Abstract
Melanoma is the most lethal skin tumor, in large part because of a propensity for early metastasis. Good models of this most clinically relevant feature of melanoma are lacking. Here we report the development of an in vivo model of metastasis that relies on orthotopic injection of GFP-tagged lines in immunodeficient mice, serial intravital imaging of tumor progression and quantification of distant spread by 2-photon laser scanning microscopy, immunohistochemistry and real-time PCR analysis. Using this system, we report an assessment of the in vivo growth and metastatic properties of 11 well-characterized human melanoma cell lines. A subset of lines demonstrated rapid in vivo growth with invasion of host vasculature and distant seeding of viscera in this system. The ability to form metastasis in vivo did not correlate with 3D collagen invasion in vitro. Surprisingly, similar lines in terms of molecular genetic events differed markedly in their propensity to metastasize to distant organs such as brain and lung. In particular, two lines harboring B-RAF mutation and high levels of phosphorylated ERK and AKT (pERK and pAKT) were reproducibly unable to form tumors after orthotopic injection. Likewise, two previously identified RAS/RAF wild-type “epithelial-like” lines that do not have elevated pERK, pAKT or express TWIST1 mRNA still demonstrated a pronounced ability for orthotopic growth and metastatic spread. All the metastatic cell lines in this model showed increased NEDD9 expression, but NEDD9 lentiviral overexpression did not convey a metastatic phenotype on non-metastatic cells. These data suggest that melanoma metastasis is a molecularly heterogeneous process that may not require epidermal-to-mesenchymal transition or ERK activation, although both may facilitate the process.
Keywords: skin cancer, metastases, HEF1, CAS-L, confocal microscopy
INTRODUCTION
A clinical hallmark of melanoma is a proclivity to early distant metastasis, even from primary tumors of small size. For example, in 5-15% of patients presenting with metastatic melanoma, no primary tumor can be found [1, 2, 3, 4]. The process of metastasis, the most lethal feature of melanoma, is not well understood. We believe this deficiency in our understanding largely stems from a lack of good model systems in which to study this complex multi-step process. To date, the majority of studies on this topic have relied on tissue culture systems or tail vein injections of immunodeficient mice (reviewed in [5, 6, 7, 8]). Although these systems have some experimental strengths, they do not faithfully recapitulate the events that are involved in metastatic spread of autochthonous tumors. Thus, there is a need for in vivo models that better mimic the human disease, in regards to tumor microenvironment and metastasis formation.
Human and murine skin differs in significant aspects, particularly in terms of melanocyte distribution. Melanocytes of human skin are predominantly located at the junction of epidermis and dermis with some melanocytes present in hair follicles, whereas in mice the melanocytes are predominantly within hair follicles or the interfollicular dermis, and are rarely present at the dermal/epidermal junction [9, 10]. Important exceptions are the murine ear and tail, which share a comparable melanocyte distribution as in humans [10]. This is of importance considering the influence that the microenvironment plays in the maintenance of cellular homeostasis as well as in the development of cancer. This fact, combined with studies showing enhanced metastases of human melanoma cell lines after subdermal implantation in nude mice ([11, 12] and reviewed in [5]), suggest that injection of human melanoma cells intradermally in the ear of immunodeficient SCID mice would constitute a more orthotopic model of human melanoma growth and metastasis than i.v. or s.c. injection. Importantly, this system also allows for non-invasive serial intravital imaging of tumors by multi-photon microscopy.
An identifiable oncogenic “driver” event can be found in most human melanoma: BRAF (~60%), N-RAS (~20%), or c-KIT (<5%) mutation [13, 14, 15, 16, 17, 18]. We previously characterized a panel of melanoma cell lines for N-RAS and B-RAF mutation status, ERK and AKT activation status, INK4a/ARF status, G1 checkpoint response, proliferation rates, and RNA expression profiles ([19], and table 1). We found that increased ERK and AKT activation correlated strongly with mutational status of RAS-RAF pathway members and showed that melanoma cell lines lacking N-RAS/B-RAF mutation had low ERK and AKT activity and decreased expression of ERK-regulated gene transcripts. Our analyses suggested the existence of an “epithelial-like” form of melanoma that is molecularly distinct from the more common form of RAS/RAF mutant melanoma and does not require ERK activation or epithelial-to-mesenchymal transformation for progression. In this work, we extend the analysis of these well-characterized cell lines to include a study of metastatic propensity in an orthotopic xenograft model.
Table 1.
Molecular characterization of the cell lines used in this work
| B-RAF | N-RAS/ C-KIT | p16-CDK4-RB | ARF-p53 | pERK status | pAKT status | Orthopic growth as xenograft | |
|---|---|---|---|---|---|---|---|
| A375 | V600E mut. | WT | Undetectable p16 mRNA | HD of ARF, p53 WT & funct. | high | low | yes |
| A2058 | V600E mut. | WT | Rb null | Undetectable ARF mRNA, p53 funct. | med | high | yes |
| SK-MEL24 | V600E mut. | WT | HD of 9p21 | HD of 9p21, p53 funct. | high | high | no |
| SK-MEL28 | V600E mut. | WT | CDK4 Arg24Cys mut. | p53 Mut (Leu145Arg) | high | high | yes |
| UACC257 | V600E mut. | WT | p16 mut. | Unknown ARF - p53 lesion | high | high | no |
| WM2664 | V600E mut. | WT | HD of 9p21 | HD of 9p21, p53 funct. | high | high | yes |
| Mel505 | WT | WT | Undetectable p16 mRNA | p53 inact. | low | low | yes |
| PMWK | WT | WT | Unknown p16 - RB lesion | p53 inact. | low | low | no |
| RPMI8322 | WT | WT | Unknown p16 - RB lesion | p53 inact. | med | low | yes |
| SK-MEL23 | WT | WT | n.d. | n.d. | n.d. | n.d. | no |
| SK-MEL187 | WT | WT | HD of 9p21 | p53 mut (Arg273His) | low | low | yes |
HD: homozygous deletion.
Metastasis is a multistep process that involves the concerted action of genes that regulate invasion of the surrounding stroma, migration towards and intravasation into the vasculature, survival in the circulation, extravasation, survival and proliferation in the target organ [20]. A characteristic of metastasis in some systems is an epithelial to mesenchymal transition (EMT) [21, 22]. Twist and NEDD9 are two genes that have been recently linked to EMT and metastasis. Twist has been shown to enhance intravasation and metastasis in different types of cancer [22, 23], while NEDD9 was identified as a melanoma metastasis gene involved in the induction of the mesenchymal-type movement in melanoma cells [24, 25]. Moreover, we recently described the regulation of NEDD9 and metastasis by LKB1/STK11 in a mouse model of lung cancer and the deletion of LKB1 in human samples [26].
