ABSTRACT
Antimicrobial resistance is a major public health threat, and there is an urgent need for new strategies to address this issue. In a recent study, a library screening strategy was developed in which an FDA-approved drug library was screened against methicillin-resistant Staphylococcus aureus (MRSA) in both its original (unmetabolized [UM]) and its human liver microsome metabolized (postmetabolized [PM]) forms and in the absence and presence of a resistant-to antibiotic. This allows the identification of agents with active metabolites and agents that can act synergistically with the resistant-to antibiotic. In this study, this strategy is applied to VanA-type vancomycin-resistant Enterococcus faecium (VREfm) in the absence and presence of vancomycin. Thirteen drugs with minimum MICs that were ≤12.5 μM under any tested condition (UM/PM vs. −/+vancomycin) were identified. Seven of these appeared to act synergistically with vancomycin, and follow-up checkerboard analyses confirmed synergy (∑FICmin ≤0.5) for six of these. Ultimately four rifamycins, two pleuromutilins, mupirocin, and linezolid were confirmed as synergistic. The most synergistic agent was rifabutin (∑FICmin = 0.19). Linezolid, a protein biosynthesis inhibitor, demonstrated relatively weak synergy (∑FICmin = 0.5). Only mupirocin showed significantly improved activity after microsomal metabolism, indicative of a more active metabolite, but efforts to identify an active metabolite were unsuccessful. Spectra of activity of several hits and related agents were also determined. Gemcitabine showed activity against a number vancomycin-resistant E. faecium and E. faecalis strains, but this activity was substantially weaker than previously observed in MRSA.
IMPORTANCE Resistance to currently used antibiotics poses a serious threat to public health. This study reports a complete screen of 1,000 FDA-approved drugs and their metabolites against vancomycin-resistant Enterococcus faecium (VREfm) in both the absence and presence of vancomycin. This identified potentially synergistic combinations of FDA-approved drugs with vancomycin, and a number of these were confirmed in follow-up checkerboard assays. Among intrinsically active FDA-approved drugs, gemcitabine was identified as having activity against a panel of VRE strains.
KEYWORDS: library screening, drug repurposing, synergy screening, Enterococcus faecium, microsome, metabolism, antibiotic drug resistance, VRE, antimicrobial resistance, cytochrome P450, drug metabolism, vancomycin resistance
INTRODUCTION
Pathogenic bacteria are becoming increasingly drug resistant, with some now virtually untreatable (1–4). There has concurrently been a lack of new antibacterial agents identified over the last 30 years to counter this threat (4–6). Enterococcus spp. are Gram-positive commensal bacteria of the intestine in humans and animals that can cause problematic infections of the gasterointestinal (GI) tract and soft tissues (7–10). Vancomycin (Vm) is one of the most important agents for the treatment of G+ bacterial infections resistant to most other antibacterial agents, including Vm-sensitive enterococcal (VSE) and methicillin-resistant Staphylococcus aureus (MRSA) infections (11–13). The emergence and spread of Vm-resistant Enterococcus spp. (VRE) are serious public health issues given the lack of alternatives for these organisms and their increasing resistance to currently used agents (8, 9, 11, 14). VRE is a WHO high priority pathogen for new agent development (15).
Chemical compound library screening is a core approach for the discovery of new bioactive agents, including for antibacterial activity. However, large untargeted (whole cell) and targeted (individual protein) library screening efforts have given overall disappointing results (16, 17). An alternative to large library screens are smaller scale efforts with high value libraries such as FDA-approved drug library screening, which has become a popular strategy for “drug repurposing” (18). This strategy can reveal novel new activities of FDA-approved drugs, which provides a greatly shortened path to clinical application. Another strategy to counter antimicrobial resistance is to identify agents that can act synergistically with or restore the activity of another antibiotic (5, 19, 20). Many drugs are also known to have active metabolites (21, 22). Comparative screening of the unmetabolized (UM) and postmetabolized (PM) libraries allows agents with increased antibacterial activity to be identified for deconvolution and active metabolite identification, as recently demonstrated using a human liver microsome-metabolized FDA-approved drug library (23).
In a prior study we have demonstrated an approach in which FDA-approved drug library screening is performed against MRSA with both FDA-approved drugs and their metabolites and simultaneously in the absence and presence of a resistant-to antibiotic (23). This allows intrinsically active agents, agents with active metabolites, and agents synergistic with the resistant-to antibiotic to be identified in a single screen. In this study, this approach was used to screen an FDA-approved library in its original (unmetabolized [UM]) and human microsome metabolized (postmetabolized [PM]) versions against vancomycin-resistant Enterococcus faecium (VREfm) in the absence and presence of sub-MIC levels of vancomycin (−/+Vm) (2 × 2 library screening design).
