ABSTRACT
Cells of Vibrio fischeri colonize the light organ of Euprymna scolopes, providing the squid bioluminescence in exchange for nutrients and protection. The bacteria encounter DNA-rich mucus throughout their transition to a symbiotic lifestyle, leading us to hypothesize a role for nuclease activity in the colonization process. In support of this, we detected abundant extracellular nuclease activity in growing cells of V. fischeri. To discover the gene(s) responsible for this activity, we screened a V. fischeri transposon mutant library for nuclease-deficient strains. Interestingly, only one strain, whose transposon insertion mapped to nuclease gene VF_1451, showed complete loss of nuclease activity in our screens. A database search revealed that VF_1451 is homologous to the nuclease-encoding gene xds in Vibrio cholerae. However, V. fischeri strains lacking xds eventually revealed slight nuclease activity on plates after 72 h. This led us to hypothesize that a second secreted nuclease, identified through a database search as VF_0437, a homolog of V. cholerae dns, might be responsible for the residual nuclease activity. Here, we show that Xds and/or Dns are involved in essential aspects of V. fischeri biology, including natural transformation, aggregation, and phosphate scavenging. Furthermore, strains lacking either nuclease were outcompeted by the wild type for squid colonization. Understanding the specific role of nuclease activity in the squid colonization process represents an intriguing area of future research.
IMPORTANCE From soil and water to host-associated secretions such as mucus, environments that bacteria inhabit are awash in DNA. Extracellular DNA (eDNA) is a nutritious resource that microbes dedicate significant energy to exploit. Calcium binds eDNA to promote cell-cell aggregation and horizontal gene transfer. eDNA hydrolysis impacts construction of and dispersal from biofilms. Strategies in which pathogens use nucleases to avoid phagocytosis or disseminate by degrading host secretions are well documented; significantly less is known about nucleases in mutualistic associations. This study describes the role of nucleases in the mutualism between V. fischeri and its squid host, Euprymna scolopes. We find that nuclease activity is an important determinant of colonization in V. fischeri, broadening our understanding of how microbes establish and maintain beneficial associations.
KEYWORDS: Euprymna scolopes, Vibrio fischeri, biofilm, genetic competence, nuclease, symbiosis
INTRODUCTION
Most host-bacterial symbiotic associations are comprised of multiple bacterial species comingling within a single host, which often makes it more difficult to resolve the complex interactions. Far less common are symbiotic relationships that contain one host and only one bacterial species, an example being the relationship between the luminous bacterium Vibrio fischeri and its squid host, Euprymna scolopes (1). Cells of V. fischeri colonize the microvilli-lined crypts of the squid light organ. In order to reach this tissue, cells aggregate near the opening of mucus-lined pores. These cells must then detach from the aggregates and swim through mucus-filled ducts into crypts of the light organ. Marine animal-secreted mucus has been found to be a source of extracellular DNA (eDNA) (2, 3). Extracellular DNA can strengthen the adhesivity of the mucus due to entanglement within the mucus mesh network (4). The mucus secreted by E. scolopes contains sugars and nucleotides, which attract V. fischeri cells (5, 6). DNA found in the mucus may help with the structure of the mucus and provide nutrients to migrating V. fischeri cells. In support of this hypothesis, interactions between V. fischeri and the epithelium-lined crypts enhance mucus production from the tissue (7). The importance of eDNA-containing squid mucin secretions in the colonization process led us to speculate on the role of nuclease activity in colonization.
Given the similarities between the light organ and human intestinal epithelium, it is likely that V. fischeri requires many of the same colonization determinants as V. cholerae (8). For example, the global regulator of luminescence and symbiotic competence, litR, in V. fischeri is homologous to hapR in V. cholerae (9). In V. cholerae, the hapR gene encodes a global regulator of many genes required for host colonization, including one encoding the secreted nuclease Dns (10, 11). That cells of V. fischeri possess a HapR homolog led us to hypothesize a potential role for a Dns homolog in V. fischeri.
Marine bacteria, including members of Vibrionaceae, produce nucleases as a survival mechanism (12). Some bacteria secrete nucleases as a mechanism to heighten colonization fitness through means of DNA degradation and repair, disrupting host defense mechanisms, biofilm formation, nutrient acquisition, and natural transformation (10, 11, 13–15). For example, infection by V. cholerae causes an inflammatory response in the human intestines by inducing proliferation of neutrophils that produce extracellular traps (i.e., NETs) to combat the bacteria. In turn, Xds and Dns are secreted by the cells in order to degrade the DNA component of the NETs, allowing evasion of phagocytosis (14). Similarly, the innate immune system of E. scolopes produces neutrophil-like hemocytes that adhere to and phagocytize bacteria, which V. fischeri must avoid in order to colonize squid (16). Based on these observations, we hypothesized that secreted nuclease activity may contribute to a variety of symbiosis-related activities (e.g., nutrient acquisition, hemocyte avoidance, etc.), thus enhancing the ability of V. fischeri to colonize the squid.
In this study, we identified homologs of both Xds and Dns in V. fischeri and provide data on important roles they play in bacterial physiology, evolution, and host colonization. Specifically, Dns appears to be more important in cell-cell aggregation, natural competence, and squid colonization, while Xds is more important for eDNA degradation and phosphate scavenging.
RESULTS
The V. fischeri nucleases are Xds and Dns homologs.
The V. fischeri genome encodes two secreted nucleases, VF_1451 and VF_0437, that are homologous to Xds and Dns (respectively) nucleases of V. cholerae (Fig. 1A and B). The Xds homologs shared 46% identity across the entire protein (62% similarity), while the Dns homologs shared 66% identity (77% similarity). Analysis of the amino acid sequences of Xds and Dns in V. fischeri using SignalP 5.0 (17) revealed a signal peptide cleavage site in each protein, suggesting that they are most likely secreted (Sec/SPI scores, 0.97 and 0.99, respectively). Genes flanking the xds homologs in V. fischeri and V. cholerae are different (Fig. 1C). Conversely, the dns homologs share genomic synteny (Fig. 1D).
FIG 1.
