Abstract
Pathogen- or microbe-associated molecular patterns (PAMPs/MAMPs) are detected as nonself by host pattern recognition receptors (PRRs) and activate pattern-triggered immunity (PTI). Microbial invasions often trigger the production of host-derived endogenous signals referred to as danger-or damage-associated molecular patterns (DAMPs), which are also perceived by PRRs to modulate PTI responses. Collectively, PTI contributes to host defense against infections by a broad range of pathogens. Remarkable progress has been made toward demonstrating the cellular and physiological responses upon pattern recognition, elucidating the molecular, biochemical, and genetic mechanisms of PRR activation, and dissecting the complex signaling networks that orchestrate PTI responses. In this review, we present an update on the current understanding of how plants recognize and respond to nonself patterns, a process from which the seemingly chaotic responses form into a harmonic defense.
Keywords: microbial elicitors, pathogen- or microbe-associated molecular patterns (PAMPs or MAMPs), damage-associated molecular patterns (DAMPs), pattern recognition receptors (PRRs), pattern-triggered immunity (PTI), plant defense
INTRODUCTION
“Knowing your enemy and yourself, you can fight a hundred battles without disaster.”
-The Art of War, Sun Tzu
The ability to discriminate self from nonself is the key for hosts to launch defense responses and win a war against microbes, which echoes the above saying from Sun Tzu. Compared to humans, sessile plants lack specialized immune cells and adaptive immunity but have developed a series of preformed defense barriers and the innate immune system to counteract nonself invasions (67, 145). The first line of plant innate immunity is triggered upon detection of microbial signatures, which were initially termed general elicitors and are now called pathogen- or microbe-associated molecular patterns (PAMPs or MAMPs) (6, 7, 14, 29, 116). Patterns originally referred to the structurally conserved and functionally essential characteristics commonly found in infectious agents but absent or disguised from the hosts in the studies of mammalian immunity (7, 116). Host-derived endogenous molecules released upon wounding or pathogen infections, named danger- or damage-associated molecular patterns (DAMPs), also trigger immune responses. These patterns are recognized by plasma membrane (PM)-resident pattern recognition receptors (PRRs) and activate immune responses against a broad spectrum of pathogens, collectively termed pattern-triggered immunity (PTI) (14, 29, 67). PTI contributes to host defense against nonadapted pathogens. Host-adapted pathogens secrete a myriad of effectors into plant cells, some of which are sensed by intracellular resistance (R) proteins, leading to effector-triggered immunity (ETI) in a pathogen race-specific manner (9, 67). Recent advances in the identification of plant PRRs and downstream signaling events suggest the blurred boundary between PTI and ETI. In addition, local induction of ETI and PTI could trigger plant resistance systemically against subsequent infections in distal tissues, a phenomenon called systemic acquired resistance (145). Recognition of nonself appears to trigger seemingly chaotic responses, yet hosts launch harmonic signaling events to warrant defense against invading pathogens. In this review, we aim to provide a current view of plant PTI, with a focus on the identification and characterization of microbial patterns and PRRs, physiological and cellular responses upon pattern recognition, and signaling events leading to PTI.
IDENTIFICATION AND CHARACTERIZATION OF PATTERNS
It has long been observed that microbial-derived compounds, including general and specific elicitors, could trigger defense responses in plants. As the genetic, molecular, and structural bases of these elicitors and the plant perception systems are being revealed, general elicitors perceived by cell-surface receptors were termed PAMPs or MAMPs, whereas pathogen-specific elicitors detected by intracellular immune sensors were often referred to as effectors (9). MAMPs from bacteria, fungi, and oomycetes, as well as plant-derived DAMPs, are diverse classes of biomolecules known to include peptides, lipophilic fatty acids, and oligosaccharides (Table 1). In this section, we review the origins and characteristics of MAMPs and DAMPs.
Table 1.
The summary of identified microbe-associated molecular patterns (MAMPs)/damage-associated molecular patterns (DAMPs) and corresponding receptors
| Category of origins | Molecules/fragments | Sources | PRRs | PRR types | References |
|---|---|---|---|---|---|
| Bacteria | flg22 | Bacterial flagellin | FLS2 (Arabidopsis and other plants) |
LRR-RLK | 23, 51 |
| flgII-28 | Flagellin from Pseudomonas syringae pathovars | FLS3 (Solanaceae) | LRR-RLK | 61 | |
| elf18 | Bacterial elongation factor Tu (EF-Tu) | EFR (Brassicaceae species) | LRR-RLK | 177 | |
| Peptidoglycan (PGN) | Bacterial cell wall component | LYM1/LYM3 (Arabidopsis) | LysM-RLP | 160 | |
| OsLYP4/OsLYP6 (Rice) | LysM-RLP | 91 | |||
| Lipopolysaccharide (LPS) | Bacterial cell wall component | LORE (putative) (Arabidopsis) | Lectin-RLK | 128 | |
| RaxX | Secreted protein from Xanthomonas oryzae pv. oryzae | XA21 (putative) (Rice) | LRR-RLK | 124, 144 | |
