotting the model-predicted probability of hypoglycemia against the observed probability in 10 discrete bins [15].
Models were developed using in Python 3.6 with widely used data science modules (pandas [16], numpy [17], tableone [18], and scikit-learn [19]). All data pre-processing, modeling, visualization, and analysis were performed with these modules. The code is publicly available at www.github.com/SreekarMantena/hypoglycemia-modeling.
3. Results
3.1. Patient cohort
In this study, data from 69,736 patients (38,121 males, 31,615 females) was analyzed.
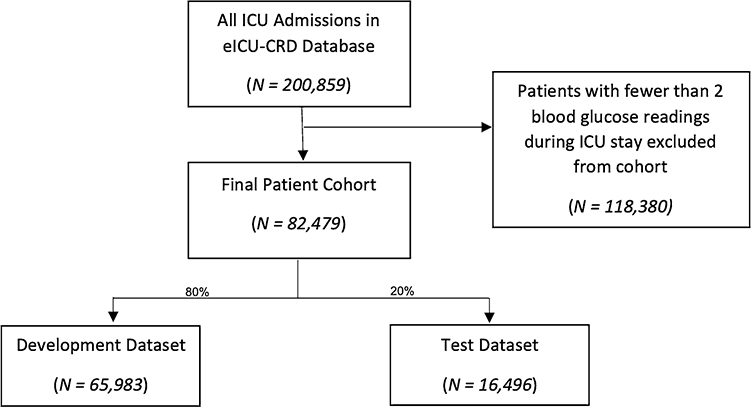
These patients underwent 82,479 hospital admissions in 199 hospitals in the US. A total of 118,380 patient admissions were excluded from the study cohort because they had fewer than two blood glucose readings, as shown in Fig. 1.
Fig. 1.
Flow diagram of patient cohort development
The incidence of hypoglycemia (blood glucose <4 mmol/L) was 19.9% in the study cohort. This is similar to the prevalence of hypoglycemia previously reported in surveys of ICU populations in US hospitals [5]. Notably, 38.7% of patients who experienced hypoglycemic events were non-diabetic. A summary of the patient demographics and predictor variables is shown in Table 1. In the patient cohort studied, the most common diagnoses at ICU admission were non-operative cardiovascular conditions (40.7%), followed by non-operative metabolic and endocrine conditions (25.2%), as presented in Table S3.
Table 1.
Demographics, lab values, vitals, and medication history of patients included in study
| Variable | Category | Overall | Non-hypoglycemic | Hypoglycemic | P Value |
|---|---|---|---|---|---|
|
| |||||
| n | 82,479 | 66,032 | 16,447 | ||
| Gender, n (%) | Female | 37,574 (45.6) | 29,440 (44.6) | 8134 (49.5) | < 0.001 |
| Male | 44,905 (54.4) | 36,592 (55.4) | 8313 (50.5) | – | |
| Age, mean (SD) | 63.7 (16.8) | 63.9 (16.7) | 62.9 (16.9) | < 0.001 | |
| Ethnicity, n (%) | African American | 10,500 (12.7) | 7690 (11.6) | 2810 (17.1) | – |
| Asian | 1018 (1.2) | 796 (1.2) | 222 (1.3) | – | |
| Caucasian | 62,258 (75.5) | 50,747 (76.9) | 11,511 (70.0) | – | |
| Hispanic | 3448 (4.2) | 2751 (4.2) | 697 (4.2) | – | |
| Native American | 662 (0.8) | 518 (0.8) | 144 (0.9) | – | |