Given the complexity of metastasis, it is important to generate experimental models in which most or all steps of the metastatic cascade are represented. In this work, we have characterized the ability of a well-characterized panel of cell lines to grow in vivo and metastasize, using a novel orthotopic xenograft system. We injected GFP-expressing cells intradermally in the ears of NOD- SCID mice and used 2-photon laser scanning microscopy for the non-invasive imaging of the skin to follow tumor cell fate in vivo in real-time [27, 28]. In this way, we have shown that melanoma cell lines markedly differ in their ability to form tumors and metastasize. Importantly, in this model, metastasis was not generally correlated with features previously reported [22, 23, 29, 30, 31, 32] to enhance metastatic spread including EMT, RAS/RAF mutation, TWIST1 expression, ERK activation, in vitro proliferation rate, 3D collagen invasion or adhesion. These data indicate that melanoma metastasis is a molecularly heterogeneous and complex process.
METHODS
Cell culture, mutational analysis and proliferation assays
Human melanoma cell lines WM2664, A375, A2058, Mel505, RPMI8322, SK-MEL187, SK-MEL23, UACC257, SK-MEL24 and SK-MEL28, and 293FT cells were obtained from the sources detailed previously and cultured as described [33]. Genomic DNA was isolated using Genomic Tips kits (Qiagen, Valencia, CA), and the mutational status of C-KIT (exons 11, 13 and 17) was determined by direct sequencing of PCR amplification products as described [16, 34]. B-RAF (exons 11 and 15) and N-RAS (codons 12, 13, and 61) mutational status were as previously described [19]. We assessed in vitro cell proliferation by crystal violet staining as previously described [35] and Cell-Titer Glo luminescence viability assay (Promega, Madison, WI) as specified by the manufacturer at days 1, 4 and 7. Cell growth was normalized to day 1 values and analyzed using GraphPad Prism software (GraphPad Software Inc., La Jolla, CA). For the 3D collagen invasion assays, 2000 cells were mixed with type I rat tail collagen (BD Biosciences, Bedford, MA) and plated on 24 well plates. 8-day-old colonies were photographed using an inverted microscope (IX-81, Olympus) and a CCD camera (OCAR-ERG, Hamamatsu) controlled by Slidebook software.
Western blot analysis
Western blot assays were done on melanoma cell lysates in RIPA buffer with protease inhibitors (Roche) and phosphatase inhibitors (Calbiochem) as described [36]. Primary antibodies used were: Twist-1 (H-81, Santa Cruz Biotechnology, Santa Cruz, CA), LKB-1 (gift from N. Bardeesi), NEDD9 (clone 2G9, gift from E. Pugacheva and E. Golemis) and actin as loading control (C-11, Santa Cruz Biotechnology, Santa Cruz, CA). We used IRDye®-conjugated secondary antibodies and imaged the membranes using an Odyssey infrared imaging system (LI-COR, Lincoln, NE).
DNA constructs, viral production, and infection of target cells
The pLentiLox 3.7 (pLL3.7) vector [37] that expresses GFP under a CMV promoter was used. The production of lentiviruses in 293FT cells and infection of target cells were as described previously [37]. For NEDD9 modulation, various shRNAs targeting human NEDD9 were generated using the pLL3.7 vector and tested for their ability to inhibit NEDD9 protein expression by western blot. The most effective NEDD9 shRNA was used to inhibit NEDD9 expression in A375, A2058 and WM2664 cells, and a nonspecific shRNA was used as control. A GFP-NEDD9 lentiviral construct (gift from E. Golemis; [38]) was used to overexpress human NEDD9 in SK-MEL24, SK-MEL28 and SK-MEL187 cells, using the pLL3.7 vector as control. Human melanoma cells were infected with the viral supernatant, and GFP+ cells were isolated by flow cytometry as described [33].
Orthotopic Tumor Injections and Intravital Microscopy
6-8 wk old female NOD/SCID mice were anesthetized with 2% isofluorane, and 5 × 105 GFP-tagged tumor cells in 10 μl of DMEM were injected intradermally (i.d.) on the dorsal side of each ear using a 10 μl syringe with a 26-gauge needle. At least six to sixteen injections per cell line were performed. For serial confocal imaging, mice were anesthetized with ketamine/xylazine and ear fur was removed using a commercially available chemical depilatory agent. Injected ears were imaged at 1, 2, 7 and 14 days post-tumor injection, and additionally at the time of euthanasia: tumors of 1.3 cm diameter or 90 days after injection. Fluorescence in the ears was excited with 900-nm light from a Chameleon Ultra Ti-Sapphire pulsed laser (Coherent Inc., Santa Clara, CA) and imaged with a Zeiss LSM 510 NLO inverted 2-photon laser scanning microscope (Thornwood, NY) using 10x 0.3 NA, 20x 0.5 NA & 40x 1.2 NA (water-immersion) objectives. Images were captured using a 12-bit cooled CCD camera (Hamamatsu, Bridgewater, NJ). Dextran red (MW 75,000, Molecular Probes, Eugene, OR) was injected via tail vein to image blood vessels and to help visualize intravasated tumor cells. All mice were housed and treated in accordance with protocols approved by the institutional care and use committees for animal research at the University of North Carolina.
Imaging of metastatic cells in organs
Mice were euthanized and organs were placed on a glass bottom chamber and imaged using the Zeiss LSM 510 NLO inverted 2-photon laser scanning microscope with the 10x, 20x & 40x objectives. Images were captured using a Hamamatsu 12-bit cooled CCD camera. Brain, lungs, spleen, liver and kidneys from all mice were analyzed. GFP-expressing cells were easily detected over the background autofluorescence present in all organs, which was used for anatomical detailing. The criteria for considering a cell GFP+ included green emission using an excitation wavelength of 900 nm, without emission in the red spectra (excitation wavelength: 543 nm), which would indicate autofluorescence as is reported in several tissues including lung macrophages [39, 40, 41].
Tissue histology and immunohistochemistry
At the time of experiment termination, ears, tumors, lungs, brains, spleens, kidneys and livers were fixed in 10% formalin, paraffin embedded, and cut into five-micrometer sections. Assistance in sample processing was provided by the University of North Carolina Center for Gastrointestinal Biology and Disease. Slides were deparaffinized and stained using a mouse anti-GFP antibody (JL-8, Clontech, Mountain View, CA) at a dilution of 1:100 with immunodetection by HRP after citric acid antigen retrieval. Epitope masking was performed using a mouse-on-mouse kit (Vector Laboratories, Burlingame, CA) per manufacturer’s instructions. Control samples were stained without primary antibody. Samples were visualized and photographed under bright field microscopy (Zeiss Axioskop 2, Thornwood, NY) using 10X 0.25 NA, 20X 0.50 NA and 40X 1.30 NA objective, and an Axiocam camera (Zeiss, Thornwood, NY).
