Abstract
Epidemic strains of Vibrio cholerae O1 are divided into two biotypes, classical and El Tor. In both biotypes, regulation of virulence gene expression depends on a cascade in which ToxR activates expression of ToxT, and ToxT activates expression of cholera toxin and other virulence genes. In the classical biotype, maximal expression of this ToxR regulon in vitro occurs at 30°C at pH 6.5 (ToxR-inducing conditions), whereas in the El Tor biotype, production of these virulence genes only occurs under very limited conditions and not in response to temperature and pH; this difference between biotypes is mediated at the level of toxT transcription. In the classical biotype, two other proteins, TcpP and TcpH, are needed for maximal toxT transcription. Transcription of tcpPH in the classical biotype is regulated by pH and temperature independently of ToxR or ToxT, suggesting that TcpP and TcpH couple environmental signals to transcription of toxT. In this study, we show a near absence of tcpPH message in the El Tor biotype under ToxR-inducing conditions of temperature and pH. However, once expressed, El Tor TcpP and TcpH appear to be as effective as classical TcpP and TcpH in activating toxT transcription. These results suggest that differences in regulation of virulence gene expression between the biotypes of V. cholerae primarily result from differences in expression of tcpPH message in response to environmental signals. We present an updated model for control of the ToxR virulence regulon in V. cholerae.
Vibrio cholerae, a curved gram-negative bacterium, causes a secretory diarrhea in humans that can produce severe dehydration and death (23). Strains of V. cholerae that cause epidemic cholera belong to serogroup O1. V. cholerae O1 is divided into two biotypes, classical and El Tor, which differ biochemically and phenotypically in such properties as susceptibility to polymyxin B, ability to hemolyze sheep erythrocytes, and the Voges-Proskauer reaction. Human infections with these two biotypes also differ clinically; infection with classical V. cholerae O1 more frequently produces severe infection than does El Tor, suggesting that the genetic and phenotypic differences between the two biotypes may also be reflected in their pathogenic potential. In 1992, a new serogroup of V. cholerae emerged as a cause of epidemic cholera in southeast Asia, V. cholerae O139 (40). Several lines of evidence suggest that this new serogroup is closely related to and derived from the El Tor biotype of V. cholerae O1 by substitution of genes encoding the O139 O antigen and acquisition of the ability to produce a capsule (1, 4, 7, 8, 48, 50).
The major virulence factor for all of these pathogenic strains of V. cholerae is cholera toxin, a heterodimeric protein exotoxin that consists of a single, enzymatically active A subunit noncovalently associated with five B subunits (13). The pentamer of B subunits binds the holotoxin to its receptor, the ganglioside GM1, on intestinal epithelial cells. Upon binding, the A subunit is nicked, reduced, and translocated intracellularly, where it acts as an ADP-ribosyl transferase, elevating cyclic AMP levels within the cell and causing a secretory diarrhea (6, 14, 15, 30). The genes for the A and B subunits of cholera toxin are contained in an operon (31) within a filamentous bacteriophage that lysogenizes V. cholerae, the CTX phage (51). In classical strains of V. cholerae, expression of cholera toxin in vitro is strongly regulated by environmental growth conditions, such as pH, temperature, and osmolarity (35, 36). How these in vitro growth conditions relate to the in vivo expression of cholera toxin is not yet certain.
A second major virulence factor of V. cholerae is the toxin-coregulated pilus (TCP), a colonization factor whose expression is regulated in classical strains by the same environmental growth conditions as those involved in the expression of cholera toxin (45). The major subunit of TCP is encoded by tcpA; several other genes involved in the biosynthesis, processing, and assembly of TcpA into the mature pilin structure are located downstream of tcpA on the V. cholerae chromosome (46). Recently, these and adjacent genes were found to be located on a V. cholerae pathogenicity island within another filamentous bacteriophage, designated VPIφ (24, 25).
Expression of cholera toxin and TCP is dependent on a regulatory gene in V. cholerae, toxR, which encodes a transmembrane protein with an amino-terminal, cytoplasmic DNA binding domain that acts as a transcriptional activator (34, 36). A second regulatory gene, toxS, is transcribed in an operon with toxR and encodes a periplasmic protein that facilitates dimerization and activation of ToxR as a DNA-binding protein (10, 33). ToxR is essential for activation of cholera toxin and TCP expression in V. cholerae. Although ToxR is sufficient to activate transcription of the ctxAB promoter in E. coli, it is not able to activate transcription from either the ctxAB or tcpA promoter directly in a V. cholerae background (2, 34). Instead, ToxR activates transcription of an additional regulatory gene, toxT, that encodes a transcriptional activator in the AraC family (12, 21, 37). ToxT then directly activates transcription at the tcpA and ctxAB promoters, as well as other genes that are part of the ToxR regulon. This suggests a regulatory cascade for expression of cholera virulence factors in which ToxR activates the gene coding for expression of ToxT, and ToxT in turn activates the genes coding for expression of cholera toxin and TCP, as well as other genes in the ToxR regulon (9).
The gene encoding ToxT is located in the tcp gene cluster downstream of tcpA, between tcpF and tcpJ (21, 26, 39). Transcription of toxT occurs both as part of the tcpA operon, as well as separately from a ToxR-dependent promoter located between tcpF and toxT (2, 20). There are several inverted repeats upstream of this toxT promoter in classical V. cholerae, and these repeats have been shown to be necessary for binding of ToxR and activation of toxT transcription. However, ToxR is not sufficient in Escherichia coli to activate transcription of an operon fusion between the classical toxT promoter and lacZ and indeed represses transcription at the toxT promoter (20). This suggests the possibility that an additional protein or proteins are necessary in classical V. cholerae for full activation at the toxT promoter.
