Abstract
Teg49 is a Staphylococcus aureus trans-acting regulatory sRNA derived from cleavage of the sarA P3 transcript. We showed by RNA-Seq here that the 5′ trident-like structure in Teg49 regulates transcriptionally (direct and indirect) 22 genes distinct from sarA. Among these, Teg49 was noted to repress spn, encoding a 102 residue preprotein which yields the mature 73 residue peptide which inhibits the catalytic activity of myeloperoxidase in human neutrophils. Teg49 was found to regulate spn mRNA post-transcriptionally in strain SH1000 through 9-nt base-pairing between hairpin loop 2 of Teg49 and an exposed bulge of the spn mRNA. Mutations of the Teg49 binding site disrupted the repression of spn, leading to reduced degradation, and increased half-life of spn mRNA in the Teg49 mutant. The spn-Teg49 interaction was also confirmed with a synonymous spn mutation to yield enhanced spn expression in the mutant vs. the parent. The Teg49 mutant with increased spn expression exhibited enhanced resistance to MPO activity in vitro. Killing assays with human neutrophils showed that the Teg49 mutant was more resistant to killing after phagocytosis. Altogether, this study shows that Teg49 in S. aureus has a distinct and important regulatory profile whereby this sRNA modulates resistance to myeloperoxidase-mediated killing by human neutrophils.
Keywords: gene expression, human neutrophils, myeloperoxidase, regulatory small RNA, RNA, SPIN, spn, Staphylococcus aureus, virulence
1 |. INTRODUCTION
Staphylococcus aureus is a human commensal and an opportunistic pathogen that can cause a myriad of infections, ranging from superficial boils and abscess to serious bacteremia (Lowy, 1998). Over the last two decades, there are increasing caseloads of S. aureus infections both in the community and hospital settings. The economic burden of these infections has been growing, not to mention the morbidity and mortality caused by this pathogen worldwide (Noskin et al., 2007; Tong et al., 2015). One reason S. aureus is highly effective in causing infections is its ability to navigate the innate immune system, which includes neutrophils (PMN for polymorphonuclear cells), macrophages, and a host of defensive proteins (Cheung et al., 2021).
Successful infections by S. aureus depend on an arsenal of virulence factors that are coordinately expressed to protect the bacteria from attack by the host’s immune systems. The expression of these virulence factors is mediated by an array of regulatory factors including two-component regulatory systems, transcription factors, and post-transcriptional modifications that sense and respond to environmental cures (Cheung et al., 2004). In contrast to protein-based regulation which requires time-consuming protein synthesis, post-transcriptional regulation via sRNAs is rapid and better at filtering high-frequency input noise (Mehta et al., 2008; Shimoni et al., 2007). It has been proposed that regulatory factors like sRNAs can become potential targets for drug development. Accordingly, we need to understand how they work to develop new therapeutics against important bacterial pathogens (Roncarati et al., 2022).
Among the myriad of transcription factors in S. aureus, SarA is a prominent DNA binding protein that binds target promoters to regulate the expression of crucial virulence factors including toxins (e.g., alpha and delta toxins), surface adhesins (e.g., protein A), and proteases (e.g., aureolysin). Deletion of sarA leads to reduced activation of the agr quorum-sensing system as well as decreased biofilm formation (Bayer et al., 1996; Chan & Foster, 1998; Cheung et al., 2008). The sarA locus has an extended 5′ UTR with an unusual architecture that contains three distinct promoters (P1, P3, and P2), driving three overlapping transcripts, all capable of making SarA (Manna et al., 1998). Nested between the proximal P1 and the P3 promoters is a small RNAs called Teg49 which was first identified by RNA-Seq of the whole S. aureus genome (Beaume et al., 2010). Additional analysis revealed that the 196 nt long Teg49 is generated from the cleavage of the sarA P3 transcript, and its expression is dependent on the stress-induced σB dependent P3 promoter (Manna et al., 2018). While some regulatory targets of Teg49 have been predicted, the spectrum of gene regulation of Teg49 independent of SarA and its exact mechanism of regulation are not known (Manna et al., 2018).
In this study, we evaluated Teg49 regulation and confirmed that it regulates genes independent of SarA, leading to specific phenotypes not attributed to SarA. Among the Teg49 specifically regulated genes, we were particularly interested in spn which encodes a 102-residue prepeptide that is processed to yield a mature 73 residue peptide (SPIN) that inhibits myeloperoxidase (MPO) in human PMNs (de Jong et al., 2017). As MPO is a critical host enzyme that converts hydrogen peroxide to HOCl, a powerful oxidant inside phagosome with potent antimicrobial properties (Hampton et al., 1996), we wanted to further understand the regulation of spn by Teg49.
We confirmed that Teg49 in S. aureus strain SH1000 repressed spn expression. Using a targeted mutagenesis approach, we deduced that Teg49 base pairs with spn to destabilize the mRNA. As a result, the spn mRNA has a longer half-life in the Teg49 mutant vs. the parent (4.994 ± 0.2592 min in the Teg49 mutant vs. 1.805 ± 0.3322 min in the parent). There are three hairpin loops within the structure of Teg49. However, only the hairpin loop 2 was predicted to bind the spn mRNA as assessed by in silico analysis. In mutagenesis studies, both hairpin 2 loop mutant of Teg49 and the reciprocal binding site spn mutant exhibited higher spn expression vs. the parental strain as determined by qRT-PCR.
Teg49-mediated repression of spn in S. aureus implied increased MPO resistance of the Teg49 mutant in vitro as compared to wild type in an enzyme mix. Teg49 mutants also had better survival after phagocytosis by human PMNs. In both of these cases, Teg49 survival was dependent on the presence of spn. With these studies, we confirm that Teg49 plays a significant role in controlling innate host response that is distinct from SarA.
2 |. RESULTS
2.1 |. The small regulatory RNA Teg49 represses the expression of spn, a virulence gene in S. aureus
Teg49 was originally identified in an exploratory whole-genome RNA-Seq study of S. aureus strain N315 (Beaume et al., 2010). This study found a 196 nt long sRNA located within the 5′-UTR of the P3 transcript of the sarA locus. Subsequent studies confirmed the existence of Teg49 located between the P3 and P1 promoters of the sarA locus (Kim et al., 2014). Teg49 is generated by the cleavage of the sarA P3 transcript in a RNase III-dependent and Hfq-independent manner (Manna et al., 2018). As part of the P3 sarA transcript, the expression of Teg49 is dependent on the P3 promoter under the control of SigB, a stress-dependent transcription factor found in Firmicutes (Manna et al., 2018). This study seeks to address the important question of whether Teg49 has a phenotype distinct from SarA.
The Teg49 knockout strain used in Manna et al. (2018) study to examine Teg49 had a 209 bp deletion that overlapped with the P1 sarA promoter; in order to study Teg49 specifically, we reconstructed the Teg49 mutant strain with a 70-base deletion at the 5′ end that is missing the three predicted hairpin loops in the structure of Teg49 (Table 1).
TABLE 1.
Strains and plasmids
| Species | Strain ID | Description | References |
|---|---|---|---|
| Cloning | |||
| Escherichia coli | IM08B | Used For Cloning (DH10B Δdcm with hsdS CC8-1of NRS384) | Monk et al. (2015) |
| Background | |||
| Staphylococcus aureus | ALC7589 | SH1000 (Functional rsbU derivative of 8325-4 rsbU+) | Horsburgh et al. (2002) |
| ALC4495 | SH1000 containing empty xylose-inducible vector, pEPSA5 | This Study | |
| ALC2727 | SH1000 containing promoterless pALC1484:gfp vector | Kahl et al. (2000) | |
| ALC9379 | SH1000 containing pALC1484:spn promoter-gfp | This Study | |
| Teg49 mutants | |||
| Staphylococcus aureus | ALC8744 | Teg49 mutant of SH1000; chromosomal deletion of 70-bp Teg49 from nt 305 to 375 from the ATG start codon of the sarA gene | This Study |
| ALC9377 | Teg49 genomic complement (ALC8744 restored with wild type Teg49) | This Study | |
| ALC8789 | ALC8744 containing an empty xylose-inducible vector, pEPSA5 | This Study | |
| ALC8790 | ALC8744 containing pEPSA5 carrying 196-bp Teg49 fragment DNA under a xylose-inducible promoter | This Study | |
| ALC9380 | Teg49 mutant (ALC8744) containing pALC1484:spn promoter-gfp | This Study | |
| ALC7289 | SH1000 Teg49 HP1 loop mutation of Teg49 (ATTGCGC→CGCTATA) | Kim et al. (2014) | |
| ALC7288 | SH1000 Teg49 HP1 stem mutation of Teg49 | Kim et al. (2014) | |
| ALC7290 | SH1000 Teg49 HP2 loop mutation of Teg49 (GTCGATT→TATCGGC) | Kim et al. (2014) | |
| ALC7291 | SH1000 Teg49 HP2 stem mutation of Teg49 | Kim et al. (2014) | |
| ALC8842 | SH1000 Teg49 HP3 Loop Mutant (ACTAAACCAAATGCTAA→GGTAGTTTTAAAAATAA) | This Study | |
| ALC9005 | SH1000 Teg49 HP3 stem mutation of Teg49 | This Study | |
| sarA mutants | |||
| Staphylococcus aureus | ALC2732 | SH1000 ΔsarA::ermC | Ballal et al. (2010) |
| ALC9378 | SH1000 sarA nonsense mutant (ATG→TAATAA) | This Study | |
| ALC8373 | SH1000 sarA P3 promoter scrambles | Manna et al. (2018) | |
| spn mutants | |||
| Staphylococcus aureus | ALC9381 | SH1000 spn deletion (382 bp genomic deletion, beginning 28 bp from the ATG start to 45 bp after the TAA stop codon) | This Study |
| ALC9382 | SH1000 spn genomic complement using pMAD | This Study | |
| ALC9383 | SH1000 with both Teg49 mutation and Δspn | This Study | |
| ALC9384 | spn synonymous mutant (TTC GCC GAT AAA to TTT TCA GAC AAG) (Amino Acid: F-A-D-N to F-A-D-N) | This Study | |
| ALC9385 | spn non-synonymous mutant (TTC GCC GAT AAA to TTT CTT CTA CGT) (Amino Acid: F-A-D-N to F-L-L-R) | This Study | |
| sae and rnc mutants | |||
| Staphylococcus aureus | ALC4473 | sae:Tn551 knockout isogenic for SH1000 | Xiong et al. (2006) |
| ALC8243 | Δrnc that is, RNase III genomic deletion of SH1000 with pMAD | Manna et al. (2018) | |
| ALC8445 | Δrnc genomic complement (ALC8243 restored with wild type rnc using pMAD) | Manna et al. (2018) | |
| pSK236 | CmR | Shuttle vector containing pUC19 cloned into the HindIII site of pC194 | This Study |
| pMAD | AmpR, ErmR | E. coli-S. aureus shuttle vector containing the temperature-sensitive origin of replication and bgaB | Arnaud et al. (2004) |
| pEPSA5 | AmpR, CmR | E. coli-S. aureus xylose-inducible shuttle vector containing xylose-inducible promoter for conditional expression of the desired gene | Forsyth et al. (2002) |
| pEPSA5::Teg49 | AmpR, CmR | pEPSA5 containing 196-bp Teg49 sRNA | Kim et al. (2014) |
| pALC1484 | AmpR, CmR | Modified pSK236 shuttle vector with a promoterless gfpuvr reporter gene preceded by an S. aureus ribosome binding site | Kahl et al. (2000) |
To understand the spectrum of genes regulated by Teg49, we first performed RNA-Seq on the wild type strain SH1000, isogenic Teg49 mutant, and Teg49 complemented with a pEPSA5:Teg49 vector. After excluding ribosomal and phage genes, we focused on genes expressed at twofold higher and or lower (Table 2). Genes regulated by SarA, ribosomal genes, phage, and transposon genes were excluded from the analysis (Manna et al., 2018; Oriol et al., 2021). In the Teg49 mutant, there were 22 genes that had notable transcriptional changes due to Teg49 independent of SarA when compared to the parental strain. Besides having a narrower selection of genes (Manna et al., 2018), this list did not include saeRS as a regulatory target for Teg49. Whether Teg49 impacts post-transcriptional event (e.g., translation of SaeRS) is not addressed in our study. Among the Teg49-repressed genes, we found SAOUHSC_00401 (i.e., spn), which codes for the SPIN protein. SPIN, secreted as a 102-residue prepeptide is then cleaved to yield the 73-residue conserved mature peptide (de Jong et al., 2017). SPIN has been previously shown to inhibit the activity of myeloperoxidase (MPO), an enzyme within the phagosome of PMNs that converts hydrogen peroxide to HOCl which is a powerful oxidant with prominent antimicrobial activity (de Jong et al., 2017; Klebanoff et al., 2013). Given its clinical relevance, we elected to pursue further the regulation of spn by Teg49.
