Abstract
The white spot disease causes significant damage to global aquaculture production. A prominent vaccine, eliciting the immunogenicity of freshwater fishes against Ichthyophthirius multifiliis yet to be developed. Thus, an Immunoinformatic drive was implemented to find out the potential epitopes from the surface immobilization antigens. B-cell derived T-cell epitopes are promiscuous elements for new generation peptide-based vaccine designing. A total of eight common B and T-cell epitopes had filtered out with no overlapping manner. Subsequently, the common epitopes are linked up with EAAAKEAAAKEAAAK linker peptides, we also added L7/L12 ribosomal protein adjuvant at the N- terminal side of peptide sequence for eliciting the immune response in a better way. The secondary and tertiary structural properties of the modeled 3D protein revealed that the protein had all the properties required for a protective immunogen. Afterward, three globally used validation server: PROCKECK, ProSA and ERRAT were used to justify the proper coordinate. NMR, Crystallographic range and error plot calculation for vaccine model also been done respectively. This was followed by molecular docking, MD simulation, NMA analysis, in silico cloning and vaccine dose-based immune response simulation to evaluate the immunogenic potency of the vaccine construct. The in silico immune simulation in response to multi-epitopes show antibody generation and elevated levels of cell-mediated immunity during repeated exposure of the vaccine. The favourable results of the in silico analysis significantly specify that the vaccine construct is really a powerful vaccine candidate and ready to proceed to the next steps of experimental validation and efficacy studies.
Graphical Abstract
Supplementary Information
The online version contains supplementary material available at 10.1007/s10989-022-10475-1.
Keywords: Spot disease, Immunoinformatics, MD simulation, NMA, B-cell, T-cell, Docking
Introduction
Aquarium fishes are widely kept as a pet and important source of fish trade communities for its gleaming aesthetic significance (Andrews 1990). These are mostly reared in the fish aquariums (Basu et al. 2012). However, pathogenic microorganisms cause harm to aquarium fishes and many fresh water fishes. Among the lots of pathogenic disease, ‘White spot disease’ play a harsh role in fresh water aquaculture and fish development (Koppang et al. 2015). Aquarium fishes like goldfish (Carassius auratus), Siamese shark (Pangasius sutchi), black angel (Pterophyllum scalare) and gold gourami (Trichopodus trichopterus) are highly susceptible (ABDULLAH-AL MAMUN, M., et al. 2021; Elsayed, et al. 2006). Culture fishes like Grass carp (Ctenopharyngodon idella), Cyprinus carpio, and catfish (Pangasianodon hypophthalmus) (Zhao, et al. 2013; Mamun, et al. 1878) are also affected.
Ichthyophthirius multifiliis, a holotrichous ciliate protozoan parasite, which is main causative agent of ‘White spot disease’ or Ichthyophthiriasis of fresh water fishes and aquarium fishes (Dickerson and Findly 2014; Stoskopf 2015; Matthews 2004; Trujillo-González et al. 2018). This parasite invades into the fish body through epithelial tissue of the skin and gills tissue where it create infections in host (Clark et al. 1988). I. multifiliis belongs to Ichthyophthiriidae family known for parasitic disease causing protozoans (Dickerson et al. 2011; Dickerson and Dawe 2006). The immobilization antigens are antigenic element that localized in surface membrane of I. multifiliis and predominantly targets the host immune responses (XU, C. et al. 1995; Wang et al. 2002).
Fish are very interesting vertebrate phylum that possesses both adaptive and innate immune system, play a great role on boosting immunity against pathogen (Magnadottir 2010). Therefore, it is easy to develop a predominant vaccine candidate that might be used against the pathogenic infections. It focuses on the design and study of vaccine candidate construction along with using different algorithms for mapping potential B cell and T-cell epitopes.
This research work focuses on computer-aided vaccine designing techniques, also known as an ‘Immunoinformatic’ approach. The method utilizes several prediction software and server to identify the specific antigenic epitopes within the targeted protein region. This approach takes account of various methodology that adopted to recognize and characterize T and B-cell epitopes for designing promiscuous multi epitope based peptide (MEBP) vaccine against protozoan parasite Ichthyophthirius multifiliis. This technique is very much easy, reliable and less expensive but effective one. MEBP vaccines are proved to be effective against cancer, filarial disease, multi-drug resistance pathogens, Tuberculosis, COVID-19, malaria and many more. These vaccines furnishes better options to provide immunogenicity, reproducibility and experimental control over antigenic protein of pathogens (Salaikumaran, et al. 2022). In the modern science, proteomic data of different pathogen which is easily and widely available in globally used software and server came into play.
Despite these advantages this Immunoinformatic technique has some limitation too; it requires highly skilled professionals to practice properly. It is widely dependent on available databases but sometimes those data found to be limited to apply this technique. The technique requires several in vitro and in vivo validation trial before implementation (Ishack and Lipner 2021).
Materials and Methods
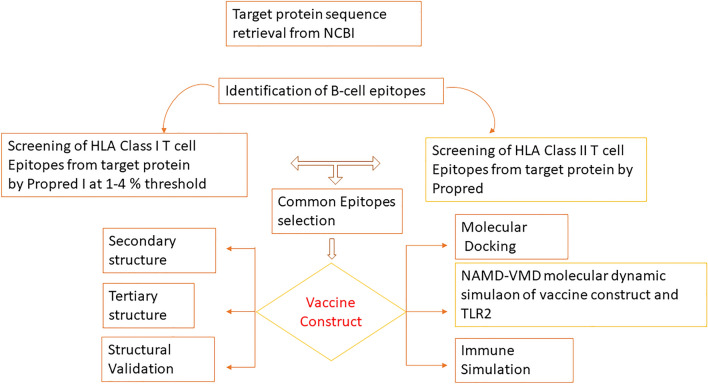
A summarized steps of the designing multi-epitope vaccine is presented as flowchart shown in Fig. 1.
Fig. 1.
Schematic representation of the workflow for the development of multi-epitope vaccine against parasite Ichthyophthirius multifiliis
Retrieval of Amino Acid Sequences of the Target Protein
The primary amino acid sequences of immobilization antigen (I-antigen) retrieved from National Center for Biotechnology Information database (https://www.ncbi.nlm.nih.gov/protein/) (Coordinators 2016). The retrieved amino acid sequences of targeted protein were used to process in computer-based server and software for designing vaccine candidate against protozoan pathogen I. multifiliis.
B-Cell Epitope Identification
The Immune Epitope Database (IEDB) was used for the prediction of B cell epitopes in the protein sequences. In this current work, we used BepiPred 2.0 prediction method integrated in IEDB database (Jespersen, et al. 2017; Kim et al. 2012). B-cell prediction is one of the crucial steps for epitopic vaccine development. B-cell epitopes stimulate host humoral immune system.
T-Cell Epitope Identification
T-cell epitopes had predicted within the B-cell epitope for enhancing the immunity against pathogenic infections. In this work, identified T-cell epitopes have the affinity towards the multiple HLA alleles. An antigen-presenting cell (APC) represents T-cell epitopes bound to major histocompatibility molecules (MHC) for boosting immune response (Patronov and Doytchinova 2013). MHC-II and MHC-I binding region in antigenic protein sequence were predicted though Propred and Propred-I web server respectively (Singh and Raghava 2001, 2003).
Multi-Epitope Subunit Vaccine Construction and Modeling
The subunit vaccine constructed by joining of the selected proposed antigenic epitope using suitable linker peptide. The subunit vaccine has played an effective role in activating both innate and adaptive immune response (Dhal et al. 2019). Here, we were using EAAAKEAAAKEAAAK linker peptides to link the epitopes, which has self-regulating immunogen as well as it will be elicit immune response (Pentel and LeSage 2014). This linker can increase the stability and folding pattern of vaccine candidate. Subsequently, tertiary structure of a protein having a promising role to regulate its functionality in sub-cellular manner (Roy et al. 2012). Therefore, here we generated 3D structure of vaccine component with help of SPARKS-X server (Yang et al. 2011).