| csp22 | Bacterial cold shock protein | NbCSPR (putative) (tobacco) | LRR-RLP | 134 | |
| eMAX | A proteinaceous component from Xanthomonas axonopodis pv. citri | ReMAX (RLP1) (putative) (Brassicaceae) | LRR-RLP | 65 | |
| PrpL | Serine protease from Pseudomonas aeruginosa PA14 | Unknown | Unknown | 22 | |
| Fungi | Chitin | Fungal cell wall | LYK5/CERK1 (Arabidopsis) | LysM-RLK | 20, 95 |
|
OsCEBiP (Rice) |
LysM-RLP | 57, 69 | |||
| EIX | Fungal xylanase | LeEix1/LeEix2 (Tomato) | LRR-RLP | 65, 131 | |
| Endopolygalacturonases (PGs) | Botrytis cinerea pectinases | RLP42 (putative) (Arabidopsis) | LRR-RLP | 169 | |
| SCFE1 | Proteinaceous component from Sclerotinia sclerotiorum | RLP30 (putative) (Arabidopsis) | LRR-RLP | 170 | |
| β-Glucan | Fungal cell wall polysaccharide | Unknown | Unknown | 129 | |
| Ergosterol | Sterol in fungal cell membrane | Unknown | Unknown | 72 | |
| BcSpl1 | B. cinerea cerato-platanin family protein | Unknown | Unknown | 47 | |
| Oomycetes | nlp20 | Necrosis- and ethylene-inducing peptide from oomycetes and other microbes | RLP23 (Arabidopsis) | LRR-RLP | 3 |
| INF1 | Oomycete elicitin | ELR (SmRLP85) (putative) (potato) | LRR-RLP | 34 | |
| Pep-13 | Oomycete transglutaminase | Unknown | Unknown | 17 | |
| XEG1 | Secreted glycoside hydrolase family 12 protein | Unknown | Unknown | 106 | |
| Hepta-β-glucoside | β-Glucan from oomycete cell wall | Unknown | Unknown | 44 | |
| Plants | AtPEPs | Secreted peptides from precursor PROPEPs | PEPR1/PEPR2 (Arabidopsis) | LRR-RLK | 75, 166 |
| PIP1/PIP2 | PAMP-induced secreted peptides | RLK7 (Arabidopsis) | LRR-RLK | 62 | |
| High mobility group box 3 (HMGB3) | Secreted plant proteins | Unknown | Unknown | 25 | |
| Oligogalacturonides (OGs) | Plant cell wall component | WAK1 (putative) (Arabidopsis) | EGF-RLK | 18 | |
| eATP | Extracellular ATP | DORN1 (Arabidopsis) | Lectin-RLK | 26 |
Abbreviations: AtPEPs, plant elicitor peptides from Arabidopsis; EGF, epidermal growth factor; LRR, leucine-rich repeat; LysM, lysine motif; PAMP, pathogen-associated molecular pattern; PROPEPs, precursor proteins of AtPEPs; PRR, pattern recognition receptor; RLK, receptor-like kinase; RLP, receptor-like protein.
Bacterial Microbe-Associated Molecular Patterns
The flagellin monomer, the major component of flagella required for bacterial mobility, is a well-understood MAMP in plants and animals. The 22-amino-acid (aa) synthetic peptide flg22 corresponds to a conserved domain of the amino (N) terminus of flagellin from Pseudomonas species and acts as a potent elicitor in Arabidopsis thaliana and other plants (40). Interestingly, flgII-28, a synthetic peptide corresponding to a motif of flagellin distinct from flg22, elicits defense responses in certain Solanaceae species but not in Arabidopsis (19). Polymorphisms in flagellin among different bacterial species appear to affect the inducibility of plant defense. A single amino acid variation in flg22 from Xanthomonas campestris pv. campestris (Xcc) leads to compromised defense responses in Arabidopsis (148). Flagellin monomers are known to be cleaved by the alkaline protease AprA secreted by Pseudomonas, which likely dampens PTI responses (120).
Cell walls of both Gram-positive and Gram-negative bacteria contain unique peptidoglycan (PGN) networks consisting of alternating sugar components N-acetylglucosamine (GlcNAc) and N-acetylmuramic acid cross-linked by oligopeptides (52). PGN fragments are released by plant-derived lysozymes and act as MAMPs to elicit plant immunity (96). Treatment with PGN but not the breakdown molecules from Staphylococcus aureus activates defense responses in Arabidopsis(52). In addition, purified PGN and muropeptides from Agrobacterium tumefaciens and Xcc elicit defense responses in Arabidopsis (37). Lipopolysaccharide (LPS), the major component of the outer membrane of Gram-negative bacteria, is a MAMP commonly used in triggering potent immune responses in mammals and also possesses elicitor activity in plants. LPS consists of lipid A, a core oligosaccharide, and an O-polysaccharide (167). Lipid A is relatively conserved and sufficient to trigger immune responses, and the core oligosaccharide is able to enhance defense in Arabidopsis (128).
The abundant bacterial protein elongation factor Tu (EF-Tu) triggers immune responses in various plant species (76). The conserved N-terminal 18-aa peptide (elf18) is sufficient to elicit responses in Brassicaceae species. In contrast, EFa50, a synthetic 50-aa peptide corresponding to the middle region of EF-Tu, functions as a MAMP in rice, suggesting that different EF-Tu recognition systems exist in different plant families (48). The N-terminal 22-aa peptide (cps22) of bacterial cold-shock proteins from S. aureus acts as a MAMP in tobacco (39). The serine protease PrpL secreted by Pseudomonas aeruginosa elicits defense responses in Arabidopsis (22). Notably, the catalytic triad required for its protease activity is essential for the elicitor activity. Whether PrpL itself or its product is perceived as a MAMP requires further investigation. An enigmatic proteinaceous eMax from Xanthomonas axonopodis pv. citri (Xac) triggers responses in several species of Brassicaceae (65). The tyrosine-sulfated protein RaxX secreted through the type I secretion system of Xanthomonas oryzae pv. oryzae (Xoo) acts as a MAMP in rice (124).