TABLE 2.
RNA-seq of ΔTeg49 mutant
| Genes | Fold change | p value | Corrected FDR | Pan genome ID | Annotation |
|---|---|---|---|---|---|
| SAOUHSC_01055a | 2.75 | 5.42E-15 | 2.36E-13 | SAUPAN003335000 | Membrane protein (A lot of genomes have a small product in that region, see SAUPAN003335000) |
| SAOUHSC_01014 | 2.72 | 7.62E-18 | 4.24E-16 | purF | Amidophosphoribosyltransferase |
| SAOUHSC_01016 | 2.61 | 4.78E-18 | 2.79E-16 | purN | Phosphoribosylglycinamide formyltransferase |
| SAOUHSC_01017 | 2.55 | 1.72E-16 | 8.94E-15 | purH | Bifunctional phosphoribosylaminoimidazolecarboxamide formyltransferase/IMP cyclohydrolase |
| SAOUHSC_01889 | 2.5 | 4.33E-20 | 2.72E-18 | ribD | Riboflavin biosynthesis protein RibD |
| SAOUHSC_01015 | 2.44 | 1.61E-14 | 6.87E-13 | purM | Phosphoribosylaminoimidazole synthetase |
| SAOUHSC_01887 | 2.33 | 6.84E-16 | 3.21E-14 | ribA | Riboflavin biosynthesis protein |
| SAOUHSC_03042 | 2.33 | 6.67E-13 | 2.53E-11 | SAUPAN006469000 | Integrase/recombinase core subunit |
| SAOUHSC_01888 | 2.31 | 2.30E-21 | 1.66E-19 | ribB | Riboflavin synthase subunit alpha |
| SAOUHSC_01018 | 2.24 | 2.73E-13 | 1.09E-11 | purD | Phosphoribosylamine--glycine ligase |
| SAOUHSC_00401 | 2.16 | 1.44E-21 | 1.08E-19 | SAUPAN002132000 | Myeloperoxidase inhibitor SPIN |
| SAOUHSC_01330 | 2.03 | 1.67E-20 | 1.14E-18 | guaC | Guanosine 5′-monophosphate oxidoreductase |
| SAOUHSC_00473 | 0.49 | 6.85E-11 | 2.37E-09 | SAUPAN002241000 | Small ORF only in NCTC8325 chromosome, between prsA and rplY |
| SAOUHSC_01403 | 0.48 | 1.32E-17 | 7.20E-16 | cspA | Cold shock protein |
| SAOUHSC_02681 | 0.47 | 5.92E-13 | 2.28E-11 | narG | Nitrate reductase subunit alpha |
| SAOUHSC_02671 | 0.47 | 5.00E-27 | 5.44E-25 | narK | Nitrite extrusion protein |
| SAOUHSC_02426 | 0.46 | 2.48E-16 | 1.26E-14 | YjiH | Membrane protein, nucleoside recognition domain (YjiH-Like) |
| SAOUHSC_00898 | 0.46 | 6.85E-17 | 3.64E-15 | argH | Argininosuccinate lyase |
| SAOUHSC_01275 | 0.45 | 2.16E-15 | 9.94E-14 | glpF | Aquaporin family protein (glycerol) |
| SAOUHSC_02130 | 0.39 | 2.15E-18 | 1.28E-16 | yebG | Hypothetical protein with NETI domain (YebG-like, DNA damage sensing) |
| SAOUHSC_00899 | 0.35 | 3.08E-23 | 2.95E-21 | argG | Argininosuccinate synthase |
| SAOUHSC_02662 | 0.29 | 3.80E-16 | 1.90E-14 | scrA | PTS system sucrose-specific transporter subunit IIBC |
Notes: RNA-Seq performed with wild type Staphylococcus aureus SH1000, a Teg49 mutant, and an inducible pEPSA:Teg49 complemented strain. It shows 22 genes differentially regulated by the Teg49 mutant but not the SH1000 isogenic parental strain. Gene numbers, given according to SH1000/NCTC8325 designation, fold change, statistical values, and Aureowiki gene information, are shown. Aureowiki annotations are based on NCBI RefSeq analyzed at (https://aureowiki.med.uni-greifswald.de). Paired-end sequencing was analyzed using CLC and edgeR. Significantly regulated genes were selected for 2× fold change, and a p-value of <.001. Genes regulated by SarA, ribosomal genes, phage, and transposon genes were excluded from the analysis.
Besides the RNA-seq data, we used Nanostring nCounter to verify the effects of Teg49 on spn. As controls, we utilized a series of genes coding for sarA P1 transcript as well as target genes of sarA such as aur, spa, and others (Figure 1a). We used the housekeeping genes hu and rpoB to normalize reads while esxB, groES, pyrP served as internal controls for SarA/Teg49-independent genes. Besides spn, other genes regulated by Teg49 including purH and ribD (Table 2) were also assayed. As shown in Figure 1a, upregulation of spn in the Teg49 mutant was observed, but not in the wild type and complemented mutant, or the sarA deletion mutant. Genes that are known to be repressed by SarA such as aur (aureolysin) and spa (protein A) (Chien et al., 1999) were up-regulated in the sarA mutant but not in the Teg49 mutant. The sarA P1 transcript, the prominent transcript within the sarA locus, was also not affected in the Teg49 mutant but was undetectable in the sarA deletion strain. Notably, the Nanostring nCounter data (Figure 1a) and the qRT-PCR results (data not shown) did not reveal differences in the expression of purH and ribD, which were up-regulated in the Teg49 mutant vs. parent SH1000 as assessed by RNA-Seq (Table 2). Likewise, expression of scrA, reduced in the Teg49 mutant vs. the parent as detected by RNA-Seq, was not altered in the isogenic Teg49 strains by Nanostring data, but the qRT-PCR results suggested that scrA expression appeared to be decreased in the Teg49 mutant (data not shown). The reason for the discrepancy between Nanostring and qRT-PCR results for scrA is not clear. Nevertheless, these data affirmed that Teg49 and SarA regulation are distinct and that Teg49 has its own distinct regulon that includes spn.
FIGURE 1.
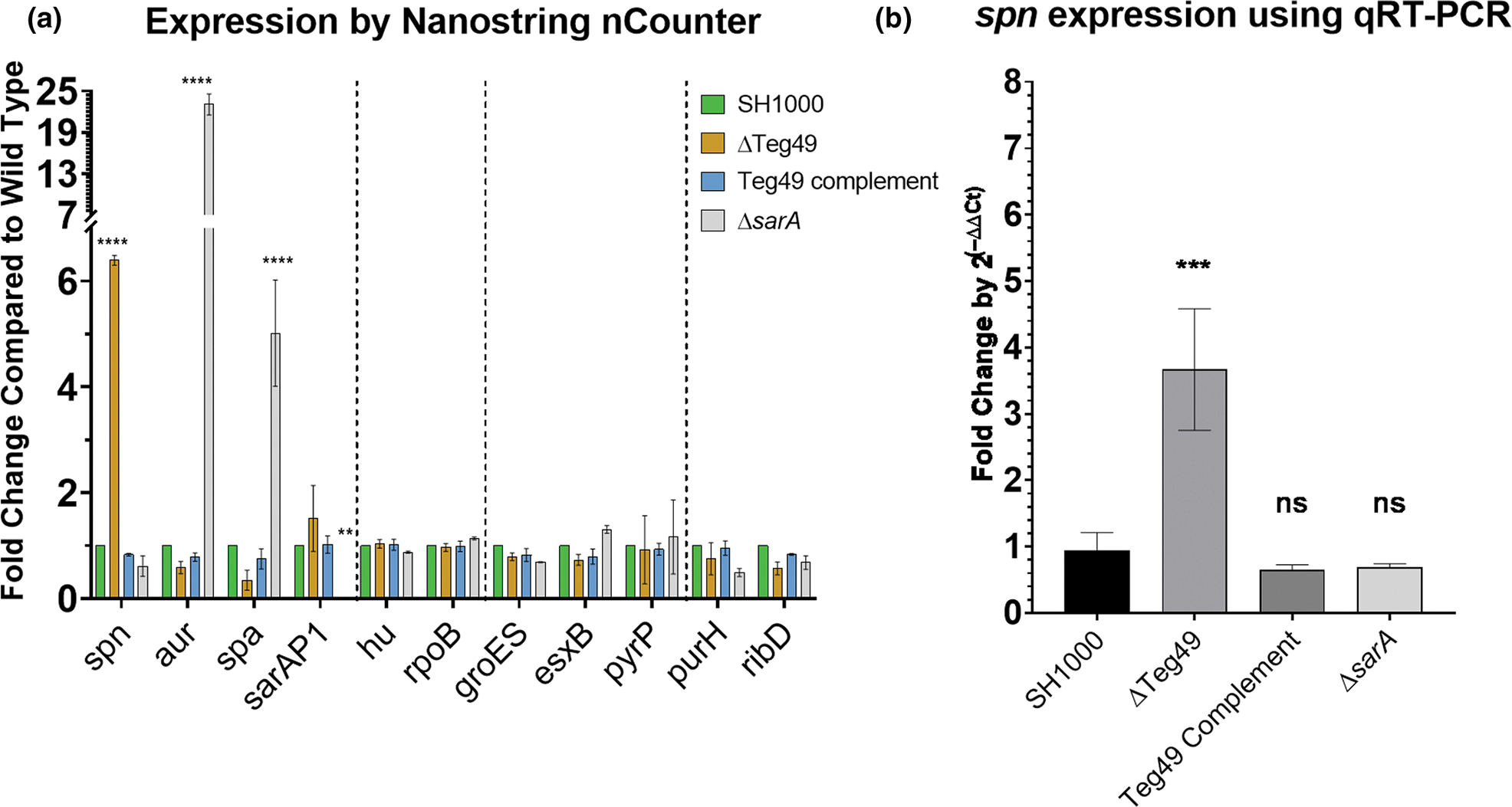
Teg49 regulates the expression of spn, a myeloperoxidase inhibitor. (a) Nanostring nCounter data for genes in Staphylococcus aureus SH1000, Teg49 mutant, complement, and sarA mutant. This array includes two housekeeping genes HU and rpoB, spn, and several genes that are known to be regulated by SarA (i.e., aureolysin gene aur and protein a gene spa). Genes are separated with vertical dotted lines to show differentially regulated genes, housekeeping genes (HU and rpoS), one group of genes not regulated by Teg49 or SarA (groES, esxB, and pyrP) and another group detected by RNA-seq but not confirmed by Nanostring (purH and ribD). Data were analyzed using a two-way ANOVA using Dunnett’s test for multiple comparisons. p values are * = .05, ** = .01, *** = .001, **** ≤ .001. (b) qRT-PCR validation of spn expression in SH1000, Teg49 mutant, Teg49 complement, and sarA mutant at 4th h of growth. Fold change was calculated by 2(−ΔΔCt). Significant differences were calculated using a one-way ANOVA followed by Tukey’s multiple comparison test compared to the Teg49 mutant. p-values indicated are * = .05, ** = .01, *** = .001.
In addition to the Nanostring data, we also confirmed regulation of spn by Teg49 using qRT-PCR by comparing wild type, Teg49 mutant/complement, and sarA mutant (Figure 1b). Again, only the Teg49 deletion mutant exhibited upregulation of spn in these strains, and deletion of sarA did not impact spn expression vs. the parent.