V. fischeri possesses two putative secreted nucleases that are homologs of Xds and Dns nucleases in V. cholerae. (A) The Xds proteins in V. fischeri ES114 and V. cholerae VC0395 share two functional domains, lamin tail domain (LTD; diagonal stripes) and the endonuclease/exonuclease/phosphatase family (stippled). (B) The Dns proteins share an endonuclease I functional domain (horizontal stripes). The numbers listed above the protein cartoon represent amino acid number. (C and D) Genes flanking the xds homologs are different (C), while the genes flanking dns in both species are homologs (D).
Xds is the predominant secreted nuclease in V. fischeri.
Inactivation of xds abolished the nuclease activity observed for wild-type strain ES114 after 24 h incubation on DNase agar. This defect could be restored by expression of xds in trans (Fig. 2A). After 72 h incubation, the xds mutant, but not the double xds dns mutant, eventually displayed secreted nuclease activity (Fig. 2B). These findings are supported by quantitative data obtained from liquid cultures. When cells were grown to optical density at 600 nm (OD600) values of 0.5 and 1.0, compared to wild type, inactivation of xds reduced average nuclease activity about 15- to 30-fold (respectively), while average nuclease activity in the dns mutant was not statistically different (Fig. 2C). Addition of EDTA completely abolished nuclease activity in cell-free supernatants of strain ES114, suggesting that Xds, and possibly Dns, requires divalent cations for activity (Fig. 3A). Furthermore, average nuclease activity was reduced by about 36% with the addition of excess phosphate to the growth medium (Fig. 3B).
FIG 2.
Nuclease activity in strains of V. fischeri. (A) Strains lacking xds (i.e., xds and xds-dns) were unable to degrade DNA in DNase agar after 24 h at 28°C, but expression of xds in trans in the xds single mutant (xds+) restored nuclease activity. (B) After 72 h, a halo around the xds mutant, but not the xds dns double mutant, suggested that Dns might eventually compensate for lack of Xds. (C) Nuclease activity in cells grown with shaking in LBS to an OD600 of either 0.5 or 1.0. Cell-free culture supernatants were mixed with DNaseAlert reagents to measure nuclease activity. Each bar represents the average of 5 replicates (with standard error); these results are representative of three separate experiments. Different letters indicate statistically significant differences based on analysis of variance (ANOVA) (P < 0.05).
FIG 3.
Effect of EDTA and phosphate on V. fischeri DNase activity. (A) Wild-type (ES114) cells were grown with shaking in LBS to an OD600 of 0.5 and 1.0. Cell-free supernatants with and without added EDTA (10 mmol/L) were mixed with DNaseAlert reagent to measure nuclease activity. Each bar represents the average of 5 replicates (with standard error); these results are representative of three separate experiments. (B) Cells were grown in minimal medium containing either 0.3 mM or 3 mM phosphate and then assayed for nuclease activity as described previously. Each bar represents the average of 5 replicates from three experiments (with standard error). Different letters indicate significant difference based on ANOVA. (P < 0.05).
Xds degrades linear and plasmid DNA.
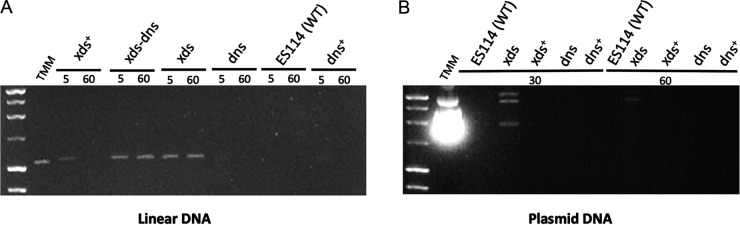
To confirm and extend these findings, we also assessed extracellular nuclease activity by mixing cell-free supernatants from V. fischeri strains with either PCR (linear) or plasmid (supercoiled; pUC19) DNA and visualizing degradation using gel electrophoresis (Fig. 4). Within 5 min, all Xds-positive (Xds+) strains (i.e., xds+, dns, ES114, and dns+) either partially or completely degraded both linear (PCR) and plasmid DNA (Fig. 4A and B). After 30 to 60 min, the Xds+ strains completely degraded either type of DNA. Strains that lacked Xds (i.e., xds and xds-dns) exhibited markedly reduced DNA degradation during the incubation periods we tested. Together, these data support the hypothesis that Xds is a key protein involved in degrading both linear and supercoiled DNA in V. fischeri.
FIG 4.
Degradation of eDNA by V. fischeri strains. Strains were cultured in TMM to an OD600 of approximately 2.0. Either sterile TMM or cell-free supernatant from each strain was mixed with ~0.2 μg of PCR DNA (linear) (A) or 1 μg of plasmid (pUC19) (B) incubated at 28°C for the times (in minutes) indicated and then visualized on a 1% agarose gel.
Cells of V. fischeri can use eDNA as their sole phosphate source.
Inactivation of xds, and to a lesser extent, dns, resulted in an overall reduced growth rate and cell yield for V. fischeri strains grown with eDNA as the sole phosphate source (Fig. 5A). Specifically, log-phase Xds− cells (i.e., xds, xds-dns) required roughly 4 h to double, while Xds+ cells (i.e., ES114, dns) doubled in just over 1 h. Furthermore, after 16 h, the Xds− strains reached an OD600 of only about half that of the Xds+ strains. However, OD600 for the dns mutant at 16 h was about 15% lower (~1.0 versus ~0.85) than that of the wild-type strain ES114, despite starting at roughly the same OD600. All strains grew similarly in Tris-minimal medium (TMM) with K2HPO4 (Fig. 5B), while none of the strains grew in the absence of a phosphate source (Fig. 5C). Taken together, V. fischeri has the capacity to use eDNA as a phosphate source that is dependent on Xds and, to a lesser extent, Dns.
FIG 5.
Secreted nucleases are essential for optimal growth when eDNA is the only source of phosphate. Cells were grown at 28°C with shaking in TMM containing sheared salmon sperm DNA (1.0 mg mL−1) (A) or K2HPO4 (B) as the sole phosphate source or no phosphate (C). Error bars represent averages (± standard error) for three separate trials.
Xds might play a role in mucin degradation.
Cells of V. fischeri encounter abundant DNA-containing mucin upon colonizing squid (5). To determine whether nuclease might play a role in degrading the DNA component of mucin, strains were grown on a medium containing mucin as the sole nutrient source. Consistent with our other findings, and unlike the wild-type strain, the Xds− strains (i.e., xds and xds-dns) were unable to degrade the DNA component of mucin after 24 h (Fig. 6). Expression of xds in trans (i.e., xds+) restored full wild-type activity.
