Abstract
The real‐time identification of protein biomarkers is crucial for the development of point‐of‐care and portable devices. Here, we use a PlyAB biological nanopore to detect haemoglobin (Hb) variants. Adult haemoglobin (HbA) and sickle cell anaemia haemoglobin (HbS), which differ by just one amino acid, were distinguished in a mixture with more than 97 % accuracy based on individual blockades. Foetal Hb, which shows a larger sequence variation, was distinguished with near 100 % accuracy. Continuum and Brownian dynamics simulations revealed that Hb occupies two energy minima, one near the inner constriction and one at the trans entry of the nanopore. Thermal fluctuations, the charge of the protein, and the external bias influence the dynamics of Hb within the nanopore, which in turn generates the unique ionic current signal in the Hb variants. Finally, Hb was counted from blood samples, demonstrating that direct discrimination and quantification of Hb from blood using nanopores, is feasible.
Keywords: Biosensors, Haemoglobin Variants, Molecular Modelling, Protein Dynamics, Realtime
The dynamics of single haemoglobin molecules trapped in the biological nanopore PlyAB are studied with single‐channel recording experiments and computational modelling. This approach enabled the differentiation of several haemoglobin variants, including haemoglobin from healthy adults (HbA) and sickle cell anaemia haemoglobin (HbS), with near 100 % accuracy and allowed for the quantification of haemoglobin directly from blood.
Introduction
The ability to measure biological molecules in real‐time or with minimal sample preparation at the point‐of‐care (POC) would save lives and reduce the cost of healthcare. The measurement of the haemoglobin (Hb) concentration is a routine blood test, [1] and it is especially important for patients with anaemia diseases. [2] Blood samples are usually sent to hospital laboratories where several tests are performed to measure the concentration of Hb.[ 3 , 4 , 5 ] In more complex tests, different Hb variants are distinguished. For instance, to diagnose the presence of sickle cell anaemia Hb (HbS), which differs by only a single amino acid (E6V) from the healthy Hb in adults (HbA), should be detected. Agarose or cellulose electrophoresis is performed to separate both variants based on their charge difference.[ 6 , 7 ] For the relative quantification of the variants, however, which could spontaneously co‐exist in heterozygote (A/S) type patients, high‐performance liquid chromatography (HPLC) or capillary electrophoresis (CE) is needed.[ 8 , 9 ] Despite these being standard techniques, they remain time‐consuming and labour‐intensive, requiring several hours from specialized personnel. As POC and personalized medicine are gaining momentum,[ 10 , 11 ] the development of a portable and low‐cost device for the rapid detection and quantification of Hb variants, amongst other proteins, becomes important. Ionic currents through nanopores are emerging as a powerful single‐molecule tool for biomacromolecule sensing,[ 12 , 13 ] and have been implemented in low‐cost portable devices for DNA sequencing.[ 14 , 15 , 16 , 17 ] More recently, nanopores have also been utilized for the detection and analysis of proteins. Solid‐state nanopores[ 18 , 19 ] measured the size and shape of chemically modified proteins anchored to the nanopore,[ 20 , 21 ] ubiquitination modifications, [22] and the conformational flexibility of proteins translocating nanopores.[ 23 , 24 ] Biological nanopores generally perform better as sensors compared their solid‐state counterparts. However, the detection of folded proteins with such pores, which typically have diameters in the 1 to 2 nm range, necessitates the attachment of a binding element at the entrance of the nanopore.[ 25 , 26 , 27 , 28 , 29 , 30 ] Although the identification of protein modifications, such as glycosylation could be readily observed,[ 28 , 29 ] it is unlikely these sensors will be capable of identifying small differences such as mutations in bound proteins. Nanopores with a larger diameter, such as cytolysin A (ClyA)[ 31 , 32 ] and fragaceatoxin C (FraC)[ 33 , 34 , 35 ] were introduced in the past decade that can sample proteins up to ≈40 kDa lodged inside the nanopore lumen.[ 34 , 36 , 37 , 38 , 39 , 40 , 41 , 42 ] The analysis of a dozen proteins, as markers for metabolomic analysis, revealed that most proteins have a unique signal that could allow their identification in solution. [41] Large post‐translational modifications such as ubiquitination have also been observed and characterised. [43] More recently, Mycobacterium smegmatis Porin A (MspA) was used to sample a few small proteins, enabling their identification and quantification from the same buffered solution. [44]
Here we show that the two‐component nanopore pleurotolysin AB [45] (PlyAB‐E1 [46] ), which has a large cylindrical shape due to its β‐barrel channel (Figure 1a), can discriminate HbA and HbS (64.5 kDa), which differ in just one amino acid (Figure 1b). Continuum modelling was used to elucidate the physical mechanisms behind the observed difference in the measured signals. Inside the nanopore, Hb fluctuates between two equilibrium positions, which arise from the delicate balance between hydrodynamic and electromechanical forces exerted on Hb. The complex protein signal arises from thermally induced swapping between the two minima, a process that could be described by stochastic simulations. Finally, we also demonstrate direct quantification of Hb from blood samples. Hence, this work reveals that PlyAB nanopores provide a new sensor for real‐time detection and identification of large, folded proteins, such as Hb.