Previous investigators have identified additional genes upstream of tcpA, including tcpP and tcpH, which are transcribed as an operon in the same direction as tcpA, and tcpI, which is transcribed divergently from tcpPH (38, 46). We and Häse and Mekalanos have previously shown that transcription of toxT in classical V. cholerae requires the gene products encoded by tcpPH (5, 18). While TcpP alone appears to be sufficient for toxT expression, activation is greatly enhanced when TcpH is coexpressed (18). The proposed membrane topology suggests that TcpP and TcpH may be localized to the cytoplasmic membrane, similar to ToxR and ToxS (18). Other studies indicate that classical TcpP and TcpH act in concert with ToxR and ToxS to activate toxT transcription, although the exact mechanism is not yet understood (27, 28).
Whereas expression of ToxR in classical V. cholerae is constitutive, transcription of toxT is regulated by the same environmental growth conditions that regulate expression of cholera toxin and TCP (9, 11, 21). We have previously shown that transcription of tcpPH in classical V. cholerae is regulated by pH and temperature, suggesting that the protein products, TcpP and TcpH, may couple environmental growth conditions to transcription of toxT and expression of the ToxR regulon in classical V. cholerae (5).
Expression of cholera toxin and TCP in El Tor strains of V. cholerae is regulated quite differently in response to environmental conditions from expression in classical strains of V. cholerae. In the El Tor biotype, production of cholera toxin and TCP occurs only under very limited in vitro growth conditions, referred to as AKI conditions (49), but not under the conditions of temperature and pH that induce expression of the ToxR regulon in classical V. cholerae. DiRita et al. (11) have previously shown that these differences between V. cholerae biotypes in expression of the ToxR regulon in response to environmental conditions are mediated at the level of toxT transcription and that overexpression of toxT from a heterologous promoter in an El Tor strain leads to expression of cholera toxin and TCP without the requirement for AKI conditions. We wished to examine whether the difference in transcription of toxT in response to environmental conditions between the two biotypes of V. cholerae reflects differences in expression of tcpPH message and/or the activity of the TcpP and TcpH proteins.
MATERIALS AND METHODS
Bacterial strains and media.
The bacterial strains and plasmids used in this study are shown in Table 1. Plasmid pUC19 was used as the vector for cloning PCR products, and pLAFR3 was used for complementation experiments. Plasmids were initially cloned in E. coli DH5α. After characterization, recombinant plasmids were electroporated into V. cholerae by using a Gene Pulser (Bio-Rad Laboratories, Richmond, Calif.) according to the manufacturer’s protocol, modified for electroporation into V. cholerae as previously described (16). The electroporation conditions were 2,500 V at 25-μF capacitance, producing time constants of 4.5 to 4.8 ms. To construct a polar mutation in the V. cholerae tcpP locus, a 230-bp internal fragment from tcpP was cloned into the suicide plasmid pGP704, resulting in plasmid pYM2-16. E. coli SM10λpir was used to mobilize plasmid pYM2-16 into V. cholerae O395 by conjugation, resulting in YM2-21. Proper integration of the suicide plasmid within tcpP was confirmed by Southern blotting. V. cholerae KSK401 was isolated following TnphoA mutagenesis of strain KSK218, by using a screen for reduced activity of the ctx::lacZ reporter activity in KSK218 (42). The TnphoA insertion in KSK401 was cloned by chromosomal capture and backcrossed to confirm that the phenotype was linked to the insertion; DNA sequencing was used to determine that TnphoA had inserted within tcpP. In order to obtain an E. coli reporter strain that could be used with clones of tcpPH in pLAFR3 (Tcr), bacteriophage P1 transduction was utilized to move the toxT::lacZ fusion, which is linked to kanamycin resistance in DH92, into E. coli MC4100 (Tcs recA+ lacZ), resulting in strain RT4146. Strains were stored in 15% glycerol at −70°C and inoculated onto Luria-Bertani (LB) agar medium prior to growth in LB liquid culture. (LB medium contained 10 g of tryptone, 5 g of yeast extract, and 5 g of NaCl per liter.) LB liquid cultures were grown on a tube roller at 30°C at pH 6.5 to induce ToxR-dependent gene expression and at 37°C at pH 8.4 to repress ToxR-dependent gene expression (35). Overnight cultures grown under ToxR-inducing conditions with moderate aeration were examined for autoagglutination as described previously (45); autoagglutination reflects expression of TCP. AKI conditions were as described previously (49). Ampicillin (25 or 100 μg/ml), streptomycin (100 μg/ml), kanamycin (45 μg/ml), tetracycline (5 or 15 μg/ml), or chloramphenicol (34 μg/ml) was added as appropriate.
TABLE 1.