FIG 6.

Xds is required to digest DNA in mucin. The absence of a halo around the xds mutants (i.e., xds, xds-dns) suggests that, for at least 24 h, no other nuclease participates in the breakdown of DNA in mucin. Complementation of xds in trans (i.e., xds+) partially restored the ability to digest mucin. Two versions of double mutant (i.e., xds-dns and xds-dns/KV8865) showed no difference in nuclease activity on mucin plates.
Dns controls cell-cell aggregation in V. fischeri.
Enhanced aggregation in strains lacking dns, but not xds, was apparent after extended incubation (i.e., 5 to 7 days) in static liquid culture (Fig. 7). Thus, we conclude that (i) eDNA may contribute to biofilm formation by V. fischeri that occurs following extended static growth, and (ii) Dns, but not Xds, appears to function to inhibit eDNA-dependent biofilm formation.
FIG 7.
Dns mutant cells display a hyperaggregation phenotype. Cells were grown in minimal medium without shaking for up to 1 week and then gently mixed to visualize aggregation. Only strains lacking dns (i.e., dns and xds-dns) showed an aggregation phenotype.
Dns acts a barrier to natural transformation.
Our previous work demonstrated a role for dns in tfoX-induced competence (18) but did not evaluate the impact of loss of Xds or of both nucleases. As previously reported, inactivation of dns increased transformation efficiency by ~2 orders of magnitude (Fig. 8). Surprisingly, despite its greater effect on extracellular nuclease activity, loss of Xds did not alter transformation efficiency: it was not statistically different from that of the wild-type strain, although it trended slightly higher in some of our experiments. Similarly, the xds-dns strain exhibited a comparable efficiency to that of the dns mutant. Complementation of the dns mutant by introduction of dns at a nonnative site substantially reduced the transformation efficiency such that no transformants were obtained (Fig. 8A). To confirm that the complemented strain was still competent to take up DNA, we introduced into the strain (and its controls) a different tfoX overexpression plasmid, one that we previously showed to induce a higher transformation frequency (18). We then repeated the competence experiments with the wild type, dns (i.e., KV8807), and complemented dns strains (i.e., dns+ KV10274). Under these genetic conditions, low numbers of transformants were obtained for the complemented dns mutant, indicating the strain was capable of DNA uptake (Fig. 8B). Together, these data reveal that Dns is the key nuclease controlling competence.
FIG 8.

The dns mutant is hypertransformable. Strains ES114, dns (KV8807), xds (KV8811), xds-dns (KV8865), and dns+ (KV10274) carrying a tfoX expression plasmid with either kanamycin (A) or chloramphenicol resistance (B) were grown at 28°C with shaking in TMM to an OD600 of approximately 0.5. Aliquots of each culture were exposed to purified DNA containing an antibiotic resistance marker. Cells were plated on selective media, transformants were counted to estimate transformation frequency, and an ANOVA test was performed assuming normal distribution. Dotted lines indicate limit of detection. *, P < 0.05; **, P < 0.005; ****, P < 0.00005; ns, not significant.
Xds and Dns are required for efficient colonization of E. scolopes.
Single-strain colonization assays performed in triplicate revealed that cells of V. fischeri lacking xds or dns were like the wild type in that they colonized squid with 100% efficiency by 24 h postinfection (Fig. 9A). Relative to wild type, the xds strain produced roughly equivalent amounts of light per squid, while the dns strain emitted, on average, about 20-fold less light per squid by 48 h post-initial exposure. Because we did not detect a luminescence defect in either the xds or dns strains (Fig. 9B), these findings suggest that inactivation of dns results in reduced colonization efficiency. In competition experiments, combining the wild type with either nuclease mutant in equal proportions, differences in colonization efficiency were magnified. The wild-type strain outcompeted the xds and dns strains in about 75% and 92% of squid, respectively; indeed, for three squid, the dns mutant was excluded (Fig. 10). These data indicate that expression of dns and, to a lesser extent, xds, provides a colonization advantage to V. fischeri.
FIG 9.
Single-strain squid colonization. (A) Closed squares, closed triangles, and open circles represent bioluminescence emission per squid infected with either xds, dns, or the wild-type strain ES114, respectively. Red lines represent average relative light units (RLU) per squid over the 48-h time period. The number of colonized squid was determined by comparing RLU emission between aposymbiotic squid and the test strains. (B) Nuclease is not required for normal growth or luminescence in culture. All results shown are typical for three independent trials.
FIG 10.

Nucleases are required for optimal squid colonization. Juvenile squid were exposed to an inoculum containing a 1:1 ratio of wild-type V. fischeri and either the xds (A) or dns (B) mutant. At 48 h postinfection, the ratio of mutant to wild-type bacteria in each light organ was assessed by plating squid light organs. The results are expressed as the relative competitive index (RCI). An RCI above 1.0 indicates that the mutant was dominant. These results are representative of three separate trials.
DISCUSSION
V. fischeri produces two nucleases that are homologs of the exonucleases Xds and Dns first characterized in V. cholerae (10, 11). Xds activity was the predominant nuclease detected in culture, with Dns activity being most apparent in an xds mutant background after extended incubation. Xds of V. cholerae requires divalent cations (i.e., Ca2+ and Mg2+) (19), and this seems to be the case for V. fischeri as well, as secreted nuclease activity was completely abolished by the addition of EDTA. Both nucleases possess signal peptide cleavage sites, suggesting that they are secreted by the general secretion (Sec) protein export pathway. Closer homology between the dns homologs in V. fischeri and V. cholerae is consistent with their genomic synteny and shared roles in biofilm formation and natural transformation (10, 11). Similarly, the relatively greater evolutionary distance in Xds proteins is consistent with their lack of synteny. Still, the role of Xds in P. aeruginosa, V. cholerae, and V. fischeri (i.e., degradation of eDNA to nucleotides for nutritional purposes) is conserved (15, 19).