Fungal Microbe-Associated Molecular Patterns
Chitin, a homopolymer of β−1,4-linked GlcNAc, is a major component of fungal cell walls. Chitin fragments, N-acetylchitooligosaccharides consisting of more than five monomers and chitosan, a deacetylated chitin derivative, have been demonstrated as MAMPs in both dicot and monocot plants (141). β-glucans are the most abundant fungal cell wall polysaccharides. The β-glucans from Magnaporthe oryzae trigger production of phytoalexins, which are immune-related antimicrobial compounds, in rice (165). Chitin and glucans could be released by plant apoplastic chitinases and glucanases that decompose fungal cell walls. Ergosterol, the most abundant sterol in fungal cell membranes, is also recognized by plants and triggers immune responses (72).
Ethylene-inducing xylanase (EIX) from Trichoderma viride induces defense responses in tomato and potato independently of its enzymatic activity. The elicitation epitope was mapped to a pentapeptide on the exposed β-strand of EIX (131). Endopolygalacturonases (PGs), a class of fungal pectinases hydrolyzing plant pectin polysaccharides, trigger defense responses in grapes independently of their enzymatic activity, suggesting that these proteins themselves are perceived and activate defense responses (122). Furthermore, in tomato, tobacco, and Arabidopsis, the ceratoplatanin family protein BcSpl1 secreted by Botrytis cinerea triggers the immunity-related cell death response called the hypersensitive response (HR) (47). In addition, the proteinaceous elicitor SCFE1 from Sclerotinia sclerotiorum culture filtrate induces typical PTI in Arabidopsis (170). The molecular identity of SCFE1 is still elusive.
Oomycete Microbe-Associated Molecular Patterns
Branched β−1,3- or β−1,6-glucans, the most abundant components of Phytophthora cell walls, trigger defense responses in soybean (44). The active fragment was identified as hepta-β-glucoside, a molecule that bears high binding affinity to the cell membranes of certain legumes. The essential fatty acids in oomycete species, such as eicosapentaenoic acid and arachidonic acid, are also able to trigger immune responses in plants (129).
Cellulose-binding elicitor lectin (CBEL) from Phytophthora cell walls is able to bind to crystalline cellulose and isolated plant cell walls. Infiltration of purified CBEL triggers necrosis and immune responses in tobacco and Arabidopsis (71). Cryptogein, a 10-kDa protein secreted by Phytophthora cryptogea, elicits HR and other defense responses in tobacco (11). Elicitins are structurally conserved lipid-binding proteins secreted by almost all pathogenic oomycetes that trigger defense responses in some species of Solanaceae and Brassicaceae (158). Pep-13, a conserved epitope derived from the calcium-dependent transglutaminase involved in cross-linking oomycete cell walls, activates defense responses in parsley and potato (17). Necrosis- and ethylene-inducing peptide 1–like proteins (NLPs), which are produced by a wide range of oomycetes, bacteria, and fungi, trigger necrosis and defense responses in dicot plants. Noncytotoxic NLPs discovered from oomycete and fungal species also confer immune activation ability (3). This elicitor activity was further pinpointed to a conserved 20-aa region (nlp20) (13). XEG1, a glycoside hydrolase 12 protein from Phytophthora sojae, is an important virulence factor and also acts as a MAMP in soybean (106).
Plant-Derived Damage-Associated Molecular Patterns
Molecules derived from the plant itself upon damage or pathogen infections are able to activate and/or amplify defense responses. Plant DAMPs include small peptides, nucleotides, and cell wall–derived oligosaccharides. Plant elicitor peptides (AtPEPs) are 23-aa peptides derived from the precursor PROPEPs in Arabidopsis. Among six family members, AtPEP1 is the first shown to be induced by wounding, jasmonic acid (JA), and ethylene (ET), and it activates defense responses(63). On the basis of transcriptional upregulation upon PTI elicitation, a group of small secreted peptides, designated PIPs, with 11 members in Arabidopsis, were proposed to function as DAMPs(62). Recently, the high mobility group box 3 (HMGB3) protein was shown to trigger PTI responses and mediate resistance against B. cinerea in Arabidopsis (25). The intracellular HMGB3 is released to the extracellular matrix when cell membranes are damaged (25).
Extracellular adenosine-5′-triphosphate (eATP) is involved in various cellular processes and triggers immune responses in plants and animals (150). The extracellular pyridine nucleotides NAD or NADP induce defense responses and resistance against bacterial pathogens in Arabidopsis (172). Oligogalacturonides (OGs), oligomers of α−1,4-linked galacturonosyl residues derived from plant cell walls, activate plant defense responses and induce resistance against B. cinerea in Arabidopsis (43). OGs could be released by the hydrolytic activity of PGs from B. cinerea, which degrade polysaccharide homogalacturonan, a major component of pectins in plant cell walls (169). Interestingly, a small O-glycosylated peptide from a group of parasitic plants belonging to Cuscuta spp. elicits typical immune responses in tomato, suggesting that plants could sense danger not only from microbes but also from invading plants (58).
CELLULAR AND PHYSIOLOGICAL RESPONSES TRIGGERED BY PATTERNS
A series of coordinated cellular and physiological responses occur in plants upon pattern recognition (Figure 1). Distinct molecular and biochemical modules mediate differential cellular responses and integrate positive or negative feedback regulations to culminate in plant PTI, which collectively contributes to plant resistance against a broad spectrum of pathogens. These responses, dictated by spatial and temporal dynamics, occur instantaneously as early as seconds to minutes or as late as hours to days (Figure 2). Despite differing in thresholds, amplitudes, and timing, shared common responses upon pattern recognition represent hallmarks of PTI elicitation (Figures 1 and 2).