Real-time PCR analysis
Human beta-actin expression in indicated samples was determined by TaqMan real-time PCR (RT-PCR) as described [42]. 18s RNA expression was used as control. Normalized Ct values were evaluated across samples, and considered negative if >37. To determine the sensitivity of this assay to detect human cells in murine tissues, we extracted RNA from a mix of mouse (3T3) and human (WM2664) cells at various ratios. Using this approach we were able to detect 64 human cells among 106 murine cells (~1 in 16,000 cells). NEDD9 expression in indicated samples was determined by RT-PCR using the primers and probe described by Zheng and McKeown-Longo [43] and normalized to human beta-actin values. Ct values were used to calculate fold change expression of NEDD9.
RESULTS
Characterization of cell lines and in vitro proliferation
For these studies, we chose a representative panel of human melanoma cell lines harboring either B-RAF mutation (UACC257, SK-MEL24, SK-MEL28, A2058, A375, WM2664) or “wild-type (WT)” lines lacking RAS/RAF mutation (PMWK, RPMI8322, Mel505, SK-MEL187, SK-MEL23) from a panel of previously described melanoma cell lines [19]. We did not analyze N-RAS mutant lines as they are less frequent than either B-RAF mutant or RAS/RAF WT lines, and we had previously noted that N-RAS and B-RAF mutant lines are similar in terms of in vitro growth, AKT/ERK activation and gene expression. Based on the growing evidence that some melanomas harbor oncogenic C-KIT mutations [16, 17], we further analyzed our entire panel of cell lines for activating C-KIT mutation. No activating C-KIT mutations were detected in any of the 11 melanoma cell lines described in this or our prior report [19]. Importantly, our prior analyses of these lines included extensive characterization of the tumor genomes, mRNA expression profiles and protein expression. In particular, we showed that most of the WT cell lines express epithelial markers such as keratins and P/E- cadherins, with decreased TWIST expression.
For intravital experiments we stably expressed GFP in all cell lines using the lentiviral vector pLL3.7, and selected for the GFP positive cells by FACS. We characterized the in vitro proliferation rates of the lines for study over a one-week period of culture. We noted heterogeneity among the cultures with regard to in vitro growth, but generally B-RAF mutant lines exhibited more rapid growth than RAS/RAF WT lines (Fig. 1a and Table 1). An exception was UACC257, a B-RAF mutant line that grew slowly in vitro. These differences in in vitro growth rates may reflect intrinsic biologic features of the tumors from which these cells were derived, or reflect enhanced adaptation to experimental culture conditions ex vivo.
Fig. 1. In vitro and in vivo growth.

A) In vitro cell proliferation rate of the different human melanoma cell lines used in this study. B) Kaplan-Mayer curve showing tumor-free survival of SCID mice injected with 5×105 GFP-tagged cell lines. UACC257, SK-MEL23, SK-MEL24 and PMWK do not form tumors even after 90 days. 3 mice were bilaterally injected per tumor cell line (6 injection sites). C) Kaplan-Mayer curve showing no difference in tumor-free survival of SCID mice injected with B-RAF mutant versus wild type cell lines.
Characterization of in vivo growth
To more physiologically analyze the ability of these cell lines to grow and form tumors, we designed an in vivo model that relies on: 1) orthotopic intradermal (i.d.) injection of GFP-tagged cell lines in the ears of NOD-SCID mice; 2) serial intravital imaging of tumor progression; and 3) quantification of distant spread by 2-photon laser scanning microscopy, immunohistochemistry (IHC) and RT-PCR analysis. We performed a time-course analysis of tumor cell growth in vivo of the 11 melanoma cell lines by imaging the ears of the melanoma cell-injected mice at 1, 2, 7, 14 days and experiment termination. We could easily detect GFP+ cells in the ear at 24 hours post-injection, and were able to observe increased cell numbers 48 hr and 1 week post-injection, at times when no tumors were detectable by palpation (Fig. 2a). Using dextran red to stain the vasculature, intravasated tumor cells could be detected in some cases by 2 weeks post-injection (Fig. 2b and 2c). These appeared as round individual GFP+ cells inside the blood vessels. In all the animals analyzed we never detected lymph node metastasis. These observations suggest that this technique is suitable for the follow-up of tumor growth even at a single cell level, and is sensitive enough to detect hematogenous spread.
Fig. 2. Serial imaging of ear tumors by intravital microscopy.
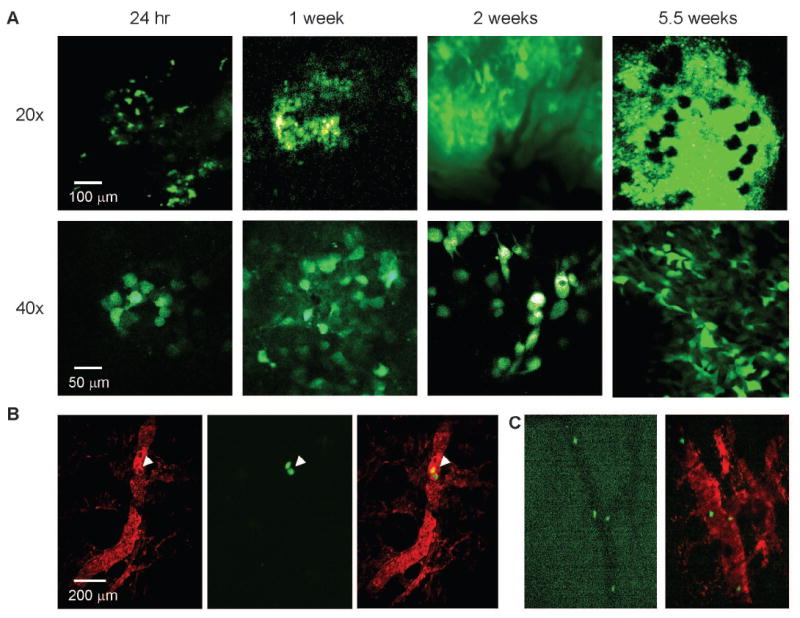
A) Representative 2-photon laser scanning microscopy images of GFP-tagged cells 1, 7, 14 and 45 days after injection of 5×105 GFP-tagged WM2664 cells i.d. in the ear of a NOD-SCID mouse, imaged at 20x (top row; scale bar: 100 μm) and 40x magnification (bottom row; scale bar: 50 μm). Single cells can be identified at all time points. B) Dextran red-labeled blood vessels showing intravasated cells 2 weeks after injection in the ear. Left: dextran red only. Note unstained area where melanoma cells are located (arrow head). Middle: GFP only. Right: merged dextran red and GFP. Scale bar: 200 μm. C) Intravasated cells can be observed by contrast in the green channel (left) or merging the red and green channels (right).