In V. cholerae, dns is negatively regulated at high cell density by HapR (10), which is homologous to LitR in V. fischeri (9). However, evidence from microarrays revealed that LitR does not affect either xds or dns expression at OD600 values of up to 2.0 (20). Consistent with those data, the loss of Dns did not substantially alter nuclease activity when assessed at either OD of 0.5 or 2. However, Dns activity might become important at later stages of growth, as evidenced by the late impact of the loss of Dns on growth on eDNA (Fig. 5A), plasmid DNA digestion (Fig. 4B), eDNA digestion on DNase agar (Fig. 2B), and biofilm aggregate formation (Fig. 7). Whether this late requirement for Dns is due to transcriptional control via LitR that was not apparent from the microarray or is due to another factor remains to be determined. In V. cholerae, xds is regulated by the two-component system PhoR/PhoB and phosphate limitation (21). Our findings suggested that nuclease activity is phosphate dependent (Fig. 3B), but additional work will be required to unravel the regulatory mechanisms for the two V. fischeri nucleases.
Both V. fischeri nuclease mutants exhibited colonization defects, with the dns mutant being more severely impaired. These mutants displayed multiple phenotypes to differing extents, including biofilm formation and ability to degrade and/or grow on eDNA. Any or all of these traits could be critical for normal symbiotic colonization. For example, biofilm formation is critical for cells of V. fischeri to initiate the squid colonization process (22). Initially, cells are collected in aggregates outside the light organ pores and must eventually detach and swim into the light organ interior (5). Deletion of rscS, encoding a sensor kinase that promotes biofilm formation, severely impaired squid colonization, while overexpression enhanced it (22, 23). In this study, the dns mutant displayed an increase in cell-cell aggregation, potentially due to the presence of excess eDNA on the bacterial surface (24). However, unlike with rscS overexpression leading to excess biofilm and superior colonization, the dns mutant was associated with a severe colonization defect, suggesting that hyperaggregation may be detrimental. While biofilm formation (aggregation) is essential in the early phases of colonization, V. fischeri cells also need to detach from the aggregate and/or squid mucus to reach the light organ interior where colonization proceeds (5). Thus, it is possible that the hyperaggregation of the dns mutant impaired cellular detachment, resulting in fewer cells reaching the light organ interior.
Alternatively, the importance of Dns in colonization could stem from a role in generating/altering a specific structure of the biofilm. For example, biofilms produced by many bacteria are laden with extracellular DNA that can be degraded and remodeled by secreted nucleases to increase colonization fitness when bacteria enter a host (11). In addition, host secretions may contribute to biofilm formation and adherence and/or inhibit dissemination (25). Pus and mucus secretions contain eDNA, which can strengthen adhesivity (25, 26). Pathogenic microbes such as V. cholerae cells are proposed to utilize extracellular nucleases to degrade mucus secreted in the host small intestine (27). V. fischeri may likewise use nucleases to degrade or alter bacterial eDNA and/or squid-secreted mucus. While our findings suggested that Xds is primarily responsible for degrading the DNA component of mucus, it is possible that symbiosis-specific conditions enhance the production or activity of Dns. Alternatively, the less severe defect of the xds mutant during competitive colonization experiments may stem from exogenous complementation by the wild-type strain present in the same location.
Other known eDNA-related host mechanisms include the production of neutrophil (PMN) extracellular traps (NETs) (28). In V. cholerae, both Xds and Dns contribute to the degradation of NETs, promoting colonization of the human gastrointestinal tract (14). In the squid symbiosis, PMN-like hemocytes are recruited to the light organ interior by newly colonizing cells of V. fischeri (29). One role of hemocytes is to remove undesirable microbes from the light organ interior, although there is currently no evidence of an interaction between V. fischeri and eDNA-containing NETs.
The Dns and Xds nucleases may also be important in symbiotic colonization due to their role in nutrient acquisition (Fig. 5A). Microbes such as V. cholerae and P. aeruginosa use Xds homologs to scavenge phosphate from eDNA in nutrient-poor environments (15, 19). Heterogenous distribution of phosphate in the squid light organ suggests that there is selective pressure to optimize phosphate scavenging, potentially via eDNA nuclease activity (30).
Last, we also investigated the relationship between nuclease secretion and natural transformation. Secreted nucleases affect natural competency in a wide range of bacteria, such as V. cholerae and Staphylococcus aureus (10, 31). For example, inactivation of dns in V. cholerae resulted in a hypertransformable phenotype (10). Similarly, inactivation of dns, but not xds, resulted in a hypertransformable phenotype by V. fischeri (Fig. 8). Xds catalyzes readily demonstrable, fairly rapid degradation of linear and plasmid eDNA (Fig. 4). Thus, on the surface, it is surprising that the major secreted nuclease Xds did not impact transformation. However, these data corroborate work in V. cholerae (10) and demonstrate that the roles of the two nucleases are distinct, potentially in part due to a difference in where these proteins localize. Our findings support a model in which Xds is secreted, while Dns is periplasmic but leaks into the environment over time. Future studies will determine whether Dns in V. fischeri is localized to the periplasm as in V. cholerae (32). Taken together, these findings contribute to our understanding of competence in V. fischeri, which has far-reaching implications for understanding how this microbe evolves and how it can be manipulated in the laboratory.
This study thus reveals a range of activities associated with secreted nucleases that are vital to the beneficial squid symbiont V. fischeri. These activities include phosphate acquisition, cell-cell aggregation, natural competence, and host colonization. Investigations into specific aspects of the squid-Vibrio association, such as aggregation and avoidance of hemocyte NETs, should allow us to bear down on the mechanisms by which nucleases might influence these critical colonization processes.
MATERIALS AND METHODS
Strains and media.