Figure 1.
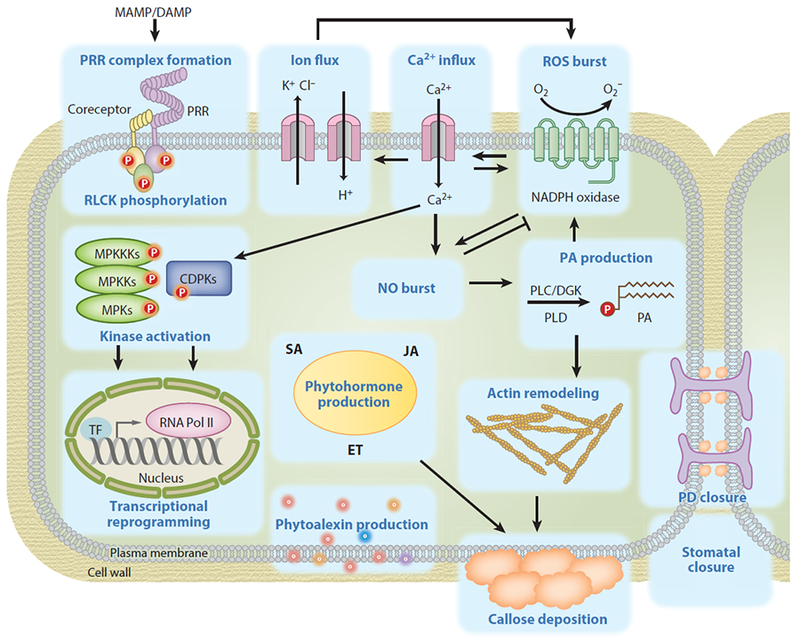
Cellular and physiological responses triggered by patterns in plants. Plant cell surface–resident pattern recognition receptors (PRRs) perceive microbe-associated molecular patterns (MAMPs) or damage-associated molecular patterns (DAMPs) and recruit the coreceptors, leading to a series of intertwined cellular and physiological responses. PRR complex formation is accompanied by rapid transphosphorylation in the complex and phosphorylation of receptor-like cytoplasmic kinases (RLCKs). Activation of PRR complexes activates mitogen-activated protein kinase (MAPK) cascades and calcium-dependent protein kinases (CDPKs), which regulate gene transcriptional changes and other cellular responses. The hallmarks of pattern-triggered immunity (PTI) responses include calcium influx, ion efflux, actin filament remodeling, plasmodesmata (PD) and stomatal closure, callose deposition, and production of reactive oxygen species (ROS), nitride oxide (NO), phosphatidic acid (PA), phytoalexins, and phytohormones. Collectively, these responses contribute to plant resistance against a variety of pathogens. The potential connections among different responses, which were mainly derived from the studies using inhibitors, are indicated with an arrowed line for positive regulation and a T-shaped line for negative regulation. Abbreviations: DGK, diacylglycerol kinase; ET, ethylene; JA, jasmonic acid; PLC, phospholipase C; PLD, phospholipase D; SA, salicylic acid; TF, transcription factor.
Figure 2.

Temporal dynamics of hallmark responses upon pattern perception. Pattern perception by pattern recognition receptors (PRRs) activates complex signaling events, culminating in a series of cellular and physiological responses with spatial and temporal dynamics. Some of the responses occur as early as seconds to minutes or as long as hours to days. Some responses are temporarily induced, whereas some are long lasting. The exemplary hallmark responses are shown with a timescale on the top. Images modified with permission from References 52 (extracellular alkalization) and 53 (stomatal closure), and adapted by permission from Macmillan Publishers Ltd: Reference 3, copyright 2015 (ethylene production), Reference 23, copyright 2007 (growth inhibition), and Reference 128, copyright 2015 (Ca2+ influx). Abbreviations: Co-IP, co-immunoprecipitation; LPS, lipopolysaccharide; MAPK, mitogen-activated protein kinase; PGN, peptidoglycan; RLCK, receptor-like cytoplasmic kinase; RLU, relative light unit; ROS, reactive oxygen species; WB, Western blot.
Increase of Cytosolic Ca2+ Concentration ([Ca2+]cyt)
The rapid and drastic increase of [Ca2+]cyt is among the earliest cellular responses upon different MAMP or DAMP treatments. At the resting state, [Ca2+]cyt is maintained at ~100–200 nanomoles, 104 times fewer than that in the apoplast or cellular organelles. Perception of MAMPs typically triggers the increase of [Ca2+]cyt with a delay of ~30–40 sec and a peak at ~2–6 min, which then lasts for at least 30 min before declining to the resting state (127). The spatial and temporal changes in [Ca2+]cyt vary in terms of the peak time, amplitude, and duration in response to different types and dosages of MAMPs (138). Cryptogein, flg22, and β-glucan trigger a biphasic [Ca2+]cyt increase, whereas a chitin-induced [Ca2+]cyt burst exhibits only a single peak. PGN triggers a relatively weaker [Ca2+]cyt increase than flg22 (52). DAMPs, including OGs, eATP, and AtPEPs, also induce typical [Ca2+]cyt increase (78, 151).
[Ca2+]cyt elevation can be due to an uptake of Ca2+ from the extracellular space and/or Ca2+ mobilization from organelles. Lines of evidence support that MAMP- triggered [Ca2+]cyt increase comes from a sustained extracellular Ca2+ influx from the apoplast (138). Chelation of extracellular Ca2+ or pretreatment with Ca2+ surrogates abolishes [Ca2+]cyt bursts, especially the first peak(78). However, the role of internal Ca2+ stores in MAMP-triggered [Ca2+]cyt increase remains ambiguous. Inhibition of Ca2+ release from internal stores by neomycin or U-73122 affects the second peak of a [Ca2+]cyt burst triggered by β-glucans, OGs, or eATP (151). External Ca2+ influx may predominantly contribute to the first peak, whereas the release of Ca2+ from internal stores is required for the second peak of MAMP-induced [Ca2+]cyt elevation.