Figure 1.
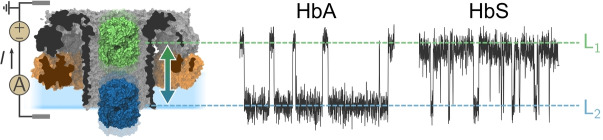
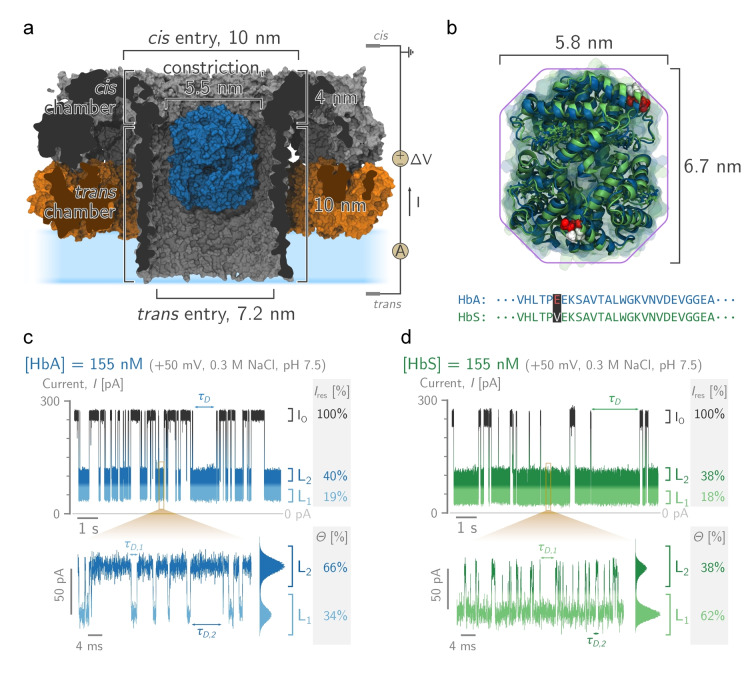
HbA and S blockades in PlyAB nanopores. a) Molecular surface representation of a single human haemoglobin A (HbA, in blue, PDBID: 2DN1 [47] ) tetramer placed within the trans chamber of a PlyAB nanopore (PlyA and PlyB in orange and grey respectively, PDB ID: 4V2T[ 45 , 46 ]). b) Structural alignment between oxygenated human HbA (in blue, PDB ID: 2DN1 [47] ) and the sickle cell disease HbS (in green, PDBID: 5E83 [48] ). The primary structures of HbA and HbS differ only by a single point mutation in the β‐subunits (E6V, coloured spheres). c) and d) are typical Hb blockades to PlyAB nanopores for HbA (blue) and HbS (green), respectively. In the enlarged trace, the deep and shallow levels are assigned to L1 (light shades) and L2 (dark shades), respectively. A whole trace histogram is shown on the side of the blockade. Hb ( ) is added to the trans side. The recordings were performed at in NaCl, buffered at pH 7.5. Currents were sampled at and filtered using a low‐pass Bessel filter. Molecular structures were rendered using VMD.[ 49 , 50 ]
Results and Discussion
Phenomenology of Hb Trapping in PlyAB
HbA represents the most abundant Hb in healthy people (over 95 % of the total Hb). HbS, which contains a single glutamic acid to valine substitution at position 6 of β‐chain subunits (E6V, Figure 1b), provokes sickle cell anaemia. Here, we added Hb to the trans side of PlyAB‐E1 (Figure 1a), an engineered variant of PlyAB that was selected for its higher solubility during nanopore preparation, [46] in a 300 mM NaCl solution at pH 7.5. Under positive bias voltages (applied to the trans reservoir), both HbA and HbS, added to the trans side of PlyAB, elicited well‐defined blockades (Figures 1c, d), whilst at negative potentials, no capture events were observed (Figure S1). The inner surface of PlyAB‐E1 is negatively charged, making the nanopore slightly cation‐selective and generating a strong electro‐osmotic flow (EOF) from trans to cis under positive applied potentials. [46] At pH 7.5 and +50 mV, the blockades ( ) of the open pore current ( ) elicited by both HbA and HbS (Figures 1c, d) contained two distinct residual current ) levels: a “deep”, low current L1 at (HbA) or (HbS) and a “shallow”, high current L2 at (HbA) or (HbS). The blockades levels rapidly switched between one another, with mean L2 dwell times ) of 1.7±0.5 ms (≈600 Hz) and 0.8±0.2 ms (≈1300 Hz) for HbA and HbS, respectively. Notably, whereas HbA lodged mainly in the shallow L2, as shown by the L2 fractional occupancy , HbS showed the opposite behaviour with (Figure 1d). Most likely, the two blockades represent the lodging of the Hb molecules in two different residence sites inside the nanopore, as previously observed for thrombin in ClyA nanopores, [32] and the small difference in charge between HbA (≈−3.74 e) and HbS (≈−1.74 e) at pH 7.5 is sufficient to shift the balance from L2‐dominant for HbA to L1‐dominant for HbS.