2.2 |. Characterizing the spn locus and its expression controlled by Teg49
Since spn expression has only been examined in USA300 strain (de Jong et al., 2017), we wanted to verify spn expression in MSSA strain SH1000 during the growth cycle. Using qRT-PCR, an increasing level of spn expression was observed in the parental strain during growth, accumulating to the late exponential/early stationary phase (~4–5 h), tapering toward the stationary phase (6 h; Figure 2a,b). We also analyzed serial Teg49 expression by densitometry of Northern blots at similar timepoints (Figure 2b, right Y axis; Figure S3d). At early timepoints (2–3 h), Teg49 expression decreased, presumably due to dilution and degradation effect from the overnight culture before robust production of Teg49 occurred. This period coincided with the up-regulation of spn expression by qRT-PCR. As the level of Teg49 increased (4–5 h), the spn expression level began to decrease (5–6 h). Overall, the trend is consistent with the repression of spn by Teg49. In previous studies, It has been shown that spn expression is not regulated by agr and quorum sensing and is instead likely to be induced by ROS stress encountered during significant increases in cell density in the early stationary phase (de Jong et al., 2017; Guerra et al., 2016).
FIGURE 2.
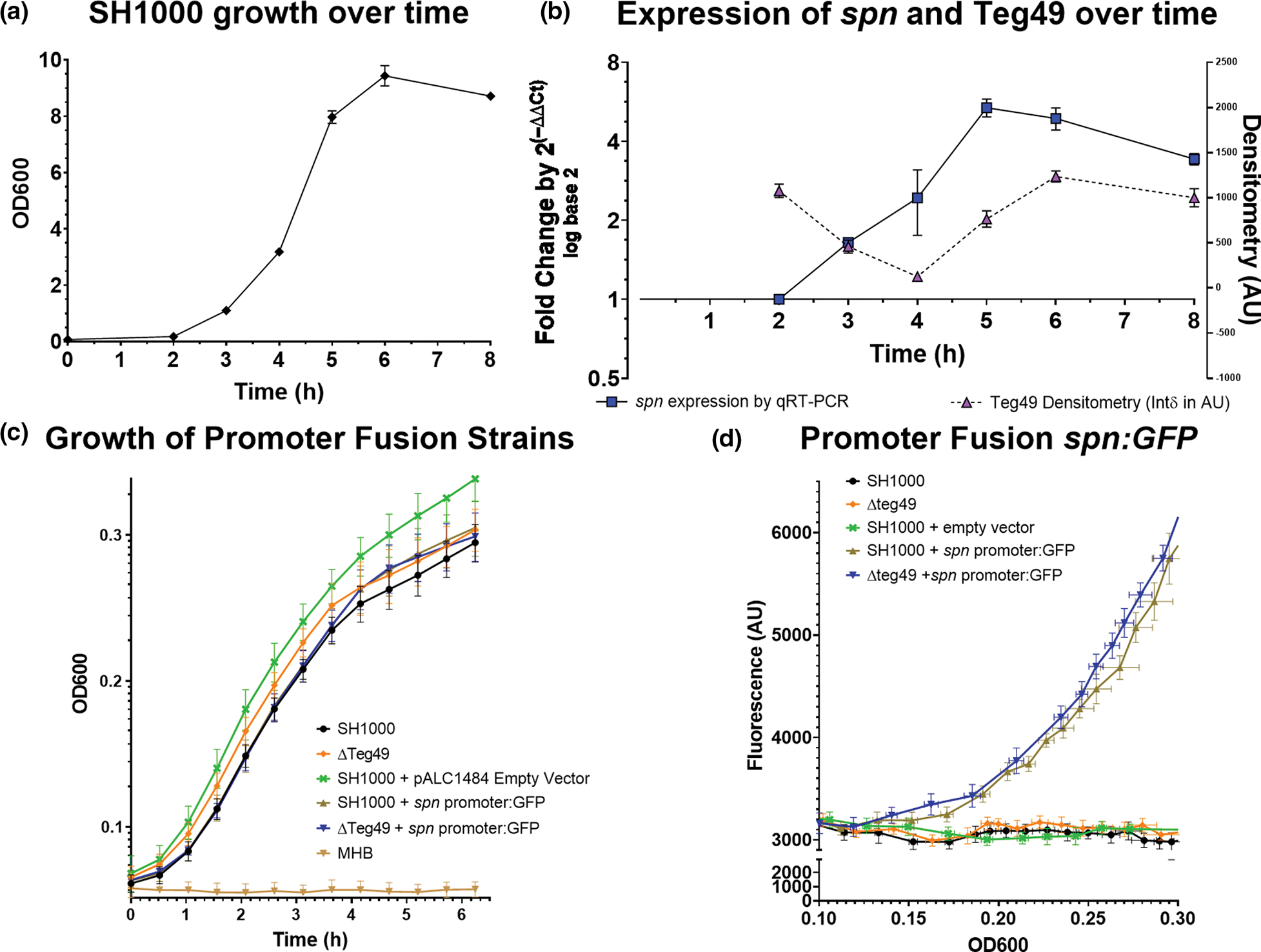
Characterization of spn expression during growth. (a) Growth of strain SH1000 as determined by OD600 in a spectrophotometer with a 1 cm cuvette. (b) The left Y-axis represents a time-course of spn expression by qRT-PCR at OD corresponding to growth in (a). Increasing levels of the spn transcript were detected as cells reached the stationary phase and then tapered. The Y axis on the right is a densitometric analysis of a northern blot of Teg49 during 8 h. growth after dilution from an overnight culture. The timepoints for the densitometry are approximate but are not identical to those in (a). (c) OD600 growth curve for wild type SH100 and ΔTeg49 with or without the pALC1484:Spn promoter GFP fusion plasmid and the control empty vector in SH1000. (d) Fluorescence of the same strains (as in (c)) was measured in AU over OD600. OD600 was measured using a Tecan infinity 1000 with a 100 μl volume in a standard 96 well plate and reached a plateau at OD600 = 0.29/0.30. There was no significant difference in fluorescence between the SH1000 and ΔTeg49 strain containing the spn promoter fusion plasmid.
We also explored the effect of Teg49 on the spn promoter by promoter fusion with GFPuvr. To map the promoter, we first determined the exact 5′ transcriptional start site for spn using 5′ RACE. Our result demonstrated only one +1 transcriptional start of spn at 24 bases upstream of the translation start (see Figure S1a). We have also deduced the putative −10 (TATGAT) and −35 (TTTTCT) boxes based on the +1 start site (Novick, 1990). Based on these data, we constructed a transcriptional fusion of a 150 bp spn promoter linked to the GFPuvr reporter in both wild type SH1000 and isogenic Teg49 deletion mutant. As shown in Figure 2c,d, there were no significant differences in growth and transcriptional activity between the wild type and the Teg49 mutant. These data indicated that sRNA Teg49 does not regulate the spn promoter and thus likely modulates spn post-transcriptionally.
2.3 |. Predicting the interaction site between Teg49 and its target spn mRNA
Both the Teg49 sRNA and the parental sarA P3 transcript retain a highly structured “trident-like” three hairpin loop structure in the region that codes for Teg49. This structure was predicted to be retained in Teg49 even after excision from the sarA P3 transcript (Figure S1b,c). The three hairpins are predicted using multiple thermodynamic structural predictions with models and software packets (see Methods section). Based on the 2004 Turner model, the nucleotides which code for Teg49 are predicted to retain the same structure after excision, and the resulting Teg49 sRNA structure is highly entropically stable with the preservation of the three hairpin loops in its 5′ region (Figure S2a) (Mathews et al., 2004). Based on this stable double-stranded structure with exposed loops, we hypothesized that Teg49 sRNA behaves like other regulatory sRNAs in S. aureus by directly targeting and degrading its target mRNA (Barrientos et al., 2021; Felden et al., 2011).
The consequence of sRNA base-pairing with cognate mRNA is to facilitate degradation by ribonucleases that target double-stranded RNA (e.g., RNase III). To explore this possibility, we utilized the IntaRNA algorithm to assess the base pair interaction between spn mRNA and Teg49 sRNA (Smith et al., 2010). This tool predicted a 9-base paring between spn mRNA (nt 242–250) with Teg49 (nt 69–61 in hairpin loop 2), with −12.23 kcal/mol hybridization energy and −8.84 kcal/mol net stability energy (Figure S2a,b). These binding energies are in the range expected for computationally predicted trans-acting RNA–RNA interactions, especially when the predicted regions are in exposed RNA regions that could readily interact with one another and seed further interaction (Brantl, 2007; Georg et al., 2020; Storz et al., 2011). In the spn mRNA, this region is located within the ORF of the encoded peptide within an exposed bulge (Figure 3a). In the Teg49 sRNA, this region is located within a hairpin loop (hairpin loop 2) of the highly structured 5′ region of Teg49, concurrent with the model that an exposed hairpin loop of a regulatory sRNA interacts with accessible regions on its target mRNA (Jørgensen et al., 2020). The interaction regions are GC-rich (5/9 nt for spn mRNA) as has been described for other “kissing loop” mRNA-sRNA interactions (Figure 3c; Geissmann et al., 2009), even though the S. aureus genome is AT-rich.
FIGURE 3.
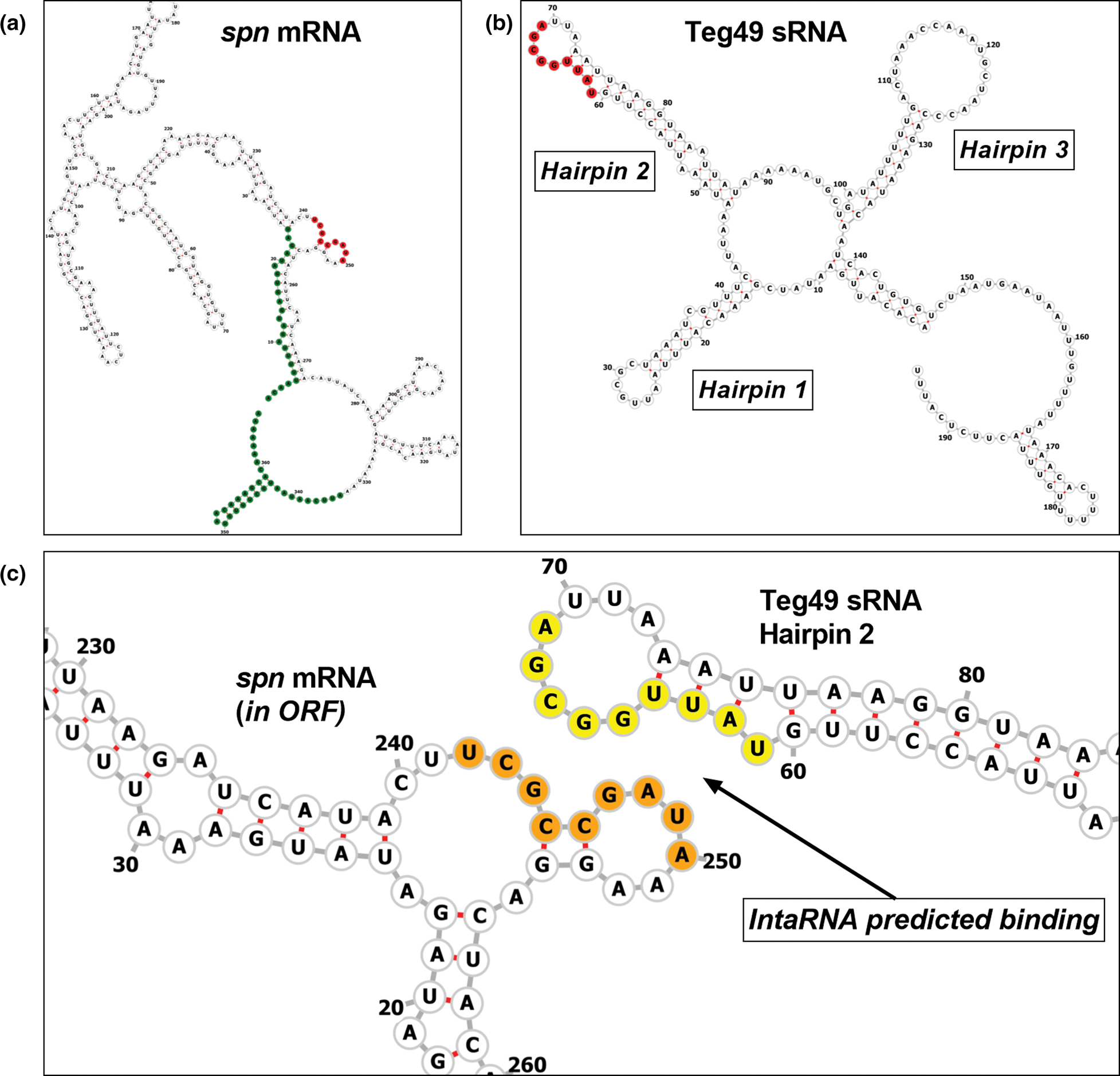
Predicted interaction between Teg49 and spn RNAs. (a) The predicted minimum free energy (MFE) RNA structure of spn mRNA from RNAfold. The spn mRNA is labeled with 5′ and 3′ UTR in green, and the IntaRNA predicted Teg49 target region is labeled in red. The region is within the ORF of spn and is predicted to be exposed. (b) The Teg49 sRNA is labeled with each of its hairpins, and the predicted spn binding region on hairpin 2 is shown in red. This binding region is located on the 5′ side of the hairpin 2 loop. (c) A closeup of the predicted interaction between spn mRNA and Teg49. In yellow are the residues in the predicted region for spn binding in Teg49, in orange is the predicted target region for binding to Teg49 in the spn coding sequence. IntaRNA prediction matches nucleotide 242 in spn mRNA to nucleotide 69 in Teg49 (U/a) and continues to nucleotide 250 in spn mRNA and nucleotide 61 in Teg49 (a/U).