[Ca2+]cyt is regulated by the coordination of passive fluxes (Ca2+ channels) and active transporters (Ca2+-ATPases and Ca2+-antiporters) across the PM and endomembranes. However, the identity of Ca2+ channels mediating MAMP-induced [Ca2+]cyt influx remains elusive. Mutation in Arabidopsis cyclic nucleotide-gated cation channel 2 (CNGC2) abolishes AtPEP3- and LPS- but not flg22-induced [Ca2+]cyt bursts (104). In addition, inhibiting the activity of ionotropic gluta-mate receptor type Ca2+ channels (iGluRs) blocks cryptogein- and flg22-induced [Ca2+]cyt bursts in tobacco and Arabidopsis (138). Moreover, rice two-pore channel 1 (OsTPC1) is involved in xylanase-induced [Ca2+]cyt bursts (56). Conversely, Ca2+-ATPases, mediating Ca2+ efflux to the extracellular matrix and endomembrane compartments, also play certain roles in MAMP-induced [Ca2+]cyt increase. Silencing of the endoplasmic reticulum (ER)-localized type 2B Ca2+-ATPase (NbCA1) in Nicotiana benthamiana increases cryptogein-triggered [Ca2+]cyt bursts in both amplitude and duration (176). Furthermore, PM-resident autoinhibited Ca2+-ATPases isoform 8 (ACA8) and ACA10 are essential for flg22-triggered [Ca2+]cyt influx in Arabidopsis (46).
Plasma Membrane Depolarization and Extracellular Alkalinization
MAMP perception induces rapid Cl−, NO3−, and K+ effluxes, as well as H+ influx across the PM, which often leads to membrane depolarization and extracellular alkalinization (66). Cytoplasmic acidification and PM depolarization are detected within 1 min in cryptogein-treated tobacco cells (125). Application of crude cell wall extracts from P. sojae to parsley cells also causes rapid alkalinization of the culture medium (64). Strong membrane potential depolarization and extracellular alkalinization in Arabidopsis mesophyll cells were recorded within 1 min after flg22 or elf18 treatment (66). Recovery of flg22-induced membrane potential to the resting state takes more than 1 h, a long-lasting process compared with that triggered by abiotic stresses (66). PGN induces a slower but more persistent increase in extracellular alkalinization than flg22 (52).
PM-resident H+-ATPases (AHAs) are considered the primary pumps to transfer protons from cytosol to the extracellular matrix to establish the PM potential (36). Upon PTI elicitation, AHA activity is likely downregulated through the sequential phosphorylation of specific residues and dynamic localization to different PM microdomains (36). Upon MAMP perception, H+ influx is accompanied by effluxes of Cl− and K+. Interestingly, flg22 treatment triggers a considerably higher peak value of Cl− efflux than of H+ influx, suggesting the involvement of anion channels in establishing PM depolarization (66). Integrin-linked kinase 1 (ILK1), which interacts with the high-affinity K+ transporter HAK5, positively regulates flg22-induced PM depolarization, implicating the involvement of K+ efflux in PM depolarization (16). Ca2+ signaling likely functions upstream of PM depolarization and extracellular alkalinization, as lanthanum, a PM Ca2+ channel blocker, inhibits flg22-induced PM depolarization (66). The role of other ion channels in PTI signaling remains to be determined.
Apoplastic Reactive Oxygen Species Bursts
Reactive oxygen species (ROS) include partially reduced forms of oxygen such as superoxide (O2−), hydrogen peroxide (H2O2), and hydroxyl radicals (•OH). Transient and rapid generation of apoplastic ROS, referred to as a ROS burst, is a hallmark of the early response to MAMP treatment (153). Typically, a ROS burst is initiated within ~4–6 min, reaches its peak ~10–15 min, then gradually declines to the resting state ~30 min after MAMP treatments in various plant species (64). ROS can act as a toxin barrier against subsequent pathogen infections. ROS are also involved in strengthening plant cell walls by oxidative cross-linking of polymers. In addition, ROS are versatile signaling molecules, mediating multiple responses (135, 153).
Both PM-localized NADPH oxidases and cell wall–associated peroxidases are involved in a MAMP-induced ROS burst. NADPH oxidases transfer electrons from cytosolic NADPH or NADH to apoplastic oxygen, leading to the production of O2−, which is then converted to H2O2 by superoxide dismutase (153). Among 10 respiratory burst oxidase homolog (RBOH) genes encoding NADPH oxidases in Arabidopsis, RBOHD is likely the major enzyme in MAMP-induced ROS production, as a MAMP-induced ROS burst is markedly compromised in the rbohD mutant (115). In addition, the class III apoplastic peroxidases that catalyze oxidoreduction between H2O2 and various reductants contribute to a MAMP-induced ROS burst. Mutations in Arabidopsis peroxidase 33 (PRX33) and PRX34 diminish a MAMP-induced ROS burst (31). The relationship between peroxidase- and RBOH-mediated ROS production remains unknown.