| Other/Unknown | 3920 (4.8) | 3016 (4.6) | 904 (5.5) | – | |
| Systolic blood pressure, mean (SD) | 123.8 (24.5) | 124.8 (24.3) | 120.0 (25.0) | < 0.001 | |
| Diastolic blood pressure, mean (SD) | 68.2 (14.7) | 68.8 (14.7) | 65.9 (14.7) | < 0.001 | |
| SpO2, mean (SD) | 96.9 (3.2) | 96.9 (3.1) | 97.0 (3.4) | < 0.001 | |
| Temperature, mean (SD) | 36.7 (0.7) | 36.7 (0.7) | 36.6 (0.9) | < 0.001 | |
| Albumin, mean (SD) | 3.2 (0.6) | 3.3 (0.6) | 3.1 (0.7) | < 0.001 | |
| Creatinine, mean (SD) | 1.7 (1.9) | 1.6 (1.8) | 2.1 (2.2) | < 0.001 | |
| BUN, mean (SD) | 27.9 (22.8) | 26.6 (21.9) | 33.3 (25.5) | < 0.001 | |
| Hemoglobin, mean (SD) | 11.7 (2.5) | 11.8 (2.5) | 11.2 (2.5) | < 0.001 | |
| Diabetes, n (%) | 22,520 (27.3) | 16,160 (24.5) | 6360 (38.7) | < 0.001 | |
| Kidney disease, n (%) | 16,295 (19.8) | 11,390 (17.2) | 4905 (29.8) | < 0.001 | |
| Pancreatitis, n (%) | 919 (1.1) | 681 (1.0) | 238 (1.4) | < 0.001 | |
| Congestive heart failure, n (%) | 7387 (9.0) | 5681 (8.6) | 1706 (10.4) | < 0.001 | |
| Hypertension, n (%) | 11,999 (14.5) | 9679 (14.7) | 2320 (14.1) | 0.074 | |
3.2. Model performance
The performance of all 15 evaluated machine-learning models is shown in Table S2. The AUROC for the models ranged from 0.64–0.85, with a mean AUROC of 0.79. The best-performing model was the XGBoost model, with the standard gradient boosting model achieving slightly lower, but similar performance. XGBoost models are more computationally efficient than standard gradient boosting models, so the XGBoost model was ultimately selected as the final model [14].
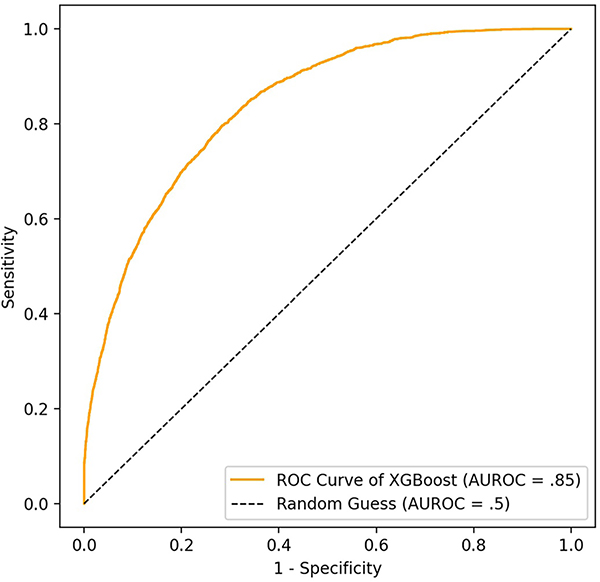
The XGBoost model achieved an AUROC of 0.85 and a sensitivity, specificity, and precision of 0.76, 0.76, and 0.44, respectively. A high sensitivity indicates that the algorithm can successfully identify the majority of patients that will experience hypoglycemia. The ROC curve for the final XGBoost model is depicted in Fig. 2, and the the feature importance of each predictor variable is presented in Fig. 3. Additionally, the XGBoost model achieved strong calibration. The predicted probabilities of hypoglycemia were similar to the observed probabilities, as shown in Fig. 4. The coefficients for the baseline logistic regression model are presented in Table S4.
Fig. 2.