Strains and plasmids used in this study
| Strain or plasmid | Relevant characteristicsa | Source or reference |
|---|---|---|
| Strains | ||
| V. cholerae | ||
| O395 | Classical strain; Smr | Laboratory collection |
| O395 toxR | O395 with mutation in toxR created by integration of suicide plasmid pVM55; Smr Apr | 52 |
| C6709 | El Tor strain; Smr | Laboratory collection |
| KTT7-1 | O395 tcpH; Smr | 5 |
| KSK218 | O395 ΔlacZ ctx::lacZ | 42 |
| KSK401 | KSK218 tcpP::TnphoA (polar on tcpH) | This study |
| YM2-21 | O395 with a polar mutation in tcpP created by integration of the suicide plasmid pYM2-16; Smr Apr | This study |
| E. coli | ||
| DH5α | F−endA1 hsdR17 supE44 thi-1 recA1 gyrA96 relA1 Δ(argF-lacZYA)U169 (Φ80d lacZΔM15) | 17 |
| SM10λpir | thi thr leu tonA lacY supE recA::RP4-2-Tc::Mu λpir R6K | 35 |
| DH92 | TE2680[Kmr-ptoxT::lacZ] | 20 |
| RT4146 | MC4100[Kmr-ptoxT::lacZ], by P1 transduction from DH92 | This study |
| CC118 | Δ(ara leu)7697 Δ(lac)X74 araD139 phoA20 galE galK thi rpsE rpoB argE recA1 | 29 |
| Plasmids | ||
| pLAFR3 | Cloning vector; Tcr | 44 |
| pACYC184 | Cloning vector; Cmr Tcr | Laboratory collection |
| pUC19 | Cloning vector; Apr | Gibco-BRL |
| pGP704 | Suicide vector; Apr | 35 |
| pPAC22 | PCR product of the wild-type tcpPH genes from V. cholerae O395, with BamHI ends, cloned into pUC19 for sequencing and subsequent constructions; Apr | 5 |
| pYM49 | BamHI fragment from pPAC22, cloned into pLAFR3 in same orientation to the lacZ promoter; Tcr | This study |
| pYM53 | PCR product of tcpPH genes from V. cholerae C6709, cloned into pUC19 for sequencing and subsequent constructions; Apr | This study |
| pYM59 | BamHI fragment from pYM53 containing tcpPH genes from C6709, cloned into pLAFR3 in same orientation to lacZ promoter; Tcr | This study |
| pYM2-16 | 230-bp tcpP PCR product from V. cholerae O395 cloned into pGP704; Apr | This study |
| pTSK | pACYC184[ptet::toxRS]; Cmr | 2 |
| pTSS-5 | pACYC184[ptet::toxT]; Cmr | 2 |
| pRAB | pRZ5202 [tcpAB::lacZ]; Apr | 2 |
| pRX | pRZ5202[ctx::lacZ]; Apr | 2 |
| pVM55 | Suicide plasmid pJM703.1, containing a 630-bp EcoRI-HpaI internal fragment of toxR; Apr | 35 |
Sm, streptomycin; Ap, ampicillin; Km, kanamycin; Tc, tetracycline; Cm, chloramphenicol.
Genetic methods.
Isolation of plasmid and bacterial chromosomal DNA, restriction enzyme digests, agarose gel electrophoresis, and Southern hybridization analysis were performed according to standard molecular biologic techniques (41). DNA restriction endonucleases, T4 DNA ligase, T4 DNA polynucleotide kinase, Klenow fragment, and shrimp alkaline phosphatase were used according to the manufacturers’ specifications. DNA fragments used as probes for Southern and Northern hybridizations were generated by PCR as previously described (5), separated on 1% agarose gels, cut out from the gel under UV illumination, and recovered with a Compass kit (American Bioanalytical, Natick, Mass.). Purified DNA fragments were radiolabelled with [α-32P]dCTP (Dupont Biotechnology Systems, NEN Research Products, Boston, Mass.) by using a random-priming labelling kit (Prime-It II; Stratagene, La Jolla, Calif.). GeneScreen and GeneScreen Plus hybridization transfer membranes (DuPont Biotechnology Systems) were used according to the manufacturer’s protocols for Northern and Southern hybridizations, respectively. DNA sequencing was performed at the Massachusetts General Hospital, Department of Molecular Biology, DNA Sequencing Core Facility, by using ABI Prism DiTerminator Cycle sequencing with AmpliTaq DNA polymerase FS and an AB1377 DNA sequencer (Perkin-Elmer Applied Biosystems Division, Foster City, Calif.).
Bacterial RNA was isolated by using CATRIMOX-14 (Qiagen, Valencia, Calif.). Five milliliters of mid-log-phase bacterial cultures grown under appropriate growth conditions was quickly added to 5 ml of crushed ice solution A (75 ml of TM buffer [see below], 1.7 ml of 1 M sodium azide) and centrifuged at 4°C. The bacterial pellet was suspended in 400 μl of TM buffer (10 mM Tris HCl [pH 7.2], 5 mM MgCl2), 4 μl of DNase I (RNase-free; Boehringer-Mannheim, Indianapolis, Ind.), and 100 μl of lysozyme (4-mg/ml final concentration). The suspension was frozen in a dry ice-ethanol bath and thawed at 37°C four times. After the fourth thaw, the suspension was frozen again, and 67 μl of 20 mM acetic acid was added to the frozen cells. The cells were then thawed at 37°C, and 500 μl of CATRIMOX-14 was added immediately upon thawing. The mixture was centrifuged for 1 min at 6,500 rpm in an Eppendorf microcentrifuge. The pellet was resuspended in 1 ml of 2 M LiCl2 and incubated at room temperature for 5 min. The suspension was then spun at top speed in a microcentrifuge for 6 min at 4°C. The pellet was again resuspended in 1 ml of LiCl2, incubated, and centrifuged as described above. The pellet was air dried and resuspended in 100 μl of RNase-free water overnight at 4°C before the RNA concentration was measured. Ten micrograms of purified RNA was denatured with glyoxal for Northern blot analysis as previously described (3).
Assays.
Cholera toxin production was assayed in culture supernatants with a GM1 enzyme-linked immunosorbent assay as described previously (22). β-Galactosidose activity was determined as described previously (32). Induction of plasmid-encoded V. cholerae tcpPH under the control of the lacZ promoter in E. coli was achieved with 5 mM isopropyl-β-d-thiogalactopyranoside (IPTG; Sigma, St. Louis, Mo.); since V. cholerae does not contain lacI, the same plasmid constructs gave constitutive expression of tcpPH in V. cholerae.
RESULTS
Transcription of tcpPH in response to pH and temperature differs between classical and El Tor strains of V. cholerae.