RESULTS AND DISCUSSION
While there has been some progress in the development of new antibacterial agents to combat the emergence and spread of antimicrobial resistance (AMR) in pathogenic bacteria (24), there has been a lack of new agents without established resistance mechanisms (5, 25–28). New agents of novel mechanism are urgently needed. Another approach to combating AMR is the identification of effective synergistic agent combinations, particularly from the repertoire of existing antibacterial agents (20, 29–31). Such synergistic agent combinations have enhanced activity against targeted organisms and can also reduce the emergence and spread of resistance. Both new agent identification and the identification of new synergistic agent combinations are potential pathways in addressing the problem of AMR. The development and demonstration of effective approaches that can perform both types of screens simultaneously would be useful for efforts to address AMR.
In a prior study, we demonstrated in MRSA how a two-dimensional screening strategy comparing an unmetabolized (UM) versus postmetabolized (PM) FDA library screen combined with a −/+ resistant-to antibiotic screen could enhance the ability to identify new agents and new synergistic combinations (23, 32). This study identified gemcitabine as having generally good anti-MRSA activity, identified strong synergy between cefoxitin and floxuridine against MRSA, and also identified capecitabine as having several anti-MRSA metabolites. Focusing this approach on an FDA-approved drug library offers the potential of identifying novel antibacterial activities in FDA-approved drugs and FDA-approved drug metabolites, and identifying novel synergistic drug combinations, all features demonstrated previously (23). The rationale for this strategy is to perform screening replication under somewhat different but informative conditions. Screening of the PM library allows FDA-approved drugs with active metabolites to be identified, and screening in the absence and presence of a resistant-to antibiotic allows agents acting synergistically with the resistant-to antibiotic to be identified.
In this study, this effort was extended to a vancomycin-resistant Enterococcus faecium strain in the absence and presence of vancomycin. Library screens were performed at 50 μM of the library compounds and in the absence or presence of 16 μg/mL vancomycin (1/8× MIC). This vancomycin level was selected as a compromise between a higher 1/4× MIC concentration of 32 μg/mL and a lower more clinically relevant concentration that would be unlikely to reveal synergies with vancomycin. Following library screening a merged hit list was made, in which any compound that gave a hit (was active) under any of the four screening conditions (UM − Vm, UM + Vm, PM − Vm, and PM + Vm) was included in the list. MICs for all the compounds in this pooled hit list were then determined for UM library compounds in both the absence and presence of 16 μg/mL vancomycin and for PM library compounds in both the absence and presence of 16 μg/mL vancomycin to give a preliminary table of MICs. MICs for compounds which gave a minimum MIC over all four tested conditions of ≤25 μM were then determined at least in triplicate. The results for agents with an MIC_min ≤12.5 (under all 4 test conditions) are summarized in Table 1. A complete list of all active and inactive agents is provided as Tables S2 and S3, respectively.
TABLE 1.
FDA library anti-VREfm (clinical isolate) hit MICs (Min_MIC ≤12.5 μM)
| UM MICs (μM) |
PM MICs (μM) |
||||||||
|---|---|---|---|---|---|---|---|---|---|
| Compound | −Vm | +Vm | −Vm | +Vm | Min_MIC | L2(UM/PM)a | AL2(UM/PM)b | L2(UM−/+Vm)c | AL2(−/+Vm)d |
| Rifampin | 0.10 | 2.4 × 10−2 | 12.5 | 12.5 | 2.4 × 10−2 | −7 | −8 | 2 e | 1 |
| Rifapentine | 0.20 | 2.4 × 10−2 | 25 | 12.5 | 2.4 × 10−2 | −7 | −8 | 3 | 2 e |
| Retapamulin | 0.20 | 4.9 × 10−2 | 3.1 | 1.6 | 4.9 × 10−2 | −4 | −4.5 | 2 | 1.5 |
| Rifaximin | 0.39 | 4.9 × 10−2 | 50 | 50 | 4.9 × 10−2 | −7 | −8.5 | 3 | 1.5 |
| Rifabutin | 0.20 | 9.8 × 10−2 | 25 | 6.25 | 9.8 × 10−2 | −7 | −6.5 | 1 | 1.5 |
| Valnemulin | 0.39 | 9.8 × 10−2 | 0.20 | 9.8 × 10−2 | 9.8 × 10−2 | 1 | 0.5 | 2 | 1.5 |
| Gemcitabine | 0.78 | 0.20 | 25 | 25 | 0.20 | −5 | −6 | 2 | 1 |
| Mupirocin | 3.1 | 0.78 | 0.78 | 0.39 | 0.39 | 2 e | 1.5 | 2 | 1.5 |
| Closantel | 1.6 | 1.6 | 12.5 | 12.5 | 1.6 | −3 | −3 | 0 | 0 |
| Novobiocin | 3.1 | 1.6 | 3.1 | 3.1 | 1.6 | 0 | −0.5 | 1 | 0.5 |
| Fidaxomicin | 12.5 | 6.25 | 100 | 100 | 6.25 | −3 | −3.5 | 1 | 0.5 |
| Florfenicol | 25 | 25 | 12.5 | 6.25 | 6.25 | 1 | 1.5 | 0 | 0.5 |
| Linezolid | 3.1 | 1.6 | 6.25 | 6.25 | 6.25 | −1 | −1.5 | 1 | 0.5 |
Values ≥2 are indicative of a significant decrease in MIC and a significant increase in potency and are in bold. For UM/PM ratios, this is indicative of a possible active metabolite, and for −/+Vm ratios this is indicative of a possible synergistic interaction between the drug and vancomycin.