ROC curve for the best-performing XGBoost model on test dataset
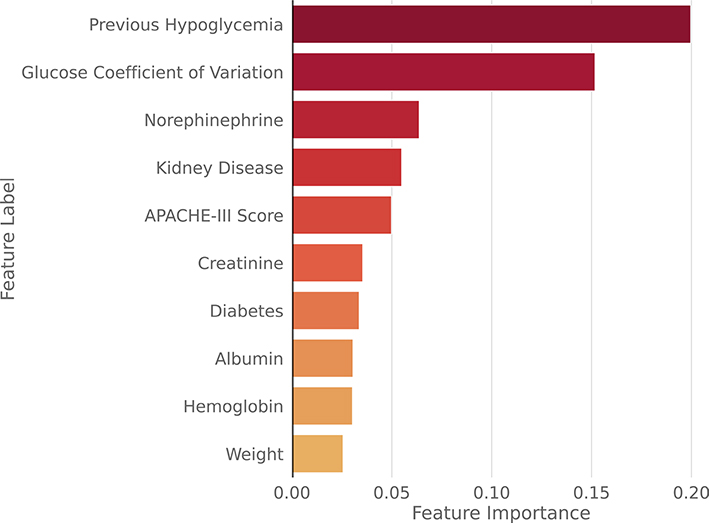
Fig. 3.
Feature importance from best-performing XGBoost model
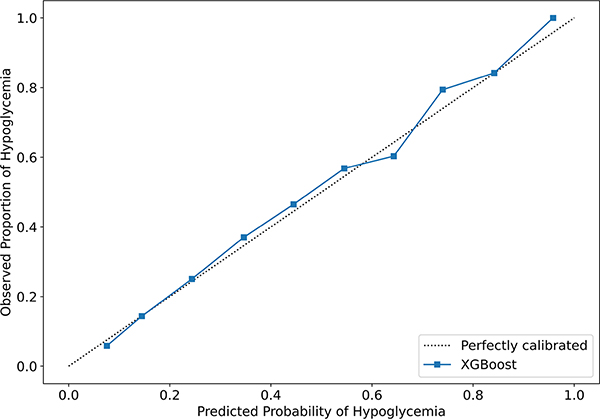
Fig. 4.
Reliability curve (calibration plot) showing predicted and observed probabilities of hypoglycemia for XGBoost model
4. Discussion
To our knowledge, this is the first study to develop a hypoglycemia prediction tool within a critically ill patient population. Developing hypoglycemia risk prediction tools for critically ill patients is particularly important, as hypoglycemia has the highest incidence and morbidity in ICU settings [20].
Using the eICU-CRD, we considered 82,479 admissions from 199 hospitals in the US and evaluated the performance of a variety of machine learning models in predicting hypoglycemic events. The final XGBoost model demonstrated strong predictive power on the held-out test set, achieving an AUROC of 0.85 for detecting hypoglycemia. The ensemble models tested (including the random-forest, gradient boosting, and XGBoost) all performed better than the baseline logistic regression model, with an improvement of 4% in AUROC. Furthermore, the final XGBoost model achieved strong calibration, as demonstrated by the reliability diagram in Fig. 4. The model-predicted probability of hypoglycemia and the observed probability of hypoglycemia are very similar across the full range of probabilities from 0 to 1. Thus, the model can be used to reliably predict a patient’s risk of hypoglycemia, rather than just providing a binary classification indicating whether or not a patient will experience a hypoglycemic event. Physicians can interpret model predictions close to 1 as indicating that a patient has a very high likelihood of experiencing hypoglycemia, while model predictions close to 0 indicate that the patient has a very low likelihood of experiencing hypoglycemia.
The most important feature for both the XGBoost model (Fig. 3) and the baseline logistic regression model (Table S4) was the incidence of a hypoglycemic event during a previous hospital admission. Albumin, creatinine, the coefficient of variation of blood glucose, kidney disease, and the administration of glucose-lowering drugs were also found to be key predictors of hypoglycemia, which is concordant with the results of previous studies [8–10, 21]. Hypoalbuminemia is a known risk factor for hypoglycemia and can be brought about by hemodilution in fluid resuscitation and capillary leak during a systemic inflammatory condition [22, 23]. Moreover, renal failure, evidenced by increased serum creatinine levels, is also known to contribute to hypoglycemia risk due to reduced insulin clearance and gluconeogenesis [2, 24]. Additionally, a diagnosis of diabetes was also among the most important features, as prior studies have shown [21].