We have previously shown that expression of tcpPH message is regulated by temperature and pH, independently of ToxR or ToxT, in classical V. cholerae (5). To determine if expression of tcpPH message could account for the differential expression of virulence factors between the two biotypes, Northern blot hybridization was utilized. When grown under ToxR-inducing conditions, classical strain O395 produced substantial amounts of tcpPH message, while El Tor strain C6709 produced significantly less (Fig. 1). Both strains produced similar levels of transcript when grown under AKI conditions (data not shown), although the levels were reduced compared to that of O395 grown under ToxR-inducing conditions. Neither strain produced significant amounts of tcpPH transcript when grown under ToxR-repressing conditions. The tcpPH mutant strain, YM2-21, did not produce any message when grown under either ToxR-inducing or ToxR-repressing conditions. Cholera toxin production mirrored tcpPH expression. Transcripts of tcpA and toxT message were absent from C6709 grown under ToxR-inducing conditions, the latter confirming observations reported by DiRita et al. (11). Neither O395 nor C6709 transcribed toxT nor tcpA under ToxR-repressing conditions.
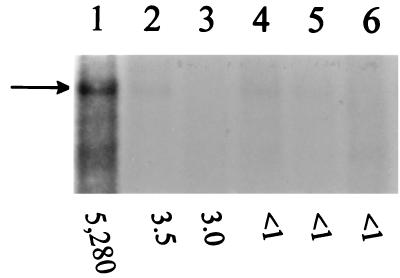
FIG. 1.
Northern blot analysis of 10 μg of total RNA from O395 (lanes 1 and 4), C6709 (lanes 2 and 5), and YM2-21 (lanes 3 and 6) grown under ToxR-inducing (lanes 1 to 3) and ToxR-repressing (lanes 4 to 6) conditions. A 458-bp fragment containing tcpPH was used as a probe. Arrow indicates 1.28 kbp of message. The values at the bottom indicate corresponding cholera toxin measurements (nanograms per milliliter at OD600).
Plasmids expressing either classical or El Tor TcpP and TcpH can complement a classical tcpPH mutant.
To determine if plasmid clones of tcpP and tcpH from each biotype were functional, we examined the ability of these constructs to complement a classical V. cholerae tcpPH mutant. The classical V. cholerae strain KSK218 contains a ctx::lacZ operon fusion in both chromosomal copies of the cholera toxin operon in classical strain O395. Inactivation of tcpP and tcpH, by TnphoA insertion within tcpP to produce strain KSK401, reduced expression at the ctx promoter, as reflected by β-galactosidase activities (Table 2). The TcpP and TcpH proteins expressed from pYM49 (classical) and pYM59 (El Tor) were each able to restore ctxAB transcription, as well as autoagglutination (a rough measure of TCP expression), in KSK401. This confirms that both clones are functional and suggests that there is little difference between the biotypes with respect to the activity of the TcpP and TcpH proteins when they are present in the classical biotype background.
TABLE 2.
Measurements of β-galactosidase activity and autoagglutination for a classical biotype ctx::lacZ V. cholerae strain grown under ToxR-inducing conditions
| Strain | β-Galactosidase activity (Miller units/OD600)a | Autoagglutination |
|---|---|---|
| KSK218 | 1,750 (1,635–1,943) | + |
| KSK401 | 97 (95–98) | − |
| KSK401(pLAFR3) | 143 (135–151) | − |
| KSK401(pYM49) | 4,848 (4,406–5,433) | + |
| KSK401(pYM59) | 3,412 (3,259–3,670) | + |
The mean and range (in parentheses) of at least three independent experiments are shown.
Differences in environmental regulation of the ToxR regulon between classical and El Tor V. cholerae are overcome by overexpressing TcpP and TcpH.
Although classical strain O395 showed a more than 4,000-fold increase in expression of cholera toxin in response to ToxR-inducing conditions, El Tor strain C6709 showed no increase (Table 3); however, both strains were able to express comparable amounts of cholera toxin in response to AKI conditions (not shown). Overexpression of TcpP and TcpH in the El Tor strain resulted in a marked increase in cholera toxin expression; plasmid pYM59, which overexpresses the El Tor tcpP and tcpH genes from the lacZ promoter, stimulated strain C6709 to produce levels of cholera toxin under ToxR-inducing conditions comparable to those of wild-type O395. Overexpression of tcpPH from this constitutive promoter resulted in expression of cholera toxin even under ToxR-repressing conditions. However, a small amount of residual regulation of expression in response to temperature and pH was still observed even when the tcpPH genes were expressed from the lacZ promoter.
TABLE 3.
Measurements of cholera toxin in culture supernatants following overnight growth under the indicated conditions
| Strain | Amt of cholera toxin (ng/ml/OD600) undera:
|
|
|---|---|---|
| ToxR-inducing conditions | ToxR-repressing conditions | |
| O395 | 4,464 (2,966–6,041) | <1 |
| C6709 | <1 | <1 |
| C6709(pLAFR3) | <1 | <1 |
| C6709(pYM59) | 4,030 (2,512–6,478) | 794 (493–1,041) |
The mean and range (in parentheses) of at least three independent experiments are shown.
TcpP and TcpH from both the classical and El Tor biotypes work synergistically with ToxR and ToxS to activate toxT transcription in E. coli.
To better understand the interaction of TcpP and TcpH with the ToxR regulon, we examined the ability of these proteins from either biotype to activate the ctx, tcpA, and toxT promoters independently of other V. cholerae proteins. Plasmids containing the appropriate regulatory genes, under the control of constitutive or inducible promoters, were tested for their ability to activate classical V. cholerae promoter fusions in E. coli (Table 4). We used a chromosomal toxT::lacZ fusion, because a plasmid-encoded fusion produced levels of β-galactosidase too high for accurate measurement. As shown previously (12), ToxRS and ToxT both activated the ctxAB promoter, while only ToxT activated the tcpA promoter; ToxRS repressed the toxT promoter approximately twofold. TcpPH from both biotypes had no effect on transcription from the ctxAB or tcpA promoters, but both activated transcription of toxT approximately twofold. The combination of ToxRS and TcpPH (from either biotype) activated transcription from the toxT promoter approximately 10-fold. This had previously been suggested to be the case for classical V. cholerae in preliminary data by Krukonis et al. (27).