As signaling molecules, ROS interact with other early responses triggered by MAMPs (Figure 1). It appears that [Ca2+]cyt increase is a prerequisite for ROS production, as lanthanum abrogates flg22- and elf18-induced ROS bursts in Arabidopsis (127). A ROS burst is also likely downstream of ion fluxes, as inhibitors of ion channels block a ROS burst (64). In addition, the rbohD mutant does not affect flg22-induced PM depolarization (66). However, a feedback regulation of [Ca2+]cyt by ROS likely exists because [Ca2+]cyt increases in response to H2O2 treatment. MAMP-induced ROS burst is important for inducing the second peak or prolonged plateau of [Ca2+]cyt (127). The causative relationship among these events is rather complex, and their genetic and biochemical bases await further elucidation.
Nitride Oxide Production
A rapid burst of nitric oxide (NO), a free radical gas that has an extremely short half-life and is a hallmark of animal innate immune responses, has also been suggested to act as an important secondary messenger and antimicrobial agent in plant defense (135). Through in vivo imaging coupled with fluorophore staining, an NO burst is detected within a few minutes upon treatment with cryptogein, xylanase, flg22, PGN, and LPS in various plant species (135). DAMPs, including eATP and AtPEPs, have also been shown to induce an NO burst (45, 105).
In animals, NO is synthesized by NO synthases (NOS) through converting L-arginine to NO and L-citrulline. However, plants possess several distinct and unique pathways for NO production in different cellular compartments with interconnected activation and feedback regulations (135). The exact sources and enzymes mediating MAMP-induced NO biosynthesis remain elusive in plants. The NOS inhibitors L-NNA or L-NAME largely block MAMP- or DAMP-induced NO bursts in tobacco and Arabidopsis, suggesting the involvement of NOS-like enzymes (4). In Arabidopsis, mutation of NOS1, a plant homolog of snail NOS, significantly reduces LPS-induced NO production and gene expression, suggesting a role of NOS1 in PTI (167).
MAMP-induced NO production seems to be tightly connected with Ca2+ signaling. Ca2+ channel blockers inhibit MAMP-induced NO production (102). Mutation in CNGC2, which encodes a nonselective cation channel, inhibits LPS-induced NO production in Arabidopsis (4). Calmodulin-like protein 24 (CML24) is also required for LPS-induced NO production (102). During MAMP-triggered stomatal closure, NO acts downstream of the ROS burst but possesses a negative feedback regulation of NADPH oxidases through S-nitrosylation (Figure 1) (5).
Phosphatidic Acid Production
Phosphatidic acid (PA), an important intermediate in lipid biosynthesis, is also considered to be a key signaling molecule regulating various cellular activities and environmental responses. In plant immunity, PA is potentially involved in the regulation of ROS production, MAP kinase (MAPK) activation, defense gene induction, and actin remodeling (152). The levels of PA and its phosphor-ylated derivative diacylglycerol pyrophosphate (DGPP) are elevated in tomato suspension cells after treatments with xylanase, chitin, and flg22. The accumulation of PA occurs in 2 min and reaches its peak approximately 8 min after treatment (157). A rapid and transient PA induction was also detected in chitosan-treated rice suspension cells (164).
Two PA biosynthetic pathways exist in plants: the direct hydrolysis of phospholipids by phospholipase D (PLD) and the combined actions of phospholipase C (PLC) and diacylglycerol kinase (DGK) to phosphorylate diacylglycerol (DAG) to PA. The flg22- and chitosan-induced PA production is predominantly from the PLC/DGK pathway in tomato (157). Consistently, application of PLC or DGK inhibitors reduces xylanase-induced PA levels (77). In addition, xylanase also slightly activates the PLD pathway in both studies. Both PLC and PLD activities are elevated in rice cells upon chitosan treatment (164). It is possible that different MAMPs induce PA production via distinct pathways yet the genetic evidence for the involvement of PA in PTI signaling remains elusive. A mutation in PLD isoform PLDδ displays delayed defense gene induction upon chitin treatment and compromised resistance against barley powdery mildew (121). In addition, PA exhibits extensive cross-talk with other signaling molecules (Figure 1). An NO burst is required for PA production, whereas inhibition of either PLC or DGK activity decreases ROS production in xylanase-treated tobacco cells (77). Consistently, PA or DAG treatment directly induces ROS production in rice cells (164). PA stimulates RBOHD activity to promote ROS production in plant hormone abscisic acid (ABA)-mediated stomatal closure (174). It remains unknown whether this is the case in plant PTI signaling.
Stomatal Closure
Stomata are the openings formed by two guard cells on the leaf surface for gas exchange and water transpiration. As the major routes for pathogen entry, stomatal opening and closure are regulated during pathogen infections. MAMP or DAMP treatment induces stomatal closure within 1 h, which serves as an important mechanism to limit pathogen entry (110). MAMP-induced stomatal closure is regulated by early signaling molecules, such as NO and ROS, as well as the plant hormone ET and several oxylipin molecules (5).
ABA is a key regulator of stomatal closure in abiotic stresses. It appears that flg22-induced stomatal closure shares common regulators with ABA-mediated stomatal closure (53). For example, the open stomata 1 (OST1) kinase regulates both ABA- and flg22-induced stomatal closure (110). ABA-activated OST1 subsequently phosphorylates S-type anion channel SLAC1, leading to SLAC1 activation and anion efflux, such as the efflux of Cl−, a major contributor of stomatal closure. SLAC1 and its homolog 3 (SLAH3) are also required for flg22-induced stomatal closure. Stomata of the slac1slah3 mutant are completely insensitive to flg22 stimulation (53). Flg22 inhibits inward K+ channel and H+-ATPase activity that regulates stomatal reopening (171). It remains unknown whether and how the OST1 kinase and anion channels are stimulated upon MAMP perception in guard cells.