This study has several key strengths and builds upon previous work in this field. All previous papers which modeled hypoglycemia prediction only trained on data from a single center. By leveraging EHR from 199 hospitals, the models developed in our study have stronger generalizability to diverse patient cohorts and are less likely to be overfitted to the patient population of a single hospital. Moreover, our study’s cohort of 82,479 ICU stays is up to ten times larger than the datasets used in previous work [8, 10, 11].
Furthermore, all predictor variables used in our models were available upon ICU admission or were collected within the first 2 h of a patient’s ICU stay. This is an important advantage of our approach that yields greater clinical feasibility. This prognostic model provides actionable information and could be built into a clinical decision support tool that automatically extracts the corresponding variables in the EHR and feeds these features into the model. Within 2 h of the patient’s ICU admission, physicians could leverage the prediction output of this model to determine if a patient has a high risk of developing a hypoglycemic event during their ICU stay and can tailor monitoring, prophylactic, and therapeutic measures accordingly. Clinicians can decide to adjust between high, medium or low insulin sliding scales, or decide to forego the scale altogether based on the risk. These interventions, which are informed by the model’s predictions, could enable physicians to reduce hypoglycemia and lower rates of hypoglycemia-associated adverse events. Improved glycemic control has also been shown to lead to economic benefits by reducing hospital length of stay, perioperative morbidity, and surgical site infections [25]. Furthermore, this model could be used to identify high-risk patients who stand to benefit the most from advanced treatment options, including closed-loop insulin delivery systems, that use continuous glucose monitoring to titrate insulin delivery [26].
Moreover, models developed in prior work used a large number of features (over 30) to predict hypoglycemic risk [8, 9]. In real-word clinical use, many of these laboratory tests may not be available for all patients, making model deployment challenging. The final XGBoost model developed in our work only uses 10 features, facilitating clinical deployment.
Additionally, 61.3% of hypoglycemic patients included in our study were non-diabetic, and many previous studies have focused on predicting hypoglycemia among diabetic patients [8, 10]. The results of our analysis suggest that prior approaches may fail to consider the significant portion of hypoglycemia cases which occur in non-diabetic patients.
As with any retrospective modeling effort, our study does have limitations. The availability of certain laboratory measurements (such as C-reactive protein) which were shown to be associated with hypoglycemia in previous studies widely varied across hospitals in the eICU-CRD database, so they were unable to be considered in the modeling.
Predictive models built using retrospective EHR data have significant potential to improve clinical care and have been shown to aid in reducing hypoglycemic events. A 2014 single-center interventional study used a linear regression model with moderate performance to identify patients with a high risk of developing hypoglycemia in real-time [7]. The implementation of the model resulted in a 68% decrease in hypoglycemic events in alerted high-risk patients [7]. Machine learning models which have greater predictive power, such as the one described in this study, could further reduce the incidence of hypoglycemia by more accurately identifying patients at who are at high-risk.
However, we recognize that there are many critical steps that must be taken before a clinical risk-prediction model can be implemented at the bedside. Multicenter prospective trials are needed to evaluate the effectiveness of these risk prediction tools in averting hypoglycemia in real-world clinical settings. Implementation research must be conducted so that these algorithms can be seamlessly integrated into a medical center’s workflow and clinicians can regularly review model predictions and optimize a patient’s treatment plan accordingly. Additionally, as these models are deployed in hospital environments, it is important to be cognizant of the fact that clinical and operational practices evolve over time. It will be critical to employ computational methods to identify dataset shift and proactively update and recalibrate models so they maintain robust performance in the years following model deployment [27].