2.4 |. Regulation of spn mRNA transcript level relies on interaction with the Teg49 hairpin loop, affecting the half-life of spn mRNA
We constructed mutants in the putative binding sites of both Teg49 and spn mRNA to assess if the model correctly predicts the interaction, leading to repression of spn mRNA by Teg49. For Teg49, we mutated each of the exposed loops (hairpins 1–3) by exchanging the nucleotides in a manner predicted to preserve RNA folding (Table 1). The results showed that only the Teg49 mutant with mutated hairpin loop 2, but not hairpin 1 and 3, displayed enhanced spn mRNA expression comparable to the Teg49 deletion mutant and much higher than the parental strain SH1000, suggesting that hairpin loop 2 of Teg49 is necessary for the interaction with spn mRNA (Figure 4a), consistent with the prediction of the IntaRNA algorithm above. Notably, mutation of hairpin 2 did not have an impact on the processing or stability of Teg49 or sarA P3 transcript (Figure S3a).
FIGURE 4.

The interaction of Teg49 and spn mRNA for proper spn regulation. (a) qRT-PCR looking at spn expression in Teg49 loop mutants at 4th h. of growth. Hairpin loop 1, 2, and 3 replacement mutants that retained the predicted structure of Teg49 were evaluated. Significance is measured using a one-way ANOVA using Dunnett’s multiple tests. p-values are * = .05, ** = .01, *** = .001, **** ≤ .001. (b) qRT-PCR looking at spn expression in spn mutants with synonymous (retaining the same amino acid -see text) and non-synonymous mutations (3 altered amino acids) in the predicted site for Teg49 interaction at 4th h. of growth. Significance is measured using a one-way ANOVA using Dunnett’s multiple tests. p-values are * = .05, ** = .01, *** = .001, **** ≤ .001. (c) Northern blots showing hybridizing spn band at serial time points in Teg49 mutant and SH1000 after addition of rifampin at time 0. (d) qRT-PCR for spn expression in SH1000 wild type, Teg49 mutant, and complement, as well as saeRS transposon mutant (sae mutant), RNase III deletion mutant (Δrnc), and complement at 4th h. of growth. Significance is measured using a one-way ANOVA using Dunnett’s multiple tests. p-values are * = .05, ** = .01, *** = .001, **** ≤ .001.
For spn, we constructed spn mutants with both synonymous and non-synonymous mutations in the putative Teg49 binding site (Table 2). The synonymous mutant did not change the peptide sequence of SPIN while the non-synonymous mutant altered the residues from Phe-Ala-Asp-Lys to Phe-Leu-Leu-Arg. In both synonymous and non-synonymous mutants, the spn mRNA levels by qRT-PCR were higher than the parent while this transcript was absent in the spn deletion mutant, consistent with the escape of spn mRNA from Teg49 regulation (Figure 4b).
One possibility of this escape from Teg49 regulation is a change in the stability of spn mRNA due to uncoupling of the sRNA-mRNA complex, leading to a lack of degradation from ribonucleases targeting double-strand RNAs (i.e., RNase III). To evaluate the stability of spn mRNA, a standard RNA stability assay was deployed wherein rifampin was used to inhibit RNA polymerase to halt transcription, thus enabling us to decipher the half-life of spn mRNA in the Teg49 mutant vs. the parent SH1000. After adding rifampin at time 0, serial RNA samples were obtained at times 0, 2.5, 5, 10, and 30 min. Northern blot analysis using these normalized RNA samples from isogenic Teg49 strains disclosed higher spn levels for the Teg49 mutant as compared to the parent (Figure 4c). Remarkably, the spn hybridizing band was still detectable in the Teg49 mutant at 10 min after the addition of rifampin, whereas it was not detectable in the wild type at the same timepoint. Using densitometry analysis, the spn mRNA in the Teg49 mutant was observed to start off at a higher concentration and was degraded slower, having a half-life of 4.994 ± 0.2592 min in the Teg49 mutant versus 1.805 ± 0.3322 min in the wild type (Figure S3b). Overall, these data fit the model where Teg49 binds spn mRNA to facilitate its degradation in the parental strain, while deletion of Teg49 reduces degradation of spn mRNA, yielding higher levels of spn transcript due to cellular accumulation. In the wild type SH1000, Teg49 was observed to degrade within 5–8 min (much like spn mRNA) and the sarA P2 and P3 transcripts were more stable and detectible at 10 min (Figure S3c), which is consistent with previous reports (Sorensen et al., 2020).
To evaluate the hypothesis that the sRNA-mRNA duplex was degraded by a RNase that targets double-stranded RNA, we looked at spn transcript levels by qRT-PCR in a SH1000 mutant with a disrupted gene for RNase III (rnc), which targeted double-strand RNAs for degradation. RNase III deletion mutant exhibited a much higher level of spn expression vs. the parent and complemented mutant (Figure 4d). This dramatic increase in spn expression level in the rnc mutant may be due to a combination of factors: (1) reduced degradation of spn mRNA in the Teg49-spn RNA duplex in the absence of RNase III; (2) decreased Teg49 level in the rnc mutant as we have shown in a prior study that RNase III likely plays a role in the cleavage of sarA P3 transcript to yield Teg49 (Manna et al., 2018). As has been previously reported, spn expression was dependent on an intact saeRS system (Figure 4d).
2.5 |. Teg49 mutants have increased resistance to human myeloperoxidase and better survival in human PMNs
Given that MPO causes bactericidal killing in phagosomes of PMNs and that SPIN interferes with MPO activity (de Jong et al., 2017; Klebanoff et al., 2013), we next asked if increased spn level in the Teg49 mutant would lead to enhanced survival of S. aureus in HOCl generated in vitro by MPO and in human PMNs ex vivo.
To simplify the complex process, we first utilized an in vitro assay to mimic the phagosomal killing of bacteria by generating HOCl in the presence of hydrogen peroxide and MPO. For this assay, we deployed an enzyme mix containing glucose and glucose oxidase (GO) to generate hydrogen peroxide (H2O2) which served as a substrate for MPO to produce hypochlorous acid (HOCl; Denys et al., 2011). This in vitro assay mimics the enzymatic pathway of MPO-generated HOCl in PMNs, except that it is done in a controlled environment to examine MPO specific effect on S. aureus survival (Figure 5a). As a control, we found there was no difference in the susceptibility of wild type and Teg49 mutants to H2O2 (Figure S4a), likely attributed to catalase production by S. aureus (Das & Bishayi, 2010). Likewise, the generation of hydrogen peroxide from glucose oxidation by glucose oxidase has a negligible effect on the killing of S. aureus (Figure S5a). Accordingly, the design of this assay examined only the enzymatic effect of MPO, and by extension, the efficacy of HOCl-mediated killing of S. aureus in a phagosome-mimicking environment. Our data showed that the Teg49 mutant was much less susceptible to MPO-mediated killing in vitro versus the parent and complement under our testing condition (Figure 5b). The survival of the sarA deletion mutant, at 60% of the inoculum, was similar to the parent. As expected, the spn deletion mutant survived much less than the parent and the complemented mutant. Importantly, the Teg49/spn double mutant displayed similar survival as the spn mutant, indicating that Teg49 relies on spn to improve survivability in HOCl generated by MPO using hydrogen peroxide as the substrate (Figure 5b). Nevertheless, once HOCl is generated (independent from MPO and H2O2), we confirmed that the absence of Teg49 does not impact bacterial survival as would be predicted from the direct inhibition of MPO by SPIN (Figure S4b).
FIGURE 5.
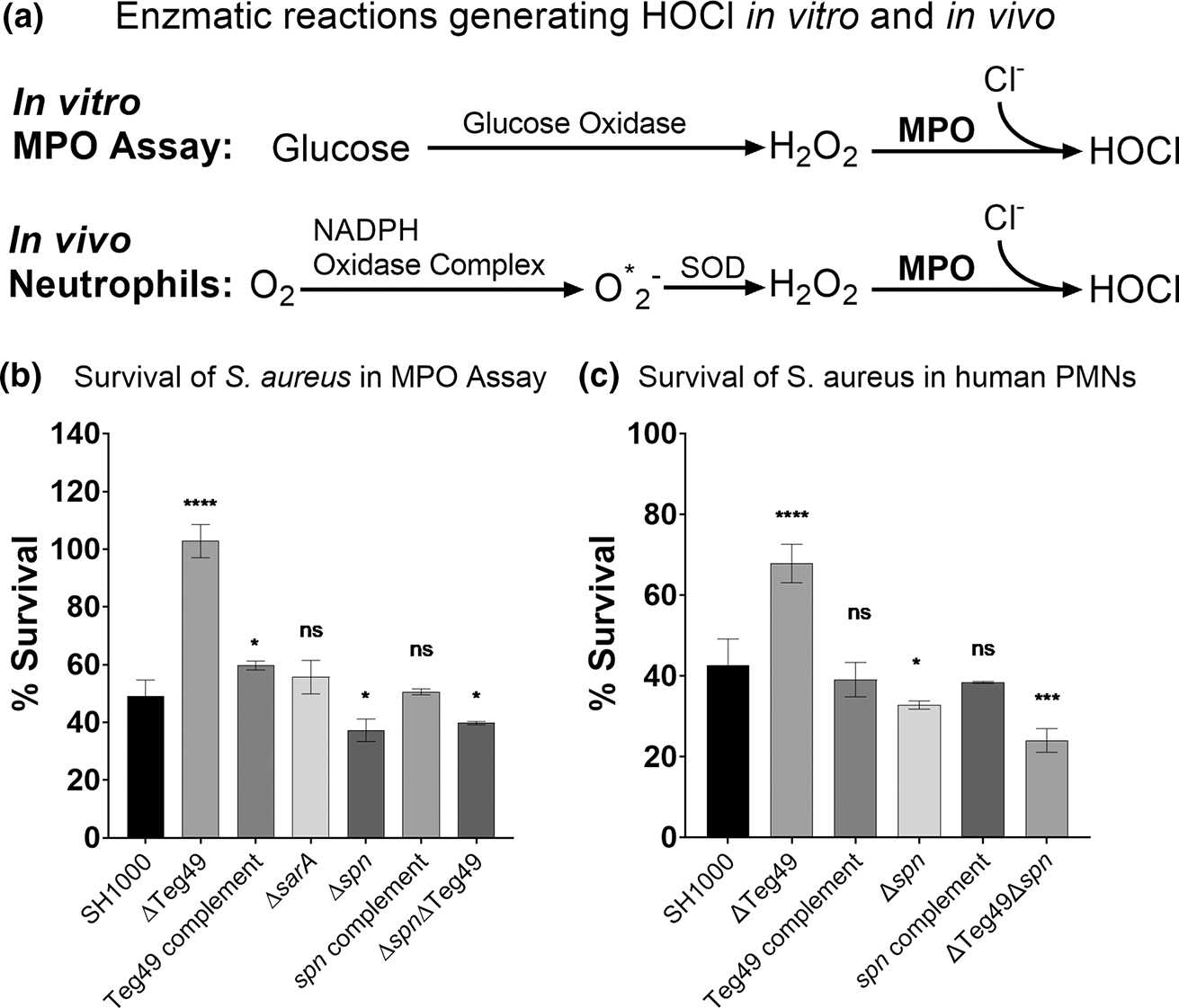
Teg49 mutants have enhanced neutrophil survival mediated through increased spn expression. (a) Diagrams of the enzymatic pathways that generate HOCl in in vitro MPO assay alongside the physiological enzymatic pathway in neutrophils that generates HOCl. In both cases, Teg49 regulation of spn affects the last step, MPO activity. In the in vitro assay, glucose oxidase is used to generate hydrogen peroxide, as compared to in vivo in neutrophils that require NADPH oxidase in the presence of oxygen to generate oxygen radicals (O*2−), and then superoxide dismutase (SOD) to yield H2O2. In both cases, MPO uses H2O2 and chloride to generate hypochlorous acid (HOCl), a powerful antimicrobial antioxidant. (b) Myeloperoxidase assay looking at cell survival in an MPO assay developed in vitro. Wild type SH1000, Teg49 mutant, complement, spn deletion mutant, complement, and double mutant were compared. A sarA mutant was added as a control. Significance is measured using a one-way ANOVA using Dunnett’s multiple tests. p-values are * = .05, ** = .01, *** = .001, **** ≤ .001. (c) Bacterial survival in human PMNs which have phagocytosed Staphylococcus aureus for a 30-min interval at an initial MOI of 8. The strains used are similar to those in (b). Significance is measured using a one-way ANOVA using Dunnett’s multiple tests. p-values are * = .05, ** = .01, *** = .001, **** ≤ .001.