Analysis of the Physicochemical and Secondary Structural Properties, Allergenicity of the Vaccine Model
The physicochemical properties of vaccine model analyzed through the Expasy Protparam tool (https://web.expasy.org/protparam/). Additionally, SOPMA; tool used for predicting Secondary structural properties of the targeted protein (Geourjon and Deleage 1995). The Expasy, Protparam server predicts the Grand Average Hydropathy (GRAVY), atomic composition, molecular weight, extinction coefficients, estimated half-life, aliphatic index and Instability index. Next, we selected Protein-Sol web server for testing of solubility of vaccine candidate in aqueous solution (Hebditch, et al. 2017).
In order to evaluate the safety measure and effectiveness of the vaccine candidate here we used AllerTOP v.2.0 and AllergenFP v.1.0 (Dimitrov, et al. 2014a, 2014b). Both servers are used auto-cross covariance (ACC) transformation algorithm for allergic prediction.
Justification of Vaccine Model
In this section, we discussed about the subsequent validation of vaccine component that was modeled via SPARKS-X. Evaluation of structural properties of vaccine candidate had carried out by Ramachandran Plot through PROCHECK server (Laskowski et al. 2006). Ramachandran plot assesses the model quality, compute with phi–psi torsion angles for each amino acid residue. Finally, those residues are categorized as favored, generously allowed and disallowed regions. Conversely, the protein folding energy was assessed by using ProSA web-server (Wiederstein and Sippl 2007). ProSA delivered us ‘Z’ score value that specifies the overall model quality. Lastly, the protein model was passed through ERRAT web server for further validation. It calculates the statistical value of non-bonded interactions between different atoms and plots different value of the error function versus position of a 9-residue sliding window, considered by a comparison with statistics from highly refined structures (Colovos and Yeates 1993).
Antigenicity Prediction of the Vaccine Element
Antigenicity of constructed vaccine component is the most significant criteria for efficient vaccine designing. The immunological studies reveal, high antigenic score value would encompasses better immunization. Here, we used VaxiJen v.2.0 protective antigen prediction server for calculating antigenic score of vaccine component (Doytchinova and Flower 2007). This server needs amino acid sequences as an input and selected the parasite as a field with the prescribed threshold value of 0.5 and the prediction accuracy of 87% (Gededzha et al. 2014). VaxiJen v2.0 permits classification of antigen depicted on the physicochemical properties of proteins.
Molecular Docking
Protein-peptide interactions regulate the several cellular processes including signalling pathways as well as regulate various morphogenic pathways (Zhong et al. 2007). Therefore, Molecular docking is a prime process for supporting the drug designing and eventually discover their cellular interactions (Pagadala et al. 2017). Here, we used PatchDock server for performing molecular docking analysis (Schneidman-Duhovny, et al. 2005). This docking server used image segmentation and object recognition technique for carrying out better computer based visioning (Pradhan and Sharma 2014). PatchDock server needs two PDB file (one for ligand and another for receptor) as an input and Clustering RMSD had selected as 4.0 Å as default.
Molecular Dynamics Simulation
Molecular Dynamics simulation is a key aspect to compute the structural stability of the protein-peptide complex (Aalten et al. 1997; Hasan, et al. 2020). Complexes of vaccine construct with TLR2 were simulated at 1200 Picoseconds (ps) time scale using NAMD 2.14 by following the energy minimization with NVT (Phillips et al. 2005). The trajectories were saved for each complex after every 2 fs and root mean square deviation (RMSD) and root mean square fluctuations (RMSF) analysis were performed using VMD 1.9.3 tools (Humphrey et al. 1996).
Immune Simulation
An in silico immune simulation was accomplished through C-IMMSIM server (https://150.146.2.1/C-IMMSIM/index.php), to validate immunological response of designing vaccine construct (Rapin et al. 2010). This server simulates mainly primary and secondary immune responses. The vaccine candidate has tested for the ability for immune response against various types of immune cells like HTL, CTL, B-cells, NK cells, dendritic cells, Immunoglobulin and cytokines. Clinical recommendation of minimum interval between two doses of vaccine is four weeks (Castiglione, et al. 2012). In this experiment, we administered three injections at a standard time period gap; using C-ImmSim immuno stimulatory server, with the recommended interval of four weeks (1, 84 and 168 time-steps parameters were set as 1 time-step is equal to eight hours of real life) for a total of 1025 steps of simulation. Other parameters were kept as default for better result.
NMA Analysis
Molecular dynamics (MD) motion is key method for analyzing the physical movements of atoms and molecules. It also demonstrates the stability of protein–protein complex. Herein, we run the iMODS server to elucidate the collective motion of protein complex via analysis of normal modes (NMA) in its internal coordinates (López-Blanco, et al. 2014). This server predicted the direction and extended of the innate motions of the protein complex in the forms of B-factors, eigen values, deformability, Variance and covariance map (Ghosh 2021). In order to described the motion stiffness of normal mode the eigen value assessed its stiffness (Saha et al. 2021). In the NMA dihedral coordinates, it naturally mimics the combined functional motions of modeled protein molecules as a set of atoms connected by harmonic springs.
Codon Optimization and In-silico Cloning
Very often, the codon uses varies in target and host species so codon adaptation is implemented to increase the translation efficiency of cloned genes within the host. Codon optimization of vaccine construct had accomplished via online web server called Java Codon Adaptation Tool (JCAT) (http://www.jcat.de/) for high-level expression of the vaccine sequence in E. coli K12 strain (Grote, et al. 2005). We had taken pET28a (+) expression vector from “addgene” vector database (Kamens 2015) to implement in silico cloning of vaccine sequence. SnapGene 5.1.7 restriction cloning software has implemented for finalizing the in silico cloning (Biotech, G.J.U.s.c. 2015).
Result
Retrieval of Amino Acid Sequences of the Target Protein
Primary amino acid sequence of I-antigen of I. multifiliis had retrieved from NCBI with the GenBank accession ID: ACH87654.2. The I-antigen has contained 452 amino acid sequences and downloaded those amino acid sequences as FASTA format.
B-cell Epitope Identification
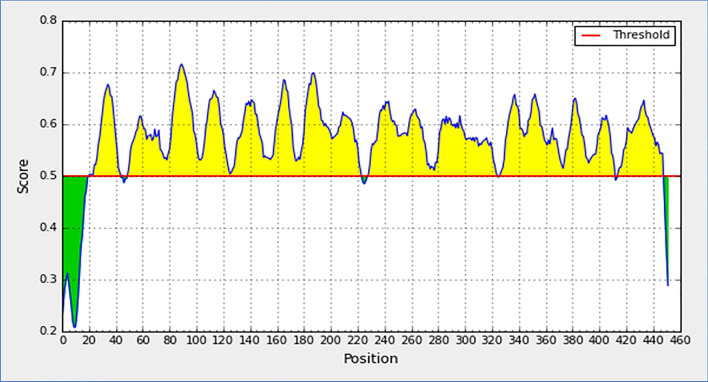
We founded a total six linear sequential B-cell epitopes along with various lengths. B-cell epitopes within I-antigen of I. multifiliis could played a robust role in provoking immunity. Here, following B-cell epitopes were positioned into Table 1 with their positional value, amino acid sequence, and length. The epitopes are graphically symbolized with the remarkable yellow color, and green color peaks represents non-epitopes one showing in Fig. 2.
Table 1.