TABLE 4.
Ability of various V. cholerae regulatory proteins to activate the ctx, tcpA, and toxT promoters fused to β-galactosidase in E. coli
| Reporter fusion | β-Galactosidase activity (Miller units/OD600) with regulatory protein expressiona
|
|||||||
|---|---|---|---|---|---|---|---|---|
| pACYC184 (control) | pLAFR3 (control) | pTSK (ToxRS) | pTSS-5 (ToxT) | pYM49 (classical TcpPH) | pYM59 (El Tor TcpPH) | pTSK + pYM49 | pTSK + pYM59 | |
| CC118(pRX) (ctx-lacZ) | 5 (4–6) | 5 (5) | 380 (360–391) | 81 (70–98) | 4 (4) | 5 (4–5) | 382 (373–389) | 380 (366–397) |
| CC118(pRAB) (tcpAB-lacZ) | 7 (6–8) | 6 (6–7) | 5 (5–6) | 342 (203–518) | 6 (6) | 6 (6) | 7 (7–8) | 7 (7) |
| RT4146 (toxT-lacZ) | 327 (314–345) | 330 (322–337) | 116 (108–128) | 247 (242–254) | 565 (553–574) | 692 (661–726) | 2,935 (2,839–3,071) | 3,554 (3,476–3,687) |
Means and ranges (in parentheses) of triplicate measurements of β-galactosidase activities are shown. Expression of TcpPH from plasmids pYM49 and pYM59 was induced with IPTG.
DISCUSSION
Production of cholera toxin and TCP is regulated differently between the two biotypes of V. cholerae O1. Classical strains of V. cholerae express these virulence factors under both ToxR-inducing conditions and AKI conditions, whereas El Tor strains of V. cholerae O1 express detectable levels only under AKI conditions in vitro. ToxR is required in both biotypes for expression of these virulence factors, although the expression of ToxR itself is independent of environmental growth conditions (9, 11). ToxT is also required for expression of these virulence factors in both biotypes, and expression of toxT in response to environmental conditions differs between the two biotypes (11). ToxT is expressed in classical strains of V. cholerae under both ToxR-inducing and AKI conditions, whereas El Tor strains of V. cholerae express ToxT only under AKI conditions. Expression of ToxT from an inducible promoter in an El Tor strain of V. cholerae allows this biotype to make cholera toxin and TCP without the requirement for AKI conditions (11). This suggests that the biotype-specific expression of cholera virulence factors is the result of differential expression of toxT in response to environmental growth conditions. DiRita et al. (11) suggested that another factor may be involved in activating toxT in V. cholerae and that this factor may be active only under AKI conditions in El Tor strains, but in a broader range of growth conditions in classical strains of V. cholerae. They also suggested that the resultant differential control of toxin production between the two biotypes may contribute to differences in their pathogenicity.
We have previously shown that transcription of toxT in classical V. cholerae requires TcpH (5). Transcription of the tcpPH operon is regulated by pH and temperature in classical strains, suggesting that TcpP and TcpH may couple the environmental signals of temperature and pH to transcription of toxT and expression of the ToxR regulon in classical V. cholerae. In classical V. cholerae, both TcpP and TcpH, in coordination with ToxR and ToxS, are required for optimal transcription of the toxT promoter (18, 27, 28). While the role of TcpP and TcpH in activation of toxT transcription has been established in classical V. cholerae, their role in the regulation of virulence gene expression in El Tor strains of V. cholerae had not been explored. Thomas et al. (47) and Ogierman et al. (38) have compared sequences of the tcp gene clusters of classical and El Tor V. cholerae and found that the major differences upstream of tcpA fall in the intergenic regions between tcpI and tcpP and between tcpH and tcpA, suggesting that differences in expression of tcpPH and/or tcpA message may underlie the differences in expression of cholera virulence factors between the two biotypes.
The data in the present study demonstrate that expression of tcpPH between the two biotypes of V. cholerae is fundamentally different in response to environmental signals of temperature and pH. In the classical strain, expression of tcpPH message is strongly induced by conditions of temperature and pH known to induce expression of the ToxR regulon, while similar conditions do not result in substantial expression of tcpPH message in the El Tor strain. toxT expression and tcpA expression were similarly divergent between the two biotypes, confirming previously reported results (11). These data suggest that a fundamental difference between the biotypes lies in the environmental regulation of tcpPH expression. Furthermore, given the role of TcpP and TcpH in expression of toxT in classical V. cholerae (5, 18), the current findings likely explain the previously reported biotype-specific expression of toxT (11).
TcpP and TcpH from both the classical and El Tor biotypes were able to activate expression of the ToxR regulon in several host backgrounds. The functionality of the El Tor TcpP and TcpH was established by demonstrating the ability of plasmid pYM59, encoding the El Tor TcpP and TcpH, to restore activation of a ctx::lacZ fusion in a tcpPH mutant of classical V. cholerae, strain KSK401. Levels of β-galactosidase were comparable to those seen with expression of classical TcpP and TcpH from plasmid pYM49. Similarly, overexpression of the El Tor tcpPH gene products in the El Tor V. cholerae background resulted in increased cholera toxin production, comparable to that of the classical biotype. Constitutive expression of the tcpPH gene products from the lacZ promoter in the El Tor background resulted in reduced regulation of cholera toxin expression in response to environmental conditions; however, some residual regulation remained. This may be the result of combined expression of TcpPH from both the tcpPH and lacZ promoters, but it may also suggest the involvement of an additional factor or factors in environmental regulation.
In E. coli, plasmids encoding either the classical or El Tor TcpP and TcpH under the control of an inducible promoter were able to activate a classical V. cholerae toxT::lacZ fusion. In both cases, the activity was greatly enhanced by the presence of ToxR and ToxS; thus, El Tor TcpP and TcpH appear to act in concert with ToxR and ToxS to amplify toxT expression, as was previously suggested for classical V. cholerae (27).