We also evaluated the survival of Teg49 mutant ex vivo with human PMNs isolated from whole blood. After mixing at an MOI of 8 bacteria per neutrophil, the bacteria were allowed to be phagocytosed for 30 min followed by removal of extracellular bacteria with gentamicin at 200 μg/ml. Intracellular survival inside PMN phagosomes were then determined after 30 min. As shown in Figure 5c, the Teg49 mutant exhibited increased survival compared to the wild type SH1000 and complement. As expected, the spn mutant had decreased survival vs. the parent. Remarkably, the Teg49/spn double mutant displayed lower survival than the Teg49 mutant and the parent, indicating that the Teg49 mutant was dependent on spn for resistance to PMN-mediated killing. This discrepancy was not likely due to cationic peptides found inside phagosomes because the Teg49 mutant and the wild type were equally susceptible to cationic membrane protein in terms of MIC to polymyxin B (Figure S5b). Overall, these results with human PMN mirrored the in vitro MPO assay in S. aureus survival.
3 |. DISCUSSION
Teg49 is a sRNA located between the P3 and the proximal P1 promoters upstream of the sarA ORF and is generated from the cleavage of the sarA P3 transcript (Manna et al., 2018). This dependence of Teg49 on sarA P3 transcription led us to wonder if Teg49 has a phenotype distinct from SarA. Accordingly, we conducted an exploratory RNA-Seq study using a Teg49 mutant, yielding a small number of genes that were altered in the Teg49 mutant of SH1000 but not in the isogenic sarA mutant (Table 2) or the parental strain. Focusing on virulence, we were particularly interested in the virulence factor spn (SAOUHSC_00401) since it was present in multiple RNA-Seq comparisons from this and previous studies (Manna et al., 2018). The gene spn encodes SPIN, a 102-residue long propeptide that is secreted and then cleaved to generate the 73-residue mature peptide which has been shown to bind MPO in its active site and thus interfere with the production of HOCl, a powerful antimicrobial oxidant in PMN phagosomes after phagocytosis (de Jong et al., 2017; Guerra et al., 2017; Kettle et al., 2007; Klebanoff et al., 2013). Besides spn, there were also several purine synthesis (pur) and riboflavin-related syntheses (rib) genes in the RNA-Seq dataset that appear to be up-regulated in the Teg49 mutant. However, additional analysis of purH and ribD by Nanostring nCounter did not reveal significant upregulation of these genes in the Teg49 mutant, so these were not pursued further.
To verify the regulation of spn by Teg49, we took advantage of the sensitivity and specificity of Nanostring nCounter. The data showed a clear delineation between SarA-regulated and Teg49-regulated genes (Figure 1a). The Teg49 mutant had no effect on known sarA regulated targets like aureolysin (aur) and protein A (spa). In contrast, the Teg49 mutant consistently displayed increased levels of spn expression vs. the parent whereas the sarA locus has no effect on spn, as corroborated by qRT-PCR (Figure 1b).
Based on our experimentally derived +1 start site, we designed a promoter fusion linking the spn promoter to the GFP reporter plasmid which was then introduced into the wild type and isogenic Teg49 mutant strain. These transcriptional studies revealed that Teg49 does not affect the spn promoter directly. We next explored if Teg49 base-pairs with spn mRNA for this interaction. Due to the observation that the interaction between two RNAs can be as few as 4 bases, we undertook an in-silico analysis with IntaRNA to reveal a potential 9-base site for interaction between Teg49 and spn mRNA (Figure 3). The interaction site is projected to be stable with negative hybridization energy of −12.23 kcal/mol. In addition, the site of interaction in Teg49 is predicted to be exposed to facilitate interaction with spn mRNA (Brantl, 2007; Georg et al., 2020; Storz et al., 2011). More specifically, the predicted interaction site for Teg49 (nt 61–69 in Figure 3b,c) resides within hairpin loop 2, one of the three stable open loops in Teg49 that interacts with an open and accessible bulge (nt 240–248) within the coding sequence of spn mRNA (Figure 3). Besides a bulging G nucleotide at position 60 in Teg49, the preceding 5′-CCUU-3′ sequence (nt 56–59) is complementary to positions 249–252 on the spn mRNA (Figure 3c). It is likely that IntaRNA did not match these nucleotides due to the high stability of the original sRNA or mRNA structures. It is also plausible that this region immediately outside the predicted interactive site may interact once an initial seed region binds since it has been found that a small region is enough to initiate seed binding before RNAs are more free to interact with each other (Brantl, 2007; Vogel & Wagner, 2007; Wagner et al., 2002). Despite the logic of the IntaRNA prediction, it was surprising that hairpin loop 1, previously shown to affect transcript stability of the sarA P3 transcript (Kim et al., 2014), was not predicted to be involved in this interaction. Overall, this mode of regulation differs from that of spn by saeRS where the cognate binding site for SaeR was found to reside within the spn promoter region (Kim et al., 2014; Nygaard et al., 2010).
As mentioned, the mechanism of direct regulation was very much suggested by structural and sequence features in Teg49. Besides being highly structured, the 5′ end of Teg49 stood out in terms of GC content. The sarA locus around Teg49 has GC content slightly below the standard ~30% GC content of the entire genome in S. aureus (Gillaspy et al., 2006; Kuroda et al., 2001). In contrast, concordant with their putative roles as interactive regions, hairpin loops 1 and 2 of Teg49 have higher than average GC content of 57% (loop 1) and 50% (loop 2), respectively. Hairpin loop 3 has a standard value of 33% (loop 3). Notably, the stems supporting the hairpin loops in the Teg49 structure have a low GC content of 26%, 16%, and 26% for stems 1–3, respectively (Figure S2b).
To verify the predicted Teg49/spn RNA–RNA interaction, we made reciprocal mutations to disrupt the binding sites of either Teg49 or spn. Besides mutation in hairpin loop 2 of Teg49, we also constructed mutants in the exposed hairpin loops 1 and 3 where nucleotides were scrambled to disrupt possible binding sites in these loops, but not the structure of the sRNA. For spn mRNA, we disrupted the putative binding regions using both synonymous and non-synonymous mutations where the third base of 4 codons in the synonymous mutant was altered to disrupt binding but not the predicted amino acid sequence or the predicted spn RNA structure. Amazingly, both the hairpin loop 2 mutant of Teg49 and the corresponding spn mutants exhibited significant upregulation of spn mRNA as would be expected in the loss of Teg49-spn mRNA interaction, providing evidence that Teg49 acts as a trans-acting sRNA to disrupt spn mRNA expression, likely via a post-transcriptional mechanism. Interestingly, the magnitude of the spn transcript levels in the spn binding site mutant (Figure 4b) was much higher than that in the Teg49 hairpin loop 2 mutant (Figure 4a), suggesting possibly other effects on spn mRNA (e.g., modification of RNA folding to obscure a RNase cleavage site on spn mRNA) since its magnitude was similar to that of a RNase III mutant (Figure 4d).
To further develop the model of direct Teg49-mediated regulation of spn, we examined the spn transcript stability as a result of the interaction. Using isogenic Teg49 strain in the SH1000 background, Northern blots revealed higher levels of the spn transcript over time in the Teg49 mutant than the parent after the addition of rifampin to halt transcription. These data implied that the spn mRNA had a longer half-life in the Teg49 mutant vs. the parent, corresponding to a model of direct regulation of spn mRNA by Teg49.
S. aureus is well adapted to overcome the human immune system. PMNs, in particular, represent the first line of defense at sites of infections (Flannagan et al., 2009; Guerra et al., 2017; Hampton et al., 1996; Klebanoff et al., 2013). Upon phagocytosis, S. aureus is trapped inside the phagosome followed by fusion with a lysosome to form the phagolysosome. Within the phagolysosomes are reactive oxygen species including superoxide anion, hydrogen peroxide, and HOCl. S. aureus secretes superoxide dismutases (Sod1 and Sod2) and catalase to counteract the effect of the superoxide anion and hydrogen peroxide, respectively (Karavolos et al., 2003). HOCl, a powerful oxidant, is the byproduct of the MPO enzyme, which is secreted into the phagosome upon fusion with the lysosome. Since HOCl possesses potent anti-staphylococcal activity, it has been surmised that S. aureus likely harbors an anti-HOCl strategy. Remarkably, de Jong et al have discovered SPIN, a protein secreted by S. aureus inside the phagosome in response to ROS stress. Besides the sRNA Teg49, it has been reported that the two-component regulatory system SaeRS in S. aureus positively regulates spn at the transcriptional level (Guerra et al., 2016; Nygaard et al., 2010). This was also observed in this study, presumably via a SaeR binding site on the spn promoter (Figure 4d).
As SPIN interferes with MPO activity in vitro, we wanted to determine the effect of Teg49 using in vitro phagosome mimicking conditions and in human PMNs ex vivo (Figure 6). In our case, we found that titration of glucose oxidase and MPO is critical to the success of the in vitro assay since a high level of hydrogen peroxide and MPO can overwhelm catalase and SPIN activity from S. aureus, respectively. Once optimized, we clearly observed that the Teg49 mutant, expressing a higher level of SPIN, can survive better than the parent/complement (Figure 5a). Remarkably, our PMN survival assay for the S. aureus strain set mirrored that of the in vitro MPO assay. There were concerns that Teg49 may affect resistance to hydrogen peroxide, HOCl, or cationic antimicrobial peptides inside the phagolysosome to account for enhanced survival (Fraunholz & Sinha, 2012; Horn et al., 2018). However, we have found that the Teg49 mutant and its isogenic parent exhibited identical MIC to polymyxin B. In addition, there were no differences between the wild type and Teg49 mutant in sensitivity to hydrogen peroxidase and HOCl (Figure S5a,b). Thus, the improved survival of Teg49 mutants in both assays is likely due to differences in MPO activity which is SPIN dependent. In particular, the Teg49/spn double mutant in both assays had reduced survival as compared to the Teg49 mutant, indicating that improved survival of the Teg49 mutant was attributable to increased spn expression (Figure 5a,b). We still found killing of some Teg49 mutant cells even though survival of the Teg49 mutant was much better than the wild type in both assays. Interestingly, Teg49 mutants seemed more sensitive to human PMN than to MPO alone, likely due to other various PMN factors known to kill bacteria in phagosomes (Collins et al., 2002; Ernst et al., 2009; Flannagan et al., 2009).