A Physical Model of Hb Trapping in PlyAB
To better understand the origin of the experimentally observed blockade levels and to estimate the forces acting on Hb, we used 2D‐axisymmetric continuum modelling to model the transport of a Hb‐like particle through PlyAB. As previously explained,[ 46 , 51 ] PlyAB, the lipid bilayer, and Hb were modelled as solid dielectric blocks. The geometry and charge distribution of the pore was constructed from an all‐atom homology model of PlyAB‐E1, effectively maintaining the corrugated surface and complex charge distribution of the nanopore. Hb proteins, however, do not show the same level of rotational symmetry (i.e., compared to PlyAB nanopores) and were hence instead approximated by a smooth cylinder‐like particle ( , ) with a uniform charge density ( , Figures 1b, 2a and S2). The relation between the protein charge and the solution pH was calculated from their molecular structures (Figure S3). Due to the axial symmetry of our model, the geometrical centre of the Hb particle was placed on the symmetry axis (i.e., ), where its z‐position could be chosen freely (i.e., ). Next, finite element analysis (COMSOL Multiphysics v5.5, COMSOL Inc., Burlington, MA, USA) was used to numerically solve the extended Poisson‐Nernst‐Planck‐Navier‐Stokes (ePNP‐NS) equations [51] for different nanopore‐Hb configurations as a function of (−100 nm≤ ≤100 nm), (0≤ ≤−20 e) and, (0≤ ≤+100 mV, see Supporting Information Sec. 3.1 for details). This produced a detailed description of the ionic and water currents flowing through the pore, together with the hydrodynamic ( ) and electromechanical ( ) forces acting on the Hb molecule. The sum of these forces, , represents the total external force exerted on Hb at equilibrium, and the cumulative integration of over yields the potential energy landscape ).
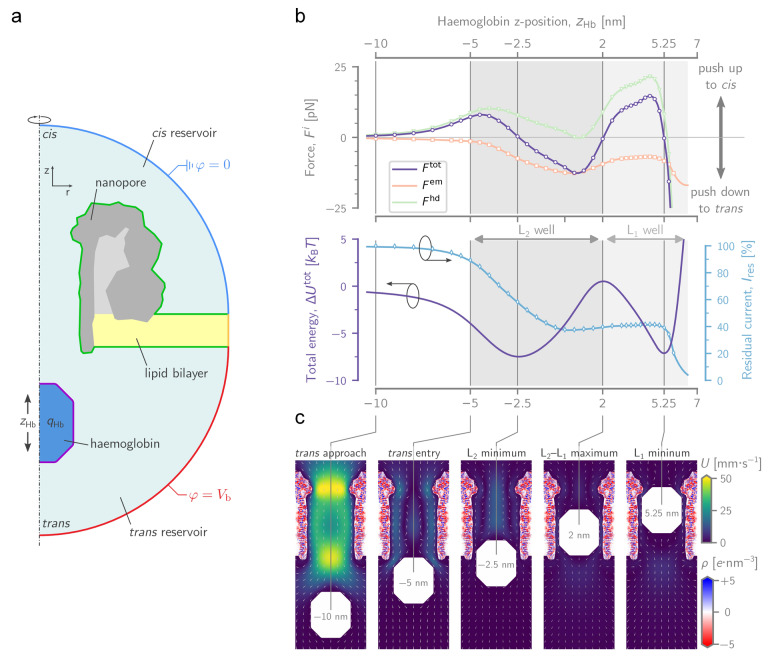
An example of a force, energy, and residual current landscape, for the case of −4 e and is shown in Figure 2b, together with several cross‐sectional heatmaps of the water velocity at several key positions (Figure 2c). As Hb traverses the pore from trans to cis, pulls Hb towards the trans entry, whereas pushes it towards the cis entrance. In our model, we found that only starts to push Hb back towards the trans side once it nears the constriction, due to a build‐up of pressure. Because both and have closely matched but opposing magnitudes ( ), their complex dependence on results in a with several equilibrium points (i.e., locations where ), notably at and (Figures 2b, c). These equilibrium points translate into local energy minima or energy wells, which in turn delimit a zone where Hb can remain trapped for a significant amount of time. Notably, the residual currents at ( ) and ( ) are highly reminiscent of the shallow L2 and the deep L1 levels observed in the experiment, and hence strongly suggest that Hb physically sits at these locations in the nanopore.
Figure 2.