Several assessments and comparisons are possible using the Table 1 MIC data (23). The first is to assess for interesting intrinsically active agents as revealed by examining the UM − Vm data in Table 1. Most of the agents listed in the UM − Vm column are well-known antibacterial agents with the exception of gemcitabine and closantel. Closantel is a veterinary antiparasitic drug that has previously been identified as having anti-MRSA and anti-VRE activity (33–36). Gemcitabine has also previously been identified as having anti-MRSA activity (37, 38), including in our own efforts (32), but its anti-VRE activity appears previously unrecognized. The spectrum of activity of several of the better agents from Table 1 was determined against a panel of VRE isolates (both VREfm and E. faecalis [VREfa]), in which gemcitabine demonstrated activity against all tested VRE strains with a median MIC of 0.78 μM (Table 2). This is a higher median MIC than against a panel of MRSA strains where the median MIC was 0.049 μM. These results indicate that gemcitabine and similar agents may have some potential for further development as anti-VRE and anti-MRSA agents. Several other nucleoside analogs were also tested for activity against these VRE strains (Table 3), but these did not exhibit the same broad anti-VRE activity as they did against MRSA (23).
TABLE 2.
Spectra of activity (UM − Vm) against VREa
| Compound | VREfm (clinical)b,c | VREfm (BAA-2317) | VREfm (BAA-2318) | VREfm (BAA-2365) | VREfa (49532) | VREfa (49533) | VREfa (51575) | VREfa (700802) |
|---|---|---|---|---|---|---|---|---|
| Rifampin | 0.10 | NAd | 0.39 | 1.6 | 6.25 | 6.25 | 6.25 | 6.25 |
| Retapamulin | 0.20 | 9.8 × 10−2 | 9.8 × 10−2 | NA | NA | NA | NA | NA |
| Rifabutin | 0.20 | NA | 0.78 | 3.1 | 25 | 25 | 25 | 25 |
| Rifapentine | 0.20 | NA | 1.6 | 3.1 | 6.25 | 6.25 | 6.25 | 12.5 |
| Rifaximin | 0.39 | 50 | 25 | 1.6 | 1.6 | 1.6 | 3.1 | 1.6 |
| Valnemulin | 0.39 | 0.39 | 0.39 | NA | NA | NA | NA | NA |
| Gemcitabine | 0.78 | 0.39 | 2.4 × 10−2 | 0.78 | 1.6 | 1.6 | 0.78 | 3.1 |
MICs are in μM.
Values from Table 1.
The VREfm used in this study was a clinical isolate from the University of Missouri-Kansas City School of Medicine. For other strains, the American Type Culture Collection (ATCC) catalog numbers are given in parentheses.
NA, not active at 50 μM, the highest concentration used in these MIC determinations.
TABLE 3.
Spectra of activity of doxifluridine, floxuridine, and 5′-fluorouracil against VREa
| Compound | VREfm (clinical) | VREfm (BAA-2317)b | VREfm (BAA-2318) | VREfa (BAA-2365) | VREfa (49532) | VREfa (49533) | VREfa (51575) | VREfa (700802) |
|---|---|---|---|---|---|---|---|---|
| DFUR | NAc | NA | NA | NA | NA | NA | 50 | NA |
| Floxuridine | NA | NA | NA | 3.1 | 25 | 50 | 6.25 | 50 |
| 5-Fluorouracil | NA | NA | NA | 3.1 | 25 | 25 | 12.5 | 50 |
MICs are in μM. DFUR, doxifluridine.
American Type Culture Collection (ATCC) catalog numbers are in parentheses.
NA, not active at 50 μM, the highest concentration used in these MIC determinations.
