Abstract
The performance of gargling for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reverse transcriptase (RT)-PCR testing has not been previously reviewed. This review systematically assessed the performance of saline and water gargling for SARS-CoV-2 RT-PCR testing in the settings of diagnosing and monitoring viral shedding.
We included original studies comparing the performance of gargling and (oropharyngeal–)nasopharyngeal swabs for SARS-CoV-2 RT-PCR testing. Studies conducted in either suspected individuals or confirmed cases were included and analysed separately. The sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) were examined using random-effects models.
Gargles achieved a high overall sensitivity (91%), specificity (97%), PPV (95%) and NPV (91%) for SARS-CoV-2 RT-PCR testing. Studies using saline gargle and water gargle have an overall sensitivity of 97% and 86%, respectively. The sensitivity values were largely maintained for saline and water gargling on stratified analysis, for both diagnosis (96% and 92%) and viral shedding monitoring (98% and 78%). A higher sensitivity was also reported by studies using sterile saline (100%), a smaller amount of gargling solution (92% versus 87%) and a longer gargling duration (95% versus 86%).
Our results supported the use of gargling as a sampling approach for SARS-CoV-2 RT-PCR testing, which achieved a high sensitivity for both diagnosis and viral shedding monitoring purposes. Further investigation on the comparative performance of different gargling mediums is needed to draw a definitive conclusion.
Short abstract
Gargling is a reliable sampling approach for SARS-CoV-2 RT-PCR test. Results suggest potential differential performance in factors including the nature, sterility and volume of gargling medium, and duration of gargling, which need further exploration. https://bit.ly/3tYau81
Introduction
Since its first emergence in late 2019, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has spread globally with the cumulated number of confirmed cases growing to more than 304 million (as of 9 January 2022) [1, 2]. The continuous reporting of new cases and the emergence of new variants of SARS-CoV-2 infections in many countries, including those already achieving a high level of vaccine uptake [1], highlighted the persistent importance of accurate and scalable diagnostic testing capacity in tackling the evolving pandemic [3]. For laboratory identification of SARS-CoV-2 in clinical diagnosis and monitoring of infection, reverse transcriptase (RT)-PCR on nasopharyngeal swab (NPS) is generally regarded as the gold-standard approach [4, 5]. In recent years, combined oropharyngeal–nasopharyngeal swab (ONPS) has increasingly been adopted as the reference gold standard because of its regarded higher sensitivity and diagnostic yield [5–10]. However, important drawbacks of these technically difficult sampling approaches, including procedural discomfort, expertise and manpower implications [11], and the requirement for high-level personal protective equipment and negative-pressure settings [12], have hindered their widespread use in ambulatory care settings.
Aiming to overcome these drawbacks and to improve the scalability of SARS-CoV-2 testing in different community settings under heightened testing demand [13], a number of alternative sampling approaches have been explored, including pooled nasal and throat swabs [14–16], saliva [17, 18], nasal swabs [19–24], oropharyngeal swabs (throat swabs) [25–27], and saline- or water-based gargle [28, 29]. In a recent systematic review, pooled nasal and throat swabs were reported to have the best testing performance, with a high sensitivity, specificity, and both positive predictive values (PPVs) and negative predictive values (NPVs) [30]. Saliva and nasal swabs had lower but still acceptable sensitivities. Self-sampling of nasal and throat swabs and nasal swabs was not associated with any major compromise of testing performance [30]. As collection of these specimens is technically easier than NPS, the lessened manpower requirement for experienced healthcare workers (HCWs) and protective facilities would facilitate their use in large-scale testing initiatives in an ambulatory setting [3], especially in low- and middle-income countries where resources may be limited [30]. Their reduced invasiveness compared with NPS would also help to enhance their acceptability to patients [17, 18, 23]. Throat swabs, on the other hand, were reported to have a much lower sensitivity and PPV, and were thus not being recommended as a reliable sampling approach [30].
The use of gargle as a testing sample, acquired by forcing the cricopharyngeal muscle to oscillate over the posterior pharyngeal wall with saline or water, has recently been explored as a self-sampling approach for SARS-CoV-2 testing. While gargling has been advocated by a number of recent narrative commentaries as a noninvasive, swab-free and virus transport medium-free sampling approach [31–34], a comprehensive review of the testing performance of gargle, however, is still lacking. Findings from original studies of gargles remain inconclusive, ranging from the reporting of satisfactory [35] to poor sensitivity for detecting SARS-CoV-2 compared to NPS/ONPS as the gold standard [36]. In particular, the comparative performance of gargling based on different mediums, including saline and water-based gargle, has not been directly compared nor previously reviewed [35, 37].
Here we report a systematic review of the performance of gargling for the detection of SARS-CoV-2. We comprehensively review the performance of gargle samples with a view to inform the choice of the most appropriate alternative sampling approach for SARS-CoV-2 testing in an evidence-based manner. The review is stratified according to populations with different infection status to examine the testing performance in two testing scenarios, including for the purpose of diagnosing SARS-CoV-2 infections in suspected populations presenting clinically in an ambulatory care setting and for the monitoring of viral shedding in populations with confirmed SARS-CoV-2 infections and discharged cases to inform downstream clinical and public health decisions in the hospital and community settings. We also assess the performance of saline gargle and water gargle for RT-PCR testing, and the determinants affecting their test performance.
Methods
Search strategy and selection criteria
A standardised search was done in PubMed, OVID Medline, Embase and Web of Science, using the search term“(((((gargle lavage) OR (gargle)) OR (gargling)) OR (mouthwash)) OR (mouth rinse)) AND ((((((((SARS-CoV-2) OR (sars cov 2)) OR (COVID-19)) OR (covid)) OR (ncov)) OR (2019-nCoV)) OR (n-CoV)) OR (coronavirus))”. Given the role of preprints in the timely dissemination of research studies during the coronavirus disease 2019 (COVID-19) pandemic, medRxiv and bioRxiv servers was also searched using the search term “((SARS CoV 2) OR COVID19 OR covid) AND (gargle OR gargling OR mouthwash OR (mouth rinse))”. The search on preprint servers was simplified because of their reduced search functionality. The search was done on 28 September 2021, to retrieve articles from 1 January 2019 with no language restrictions. Additional relevant articles from the reference sections were also reviewed.
Original clinical studies examining the performance of gargles for SARS-CoV-2 RT-PCR testing using either NPS or ONPS as the reference gold standard were included. Studies conducted in either suspected individuals or confirmed cases of SARS-CoV-2 infection were both included, but were analysed separately to address the performance of gargling in the two corresponding contexts. Studies on suspected individuals are useful for examining the test performance for correctly diagnosing individuals having the infection [30]. On the other hand, studies on confirmed cases are useful for assessing test performance for continuous monitoring of virus shedding in patients in different stages to inform related clinical management and public health decisions. Studies without data on performance of gargles for identifying SARS-CoV-2 by RT-PCR, with a detailed collection method that deviated from the definition of gargling, or those that only examined viricidal effect of gargles, were excluded.
Two authors (N.N.Y.T. and H.C.S.) screened all the articles, with disagreement resolved by consensus together with a third author (D.K.M.I.). Studies identified from different databases were de-duplicated after screening. Two authors (N.N.Y.T. and H.C.S.) independently extracted data from the included studies, with disagreement resolved by consensus with a third author (D.K.M.I.). Two authors (N.N.Y.T. and D.K.M.I.) assessed studies for methodological quality, including risk of bias and applicability, adapted from the Quality Assessment of Diagnostic Accuracy Studies tool for diagnostic studies [38].
Data analysis
For the included studies, either individual data or summary estimates of sample size, number of true-positive, true-negative, false-positive and false-negative results in each study were extracted. By use of a standardised data extraction chart, we also retrieved information on study period, country, setting, positivity rate of the corresponding gold-standard test adopted (NPS/ONPS), symptomatic status of population, infection status of population, criteria for SARS-CoV-2 infection among confirmed cases, sampling approaches, gargling technique and procedures, peer-review status, age of population, inclusion of children population, target genes assessed, and the cycle threshold value for positive test result. These findings were checked for agreement.
The sensitivity, specificity, PPV and NPV of RT-PCR tests and the 95% confidence intervals were calculated for gargles, with random-effects meta-analyses using the inverse variance method [39] and a restricted maximum-likelihood estimator for heterogeneity [40, 41], with NPS or ONPS as the reference. Studies capturing both positive and negative individuals for the reference gold standard were included for the calculation of all four performance indicators, while studies including only individuals positive for the reference gold standard were only included in the calculation of sensitivity, as the calculation of specificity, PPV and NPV was not allowable with the absence of negative cases. Freeman–Tukey double arcsine transformation was incorporated for normalising and stabilising the variance of sampling distribution of proportions, and pooled estimates were back-transformed using the harmonic mean [42]. The Q statistic and its p value were calculated to test whether effect sizes departed from homogeneity, and the I2 statistic and its 95% CI were calculated to examine the proportion of dispersion due to heterogeneity [43, 44]. Heterogeneity was assessed as per the definition in the Cochrane Handbook for Systematic Reviews of Interventions [45]. To assess possible factors contributing to heterogeneity, subgroup analyses were done for gargles in individuals with and without confirmed SARS-CoV-2 infection, respectively. Stratified study-level characteristics included positivity rate of ONPS (<20% or ≥20%), geographical regions (European countries or non-European countries), symptomatic status of population (symptomatic only or symptomatic and asymptomatic), infection status of population (suspected only or suspected and confirmed), peer-review status (yes or no), inclusion of children population (yes or no), and number of target genes assessed in RT-PCR (one gene or two or more genes). To examine the optimal practical implementation of gargling, the testing performance was stratified by gargle media (saline or water), sterilisation of gargles (yes or no), volume of gargling medium (≤5 mL or >5 mL) and duration of gargling procedures (≤10 s or >10 s). Comparisons between samples evaluated against NPS and ONPS were presented using forest plots for gargle, as allowable by the availability of studies adopting two different referential approaches. Sensitivity analysis of using a composite reference comparator, defined as positive if either the NPS, ONPS or gargle was positive, was conducted. Data were analysed with the metafor (version 2.4.0) packages in R (version 3.6.0). The protocol was registered on PROSPERO (CRD42022309314).