As was the case with in vitro proliferation, we noted marked heterogeneity among melanoma cell lines with regard to their ability to grow in this in vivo system. While some cell lines were noted to grow well in vivo (SK-MEL28, SK-MEL187, A375, WM2664, Mel505 and RPMI8322), others were unable to form tumors in the ears of NOD-SCID mice (UACC257, SK-MEL24, SK-MEL23, PMWK; Table 1, Fig. 1b). Using this in vivo model of tumor growth, we observed no correlation between in vitro and in vivo growth (data not shown), suggesting the importance of the in vivo model in the characterization of tumor cell lines. Based on our prior and present analysis of these cell lines [19], the ability to form orthotopic tumors did not correlate with known molecular features of the cell lines (Fig. 1b and 1c, Table 1). For example, a few WT lines harboring low levels of pERK and pAKT activation (SK-MEL187, Mel505, RPMI8322) grew in vivo as well as, or faster than some B-RAF mutant cell lines that have increased pERK and pAKT levels (SK-MEL28, A2058, A375, WM2664; Fig. 1b). Likewise, growth in this assay did not correlate with INK4a/ARF mutational status, expression of pAKT, TWIST1 expression, or an “EMT phenotype” (indicated by expression of keratins, TWIST and P/E-cadherins), despite literature suggesting roles for these genetic events in tumor progression and invasion [22, 23, 44, 45, 46, 47]. While our sample is too small to exclude the possibility that these genetic events enhance tumor growth, these data demonstrate that none of these molecular features is strictly required for local orthotopic growth of melanoma cell lines in immunodeficient mice.
Detection of metastasis
In order to determine the relationship between primary orthotopic growth and distant spread, tumor-bearing mice were analyzed for metastasis. Mice were sacrificed at 90 days post-injection or when primary ear tumors reached 1.3 cm in size, and organs were harvested for analysis. Fresh organs were imaged by 2-photon laser scanning microscopy, and formalin-fixed paraffin-embedded organs were analyzed by IHC for GFP. Additionally, RT-PCR for human beta-actin mRNA was used to confirm the presence of human melanoma cells in the studied organs. While no metastases were observed macroscopically in any distant organ, using 2-photon laser scanning imaging we detected small GFP+ foci in the lungs and brains of mice harboring tumor derived from WM2664, A2058, A375 and RPMI8322 and in the brains of Mel505 tumor-bearing mice (Table 2, Fig. 3). RT-PCR amplification of human beta-actin mRNA (Table 2) confirmed the presence of metastatic spread in lungs and brains. Moreover, by RT-PCR, we detected the presence of human cells in the lungs and brains of tumor-bearing mice that did not show signs of GFP+ cells by 2-photon laser scanning microscopy (Table 2), suggesting RT-PCR amplification is more sensitive to detect rare, distantly spread cells (e.g. identifying a single human cell ~16,000 mouse cells, see methods). Importantly, however, the physiological significance of rare tumor cells detected by 2-photon microscopy or RT-PCR is unknown, and could merely represent intravascular circulating tumor cells rather than nascent metastatic foci.
Table 2.
Summary of the metastatic properties of the different cell lines by 2-photon laser scanning microscopy and RT-PCR analysis.
| Cell line | Tumor growth* | Green cells in lungs by 2-P** | RT-PCR hu-β-actin in lungsˆ | Green cells in brain by 2-P** | RT-PCR hu-β-actin in brainˆ | Green cells in liver, spleen, kidney by 2-P** | RT-PCR hu-β-actin in liver, spleen, kidneyˆ |
|---|---|---|---|---|---|---|---|
| A375 | 22/22 | 6/7 (m) | 1/4 | 4/7 | 1/3 | 0/7 | 0/3 |
| A2058 | 15/18 | 7/10 (M) | 2/3 | 6/10 | 3/4 | 0/10 | 0/3 |
| SK-MEL24 | 0/18 | 0/6 | 0/4 | 0/6 | 0/4 | 0/6 | 0/3 |
| SK-MEL28 | 16/16 | 0/5 | 1/5 | 0/8 | 0/5 | 0/5 | 0/3 |
| UACC257 | 0/12 | 0/6 | 0/3 | 0/6 | 0/3 | 0/6 | 0/3 |
| WM2664 | 17/20 | 9/9 (M) | 9/10 | 4/8 | 1/4 | 0/10 | 0/3 |
| Mel505 | 12/12 | 0-2/6 (m) | 2/6 | 1/3 | 2/3 | 0/6 | 0/3 |
| PMWK | 0/12 | 0/6 | 0/6 | 0/6 | 0/6 | 0/6 | 0/3 |
| RPMI8322 | 3/4 | 2/2 (m) | 2/2 | 2/3 | 2/2 | 0/3 | 0/3 |
| SK-MEL23 | 0/6 | 0/3 | 0/3 | 0/3 | 0/3 | 0/3 | 0/3 |
| SK-MEL187 | 10/12 | 0/6 | 0/3 | 0/3 | 0/3 | 0/6 | 0/3 |
=# tumors formed/ # of sites injected
= # of organs with metastases/ total organs analyzed
= # of + organs by RT-PCR /total organs analyzed
m= micrometastases (1-3 cells)
M= gross metastasis
2-P= 2-photon laser scanning microscopy
Fig. 3. Detection of metastatic cells in whole organs.
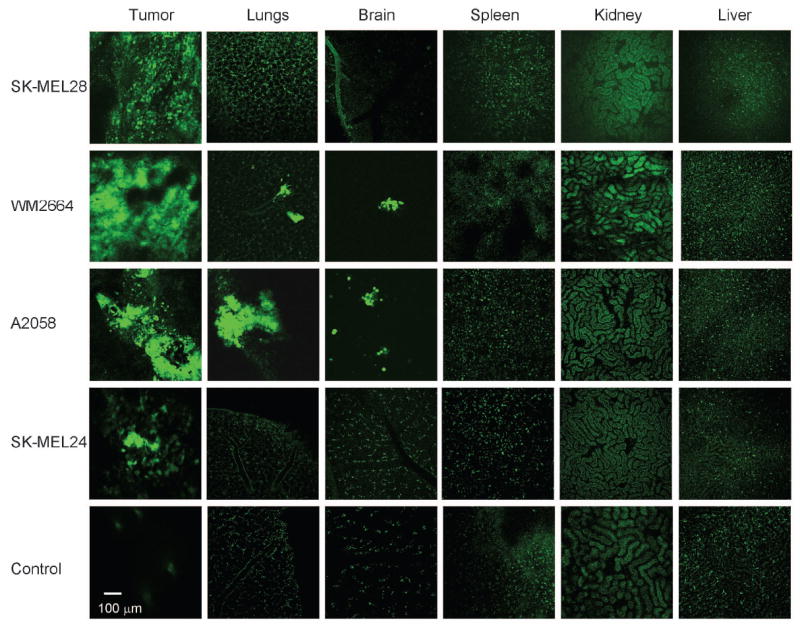
Representative 2-photon images of tumors and organs of mice injected with the different human melanoma cell lines at end-point (when tumors reached 1.3 mm diameter or 90 days post-injection). SK-MEL28 cells form rapidly growing tumors, but show no signs of metastasis in any organs. WM2664 and A2058 cells form slower growing tumors that metastasize to lungs and brain. SK-MEL24 cells were not able to form overt tumors and did not give rise to metastasis in any organs, but cells were still present in the ear 90 days after injection. Note anatomical detailing of organs by autofluorescence. Scale bar: 100 μm.