Actin Filament Remodeling
The actin cytoskeleton, which supports cell rigidity and cytoplasmic streaming among different compartments, plays a central role in the activation of host defenses in animals. In plants, a growing body of evidence indicates that MAMP perception induces rapid and transient changes in actin organization, including an increase of both actin filament abundance and filament bundling (32). The increase in the actin filament abundance occurs as early as 15 min, whereas the enhanced filament bundling is detectable at 18 h after Pseudomonas syringae inoculation. Treatment of flg22 or chitin only triggers a rapid increase in actin filament abundance and does not enhance the extent of filament bundling (60). Detailed analyses of single-filament dynamics suggest that MAMP treatment increases the maximal length and lifetime of the growing actin filaments; meanwhile, it slows down the filament destruction process by inhibiting the severing activity, likely via actin depolymerization factor 4 (ADF4). It appears that the actin filament elongation rate and the regrowth frequency are not altered upon MAMP treatments (84).
Consistent with the above observations, inhibitors blocking actin polymerization promote Arabidopsis susceptibility to P. syringae, implicating an important role of actin cytoskeleton remodeling in plant immunity (60). Moreover, MAMP-triggered callose deposition is impaired when either the actin cytoskeleton or actin dynamics is perturbed (59). Interestingly, ADF4 is required for elf26 (a longer version of elf18) but not for chitin-induced actin filament abundance, suggesting that distinct signaling pathways contribute to different MAMP-induced actin dynamics (59). Capping proteins (CPs), key regulators in controlling the number of filament ends and the amount of filament-filament annealing, are also involved in MAMP-induced actin cytoskeleton remodeling and callose deposition (84). Notably, PA binds and inhibits CP activity in vitro, and exogenous PA treatment increases the actin filament abundance. It is postulated that MAMP-induced actin remodeling is due to the negative regulation of CP by PA signaling (84). The detailed regulation between PA and CP in MAMP-induced actin dynamics still awaits elucidation.
Production of Antimicrobial Compounds and Phytohormones
Production of plant antimicrobial compounds referred to as phytoalexins has long been observed during plant-pathogen or -insect interactions. Camalexin, a characteristic indolic phytoalexin derived from tryptophan in Arabidopsis and related Brassicaceae species, is strongly induced by various pathogens, including bacteria, fungi, viruses, and oomycetes (1). However, MAMP-induced camalexin production is often not as pronounced as that induced by pathogen infections. It was reported that camalexin can be induced upon flg22 treatment in Arabidopsis roots (112). Indoleglucosinolates, another group of tryptophan-derived secondary metabolites, also participate in Arabidopsis defense against bacteria and fungi and are required for flg22-induced callose deposition (27). The tryptophan-derived cyanogenic compound, 4-hydroxyindole-3-carbonyl nitrile, is induced by flg22 treatment and contributes to defense responses in Arabidopsis (126). In rice, chitin elicits accumulation of terpenoids, the major phytoalexins in monocots (136). Phenolic compounds, such as phenylamides, are another type of phytoalexins that accumulate in response to MAMP treatment and mainly participate in cell wall reinforcement upon pathogen attacks in rice(24). With the advancement of mass spectrometry–based metabolomics and analytical chemistry, involvement of various types of phytoalexins in PTI will likely be elucidated.
Plant defense hormones, including salicylic acid (SA), ET, and JA, have been implicated in PTI responses (50). Consistently, the biosynthesis of these hormones is induced upon MAMP treatments. ET production has been established as a sensitive bioassay to monitor PTI responses triggered by MAMPs, including flg22, elf18, EIX, and nlp20 (3, 8, 40, 76) (Figure 2). Flg22 induces ET production at 1 h and peaks at 4 h in Arabidopsis seedlings (97). A moderate accumulation of SA was detected upon flg22 or LPS treatment in Arabidopsis (154). In addition, JA production, which is usually induced by necrotrophic pathogen infections, is elevated upon oomycete-derived Pep-13 treatment in potato (54). By analyzing the Arabidopsis mutants deficient in these three hormone pathways, it was shown that SA, JA, and ET act positively in flg22- and elf18-mediated PTI and mainly contribute to some of the late PTI responses (155). Additionally, other plant hormones, including brassinosteroids, auxins, ABA, cytokinins, and gibberellins, are also implicated in plant immunity (29).
Callose Deposition
Callose is a high molecular weight β−1,3 glucan polymer that strengthens weak or compromised sections of plant cell walls (101). Callose deposits in a timely manner at the site of infections, forming a prominent physical barrier for pathogen attacks and often leading to the formation of papillae. Various MAMPs or DAMPs, including flg22, elf18, chitin, PGN, and OG, induce callose deposits in plant roots, cotyledons, and leaves (101). Powdery mildew resistant 4 (PMR4), a glucan synthase-like protein, is involved in callose synthesis and plays an important role in plant resistance against fungal infections. However, flg22- or chitosan-induced callose deposition is not completely blocked in the pmr4 mutant, suggesting the involvement of other callose synthases in response to MAMPs (27, 101).
Callose deposition is regulated at multiple levels. Loss of RBOHD, a key enzyme in generating apoplastic ROS, results in compromised flg22- and OG-induced callose deposits (49). In addition, MAMP-induced callose deposition is almost completely abolished in the prx33prx34 mutants(31), suggesting both NADPH oxidase– and peroxidase–produced ROS are required for MAMP-induced callose biosynthesis. Mutations in ethylene response 1 (ETR1) or ethylene insensitive 2 (EIN2), which block ET perception or signaling, respectively, largely diminish MAMP-induced callose deposition, pointing to a critical role of ET in MAMP-induced callose deposition (27).