Results
From the 757 studies identified in our search, 342 duplicates were excluded. After screening the titles and abstracts of the remaining articles, 45 full texts were screened (figure 1). On the basis of our selection criteria, 27 of those studies were excluded and 18 studies [28, 29, 35, 36, 46–59] met our inclusion criteria (table 1). One study examined two study populations, of suspected infection and of confirmed cases, which were analysed separately [59]. Of these 19 study populations, seven assessed the diagnostic performance of gargle in populations with suspected infection [28, 49, 50, 54, 55, 57, 59], and 12 assessed the use of gargle for monitoring viral shedding in populations already confirmed to have SARS-CoV-2 infections, either as hospital inpatients or after being discharged (table 1) [29, 35, 36, 46–48, 51–53, 56, 58, 59]. Regarding the study location, there were five studies from Canada [28, 47, 50, 52, 57], six from Germany [36, 48, 49, 51, 54, 56] and with the rest (seven) from Finland [29], Turkey [55], China [46], Korea [58], Indonesia [59], Israel [53] and India [35]. These studies spanned from 23 March 2020 to 19 July 2021, covering the period of different prevailing SARS-CoV-2 variants, from the ancestral strain, alpha variant, to its delta variant. The main prevailing strains of SARS-CoV-2 included the ancestral strain for studies in North America, the ancestral and alpha strains for studies in European countries, the ancestral and delta strains for studies in Asian countries, and the ancestral strain for the study in Middle East countries. A total of 4444 individuals were included in the 18 eligible studies. Besides two that did not specify participants’ age [50, 56], all reviewed studies included an adult population, among which eight also included children aged below 18 years [28, 36, 47, 48, 52, 54, 57, 59]. 12 studies [28, 29, 35, 46, 47, 49, 51, 52, 54–56, 58] used NPS and six studies [36, 48, 50, 53, 57, 59] used ONPS, all collected by HCWs, as the reference gold standard. A total of 8888 clinical samples were included in our analysis, including 4444 NPS/ONPS as control, and 4444 gargle samples for testing (1187 saline gargles, 2711 water gargles and 546 gargles without specification).
FIGURE 1.
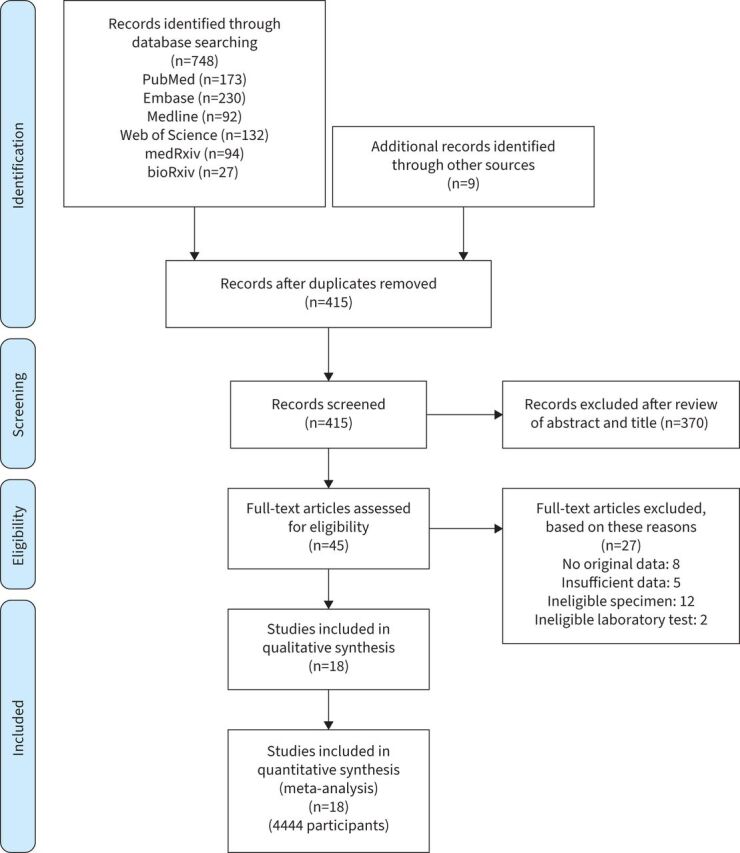
Study profile.
TABLE 1.
Characteristics of included studies of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reverse transcriptase (RT)-PCR testing
| Author (year) [ref.] | Study period | Location | Setting | Positivity rate of (O)NPS (95% CI), % | Patient characteristics | Criteria for SARS-CoV-2 infection | Sampling approaches | Gargling technique | Age (years) | Includes children | Target gene | Cycle threshold | Peer reviewed | Evidence rating |
| Kandel et al. (2021) [ 28 ] | 28 August 2020–25 September 2020 | Canada | Three outpatient COVID-19 assessment centres | 10.53 | Suspected, symptomatic and asymptomatic outpatients | NA | NPS, 0.9% saline gargle | Swish and gargle three times with 3 mL of 0.9% normal saline | Median: 33 IQR: 24–51 |
Yes | E, 5′UTR#, S¶, ORF1ab¶ | 38#, 37¶ | Yes | Moderate |
| Dumaresq et al. (2021) [ 57 ] | 8 October 2020–23 October 2020 | Canada | Drive-through test centre | 12.04 | Suspected, symptomatic and asymptomatic individuals | NA | Combined ONPS, natural spring water gargle | Rinse the mouth with 5 mL of natural spring water for 5 s, tilt the head back and gargle for 5 s, repeat this cycle once, expel the water back in the plastic cup and empty it in a 15 mL conical polypropylene centrifuge tube | Median: 40 Range: 6–91 |
Yes | RdRP, N, E | NA | Yes | High |
| Olearo et al. (2021) [ 49 ] | 1 December 2020–28 February 2021 | Germany | University hospital | 3.45 | Asymptomatic HCW | NA | NPS, tap water gargle | Gargle at home with 5–7 mL of tap water for 30 s, spit into an empty 10 mL sterile container using a straw, with instructions in flyers and videos | NA | No | ORF1ab, E | NA | Yes | Moderate |
| Zander et al. (2021) [ 54 ] | October, 2020–December, 2020 | Germany | Doctors' offices | 32.50 | Suspected mostly symptomatic outpatients | NA | NPS, saline gargle | 30 s gargling of 5 mL 0.9% NaCl | Range: 13–89 | Yes | E | 35 | Yes | High |
| Kocagoz et al. (2021) [ 55 ] | NA | Turkey | Hospital | 20.94 | Suspected symptomatic inpatients | NA | NPS, drinking water gargle | Take a few sips of regular drinking water, gargle and rigorously rinse the mouth forcefully with drinking water for at least 10 s, spit into an empty cup | Range: ≥18 | No | ORF1ab | NA | Yes | Moderate |
| Benoit et al. (2021) [ 50 ] | November 2020–December 2020 | Canada | Two COVID-19 screening clinics | 20.03 | Suspected, symptomatic outpatients or asymptomatic close contacts | NA | Combined ONPS, natural spring water gargle | Gargle with 5 mL of natural spring water for 5 s in the mouth, 5 s in the throat, to repeat the cycle once, then spit in the initial goblet | NA | NA | ORF1ab, E | NA | Yes | High |
| Utama et al. (2022) [ 59 ] | 3 March 2021–19 July 2021 | Indonesia | Hospital | 60.66 | Suspected symptomatic and asymptomatic outpatients; confirmed symptomatic and asymptomatic COVID-19 inpatients and SARS-CoV-2-negative inpatients | Previously diagnosed COVID-19-positive and admitted to hospitals+ | Combined ONPS, gargle§ | Satisfy 45-min fasting period, deeply inhale 5–6 times, throat cough 5–6 times, gargle 2.5 mL of gargle solution, spit into a collection tube with 3 mL BioSaliva Collection Buffer, with instructions in the protocol, and supervised by HCW | Mean: 34.3±12.5 (outpatients); 45.4±16.5 (inpatients) | Yes | Helicase, RdRP | 40 | Yes | Moderate |
| Goldfarb et al. (2021) [ 47 ] | 8 May 2020–11 September 2020 | Canada | COVID-19 collection centre and residence | 74.47 | Confirmed and suspected, symptomatic and asymptomatic COVID-19 outpatients | Had SARS-CoV-2 detected in any clinical sample within a median of 3 days from infection confirmation+ | NPS, sterile 0.9% saline gargle | Swish 5 mL of sterile 0.9% saline in the mouths for 5 s followed by tilting the head back and gargling for 5 s. This gargle cycle was repeated two more times and the saline was expelled into a sterile empty container | Range: 4–71 | Yes | E, RdRp | 40 | Yes | Moderate |
| Michel et al. (2021) [ 36 ] | May 2020 | Germany | Refugee facility | 51.32 | Confirmed and suspected, asymptomatic individuals | Had a previous positive PCR-test for SARS-CoV-2 at a median time of 14 days prior+ | Combined ONPS, sterile water gargle | Gargling for at least 5 s with 10 mL sterile water under the supervision of medical personnel; gargling was demonstrated if necessary | Mean: 29 Range: 7–59 |
Yes | E, ORF1a | 40 | Yes | Moderate |