The second assessment is to compare Table 1 MICs to identify agents with increased activity after metabolism, indicative of more active metabolites. Nearly all drugs are transformed into at least one metabolite, and such metabolites frequently have distinct biological activities (21, 22). Our prior study demonstrated the potential of a UM versus PM library screen to identify active drug metabolites (23). The effect of microsomal metabolism on compound activity against VREfm is highlighted in Table 1 in the L2(UM/PM−Vm) value. L2(UM/PM−Vm) is the log base 2 value of the ratio between the MICUM−Vm and MICPM−Vm values, as defined in Table 1. The AL2(UM/PM) represents the average of log 2 value of MICUM−Vm/MICPM−Vm and log 2 value of MICUM+Vm/MICPM+Vm as defined in Table 1 (23). These values place the fold change in MIC values between various treatments on a log scale. Values of L2(UM/PM−Vm) ≥2 (i.e., 4-fold reduction in MIC) are in bold and indicate substantially increased potency (lower MIC) after metabolism. Only mupirocin met this standard after metabolism (Table 1), suggesting the possibility of an active metabolite. However, we were unable to identify an active metabolite. While library metabolism was useful in identifying novel anti-MRSA compounds (23), it was not successful when applied to VREfm.
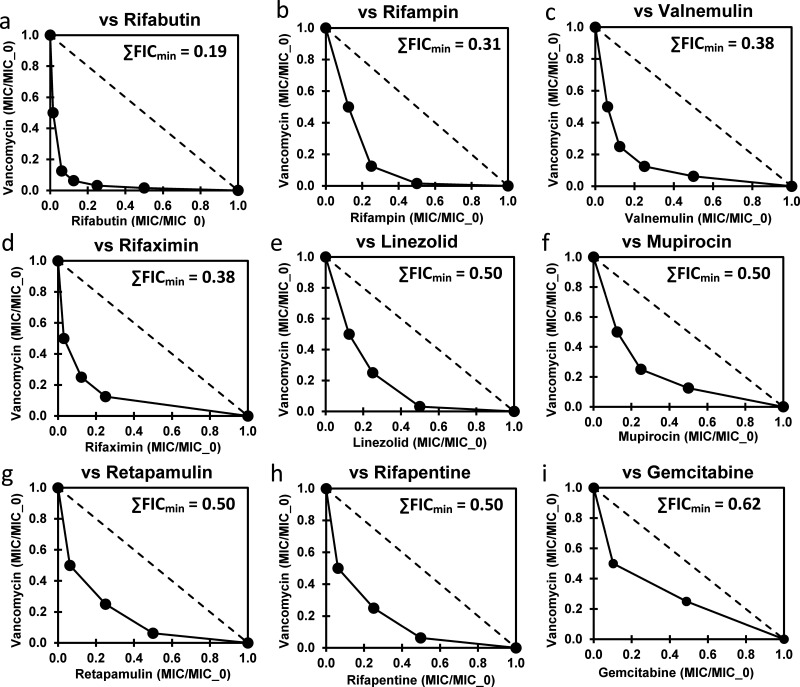
The third assessment from Table 1 data is comparisons to reveal possible synergistic agent combinations with vancomycin (Table 1; L2(UM−/+Vm) and AL2(−/+Vm) columns). This identified seven potential agents with vancomycin synergy: rifampin, rifapentine, retapamulin, rifaximin, valnemulin, gemcitabine, and mupirocin. Checkerboard assays revealed significant synergies (∑FICmin ≤0.5) for six of these, but not for gemcitabine (Fig. 1). Since several of these were rifamycins, synergy for rifabutin was also tested for and confirmed (Fig. 1a). The observation of synergy of these rifamycins, RNA biosynthesis inhibitors, with vancomycin is interesting. Rifampicin synergy with vancomycin has been previously observed with several VRE strains and with no synergy for vancomycin-sensitive enterococcal strains (39). Both pleuromutilin (retapamulin and valnemulin) protein biosynthesis inhibitors also demonstrated synergy with vancomycin (Fig. 1g and c, respectively). A checkerboard assay was then performed between linezolid and vancomycin, and this was also confirmed as a synergistic combination (Fig. 1e). Mupirocin, an Ile-tRNA biosynthesis inhibitor that induces (p)ppGpp biosynthesis (stringent response) (40, 41), also demonstrated modest synergy (Fig. 1f). These observations demonstrate that direct RNA biosynthesis inhibitors (rifamycins), indirect RNA biosynthesis inhibitors (mupirocin), and protein biosynthesis inhibitors (pleuromutilins and linezolid) can all act synergistically with vancomycin on VREfm. Vancomycin resistance in this strain of VREfm is inducible (42), and the ability of these agents to block RNA and protein biosynthesis likely blocks the ability of this VREfm strain to express high level vancomycin resistance. Rifabutin was the most synergistic of these agents for reasons which are currently unknown. The report of rifampin-vancomycin synergy in VRE strains, but not in VSE strains (39), also supports this conclusion.
FIG 1.
(a to i) Isobolograms from checkerboard assay results for combinations of vancomycin with potentially synergistic agents against VREfm (clinical isolate). The dashed line in the isobolograms is for the no interaction (additive MICs) curve. MICs for other agents alone as given in Table 1.