Selected B-cell epitopes from IEDB and BepiPred 2.0 prediction method
| Sl no | Start point | End point | Amino acid sequence | Length |
|---|---|---|---|---|
| 1 | 1 | 44 | MKFNILIILIISLFINELRAVNCPNGAAIANGQSDTGAADINTC | 44 |
| 2 | 46 | 46 | H | 1 |
| 3 | 50 | 223 | HFYFNGGNPAGQAPGAGQFNPGVSQCIACQVHKADSQHRQGGDANLAAQCSNLCPAGTAVEDGSPTFTQSLTQCVNCKPNFYFNGGNPTGQAPGAGQFDPTQLIANPDLANNPEVPNVSSPNGQCVACQVNKSDSQLRPGAQANLATQCNNECPTGTAIQDGAIFIYTQSISQC | 174 |
| 4 | 229 | 324 | DFYFNGGNPSAQNPGNGQFTPGQLIANPDAATSAQIPMVPGPNSKCVACESKKTNSQSRSGLEANLAAQCGTECPAGTLVTDGVTPTYTVSLSQCV | 96 |
| 5 | 327 | 412 | KAGFYQNSNFEAGKSQCNKCAVSKTGSASVPGNSATSATQCQNDCPAGTVVDDGTSTNFVALASECTKCQANFYASKTSGFAAGTD | 86 |
| 6 | 415 | 452 | TECSKKLTSGATAKVYAEATQKAQCASSTFAKFLSMSL | 34 |
Fig. 2.
The threshold level (antigenicity) of top B-cell linear epitopes
T-Cell Epitope Identification
Surface-exposed B-cell epitopes of the prioritized proteins with high antigenicity were taken into consideration to predict B-cell derived T-cell epitope. T-cell epitopes prediction was based on the binding of epitopes to both MHC-I and MHC-II molecules. Here we filtered out common B and T cell epitopes that mostly binding with MHC alleles tabulated in Table 2 and graphically reflects in Fig. 3 for better visualization.
Table 2.
Selected T-cell epitopes common for both MHC molecules predicted from B-cell epitopic regions
| Sl No | Sequence | MHC-I allele | MHC-II allele | Position |
|---|---|---|---|---|
| 1 | FNILIILII |
HLA-A2.1 HLA-B*5101 HLA-B*5102 HLA-B*5201 HLA-B*5301 HLA-B*5401 HLA-B*51 HLA-Cw*0602 MHC-Db MHC-Kd MHC-Kk |
DRB1_0101 DRB1_0102 DRB1_0401 |
3–11 |
| 2 | FYFNGGNPA |
HLA-B*5301 HLA-B*5401 HLA-B*51 HLA-Cw*0401 MHC-Kd |
DRB1_0101 DRB1_0102 DRB1_0401 DRB1_0426 |
51–59 |
| 3 | FTQSLTQCV |
HLA-A*0201 HLA-A2.1 HLA-B*2702 HLA-B*3501 HLA-B*5301 HLA-B*51 HLA-B*5801 HLA-B61 MHC-Db revised MHC-Ld |
DRB1_0401 DRB1_0421 DRB1_0426 DRB1_0701 DRB1_0703 |
116–124 |
| 4 | FIYTQSISQ |
HLA-A3 HLA-B*2702 HLA-B*2705 HLA-B*5301 HLA-B*5401 HLA-B*51 HLA-B*5801 HLA-B62 MHC-Kb |
DRB1_0305 DRB1_0401 DRB1_0402 DRB1_0405 DRB1_0408 DRB1_0701 DRB1_0703 DRB1_0804 DRB1_1101 DRB1_1307 DRB1_1321 DRB5_0101 DRB5_0105 DRB1_0817 |
214–222 |
| 5 | MVPGPNSKC |
HLA-A*0205 HLA-A3 HLA-B*3501 HLA-B*5101 HLA-B*5102 HLA-B*5103 HLA-B*51 HLA-B61 HLA-B7 HLA-B*0702 MHC-Ld |
DRB1_0306 DRB1_0307 DRB1_0308 DRB1_0311 DRB1_0402 DRB1_0404 DRB1_0423 DRB1_1107 DRB1_1501 DRB1_1506 |
266–274 |
| 6 | VVDDGTSTN |
HLA-B*3701 HLA-B*4403 HLA-B61 MHC-Kk |
DRB1_0301 DRB1_0306 DRB1_0307 DRB1_0308 DRB1_0311 DRB1_0410 DRB1_1107 DRB1_0401 DRB1_0421 |
376–384 |
| 7 | FAKFLSMSL |
HLA-B*5101 HLA-B*5102 HLA-B*5103 HLA-B*5301 HLA-B*5401 HLA-B*51 HLA-B*5801 HLA-B60 HLA-B7 HLA-B*0702 HLA-B8 |
DRB1_0308 DRB1_0311 DRB1_0410 DRB1_1107 DRB1_0401 |
444–452 |
| 8 | FYFNGGNPS |
HLA-B*5401 HLA-B*51 HLA-Cw*0401 MHC-Kd |
DRB1_0101 DRB1_0305 DRB1_0401 DRB1_0421 DRB1_0426 DRB1_0813 DRB1_1114 DRB1_1120 DRB1_1302 DRB1_1323 |
230–238 |
Fig. 3.
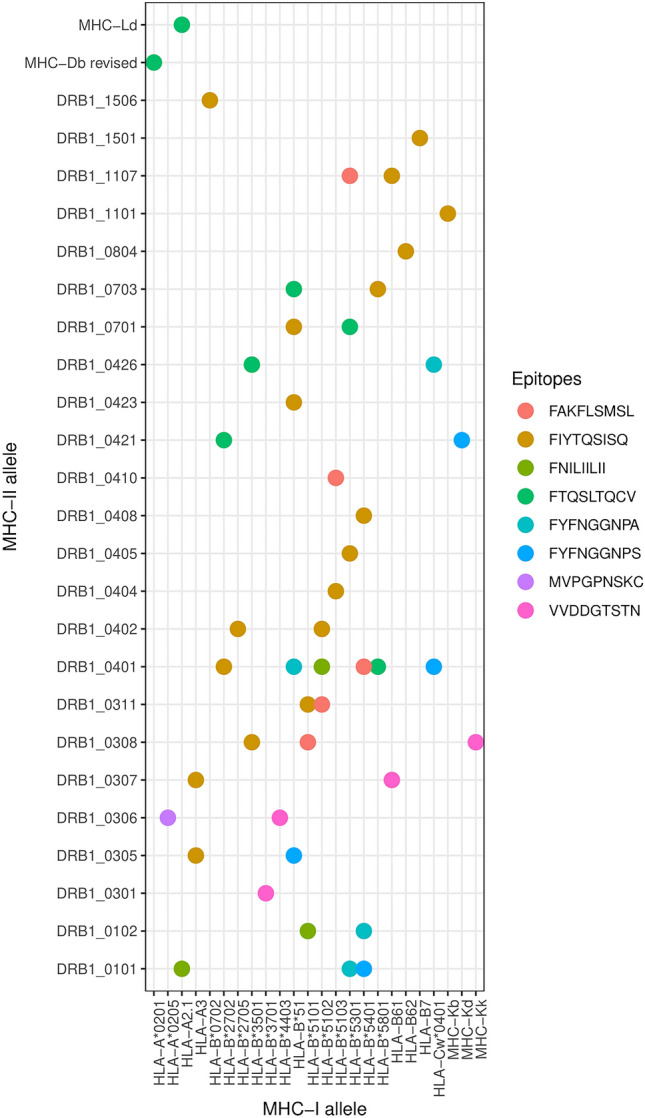
Graphical representation of T − cell epitopes along with binding alleles
Multi-Epitope Subunit Vaccine Construction and Modeling
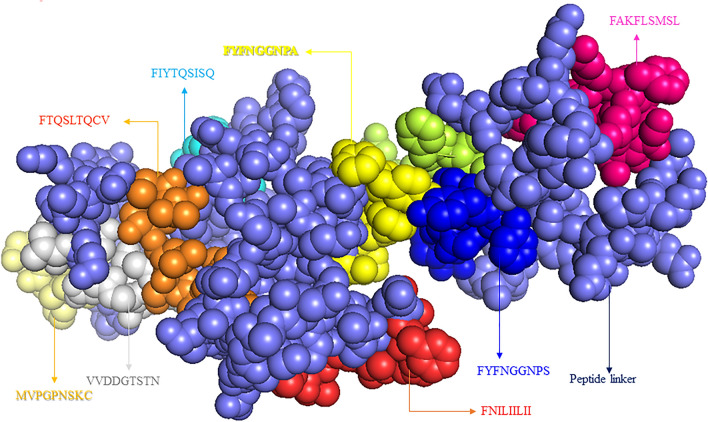
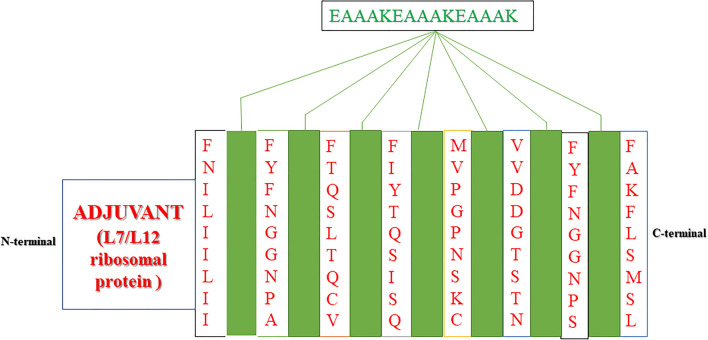
Multi-epitopic peptide-based vaccine comprises antigenic component of a pathogenic organism to induce an immunogenic reaction in the host body. In this study, the predicted T cell within B cell epitopes were combined in a sequential manner to construct the final vaccine candidate and server generated 3D structure Vaccine model showing in the Fig. 4. Sequentially, final vaccine construct has been associated with adjuvant L7/L12 ribosomal protein at the N-terminal side of vaccine model Fig. 5. Interactions of adjuvant with goldfish (Carassius auratus) toll like receptor-2 (TLR2) and common vaccine epitopes stimulate robust immune-reaction.