| Guo et al. (2020) [ 46 ] | 23 March 2020–31 March 2020 | China | Hospital | 4.17 | Confirmed COVID-19 hospitalised and discharged patients | Laboratory-confirmed COVID-19, met the diagnosis criteria of COVID-19 as per national guideline, within 48–57 days of symptom onset | NPS, sterile normal saline gargle | Oscillate over the posterior pharyngeal wall with 20 mL of sterile normal saline for 5–10 s, then spit out the saline into a sterile container | Range: 26–83 | No | N, ORF1ab | NA | Yes | Moderate |
| Poukka et al. (2021) [ 29 ] | NA | Finland | Acute COVID-19 outpatients | 95 | Confirmed symptomatic and asymptomatic COVID-19 outpatients | Had been diagnosed with SARS-CoV-2 infection by RT-PCR on NPS within 1–2 days of diagnosis | NPS, water gargle | Gargle sip of water (1/4 of a glass) for 5–20 s before spitting it into a container | Mean: 38.7±12.6 Range: ≥3 |
NA | E | NA | Yes | Moderate |
| Gertler et al. (2021) [ 48 ] | 7 December 2020–11 January 2021 | Germany | University hospital and residence | 100 | Confirmed symptomatic COVID-19 outpatients | Initial laboratory RT-PCR confirmation of SARS-CoV-2 by professional-collected, combined ONPS in a testing centre, enrolled within 2–15 days from symptom onset | Combined ONPS, tap water gargle | Gargle 10 mL tap water and spit into a plastic container; written instructions and observed without any additional verbal instructions | Median: 31.5 Range: 17–66 |
Yes | E, ORF1ab | NA | Yes | Moderate |
| Mittal et al. (2020) [ 35 ] | May 2020–June 2020 | India | Tertiary care hospital | 100 | Confirmed symptomatic and asymptomatic COVID-19 inpatients | RT-PCR-confirmed patients with SARS-CoV-2 infection within 72 h of diagnosis by a trained HCW, as per the national and hospital policies | NPS, sterile normal saline gargle | Perform gargle with 8–10 mL sterile normal saline for 15–20 s and spit back to a screw-capped container | Mean: 45.08±12.78 | No | N1, N2 | 40 | Yes | Moderate |
| Kwon et al. (2022) [ 58 ] | 7 May 2021–7 July 2021 | Korea | Medical centres and hospital | 50.50 | Confirmed symptomatic and asymptomatic COVID-19 inpatients and healthy individuals | Previously confirmed COVID-19 patients hospitalised in medical centres and hospital+ | NPS, Bean extract gargle | Swirl and gargle 5 mL of bean extract-based gargle for 2 min, spit into a tube | Mean: 43.7, Range: 18–83 | No | RdRP, E, N | NA | Yes | Moderate |
| Hitzenbichler et al. (2021) [ 51 ] | 27 April 2020–8 December 2020 | Germany | Medical centre | 85.29 | Confirmed symptomatic and asymptomatic COVID-19 inpatients | Patients with COVID-19 admitted to nonintensive care medical ward, with laboratory-confirmed SARS-CoV-2 infection (defined by a positive RT-qPCR result before inclusion) | NPS, sterile saline or water gargle | Gargle 10 mL medical grade saline or water for 5–10 s, oscillating over the posterior pharyngeal wall, transfer the liquid into a sterile test tube | Mean: 57.5±14.9, Range: 22–83 | No | E | NA | Yes | Moderate |
| Kinshella et al. (2021) [ 52 ] | 4 September 2020–29 October 2020 | Canada | Hospital COVID-19 collection centre | 100 | Confirmed symptomatic COVID-19 outpatients | Patients with positive RT-PCR result of NPS | NPS, sterile saline gargle | Swish 5 mL of sterile 0.9% saline for 5 s, tilt the head back and gargle for 5 s. Repeat the swish and gargle cycle two more times, expel the saline into a sterile empty container. Saline gargle was self-collected with collection observed by nursing staff, or with written and visual collection instructions, but no observation and no guidance provided by an HCW | Median: 29.5 IQR: 14–41 |
Yes | E, RdRP | 40 | Yes | Moderate |
| Biber et al. (2021) [ 53 ] | July 2020–September 2020 | Israel | Nonhospital facilities dedicated to COVID-19 patients in isolation | 84.67 | Confirmed symptomatic and asymptomatic COVID-19 outpatients in isolation | COVID-19 outpatients previously RT-PCR-confirmed by combined ONPS | Combined ONPS, normal saline (0.9%) gargle | Rinse and gargle 10 cc (10 mL) of normal saline (0.9%) for 10–20 s, spit into a sterile container | Range: 18–73 | No | N, E, RdRP | 40 | Yes | High |
| Casati et al. (2022) [ 56 ] | NA | Germany | Medical university | 48.72 | Confirmed symptomatic COVID-19 ambulatory patients and negative close contacts | Previously tested positive for SARS-CoV-2 via RT-PCR on NPS+ | NPS, sterile saline gargle | Perform gargling with 8 mL of sterile 0.9% saline, and spit into a sterile container without additives, after verbal and visual instructions | NA | NA | E | NA | Yes | Moderate |
CI: confidence interval; COVID-19: coronavirus disease 2019; E: envelope gene; HCW: healthcare worker; IQR: interquartile range; ONPS: oropharyngeal–nasopharyngeal swab; ORF: open reading frame gene; N: nucleocapsid gene; NA: not available; NPS: nasopharyngeal swab; RdRP: RNA-dependent RNA polymerase gene; RT-qPCR: quantitative reverse transcription PCR; S: spike protein gene; UTR: untranslated region. Normal saline denotes 0.9% saline. #: for NPS only. ¶: for gargle only. +: for confirmed cases only. §: no information on gargling medium.
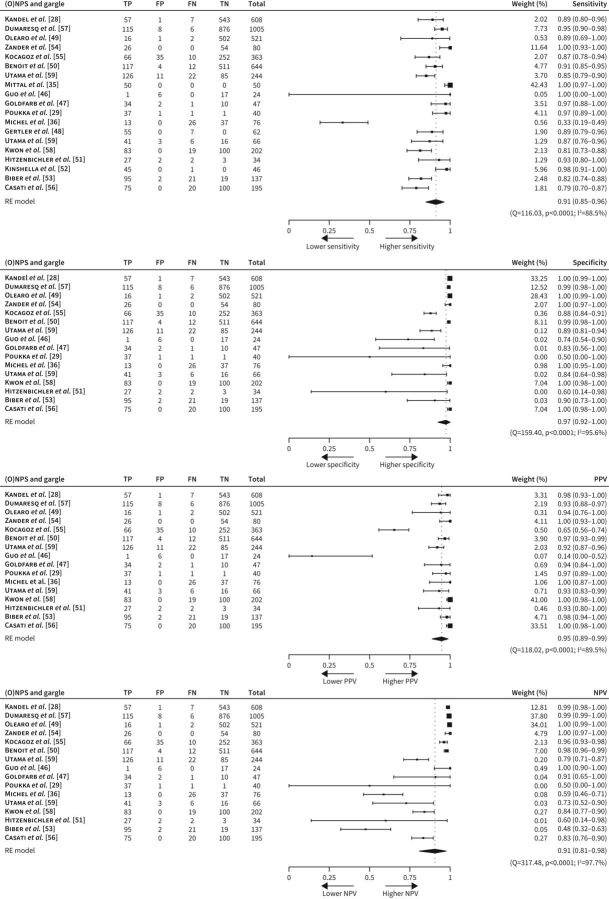
Overall pooled performance estimates of gargles among these studies included a sensitivity of 91% (95% CI 85–96), a specificity of 97% (92–100), a PPV of 95% (89–99) and an NPV of 91% (81–98; figure 2). When stratified by the testing setting, the overall performance generally remained similar in the two testing scenarios. Among the seven studies consisting of suspected individuals presenting with symptoms [28, 50, 54, 55, 57, 59] or asymptomatic individuals with known exposure to confirmed cases [28, 49, 50, 54, 57, 59], the diagnostic performance of gargle for SARS-CoV-2 infection included a sensitivity of 91% (95% CI 87–95), a specificity of 98% (94–100), a PPV of 93% (85–99) and an NPV of 98% (93–100; figure 3). Among the 12 studies on individuals confirmed to have SARS-CoV-2 infection [29, 35, 36, 46–48, 51–53, 56, 58, 59], the testing performance of gargle for monitoring viral shedding of SARS-CoV-2 included a sensitivity of 91% (95% CI 80–98), a specificity of 94% (81–100), a PPV of 95% (84–100) and an NPV of 78% (63–90; figure 4).
FIGURE 2.
Meta-analysis of the sensitivity, specificity, positive predictive value and negative predictive value using (oropharyngeal–)nasopharyngeal swab ((O)NPS) as a reference standard. FN: false negative; FP: false positive; NPV: negative predictive value; PPV: positive predictive value; TN: true negative; TP: true positive; RE: random effects.
FIGURE 3.
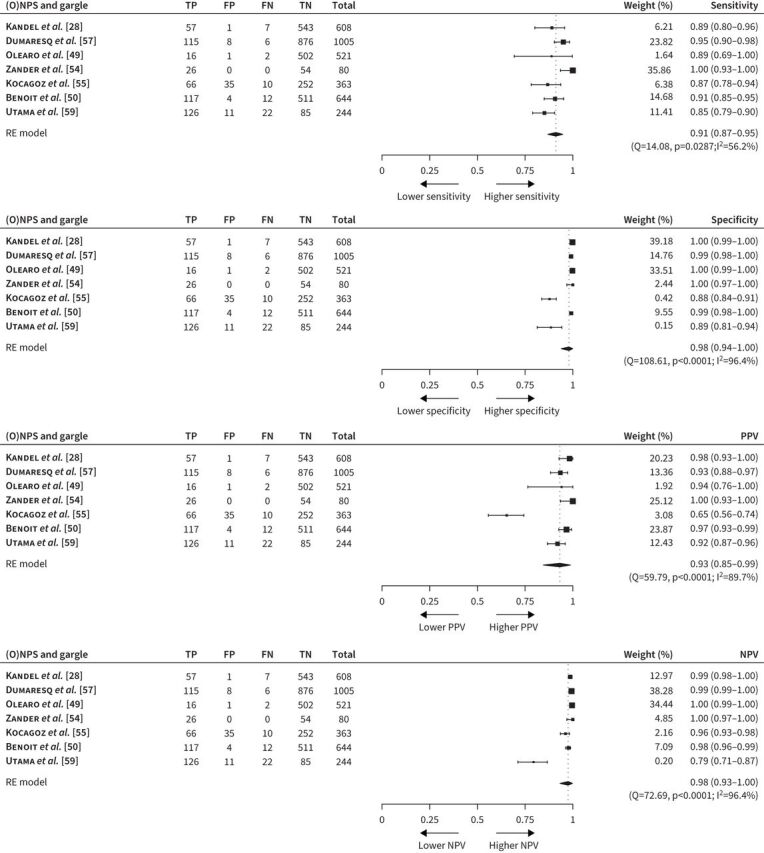
Meta-analysis of the sensitivity, specificity, positive predictive value and negative predictive value using (oropharyngeal–)nasopharyngeal swab ((O)NPS) as a reference standard for diagnostic testing in suspected individuals. FN: false negative; FP: false positive; NPV: negative predictive value; PPV: positive predictive value; TN: true negative; TP: true positive; RE: random effects.