The overall goal of this effort was to further demonstrate the utility of enhanced library screening approaches in which replicate library screens are performed with variation between the replicates. The basic screen (UM − Vm) identified both closantel and gemcitabine as nontypical anti-VRE agents. The spectrum of activity of closantel against a number of VRE strains has previously been reported (36). Gemcitabine was demonstrated in this study (Table 2) to also have activity against a number of VRE strains. The molecular target of gemcitabine is unknown, but further studies of gemcitabine and homologs seem justified from these observations. No agents with identifiable active metabolites were discovered in this screen, in contrast to the identification of active capecitabine metabolites in MRSA (23). Screening for synergistic combinations with vancomycin revealed a number of synergistic agents (Fig. 1). These were all either RNA or protein biosynthesis inhibitors, suggesting a common mechanistic basis for these synergies. Some of the agents and agent combinations identified in this effort may be suitable candidates for further in vitro and in vivo studies, and ultimately clinical application.
MATERIALS AND METHODS
General.
The reagents and materials used in this study were as described previously (23). Bacterial strains were obtained from American Type Culture Collection (ATCC; Manasas, VA) and the Biodefense and Emerging Infections Research Resources Repository (BEI; Manasas, VA). The bacterial strain used for library screening was a VanA-type vancomycin-resistant Enterococcus faecium (VREfm) clinical strain we have used in other studies (42). Other bacterial strains used in this study were as indicated in the appropriate tables. UM and PM working library plates including control antibiotics and control microsomal metabolism substrates were prepared as described previously (23) at 0.5 mM. VRE growth medium consisted of brain heart infusion (37.5 g/L), hemin (10 mg/L), and NAD+ (10 mg/L).
UM/PM versus −/+Vm library screen against VREfm.
Four sets of library screening plates were prepared for the following screens; UM − Vm, UM + Vm, PM − Vm, and PM + Vm. Two sets of UM plates and two sets of PM plates were first prepared from working library samples (2 μL at 0.5 mM working library samples per well) in 384-well Corning microtiter plates (catalog no. 3680) using a Biomek 3000 liquid handing workstation. Plates were frozen at −80°C and dried under strong vacuum (<50 μm Hg) in a Genevac Quatro centrifugal concentrator. To each well in each set was added 20 μL of VRE growth medium containing 4,000 CFU VRE and containing either no Vm for −Vm screens or +16 μg mL−1 Vm (1/8× MIC) for +Vm screens. These additions were performed using an Integra Viaflo Assist automated multichannel pipette (Hudson, NH) in a Labconco (Kansas City, MO) BSL-2 biosafety cabinet. Plates were incubated for 48 h at 35°C. Fresh VRE growth medium (10 μL) was added to the wells of these four sets of plates, followed by incubation for 2 h at 35°C to restart active cell growth. To the wells of these plates was then added 10 μL of 100 μg mL−1 resazurin (sodium salt) (43–45). The plates were incubated for another 2 h at 35°C, and the A610 – A450 absorbance difference (Promega Technical Bulletin TB317) was measured in a Molecular Devices SpectraMax M5 multimode microplate reader (San Jose, CA).
Hit picking and MIC determination.
Library screening data were processed and analyzed using homemade Matlab scripts (The Mathworks, Natick, MA). Based on the values for known active and inactive antibacterial agent controls, a cut-off value between active and inactive compounds was selected and lists of active wells in each screening set (UM − Vm, UM + Vm, PM − Vm, and PM + Vm) were generated. These lists were merged to give a pooled hit list. Rows were added to this pooled hit list to include known active and inactive antibiotics containing wells as controls. MICs were determined by hit picking 2-μL samples from both UM and PM working plates (two sets from each) into the first columns of 384-well plates (four sets total, for UM − Vm, UM + Vm, PM − Vm, and PM + Vm MIC determinations). These samples were then serially diluted in steps of two across the plates with DMSO using an Integra Viaflo Assist automated multichannel pipettor. The last column was left blank (DMSO only). These plates were frozen at −80°C and dried under strong vacuum as described above. To each well in each set was added 20 μL VRE growth medium containing 4000 CFU VREfm and containing either no Vm for −Vm MICs or 16 μg mL−1 Vm for +Vm MICs. (This provided MIC plates with 50 μM as the highest test agent concentration.) Plates were incubated for 48 h at 35°C. Fresh VRE growth medium (10 μL) was added to the wells of these four sets of plates, followed by incubation for 2 h at 35°C to restart active cell growth. To the wells of these plates was then added 10 μL of 100 μg mL−1 resazurin (43–45). The plates were incubated for another 2 h at 35°C, and the A610 – A450 absorbance difference was measured as described above. MICs were determined using a cutoff midway between known active and inactive samples. All MICs were determined at least in triplicate and at least in quadruplicate for MIC_min ≤ 25 μM to ensure reproducibility.