In summary, our study uses a large, multi-center database to develop and validate a predictive model for hypoglycemia which achieves strong performance. Such tools could alert clinicians of patients at high-risk of hypoglycemia in real-time, informing clinical decision-making and reducing the occurrence of hypoglycemia-associated adverse events.
Supplementary Material
Funding
The work of ARA was supported by the PhD fellowship PD/BD/114107/2015 from Fundação da Ciência e da Tecnologia (FCT). The work of ARA, SMSV, and JMCS was supported through IDMEC, under LAETA, project UIDB/50022/2020; also, by the European Regional Development Fund (LISBOA-01-0145-FEDER-031474) and FCT through Programa Operacional Regional de Lisboa (PTDC/EMESIS/31474/2017). LAC is funded by the National Institute of Health through NIBIB Grant R01 E017215.
Footnotes
Conflict of interest The authors have no relevant financial or non-financial interests to disclose.
Ethical approval The study is exempt from institutional review board approval due to its analysis of data that has been de-identified and its security schema, for which the re-identification risk was certified as meeting safe harbor standards by an independent privacy expert (Privacert, Cambridge, MA) (Health Insurance Portability and Accountability Act Certification no. 1031219-2).
Supplementary Information The online version contains supplementary material available at https://doi.org/10.1007/s10877-021-00760-7.
Data availability
The data used in this paper is from the eICU-CRD database. The access to this dataset is controlled and researchers should request access on the PhysioNet website (https://physionet.org/about/database/). All code used in this study is available on a GitHub repository (www.github.com/SreekarMantena/hypoglycemia-modeling).
References
- 1.The NICE-SUGAR Study Investigators. Hypoglycemia and Risk of Death in Critically Ill Patients. New Engl J Med. 2012; 367:1108–18. [DOI] [PubMed] [Google Scholar]
- 2.Hulkower RD, Pollack RM, Zonszein J. Understanding hypoglycemia in hospitalized patients. Diabetes Manag. 2014;4:165–76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brutsaert E, Carey M, Zonszein J. The clinical impact of inpatient hypoglycemia. J Diabetes Complications. 2014;28:565–72. [DOI] [PubMed] [Google Scholar]
- 4.Krinsley JS, Grover A. Severe hypoglycemia in critically ill patients: Risk factors and outcomes. Crit Care Med. 2007;35:2262–7. [DOI] [PubMed] [Google Scholar]
- 5.Cook CB, Kongable GL, Potter DJ, Abad VJ, Leija DE, Anderson M. Inpatient glucose control: a glycemic survey of 126 U.S. hospitals. J Hosp Med. 2009;4:E7–14. [DOI] [PubMed] [Google Scholar]
- 6.Brodovicz KG, Mehta V, Zhang Q, Zhao C, Davies MJ, Chen J, et al. Association between hypoglycemia and inpatient mortality and length of hospital stay in hospitalized, insulin-treated patients. Curr Med Res Opin. 2013;29:101–7. [DOI] [PubMed] [Google Scholar]
- 7.Kilpatrick CR, Elliott MB, Pratt E, Schafers SJ, Blackburn MC, Heard K, et al. Prevention of inpatient hypoglycemia with a real-time informatics alert. J Hosp Med. 2014;9:621–6. [DOI] [PubMed] [Google Scholar]
- 8.Ruan Y, Bellot A, Moysova Z, Tan GD, Lumb A, Davies J, et al. Predicting the risk of inpatient hypoglycemia with machine learning using electronic health records. Diabetes Care. 2020;43:1504–11. [DOI] [PubMed] [Google Scholar]
- 9.Mathioudakis NN, Everett E, Routh S, Pronovost PJ, Yeh H-C, Golden SH, et al. Development and validation of a prediction model for insulin-associated hypoglycemia in non-critically ill hospitalized adults. BMJ Open Diabetes Res Care. 2018;6:e000499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Stuart K, Adderley NJ, Marshall T, Rayman G, Sitch A, Manley S, et al. Predicting inpatient hypoglycaemia in hospitalized patients with diabetes: a retrospective analysis of 9584 admissions with diabetes. Diabet Med. 2017;34:1385–91. [DOI] [PubMed] [Google Scholar]