FIGURE 4.
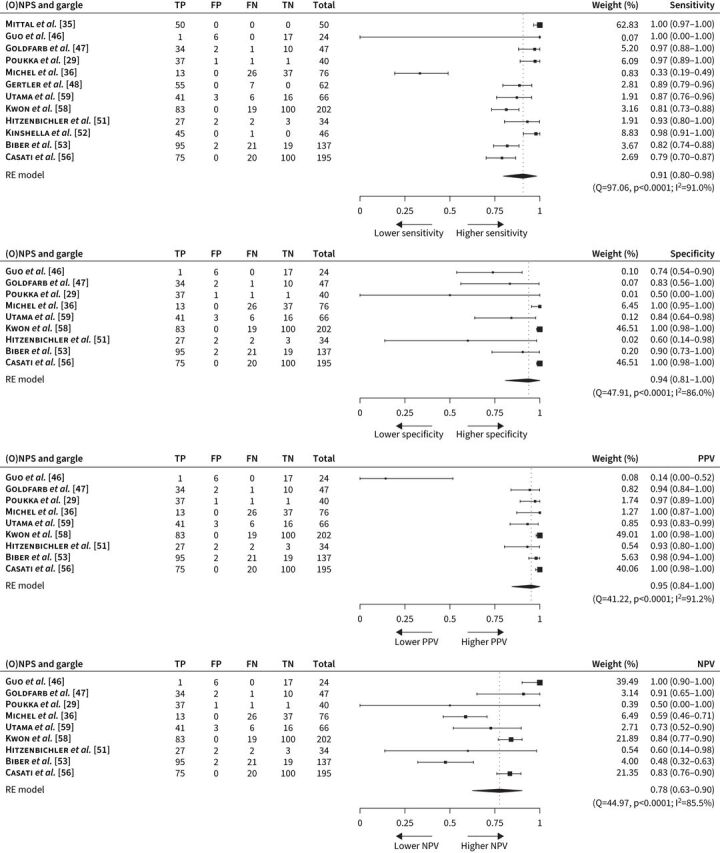
Meta-analysis of the sensitivity, specificity, positive predictive value and negative predictive value using (oropharyngeal–)nasopharyngeal swab ((O)NPS) as a reference standard among studies consisting of confirmed cases. FN: false negative; FP: false positive; NPV: negative predictive value; PPV: positive predictive value; TN: true negative; TP: true positive; RE: random effects.
All performance indicators of gargles for RT-PCR testing were in general heterogeneous (I2 ranged from 88.5% to 97.7%; figure 2), with diverse population characteristics and study settings. Overall, a lower testing sensitivity was reported in studies using ONPS as the reference standard (83% versus 96% in studies using NPS), or studies involving asymptomatic individuals only (62% versus 91% in studies with symptomatic and asymptomatic individuals and 91% in symptomatic individuals) (table 2). Studies including a child population had a similar pooled sensitivity (89%) to the overall figure. A lower PPV was found for studies with a lower disease prevalence (positive rate of less than 20%) (84% versus 96%), or studies done in Asia (85% versus 95% in European countries and 96% in North America; table 2). A lower NPV was found for studies with ONPS as the reference standard (81% versus 97%), studies with a disease prevalence greater or equal to 20% (84% versus 100%) and studies done in Middle East countries (47% versus 99% in North America, 90% in European countries and 85% in Asia; table 2). Heterogeneity generally remained moderate to substantial (I2≥30%) in most stratified analyses.
TABLE 2.
Stratified pooled estimates on diagnostic indicators for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) using gargling as an alternative sampling approach among all populations by study characteristics
| Study characteristic | Studies, n | Sensitivity estimate (95% CI) | I2 (95% CI), % | Studies, n | Specificity estimate (95% CI) | I2 (95% CI), % | PPV estimate (95% CI) | I2 (95% CI), % | NPV estimate (95% CI) | I2 (95% CI), % |
| Gargle | 19 | 91 (85–96) | 88 (86–91) | 16 | 97 (92–100) | 96 (95–96) | 95 (89–99) | 89 (87–91) | 91 (81–98) | 98 (97–98) |
| Population | ||||||||||
| Suspected only | 7 | 91 (87–95) | 56 (33–71) | 7 | 98 (94–100) | 96 (95–97) | 93 (85–99) | 90 (86–92) | 98 (93–100) | 96 (95–97) |
| Confirmed and suspected | 12 | 91 (80–98) | 91 (89–93) | 9 | 94 (81–100) | 86 (81–90) | 95 (84–100) | 91 (88–94) | 78 (63–90) | 86 (80–89) |
| Reference standard | ||||||||||
| NPS | 12 | 96 (90–99) | 74 (65–81) | 10 | 97 (88–100) | 96 (95–97) | 93 (80–100) | 93 (91–95) | 97 (89–100) | 94 (92–95) |
| ONPS | 7 | 83 (68–94) | 95 (93–96) | 6 | 96 (90–100) | 93 (90–95) | 96 (93–98) | 27 (0–52) | 81 (59–96) | 99 (98–99) |
| Gargle medium# | ||||||||||
| Saline | 8 | 97 (90–100) | 82 (76–87) | 6 | 96 (86–100) | 92 (89–94) | 94 (72–100) | 96 (94–97) | 92 (74–100) | 96 (95–97) |
| Water | 7 | 86 (70–97) | 94 (92–95) | 6 | 100 (96–100) | 96 (94–97) | 93 (82–99) | 88 (84–92) | 96 (79–100) | 99 (99–99) |
| Sterility of gargle | ||||||||||
| Saline gargle (n=8) | ||||||||||
| Sterile | 5 | 100 (90–100) | 81 (72–88) | 3 | 91 (64–100) | 89 (82–93) | 82 (20–100) | 97 (96–98) | 92 (78–100) | 64 (31–81) |
| Nonsterile/not mentioned | 3 | 91 (77–99) | 85 (73–92) | 3 | 99 (93–100) | 80 (63–89) | 99 (96–100) | 0 (0–86) | 90 (49–100) | 98 (98–99) |
| Water gargle (n=7) | ||||||||||
| Sterile | 1 | 33 (19–49) | - | 1 | 100 (95–100) | - | 100 (87–100) | - | 59 (46–71) | - |
| Nonsterile/not mentioned | 6 | 92 (88–95) | 30 (0–55) | 5 | 100 (93–100) | 97 (96–98) | 91 (79–99) | 91 (87–93) | 100 (100–100) | 83 (74–89) |
| Volume of gargle¶ | ||||||||||
| ≤5 mL | 10 | 92 (87–95) | 67 (53–77) | 9 | 99 (95–100) | 95 (93–96) | 97 (94–99) | 54 (32–69) | 95 (88–99) | 97 (97–98) |
| >5 mL | 7 | 87 (66–100) | 94 (92–95) | 5 | 93 (75–100) | 88 (82–92) | 91 (60–100) | 96 (94–97) | 74 (49–93) | 92 (88–94) |
| Duration of gargling+ | ||||||||||
| ≤10 s | 6 | 86 (62–100) | 93 (90–95) | 6 | 89 (72–99) | 81 (71–88) | 86 (60–100) | 95 (93–96) | 78 (50–98) | 93 (91–95) |
| >10 s | 8 | 95 (90–99) | 76 (66–83) | 6 | 100 (100–100) | 10 (0–44) | 98 (94–100) | 56 (30–72) | 98 (92–100) | 96 (94–97) |
| Positivity rate of ONPS | ||||||||||
| <20% | 4 | 99 (93–100) | 27 (9–42) | 4 | 98 (87–100) | 99 (98–99) | 84 (44–100) | 96 (94–98) | 100 (99–100) | 4 (0–100) |
| ≥20% | 15 | 89 (82–95) | 91 (89–93) | 12 | 97 (90–100) | 91 (88–93) | 96 (91–99) | 86 (82–89) | 84 (71–94) | 95 (94–96) |
| Region | ||||||||||
| European countries | 8 | 86 (72–97) | 92 (89–94) | 7 | 99 (91–100) | 94 (92–95) | 95 (85–100) | 86 (81–89) | 90 (71–100) | 97 (96–98) |
| North America | 5 | 94 (90–96) | 26 (0–54) | 4 | 100 (100–100) | 34 (0–61) | 96 (93–98) | 0 (0–100) | 99 (98–100) | 61 (33–78) |
| Asia | 5 | 96 (83–100) | 85 (77–90) | 4 | 91 (74–100) | 90 (84–93) | 85 (44–100) | 98 (97–98) | 85 (73–95) | 79 (64–88) |
| Middle East | 1 | 82 (74–88) | - | 1 | 90 (73–100) | - | 98 (94–100) | - | 47 (32–63) | - |
| Symptomology§ | ||||||||||
| Symptomatic | 3 | 91 (83–97) | 62 (28–80) | 1 | 88 (84–91) | - | 65 (56–74) | - | 96 (93–98) | - |
| Symptomatic and asymptomatic | 13 | 91 (87–95) | 79 (73–84) | 12 | 98 (93–100) | 95 (93–96) | 97 (95–99) | 56 (38–68) | 89 (77–98) | 97 (97–98) |
| Asymptomatic | 2 | 62 (9–100) | 94 (89–97) | 2 | 100 (100–100) | 0 (0–98) | 98 (87–100) | 0 (0–81) | 86 (29–100) | 99 (98–99) |
| Included childrenƒ | ||||||||||
| Yes | 9 | 89 (77–97) | 93 (91–95) | 7 | 97 (91–100) | 94 (91–95) | 95 (93–97) | 6 (0–29) | 90 (76–99) | 98 (97–98) |
| No | 7 | 94 (85–100) | 77 (66–84) | 6 | 93 (79–100) | 96 (94–97) | 86 (59–100) | 96 (95–97) | 89 (68–100) | 98 (97–98) |
| Peer reviewed | ||||||||||
| Yes | 19 | 91 (85–96) | 88 (86–91) | 16 | 97 (92–100) | 96 (95–96) | 95 (89–99) | 89 (87–91) | 91 (81–98) | 98 (97–98) |
| No | 0 | – | – | 0 | – | – | – | – | – | – |
| Target gene | ||||||||||
| 1 gene | 5 | 92 (83–98) | 78 (65–86) | 5 | 96 (74–100) | 94 (92–96) | 94 (80–100) | 91 (88–94) | 93 (72–100) | 93 (90–95) |
| ≥2 genes | 14 | 91 (82–97) | 91 (88–93) | 11 | 97 (91–100) | 96 (95–97) | 95 (89–99) | 87 (82–90) | 89 (77–97) | 98 (98–99) |
PPV: positive predictive value; NPV: negative predictive value; ONPS: oropharyngeal–nasopharyngeal swab. Other regions included European countries, Mediterranean countries and Asian countries. Target gene refers to number of gene regions tested by reverse transcriptase PCR assays, i.e. assays detecting both nucleocapsid (N)1 and N2 were counted as dual targets. #: three studies did not specify the gargling medium (water or saline), and one study used bean-extract gargle. One study used sterile gargle, without specifying water or saline gargle. ¶: two studies did not report the volume of gargling solutions. +: five studies (six study populations) did not report the duration of gargling. §: one study did not report the symptomatic status of the population. ƒ: three studies did not specify the inclusion of children population.