Fig. 4.
The tertiary structure of the designed vaccine construct
Fig. 5.
Structural arrangement of the final vaccine construct
Analysis of the Physicochemical and Secondary Structural Properties, Allergenicity of the Vaccine Protein
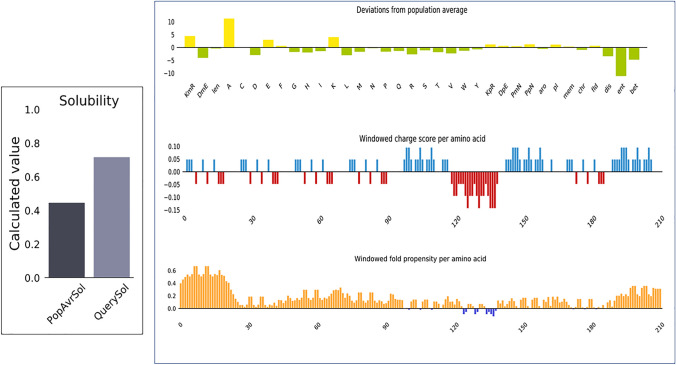
In this subsection, we assumed the physicochemical properties of vaccine construct using Expasy Protparam server, calculating various values of incorporated parameters. The Molecular weight of vaccine model is 23055.04 Dalton, extinction coefficients values are 6085 M−1 cm−1(assuming all pairs of Cys residues form cystines) and 5960 M−1 cm−1(assuming all Cys residues are reduced), instability index (II) value 26.87, estimated half-life was 30 h, the value of aliphatic index is 66.80 and GRAVY value of I-antigen of I. multifiliis was − 0.140 (Table 3). As well as Protein-Sol server calculated the soluble capacity of vaccine candidate, and the results show that the vaccine candidate soluble with water (Fig. 6).
Table 3.
Antigenicity, allergenicity, solubility, and physicochemical property assessments of the primary sequence of multi-epitope-based vaccine construct
| Sl. No | Features | Assessment |
|---|---|---|
| 1 | Antigenicity | 0.5914 (probable ANTIGEN) |
| 2 | Allergenicity |
Probable non-allergen (AllerTOP v.2.0) Probable non-allergen (AllergenFP v.1.0) |
| 3 | Solubility | 0.717 (Soluble) |
| 4 | Number of amino acids | 231 |
| 5 | Molecular weight | 23055.04 Dalton |
| 6 | Theoretical isoelectric point (pI) | 7.83 |
| 7 | Total number of atoms | 3251 |
| 8 | Formula | C1012H1631N277O326S5 |
| 9 | Estimated half-life |
1 h (mammalian reticulocytes, in vitro) > 30 min (yeast, in vivo) > 10 h (Escherichia coli, in vivo) |
| 10 | Instability index | 26.87 (Stable) |
| 11 | Aliphatic index | 66.80 |
| 12 | Grand average of hydropathicity (GRAVY) | − 0.140 |
Fig. 6.
Solubility index of Vaccine construct showing in the plot
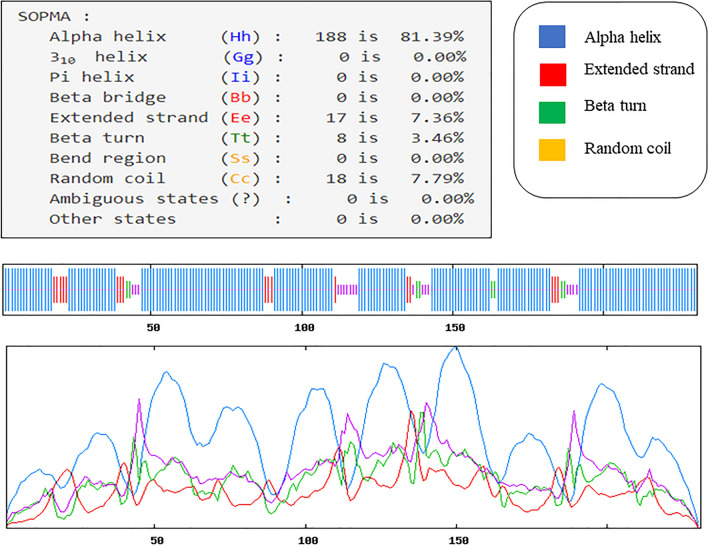
From the SOPMA; we were getting the secondary structural values like the alpha helix, extended strand, beta turn and random coil were calculating 81.39%, 7.36%, 3.46%, and 7.79%, respectively depicted in the Fig. 7. This server also calculates window width value that was 17, Similarity threshold value-8 and numbers of states was four.
Fig. 7.
Probability score graph of occurrence of helix (Purple), strand (Green), turn (Red), and coil (Light blue) at each amino acid position in the secondary structure of the final Vaccine construct. Each residue position is characterized by the greater probability score associated secondary structure
Both two servers predicted that the vaccine showed non-allergenic in nature and it’s safe in respect of allergen (Table 3).
Justification of Vaccine Model
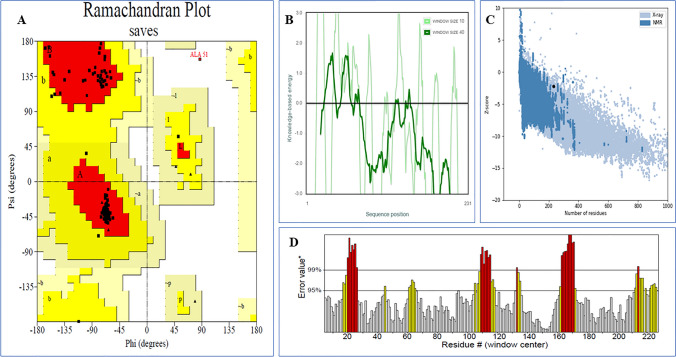
The modeled vaccine candidate had justified and validated through ERRAT, PROCHECK and ProSA web server. Here, the distribution of amino acid showed within Ramachandran map and exposed after homology modeling processes in order to validate the 3D vaccine protein model Fig. 8A. The Ramachandran plot of vaccine protein model revealed that 96.8% of residues lies in the most favorable zones, 2.8% of residues lies in allowed zones and only 0.5%in disallowed areas and statistical data plotted in the Table 4. ProSA-web and ERRAT confirmed the quality, energy value and potential errors in a crude 3D model respectively. ERRAT calculates 75.336% of quality factor that evaluates modeled protein’s overall quality reflects in Fig. 8D. Though the ProSA-web exposed a Z-score of − 0.93, Fig. 8C, for the query 3D model of vaccine, which lies outside the range of scores of NMR and lies between X-ray value commonly found in comparable size native Proteins. In the Fig. 8B represented the local energy model that have another significant sign for model quality assessment.