We further confirmed the presence of tumor foci in lungs of A2058 and WM2664 tumor-bearing mice by IHC staining for GFP (Fig. 4), but we were unable to detect GFP+ cells in the lungs of A375 or RPMI8322 tumor-bearing mice by this method (Fig. 4 and data not shown). We believe this demonstrates an increased sensitivity of 2-photon imaging of whole organs over commonly employed techniques of serial paraffin sectioning with IHC analysis. Three animals demonstrated lymph node enlargement, but metastatic tumor was not observed in these nodes suggesting enlargement reflected a reactive process. A comprehensive investigation of lymph node metastasis was not performed in the absence of nodal enlargement. Finally, no melanoma cells were observed in the liver, spleen or kidney of any tumor-bearing mice by 2-photon, IHC or RT-PCR (Fig. 3, Table 2 and data not shown). In aggregate, these data suggest a predilection of certain of these melanoma cell lines for metastatic spread to brain and lung, but not other organs tested.
Fig. 4. Immunohistochemical detection of GFP metastasis in lungs.

Representative images of paraffin sections of tumors, lungs and brains of mice injected with the different human melanoma cell lines at end-point (when tumors reached 1.3 mm diameter or after 90 days). SK-MEL28 tumors stained positively for GFP (brown), but no GFP staining was observed in any organs from these mice by IHC. WM2664 and A2058 tumors and lungs were positive for GFP, confirming the 2-photon data. All other organs were GFP negative. Scale bar: 200 μm.
Gene expression analysis
We sought to determine if tumor molecular genetics or gene expression correlated with metastatic spread in this system. As with primary tumor growth, no correlation was noted between distant spread and RAS/RAF mutation, INK4a/ARF or p53 status, expression of pAKT or pERK, TWIST1 expression, or expression of markers associated with EMT. Particularly, the WT cell lines MEL505 and RPMI8322, which have metastatic properties in the absence of RAS-RAF mutations, express keratins, P/E-cadherin and CD24 [19]. We have previously analyzed genome-wide mRNA expression of these lines by high-density microarray analysis [19], and we also performed several types of supervised analyses using SAM [48] to compare mRNA expression with tumor growth or metastasis. Although we performed independent SAM analysis by cell line growth, cell line metastasis to any organ, cell line metastasis to lung, and cell line metastasis to brain; no robust gene signature with low false-discovery rate could be identified that correlated with local growth or metastatic spread in any of these analyses. Although it is worth noting that 11 samples provides only limited statistical power for supervised genome-wide expression analysis, these data suggest heterogeneity of gene expression among lines capable of distant spread in this system.
Melanoma metastasis protein expression
We analyzed the expression of metastasis genes in this system. Given previous findings [22, 23, 24, 25, 26], we studied TWIST, LKB-1 and NEDD9 protein expression in our panel of cell lines. While no correlation was observed between TWIST or LKB-1 expression in vitro and tumorigenesis or metastasis in vivo, all the cell lines that were able to metastasize in vivo expressed NEDD9 protein (Fig. 5a), consistent with previous reports of NEDD9 being involved in melanoma metastasis [24]. Based on this data, we further evaluated the role of NEDD9 in metastasis formation in our model.
Fig. 5. Twist1, LKB1 and NEDD9 expression.

A) Protein expression was assessed in our panel of melanoma cell lines by western blot. Note expression of NEDD9 protein by cells that are metastatic in vivo (in blue). B) Western blot confirming GFP-NEDD9 overexpression (SK-MEL24, SK-MEL28 and SK-MEL187) or shRNA downregulation of NEDD9 protein (A375, A2058 and WM2664) compared to controls.
For this purpose, we overexpressed GFP-NEDD9 using a lentiviral construct [38] in cells that do not express NEDD9 and do not metastasize in our model (SK-MEL24, SK-MEL28 and SK-MEL187). Conversely, we inhibited NEDD9 expression in cells that are metastaic in our model (A375, A2058 and WM2664) using a lentiviral shRNA system (pLL3.7). We confirmed overexpression or downregulation of NEDD9 protein by western blot and RT-PCR (Fig. 5b and supplementary Fig. 2). We then analyzed the effects of NEDD9 modulation on cell proliferation, cell adhesion and cell invasion into 3D collagen in vitro (Supplementary Fig. 3 and data not shown). Inconsistent effects of NEDD9 overexpression and knockdown were noted in vitro (Supplementary Fig. 3), consistent with prior reports of NEDD9 enhancing and reducing in vitro proliferation and regulating cell cycle progression (reviewed in [49], [50]). In accord with results from Kim et al. [24], NEDD9 downregulation in A2058 cells abrogated tumor growth in vivo, but did not affect tumor growth or metastasis in two other RAF-mutant cell lines (A375 and WM2664, data not shown). Using both the orthotopic intradermal ear injection model and the 3D collagen invasion assay we found that overexpression of NEDD9 in SK-MEL28, SK-MEL24 and SK-MEL187 cells was not sufficient to affect tumor growth or to induce metastasis. Although the correlation of NEDD9 expression and results of NEDD9 knockdown in A2058 cells are consistent with an important role of this gene in melanoma metastasis as reported [24], our data suggest that transient knockdown or overexpression of this gene is not universally sufficient to modulate melanoma metastasis.
DISCUSSION
In the present study, we have expanded our previous analysis and characterization of a panel of human melanoma cell lines [19] using an in vivo model of melanoma metastasis in which we injected GFP-tagged human melanoma cells intradermally into the ears of NOD-SCID mice. With this model, we were able to detect orthotopically growing single GFP+ cells as early as 24 hours post-injection, and were also able to observe intravasated GFP+ cells in the vasculature at 2 weeks after injection. We found that several, but not all of the 11 human melanoma cell lines tested were able to grow rapidly in vivo, invade the vasculature and form distant metastasis in the brains and lungs of NOD-SCID mice. We could detect metastatic foci in lungs and brains by 2-photon laser scanning microscopy and confirmed these results through IHC and RT-PCR based methods. We noted that 2-photon imaging was more sensitive than IHC, but less sensitive than RT-PCR for detecting tumor cells in a distant organ. Hematogenous spread in this system was observed as micrometastases. It is possible that these tumor deposits were unable to form macrometastases because of some biological limit to growth, but more likely, we believe this observation reflects the fact that the experimentsmost mice had to be sacrificed because of growth of the primary ear tumor at a time before macrometastases could form. Possible means to overcome this issue might be to inject a smaller number of cells intradermally, or to resect the primary tumor after some initial period of growth.