Regarding the medium of the gargle, saline gargle was used in eight studies [28, 35, 46, 47, 52–54, 56], with five specifying sterile saline gargle [35, 46, 47, 52, 56] while the other three did not specify on medium sterility; [28, 53, 54] water gargle was used in seven studies [29, 36, 48–50, 55, 57], with one specifying sterile water gargle [36] and the rest used nonsterile water gargle, including natural spring water gargle and tap water gargle [29, 48–50, 55, 57]. One study used bean-extract gargle [58], while the remaining two did not specify the gargling mediums [51, 59]. Studies using saline gargle and using water gargle reported an overall sensitivity of 97% and 86% respectively, and a broadly similar overall specificity (96% versus 100%), PPV (94% versus 93%) and NPV (92% versus 96%) (table 2). The respective sensitivity of saline gargle and water gargle was maintained on stratification by testing settings, including for diagnostic testing on individuals with suspected infection (96% versus 92%) (supplementary table 2), and for monitoring viral shedding of SARS-CoV-2 in individuals with confirmed SARS-CoV-2 infection (98% versus 78%) (supplementary table 3). Among the eight studies using saline gargle, five studies specified using sterile saline gargle and generally suggested a potential of higher overall sensitivity (100% versus 91%) compared to the three that did not specify if the gargle was sterile or not (table 2).
The volume of gargling solution used in the studies ranged from 2 mL to 20mL, and the duration of gargling during the sampling procedure varied from 5 s to 2 min. Regarding the volume of gargle, studies using ≤5 mL of gargling solution reported a higher overall testing sensitivity (92% versus 87%), specificity (99% versus 93%), PPV (97% versus 91%) and NPV (95% versus 74%), compared to those using >5 mL of gargling solution (table 2). For the duration of gargling during the sampling procedure, studies requiring a total gargling time of >10 s reported a higher overall testing sensitivity (95% versus 86%), specificity (100% versus 89%), PPV (98% versus 86%) and NPV (98% versus 78%), compared to those requiring ≤10 s of gargling (table 2). A robust examination of the impact of medium sterility, volume and gargling duration in different testing settings, however, was not possible due to the small number of studies on further stratification.
Sensitivity analysis of gargling using a composite reference comparator showed a similar sensitivity (90% versus 91%) and NPV (91% versus 91%) to full analysis using ONPS as the reference standard (supplementary figure 1, table 4). Quality assessment showed that all included studies were of good or acceptable quality and low risk of bias (supplementary figure 2). Sensitivity analysis was not performed regarding peer-review status as all included studies were peer-reviewed (table 2). Publication bias was examined by funnel plots of the performance indicators, which suggested the presence of a low risk of bias for sensitivity and PPV, but some risk of bias for specificity and NPV (supplementary figure 3).
Discussion
This is the first systematic review and meta-analysis examining self-collection of gargles for SARS-CoV-2 RT-PCR testing. We comprehensively evaluated the performance and reported pooled estimates of sensitivity, specificity, PPV and NPV of gargling approaches for the detection of SARS-CoV-2, and compared the differential performance of different gargling medium, volume and duration, and the contextual performance in diagnostic testing among suspected individuals and clinical virologic monitoring among confirmed COVID-19 patients. Our review included studies conducted from 23 March 2020 to 19 July 2021, covering the period of different prevailing SARS-CoV-2 variants, from the ancestral strain, alpha variant, to its delta variant. Our review minimised potential bias from different testing techniques by including only studies with RT-PCR testing procedures and excluding rapid diagnostic tests using other technologies. Studies with different positivity rate and population characteristics were included in our studies to increase the generalisability of our findings, while stratified analysis provide a context-specific estimate for different gargling mediums and procedures, and test application settings. By stratifying the performance indicators of gargling with respect to their reference gold standard, status of infection, gargling mediums, procedural implementation and study settings, our review identified the factors of heterogeneity and examined their potential impact on testing performance. The bias of using different reference comparators were minimal in our review, with similar results generated in the sensitivity analysis using a composite reference comparator and in the overall analysis using ONPS as a reference comparator.
Our findings suggest that gargling as a sampling approach offered a very good performance for detecting SARS-CoV-2 by RT-PCR, with an overall pooled sensitivity of 91%, specificity of 97%, PPV of 95% and NPV of 91%, making it a reliable alternative sampling option for SARS-CoV-2 testing. This is compatible with the reporting of the release of SARS-CoV-2 in a high quantity in the airway epithelial cells of an infected person [60]. The higher overall sensitivity form studies using saline gargle compared to those using water medium (97% versus 86%) suggested the potential dependency of testing performance on the nature of gargle medium, which is compatible with the suggestion that hypotonic solutions may prematurely lyse cells or disrupt lysosomes and degrade RNA and result in a sharp decline in the RT-PCR product [61]. However, further studies employing a head-to-head comparison of the two media would be needed before a definitive conclusion can be drawn regarding their comparative performance. Likewise, although the near-perfect overall sensitivity (100%) reported in studies exclusively using sterile saline gargle is compatible with previous reports of giving a similar viral yield for detection of SARS-CoV-2 by sterile normal saline and viral transport media (VTM) [62], and in agreement with the World Health Organization recommendation to use sterile normal saline as an alternative for RT-PCR testing when VTM is not available [4], this would need to be confirmed by further studies.
The near-perfect sensitivity achieved by saline gargle (97%) and sterile saline gargle (100%) implied the great advantage of reducing potential false negativity, which is of utmost importance and has significant implications for the control of an evolving COVID-19 epidemic in a community. As infected individuals not detected by the screening test will fail to be properly channelled to the required clinical management and isolation, they may seed further transmission and infection in the community. Likewise, false negativity of virologic monitoring among confirmed COVID-19 patients may result in premature discharge from hospitals or isolation facilities when still actively shedding viruses and risk further transmission. The maintained high sensitivity of saline gargle both for diagnosing SARS-CoV-2 infection in suspected individuals (96%) and for monitoring of viral shedding in confirmed COVID-19 patients (98%) supported its suitability for both testing purposes. For use as diagnostic testing in suspected individuals, the pooled sensitivity of saline gargle (96%) is slightly higher than those achieved by most other common sampling approaches, including nasal swab (86%), saliva (85%) and throat swab (68%), and only lower than that reported for pooled nasal and throat swab (97%) by a recent systematic review [30]. Although our results suggested that sensitivity may be further improved to near perfection by the exclusive use of sterile saline gargle, robust stratified examination for different testing settings was not allowable in this study. On the other hand, the sensitivity of 86% achieved by water gargle indicated that almost one seventh of infected cases may be missed, which suggested that it may not be an optimal sampling approach for the detection of SARS-CoV-2 in settings where potentially missing cases may be consequentially serious, and further studies with head-to-head comparisons are needed to better clarify the comparative performance of different gargle media.
In a disease screening setting, false-positive results may incur unnecessary confirmatory tests and risk wrongful hospitalisation with potential unnecessary risk exposure. False-positive results in a virologic monitoring setting are associated with delayed discharge. In the COVID-19 epidemic, cases with positive screening results are generally followed-up by health agencies and verified by further confirmatory testing. However, the concerns associated with false positivity would be low for saline gargle owing to its high specificity (100%) and PPV (99%) in diagnostic testing, and high overall specificity (96%) and high overall PPV (94%) in different settings.
Our findings also help to refine the practical procedural considerations for using gargle as a sampling medium, by highlighting that using a larger amount of gargling solution (>5 mL) or a shorter gargling duration (≤10 s) may jeopardise testing sensitivity, potentially due to a dilution effect or an inability to capture sufficient viral particles. The need to avoid using an excessive amount of gargle is compatible with the usual usage of a relatively small amount (2–3 mL) of transport medium for swabs as per the US Centers for Disease Control and Prevention's recommendation [63]. According to our findings, adopting a certain minimal duration (>10 s) and a maximal allowable volume of saline (≤5 mL) would help to ensure an adequate amount of viral particles to be captured, and to give an optimal concentration for a sufficiently sensitive result.