Strains and plasmids used in this study are listed in Table 1. The parent V. fischeri strain used in this study (e.g., ES114) was isolated from E. scolopes (33) and grown aerobically using various media at 24 to 28°C. For molecular biology and enzyme assay applications, V. fischeri cells were cultured in lysogeny broth salt (LBS) containing 10 g of tryptone, 5 g of yeast extract, 20 g of NaCl, and 20 mM Tris-hydrochloride (Tris-HCl, pH 7.5) per L of sterile water. All squid colonization and bioluminescence assays used seawater tryptone (SWT), which contained 5 g of tryptone, 3 g of yeast extract, 3 mL of glycerol, and 700 mL of artificial seawater (ASW) per L of sterile water. Escherichia coli was cultured in LB-Miller, which contained 10 g tryptone, 5 g yeast extract, and 10 g NaCl. Agar was added at 15 mg mL−1 for medium solidification for plating. Where necessary, antibiotics chloramphenicol, erythromycin, and kanamycin were added to growth media at 2.5, 5, and 100 μg mL−1, respectively, for V. fischeri and 30, 100, and 150 μg mL−1, respectively, for E. coli. DNase activity was assessed using DNase agar (Thermo Fisher), as well as porcine mucin agar containing 1% mucin (Sigma-Aldrich), 500 mL of ASW per L of sterile water, 5% Tris-HCl (1 M, pH 7.4), 0.3% glycerol, 0.006% K2HPO4, and 1.5% agar. To visualize nuclease activity, DNase and mucin agar plates were treated with either 1 N HCl or 0.1% amido black in 3.5 M acetic acid, respectively. Mucin plates were destained with 1.2 M acetic acid. Liquid medium for aggregation assays was a modified HEPES-minimal medium that contained artificial seawater, 0.1% (wt/vol) ammonium chloride, 0.0058% K2HPO4, 10 μM ferrous ammonium sulfate, 100 mM HEPES, pH 7.5, and 0.03% Casamino Acids. For natural transformation and in vitro luminescence experiments, V. fischeri strains were grown in Tris-minimal medium (TMM) (100 mM Tris, pH 7.5, 300 mM NaCl, 0.1% ammonium chloride, 10 mM N-acetylglucosamine, 50 mM MgSO4, 10 mM KCl, 10 mM CaCl2, 0.0058% K2HPO4, and 10 μM ferrous ammonium sulfate).
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Description | Reference or source |
|---|---|---|
| Bacterial strains | ||
| ES114 | Wild type | 33 |
| xds | xds::miniTn5 ermr | This study |
| xds (KV8811) | Δxds::FRT-erm | This study |
| dns | Δdns::FRT | This study |
| dns (KV8807) | Δdns::FRT-erm | 18 |
| xds+ | xds + pVSV105/xds cmr | This study |
| dns+ | dns + pVSV105/dns cmr | This study |
| dns+ | Δdns::FRT IG (yeiR-FRT-Erm/glmS)::PnrdR dns | This study |
| xds-dns | xds::miniTn5 ermr/Δdns::FRT | This study |
| xds-dns (KV8865) | Δxds::FRT Δdns::FRT | This study |
| Plasmids | ||
| pESV104 | Helper plasmid (tra+, mob+, knr) | 38 |
| pVSV105 | Shuttle vector (cmr) | 40 |
| pEVS170 | miniTn5 delivery plasmid (ermr) | 38 |
| plosTfoX-Cm | TfoX overexpression (cmr) | 36 |
| plosTfoX-Kn | TfoX overexpression (knr) | 37 |
Constructing nuclease mutants and complementation.
Mutants were constructed either by random transposon mutagenesis (xds) (34, 35) or by TfoX-mediated transformation and in-frame allelic replacement (dns [KV8807], xds [KV8811], xds-dns, and xds-dns [KV8865]), and complemented dns+ (KV10274) (36, 37) strains were constructed by TfoX-mediated transformation. Transposon delivery to create the xds strain was carried out using an E. coli donor strain (carrying pEVS170) and E. coli helper strain CC118 λpir carrying pEVS104 (38). Two thousand mutants were streak purified on LBS and then patched onto DNase agar. After 24 h growth, plates were flooded with 1 N HCl in order to detect a patch lacking nuclease activity. For Tfox-mediated transformation and allelic exchange, we introduced a tfoX overexpression plasmid from E. coli via triparental conjugations using an E. coli strain containing the tfoX overexpression plasmid, a pEVS104 (38)-containing E. coli, and the recipient V. fischeri strain. Strains containing the tfoX plasmid were then transformed with either PCR DNA derived from PCR splicing by overlap extension (SOE) reactions or from genomic DNA obtained from a strain carrying an antibiotic-marked deletion or insertion. DNA fragments for PCR SOE were constructed as previously described (39) using high-fidelity PCR enzyme EMD Millipore KOD DNA polymerase, and primers are listed in Table 2. The reactions allowed for construction of a fragment that contains an antibiotic resistance cassette flanked by sequences homologous to ~500-bp sequences up- and downstream of the gene or region of interest. The PCR or genomic DNA was transformed into tfoX overexpression strains, and the resulting cultures were plated onto a selective medium. Colonies that arose were passaged onto the same selective medium once more and then onto nonselective medium to facilitate loss of the unstable overexpression plasmid. Strains were confirmed to have the insertion or deletion of interest via a PCR using outside primers. The dns complementation cassette was sequenced (ACGT, Inc., Wheeling, IL) to ensure no mutations arose. To remove antibiotic cassettes (Ermr or Trimr) from the insertion mutants and complementation strains, we performed a triparental mating with the V. fischeri strain of interest, an E. coli strain bearing pEVS104, and an E. coli strain bearing pKV496 containing the flippase that recognizes FRT sites as previously described (38), followed by selection for pKV496 (Kanr). Conjugants were then assessed for sensitivity to erythromycin (Erm) or trimethoprim (Trim). Sensitive colonies were confirmed by PCR to have lost the antibiotic cassette and were passaged nonselectively to facilitate loss of pKV496.
TABLE 2.