Although there are a small number of amino acid differences between the El Tor and classical TcpP and TcpH proteins, the fact that the El Tor proteins were able to activate transcription of toxT and expression of the ToxR regulon in either biotype and in E. coli indicates little difference in the functionality of these proteins compared to that of the classical TcpP and TcpH proteins. Recently, Häse and Mekalanos showed that the activity of classical TcpP and TcpH is influenced by Na+ concentration or osmolarity (19). While our results indicate that the effects of pH and temperature on TcpP and TcpH activity do not differ between the two biotypes, we did not examine whether the effects of Na+ concentration or osmolarity on TcpPH activity differ between the two biotypes.
The near absence of tcpPH message in El Tor V. cholerae strains when grown under ToxR-inducing conditions suggests (i) that there are differences in the tcpPH promoters between the two biotypes that result in altered environmental regulation, (ii) that a trans-acting factor(s) that is present in classical strains but absent or altered in El Tor strains is the cause of the observed differential expression in response to temperature and pH, or (iii) that there are differences in tcpPH transcript stability between the two biotypes. Experiments are currently under way to investigate these hypotheses.
Recently, a protein designated AphA was found to be required for activation of tcpPH expression in classical V. cholerae, although the mechanism of activation is not known (43). Expression of aphA does not appear to be influenced by environmental conditions that activate tcpPH expression in classical V. cholerae. It is not yet known whether AphA plays a role in the differential expression of tcpPH in the classical and El Tor biotypes.
The results presented here, in addition to previously reported results (5, 18, 27), are consistent with a model in which TcpP and TcpH, in conjunction with ToxR and ToxS, activate transcription of toxT in classical and El Tor strains of V. cholerae, with subsequent expression of the ToxR regulon. An updated model for regulation of virulence gene expression in classical V. cholerae is depicted in Fig. 2.
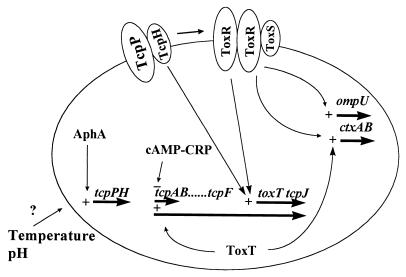
FIG. 2.
Model for the regulation of the ToxR regulon in classical V. cholerae, incorporating previous observations as well as those in the present study (see text for discussion). +, transcriptional activation; −, repression. Transcription is represented by thick solid arrows. Additional factors regulating tcpPH transcription (indicated by ?) are currently under investigation. cAMP-CRP, cyclic AMP (cAMP)-cyclic AMP receptor protein.
ACKNOWLEDGMENTS
This work was supported by grants from the National Institute of Allergy and Infectious Diseases RO1AI34968 (to S.B.C.), RO1AI39654 (to R.K.T.), and R29AI41558 (to K.A.S.).
We thank Claudia Häse and John J. Mekalanos for helpful discussions and Jennifer Y. Thibert for constructing strains and performing assays.
REFERENCES
- 1.Bik E M, Bunschoten A E, Willems R J, Chang A C, Mooi F R. Genetic organization and functional analysis of the otn DNA essential for cell-wall polysaccharide synthesis in Vibrio cholerae O139. Mol Microbiol. 1996;20:799–811. doi: 10.1111/j.1365-2958.1996.tb02518.x. [DOI] [PubMed] [Google Scholar]
- 2.Brown R C, Taylor R K. Organization of tcp, acf, and toxT genes within a ToxT-dependent operon. Mol Microbiol. 1995;16:425–439. doi: 10.1111/j.1365-2958.1995.tb02408.x. [DOI] [PubMed] [Google Scholar]
- 3.Burnett W V. Northern blotting of RNA denatured in glyoxal without buffer recirculation. BioTechniques. 1997;22:668–671. doi: 10.2144/97224st01. [DOI] [PubMed] [Google Scholar]
- 4.Calia K E, Murtagh M, Ferraro M J, Calderwood S B. Comparison of Vibrio cholerae O139 with V. cholerae O1 classical and El Tor biotypes. Infect Immun. 1994;62:1504–1506. doi: 10.1128/iai.62.4.1504-1506.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Carroll P A, Tashima K T, Rogers M B, DiRita V J, Calderwood S B. Phase variation in tcpH modulates expression of the ToxR regulon in Vibrio cholerae. Mol Microbiol. 1997;25:1099–1111. doi: 10.1046/j.1365-2958.1997.5371901.x. [DOI] [PubMed] [Google Scholar]
- 6.Cassel D, Pfeuffer T. Mechanism of cholera toxin action: covalent modification of the guanyl nucleotide-binding protein of the adenylate cyclase system. Proc Natl Acad Sci USA. 1978;75:2669–2673. doi: 10.1073/pnas.75.6.2669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Comstock L E, Johnson J A, Michalski J M, Morris J G, Jr, Kaper J B. Cloning and sequence of a region encoding a surface polysaccharide of Vibrio cholerae O139 and characterization of the insertion site in the chromosome of Vibrio cholerae O1. Mol Microbiol. 1996;19:815–826. doi: 10.1046/j.1365-2958.1996.407928.x. [DOI] [PubMed] [Google Scholar]
- 8.Comstock L E, Maneval D, Jr, Panigrahi P, Joseph A, Levine M M, Kaper J B, Morris J G, Jr, Johnson J A. The capsule and O antigen in Vibrio cholerae O139 Bengal are associated with a genetic region not present in Vibrio cholerae O1. Infect Immun. 1995;63:317–323. doi: 10.1128/iai.63.1.317-323.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.DiRita V J. Co-ordinate expression of virulence genes by ToxR in Vibrio cholerae. Mol Microbiol. 1992;6:451–458. doi: 10.1111/j.1365-2958.1992.tb01489.x. [DOI] [PubMed] [Google Scholar]