Spectrum-of-activity of VRE hits.
MICs were determined against a panel of VREfm and VREfa strains (Table 2). Plates were prepared by serial dilution of compounds in DMSO across 384-well plates and drying under high vacuum as described above.
Checkerboard assays to confirm synergy.
Checkerboard assays (29) were performed to confirm synergy for prospective synergistic agents. Checkerboard assays were performed in 96-well plates from DMSO compound stocks using serial dilutions in steps of two in both dimensions in DMSO, and plates were then dried under vacuum as described above. To these plates was added 100 μL VRE growth medium containing 4,000 CFU VRE to each well, and plates were incubated for 48 h at 35°C. Fresh VRE growth medium (50 μL) was added, plates were incubated at 35°C for 2 h, 50 μL of 100 μg mL−1 resazurin was added, plates were incubated an additional 2 h, and the resazurin absorbance difference was measured as described above. All checkerboard assays were performed at least in triplicate and averaged.
ACKNOWLEDGMENTS
The authors acknowledge support from the National Institutes of Health (R15-GM126502) to W.G.G.
Footnotes
Supplemental material is available online only.
Contributor Information
William G. Gutheil, Email: gutheilw@umkc.edu.
Amit Singh, Indian Institute of Science Bangalore.
REFERENCES
- 1.World Health Organization. 2014. Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva, Switzerland. [Google Scholar]
- 2.Ventola CL. 2015. The antibiotic resistance crisis: part 1: causes and threats. P T 40:277–283. [PMC free article] [PubMed] [Google Scholar]
- 3.CDC. 2019. Antibiotic resistance threats in the United States, 2019. CDC, Atlanta, GA. [Google Scholar]
- 4.De Oliveira DMP, Forde BM, Kidd TJ, Harris PNA, Schembri MA, Beatson SA, Paterson DL, Walker MJ. 2020. Antimicrobial resistance in ESKAPE pathogens. Clin Microbiol Rev 33:e00181-19. doi: 10.1128/CMR.00181-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brown ED, Wright GD. 2016. Antibacterial drug discovery in the resistance era. Nature 529:336–343. doi: 10.1038/nature17042. [DOI] [PubMed] [Google Scholar]
- 6.Butler MS, Blaskovich MA, Cooper MA. 2017. Antibiotics in the clinical pipeline at the end of 2015. J Antibiot (Tokyo) 70:3–24. doi: 10.1038/ja.2016.72. [DOI] [PubMed] [Google Scholar]
- 7.Teixeira LM, Carvalho MDS, Facklam RR. 2007. Enterococcus, p 430–442. In Manual of clinical microbiology, 9th ed. American Society for Microbiology, Washington, DC. [Google Scholar]
- 8.Arias CA, Murray BE. 2012. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol 10:266–278. doi: 10.1038/nrmicro2761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Barber KE, King ST, Stover KR, Pogue JM. 2015. Therapeutic options for vancomycin-resistant enterococcal bacteremia. Expert Rev Anti Infect Ther 13:363–377. doi: 10.1586/14787210.2015.1001839. [DOI] [PubMed] [Google Scholar]
- 10.Mancuso G, Midiri A, Gerace E, Biondo C. 2021. Bacterial antibiotic resistance: the most critical pathogens. Pathogens 10:1310. doi: 10.3390/pathogens10101310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wang JL, Hsueh PR. 2009. Therapeutic options for infections due to vancomycin-resistant enterococci. Expert Opin Pharmacother 10:785–796. doi: 10.1517/14656560902811811. [DOI] [PubMed] [Google Scholar]
- 12.Koomanachai P, Crandon JL, Nicolau DP. 2009. Newer developments in the treatment of Gram-positive infections. Expert Opin Pharmacother 10:2829–2843. doi: 10.1517/14656560903357491. [DOI] [PubMed] [Google Scholar]
- 13.Rivera AM, Boucher HW. 2011. Current concepts in antimicrobial therapy against select gram-positive organisms: methicillin-resistant Staphylococcus aureus, penicillin-resistant pneumococci, and vancomycin-resistant enterococci. Mayo Clin Proc 86:1230–1243. doi: 10.4065/mcp.2011.0514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Miller WR, Murray BE, Rice LB, Arias CA. 2020. Resistance in vancomycin-resistant Enterococci. Infect Dis Clin North Am 34:751–771. doi: 10.1016/j.idc.2020.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tacconelli E, Carrara E, Savoldi A, Harbarth S, Mendelson M, Monnet DL, Pulcini C, Kahlmeter G, Kluytmans J, Carmeli Y, Ouellette M, Outterson K, Patel J, Cavaleri M, Cox EM, Houchens CR, Grayson ML, Hansen P, Singh N, Theuretzbacher U, Magrini N, WHO Pathogens Priority List Working Group . 2018. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis 18:318–327. doi: 10.1016/S1473-3099(17)30753-3. [DOI] [PubMed] [Google Scholar]