Primers used in this study
| Name | Sequence | Purpose |
|---|---|---|
| Xds-NdeI-F | CCCCATATGGAGCTCTGCCAGATTTGTAG | Complement VF_1451 (xds) |
| Xds-NdeI-R | CCCCATATGTGCAAAGTGGTGCAGAAACC | Complement VF_1451 (xds) |
| Dns-NdeI-F | CCCCATATGCGGTACGTCGCCATAATCGA | Complement VF_0437 (dns) |
| Dns-NdeI-R | CCCCATATGTTCGCATTACACGACCAACG | Complement VF_0437 (dns) |
| VIPCR-F | CCTAGAGCGGCCGCAGA | Tn localization in VF_1451 (xds) |
| VIPCR-RBio | Biotin-ACTGGCCGTCGTTTTACAG | Tn localization in VF_1451 (xds) |
| MoSeq-F | AGATGTGTATAAGAGAC | Tn localization in VF_1451 (xds) |
| 2508 | CTTCCAGCCCATAACTCTCC | Delete VF_1451 (xds); strain KV8811 |
| 2509 | taggcggccgcactaagtatggACTTTCCATAGTTGAGTTATATGAA | Delete VF_1451 (xds); strain KV8811 |
| 2089 | CCATACTTAGTGCGGCCGCCTA | Delete VF_1451 (xds); strain KV8811 |
| 2090 | CCATGGCCTTCTAGGCCTATCC | Delete VF_1451 (xds); strain KV8811 |
| 2510 | ggataggcctagaaggccatggACGTAAATAACCGTTATAAAGAGC | Delete VF_1451 (xds); strain KV8811 |
| 2511 | GCAAGATTACATCGTTAAAGCC | Delete VF_1451 (xds); strain KV8811 |
| 2090 | CCATGGCCTTCTAGGCCTATCC | Complement VF_0437 (dns); strain KV10274 |
| 2290 | AAGAAACCGATACCGTTTACG | Complement VF_0437 (dns); strain KV10274 |
| 4033 | ggataggcctagaaggccatggAGGAGGAAGTTATTATGAAATTAATCCGCTTAC | Complement VF_0437 (dns); strain KV10274 |
| 3353 | taggcggccgcactaagtatggaGGAGGTTAAAGTATGCTTTTTCAACC | Complement VF_0437 (dns); strain KV10274 |
| 2196 | tccatacttagtgcggccgccta | Complement VF_0437 (dns); strain KV10274 |
| 1487 | GGTCGTGGGGAGTTTTATCC | Complement VF_0437 (dns); strain KV10274 |
The dns and xds dns mutants were constructed using PCR splicing by overlap extension (PCR SOE) to remove dns through a series of PCRs (39). For in-frame allelic replacement, resolution of the FRT-flanked antibiotic resistance cassettes was achieved using the flippase plasmid pKV496 (39). For complementation of either xds or dns, the gene (including the ribosome binding site through the stop codon) was amplified, inserted into pVSV105 plasmid (40), and then mobilized into the recipient cells by triparental mating. Primers for Xds (Xds-NdeI-F/R) and Dns (Dns-NdeI-F/R) complementation are listed in Table 2.
Determining the site of Tn5 insertion in V. fischeri strains.
Transposon insertion was localized in nuclease mutants as described previously (35). Genomic DNA was extracted from nuclease mutants, cut with Sau3AI, and self-ligated to form closed circular DNA fragments. Next, fragments were subjected to inverse PCR (iPCR) using primers VIPCR-F and VIPCR-RBio to amplify DNA flanking the transposon insertion, and then amplicons were sequenced using primer MoSeq-F and a pyrosequencer to identify the flanking DNA (Table 2).
eDNA degradation assay.
All strains of V. fischeri were incubated in TMM overnight with shaking at 225 rpm at 28°C to an optical density at 600 nm (OD600) of about 2.0. One milliliter of culture was pelleted by centrifugation. The cell-free supernatant was withdrawn and combined with purified PCR or plasmid DNA. The mixtures were incubated from 5 to 60 min at 28°C. The degree of DNA degradation was assessed on a 1% agarose gel.
Quantitative nuclease assay.
Nuclease activity was quantified using the DNaseAlert QC system (Thermo Fisher), according to the manufacturer’s instructions. V. fischeri strains were grown with shaking in LBS at 28°C to OD600 readings of 0.5 and 1.0. One-milliliter aliquots of culture were pelleted by centrifugation. Eighty microliters of each supernatant was combined with 10 μL of 10× NucleaseAlert buffer and 10 μL DNaseAlert substrate in wells of a microtiter plate and incubated at 37°C for 60 min. Fluorescence was measured using a Cytation 5 imaging reader (BioTek Instruments, Inc.). DNase I (2 U μL−1) was utilized as a positive control. To measure the effects of a chelating agent on nuclease activity, some supernatants were also combined with 10 mmol L−1 EDTA.
Role of nuclease activity in phosphate scavenging.
Strains were grown with shaking at 225 rpm in TMM overnight. TMM was prepared as a complete medium (i.e., containing all nutrients to support V. fischeri cells) or without a phosphate (i.e., K2HPO4) source. Media without a phosphate source were supplemented with exogenous salmon sperm DNA to a final concentration of 1.0 mg mL−1. Cells of nuclease mutants and wild-type V. fischeri were inoculated into separate wells of a microtiter plate. Initial OD600 was ~0.015 for all cultures. The cells were grown at 28°C in a Cytation 5 imaging reader (BioTek Instruments, Inc.) with shaking at 237 rpm for 20 h. Optical density readings at 600 nm were taken every 10 min.
Transformation assay.
Transformation assays were performed as described in Cohen et al. (18), using ES114 (wild-type strain) and the following genetically modified strains: dns (KV8807), xds (KV8811), xds-dns (KV8865), and dns+ (KV10274). In brief, V. fischeri strains carrying tfoX overexpression plasmids with kanamycin (plostfoX-Kn) (37) or chloramphenicol (plostfoX-Cm) (36) resistance were grown overnight in TMM containing the appropriate antibiotic. In ES114-derived strains, plostfoX-Kn promotes a transformation efficiency that is over 10× lower than that induced by plostfoX-Cm and thus permits experimental observations of increased transformation efficiency such as seen with a dns mutant (18). Cells were subcultured into fresh TMM and grown to an OD600 of about 0.5. Cells were inoculated with chromosomal DNA (~0.5 μg) from strain KV8300 (ΔfliQ::Frt-Trim) (39) or left untreated and incubated at room temperature for 30 min without shaking. LBS (0.5 mL) was added to the cells in TMM, and the cultures were grown with shaking at 28°C for at least 90 min. The cultures were then serial diluted or left undiluted, and 100 μL of each serial dilution was plated onto antibiotic plates (LBS containing trimethoprim) for selection of transformants. The plates were incubated overnight at 28°C. Total colony counts were performed to assess transformation efficiency at each dilution. Untreated control samples were diluted and plated onto LBS plates alone to obtain total CFU. As in described in Cohen et al. (18), we calculated transformation frequency by multiplying the number of antibiotic-resistant transformants by the dilution factor and dividing by the total calculated CFU.
Squid colonization.