Fig. 8.
Validation of the tertiary structure of the vaccine. A The Ramachandran plot statistics represent the most favorable, accepted, and disallowed regions with a percentage of 96.8%, 2.8%, and 0.5%, respectively, B Local quality assessment plot C The ProSA-web representing the Z-score of − 0.93 for the vaccine model. D ERRAT error plot showing percentage of error of the vaccine model protein
Table 4.
Distribution of amino acid residues showing in Ramachandran plot
| Type of amino acid residue(s) | No. of amino acid residues | Percentage |
|---|---|---|
| Most favoured region [A, B, L] | 211 | 96.8% |
| Additional allowed region [a, b, l, p] | 6 | 2.8% |
| Generously allowed region [~a, ~b, ~l, ~p] | 0 | 0.0% |
| Disallowed region | 1 | 0.5% |
| Total Number of non-glycine and non-proline residues | 218 | 100% |
| End-residues except glycine and proline | 2 | – |
| Glycine residues | 7 | – |
| Proline residues | 4 | – |
| Total number of residues | 231 | |
Antigenicity Prediction of the Vaccine Element
The VaxiJen server predicted the vaccine candidate has the antigenic property. The antigenic score for the vaccine candidate is 0.5914. This score crosses the threshold value and confirms antigenic property of vaccine component. We concluded the significant antigenic propensity from the antigenic value.
Molecular Docking
The molecular interaction between TLR2 and the designed vaccine candidate was studied by molecular docking method. The server predicted the complex structure based on complementary score, ACE (Atomic Contact Energy) and estimated interface area of the complex protein (Supplementary table-1). Among the first 20 results, only the top rank complex model depending on ACE value had taken. The ACE of the selected docking complex was -160.05 kcal/mol, the negative value indicates spontaneous molecular affinity between the putative vaccine molecules and immune receptors TLR2. The result showed that vaccine construct interacted with TLR2 receptor with significantly lower binding energy Fig. 9.
Fig. 9.
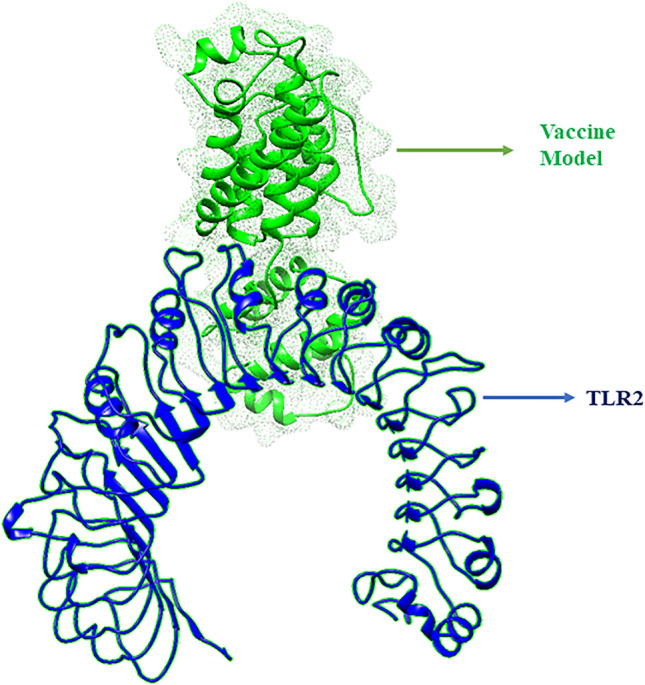
Molecular docking between the vaccine and the TLR-2 receptor
Molecular Dynamics Simulation
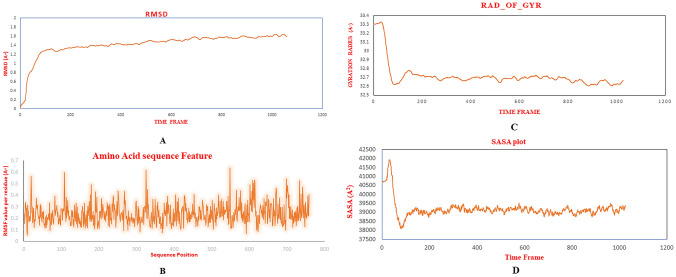
The molecular dynamics simulation was studied on the basis of Root mean square fluctuation (RMSF), Root mean square deviation (RMSD), Radius of gyration (Rg) and Solvent-accessible surface area (SASA) values present as a fraction of Picoseconds (Aspects of time) in Fig. 10A–D respectively. The conformational changes of the complex structure was calculated by RMSD values with the range of 0–1200 ps. The RMSD values steadily increased from 0 to 200 ps, and reached stable state throughout the simulation.
Fig. 10.
Molecular dynamics simulation A Root mean square deviation (RMSD) B root mean square fluctuation (RMSF) analysis of protein backbone and side chain residues of MD simulated vaccine construct C Radius of Gyration (Rg) plot in during MD simulation and D SASA plot
NMA Assay
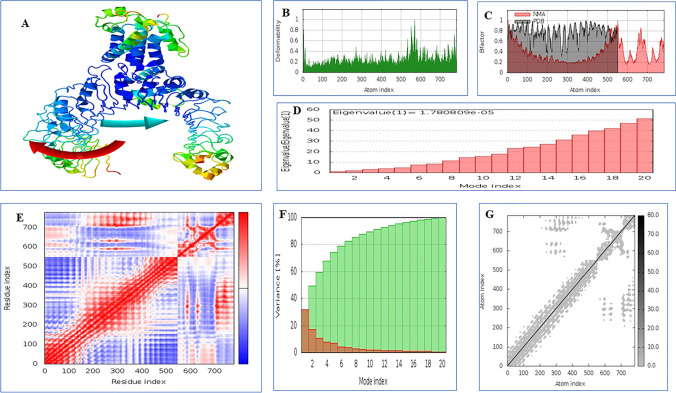
Vaccine construct stability investigated through NMA mobility analysis (Fig. 11A), deformability analysis, B-factor (Fig. 11C), eigen value analysis, covariance map, variance map and elastic network plot. Results revealed that the placements of hinges in the chain was insignificant (Fig. 11B) and the B-factor column gave an averaged RMS (Fig. 11C). The estimated higher eigen value 1.780809e-06 (Fig. 11D) indicated low chance of deformation of vaccine candidate. The correlation matrix and elasticity of the vaccine candidate was shown in Fig. 11E and G, respectively. The variance associated to each normal mode is inversely related to the eigen value. Colored bars show the individual (red) and cumulative (green) variances showing in Fig. 11F.
Fig. 11.
Molecular dynamics analysis of vaccine protein-TLR2 complex; stability of the protein–protein complex was investigated through A NMA mobility showing with arrows B B-factor values C deformability plot D eigen value E covariance of residue index F Variance map and G elastic network analysis
Immune Simulation
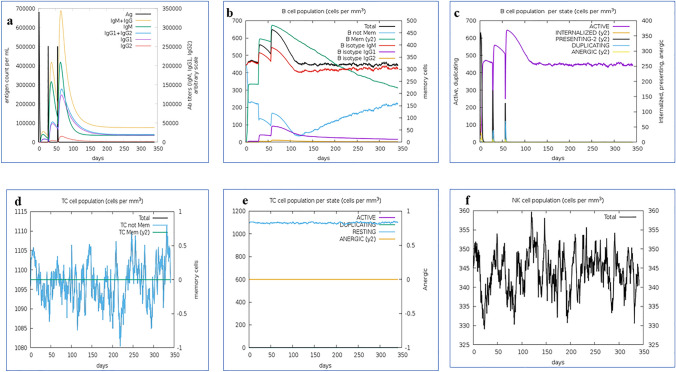
The in silico simulated immune response of the vaccine construct was eliciting high primary and secondary immune responses by triggering the immune system, including CTL, HTL, sustainable memory cells and other cells. A high level of (IgM + IgG) antibodies, as well as other immune cells were drastically rising after administration of vaccine and primary immune cells like IgG and IgM immunoglobulins showed the prolonged effects against pathogens reflects in Fig. 12a. Moreover, the elevation of CTL cells, which reaches a maximum of 1207 cells/mm3 after 10 days of vaccine administration and decreases slowly after 23 days showing in Fig. 12d. Elevated CTL cells subsequently evoked to generate a high number of memory cells presented in Fig. 12e. The innate immunity mainly increased along with B-cell population was found to be increased with B isotype IgM and B-memory cell also be increased up to 600–700 cells/mm3(Fig. 12b–c). Memory cells play a central role to regulate prevention of viral infection/ re-infection through self-memorization upon encountering pathogens. Subsequently, administration of vaccine found to be elevated other central regulators of immune system (cytokines, interleukins, and NK cells) (Fig. 12f–i). These results signify the designed vaccine as a potent candidate to elicit a robust immune response to fight against fish pathogen.