- 16.Payne DJ, Gwynn MN, Holmes DJ, Pompliano DL. 2007. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nat Rev Drug Discov 6:29–40. doi: 10.1038/nrd2201. [DOI] [PubMed] [Google Scholar]
- 17.Tommasi R, Brown DG, Walkup GK, Manchester JI, Miller AA. 2015. ESKAPEing the labyrinth of antibacterial discovery. Nat Rev Drug Discov 14:529–542. doi: 10.1038/nrd4572. [DOI] [PubMed] [Google Scholar]
- 18.Miro-Canturri A, Ayerbe-Algaba R, Smani Y. 2019. Drug repurposing for the treatment of bacterial and fungal infections. Front Microbiol 10:41. doi: 10.3389/fmicb.2019.00041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ejim L, Farha MA, Falconer SB, Wildenhain J, Coombes BK, Tyers M, Brown ED, Wright GD. 2011. Combinations of antibiotics and nonantibiotic drugs enhance antimicrobial efficacy. Nat Chem Biol 7:348–350. doi: 10.1038/nchembio.559. [DOI] [PubMed] [Google Scholar]
- 20.Coates ARM, Hu Y, Holt J, Yeh P. 2020. Antibiotic combination therapy against resistant bacterial infections: synergy, rejuvenation and resistance reduction. Expert Rev Anti Infect Ther 18:5–15. doi: 10.1080/14787210.2020.1705155. [DOI] [PubMed] [Google Scholar]
- 21.Kang MJ, Song WH, Shim BH, Oh SY, Lee HY, Chung EY, Sohn Y, Lee J. 2010. Pharmacologically active metabolites of currently marketed drugs: potential resources for new drug discovery and development. Yakugaku Zasshi 130:1325–1337. doi: 10.1248/yakushi.130.1325. [DOI] [PubMed] [Google Scholar]
- 22.Obach RS. 2013. Pharmacologically active drug metabolites: impact on drug discovery and pharmacotherapy. Pharmacol Rev 65:578–640. doi: 10.1124/pr.111.005439. [DOI] [PubMed] [Google Scholar]
- 23.Ayon NJ, Gutheil WG. 2019. Dimensionally enhanced antibacterial library screening. ACS Chem Biol 14:2887–2894. doi: 10.1021/acschembio.9b00745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Butler MS, Gigante V, Sati H, Paulin S, Al-Sulaiman L, Rex JH, Fernandes P, Arias CA, Paul M, Thwaites GE, Czaplewski L, Alm RA, Lienhardt C, Spigelman M, Silver LL, Ohmagari N, Kozlov R, Harbarth S, Beyer P. 2022. Analysis of the clinical pipeline of treatments for drug-resistant bacterial infections: despite progress, more action is needed. Antimicrob Agents Chemother 66:e0199121. doi: 10.1128/AAC.01991-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Campanini-Salinas J, Andrades-Lagos J, Mella-Raipan J, Vasquez-Velasquez D. 2018. Novel classes of antibacterial drugs in clinical development, a hope in a post-antibiotic era. Curr Top Med Chem 18:1188–1202. doi: 10.2174/1568026618666180816162846. [DOI] [PubMed] [Google Scholar]
- 26.Theuretzbacher U, Bush K, Harbarth S, Paul M, Rex JH, Tacconelli E, Thwaites GE. 2020. Critical analysis of antibacterial agents in clinical development. Nat Rev Microbiol 18:286–298. doi: 10.1038/s41579-020-0340-0. [DOI] [PubMed] [Google Scholar]
- 27.Paulin S, Alm RA, Beyer P. 2020. A novel pre-clinical antibacterial pipeline database. PLoS One 15:e0236604. doi: 10.1371/journal.pone.0236604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Burki TK. 2021. Development of new antibacterial agents: a sense of urgency needed. Lancet Respir Med 9:e54. doi: 10.1016/S2213-2600(21)00230-7. [DOI] [PubMed] [Google Scholar]
- 29.Pillai SK, Moellering RCJ, Eliopoulos GM. 2005. Chapter 9: Antimicrobial combinations, p 365–440. In Lorian V (ed), Antibiotics in laboratory medicine. Wolters Kluwer Health, Philadelphia, PA. [Google Scholar]
- 30.Roemer T, Boone C. 2013. Systems-level antimicrobial drug and drug synergy discovery. Nat Chem Biol 9:222–231. doi: 10.1038/nchembio.1205. [DOI] [PubMed] [Google Scholar]
- 31.Tyers M, Wright GD. 2019. Drug combinations: a strategy to extend the life of antibiotics in the 21st century. Nat Rev Microbiol 17:141–155. doi: 10.1038/s41579-018-0141-x. [DOI] [PubMed] [Google Scholar]