A computational model of Hb trapping in PlyAB. a) Cartoon of the setup used in finite element modelling simulations, showing the geometry of the PlyAB pore (grey), the lipid bilayer (yellow), and the Hb particle (dark blue). b, top) Force landscape of Hb as it moves through PlyAB from trans to cis. The total force ( , purple) is the sum of the hydrodynamic ( , green) and the electromechanical ( , orange) force. Because both forces mostly oppose each other and are of equal magnitude, their fluctuations result in several locations where the total force on Hb is zero (a positive force means that Hb is pushed up, i.e., towards cis, and vice versa). b, bottom) On the energy landscape (purple), these points of zero force are translated into two local energy minima ( and ), separated by a local maximum ). The minima are presumed to correspond to the L2 and L1 current levels observed experimentally, respectively. The residual current landscape (blue) shows a rapid decline from 100 to 40 % as the Hb particle enters the lumen, after which it remains stable at until Hb reaches the cis constriction. c) Cross‐sectional heatmaps of the PlyAB‐Hb simulation at several key Hb locations ). The colouring shows the magnitude of the water velocity field ( ), whose direction is indicated by the arrows. The complex charge density of PlyAB ( ) is also drawn within the solid dielectric of the pore. Data in (c and Figure 2b) were for ( for HbA, for HbS, see Figure S8) and at .
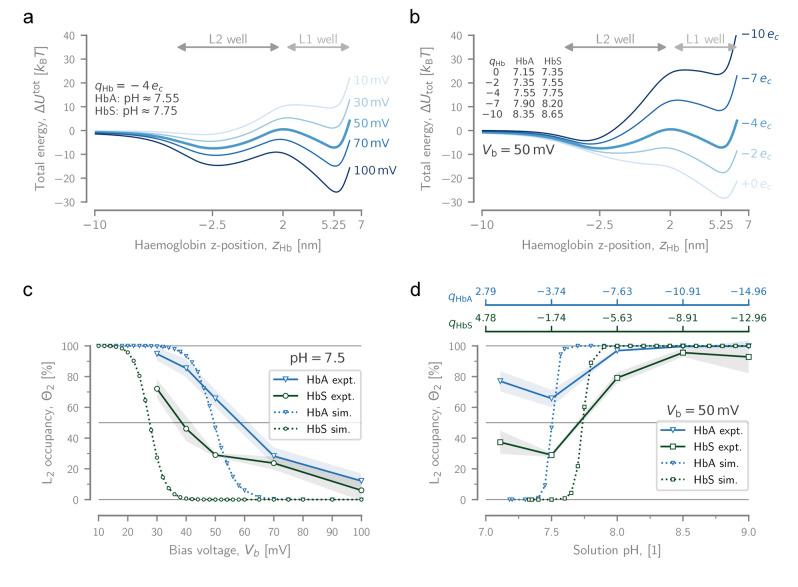
Under virtually all conditions, the simulated energy landscapes contain these two distinct energy minima. Their exact location, width, and depth, however, exhibit a strong dependence on both the applied bias voltage (Figures 3a and S4) and the Hb charge (Figures 3b and S5). Increasing the positive applied bias voltage effectively tilts the total energy landscape towards the cis side of the pore in favour of the deeper L1 position (Figure 3a). This matches the experimentally observed decrease in L2 occupancy as a function of applied bias (Figure 3c and Table S1). Interestingly, experiments also showed that the of L1 remained virtually constant (Figure S6 and Table S1), and that of L2 decreased with increasing positive applied bias (Figure S7a and Table S1). This suggests that the L1 and L2 binding sites are located at a rigid (e.g., near the constriction of the nanopore) and a malleable (e.g., near the trans entry of the pore) position, respectively. This is indeed corroborated by the energy landscapes where the width and position of the deep energy minimum at barely changes with voltage, whereas the location of the shallow minimum at shifts deeper into the pore, towards lower residual currents. Likewise, higher pH values, which increase the negative charge of Hb, tilt the energy landscape of Hb inside PlyAB towards the trans side of the pore (Figure 3b). As experimentally observed, this both increases the likelihood of L2 occupation (Figure 3d and Table S2), but also increases the observed for that level with increasing pH (Figure S7b and Table S2).
Figure 3.
Experimental and modelled dynamics of Hb. Dependence of the total potential energy for a Hb particle, , as it translocates through PlyAB from trans to cis on a) the applied bias voltage (at a fixed Hb charge: ), and b) the particle charge (at a fixed voltage: ). Note that corresponds to a solution pH‐values of and for HbA and HbS, respectively. The approximate pH values corresponding to the plotted charges listed on the graph, and a titration curve can be found in the Supporting Information (Figure S8). Dependence of the fractional occupation of L2 for HbA and HbS, as measured experimentally and estimated with simulation, on c) the applied bias voltage (at pH 7.5), and d) the solution pH (at ). At pH 7.5, the net charges of HbA and HbS are and , respectively. The shaded areas behind the experimental curves represent the standard deviation ( ). The simulation data only show conditions in which the L2 state is present as a distinct local minimum.
Furthermore, the overall dwell time of Hb blockades reached a maximum at and then decreased again (Figure S8a and Table S1), a behaviour associated with the translocation of proteins across nanopores after a threshold potential. [40] Even though we modelled both PlyAB and Hb as rigid bodies, whose geometries prohibit full translocation, our simulations corroborated these findings. Because higher positive voltages effectively tilt the energy landscape towards the cis side (Figure 3a), it becomes more difficult for Hb to escape back to trans, which facilitates the translocation through the constriction. A mirrored dwell time dependency is observed for the pH, where a maximum is observed around pH 7.5 (Figure S8b and Table S2). As the pH of the solution is increased, Hb acquires a more negative charge, which under positive potentials tilts the energy landscape towards trans. In turn, the likelihood of escaping back to trans increases.