Fig. 12.
In silico immune response simulation of the multi-epitope-based vaccine construct. a Production of immuno globulins upon antigen exposure, b Population of B lymphocytes after three injections, c Population of B-cell per cell state, d cytotoxic T lymphocytes population, e Amount of Cytotoxic T lymphocytes population per state, f Population of Natural Killer cells, g Population of Macrophages, h Population of Dendritic cells, i Concentration of cytokines and interleukins with Simpson index [D]
Codon Optimization and In-silico Cloning
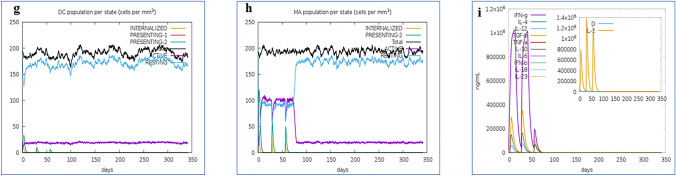
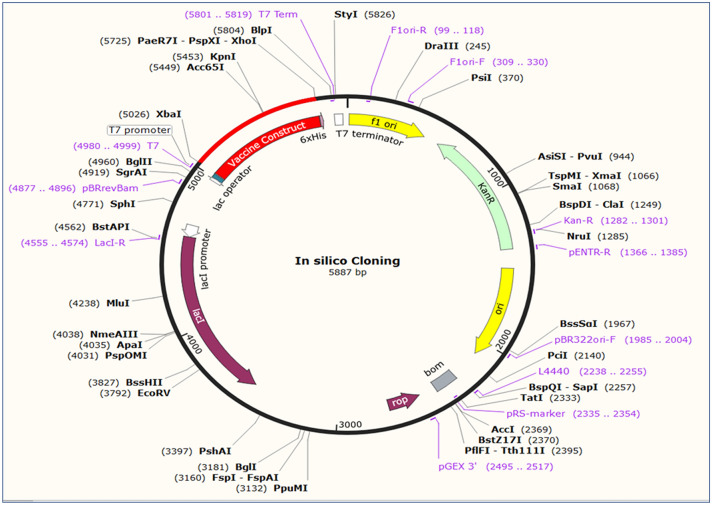
Due to dissimilarity in the regulatory systems of E. coli and fishes, codon adaptation was accomplished within the host to justify its expression. Vaccine candidate was reverse transcribed for the codon adaptation, and the server predicted codon adaptation index (CAI) was 1; ensuring the higher proportion of most abundant adapted codons. The GC content of the optimized codons was 50.73%. Finally, the optimized codon sequence inserted into pET28a ( +) plasmid vector along with XhoI and XbaI restriction sites. A clone of 5887 base pair was produced comprising 778 bp desired sequence and shown in red color in between the sequence of pET28a ( +) vector and the rest belonging to the vector only (Fig. 13).
Fig. 13.
In silico restriction cloning of the multi-epitope vaccine sequence into the pET28a (+) expression vector. The red region represents the vaccine coding gene and the black circle represents the vector backbone
Discussion
Ichthyophthirius multifiliis is a protozoan parasite (an obligate parasite of fish), causing the ‘White spot disease’, major burden for fisherman and aqua culturist globally (Gersdorff Jørgensen 2017). The immobilization antigen are the highly abundant surface protein, mainly observed on a number of holotrich ciliates. At least five different serotypes of this parasite exists, characterized by differences in the surface I-antigen (Dickerson et al. 1993). The protozoan parasite infects (infective theront stage of the parasite invades) the skins and the gills of freshwater fishes (Ewing et al. 1985; Ventura and Paperna 1985). Recently the growing economic status of fish parasitosis for aquaculture sector and as well as fisheries sector has boosted the interest to develop the defense mechanisms against White spot disease. Interestingly, fish has both innate and adaptive immune responses to control protozoan parasite infections (Alvarez-Pellitero 2008). Innate immune response imitates immune recognition controlled by the activation of pathogen recognizing receptors (PRRs) which recognized pathogen-associated molecular patterns (PAMPs) (Medzhitov and Janeway 2002). Fishes have a number of PRRs, mainly Toll-like receptors (TLRs) that mainly activated and relies cascade mechanism through PAMP binding within the receptor (Roach et al. 2005; Purcell et al. 2006).
Hitherto, there is no prophylactic treatment or putative vaccines available while repetitive treatments with supplementary medicines are needed to control the infection. Historically, a large number of chemicals and drugs have been applied to combat White spot disease but due to changing norms and regulations with recognition of carcinogenic effects on environment the most efficient compounds are now prohibited. Researchers continuously search for novel substances like vaccine or non-carcinogenic drugs, which might be highly effective against the parasite as well protozoan organisms and harmless for the fish at the same time. These compounds should be ecofriendly and cost-effective.
Present research design was carried out to develop a multi-epitope based potential vaccine targets against protozoan I. multifiliis the causative agent of White spot disease (Ichthyophthiriasis) which have burden in the global fish culture industry. The multi-epitope-based vaccination is the new approach because of the epitope-based vaccine can generate precise immune responses against pathogens and enhance binding interaction with the target molecules. Now, in this work we selected I-antigen as target protein to design a MEBP vaccine through next generation peptide vaccine formulation methods. For the better response or the dual-purpose response (both humoral and cytotoxic immune response), T-cell epitopes derived from B-cell epitopes were identified in this study. Afterwards, both MHC-I and MHC-II epitopes were predicted successfully by the help of ProPed and ProPed-I server respectively. Next, we have identified common B and T-cell epitope and linked them up with suitable peptide linkers for construction of this vaccine candidate. 50S ribosomal protein L7/L12 adjuvant has been linked at N terminal with proper linker to boost up immunogenicity at cellular level which was used by other researcher for vaccine development.
In this study the 3D structure of vaccine candidate developed through SPARKS-X web server, and it’s validated through ProSA, PROCHECK and ERRAT error plot web-server. Subsequently ‘Z’ score, local energy plot was evaluated to comprehend its proper folding. Ramachandran plot of each amino acid residues of vaccine candidate was studied. Whereas, ProSA predicted ‘Z’ score negative value of − 0.93, indicating a good sign of model quality assessment (MacKerell et al. 2000). Along with the local quality model analysis which indicates its reliable structural quality.
PatchDock automatic web server demonstrated the binding interaction of the design vaccine candidate and TLR2, and calculated ACE score is noted as − 160.05 kcal/mol. The negative ACE value indicates the spontaneous reaction and proper interaction between the protein–peptide complex (Patra, et al. 2019) (Fig. 9). The stability of the vaccine structure was accessed through molecular dynamics simulation with the help of NAMD software package and it is CHARMM topology-based server (MacKerell et al. 2000). With the help of VMD 1.9.3 graphical software we analysed the RMSF, RMSD, radius of gyration and SASA plot of vaccine candidate (Fig. 10). From the result of the RMSD plot we concluded that the binding interaction is stable and flexible with TLR2. Whereas, NMA study performed to analyse the molecular mobility, comparative deformability, and B factor with PDB value. The eigen value was calculated as 1.780809e-06 which indicates the better flexibility of the vaccine candidate with receptor molecule. Finally, the Snap Gene cloning software appointed to amplify the desired vaccine sequence within the pET28b ( +) expression vector.