Juvenile E. scolopes were collected within minutes of hatching. Hatchlings were removed with plastic eye droppers and held in artificial seawater at 22°C for at least 30 min before inoculating the water with 3,000 CFU of either wild-type V. fischeri, the xds mutant, or the dns mutant. Individual squid were then washed in sterile seawater placed in their own scintillation vial filled with 5 mL of artificial seawater. Luminescence emission for each squid was measured for up to 48 h using a GloMax 20/20 luminometer. Luminescence emission was used to assess colonization effectiveness. For cocolonization assays, squid were exposed to roughly a 1:1 mixture of the wild-type strain and either mutant for 12 h; squid were then washed and transferred to symbiont-free seawater for the remainder of the experiment. After 48 h, squid light organs were plated on selective and nonselective media to assess the relative proportions of each strain. All animal experiments were conducted in compliance with protocol number A18-029 approved by the Institutional Animal Care and Use Committee, University of Connecticut.
In vitro luminescence and growth.
Strains were grown with shaking at 28°C in SWT. At regular intervals, aliquots of each culture were assayed for cell density in a spectrophotometer and luminescence in a GloMax 20/20 luminometer.
Data availability.
Supporting data will be made available upon reasonable request.
ACKNOWLEDGMENTS
We thank Josh Cohen for constructing and contributing strains for this project.
This work was supported by funding from the NIH, R35 GM130355, awarded to K.L.V.
Contributor Information
Pat M. Fidopiastis, Email: pfidopia@calpoly.edu.
Karyn N. Johnson, University of Queensland
REFERENCES
- 1.Visick K, Ruby E. 2006. Vibrio fischeri and its host: it takes two to tango. Curr Opin Microbiol 9:632–638. 10.1016/j.mib.2006.10.001. [DOI] [PubMed] [Google Scholar]
- 2.Livia L, Antonella P, Hovirag L, Mauro N, Panara F. 2006. A nondestructive, rapid, reliable and inexpensive method to sample, store and extract high-quality DNA from fish body mucus and buccal cells. Mol Ecol Notes 6:257–260. 10.1111/j.1471-8286.2005.01142.x. [DOI] [Google Scholar]
- 3.Domingues RR, Garrone-Neto D, Hilsdorf AWS, Gadig OBF. 2019. Use of mucus as a non-invasive sampling method for DNA barcoding of stingrays and skates (batoid elasmobranchs). J Fish Biol 94:512–516. 10.1111/jfb.13919. [DOI] [PubMed] [Google Scholar]
- 4.Duncan G, Jung J, Hanes J, Suk J. 2016. The mucus barrier to inhaled gene therapy. Mol Ther 24:2043–2053. 10.1038/mt.2016.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nyholm SV, Stabb EV, Ruby EG, McFall-Ngai MJ. 2000. Establishment of an animal–bacterial association: recruiting symbiotic vibrios from the environment. Proc Natl Acad Sci USA 97:10231–10235. 10.1073/pnas.97.18.10231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.DeLoney-Marino C, Wolfe A, Visick K. 2003. Chemoattraction of Vibrio fischeri to serine, nucleosides, and N-acetylneuraminic acid, a component of squid light-organ mucus. Appl Environ Microbiol 69:7527–7530. 10.1128/AEM.69.12.7527-7530.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nyholm S, Deplancke B, Gaskins H, Apicella M, McFall-Ngai M. 2002. Roles of Vibrio fischeri and nonsymbiotic bacteria in the dynamics of mucus secretion during symbiont colonization of the Euprymna scolopes light organ. Appl Environ Microbiol 68:5113–5122. 10.1128/AEM.68.10.5113-5122.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ruby EG, Urbanowski M, Campbell G, Dunn A, Faini M, Gunsalus R, Lostroh P, Lupp C, McCann J, Millikan D, Schaefer A, Stabb E, Stevens A, Visick K, Whistler C, Greenberg EP. 2005. Complete genome sequence of Vibrio fischeri: symbiotic bacterium with pathogenic congeners. Proc Natl Acad Sci USA 102:3004–3009. 10.1073/pnas.0409900102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fidopiastis PM, Miyamoto CM, Jobling MG, Meighen EA, Ruby EG. 2002. LitR, a new transcriptional activator in Vibrio fischeri, regulates luminescence and symbiotic light organ colonization. Mol Microbiol 45:131–143. 10.1046/j.1365-2958.2002.02996.x. [DOI] [PubMed] [Google Scholar]
- 10.Blokesch M, Schoolnik GK. 2008. The extracellular nuclease Dns and its role in natural transformation of Vibrio cholerae. J Bacteriol 190:7232–7240. 10.1128/JB.00959-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Seper A, Fengler V, Roier S, Wolinski H, Kohlwein S, Bishop AL, Camilli A, Reidl J, Schild S. 2011. Extracellular nucleases and extracellular DNA play important roles in Vibrio cholerae biofilm formation. Mol Microbiol 82:1015–1037. 10.1111/j.1365-2958.2011.07867.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Al-Wahaibi ASM, Lapinska E, Rajarajan N, Dobretsov S, Upstill-Goddard R, Burgess JG. 2019. Secretion of DNases by marine bacteria: a culture based and bioinformatics approach. Front Microbiol 10:969. 10.3389/fmicb.2019.00969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Berends ET, Horswill AR, Haste NM, Monestier M, Nizet V, von Kockritz, Blickwede M. 2010. Nuclease expression by Staphylococcus aureus facilitates escape from neutrophil extracellular traps. J Innate Immun 2:576–586. 10.1159/000319909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Seper A, Hosseinzadeh A, Gorkiewicz G, Lichtenegger S, Roier S, Leitner DR, Rohm M, Grutsch A, Reidl J, Urban CF, Schild S. 2013. Vibrio cholerae evades neutrophil extracellular traps by the activity of two extracellular nucleases. PLoS Pathog 9:e1003614. 10.1371/journal.ppat.1003614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mulcahy H, Charron-Mazenod L, Lewenza S. 2010. Pseudomonas aeruginosa produces an extracellular deoxyribonuclease that is required for utilization of DNA as a nutrient source. Env Microbiol 12:1621–1629. 10.1111/j.1462-2920.2010.02208.x. [DOI] [PubMed] [Google Scholar]