Although a stratified comparison of performance in adults versus children is not permissible by the data, the lack of notable difference in sensitivity (91% versus 89%) between the pooled overall estimate from all 18 studies and eight of the studies including children aged 3–18 years in their study population potentially suggested a comparable performance of gargling in adults and children, and that gargle would be an appropriate sampling option for children. The higher pooled sensitivity (89%) by gargle in studies including children as compared to that reported previously for saliva (83.6%) [64] further suggests gargling to be a viable option for a child population. Although some caution has been advocated for its use in infants due to the technical difficulty for this age group [65], gargling has also been used in some settings for the molecular testing of common respiratory infections in children [66, 67]. Further large-scale study with different age groups would help to better address the differential performance of gargling in extremities of age.
The observed lower PPV (84%) in studies with lower disease prevalence was compatible with the general understanding. On stratified analysis by study areas, a much lower PPV (85%) was observed in the subset of three studies conducted in Asia compared with those from other continents (95%-98%), this may be related to the particularly low disease prevalence (4.17%) reported in a study from China [46].
Besides the analytic sensitivity of a test, its utility also depends on its availability, accessibility, acceptability, affordability, frequency of testing, and turnaround time [68, 69]. Moreover, the ease of sample collection and delivery, availability of consumables, and need for special protective equipment are critical factors that may further constrain its use in various community settings [57]. As many SARS-CoV-2-infected individuals can be asymptomatic [70] and potentially infectiousness before symptom onset [71, 72], a test that can be easily repeated would be highly desirable for testing suspected individuals, including potential contacts in the community and people under quarantine [68, 69]. Although generally regarded as a reference clinical sample for SARS-CoV-2 testing, NPS has limited utility in public health practice because of its technical difficulty, requirement of skilled HCWs for its collection [11], procedural discomfort, and expertise and facilities constraints [12]. Under the evolving COVID-19 pandemic situation, gargling as an alternative noninvasive swab-free self-sampling approach may offer a number of practical and logistical advantages if used in a suitable context in different countries. The simple execution of the gargle self-collection procedure allowed for repeated testing in the absence of trained HCWs and high-level personal protective equipment and negative-pressure facilities. The high stability of viral RNA in gargle at room temperature for several days [47, 57] without requiring cold-chain delivery [73] allowed for the option of late and flexible drop-off specimen delivery. Besides being technically simpler to collect [28] than deep-throat saliva, gargling also offered a number of technical advantages. The higher homogeneity and lower viscosity would help to reduce the risk of clogging of liquid handling systems and testing failure [36, 74, 75]. The reduced steps needed for extraction, resuspension and transfer would help to minimise the risk of cross-contamination and false-positivity [76]. Together, these factors lead to advantages of easy deployment and scalable capacity for any screening or surveillance regimens where frequent and large-scale mass testing would be needed [69], especially in low-to-middle-income countries and remote settings with limited healthcare infrastructure and clinic and hospital accessibility [3]. The minimised manpower and zero swab requirement also allow limited resources to be redeployed to other competing needs in resource-constrained settings. The high acceptability of self-collected gargle as reported by some studies suggested that it may also potentially help to abate the testing hesitancy that is not uncommonly encountered in mandatory testing scenarios [35, 47, 77]. Our findings provide an evidence-based justification for COVID-19 testing approaches in national and international guidelines, and support the recommendation of gargling as an inexpensive and easy sampling approach for routine repeated testing in various community settings [78], including primary and secondary schools, universities [77, 79, 80], nursing homes [81], daycare facilities for children aged 1–7 years [67], children's hospitals [82], homes [83] and in other specific high-risk populations having medically indicated needs for repeated testing [78].
This review has several limitations. First, publication bias might have resulted in an overestimation of some of the performance indicators we examined if studies with null or negative results were less likely to be published [84]. However, the presence of substantial risk of bias was not supported by the funnel plots of different performance indicators. Although only a small number of relevant primary studies are available, our extended search of multiple literature databases and the inclusion of preprints should have helped to minimise the risk of publication bias in our review. Second, substantial heterogeneity was observed in the performance indicators of gargles. Various factors for heterogeneity, including gargling media, volume of gargling solution, duration of gargling, positivity rate of ONPS, reference comparators, number of genes tested and study settings, were adjusted in stratified analysis and should have helped to assess their impact on the performance indicators. Third, various factors potentially contributing to the residual heterogeneity, including case definitions of SARS-CoV-2 infection, RNA extraction methods, RT-PCR targeted genes and time delay between symptom onset and sample collection, were not accounted for due to the limited number of studies available and may have affected our observed results [53]. Detailed exploration of the adjusted impact of these factors identified in the subgroup analyses by use of a multivariate meta-regression approach was not possible with the small sample size and reduced power due to a small number of available studies. Therefore, caution regarding the generalisability of the findings should be exercised when applying the results to real-life programmes with varying sampling and testing logistics in different settings. For instance, the long delay between sample collection and symptom onset in some studies may have impacted the observed sensitivity and may not be typical of real-life situations [36]. Fourth, studies examining diagnostic performance of gargle in asymptomatic individuals only was underrepresented compared to studies on symptomatic individuals, which might have limited the generalisability of our results to the situation of general population screening. Although studies with different prevailing SARS-CoV-2 strains were included, covering the ancestral strain to delta variants, the performance of gargling, like all other sampling methods, may vary across variants, and our findings may not be fully applicable to new evolving variants. As studies examining the performance of saline gargle or water gargle were not compared head-to-head, the result may be affected by a number of factors including study population, prevalence, volume of gargling solution, instructions given and PCR methodology. The performance of saline and water gargle observed in our review would need to be further confirmed by studies with a head-to-head comparison. Further study examining the impact of duration and intensity of gargling, different volumes of gargling solutions, populations of different symptomatology, different RNA extraction methods, different cycle thresholds, different case definitions and different reference comparators is warranted for further optimising the testing performance of gargling.
Conclusions and future perspectives
In summary, in this review we synthesised the pooled performance estimates of gargling and found that self-collection of gargle achieved a high performance compared with the gold standard of ONPS and represented a reliable alternative sampling approach for both the purposes of diagnosing SARS-CoV-2 infection and monitoring viral shedding. A potential dependency of testing performance on the nature and sterility of gargle media was highlighted, which suggested the need for further studies with head-to-head comparisons for assessing the use of different gargling media in SARS-CoV-2 testing. Our study also informed the practical implementation of gargling, in terms of duration of gargling and volume of solution.
As a simple and noninvasive sampling approach that can be easily conducted, self-collection of gargle appears to be a viable option to boost acceptability and accessibility, and to enable efficient, timely and frequent SARS-CoV-2 testing in different community settings. Its lower swab, collection expertise and manpower requirements, and less stringent requirements for specimen collection venue or delivery process may help to overcome the traditional technical and resource limitations of nasopharyngeal swabbing. It may represent a suitable option for easy deployment and upscaling for mass testing in remote or resource-limited settings during the evolving pandemic. Our results provide a useful reference framework for the proper implementation of gargling and the interpretation of SARS-CoV-2 testing results using gargling, both for diagnosing SARS-CoV-2 infection and virologic monitoring in different testing settings.
Supplementary material
Please note: supplementary material is not edited by the Editorial Office, and is uploaded as it has been supplied by the author.
Supplementary material ERR-0014-2022.SUPPLEMENT (411.9KB, pdf)
Footnotes
Provenance: Submitted article, peer reviewed.
Author contributions: G.M. Leung and D.K.M. Ip conceived and designed the study. N.N.Y. Tsang and H.C. So screened literature and extracted data. N.N.Y. Tsang and H.C. So had access to and verified the data. B.J. Cowling advised on the statistical analysis. N.N.Y. Tsang did the statistical analysis. N.N.Y. Tsang, H.C. So, and D.K.M. Ip interpreted the data. N.N.Y. Tsang and D.K.M. Ip wrote the first draft of the manuscript, and all authors provided critical review and revision of the text and approved the final version. D.K.M. Ip had final responsibility for the decision to submit for publication.
Data sharing: The data supporting this meta-analysis are from previously reported studies and datasets, which have been cited. The processed data are available from the corresponding author, upon reasonable request.
Conflict of interest: B.J. Cowling consults for Roche and Sanofi Pasteur. All other authors declare no competing interests.
Support statement: This project was supported by the Theme-based Research Scheme (T11-712/19-N) of the Research Grants Council of the Hong Kong Special Administrative Region Government. The funder of the study had no role in study design, data collection, data analysis, data interpretation, or writing of the report. Funding information for this article has been deposited with the Crossref Funder Registry.
References
- 1.World Health Organization . Weekly epidemiological update on COVID-19 – 6 July 2021. www.who.int/publications/m/item/weekly-epidemiological-update-on-covid-19---6-july-2022 Date last updated: 6 July 2022. Date accessed: 4 August 2022.
- 2.World Health Organization . Weekly epidemiological update on COVID-19 – 11 January 2022. www.who.int/publications/m/item/weekly-epidemiological-update-on-covid-19---11-january-2022 Date last updated: 11 January 2022. Date accessed: 4 August 2022.
- 3.Boum Y, Eyangoh S, Okomo M-C. Beyond COVID-19–will self-sampling and testing become the norm? Lancet Infect Dis 2021; 21: 1194–1195. doi: 10.1016/S1473-3099(21)00197-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.World Health Organization . Laboratory Testing for Coronavirus Disease (COVID-19) in Suspected Human Cases: Interim Guidance, 19 March 2020. World Health Organization, Geneva; 2020. [Google Scholar]
- 5.Centers for Disease Control and Prevention . Interim guidelines for collecting and handling of clinical specimens for COVID-19 testing. www.cdc.gov/coronavirus/2019-nCoV/lab/guidelines-clinical-specimens.html Date accessed: 7 June 2021. Date last updated: 15 July 2022.