- 10.DiRita V J, Mekalanos J J. Periplasmic interaction between two membrane regulatory proteins, ToxR and ToxS, results in signal transduction and transcriptional activation. Cell. 1991;64:29–37. doi: 10.1016/0092-8674(91)90206-e. [DOI] [PubMed] [Google Scholar]
- 11.DiRita V J, Neely M, Taylor R K, Bruss P M. Differential expression of the ToxR regulon in classical and El Tor biotypes of Vibrio cholerae is due to biotype-specific control over toxT expression. Proc Natl Acad Sci USA. 1996;93:7991–7995. doi: 10.1073/pnas.93.15.7991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.DiRita V J, Parsot C, Jander G, Mekalanos J J. Regulatory cascade controls virulence in Vibrio cholerae. Proc Natl Acad Sci USA. 1991;88:5403–5407. doi: 10.1073/pnas.88.12.5403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gill D M. The arrangement of subunits in cholera toxin. Biochemistry. 1976;15:1242–1248. doi: 10.1021/bi00651a011. [DOI] [PubMed] [Google Scholar]
- 14.Gill D M, King C A. The mechanism of action of cholera toxin in pigeon erythrocyte lysates. J Biol Chem. 1975;250:6424–6432. [PubMed] [Google Scholar]
- 15.Gill D M, Meren R. ADP-ribosylation of membrane proteins catalyzed by cholera toxin: basis of the activation of adenylate cyclase. Proc Natl Acad Sci USA. 1978;75:3050–3054. doi: 10.1073/pnas.75.7.3050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Goldberg M B, Boyko S A, Calderwood S B. Positive transcriptional regulation of an iron-regulated virulence gene in Vibrio cholerae. Proc Natl Acad Sci USA. 1991;88:1125–1129. doi: 10.1073/pnas.88.4.1125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983;166:557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- 18.Hase C C, Mekalanos J J. TcpP protein is a positive regulator of virulence gene expression in Vibrio cholerae. Proc Natl Acad Sci USA. 1998;95:730–734. doi: 10.1073/pnas.95.2.730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hase C C, Mekalanos J J. Effects of changes in membrane sodium flux on virulence gene expression in Vibrio cholerae. Proc Natl Acad Sci USA. 1999;96:3183–3187. doi: 10.1073/pnas.96.6.3183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Higgins D E, DiRita V J. Transcriptional control of toxT, a regulatory gene in the ToxR regulon of Vibrio cholerae. Mol Microbiol. 1994;14:17–29. doi: 10.1111/j.1365-2958.1994.tb01263.x. [DOI] [PubMed] [Google Scholar]
- 21.Higgins D E, Nazareno E, DiRita V J. The virulence gene activator ToxT from Vibrio cholerae is a member of the AraC family of transcriptional activators. J Bacteriol. 1992;174:6974–6980. doi: 10.1128/jb.174.21.6974-6980.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Holmgren J. Comparison of the tissue receptors for Vibrio cholerae and Escherichia coli enterotoxins by means of gangliosides and natural cholera toxoid. Infect Immun. 1973;8:851–859. doi: 10.1128/iai.8.6.851-859.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kaper J B, Morris J G, Jr, Levine M M. Cholera. Clin Microbiol Rev. 1995;8:48–86. doi: 10.1128/cmr.8.1.48. . (Erratum, 8:316.) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Karaolis D K, Johnson J A, Bailey C C, Boedeker E C, Kaper J B, Reeves P R. A Vibrio cholerae pathogenicity island associated with epidemic and pandemic strains. Proc Natl Acad Sci USA. 1998;95:3134–3139. doi: 10.1073/pnas.95.6.3134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Karaolis D K, Somara S, Maneval D R, Jr, Johnson D A, Kaper J B. A bacteriophage encoding a pathogenicity island, a type-IV pilus and a phage receptor in cholera bacteria. Nature. 1999;399:375–379. doi: 10.1038/20715. [DOI] [PubMed] [Google Scholar]
- 26.Kaufman M R, Shaw C E, Jones I D, Taylor R K. Biogenesis and regulation of the Vibrio cholerae toxin-coregulated pilus: analogies to other virulence factor secretory systems. Gene. 1993;126:43–49. doi: 10.1016/0378-1119(93)90588-t. [DOI] [PubMed] [Google Scholar]
- 27.Krukonis E S, Crawford J A, DiRita V J. Abstracts of the 98th General Meeting of the American Society for Microbiology 1998. Washington, D.C: American Society for Microbiology; 1998. Vibrio cholerae ToxR/ToxS and TcpP/TcpH cooperatively activate transcription from the toxT promoter, abstr. B-114; p. 74. [Google Scholar]
- 28.Krukonis E S, DiRita V J. Abstracts of the 99th General Meeting of the American Society for Microbiology 1999. Washington, D.C: American Society for Microbiology; 1999. Analysis of the mechanism of Vibrio cholerae ToxT activation by TcpP and the role of ToxR, abstr. B/D-223; p. 73. [Google Scholar]
- 29.Manoil C, Beckwith J. TnphoA: a transposon probe for protein export signal. Proc Natl Acad Sci USA. 1985;82:8129–8133. doi: 10.1073/pnas.82.23.8129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mekalanos J J, Collier R J, Romig W R. Enzymic activity of cholera toxin. II. Relationships to proteolytic processing, disulfide bond reduction, and subunit composition. J Biol Chem. 1979;254:5855–5861. [PubMed] [Google Scholar]
- 31.Mekalanos J J, Swartz D J, Pearson G D, Harford N, Groyne F, de Wilde M. Cholera toxin genes: nucleotide sequence, deletion analysis and vaccine development. Nature. 1983;306:551–557. doi: 10.1038/306551a0. [DOI] [PubMed] [Google Scholar]