Approximately 80% of metastatic melanomas harbor mutually exclusive activating mutations of either N-RAS or B-RAF (reviewed in [51]). These lesions lead to activation of the RAF/MEK/ERK/MAPK pathway, which in turn controls the transcription of hundreds to thousands of genes related to cellular proliferation, survival, and motility [19]. There is extensive literature proposing ERK activation [29, 52], RAS/RAF mutation [30, 31], TWIST1 expression [23], NEDD9 expression [24] and EMT [21, 22, 53] as facilitators of metastasis, mainly for their roles in modulating cell-cell contacts, cell motility, cell adhesion, and inducing an invasive phenotype [32]. In our previous work we found increased ERK activation and TWIST1 expression in most RAS/RAF mutant melanoma cell lines when compared to epithelial-like cell lines (which express keratins and P/E- cadherins) and normal human melanocytes [19]. As some WT lines demonstrated rapid metastasis in this system, our results suggest that RAS/RAF mutations, ERK activation and expression of transcripts associated with EMT are not strictly required for metastasis, although these features may possibly facilitate the process. Our results are also consistent with the finding that epithelial markers keratin 18 and CD24 expression do not correlate with improved univariate survival in a tissue microarray data set [19]. It is important to note, however, that EMT has been reported to be a reversible process, and therefore we can not exclude the possibility that this occurs in vivo to facilitate metastasis even in the WT lines, despite their in vitro patterns of gene expression.
In summary, we describe an orthotopic melanoma mouse model for the study of early events contributing to the development of melanoma from human cell lines. Using this model, we describe the propensity of several commonly studied RAF-mutant and WT human melanoma cell lines to grow locally and spread distantly when injected orthotopically, and observe a correlation of NEDD9 expression with this process. The observed differences in growth and metastasis did not correlate with any of the other characterized molecular genetic features of the cell lines including several candidate genes thought to be involved in melanoma metastasis. Moreover, the ability to form metastasis in vivo did not correlate with 3D collagen invasion in vitro. One possible explanation for the poor correlation between assays is the lack of other stromal components, including cells such as fibroblasts and keratinocytes in the in vitro assay. While the heterogeneity and small number of cell lines analyzed in our sample prevents us from drawing definitive conclusions about the relative role of these processes generally in melanoma spread, we consider it remarkable that certain WT melanoma cell lines without ERK activation or expression of transcripts associated with EMT demonstrated rapid hematogenous spread to the brain and lung after orthotopic injection. Therefore, our work suggests the conclusion that melanoma metastasis is a complex and heterogeneous process in vivo.
Supplementary Material
A) 2-photon images of lungs from an A2058 (top) and a SK-MEL28 tumor- bearing mouse (bottom) comparing real GFP fluorescence versus autofluorescence. GFP+ cells (top) do not emit in the red channel (middle panel), while autofluorescence (bottom) can be detected in both the green (left panel) and red channels. Right panel: GFP detection by IHC. Scale bar: 200 μm. B) Detail of lung and brain metastasis. 2-photon laser scanning microscopy images from representative tumors, lungs and brains from GFP-tagged human melanoma cells cropped from 20x magnification. Scale bar: 50 μm.
Fold change expression of NEDD9 RNA by RT-PCR confirming GFP-NEDD9 overexpression (SK-MEL24, SK-MEL28 and SK-MEL187) or shRNA downregulation of NEDD9 protein (A375, A2058 and WM2664) compared to controls.
A) In vitro cell proliferation rate of GFP-NEDD9 and NEDD9 shRNA (shN9) expressing cells compared to control cells. B) Invasion into 3D collagen by these cell lines. Photos of 8-days-old cultures were taken on a DIC microscope at 10x magnification. Scale bar: 200 μm.
Acknowledgments
We wish to thank Gerald Gordon and Robert T. Currin for their assistance with the intravital microscopy, the UNC Histology Core (NIH P30DK034987), Janiel Shields and Thomas Marshall for reagents and assistance, and Scott McPhee, Nabeel Bardeesy and Christin Petre for comments on the manuscript.
Financial support: NCI and NIEHS (CA106991, CA90679, CA105837, and ES14635), the Golfers Against Cancer Foundation (to NES and JEB), ACS (RSG-08-154-01-CSM to JEB) and the National Cancer Center (to KBM).
References
- 1.Jain D, Singh T, Kumar N, Daga MK. Metastatic malignant melanoma in bone marrow with occult primary site – a case report with review of literature. Diagn Pathol. 2007;2:38–41. doi: 10.1186/1746-1596-2-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Anbari KK, Schuchter LM, Bucky LP, Mick R, Synnestvedt M, Guerry D, et al. Melanoma of unknown primary site: presentation, treatment, and prognosis--a single institution study. University of Pennsylvania Pigmented Lesion Study Group. Cancer. 1997;79:1816–21. doi: 10.1002/(sici)1097-0142(19970501)79:9<1816::aid-cncr26>3.0.co;2-#. [DOI] [PubMed] [Google Scholar]
- 3.Baab GH, McBride CM. Malignant melanoma: the patient with an unknown site of primary origin. Arch Surg. 1975;110:896–900. doi: 10.1001/archsurg.1975.01360140040008. [DOI] [PubMed] [Google Scholar]
- 4.Chang P, Knapper WH. Metastatic melanoma of unknown primary. Cancer. 1982;49:1106–11. doi: 10.1002/1097-0142(19820315)49:6<1106::aid-cncr2820490607>3.0.co;2-0. [DOI] [PubMed] [Google Scholar]
- 5.Gattoni-Celli S, Byers SH, Calorini L, Ferrone S. Organ-Specific Metastases in Melanoma: Experimental Animal Models. Pigment Cell Res. 1993;6:981–384. doi: 10.1111/j.1600-0749.1993.tb00619.x. [DOI] [PubMed] [Google Scholar]
- 6.Fodstad O, Kjønniksen I, Aamdal S, Nesland JM, Boyd MR, Pihl A. Extrapulmonary, tissue-specific metastasis formation in nude mice injected with FEMX-I human melanoma cells. Cancer Res. 1988;48:4382–8. [PubMed] [Google Scholar]