To quantify the dynamic exchange between the two energy minima, 1D Brownian dynamics (BD) simulations were used, in which thermal fluctuations were added on top of the energy landscapes (Figure S9), allowing the particle to move as a function of time. Similar to the experiment, the BD simulations yielded a current trace, but also a position trace (Figure S10). Note that given the simplifications and assumptions of our PlyAB‐Hb model, this system is expected to capture the essence rather than the details of the current blockades measured experimentally (see Supporting Information for more details). Experimentally, the relative occupancy of L2 ( ) decreased with increasing voltage from to , from to , and from to for HbA and HbS, respectively (Figure 3c and Table S1). As a function of pH (i.e., Hb charge) the effect is reversed, with rising from to for HbA and from to when increasing the pH from 7.5 to 9.0 (Figure 3d and Table S2). In both cases, the BD simulations predicted a similar outcome, where the Hb landscapes favour the deeper L1 with increasing voltage (Figure 3a), whereas the shallower L2 is preferred when the pH is increased (Figure 3b). Notably, even though the transitions from L2‐ to L1‐dominant occupancy (and vice versa) are more gradual in the experiment compared to the simulation, the threshold voltage/pH at which the level preference begins to change, and the relative offsets between HbA and HbS, do match closely.
Identification and Quantification of Hb Variants
A precise Hb quantification in solution is important because both HbS and HbA types exist in the blood of heterozygote (A/S) type sickle cell anaemia patients. To identify both proteins in the solution, HbA or HbS should be identifiable at the single blockade level. We found that the L2 values of HbA were higher than HbS, under all potentials and pH values tested (Figure S7 and Tabs. S1 and S2), the reasons for which were outlined above. At pH 7.5 and , HbA and HbS showed the largest difference, and under these conditions, most blockades could be correctly assigned to either HbA or HbS using an threshold value of for L2. Analysing 600 blockades using three nanopores (Figure S11), we found that of single blockades collected from HbA events had an higher than the threshold, while of HbS events showed lower values. By contrast, the fractional occupancy of L2 was a less accurate predictor, as only of HbA blockades showed higher fractional L2 occupancy compared to L1. These data indicate that PlyAB nanopores are remarkably sensitive to single (charged) amino acid mutations in large, folded proteins.
In infants, oxygen is carried by human foetal Hb (HbF, α2γ2), which is more efficient as an oxygen carrier than adult Hb. In children HbF and HbA/HbS might both be present, and therefore the early diagnosis of sickle cell anaemia is complex, as HbF compensates for the lower oxygen transport of HbS. Because generating a high level of HbF in the patient's blood is advantageous during the treatment, [54] it is important to discriminate HbF from other variants for early sickle cell anaemia diagnosis and prognosis. HbF, HbA, and HbS have the same overall shape (Figure 4a), given that they contain two identical α‐subunits, but different γ‐/β‐chains with of sequence identity (Figure 4b). When HbF was added to the trans side of a PlyAB nanopore, it showed two distinctive levels, as previously observed by HbA or HbS. However, with residual currents of and for L1 and L2, respectively, the blockades were less deep compared to either HbA or HbS (Figures 4c and S12). All HbF blockades could be distinguished from HbA or HbS blockades. Therefore, PlyAB‐E1 nanopores can discriminate mixtures of the three different Hb variants (Figure 4d). Interestingly, unlike the single substitution in HbS, the numerous amino acid differences between HbA and HbF do not give rise to a change in the net charge (Figure S8). They do impact, however, the distribution of those charges within Hb. Given that we use a uniform charge density for Hb, our model can only differentiate based on an absolute charge, and hence it cannot discern HbF from HbA.
Figure 4.