- 32.Miller J H. A short course in bacterial genetics. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1992. [Google Scholar]
- 33.Miller V L, DiRita V J, Mekalanos J J. Identification of toxS, a regulatory gene whose product enhances ToxR-mediated activation of the cholera toxin promoter. J Bacteriol. 1989;171:1288–1293. doi: 10.1128/jb.171.3.1288-1293.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Miller V L, Mekalanos J J. Synthesis of cholera toxin is positively regulated at the transcriptional level by toxR. Proc Natl Acad Sci USA. 1984;81:3471–3475. doi: 10.1073/pnas.81.11.3471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Miller V L, Mekalanos J J. A novel suicide vector and its use in construction of insertion mutations: osmoregulation of outer membrane proteins and virulence determinants in Vibrio cholerae requires toxR. J Bacteriol. 1988;170:2575–2583. doi: 10.1128/jb.170.6.2575-2583.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Miller V L, Taylor R K, Mekalanos J J. Cholera toxin transcriptional activator ToxR is a transmembrane DNA binding protein. Cell. 1987;48:271–279. doi: 10.1016/0092-8674(87)90430-2. [DOI] [PubMed] [Google Scholar]
- 37.Ogierman M A, Manning P A. Homology of TcpN, a putative regulatory protein of Vibrio cholerae, to the AraC family of transcriptional activators. Gene. 1992;116:93–97. doi: 10.1016/0378-1119(92)90634-2. [DOI] [PubMed] [Google Scholar]
- 38.Ogierman M A, Voss E, Meaney C, Faast R, Attridge S R, Manning P A. Comparison of the promoter proximal regions of the toxin-co-regulated tcp gene cluster in classical and El Tor strains of Vibrio cholerae O1. Gene. 1996;170:9–16. doi: 10.1016/0378-1119(95)00744-x. [DOI] [PubMed] [Google Scholar]
- 39.Ogierman M A, Zabihi S, Mourtzios L, Manning P A. Genetic organization and sequence of the promoter-distal region of the tcp gene cluster of Vibrio cholerae. Gene. 1993;126:51–60. doi: 10.1016/0378-1119(93)90589-u. [DOI] [PubMed] [Google Scholar]
- 40.Ramamurthy T, Garg S, Sharma R, Bhattacharya S K, Nair G B, Shimada T, Takeda T, Karasawa T, Kurazano H, Pal A, et al. Emergence of novel strain of Vibrio cholerae with epidemic potential in southern and eastern India. Lancet. 1993;341:703–704. doi: 10.1016/0140-6736(93)90480-5. . (Letter.) [DOI] [PubMed] [Google Scholar]
- 41.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 42.Skorupski K, Taylor R K. Cyclic AMP and its receptor protein negatively regulate the coordinate expression of cholera toxin and toxin-coregulated pilus in Vibrio cholerae. Proc Natl Acad Sci USA. 1997;94:265–270. doi: 10.1073/pnas.94.1.265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Skorupski K, Taylor R K. A new level in the Vibrio cholerae ToxR virulence cascade: AphA is required for transcriptional activation of the tcpPH operon. Mol Microbiol. 1999;31:763–771. doi: 10.1046/j.1365-2958.1999.01215.x. [DOI] [PubMed] [Google Scholar]
- 44.Staskawicz B, Dahlbeck D, Keen N, Napoli C. Molecular characterization of cloned avirulence genes from race 0 and race 1 of Pseudomonas syringae pv. glycinea. J Bacteriol. 1987;169:5789–5794. doi: 10.1128/jb.169.12.5789-5794.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Taylor R K, Miller V L, Furlong D B, Mekalanos J J. Use of phoA gene fusions to identify a pilus colonization factor coordinately regulated with cholera toxin. Proc Natl Acad Sci USA. 1987;84:2833–2837. doi: 10.1073/pnas.84.9.2833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Taylor R K, Shaw C E, Peterson K M, Spears P, Mekalanos J J. Safe, live Vibrio cholerae vaccines? Vaccine. 1988;6:151–154. doi: 10.1016/s0264-410x(88)80019-7. [DOI] [PubMed] [Google Scholar]
- 47.Thomas S, Williams S G, Manning P A. Regulation of tcp genes in classical and El Tor strains of Vibrio cholerae O1. Gene. 1995;166:43–48. doi: 10.1016/0378-1119(95)00610-x. [DOI] [PubMed] [Google Scholar]
- 48.Waldor M K, Colwell R, Mekalanos J J. The Vibrio cholerae O139 serogroup antigen includes an O-antigen capsule and lipopolysaccharide virulence determinants. Proc Natl Acad Sci USA. 1994;91:11388–11392. doi: 10.1073/pnas.91.24.11388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Waldor M K, Mekalanos J J. ToxR regulates virulence gene expression in non-O1 strains of Vibrio cholerae that cause epidemic cholera. Infect Immun. 1994;62:72–78. doi: 10.1128/iai.62.1.72-78.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Waldor M K, Mekalanos J J. Emergence of a new cholera pandemic: molecular analysis of virulence determinants in Vibrio cholerae O139 and development of a live vaccine prototype. J Infect Dis. 1994;170:278–283. doi: 10.1093/infdis/170.2.278. [DOI] [PubMed] [Google Scholar]
- 51.Waldor M K, Mekalanos J J. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science. 1996;272:1910–1914. doi: 10.1126/science.272.5270.1910. [DOI] [PubMed] [Google Scholar]
- 52.Wong S M, Carroll P A, Rahme L G, Ausubel F M, Calderwood S B. Modulation of expression of the ToxR regulon in Vibrio cholerae by a member of the two-component family of response regulators. Infect Immun. 1998;66:5854–5861. doi: 10.1128/iai.66.12.5854-5861.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]