- 6.Ek P, Böttiger B, Dahlman D, et al. A combination of naso- and oropharyngeal swabs improves the diagnostic yield of respiratory viruses in adult emergency department patients. Infect Dis 2019; 51: 241–248. doi: 10.1080/23744235.2018.1546055 [DOI] [PubMed] [Google Scholar]
- 7.Labbé A-C, Benoit P, Gobeille Paré S, et al. Comparison of saliva with oral and nasopharyngeal swabs for SARS-CoV-2 detection on various commercial and laboratory-developed assays. J Med Virol; 93: 5333–5338. doi: 10.1002/jmv.27026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yan Y, Chang L, Wang L. Laboratory testing of SARS-CoV, MERS-CoV, and SARS-CoV-2 (2019-nCoV): current status, challenges, and countermeasures. Rev Med Virol 2020; 30: e2106. doi: 10.1002/rmv.2106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pan Y, Zhang D, Yang P, et al. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect Dis 2020; 20: 411–412. doi: 10.1016/S1473-3099(20)30113-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sharma K, Aggarwala P, Gandhi D, et al. Comparative analysis of various clinical specimens in detection of SARS-CoV-2 using rRT-PCR in new and follow up cases of COVID-19 infection: quest for the best choice. PLoS One 2021; 16: e0249408. doi: 10.1371/journal.pone.0249408 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Marty FM, Chen K, Verrill KA. How to obtain a nasopharyngeal swab specimen. N Engl J Med 2020; 382: e76. doi: 10.1056/NEJMvcm2010260 [DOI] [PubMed] [Google Scholar]
- 12.Karligkiotis A, Arosio A, Castelnuovo P. How to obtain a nasopharyngeal swab specimen. N Engl J Med 2020; 383: e14. doi: 10.1056/NEJMc2015949 [DOI] [PubMed] [Google Scholar]
- 13.Pettit SD, Jerome KR, Rouquié D, et al. ‘All In': a pragmatic framework for COVID-19 testing and action on a global scale. EMBO Mol Med 2020; 12: e12634. doi: 10.15252/emmm.202012634 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Shakir SM, Barker AP, Hillyard DR, et al. Combined self-collected anterior nasal and oropharyngeal specimens versus provider-collected nasopharyngeal swabs for the detection of SARS-CoV-2. J Clin Microbiol 2020; 59: e02291-20. doi: 10.1128/JCM.02291-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Vlek ALM, Wesselius TS, Achterberg R, et al. Combined throat/nasal swab sampling for SARS-CoV-2 is equivalent to nasopharyngeal sampling. Eur J Clin Microbiol Infect Dis 2020; 40: 193–195. doi: 10.1007/s10096-020-03972-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.LeBlanc JJ, Heinstein C, MacDonald J, et al. A combined oropharyngeal/nares swab is a suitable alternative to nasopharyngeal swabs for the detection of SARS-CoV-2. J Clin Virol 2020; 128: 104442. doi: 10.1016/j.jcv.2020.104442 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Valentine-Graves M, Hall E, Guest JL, et al. At-home self-collection of saliva, oropharyngeal swabs and dried blood spots for SARS-CoV-2 diagnosis and serology: post-collection acceptability of specimen collection process and patient confidence in specimens. PLoS One 2020; 15: e0236775. doi: 10.1371/journal.pone.0236775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tajima Y, Suda Y, Yano K. A case report of SARS-CoV-2 confirmed in saliva specimens up to 37 days after onset: Proposal of saliva specimens for COVID-19 diagnosis and virus monitoring. J Infect Chemother 2020; 26: 1086–1089. doi: 10.1016/j.jiac.2020.06.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pinninti S, Trieu C, Pati SK, et al. Comparing nasopharyngeal and mid-turbinate nasal swab testing for the identification of SARS-CoV-2. Clin Infect Dis 2020; 72: 1253–1255. doi: 10.1093/cid/ciaa882 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Harrington A, Cox B, Snowdon J, et al. Comparison of Abbott ID Now and Abbott m2000 methods for the detection of SARS-CoV-2 from nasopharyngeal and nasal swabs from symptomatic patients. J Clin Microbiol 2020; 58: e00798-00720. doi: 10.1128/JCM.00798-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rhoads DD, Cherian SS, Roman K, et al. Comparison of Abbott ID Now, DiaSorin Simplexa, and CDC FDA emergency use authorization methods for the detection of SARS-CoV-2 from nasopharyngeal and nasal swabs from individuals diagnosed with COVID-19. J Clin Microbiol 2020; 58: e00760-00720. doi: 10.1128/JCM.00760-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nachtigall FM, Pereira A, Trofymchuk OS, et al. Detection of SARS-CoV-2 in nasal swabs using MALDI-MS. Nat Biotechnol 2020; 38: 1168–1173. doi: 10.1038/s41587-020-0644-7 [DOI] [PubMed] [Google Scholar]
- 23.Palmas G, Moriondo M, Trapani S, et al. Nasal swab as preferred clinical specimen for COVID-19 testing in children. Pediatr Infect Dis J 2020; 39: e267–e270. doi: 10.1097/INF.0000000000002812 [DOI] [PubMed] [Google Scholar]
- 24.Basu A, Zinger T, Inglima K, et al. Performance of Abbott ID Now COVID-19 rapid nucleic acid amplification test using nasopharyngeal swabs transported in viral transport media and dry nasal swabs in a New York City academic institution. J Clin Microbiol 2020; 58: e01136-01120. doi: 10.1128/JCM.01136-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Petruzzi G, De Virgilio A, Pichi B, et al. COVID-19: nasal and oropharyngeal swab. Head Neck 2020; 42: 1303–1304. doi: 10.1002/hed.26212 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Berenger BM, Fonseca K, Schneider AR, et al. Sensitivity of nasopharyngeal, nasal and throat swab for the detection of SARS-CoV-2. medRxiv 2020; preprint [ 10.1101/2020.05.05.20084889]. [DOI] [Google Scholar]
- 27.Guest JL, Sullivan PS, Valentine-Graves M, et al. Suitability and sufficiency of telehealth clinician-observed, participant-collected samples for SARS-CoV-2 testing: the iCollect cohort pilot study. JMIR Public Health Surveill 2020; 6: e19731. doi: 10.2196/19731 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kandel CE, Young M, Serbanescu MA, et al. Detection of severe acute respiratory coronavirus virus 2 (SARS-CoV-2) in outpatients: a multicenter comparison of self-collected saline gargle, oral swab, and combined oral-anterior nasal swab to a provider collected nasopharyngeal swab. Infect Control Hosp Epidemiol 2021; 42: 1340–1344. doi: 10.1017/ice.2021.2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Poukka E, Mäkelä H, Hagberg L, et al. Detection of SARS-CoV-2 infection in gargle, spit, and sputum specimens. Microbiol Spectr 2021; 9: e0003521. doi: 10.1128/Spectrum.00035-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tsang NNY, So HC, Ng KY, et al. Diagnostic performance of different sampling approaches for SARS-CoV-2 RT-PCR testing: a systematic review and meta-analysis. Lancet Infect Dis 2021; 21: 1233–1245. doi: 10.1016/S1473-3099(21)00146-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.See A, Toh ST. Respiratory sampling for severe acute respiratory syndrome coronavirus 2: an overview. Head Neck 2020; 42: 1652–1656. doi: 10.1002/hed.26232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mawaddah A, Genden HS, Lum SG, et al. Upper respiratory tract sampling in COVID-19. Malays J Pathol 2020; 42: 23–35. [PubMed] [Google Scholar]
- 33.McLennan M, Goldfarb D, Donoghue M, et al. Implementing saline gargle sample collection for COVID-19 testing. BC Med J 2020; 62: 384–385. [Google Scholar]
- 34.Tsai CL, Wu PC. Possible beneficial role of throat gargling in the coronavirus disease pandemic. Public Health 2020; 185: 45–46. doi: 10.1016/j.puhe.2020.05.055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mittal A, Gupta A, Kumar S, et al. Gargle lavage as a viable alternative to swab for detection of SARS-CoV-2. Indian J Med Res 2020; 152: 77–81. doi: 10.4103/ijmr.IJMR_2384_19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Michel W, Färber J, Dilas M, et al. A combined oro-nasopharyngeal swab is more sensitive than mouthwash in detecting SARS-CoV-2 by a high-throughput PCR assay. Infection 2021; 49: 527–531. doi: 10.1007/s15010-021-01600-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Saito M, Adachi E, Yamayoshi S, et al. Gargle lavage as a safe and sensitive alternative to swab samples to diagnose COVID-19: a case report in Japan. Clin Infect Dis 2020; 71: 893–894. doi: 10.1093/cid/ciaa377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Whiting PF, Rutjes AW, Westwood ME, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med 2011; 155: 529–536. doi: 10.7326/0003-4819-155-8-201110180-00009 [DOI] [PubMed] [Google Scholar]
- 39.Veroniki AA, Jackson D, Bender R, et al. Methods to calculate uncertainty in the estimated overall effect size from a random-effects meta-analysis. Res Synth Methods 2019; 10: 23–43. doi: 10.1002/jrsm.1319 [DOI] [PubMed] [Google Scholar]
- 40.Thompson SG, Sharp SJ. Explaining heterogeneity in meta-analysis: a comparison of methods. Stat Med 1999; 18: 2693–2708. doi: [DOI] [PubMed] [Google Scholar]
- 41.Langan D, Higgins JPT, Jackson D, et al. A comparison of heterogeneity variance estimators in simulated random-effects meta-analyses. Res Synth Methods 2019; 10: 83–98. doi: 10.1002/jrsm.1316 [DOI] [PubMed] [Google Scholar]
- 42.Miller JJ. The inverse of the Freeman–Tukey double arcsine transformation. Am Stat 1978; 32: 138–138. [Google Scholar]
- 43.Higgins JPT, Thompson SG, Deeks JJ, et al. Measuring inconsistency in meta-analyses. BMJ 2003; 327: 557–560. doi: 10.1136/bmj.327.7414.557 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med 2002; 21: 1539–1558. doi: 10.1002/sim.1186 [DOI] [PubMed] [Google Scholar]
- 45.Higgins JP, Thomas J, Chandler J, et al. Cochrane Handbook for Systematic Reviews of Interventions. Hoboken, John Wiley & Sons, 2019. [Google Scholar]