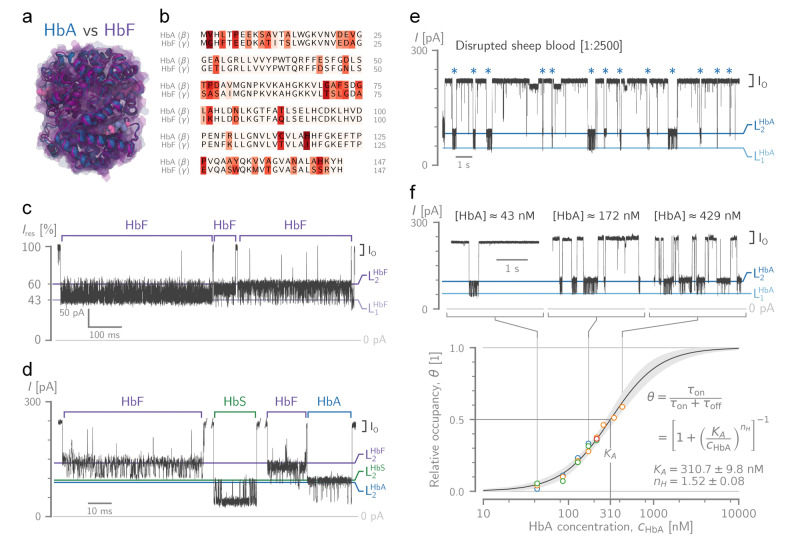
Discrimination of human foetal haemoglobin (HbF) from HbA and HbS and direct quantification of Hb from blood. a) Structural alignment between human HbA (α2β2, in blue, PDB ID: 2DN1 [47] ) and HbF (α2γ2, in purple, PDBID: 4MQJ [52] ). The quaternary structures of HbA and HbF share the same α subunits, but in HbF, the γ‐globin chains replace the β ones. b) The sequence alignment of the β‐globin chain of HbA and the γ‐chain of HbF. Their amino acid differences are highlighted with red shading. c) Typical traces of HbF trapping ( in trans). d) Discrimination of Hb variants in a mixture of HbF, HbA, and HbS (all at in trans). e) Typical protein blockades from sampling sheep blood. Red blood cells were disrupted by adding n‐dodecyl‐β‐D‐maltoside (DDM) and then a 1 : 2500 blood dilution (final) was sampled by the PlyAB nanopore. Hb events are indicated by a blue asterisk. f) Hb concentration ( ) calibration traces (top) and the resulting curve (bottom) using Hb isolated from sheep blood and quantified spectroscopically. Using least‐squares non‐linear regression, the Hill–Langmuir equation was fitted to the ‐dependence of the fraction of time the pore was blocked by Hb ( , yielding a half occupation concentration and a Hill coefficient of . The shading indicates the 3σ confidence interval of the fit. All recordings were performed at in NaCl, pH with a sampling rate and low‐pass Bessel filter. Molecular structures were rendered using VMD.[ 49 , 50 ] The sequence alignment was created with the Biotite software package. [53]
We further expanded our work by showing that Hb can be detected directly from disrupted red blood cells. Diluted sheep blood was disrupted using 0.02 % n‐dodecyl‐β‐D‐maltoside (DDM) and directly added to the chamber. After the addition of Hb, the current trace shows a combination of longer Hb events and short‐lived spikes from other blood proteins (Figure 4e). Hb constitutes most of the protein mass in red blood cells, hence Hb blockades ( ) were not obscured by the presence of other proteins. The pore showed more frequent Hb blockades with increasing amounts of diluted sheep blood in the chamber (Figure 4f). The fraction of time the pore was blocked by Hb ( , where and denote the time for which the pore is blocked by Hb and open, respectively) shows a sigmoidal response, with a quasi‐linear region between to (Figure 4f). Note that the concentration of the Hb in diluted sheep blood was quantified spectroscopically. Fitting of the Hill‐Langmuir equation to the data yields a half occupation concentration and a Hill coefficient of . Note that the concentration of Hb from disrupted blood ( ) falls within the quasi‐linear range of the binding curve, demonstrating that our nanopore system can quantify the Hb concentration directly from blood.
Conclusion
In conclusion, we employed PlyAB‐E1 biological nanopores to count and differentiate human Hb variants. Remarkably, despite the shape and volume of the Hb variants being almost identical, ionic currents allowed identifying differences in Hb proteins of one amino acid with more than 97 % accuracy.
Our continuum and BD models, based on simplified 2D‐axisymmetric geometries and the ePNP‐NS equations, can explain the nature of the exceptional ability of the nanopore to distinguish between proteins differing by as little as one charged residue (HbA and HbS). In PlyAB there are two stable equilibrium configurations of Hb, which correspond to the L1 and L2 levels observed in experiments. The model shows that the relative occupancy of the two levels is very sensitive to the net protein charge (i.e., the pH) and the applied bias, providing a physical explanation of the complex signal observed in PlyAB and the resulting ability to identify Hb variants with high accuracy. Hence, just as in isoelectric focusing techniques, nanopore currents can identify single charge differences in individual proteins if they are lodged inside the nanopore. As the model was built from physical first principles, it also suggests that multiple equilibria could be found for other proteins trapped in narrow pores.
Normal Hb counts are to ( to ), concentrations that are easily accessible to nanopore analysis (here we measure or ). Hence, nanopores integrated into low‐cost nanofluidic devices could be used for automated blood analysis in point‐of‐care devices. The exquisite detection and discrimination of Hb variants down to one amino acid would allow the counting and identification of Hb variants, with immediate application in the diagnosis of a variety of Hb dysfunctions such as those in sickle cell anaemia patients.
Conflict of interest
G.M. is a founder, director, and shareholder of Portal Biotech Limited, a company engaged in the development of nanopore technologies. This work was not supported by Portal Biotech Limited..
1.
Supporting information
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Acknowledgements
The authors thank the University of Groningen and the ERC consolidator grant (726151, DeE‐Nano) for funding G.H. and G.M.