- 32.Ayon NJ, Gutheil WG. 2021. Correction to “Dimensionally Enhanced Antibacterial Library Screening”. ACS Chem Biol 16:1610–1611. doi: 10.1021/acschembio.1c00435. [DOI] [PubMed] [Google Scholar]
- 33.Macielag MJ, Demers JP, Fraga-Spano SA, Hlasta DJ, Johnson SG, Kanojia RM, Russell RK, Sui Z, Weidner-Wells MA, Werblood H, Foleno BD, Goldschmidt RM, Loeloff MJ, Webb GC, Barrett JF. 1998. Substituted salicylanilides as inhibitors of two-component regulatory systems in bacteria. J Med Chem 41:2939–2945. doi: 10.1021/jm9803572. [DOI] [PubMed] [Google Scholar]
- 34.Rajamuthiah R, Fuchs BB, Jayamani E, Kim Y, Larkins-Ford J, Conery A, Ausubel FM, Mylonakis E. 2014. Whole animal automated platform for drug discovery against multi-drug resistant Staphylococcus aureus. PLoS One 9:e89189. doi: 10.1371/journal.pone.0089189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Niu H, Yee R, Cui P, Tian L, Zhang S, Shi W, Sullivan D, Zhu B, Zhang W, Zhang Y. 2017. Identification of agents active against methicillin-resistant Staphylococcus aureus USA300 from a clinical compound library. Pathogens 6:44. doi: 10.3390/pathogens6030044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Mohammad H, AbdelKhalek A, Abutaleb NS, Seleem MN. 2018. Repurposing niclosamide for intestinal decolonization of vancomycin-resistant enterococci. Int J Antimicrob Agents 51:897–904. doi: 10.1016/j.ijantimicag.2018.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jordheim LP, Ben Larbi S, Fendrich O, Ducrot C, Bergeron E, Dumontet C, Freney J, Doleans-Jordheim A. 2012. Gemcitabine is active against clinical multiresistant Staphylococcus aureus strains and is synergistic with gentamicin. Int J Antimicrob Agents 39:444–447. doi: 10.1016/j.ijantimicag.2012.01.019. [DOI] [PubMed] [Google Scholar]
- 38.Sandrini MP, Shannon O, Clausen AR, Bjorck L, Piskur J. 2007. Deoxyribonucleoside kinases activate nucleoside antibiotics in severely pathogenic bacteria. Antimicrob Agents Chemother 51:2726–2732. doi: 10.1128/AAC.00081-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Skinner K, Sandoe JAT, Rajendran R, Ramage G, Lang S. 2017. Efficacy of rifampicin combination therapy for the treatment of enterococcal infections assessed in vivo using a Galleria mellonella infection model. Int J Antimicrob Agents 49:507–511. doi: 10.1016/j.ijantimicag.2016.12.006. [DOI] [PubMed] [Google Scholar]
- 40.Haseltine WA, Block R. 1973. Synthesis of guanosine tetra- and pentaphosphate requires the presence of a codon-specific, uncharged transfer ribonucleic acid in the acceptor site of ribosomes. Proc Natl Acad Sci USA 70:1564–1568. doi: 10.1073/pnas.70.5.1564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Abranches J, Martinez AR, Kajfasz JK, Chávez V, Garsin DA, Lemos JA. 2009. The molecular alarmone (p)ppGpp mediates stress responses, vancomycin tolerance, and virulence in Enterococcus faecalis. J Bacteriol 191:2248–2256. doi: 10.1128/JB.01726-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Gargvanshi S, Vemula H, Gutheil WG. 2021. Effect of vancomycin on cytoplasmic peptidoglycan intermediates and van Operon mRNA levels in VanA-Type vancomycin-resistant Enterococcus faecium. J Bacteriol 203:e0023021. doi: 10.1128/JB.00230-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Baker CN, Tenover FC. 1996. Evaluation of Alamar colorimetric broth microdilution susceptibility testing method for staphylococci and enterococci. J Clin Microbiol 34:2654–2659. doi: 10.1128/jcm.34.11.2654-2659.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Rampersad SN. 2012. Multiple applications of Alamar Blue as an indicator of metabolic function and cellular health in cell viability bioassays. Sensors (Basel) 12:12347–12360. doi: 10.3390/s120912347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tyc O, Tomás-Menor L, Garbeva P, Barrajón-Catalán E, Micol V. 2016. Validation of the AlamarBlue assay as a fast screening method to determine the antimicrobial activity of botanical extracts. PLoS One 11:e0169090. doi: 10.1371/journal.pone.0169090. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material. Download spectrum.01412-22-s0001.pdf, PDF file, 0.1 MB (139.7KB, pdf)