- 11.Elliott MB, Schafers SJ, McGill JB, Tobin GS. Prediction and prevention of treatment-related inpatient hypoglycemia. J Diabetes Sci Technol. 2012;6:302–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pollard TJ, Johnson AEW, Raffa JD, Celi LA, Mark RG, Badawi O. The eICU collaborative research database, a freely available multi-center database for critical care research. Sci Data. 2018. 10.1038/sdata.2018.178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bertsimas D, Pawlowski C, Zhuo YD. From predictive methods to missing data imputation: an optimization approach. J Machine Learning Res. 2018;18(1):7133–71. [Google Scholar]
- 14.Chen T and Guestrin C. XGBoost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. 2016; 785–794. [Google Scholar]
- 15.Steyerberg EW. Clinical prediction models: a practical approach to development, validation and updating. Cham: Springer; 2019. [Google Scholar]
- 16.McKinney W Data Structures for Statistical Computing in Python. Proceedings of the 9th Python in Science Conference. 2010; 56–61. [Google Scholar]
- 17.van der Walt S, Colbert SC, Varoquaux G. The NumPy array: a structure for efficient numerical computation. Computing in Science and Engineering. 2011;13:22–30. [Google Scholar]
- 18.Pollard TJ, Johnson AEW, Raffa JD, Mark RG. tableone: an open source Python package for producing summary statistics for research papers. JAMIA Open. 2018;1:26–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pedregosa F, Michel V, Grisel O, Blondel M, Prettenhofer P, Weiss R, et al. Scikit-learn: machine learning in Python. J Mach Learn Res. 2011;12:2825–30. [Google Scholar]
- 20.Lacherade JC, Jacqueminet S, Preiser JC. An overview of hypoglycemia in the critically Ill. J Diabetes Science and Technology. 2009;3:1242–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pratiwi C, Mokoagow MI, Made Kshanti IA, Soewondo P. The risk factors of inpatient hypoglycemia: a systematic review. Heliyon. 2020;6(5):e03913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Leibovitz E, Wainstein J, Boaz M. Association of albumin and cholesterol levels with incidence of hypoglycaemia in people admitted to general internal medicine units. Diabet Med. 2018;35:1735–41. [DOI] [PubMed] [Google Scholar]
- 23.McCluskey A, Thomas AN, Bowles BJM, Kishen R. The prognostic value of serial measurements of serum albumin concentration in patients admitted to an intensive care unit. Anaesthesia. 1996;51:724–7. [DOI] [PubMed] [Google Scholar]
- 24.Arem R Hypoglycemia associated with renal failure. Endocrinol Metab Clin North Am. 1989;18:103–21. [PubMed] [Google Scholar]
- 25.Krinsley J, Schultz MJ, Spronk PE, van Braam Houckgeest F, van der Sluijs JP, Mélot C, et al. Mild hypoglycemia is strongly associated with increased intensive care unit length of stay. Ann Intensive Care. 2011. 10.1186/2110-5820-1-49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Salinas PD, Mendez CE. Glucose management technologies for the critically ill. J Diabetes Sci Technol. 2019;13(4):682–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Finlayson SG, Subbaswamy A, Singh K, Bowers J, Kupke A, Zittrain J, et al. The clinician and dataset shift in artificial intelligence. N Engl J Med. 2021;385(3):283–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data used in this paper is from the eICU-CRD database. The access to this dataset is controlled and researchers should request access on the PhysioNet website (https://physionet.org/about/database/). All code used in this study is available on a GitHub repository (www.github.com/SreekarMantena/hypoglycemia-modeling).