G. Huang, A. Voorspoels, R. C. A. Versloot, N. J. van der Heide, E. Carlon, K. Willems, G. Maglia, Angew. Chem. Int. Ed. 2022, 61, e202206227; Angew. Chem. 2022, 134, e202206227.
A previous version of this manuscript has been deposited on a preprint server (https://doi.org/10.26434/chemrxiv‐2021‐7gb4h).
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
- 1. Yang X., Piety N. Z., Vignes S. M., Benton M. S., Kanter J., Shevkoplyas S. S., Clin. Chem. 2013, 59, 1506–1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Weiss G., Goodnough L. T., N. Engl. J. Med. 2005, 352, 1011–1023. [DOI] [PubMed] [Google Scholar]
- 3. Zulstra W. G., Buursma A., Comp. Biochem. Physiol. Part B 1987, 88, 251–255. [DOI] [PubMed] [Google Scholar]
- 4. Bhaskaram P., Balakrishna N., Radhakrishna K. V., Krishnaswamy K., Indian J. Pediatr. 2003, 70, 25–28. [DOI] [PubMed] [Google Scholar]
- 5. Weykamp C. W., Penders T. J., Siebelder C. W., Muskiet F. A., van der Slik W., Clin. Chem. 1993, 39, 138–142. [PubMed] [Google Scholar]
- 6. Louderback A. L., JAMA J. Am. Med. Assoc. 1967, 202, 718–719. [PubMed] [Google Scholar]
- 7. Schmidt R. M., Holland S., Clin. Chem. 1974, 20, 591–594. [PubMed] [Google Scholar]
- 8. Schnedl W. J., Lahousen T., Wallner S. J., Krause R., Lipp R. W., Clin. Biochem. 2005, 38, 88–91. [DOI] [PubMed] [Google Scholar]
- 9. Mais D. D., Gulbranson R. D., Keren D. F., Am. J. Clin. Pathol. 2009, 132, 34–38. [DOI] [PubMed] [Google Scholar]
- 10. Wang J., Biosens. Bioelectron. 2006, 21, 1887–1892. [DOI] [PubMed] [Google Scholar]
- 11. Yager P., Domingo G. J., Gerdes J., Annu. Rev. Biomed. Eng. 2008, 10, 107–144. [DOI] [PubMed] [Google Scholar]
- 12. Howorka S., Siwy Z., Chem. Soc. Rev. 2009, 38, 2360. [DOI] [PubMed] [Google Scholar]
- 13. Ying Y.-L., Long Y.-T., J. Am. Chem. Soc. 2019, 141, 15720–15729. [DOI] [PubMed] [Google Scholar]
- 14. Kasianowicz J. J., Brandin E., Branton D., Deamer D. W., Proc. Natl. Acad. Sci. USA 1996, 93, 13770–13773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Maglia G., Restrepo M. R., Mikhailova E., Bayley H., Proc. Natl. Acad. Sci. USA 2008, 105, 19720–19725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Derrington I. M., Butler T. Z., Collins M. D., Manrao E., Pavlenok M., Niederweis M., Gundlach J. H., Proc. Natl. Acad. Sci. USA 2010, 107, 16060–16065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Deamer D., Akeson M., Branton D., Nat. Biotechnol. 2016, 34, 518–524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Dekker C., Nat. Nanotechnol. 2007, 2, 209–215. [DOI] [PubMed] [Google Scholar]
- 19. Miles B. N., Ivanov A. P., Wilson K. A., Doğan F., Japrung D., Edel J. B., Chem. Soc. Rev. 2013, 42, 15–28. [DOI] [PubMed] [Google Scholar]
- 20. Yusko E. C., Bruhn B. R., Eggenberger O. M., Houghtaling J., Rollings R. C., Walsh N. C., Nandivada S., Pindrus M., Hall A. R., Sept D., Li J., Kalonia D. S., Mayer M., Nat. Nanotechnol. 2017, 12, 360–367. [DOI] [PubMed] [Google Scholar]
- 21. Houghtaling J., Ying C., Eggenberger O. M., Fennouri A., Nandivada S., Acharjee M., Li J., Hall A. R., Mayer M., ACS Nano 2019, 13, 5231–5242. [DOI] [PubMed] [Google Scholar]
- 22. Nir I., Huttner D., Meller A., Biophys. J. 2015, 108, 2340–2349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Hu R., Rodrigues J. V., Waduge P., Yamazaki H., Cressiot B., Chishti Y., Makowski L., Yu D., Shakhnovich E., Zhao Q., Wanunu M., ACS Nano 2018, 12, 4494–4502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Waduge P., Hu R., Bandarkar P., Yamazaki H., Cressiot B., Zhao Q., Whitford P. C., Wanunu M., ACS Nano 2017, 11, 5706–5716. [DOI] [PubMed] [Google Scholar]
- 25. Movileanu L., Howorka S., Braha O., Bayley H., Nat. Biotechnol. 2000, 18, 1091–1095. [DOI] [PubMed] [Google Scholar]
- 26. Rotem D., Jayasinghe L., Salichou M., Bayley H., J. Am. Chem. Soc. 2012, 134, 2781–2787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Wang S., Haque F., Rychahou P. G., Evers B. M., Guo P., ACS Nano 2013, 7, 9814–9822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Fahie M. A., Yang B., Mullis M., Holden M. A., Chen M., Anal. Chem. 2015, 87, 11143–11149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Fahie M., Chisholm C., Chen M., ACS Nano 2015, 9, 1089–1098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Thakur A. K., Movileanu L., Nat. Biotechnol. 2019, 37, 96–101. [Google Scholar]