Conclusion
There is no completely effective vaccine reported against spot disease yet. In this work, a successful attempt was made to design a multi epitope-based protein (MEBP) vaccine against spot disease. Immunoinformatics and reverse vaccinology approaches were used to develop a potential and safe vaccine candidate that could trigger two types of immune responses: humoral and cellular. Our vaccine candidate is highly immunogenic, safe, stable and strongly interacting with fish TLR2 receptors. Here, eight common epitopes (FNILIILII, FYFNGGNPA, FTQSLTQCV, FIYTQSISQ, MVPGPNSKC, VVDDGTSTN, FYFNGGNPS, and FAKFLSMSL) filtered out and linked with stable peptide linker to generate an effective vaccine candidate that predominantly binds with fish TLR2 that facilitate the innate as well as adaptive immune systems and accelerates the immune response against pathogens. It was fascinating that our designed vaccine may able to stimulate neutralizing antibody as well as other central cellular responses well up to 350 days after the last third booster shot as computationally derived. Though our study has some limitations and recommended that the study may performed with in vitro and in vivo techniques to ensure the exact effectiveness.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (DOCX 13 kb) Supplementary table 1: PatchDock predicted docking score (top 20) which predicted the rank based on complementary score, ACE (Atomic Contact Energy) and estimated interface area. (Here bold model has been taken for its high negative ACE value).
Acknowledgements
This research work is supported by the Council of Scientific & Industrial Research (CSIR) sponsored Junior Research Fellowship Program (File No. 09/599(0087)/2019-EMR-I).
Author contributions
PG and PP designed the model of the computational framework, in silico analysis and wrote the manuscript. PG, NM, DSC and BCP carried out the implementation and validations. BCP, NM and DSC helped with the analysis and editing the manuscript.
Data Availability
All data generated or analyzed during this study are included in this published article.
Declarations
Conflict of interest
The authors declare no conflict of interest.
Ethics approval
Not Applicable.
Consent for publication
Not applicable.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- ABDULLAH-AL MAMUN M, et al. Mass infection of Ichthyophthirius multifiliis in two ornamental fish and their control measures. Ann Biol. 2021;37(2):209–214. [Google Scholar]
- Alvarez-Pellitero P. Fish immunity and parasite infections: from innate immunity to immunoprophylactic prospects. Vet Immunol Immunopathol. 2008;126(3–4):171–198. doi: 10.1016/j.vetimm.2008.07.013. [DOI] [PubMed] [Google Scholar]
- Andrews C. The ornamental fish trade and fish conservation. J Fish Biol. 1990;37:53–59. doi: 10.1111/j.1095-8649.1990.tb05020.x. [DOI] [Google Scholar]
- Basu A, Dutta D, Banerjee S. Indigenous ornamental fishes of west Bengal. Recent Res Sci Technol. 2012;4(11):12–21. [Google Scholar]
- Biotech, G.J.U.s.c., SnapGene software [web]. 2015
- Castiglione F, et al. How the interval between prime and boost injection affects the immune response in a computational model of the immune system. Computational Math Method Med. 2012 doi: 10.1155/2012/842329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark TG, Dickerson HW, Findly RC. Immune response of channel catfish to ciliary antigens of Ichthyophthirius multifiliis. Dev Comp Immunol. 1988;12(3):581–594. doi: 10.1016/0145-305X(88)90074-2. [DOI] [PubMed] [Google Scholar]
- Colovos C, Yeates TO. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci. 1993;2(9):1511–1519. doi: 10.1002/pro.5560020916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coordinators NR. Database resources of the national center for biotechnology information. Nucleic acids research. 2016;44(Database issue):7–22. doi: 10.1093/nar/gkt1146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhal AK, et al. An immunoinformatics approach for design and validation of multi-subunit vaccine against Cryptosporidium parvum. Immunobiology. 2019;224(6):747–757. doi: 10.1016/j.imbio.2019.09.001. [DOI] [PubMed] [Google Scholar]
- Dickerson HW, Clark TG, Leff AA. Serotypic variation among isolates of Ichthyophthirius multifiliis based on immobilization. J Eukaryot Microbiol. 1993;40(6):816–820. doi: 10.1111/j.1550-7408.1993.tb04480.x. [DOI] [PubMed] [Google Scholar]
- Dickerson HW, Woo PTK, Buchmann K (2011) Ichthyophthirius multifiliis. In: Woo PTK (ed) Fish parasites: pathobiology and protection. CABI, Wallingford, pp 55–72
- Dickerson H, Findly R. Immunity to Ichthyophthirius infections in fish: a synopsis. Dev Comp Immunol. 2014;43(2):290–299. doi: 10.1016/j.dci.2013.06.004. [DOI] [PubMed] [Google Scholar]
- Dickerson HW, Woo PTK, Buchmann K (2011) Ichthyophthirius multifiliis. In: Woo PTK (ed) Fish parasites: pathobiology and protection, CABI, Wallingford, pp 55–72
- Dimitrov I, et al. AllergenFP: allergenicity prediction by descriptor fingerprints. Bioinformatics. 2014;30(6):846–851. doi: 10.1093/bioinformatics/btt619. [DOI] [PubMed] [Google Scholar]
- Dimitrov I, et al. AllerTOP v. 2—a server for in silico prediction of allergens. J mol model. 2014;20(6):1–6. doi: 10.1007/s00894-014-2278-5. [DOI] [PubMed] [Google Scholar]
- Doytchinova IA, Flower DR. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007;8(1):4. doi: 10.1186/1471-2105-8-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elsayed EE, et al. Ichthyophthiriasis various fish susceptibility or presence of more than one strain of the parasite. Nat Sci. 2006;4(3):5–13. [Google Scholar]
- Ewing MS, Kocan KM, Ewing SA. Ichthyophthirius multifiliis (Ciliophora) invasion of gill epithelium 1. J Protozool. 1985;32(2):305–310. doi: 10.1111/j.1550-7408.1985.tb03055.x. [DOI] [PubMed] [Google Scholar]
- Gededzha MP, Mphahlele MJ, Selabe SG. Prediction of T-cell epitopes of hepatitis C virus genotype 5a. Virol J. 2014;11(1):187. doi: 10.1186/1743-422X-11-187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geourjon C, Deleage G. SOPMA: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics. 1995;11(6):681–684. doi: 10.1093/bioinformatics/11.6.681. [DOI] [PubMed] [Google Scholar]
- Ghosh P, et al. A novel multi-epitopic peptide vaccine candidate against helicobacter pylori in-silico identification, design cloning and validation through molecular dynamics. Int J Peptide Res Ther. 2021;27(2):1–18. doi: 10.1007/s10989-020-10157-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grote A, et al. JCat a novel tool to adapt codon usage of a target gene to its potential expression host. Nuclei Acids Res. 2005;33(suppl2):526–531. doi: 10.1093/nar/gki376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hasan M, et al. Contriving a chimeric polyvalent vaccine to prevent infections caused by herpes simplex virus (type-1 and type-2): an exploratory immunoinformatic approach. J Biomol struct dynamics. 2020;38(10):2898–2915. doi: 10.1080/07391102.2019.1647286. [DOI] [PubMed] [Google Scholar]
- Hebditch M, et al. Protein-Sol: a web tool for predicting protein solubility from sequence. Informatics. 2017;33(19):3098–3100. doi: 10.1093/bioinformatics/btx345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Humphrey W, Dalke A, Schulten KJJomg. VMD visual molecular dynamics. J Mol Gr. 1996;14(1):33–38. doi: 10.1016/0263-7855(96)00018-5. [DOI] [PubMed] [Google Scholar]