- 46.Guo W-L, Jiang Q, Ye F, et al. Effect of throat washings on detection of 2019 novel coronavirus. Clin Infect Dis 2020; 71: 1980–1981. doi: 10.1093/cid/ciaa416 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Goldfarb DM, Tilley P, Al-Rawahi GN, et al. Self-collected saline gargle samples as an alternative to health care worker-collected nasopharyngeal swabs for COVID-19 diagnosis in outpatients. J Clin Microbiol 2021; 59: e02427-20. doi: 10.1128/JCM.02427-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gertler M, Krause E, van Loon W, et al. Self-collected oral, nasal and saliva samples yield sensitivity comparable to professionally collected oro-nasopharyngeal swabs in SARS-CoV-2 diagnosis among symptomatic outpatients. Int J Infect Dis 2021; 110: 261–266. doi: 10.1016/j.ijid.2021.07.047 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Olearo F, Nörz D, Hoffman A, et al. Clinical performance and accuracy of a qPCR-based SARS-CoV-2 mass-screening workflow for healthcare-worker surveillance using pooled self-sampled gargling solutions: a cross-sectional study. J Infect 2021; 83: 589–593. doi: 10.1016/j.jinf.2021.08.047 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Benoit P, Labbé AC, Lalancette L, et al. Comparison of SARS-CoV-2 detection with the Cobas® 6800/8800 system on gargle samples using two sample processing methods with combined oropharyngeal/nasopharyngeal swab. J Med Virol 2021; 93: 6837–6840. doi: 10.1002/jmv.27245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hitzenbichler F, Bauernfeind S, Salzberger B, et al. Comparison of throat washings, nasopharyngeal swabs and oropharyngeal swabs for detection of SARS-CoV-2. Viruses 2021; 13: 653. doi: 10.3390/v13040653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kinshella M-LW, Tilley P, Al-Rawahi GN, et al. Evaluation of observed and unobserved self-collection of saline gargle samples for the detection of SARS-CoV-2 in outpatients. Diagn Microbiol Infect Dis 2021; 102: 115566. doi: 10.1016/j.diagmicrobio.2021.115566 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Biber A, Lev D, Mandelboim M, et al. The role of mouthwash sampling in SARS-CoV-2 diagnosis. Eur J Clin Microbiol Infect Dis 2021; 40: 2199–2206. doi: 10.1007/s10096-021-04320-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zander J, Scholtes S, Ottinger M, et al. Self-collected gargle lavage allows reliable detection of SARS-CoV-2 in an outpatient setting. Microbiol Spectr 2021; 9: e0036121. doi: 10.1128/Spectrum.00361-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kocagoz T, Can O, Yurttutan Uyar N, et al. Simple concentration method enables the use of gargle and mouthwash instead of nasopharyngeal swab sampling for the diagnosis of COVID-19 by PCR. Eur J Clin Microbiol Infect Dis 2021; 40: 2617–2622. doi: 10.1007/s10096-021-04326-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Casati B, Verdi JP, Hempelmann A, et al. Rapid, adaptable and sensitive Cas13-based COVID-19 diagnostics using ADESSO. Nature Comms 2022; 13: 3308. doi: 10.1038/s41467-022-30862-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Dumaresq J, Coutlée F, Dufresne PJ, et al. Natural spring water gargle and direct RT-PCR for the diagnosis of COVID-19 (COVID-SPRING study). J Clin Virol 2021; 144: 104995. doi: 10.1016/j.jcv.2021.104995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kwon J, Ko E, Cho S-Y, et al. Bean extract-based gargle for efficient diagnosis of active COVID-19 infection using rapid antigen tests. Microbiology Spectrum 2022; 10: e01614–e01621. doi: 10.1128/spectrum.01614-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Utama R, Hapsari R, Puspitasari I, et al. Self-collected gargle specimen as a patient-friendly sample collection method for COVID-19 diagnosis in a population context. Sci Rep 2022; 12: 3706. doi: 10.1038/s41598-022-07690-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ehre C. SARS-CoV-2 infection of airway cells. N Engl J Med 2020; 383: 969. doi: 10.1056/NEJMicm2023328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Klebe RJ, Grant GM, Grant AM, et al. RT-PCR without RNA isolation. Biotechniques 1996; 21: 1094–1100. doi: 10.2144/96216rr02 [DOI] [PubMed] [Google Scholar]
- 62.Garnett L, Bello A, Tran KN, et al. Comparison analysis of different swabs and transport mediums suitable for SARS-CoV-2 testing following shortages. J Virol Methods 2020; 285: 113947. doi: 10.1016/j.jviromet.2020.113947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Centers for Disease Control and Prevention . Interim guidelines for collecting and handling of clinical specimens for COVID-19 testing. www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html Date accessed: 25 October 2021. Date last updated: 15 July 2022.
- 64.Dhillon SK, Schelstraete P, Cornelissen L, et al. The accuracy of saliva versus nasopharyngeal and/or oropharyngeal samples for the detection of SARS-CoV-2 in children – A rapid systematic review and meta-analysis. medRxiv 2021; preprint [ 10.1101/2021.06.21.21259284]. [DOI] [Google Scholar]
- 65.Malecki M, Lüsebrink J, Teves S, et al. Pharynx gargle samples are suitable for SARS-CoV-2 diagnostic use and save personal protective equipment and swabs. Infect Control Hosp Epidemiol 2021; 42: 248–249. doi: 10.1017/ice.2020.229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Morikawa S, Hiroi S, Kase T. Detection of respiratory viruses in gargle specimens of healthy children. J Clin Virol 2015; 64: 59–63. doi: 10.1016/j.jcv.2015.01.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Lübke N, Schupp A-K, Bredahl R, et al. Screening for SARS-CoV-2 infections in daycare facilities for children in a large city in Germany. medRxiv 2021; preprint [ 10.1101/2021.02.26.21252510]. [DOI] [Google Scholar]
- 68.Larremore DB, Wilder B, Lester E, et al. Test sensitivity is secondary to frequency and turnaround time for COVID-19 screening. Sci Adv 2021; 7: eabd5393. doi: 10.1126/sciadv.abd5393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Mina MJ, Parker R, Larremore DB. Rethinking Covid-19 test sensitivity – a strategy for containment. N Engl J Med 2020; 383: e120. doi: 10.1056/NEJMp2025631 [DOI] [PubMed] [Google Scholar]
- 70.Oran DP, Topol EJ. Prevalence of asymptomatic SARS-CoV-2 infection: a narrative review. Ann Intern Med 2020; 173: 362–367. doi: 10.7326/M20-3012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.He X, Lau EH, Wu P, et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med 2020; 26: 672–675. doi: 10.1038/s41591-020-0869-5 [DOI] [PubMed] [Google Scholar]
- 72.Wölfel R, Corman VM, Guggemos W, et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020; 581: 465–469. doi: 10.1038/s41586-020-2196-x [DOI] [PubMed] [Google Scholar]
- 73.Bosworth A, Whalley C, Poxon C, et al. Rapid implementation and validation of a cold-chain free SARS-CoV-2 diagnostic testing workflow to support surge capacity. J Clin Virol 2020; 128: 104469. doi: 10.1016/j.jcv.2020.104469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Yelagandula R, Bykov A, Vogt A, et al. SARSeq, a robust and highly multiplexed NGS assay for parallel detection of SARS-CoV2 and other respiratory infections. medRxiv 2020; preprint [ 10.1101/2020.10.28.20217778]. [DOI] [Google Scholar]
- 75.Lai CK, Lui GC, Chen Z, et al. Comparison of self-collected mouth gargle with deep-throat saliva samples for the diagnosis of COVID-19: mouth gargle for diagnosis of COVID-19. J Infect 2021; 83: 496–522. doi: 10.1016/j.jinf.2021.07.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Vogels CBF, Watkins AE, Harden CA, et al. SalivaDirect: a simplified and flexible platform to enhance SARS-CoV-2 testing capacity. Med 2021; 2: 263–280.e266. doi: 10.1016/j.medj.2020.12.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Sweeney-Reed CM, Wolff D, Hörnschemeyer S, et al. Feasibility of a surveillance programme based on gargle samples and pool testing to prevent SARS-CoV-2 outbreaks in schools. Sci Rep 2021; 11: 19521. doi: 10.1038/s41598-021-98849-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Fisher B, Seese L, Sultan I, et al. The importance of repeat testing in detecting coronavirus disease 2019 (COVID-19) in a coronary artery bypass grafting patient. J Card Surg 2020; 35: 1342–1344. doi: 10.1111/jocs.14604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Koirala A, Wood N, Macartney K. Testing for SARS-CoV-2 infection: a key strategy to keeping schools and universities open. Lancet Child Adolesc Health 2021; 5: 387–389. doi: 10.1016/S2352-4642(21)00087-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Willeit P, Krause R, Lamprecht B, et al. Prevalence of RT-qPCR-detected SARS-CoV-2 infection at schools: first results from the Austrian School-SARS-CoV-2 prospective cohort study. Lancet Reg Health Eur 2021; 5: 100086. doi: 10.1016/j.lanepe.2021.100086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Maricic T, Nickel O, Aximu-Petri A, et al. A direct RT-qPCR approach to test large numbers of individuals for SARS-CoV-2. PLoS One 2020; 15: e0244824. doi: 10.1371/journal.pone.0244824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Gadkar VJ, Goldfarb DM, Young V, et al. Gargle-Direct: extraction-free detection of SARS-CoV-2 using real-time PCR (RT-qPCR) of saline gargle rinse samples. medRxiv 2020; preprint [ 10.1101/2020.10.09.20203430]. [DOI] [Google Scholar]
- 83.Kellner MJ, Ross JJ, Schnabl J, et al. A rapid, highly sensitive and open-access SARS-CoV-2 detection assay for laboratory and home testing. Front Mol Biosci 2022; 9: 801309. doi: 10.3389/fmolb.2022.801309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Song F, Khan KS, Dinnes J, et al. Asymmetric funnel plots and publication bias in meta-analyses of diagnostic accuracy. Int J Epidemiol 2002; 31: 88–95. doi: 10.1093/ije/31.1.88 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Please note: supplementary material is not edited by the Editorial Office, and is uploaded as it has been supplied by the author.
Supplementary material ERR-0014-2022.SUPPLEMENT (411.9KB, pdf)