FIGURE 6.
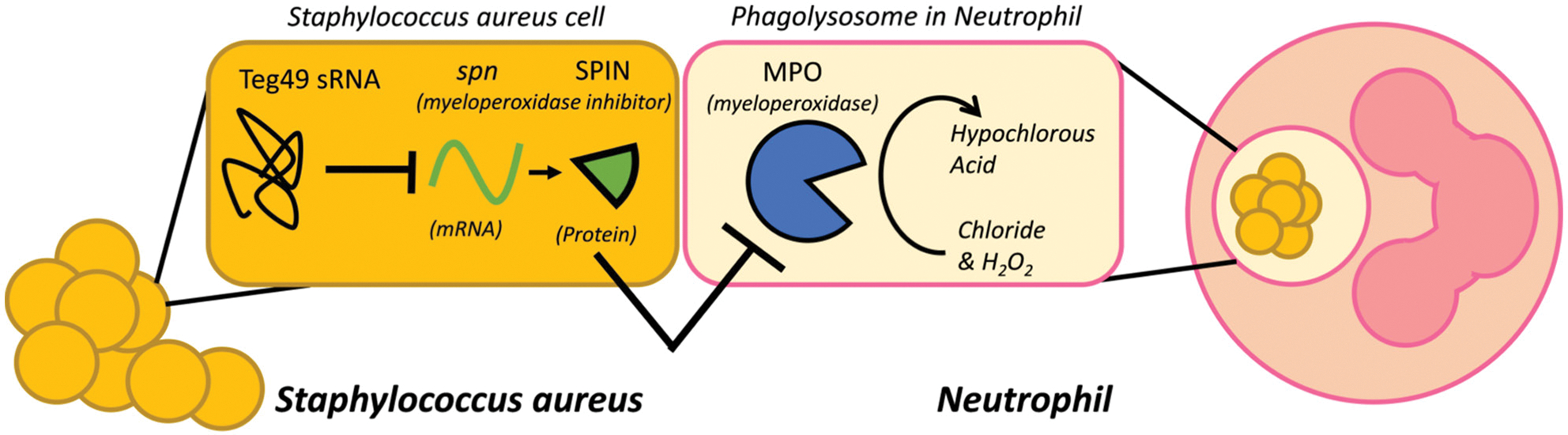
Model for Teg49 function in neutrophil myeloperoxidase tolerance. Model of how regulation of spn mRNA by Teg49 in S. aureus increases survival in PMN phagolysosomes. In this representative image, myeloperoxidase activity within the phagolysosome of phagocytes (PMNs in this case) generates hypochlorous acid to kill bacterial cells. Myeloperoxidase inhibitor (encoded by spn) inhibits MPO activity to reduce HOCl production. In this study, we provide an extra layer of regulation by showing that sRNA Teg49 can repress spn expression post-transcriptionally by base-pairing with spn mRNA to facilitate degradation and hence reduce translation.
The sarA P3 transcript, as the precursor to Teg49, is activated as the cells enter the stationary phase due to its dependency on the stress-induced SigB promoter. That would mean under exponential growth, the expression of Teg49 is low and SPIN high. Upon stress, Teg49 is expressed to repress SPIN, except under a ROS generating environment, a condition known to activate the spn promoter (Guerra et al., 2016). As Teg49 does not impact growth in TSB or MHB (Figure S1a), it appears that Teg49 may modulate intracellular levels of spn to prevent overexpression during the stationary phase, falling in line with the idea that sRNAs often act more as dials that can limit significant mRNA fluctuations rather than on/off switches in gene regulation (Kelly et al., 2018; Mehta et al., 2008; Waters & Storz, 2009).
Teg49 is shown to be a trans-acting regulatory sRNA that directly binds spn mRNA and hence plays a crucial role to the survival of S. aureus within a phagolysosome. While the folding and structure of this sRNA are clearly important to its function, more work need to be done to explore its structure–function relationship. Like many other sRNAs, the exposed GC-rich sequence on hairpin-loops seems to be the key interaction site of this sRNA for its function (Jørgensen et al., 2020). Apart from previous studies examining the ability of the hairpin loop 1 to stabilize the parental sarA P3 mRNA, hairpin 2 loop has now been shown to play an interacting role with its target spn mRNA. Interestingly, neither of these loops has been shown to be crucial to its excision and maturation of Teg49 from the parental sarA P3 mRNA. Nevertheless, it remains to be seen whether the remaining features of Teg49 play other functional roles including the broader interactions of Teg49 with other regulatory targets.
4 |. EXPERIMENTAL PROCEDURES
4.1 |. Bacterial strains, plasmids, primers, and growth conditions
The bacterial strains, plasmids, and primers used in this study are summarized in Table 1. Mutants were made in the S. aureus SH1000 background. Escherichia coli IM08B was used as the initial recipient for the transformation of shuttle plasmids (Monk et al., 2015). E. coli strains were grown with shaking at 250 rpm at 37°C in LB (Becton Dickinson), or LB with 50 μg/ml ampicillin. S. aureus strains were grown with shaking at 250 rpm at 37 or 30°C in either tryptic soy broth (TSB) or Muller Hinton broth (MHB) (Becton Dickinson), with antibiotics added when needed (10 μg/ml erythromycin, 5 μg/ml chloramphenicol, 10 μg/ml tetracycline, or 40 μg/ml kanamycin).
4.2 |. Genetic manipulations in E. coli and S. aureus
Temperature-sensitive shuttle vector pMAD was used for homologous recombination in S. aureus as previously described (Arnaud et al., 2004). The Teg49 deletion mutant was confirmed to not express Teg49 by Northern blots and subsequently complemented using pMAD::Teg49 for insertion into the chromosome via homologous recombination. Complementation in trans- for Teg49 was performed with the recombinant vector pEPSA5::Teg49 and induced with 2% xylose in broth for expression (Kim et al., 2014). Cloning and plasmid mutagenesis was performed using PCR and Pfu and Taq polymerases (NEB), and clones were verified by restriction digest and DNA sequencing as described previously.
To determine the contribution of Teg49 sRNA in regulation, we have constructed 70-bp chromosomal deletion of the Teg49 region in S. aureus strain SH1000 using routine procedures as described (Manna et al., 2018). For the Teg49 deletion construct, a 2.0 kb DNA fragment was amplified that contained the intact sarA locus by PCR using primers flanking with BamHI site at both ends and chromosomal DNAs as the template for SH1000. The DNA fragment was cloned into a shuttle vector pMAD (β-gal, ermR). Site-specific mutagenesis was performed with KLD enzymes mix (NE Biolabs Inc, USA) and transformed to E. coli IM08B for shuttling into SH1000 strain by electroporation. Homologous recombination using temperature and colorimetric selection were done to achieve the final chromosomal deletion mutant. Plasmid DNA was isolated and restriction digested with BamHI and sequencing was used during each crucial step to confirm constructs and mutants. Similarly, mutants for Teg49 and sarA start site were constructed using the KLD enzyme mix and 2.0 kB DNA fragments of the sarA locus. Complementation for the sarA locus was done using an allelic exchange with a wild type sarA locus in pMAD. Mutants in spn were constructed by amplifying a 2.2 kb DNA fragment of the spn locus and cloning it into pMAD shuttle vector using the NEBuilder HiFi DNA assembly kit (NEB) with the Gibson assembly method. Deletion of spn was made using inserted flanking SalI and XhoI restriction digest sites, and mutations in the spn locus were done using KLD and allelic exchange as described for Teg49. The pALC1484:spn promoter-gfp vector was made using inserted EcoRI and XbaI restriction digest sites as described for Teg49. Primers for the above recombinant clones are listed in Table S1.
4.3 |. Isolation of total cellular RNA, northern blot hybridization, and RNA stability assay
RNA isolation and purification and subsequent Northern blot analysis were performed as described previously (Cheung et al., 1994; Manna et al., 2018; Oriol et al., 2021). Briefly, 15 μg of total RNAs were run on 8% polyacrylamide-7 M urea gels and transferred onto Hybond-N+ membranes (Amersham, USA) to detect sRNA. For mRNA, 10 μg total RNA was run on 1% agarose-formaldehyde gels and transferred onto Hybond-H+ membranes (Amersham, USA). For probes, DNA fragments containing the coding sequences of sarA and spn and sequence of Teg49 were amplified by PCR (Table S1) and gel purified. DNA probes were radiolabeled with [α-32 P] dCTP using the random-primed DNA-labeling kit (Roche Diagnostics GmbH) and hybridized under aqueous-phase conditions at 65°C. Blots were washed, exposed overnight on a phosphorimager screen, and scanned by a Typhoon FLA 9500 Molecular Imager (GE, USA). Loading controls were performed with radiolabeled transfer-messenger RNA (Tm RNA) or 16S RNA ethidium bromide labeling. RNA stability assays were done by stopping transcription at 4 h growth using 200 μg/ml rifampicin and quick-freezing cells at given timepoints. RNA extraction and preparation were done immediately as described above. Densitometry with ImageJ was used to measure density counts in lane measurements, and compare density using integrated lane density over a defined region of interest with ImageJ/Fiji Tools (Belasco & Brawerman, 2012; Chen et al., 2008; Villa et al., 2021).
4.4 |. RNA-Seq analysis
Overnight cultures were diluted 1:1000 in fresh TSB to an initial OD600 of about 0.05 and grown at 37°C in TSB with or without 5 μg/ml chloramphenicol for 3 h with 2% xylose in broth. Two replicate cultures for each sample were grown, and total cellular RNAs from two biological replicates were extracted as described previously (Kim et al., 2014). Purelink RNA Mini Kit (Invitrogen) column purification was performed for all samples. The absence of DNA was checked by qPCR, RNA was quantified using Qubit (Life Technologies), and RNA integrity was verified on a Bioanalyzer (Agilent). One microgram of total RNA was ribo-depleted with a bacterial Ribo-Zero kit from Illumina. A TruSeq RNA stranded kit from Illumina was used for cDNA library preparation. The library quantity was measured with Qubit, and quality was assessed on a Tapestation on a DNA high-sensitivity chip (Agilent Technologies). The libraries were pooled at equimolarity and loaded at 2 nM for clustering.
Oriented 75-base single-read sequencing was performed on an Illumina HiSeq 4000 sequencer, yielding a ~ 40 M reads/sample reads per sample. Adaptor trimming was done before reads were mapped to S. aureus NCTC8325 (NC_007795.1) and gene expression was determined using CLC Genomics Workbench 11 (QIAGEN CLC Genomics Workbench 20.0).
Final RNA-Seq analysis and data analysis were carried out using previously described procedures (Manna et al., 2018). Gene expression analysis and statistics were done in R v3.36.0 using the edgeR package (Robinson et al., 2010) following a bioinformatics protocol described previously (Anders et al., 2013). Genes whose expression was less than one read per million in two replicates of the samples were filtered out. Pairwise comparisons were performed with exact tests for statistical significance (FDR-adjusted p < .05) (Anders et al., 2013; Robinson et al., 2010).
4.5 |. Nanostring nCounter
Nanostring nCounter system (Nanostring Technologies, Seattle, WA) was used with the methods described previously (Geiss et al., 2008). Briefly, cells were grown overnight, re-seeded 1:1000 in fresh media, grown at 37°C with shaking, and harvested at 4 h. RNA was extracted, treated with TurboDNase (Invitrogen), and 100 ng of total RNA was hybridized overnight with reporter and capture probes at 65°C according to the manufacturer’s protocol. A custom-designed code-set S. aureus genes, with code-set design and construction provided by Nanostring Technologies, was used to measure samples performed in biologic duplicates to simultaneously detect the 12 genes assayed in this study. Targets were counted on the nCounter machine. nSolver Analysis Software 4.0 (Nanostring Technologies, Seattle, WA) was used to calculate differences in gene expression. Statistical analysis for data sets was analyzed with GraphPad Prism 8.0 (GraphPad Software, San Diego, CA).