- 7.Ishikawa M, Fernandez B, Kerbel RS. Highly pigmented human melanoma variant which metastasizes widely in nude mice, including to skin and brain. Cancer Res. 1988;48:4897–903. [PubMed] [Google Scholar]
- 8.Kerbel RS, Cornil I, Theodorescu D. Importance of orthotopic transplantation procedures in assessing the effects of transfected genes on human tumor growth and metastasis. Cancer Metastasis Rev. 1991;10:201–15. doi: 10.1007/BF00050792. [DOI] [PubMed] [Google Scholar]
- 9.Bleehen S. Normal pigmentation, radiation and response to sun exposure. In: Champion RH, Burns DA, Breathnach SM, editors. Textbook of Dermatology. London: Blackwell-Science; 1998. pp. 1765–7. [Google Scholar]
- 10.Silvers W. The coat colors of mice: A model for mammalian gene action and interaction. In: Altman PL, K DD, editors. Inbred and Genetically Defined Strains of Laboratory Animals, Part1 Mouse and Rat. New York: Springer-Verlag; 1979. pp. 4–5. [Google Scholar]
- 11.Chishima T, Miyagi Y, Wang X, Yamaoka H, Shimada H, Moossa AR, et al. Cancer invasion and micrometastasis visualized in live tissue by green fluorescent protein expression. Cancer Res. 1997;57:2042–7. [PubMed] [Google Scholar]
- 12.Cornil I, Man S, Fernandez B, Kerbel RS. Enhanced tumorigenicity, melanogenesis, and metastases of a human malignant melanoma after subdermal implantation in nude mice. J Natl Cancer Inst. 1989;81:938–44. doi: 10.1093/jnci/81.12.938. [DOI] [PubMed] [Google Scholar]
- 13.Dong J, Phelps RG, Qiao R, Yao S, Benard O, Ronai Z, et al. BRAF oncogenic mutations correlate with progression rather than initiation of human melanoma. Cancer Res. 2003;63:3883–5. [PubMed] [Google Scholar]
- 14.Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949–54. doi: 10.1038/nature00766. [DOI] [PubMed] [Google Scholar]
- 15.Herlyn M, Satyamoorthy K. Activated ras. Yet another player in melanoma? Am J Pathol. 1996;149:739–44. [PMC free article] [PubMed] [Google Scholar]
- 16.Willmore-Payne C, H J, Tripp S, Layfield LJ. Human malignant melanoma: detection of BRAF- and c-kit activating mutations by high-resolution amplicon melting analysis. Human Pathology. 2005;36:486–93. doi: 10.1016/j.humpath.2005.03.015. [DOI] [PubMed] [Google Scholar]
- 17.Curtin JA, Busam K, Pinkel D, Bastian BC. Somatic activation of KIT in distinct subtypes of melanoma. J Clin Oncol. 2006;24:4340–6. doi: 10.1200/JCO.2006.06.2984. [DOI] [PubMed] [Google Scholar]
- 18.Daniotti M, Oggionni M, Ranzani T, Vallacchi V, Campi V, Di Stasi D, et al. BRAF alterations are associated with complex mutational profiles in malignant melanoma. Oncogene. 2004;23:5968–77. doi: 10.1038/sj.onc.1207780. [DOI] [PubMed] [Google Scholar]
- 19.Shields JM, Thomas NE, Cregger M, Berger AJ, Leslie M, Torrice C, et al. Lack of Extracellular Signal-Regulated Kinase Mitogen-Activated Protein Kinase Signaling Shows a New Type of Melanoma. Cancer Res. 2007;67:1502–12. doi: 10.1158/0008-5472.CAN-06-3311. [DOI] [PubMed] [Google Scholar]
- 20.Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003;3:453–8. doi: 10.1038/nrc1098. [DOI] [PubMed] [Google Scholar]
- 21.Alonso SR, Tracey L, Ortiz P, Perez-Gomez B, Palacios J, Pollan M, et al. A high-throughput study in melanoma identifies epithelial-mesenchymal transition as a major determinant of metastasis. Cancer Res. 2007;67:3450–60. doi: 10.1158/0008-5472.CAN-06-3481. [DOI] [PubMed] [Google Scholar]
- 22.Kang Y, Massague J. Epithelial-mesenchymal transitions: twist in development and metastasis. Cell. 2004;118:277–9. doi: 10.1016/j.cell.2004.07.011. [DOI] [PubMed] [Google Scholar]
- 23.Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C, et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell. 2004;117:927–39. doi: 10.1016/j.cell.2004.06.006. [DOI] [PubMed] [Google Scholar]
- 24.Kim M, Gans JD, Nogueira C, Wang A, Paik JH, Feng B, et al. Comparative oncogenomics identifies NEDD9 as a melanoma metastasis gene. Cell. 2006;125:1269–81. doi: 10.1016/j.cell.2006.06.008. [DOI] [PubMed] [Google Scholar]
- 25.Sanz-Moreno V, Gadea G, Ahn J, Paterson H, Marra P, Pinner S, et al. Rac activation and inactivation control plasticity of tumor cell movement. Cell. 2008;135:510–23. doi: 10.1016/j.cell.2008.09.043. [DOI] [PubMed] [Google Scholar]
- 26.Ji H, Ramsey MR, Hayes DN, Fan C, McNamara K, Kozlowski P, et al. LKB1 modulates lung cancer differentiation and metastasis. Nature. 2007;448:807–10. doi: 10.1038/nature06030. [DOI] [PubMed] [Google Scholar]
- 27.Centonze VE, White JG. Multiphoton excitation provides optical sections from deeper within scattering specimens than confocal imaging. Biophys J. 1998;75:2015–24. doi: 10.1016/S0006-3495(98)77643-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang W, Wyckoff JB, Frohlich VC, Oleynikov Y, Hüttelmaier S, Zavadil J, et al. Single cell behavior in metastatic primary mammary tumors correlated with gene expression patterns revealed by molecular profiling. Cancer Res. 2002;62:6278–88. [PubMed] [Google Scholar]
- 29.Ward Y, Wang W, Woodhouse E, Linnoila I, Liotta L, Kelly K. Signal pathways which promote invasion and metastasis: critical and distinct contributions of extracellular signal-regulated kinase and Ral-specific guanine exchange factor pathways. Mol Cell Biol. 2001;21:5958–69. doi: 10.1128/MCB.21.17.5958-5969.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Houben R, Becker JC, Kappel A, Terheyden P, Brocker EB, Goetz R, et al. Constitutive activation of the Ras-Raf signaling pathway in metastatic melanoma is associated with poor prognosis. J Carcinog. 2004;3:6. doi: 10.1186/1477-3163-3-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Giehl K. Oncogenic Ras in tumour progression and metastasis. Biological chemistry. 2005;386:193–205. doi: 10.1515/BC.2005.025. [DOI] [PubMed] [Google Scholar]
- 32.Hoon DS, Kitago M, Kim J, Mori T, Piris A, Szyfelbein K, et al. Molecular mechanisms of metastasis. Cancer metastasis reviews. 2006;25:203–20. doi: 10.1007/s10555-006-8500-x. [DOI] [PubMed] [Google Scholar]