- Ishack S., Lipner S.R. Bioinformatics and immunoinformatics to support COVID-19 vaccine development. J Med Virol. 2021;93(9):5209–5211. doi: 10.1002/jmv.27017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jespersen MC, et al. BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017;45(W1):W24–W29. doi: 10.1093/nar/gkx346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamens JJNAR. The Addgene repository an international nonprofit plasmid and data resource. Nuclei Acids Research. 2015;43(D1):D1152–D1157. doi: 10.1093/nar/gku893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Y, et al. Immune epitope database analysis resource. Nucleic Acids Res. 2012;40(W1):W525–W530. doi: 10.1093/nar/gks438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koppang EO, Kvellestad A, Fischer U. Mucosal health in aquaculture. Elsevier; 2015. Fish mucosal immunity: gill; pp. 93–133. [Google Scholar]
- Laskowski RA, MacArthur MW, Thornton JM (2006) PROCHECK: validation of protein-structure coordinates
- López-Blanco JR, et al. iMODS internal coordinates normal mode analysis server. Nucleic Acids Res. 2014;42(W1):W271–W276. doi: 10.1093/nar/gku339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacKerell AD, Jr, Banavali N, Foloppe NJBORoB. Development and current status of the CHARMM force field for nucleic acids. Biopolymers. 2000;56(4):257–265. doi: 10.1002/1097-0282(2000)56:4<257::AID-BIP10029>3.0.CO;2-W. [DOI] [PubMed] [Google Scholar]
- Magnadottir B. Immunological control of fish diseases. Mar Biotechnol. 2010;12(4):361–379. doi: 10.1007/s10126-010-9279-x. [DOI] [PubMed] [Google Scholar]
- Mamun M, et al. Heavy infection of Ichthyophthirius multifiliis in striped catfish (Pangasianodon hypophthalmus, Sauvage 1878) and its treatment trial by different therapeutic agents in a control environment. J Appl Aquac. 2020;32(1):81–93. doi: 10.1080/10454438.2019.1610541. [DOI] [Google Scholar]
- Matthews JL. Methods in cell biology. Elsevier; 2004. Common diseases of laboratory zebrafish; pp. 617–643. [DOI] [PubMed] [Google Scholar]
- Medzhitov R, Janeway CAJS. Decoding the patterns of self and nonself by the innate immune system. Science. 2002;296(5566):298–300. doi: 10.1126/science.1068883. [DOI] [PubMed] [Google Scholar]
- Pagadala NS, Syed K, Tuszynski J. Software for molecular docking: a review. Biophys Rev. 2017;9(2):91–102. doi: 10.1007/s12551-016-0247-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patra P, et al. Biocomputational analysis and in silico characterization of an angiogenic protein RNase5 in zebrafish Danio rerio. Int J Peptide Res Ther. 2019;26(4):1–11. [Google Scholar]
- Patronov A, Doytchinova I. T-cell epitope vaccine design by immunoinformatics. Open Biol. 2013;3(1):120139. doi: 10.1098/rsob.120139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pentel PR, LeSage MG. Advances in pharmacology. Elsevier; 2014. New directions in nicotine vaccine design and use; pp. 553–580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips JC, et al. Scalable Molecular Dynamics with NAMD. J Computational Chem. 2005;26(16):1781–1802. doi: 10.1002/jcc.20289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pradhan N, Sharma K (2014) Understanding Ubl-rpn1 intermolecular interaction. J Adv Pharm Technol Res 1(3):1
- Purcell MK, et al. Conservation of toll-like receptor signaling pathways in teleost fish. Genom Proteom. 2006;1(1):77–88. doi: 10.1016/j.cbd.2005.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rapin N, et al. Computational immunology meets bioinformatics: the use of prediction tools for molecular binding in the simulation of the immune system. PLOS One. 2010;5(4):e9862. doi: 10.1371/journal.pone.0009862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roach JC, et al. The evolution of vertebrate toll-like receptors. Proc Natl Acad Sci. 2005;102(27):9577–9582. doi: 10.1073/pnas.0502272102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roy A, Yang J, Zhang Y. COFACTOR: an accurate comparative algorithm for structure-based protein function annotation. Nucleic Acids Res. 2012;40(W1):W471–W477. doi: 10.1093/nar/gks372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saha R, Ghosh P, Burra VPJB. Designing a next generation multi-epitope based peptide vaccine candidate against SARS-CoV-2 using computational approaches. Biotech. 2021;11(2):1–14. doi: 10.1007/s13205-020-02574-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salaikumaran MR, et al. An in silico approach to study the role of epitope order in the multi-epitope-based peptide (MEBP) vaccine design. Sci Rep. 2022;12(1):1–18. doi: 10.1038/s41598-022-16445-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneidman Duhovny D, et al. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 2005;33(Web Server issue):363–7. doi: 10.1093/nar/gki481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh H, Raghava G. ProPred: prediction of HLA-DR binding sites. Bioinformatics. 2001;17(12):1236–1237. doi: 10.1093/bioinformatics/17.12.1236. [DOI] [PubMed] [Google Scholar]
- Singh H, Raghava G. ProPred1: prediction of promiscuous MHC Class-I binding sites. Bioinformatics. 2003;19(8):1009–1014. doi: 10.1093/bioinformatics/btg108. [DOI] [PubMed] [Google Scholar]
- Stoskopf MK. Laboratory Animal Medicine. Elsevier; 2015. Biology and management of laboratory fishes; pp. 1063–1086. [Google Scholar]
- Trujillo-González A, Becker JA, Hutson KS. Advances in parasitology. Elsevier; 2018. Parasite dispersal from the ornamental goldfish trade; pp. 239–281. [DOI] [PubMed] [Google Scholar]
- Van Aalten DM, et al. A comparison of techniques for calculating protein essential dynamics. J Computational Chem. 1997;18(2):169–181. doi: 10.1002/(SICI)1096-987X(19970130)18:2<169::AID-JCC3>3.0.CO;2-T. [DOI] [Google Scholar]
- Ventura MT, Paperna I. Histopathology of Ichthyophthirim multifiliis infections in fishes. J Fish Biol. 1985;27(2):185–203. doi: 10.1111/j.1095-8649.1985.tb04020.x. [DOI] [Google Scholar]
- von Gersdorff Jørgensen L. The fish parasite Ichthyophthirius multifiliis–host immunology, vaccines and novel treatments. Fish Shellfish Immunol. 2017;67:586–595. doi: 10.1016/j.fsi.2017.06.044. [DOI] [PubMed] [Google Scholar]
- Wang X, et al. Immunisation of channel catfish, Ictalurus punctatus, with Ichthyophthirius multifiliis immobilisation antigens elicits serotype-specific protection. Fish Shellfish Immunol. 2002;13(5):337–350. doi: 10.1006/fsim.2001.0410. [DOI] [PubMed] [Google Scholar]
- Wiederstein M, Sippl MJ. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007;35(suppl2):W407–W410. doi: 10.1093/nar/gkm290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- XU C, et al. Analysis of the soluble and membrane-bound immobilization antigens of Ichthyophthirius multifiliis. J Eukaryot Microbiol. 1995;42(5):558–564. doi: 10.1111/j.1550-7408.1995.tb05905.x. [DOI] [Google Scholar]
- Yang Y, et al. Improving protein fold recognition and template-based modeling by employing probabilistic-based matching between predicted one-dimensional structural properties of query and corresponding native properties of templates. Bioinformatics. 2011;27(15):2076–2082. doi: 10.1093/bioinformatics/btr350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao F, et al. Grass carp (Ctenopharyngodon idella) TRAF6 and TAK1: molecular cloning and expression analysis after Ichthyophthirius multifiliis infection. Fish Shellfish Immunol. 2013;34(6):1514–1523. doi: 10.1016/j.fsi.2013.03.003. [DOI] [PubMed] [Google Scholar]
- Zhong S, Macias AT, MacKerell AD. Computational identification of inhibitors of protein-protein interactions. Curr Top Med Chem. 2007;7(1):63–82. doi: 10.2174/156802607779318334. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary file1 (DOCX 13 kb) Supplementary table 1: PatchDock predicted docking score (top 20) which predicted the rank based on complementary score, ACE (Atomic Contact Energy) and estimated interface area. (Here bold model has been taken for its high negative ACE value).
Data Availability Statement
All data generated or analyzed during this study are included in this published article.