4.6 |. qRT-PCR analysis
qRT-PCR was performed as previously described (Manna et al., 2018). Overnight cultures of S. aureus were re-seeded in fresh media at 1:1000 and grown to the indicated time. Total cellular RNAs were extracted and treated with Turbo DNase (Invitrogen) as described previously (Kim et al., 2014). cDNA was generated using RevertAid-H minus First-Strand cDNA synthesis mix (ThermoScientific) with the total RNA sample that has been treated with DNase I. Ct values were measured on a C1000 Thermal Cycler with a CFX96 Real-Time System (BioRad). qRT-PCR analysis was used to amplify cDNA using the primers (Table S1) for spn and the reference gene rpoB. Triplicate samples were analyzed, and relative fold expression was calculated using the threshold cycle (2−ΔΔCT).
4.7 |. 5′ RACE assay
Rapid amplification of 5′ complementary DNA ends (5′-RACE) was performed as previously described (Scotto–Lavino et al., 2006). Briefly, RNA was extracted and purified using the geneJet RNA purification kit (ThermoScientific), treated with TurboDNAse (Invitrogen), modified with terminal transferase (New England BioLabs), and re-purified. The SMARter 5′/3′ RACE kit (Takara) was used to generate cDNA, and gene-specific primers (Table S1) were used to amplify spn before sequencing.
4.8 |. Promoter fusion
The promoter region comprising 150-bp upstream of the spn translation start site was cloned into a promoterless GFPuvr plasmid (pALC1484) as previously described (Cheung et al., 1997) and transformed into E. coli IM08B. Recombinant plasmids were verified by restriction digest and DNA sequencing. Promoter fusion constructs were then electroporated into wild type SH1000 or Teg49 mutant. Overnight cultures were diluted 1:100 in 10 ml of TSB and grown at 37°C with shaking at 250 rpm. At designated times, 200 μl of culture was removed from each tube, transferred to a 96-well microtiter plate and read for optical density and florescence simultaneously with a Tecan Infinite M1000 microplate reader, using OD600 for growth and excitation and emission wavelength of 488 and 510 nm, respectively, for fluorescence.
4.9 |. RNA folding and IntaRNA prediction
RNA structures were generated using RNAfold from ViennaRNA Package 2.0, using the Turner 2004 model at 37°C with standard folding parameters (Lorenz et al., 2011; Mathews et al., 2004). This was used as part of the FreiburgRNA Tools suite (Smith et al., 2010). Additional validation using ViennaRNA was done with Turner 1999 and Andronescu 2007 models, both predicting almost identical structures (Andronescu et al., 2007; Zuker et al., 1999). RNA structure software was also used to validate and predict a similar folding of RNA using standard conditions (Reuter & Mathews, 2010). The interactions between Teg49 and spn mRNA were predicted using IntaRNA (Mann et al., 2017). Input RNAs were the complete Teg49 sequence and the spn sequence starting from the 5′ transcriptional start site to the predicted 3′ terminator 35 bp downstream from the stop codon.
4.10 |. MIC and stress tests
For H2O2 and NaOCl stress tests, cells were grown overnight in MHB. These cells were serially diluted 1:1000 in fresh MHB to an OD600 of ~0.05. Cultures were then grown for 2 h in Tecan infinity 1000 at 37°C with shaking, at which point H2O2 (18 mM or 1.84 M) or NaOCl (35 mM or 3.5 M) was added into the culture. Plates were then returned to Tecan for further readings.
MIC assay with PMB (Sigma) was done according to the CLSI standard (CLSI, 2015). Cells were grown in calcium and magnesium adjusted MHB overnight, diluted 10-folds to 1 × 106 CFU/ml. These samples were mixed with serial dilutions of PMB in 96-well plates and incubated for 18 h at 37°C to determine MIC.
4.11 |. Myeloperoxidase assay
The myeloperoxidase killing assay was adapted from protocols as described previously (de Jong et al., 2017; Denys et al., 2011). Glucose oxidase (GO) and MPO concentrations were adjusted to have almost 0% and ~50% bactericidal effects respectively in SH1000. Briefly, overnight cultures of S. aureus grown in TSB were re-seeded in fresh MHB media 1:1000 to an approximate OD600 of 0.05 and grown to an OD600 of 0.5 with shaking at 37°C. Cells were then washed with HBSS (Corning) and diluted 1:100 in 300 mM glucose HBSS. This solution was then added 1:1 to an enzyme solution of 20 nM glucose oxidase from Aspergillus (Sigma–Aldrich), and 220 nM myeloperoxidase from human leukocytes (Sigma Aldrich). Samples were incubated with shaking at 37°C for 1 h, after which excess catalase from bovine liver (Sigma Aldrich) was added to stop the reaction. Samples were serially diluted in TSB, plated on TSA plated overnight at 37°C, and enumerated the next day. Statistical analysis was done with GraphPad Prism 8.0 (GraphPad Software, San Diego, CA).
4.12 |. Polymorphonuclear cell killing assay
Human neutrophils were extracted according to previous studies (Cho et al., 2021). Venous blood from healthy human donors was obtained with the consent and protocols approved by the Committee for the Protection of Human Subjects of the Geisel School of Medicine at Dartmouth (Approval #: STUDY00029984). Formal verbal consent was obtained from the studied subjects. Only adults are included in this study.
Polymorphonuclear cell assays were done in accordance with previously described protocols (Mikheyeva et al., 2019; Nauseef, 2007). Briefly, fresh venous blood was diluted within prewarmed RPMI and overlaid on 10 ml of Ficoll–Hypaque. Blood components were separated by 30 min spin at 500 g at room temperature. All layers except the PMN-erythrocyte pellet were discarded. Pellet was resuspended in 20 ml of Hank’s balanced salt solution (HBSS) without Mg2+/Ca2+, and 20 ml of 6% dextran was added. The suspension was incubated for 20 min at room temperature, and the PMN layer was collected and centrifuged. Contaminating erythrocytes were lysed with sterile H2O and tonicity restored with saline solution. PMNs were washed with HBSS and resuspended to 1 × 105 cells ml−1 in RPMI with 5% serum.
Bacterial cells were seeded from overnight culture and grown to OD600 0.5 in MHB, then opsonized in 5% human serum for 15 min at 37°C in pre-warmed RPMI. Infection of PMNs was done at a multiplicity of infection of eight bacteria per PMN and reaction mixtures were rotated at 37°C for 30 min. Non-opsonized bacteria that were not phagocytosed were killed using 200 μg/ml gentamicin in RPMI and PMNs with engulfed bacteria were then further incubated in RPMI at 37°C. The PMN-bacteria mixtures were sampled at 0 and 30 min post-gentamicin treatment. PMN samples were then lysed in a 0.1% Triton X-100 in PBS, and serial dilutions were plated on TSB. CFUs were counted the next day after overnight growth at 37°C. Statistical analysis was done with GraphPad Prism 8.0 (GraphPad Software, San Diego, CA).
Supplementary Material
ACKNOWLEDGMENTS
We acknowledge the Genomics and Molecular Biology Shared Resources at the Norris Cotton Cancer Center at Dartmouth. This research was supported by a training grant (T32HL134598) from the NIH. This research was also supported by research grants from the CF foundation to A.L.C (CHEUNG20G0) and W.F.C.R (Rigby20GO) as well as from NCI (5P30 CA023108-40). We thank our colleagues from the M2P2 department for advice, especially Zhongyou Li for expertise in RNA-Seq analysis, and the Dartmouth analytics core (DAC) for help wrangling data in the RNA-Seqdataset.
Funding information
Cystic Fibrosis Foundation, Grant/Award Number: CHEUNG20G0 and rigby20G0; National Cancer Institute, Grant/Award Number: 5P30CA023108-40; NIH Heart and Lung Institute, Grant/Award Number: T32 HL134598
Footnotes
ETHICS STATEMENT
All studies with human subjects have obtained IRB approval from Dartmouth College concerning ethical treatment of human participants and gathering of informed consent, as appropriate. There are no conflicts of interests among all authors. All authors confirm to agree with the final ethics statement.
SUPPORTING INFORMATION
Additional supporting information may be found in the online version of the article at the publisher’s website.
DATA AVAILABILITY STATEMENT
Data available on request due to privacy/ethical restrictions.
REFERENCES
- Anders S, McCarthy DJ, Chen Y, Okoniewski M, Smyth GK, Huber W et al. (2013) Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nature Protocols, 8, 1765–1786. [DOI] [PubMed] [Google Scholar]
- Andronescu M, Condon A, Hoos HH, Mathews DH & Murphy KP (2007) Efficient parameter estimation for RNA secondary structure prediction. Bioinformatics, 23, i19–i28. [DOI] [PubMed] [Google Scholar]
- Arnaud M, Chastanet A & Debarbouille M (2004) New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive bacteria. Applied and Environmental Microbiology, 70, 6887–6891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballal A & Manna AC (2010) Control of thioredoxin reductase gene (trxB) transcription by SarA in Staphylococcus aureus. Journal of Bacteriology, 192, 336–345. 10.1128/jb.01202-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrientos L, Mercier N, Lalaouna D & Caldelari I (2021) Assembling the current pieces: the puzzle of RNA-mediated regulation in Staphylococcus aureus. Frontiers in Microbiology, 12, 706690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bayer MG, Heinrichs JH & Cheung AL (1996) The molecular architecture of the sar locus in Staphylococcus aureus. Journal of Bacteriology, 178, 4563–4570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beaume M, Hernandez D, Farinelli L, Deluen C, Linder P, Gaspin C et al. (2010) Cartography of methicillin-Resistant S. aureus transcripts: detection, orientation and temporal expression during growth phase and stress conditions. PLoS One, 5, e10725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belasco JG & Brawerman G (2012) Control of messenger RNA stability. New York, NY: Elsevier. [Google Scholar]
- Brantl S (2007) Regulatory mechanisms employed by cis-encoded antisense RNAs. Current Opinion in Microbiology, 10, 102–109. [DOI] [PubMed] [Google Scholar]
- Chan PF & Foster SJ (1998) Role of SarA in virulence determinant production and environmental signal transduction in Staphylococcus aureus. Journal of Bacteriology, 180, 6232–6241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C-YA, Ezzeddine N & Shyu A-B. (2008) Messenger RNA half-life measurements in mammalian cells. Methods in Enzymology, 448, 335–357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung AL, Bayer AS, Zhang G, Gresham H & Xiong Y-Q (2004) Regulation of virulence determinants in vitro and in vivo in Staphylococcus aureus. FEMS Immunology & Medical Microbiology, 40, 1–9. [DOI] [PubMed] [Google Scholar]
- Cheung AL, Bayer MG & Heinrichs JH (1997) Sar genetic determinants necessary for transcription of RNAII and RNAIII in the agr locus of Staphylococcus aureus. Journal of Bacteriology, 179, 3963–3971. Available from: 10.1128/jb.179.12.3963-3971.1997 Accessed 17 February 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung AL, Eberhardt KJ & Fischetti VA (1994) A method to isolate RNA from gram-positive bacteria and mycobacteria. Analytical Biochemistry, 222, 511–514. [DOI] [PubMed] [Google Scholar]
- Cheung AL, Nishina KA, Trotonda MP & Tamber S (2008) The SarA protein family of Staphylococcus aureus. The International Journal of Biochemistry & Cell Biology, 40, 355–361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung GYC, Bae JS & Otto M (2021) Pathogenicity and virulence of Staphylococcus aureus. Virulence, 12, 547–569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chien Y, Manna AC, Projan SJ & Cheung AL (1999) SarA, a global regulator of virulence determinants in Staphylococcus aureus, binds to a conserved motif essential for Sar-dependent gene regulation. The Journal of Biological Chemistry, 274, 37169–37176. [DOI] [PubMed] [Google Scholar]
- Cho J, Costa SK, Wierzbicki RM, Rigby WFC & Cheung AL (2021) The extracellular loop of the membrane permease VraG interacts with GraS to sense cationic antimicrobial peptides in Staphylococcus aureus. PLoS Pathogens, 17, e1009338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CLSI. (2015) Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard. CLSI document M07–A10. [Google Scholar]