- 33.Petermann KB, Rozenberg GI, Zedek D, Groben P, McKinnon K, Buehler C, et al. CD200 is induced by ERK and is a potential therapeutic target in melanoma. The Journal of clinical investigation. 2007;117:3922–9. doi: 10.1172/JCI32163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Willmore C, Holden JA, Zhou L, Tripp S, Wittwer CT, Layfield LJ. Detection of c-kit-activating mutations in gastrointestinal stromal tumors by high-resolution amplicon melting analysis. Am J Clin Pathol. 2004;122:206–16. doi: 10.1309/4E6U-YBY6-2N2F-CA6N. [DOI] [PubMed] [Google Scholar]
- 35.Sharpless NE, Bardeesy N, Lee KH, Carrasco D, Castrillon DH, Aguirre AJ, et al. Loss of p16Ink4a with retention of p19Arf predisposes mice to tumorigenesis. Nature. 2001;413:86–91. doi: 10.1038/35092592. [DOI] [PubMed] [Google Scholar]
- 36.Ramsey MR, Krishnamurthy J, Pei XH, Torrice C, Lin W, Carrasco DR, et al. Expression of p16Ink4a compensates for p18Ink4c loss in cyclin-dependent kinase 4/6-dependent tumors and tissues. Cancer Res. 2007;67:4732–41. doi: 10.1158/0008-5472.CAN-06-3437. [DOI] [PubMed] [Google Scholar]
- 37.Rubinson DA, Dillon CP, Kwiatkowski AV, Sievers C, Yang L, Kopinja J, et al. A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nat Genet. 2003;33:401–6. doi: 10.1038/ng1117. [DOI] [PubMed] [Google Scholar]
- 38.Pugacheva EN, Golemis EA. The focal adhesion scaffolding protein HEF1 regulates activation of the Aurora-A and Nek2 kinases at the centrosome. Nature cell biology. 2005;7:937–46. doi: 10.1038/ncb1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Weyde J, Wassermann K, Schell-Frederick E. Analysis of single and double-stained alveolar macrophages by flow cytometry. J Immunol Methods. 1997;207:115–23. doi: 10.1016/s0022-1759(97)00103-8. [DOI] [PubMed] [Google Scholar]
- 40.Bunn HJ, Woltmann G, Grigg J. Applicability of laser scanning cytometry to study paediatric alveolar macrophages. Eur Respir J. 2002;20:1437–43. doi: 10.1183/09031936.02.00033502. [DOI] [PubMed] [Google Scholar]
- 41.Wyckoff JB, Jones JG, Condeelis JS, Segall JE. A Critical Step in Metastasis: In Vivo Analysis of Intravasation at the Primary Tumor. Cancer Res. 2000;60:2504–11. [PubMed] [Google Scholar]
- 42.Penland SK, Keku TO, Torrice C, He X, Krishnamurthy J, Hoadley KA, et al. RNA expression analysis of formalin-fixed paraffin-embedded tumors. Lab Invest. 2007 doi: 10.1038/labinvest.3700529. [DOI] [PubMed] [Google Scholar]
- 43.Zheng M, McKeown-Longo PJ. Regulation of HEF1 expression and phosphorylation by TGF-beta 1 and cell adhesion. The Journal of biological chemistry. 2002;277:39599–608. doi: 10.1074/jbc.M202263200. [DOI] [PubMed] [Google Scholar]
- 44.Govindarajan B, Sligh JE, Vincent BJ, Li M, Canter JA, Nickoloff BJ, et al. Overexpression of Akt converts radial growth melanoma to vertical growth melanoma. J Clin Invest. 2007;117:719–29. doi: 10.1172/JCI30102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sharpless E, Chin L. The INK4a/ARF locus and melanoma. Oncogene. 2003;22:3092–8. doi: 10.1038/sj.onc.1206461. [DOI] [PubMed] [Google Scholar]
- 46.Alonso SR, Ortiz P, Pollan M, Perez-Gomez B, Sanchez L, Acuna MJ, et al. Progression in cutaneous malignant melanoma is associated with distinct expression profiles: a tissue microarray-based study. Am J Pathol. 2004;164:193–203. doi: 10.1016/s0002-9440(10)63110-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Altomare DA, Testa JR. Perturbations of the AKT signaling pathway in human cancer. Oncogene. 2005;24:7455–64. doi: 10.1038/sj.onc.1209085. [DOI] [PubMed] [Google Scholar]
- 48.Tusher VG, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A. 2001;98:5116–21. doi: 10.1073/pnas.091062498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Singh M, Cowell L, Seo S, O’Neill G, Golemis E. Molecular basis for HEF1/NEDD9/Cas-L action as a multifunctional co-ordinator of invasion, apoptosis and cell cycle. Cell biochemistry and biophysics. 2007;48:54–72. doi: 10.1007/s12013-007-0036-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dadke D, Jarnik M, Pugacheva EN, Singh MK, Golemis EA. Deregulation of HEF1 impairs M-phase progression by disrupting the RhoA activation cycle. Molecular biology of the cell. 2006;17:1204–17. doi: 10.1091/mbc.E05-03-0237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Chin L, Garraway LA, Fisher DE. Malignant melanoma: genetics and therapeutics in the genomic era. Genes Dev. 2006;20:2149–82. doi: 10.1101/gad.1437206. [DOI] [PubMed] [Google Scholar]
- 52.Mirmohammadsadegh A, Mota R, Gustrau A, Hassan M, Nambiar S, Marini A, et al. ERK1/2 is highly phosphorylated in melanoma metastases and protects melanoma cells from cisplatin-mediated apoptosis. J Invest Dermatol. 2007;127:2207–15. doi: 10.1038/sj.jid.5700870. [DOI] [PubMed] [Google Scholar]
- 53.Grunert S, Jechlinger M, Beug H. Diverse cellular and molecular mechanisms contribute to epithelial plasticity and metastasis. Nat Rev Mol Cell Biol. 2003;4:657–65. doi: 10.1038/nrm1175. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
A) 2-photon images of lungs from an A2058 (top) and a SK-MEL28 tumor- bearing mouse (bottom) comparing real GFP fluorescence versus autofluorescence. GFP+ cells (top) do not emit in the red channel (middle panel), while autofluorescence (bottom) can be detected in both the green (left panel) and red channels. Right panel: GFP detection by IHC. Scale bar: 200 μm. B) Detail of lung and brain metastasis. 2-photon laser scanning microscopy images from representative tumors, lungs and brains from GFP-tagged human melanoma cells cropped from 20x magnification. Scale bar: 50 μm.
Fold change expression of NEDD9 RNA by RT-PCR confirming GFP-NEDD9 overexpression (SK-MEL24, SK-MEL28 and SK-MEL187) or shRNA downregulation of NEDD9 protein (A375, A2058 and WM2664) compared to controls.
A) In vitro cell proliferation rate of GFP-NEDD9 and NEDD9 shRNA (shN9) expressing cells compared to control cells. B) Invasion into 3D collagen by these cell lines. Photos of 8-days-old cultures were taken on a DIC microscope at 10x magnification. Scale bar: 200 μm.