- 16.Nyholm S, Stewart J, Ruby E, McFall-Ngai M. 2009. Recognition between symbiotic Vibrio fischeri and the haemocytes of Euprymna scolopes. Environ Microbiol 11:483–493. 10.1111/j.1462-2920.2008.01788.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Armenteros JJA, Tsirigos KD, Sønderby CK, Petersen TN, Winther O, Brunak S, Heijne GV, Nielsen H. 2019. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol 37:420–423. 10.1038/s41587-019-0036-z. [DOI] [PubMed] [Google Scholar]
- 18.Cohen JJ, Eichinger SJ, Witte DA, Cook CJ, Fidopiastis PM, Tepavcevic J, Visick KL. 2021. Control of competence in Vibrio fischeri. Appl Environ Microbiol 87:e01962-20. 10.1128/AEM.01962-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pressler K, Mitterer F, Vorkapic D, Reidl J, Oberer M, Schild S. 2019. Characterization of Vibrio cholerae's extracellular nuclease Xds. Front Microbiol 10:2057. 10.3389/fmicb.2019.02057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Siegl A. 2004. The regulatory network of LitR in Vibrio fischeri: a microarray-based characterization. Master’s thesis. University of Hawaii, Manoa, Honolulu, HI. [Google Scholar]
- 21.McDonough E, Lazinski DW, Camilli A. 2014. Identification of in vivo regulators of the Vibrio cholerae xds gene using a high-throughput genetic selection. Mol Microbiol 92:302–315. 10.1111/mmi.12557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yip E, Geszvain K, DeLoney-Marino C, Visick K. 2006. The symbiosis regulator RscS controls the syp gene locus, biofilm formation and symbiotic aggregation by Vibrio fischeri. Mol Microbiol 62:1586–1600. 10.1111/j.1365-2958.2006.05475.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Visick KL, Skoufos LM. 2001. Two-component sensor required for normal symbiotic colonization of Euprymna scolopes by Vibrio fischeri. J Bacteriol 183:835–842. 10.1128/JB.183.3.835-842.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Das T, Sehar S, Koop L, Wong YK, Ahmed S, Siddiqui K, Manefield M. 2014. Influence of calcium in extracellular DNA mediated bacterial aggregation and biofilm formation. PLoS One 9:e91935. 10.1371/journal.pone.0091935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lister JL, Horswill AR. 2014. Staphylococcus aureus biofilms: recent developments in biofilm dispersal. Front Cell Infect Microbiol 4:178. 10.3389/fcimb.2014.00178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sherry S, Tillett WS, Christensen LR. 1948. Presence and significance of desoxyribose nucleoprotein in the purulent pleural exudates of patients. Proc Soc Exp Biol Med 68:179–184. 10.3181/00379727-68-16428. [DOI] [PubMed] [Google Scholar]
- 27.Focareta T, Manning PA. 1991. Distinguishing between the extracellular DNases of Vibrio cholerae and development of a transformation system. Mol Microbiol 5:2547–2555. 10.1111/j.1365-2958.1991.tb02101.x. [DOI] [PubMed] [Google Scholar]
- 28.Storisteanu DML, Pocock JM, Cowburn AS, Juss JK, Nadesalingam A, Nizet V, Chilvers ER. 2017. Evasion of neutrophil extracellular traps by respiratory pathogens. Am J Respir Cell Mol Biol 56:423–431. 10.1165/rcmb.2016-0193PS. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.McAnulty SJ, Nyholm SV. 2016. The role of hemocytes in the Hawaiian bobtail squid, Euprymna scolopes: a model organism for studying beneficial host–microbe interactions. Front Microbiol 7:2013. 10.3389/fmicb.2016.02013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Stoudenmire JL, Essock-Burns T, Weathers EN, Solaimanpour S, Mrazek J, Stabb EV. 2018. An iterative, synthetic approach to engineer a high-performance PhoB-specific reporter. App Env Microbiol 84:e00603-18. 10.1128/AEM.00603-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rudin L, Sjostrom J, Lindberg M, Philipson L. 1974. Factors affecting competence for transformation in Staphylococcus aureus. J Bacteriol 118:155–164. 10.1128/jb.118.1.155-164.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Seitz P, Blokesch M. 2014. DNA transport across the outer and inner membranes of naturally transformable Vibrio cholerae is spatially but not temporally coupled. mBio 5:e01409-14. 10.1128/mBio.01409-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Boettcher KJ, Ruby EG. 1990. Depressed light emission by symbiotic Vibrio fischeri of the sepiolid squid Euprymna scolopes. J Bacteriol 172:3701–3706. 10.1128/jb.172.7.3701-3706.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lyell NL, Septer AN, Dunn AK, Duckett D, Stoudenmire JL, Stabb EV. 2017. An expanded transposon mutant library reveals that Vibrio fischeri δ-Aminolevulinate auxotrophs can colonize Euprymna scolopes. Appl Environ Microbiol 83:e02470-16. 10.1128/AEM.02470-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Stoudenmire JL, Black M, Fidopiastis PM, Stabb EV. 2019. Mutagenesis of Vibrio fischeri and other marine bacteria using hyperactive Mini-Tn5 derivatives. Methods Mol Biol 2016:87–104. 10.1007/978-1-4939-9570-7_9. [DOI] [PubMed] [Google Scholar]
- 36.Pollack-Berti A, Wollenberg MS, Ruby EG. 2010. Natural transformation of Vibrio fischeri requires tfoX and tfoY. Env Microbiol 12:2302–2311. 10.1111/j.1462-2920.2010.02250.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Brooks JF, Gyllborg MC, Cronin DC, Quillin SJ, Mallama CA, Foxall R, Whistler C, Goodman AL, Mandel MJ. 2014. Global discovery of colonization determinants in the squid symbiont Vibrio fischeri. Proc Natl Acad Sci USA 111:17284–17289. 10.1073/pnas.1415957111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Stabb EV, Ruby EG. 2002. RP4-based plasmids for conjugation between Escherichia coli and members of the Vibrionaceae. Methods Enzymol 358:413–426. 10.1016/s0076-6879(02)58106-4. [DOI] [PubMed] [Google Scholar]
- 39.Visick K, Hodge-Hanson K, Tischler A, Bennett A, Mastrodomenico V. 2018. Tools for rapid genetic engineering of Vibrio fischeri. Appl Environ Microbiol 84:e00850-18. 10.1128/AEM.00850-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Dunn AK, Millikan DS, Adin DM, Bose JL, Stabb EV. 2006. New rfp- and pES213-derived tools for analyzing symbiotic Vibrio fischeri reveal patterns of infection and lux expression in situ. Appl Environ Microbiol 72:802–810. 10.1128/AEM.72.1.802-810.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Supporting data will be made available upon reasonable request.