- Collins LV, Kristian SA, Weidenmaier C, Faigle M, van Kessel KPM, Strijp JAG et al. (2002) Staphylococcus aureus strains lacking d-alanine modifications of teichoic acids are highly susceptible to human neutrophil killing and are virulence attenuated in mice. The Journal of Infectious Diseases, 186, 214–219. [DOI] [PubMed] [Google Scholar]
- Das D & Bishayi B (2010) Contribution of catalase and superoxide dismutase to the intracellular survival of clinical isolates of Staphylococcus aureus in murine macrophages. Indian Journal of Microbiology, 50, 375–384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denys GA, Grover P, O’Hanley P & Stephens JT Jr. (2011) In vitro antibacterial activity of E-101 solution, a novel myeloperoxidase-mediated antimicrobial, against gram-positive and gram-negative pathogens. Journal of Antimicrobial Chemotherapy, 66, 335–342. [DOI] [PubMed] [Google Scholar]
- Ernst CM, Staubitz P, Mishra NN, Yang S-J, Hornig G, Kalbacher H et al. (2009) The bacterial defensin resistance protein MprF consists of separable domains for lipid Lysinylation and antimicrobial peptide repulsion. PLoS Pathogens, 5, e1000660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Felden B, Vandenesch F, Bouloc P & Romby P (2011) The Staphylococcus aureus RNome and its commitment to virulence. PLoS Pathogens, 7, e1002006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flannagan RS, Cosío G & Grinstein S (2009) Antimicrobial mechanisms of phagocytes and bacterial evasion strategies. Nature Reviews. Microbiology, 7, 355–366. [DOI] [PubMed] [Google Scholar]
- Fraunholz M & Sinha B (2012) Intracellular Staphylococcus aureus: live-in and let die. Frontiers in Cellular and Infection Microbiology, 2 10.3389/fcimb.2012.00043 Accessed 23 January 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL et al. (2008) Direct multiplexed measurement of gene expression with color-coded probe pairs. Nature Biotechnology, 26, 317–325. [DOI] [PubMed] [Google Scholar]
- Geissmann T, Chevalier C, Cros MJ, Boisset S, Fechter P et al. (2009) A search for small noncoding RNAs in Staphylococcus aureus reveals a conserved sequence motif for regulation. Nucleic Acids Research, 37, 7239–7257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georg J, Lalaouna D, Hou S, Lott SC, Caldelari I, Marzi S et al. (2020) The power of cooperation: experimental and computational approaches in the functional characterization of bacterial sRNAs. Molecular Microbiology, 113, 603–612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gillaspy AF, Worrell V, Orvis J, Roe BA, Dyer DW & Iandolo JJ (2006) The Staphylococcus aureus NCTC 8325 genome. In: Gram-Positive Pathogens. Washington, DC: John Wiley & Sons, Ltd, pp. 381–412. 10.1128/9781555816513.ch32 Accessed 30 January 2022. [DOI] [Google Scholar]
- Guerra FE, Addison CB, de Jong NWM, Azzolino J, Pallister KB, van Strijp J et al. (2016) Staphylococcus aureus SaeR/S-regulated factors reduce human neutrophil reactive oxygen species production. Journal of Leukocyte Biology, 100, 1005–1010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerra FE, Borgogna TR, Patel DM, Sward EW & Voyich JM (2017) Epic immune battles of history: neutrophils vs. Staphylococcus aureus. Frontiers in Cellular and Infection Microbiology, 7, 286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hampton MB, Kettle AJ & Winterbourn CC (1996) Involvement of superoxide and myeloperoxidase in oxygen-dependent killing of Staphylococcus aureus by neutrophils. Infection and Immunity, 64, 3512–3517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horn J, Stelzner K, Rudel T & Fraunholz M (2018) Inside job: Staphylococcus aureus host-pathogen interactions. International Journal of Medical Microbiology, 308, 607–624. [DOI] [PubMed] [Google Scholar]
- Horsburgh MJ, Aish JL, White IJ, Shaw L, Lithgow JK & Foster SJ (2002) σ B modulates virulence determinant expression and stress resistance: characterization of a functional rsbU strain derived from staphylococcus aureus 8325–4. Journal of Bacteriology, 184, 5457–5467. 10.1128/jb.184.19.5457-5467.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Jong NWM, de Ramyar KX, Guerra FE, Nijland R, Fevre C, Voyich JM et al. (2017) Immune evasion by a staphylococcal inhibitor of myeloperoxidase. Proceedings of the National Academy of Sciences of the United States of America, 114, 9439–9444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jørgensen MG, Pettersen JS & Kallipolitis BH (2020) sRNA-mediated control in bacteria: an increasing diversity of regulatory mechanisms. Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms, 1863, 194504. [DOI] [PubMed] [Google Scholar]
- Kahl BC, Goulian M, van Wamel W, Herrmann M, Simon SM, Kaplan G, et al. (2000) Staphylococcus aureus RN6390 replicates and induces apoptosis in a pulmonary epithelial cell line. Infection and Immunity, 68, 5385–5392. 10.1128/iai.68.9.5385-5392.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karavolos MH, Horsburgh MJ, Ingham E & Foster SJ (2003) Role and regulation of the superoxide dismutases of Staphylococcus aureus. Microbiology, 149, 2749–2758. [DOI] [PubMed] [Google Scholar]
- Kelly CL, Harris AWK, Steel H, Hancock EJ, Heap JT & Papachristodoulou A (2018) Synthetic negative feedback circuits using engineered small RNAs. Nucleic Acids Research, 46, 9875–9889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kettle AJ, Anderson RF, Hampton MB & Winterbourn CC (2007) Reactions of superoxide with myeloperoxidase. Biochemistry, 46, 4888–4897. [DOI] [PubMed] [Google Scholar]
- Kim S, Reyes D, Beaume M, Francois P & Cheung A (2014) Contribution of teg49 small RNA in the 5′ upstream transcriptional region of sarA to virulence in Staphylococcus aureus. Infection and Immunity, 82, 4369–4379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klebanoff SJ, Kettle AJ, Rosen H, Winterbourn CC & Nauseef WM (2013) Myeloperoxidase: a front-line defender against phagocytosed microorganisms. Journal of Leukocyte Biology, 93, 185–198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroda M, Ohta T, Uchiyama I, Baba T, Yuzawa H, Kobayashi I et al. (2001) Whole genome sequencing of meticillin-resistant Staphylococcus aureus. The Lancet, 357, 1225–1240. [DOI] [PubMed] [Google Scholar]
- Lorenz R, Bernhart SH, Hönerzu Siederdissen C, Tafer H, Flamm C, Stadler PF et al. (2011) ViennaRNA Package 2.0. Algorithms for Molecular Biology, 6, 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowy FD (1998) Staphylococcus aureus infections. New England Journal of Medicine, 339, 520–532. [DOI] [PubMed] [Google Scholar]
- Mann M, Wright PR & Backofen R (2017) IntaRNA 2.0: enhanced and customizable prediction of RNA–RNA interactions. Nucleic Acids Research, 45, W435–W439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manna AC, Bayer MG & Cheung AL (1998) Transcriptional analysis of different promoters in the Sar locus in Staphylococcus aureus. Journal of Bacteriology, 180, 3828–3836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manna AC, Kim S, Cengher L, Corvaglia A, Leo S, Francois P et al. (2018) Small RNA teg49 is derived from a sarA transcript and regulates virulence genes independent of SarA in Staphylococcus aureus. Infection and Immunity, 86, 617–635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mathews DH, Disney MD, Childs JL, Schroeder SJ, Zuker M & Turner DH (2004) Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. PNAS, 101, 7287–7292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta P, Goyal S & Wingreen N (2008) A quantitative comparison of sRNA-based and protein-based gene regulation. Molecular Systems Biology, 4, 221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mikheyeva IV, Thomas JM, Kolar SL, Corvaglia A-R, Gaϊa N, Leo S et al. (2019) YpdA, a putative bacillithiol disulfide reductase, contributes to cellular redox homeostasis and virulence in Staphylococcus aureus. Molecular Microbiology, 111, 1039–1056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monk IR, Tree JJ, Howden BP, Stinear TP & Foster TJ (2015) Complete bypass of restriction systems for major Staphylococcus aureus lineages. mBio, 6, e00308–00315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nauseef WM (2007) Isolation of human neutrophils from venous blood. In: Quinn MT, DeLeo FR & Bokoch GM (Eds.) Neutrophil methods and protocols. Totowa, NJ: Humana Press, pp. 15–20. 10.1007/978-1-59745-467-4_2 Accessed 14 February 2022. [DOI] [Google Scholar]
- Noskin GA, Rubin RJ, Schentag JJ, Kluytmans J, Hedblom EC, Jacobson C et al. (2007) National trends in Staphylococcus aureus infection rates: impact on economic burden and mortality over a 6-year period (1998–2003). Clinical Infectious Diseases, 45, 1132–1140. [DOI] [PubMed] [Google Scholar]
- Novick RP (Ed.). (1990) Molecular biology of the staphylococci. New York, NY: VCH Publishers. [Google Scholar]
- Nygaard TK, Pallister KB, Ruzevich P, Griffith S, Vuong C & Voyich JM (2010) SaeR binds a consensus sequence within virulence gene promoters to advance USA300 pathogenesis. The Journal of Infectious Diseases, 201, 241–254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oriol C, Cengher L, Manna AC, Mauro T, Pinel-Marie M-L, Felden B et al. (2021) Expanding the Staphylococcus aureus SarA regulon to small RNAs. mSystems, 6, 713–721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reuter JS & Mathews DH (2010) RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics, 11, 129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ & Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26, 139–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roncarati D, Scarlato V & Vannini A (2022) Targeting of regulators as a promising approach in the search for novel antimicrobial agents. Microorganisms, 10, 185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scotto–Lavino E, Du G & Frohman MA (2006) 5′ end cDNA amplification using classic RACE. Nature Protocols, 1, 2555–2562. [DOI] [PubMed] [Google Scholar]
- Shimoni Y, Friedlander G, Hetzroni G, Niv G, Altuvia S, Biham O et al. (2007) Regulation of gene expression by small non-coding RNAs: a quantitative view. Molecular Systems Biology, 3, 138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith C, Heyne S, Richter AS, Will S & Backofen R (2010) Freiburg RNA tools: a web server integrating INTARNA, EXPARNA and LOCARNA. Nucleic Acids Research, 38, W373–W377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sorensen HM, Keogh RA, Wittekind MA, Caillet AR, Wiemels RE, Laner EA et al. (2020) Reading between the lines: utilizing RNA-seq data for global analysis of sRNAs in Staphylococcus aureus. mSphere, 5, e00439–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Storz G, Vogel J & Wassarman KM (2011) Regulation by small RNAs in bacteria: expanding frontiers. Molecular Cell, 43, 880–891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong SYC, Davis JS, Eichenberger E, Holland TL & Fowler VG (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clinical Microbiology Reviews, 28, 603–661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villa JK, Han R, Tsai C-H, Chen A, Sweet P, Franco G et al. (2021) A small RNA regulates pprM, a modulator of pleiotropic proteins promoting DNA repair, in Deinococcus radiodurans under ionizing radiation. Scientific Reports, 11, 12949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogel J & Wagner EGH (2007) Target identification of small noncoding RNAs in bacteria. Current Opinion in Microbiology, 10, 262–270. [DOI] [PubMed] [Google Scholar]
- Wagner EGH, Altuvia S & Romby P (2002) Antisense RNAs in bacteria and their genetic elements. Advances in Genetics, 46, 361–398. [DOI] [PubMed] [Google Scholar]
- Waters LS & Storz G (2009) Regulatory RNAs in bacteria. Cell, 136, 615–628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiong YQ, Willard J, Yeaman MR, Cheung AL & Bayer AS (2006) Regulation of Staphylococcus aureus α-toxin gene (hla) expression by agr, sarA, and sae in vitro and in experimental infective endocarditis. The Journal of Infectious Diseases, 194, 1267–1275. 10.1086/508210 [DOI] [PubMed] [Google Scholar]
- Zuker M, Mathews DH & Turner DH (1999) Algorithms and thermodynamics for RNA secondary structure prediction: a practical guide. In: Barciszewski J & Clark BFC (Eds.) RNA biochemistry and biotechnology. Dordrecht: Springer Netherlands, pp. 11–43. 10.1007/978-94-011-4485-8_2 Accessed 14 February 2022. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data available on request due to privacy/ethical restrictions.