- 31. Soskine M., Biesemans A., Moeyaert B., Cheley S., Bayley H., Maglia G., Nano Lett. 2012, 12, 4895–4900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Soskine M., Biesemans A., de Maeyer M., Maglia G., J. Am. Chem. Soc. 2013, 135, 13456–13463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Wloka C., Mutter N. L., Soskine M., Maglia G., Angew. Chem. Int. Ed. 2016, 55, 12494–12498; [DOI] [PubMed] [Google Scholar]; Angew. Chem. 2016, 128, 12682–12686. [Google Scholar]
- 34. Huang G., Willems K., Soskine M., Wloka C., Maglia G., Nat. Commun. 2017, 8, 935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Huang G., Voet A., Maglia G., Nat. Commun. 2019, 10, 835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Biesemans A., Soskine M., Maglia G., Nano Lett. 2015, 15, 6076–6081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Soskine M., Biesemans A., Maglia G., J. Am. Chem. Soc. 2015, 137, 5793–5797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. van Meervelt V., Soskine M., Singh S., Schuurman-Wolters G. K., Wijma H. J., Poolman B., Maglia G., J. Am. Chem. Soc. 2017, 139, 18640–18646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Galenkamp N. S., Soskine M., Hermans J., Wloka C., Maglia G., Nat. Commun. 2018, 9, 4085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Willems K., Ruić D., Biesemans A., Galenkamp N. S., van Dorpe P., Maglia G., ACS Nano 2019, 13, 9980–9992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Zernia S., van der Heide N. J., Galenkamp N. S., Gouridis G., Maglia G., ACS Nano 2020, 14, 2296–2307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Lucas F. L. R., Sarthak K., Lenting E. M., Coltan D., van der Heide N. J., Versloot R. C. A., Aksimentiev A., Maglia G., ACS Nano 2021, 15, 9600–9613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Wloka C., van Meervelt V., van Gelder D., Danda N., Jager N., Williams C. P., Maglia G., ACS Nano 2017, 11, 4387–4394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Liu Y., Wang K., Wang Y., Wang L., Yan S., Du X., Zhang P., Chen H.-Y., Huang S., J. Am. Chem. Soc. 2022, 144, 757–768. [DOI] [PubMed] [Google Scholar]
- 45. Lukoyanova N., Kondos S. C., Farabella I., Law R. H. P., Reboul C. F., Caradoc-Davies T. T., Spicer B. A., Kleifeld O., Traore D. A. K., Ekkel S. M., Voskoboinik I., Trapani J. A., Hatfaludi T., Oliver K., Hotze E. M., Tweten R. K., Whisstock J. C., Topf M., Saibil H. R., Dunstone M. A., PLoS Biol. 2015, 13, e1002049-. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Huang G., Willems K., Bartelds M., van Dorpe P., Soskine M., Maglia G., Nano Lett. 2020, 20, 3819–3827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Park S.-Y., Yokoyama T., Shibayama N., Shiro Y., Tame J. R. H., J. Mol. Biol. 2006, 360, 690–701. [DOI] [PubMed] [Google Scholar]
- 48. Oksenberg D., Dufu K., Patel M. P., Chuang C., Li Z., Xu Q., Silva-Garcia A., Zhou C., Hutchaleelaha A., Patskovska L., Patskovsky Y., Almo S. C., Sinha U., Metcalf B. W., Archer D. R., Br. J. Haematol. 2016, 175, 141–153. [DOI] [PubMed] [Google Scholar]
- 49. Humphrey W., Dalke A., Schulten K., J. Mol. Graphics 1996, 14, 33–38. [DOI] [PubMed] [Google Scholar]
- 50.J. Stone, An Efficient Library for Parallel Ray Tracing and Animation, MSc thesis, University of Missouri-Rolla, 1998.
- 51. Willems K., Ruić D., Lucas F. L. R., Barman U., Verellen N., Hofkens J., Maglia G., van Dorpe P., Nanoscale 2020, 12, 16775–16795. [DOI] [PubMed] [Google Scholar]
- 52.J. Soman, J. Olson, “Structure of Wild-type Fetal Human Hemoglobin HbF” 2013, https://doi.org/10.2210/pdb4mqj/pdb .
- 53. Kunzmann P., Hamacher K., BMC Bioinf. 2018, 19, 346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Akinsheye I., Alsultan A., Solovieff N., Ngo D., Baldwin C. T., Sebastiani P., Chui D. H. K., Steinberg M. H., Blood 2011, 118, 19–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.