Plasmodesmata Closure
Plasmodesmata (PD) are cytoplasmic channels that connect adjacent cells across the cell walls and mediate molecular exchanges among different cells in response to developmental, environmental, and pathogen signals (79). Regulation of PD permeability plays a significant role in plant-pathogen interactions (80). Chitin and flg22 induce PD closure by stimulating PD callose deposition in Arabidopsis (38). PD-located protein 5 (PDLP5), residing in the central region of PD channels, inhibits PD trafficking in Arabidopsis. Interestingly, PDLP5 is induced upon bacterial infections and plays a positive role in plant defense against P. syringae infections, indicating the importance of PD closure in plant disease resistance (80). It is also plausible that signaling molecules and/or defense-related metabolites are transmitted via PD from infected cells. Recently, it has been shown that callose synthase 1 (CALS1) and CALS8 contribute to SA-mediated and ROS-dependent PD callose accumulations, respectively, both of which require PDLP5 (30). Apparently, plants employ specialized components and pathways to regulate PD dynamics in response to different signaling molecules.
PATTERN RECOGNITION RECEPTOR COMPLEXES PERCEIVING PATTERNS
Plants perceive patterns by cell surface–resident PRRs, which are mainly transmembrane receptor-like kinases (RLKs) or receptor-like proteins (RLPs). PRRs often dynamically complex with coreceptors and other regulatory proteins to ensure prompt signaling activation and attenuation (Figure 3). Both RLKs and RLPs comprise an extracellular domain and a transmembrane domain, but RLPs lack an intracellular kinase domain. The various extracellular domains, including leucine-rich repeats (LRRs), lysine motifs (LysMs), lectin motifs, and epidermal growth factor (EGF)-like domains, mainly determine the specificity of ligand binding (12, 29).
Figure 3.

Elicitation of early pattern recognition receptor (PRR) signaling in Arabidopsis and rice. (a) flg22-induced early signaling in Arabidopsis. (i) In the resting state, FLS2 does not form a stable complex with BAK1 but interacts with several receptor-like cytoplasmic kinases (RLCKs) (including BIK1, PCRK1/2, and BSK1), NADPH oxidase RBOHD, and heterotrimeric G proteins (XLG2/Gβ/Gγ). BAK1 interacts with the pseudokinase BIR2 to block the interaction between BAK1 and FLS2, the phosphatase PP2A to reduce its kinase activity, and the E3 ligases PUB12 and PUB13. BIK1 also associates with BAK1, the heterotrimeric G proteins, the phosphatase PP2C38, and CPK28. CPK28 phosphorylates BIK1, likely promoting BIK1 degradation through the 26S proteasome, whereas the heterotrimeric G proteins may suppress BIK1 degradation. (ii) Upon flg22 perception, FLS2 associates and transphosphorylates with BAK1, subsequently releasing BIK1, BSK1, BIR2, and the heterotrimeric G proteins from the FLS2-BAK1 complex. BIK1 phosphorylates RBOHD to regulate reactive oxygen species (ROS) production. RBOHD activity is also regulated by CDPKs and XLG2. The activated PRR complex further activates two mitogen-activated protein kinase (MAPK) cascades (MEKK1-MKK1/2-MPK4 and MEKK-MKK4/5-MPK3/6) and CPK4/5/6/11, which regulate gene transcriptional changes through phosphoregulation of transcription factors and the general transcription machinery. The FLS2 signaling is also positively regulated by PCRK1/2 and negatively regulated by MKKK7. (iii) To attenuate the activated PRRs, the plant E3 ubiquitin ligases PUB12 and PUB13 are phosphorylated by BAK1 and recruited to FLS2 for FLS2 ubiquitination and degradation in the vacuole or 26S proteasome. flg22 perception also induces FLS2 endocytosis and intracellular trafficking, which may lead to FLS2 degradation or full activation of defense responses. Other PRR complex components, such as BAK1 and BIK1, may also undergo endocytosis and degradation. Similar signaling mechanisms also apply to other PRRs, in particular, leucine-rich repeat (LRR)–receptor-like kinase (RLK)-encoded PRRs. Abbreviations: LE/MVB, late endosome/multivesicular body; TGN/EE, trans-Golgi network/early endosome. (b) Chitin-induced early signaling in Arabidopsis and rice. (i) In Arabidopsis, LYK5 heterodimerizes with CERK1 upon chitin perception. Phosphorylation of CERK1 further phosphorylates and activates PBL27, which directly interacts with MAPKKK5 and phosphorylates and activates the MAPK cascade (MAPKKK5-MKK4/5-MPK3/6). CERK1, to a lesser extent, also phosphorylates BIK1, which contributes to a ROS burst. (ii) In rice, OsCERK1 is recruited to chitin elicitor-binding protein (CEBiP) upon chitin perception. Activated OsCERK1 phosphorylates OsRLCK185 and likely OsRLCK176, which are required for the activation of chitin-induced MAPK cascades consisting of OsMAPKKK-OsMKK4-OsMPK3/6. OsCERK1 also phosphorylates OsRacGEF1, leading to the activation of OsRac1. OsRac1 is required for the chitin-induced MAPK activation and regulates ROS burst by interacting with OsRBOH.
Leucine-Rich Repeat Domain–Containing Pattern Recognition Receptors
LRR domain–containing PRRs include both LRR-RLKs and LRR-RLPs, which primarily function in peptide ligand perception and signaling. LRR-RLKs constitute the largest group of RLKs, with more than 200 members in Arabidopsis, and include some well-studied PRRs, such as flagellin-sensing 2 (FLS2) recognizing flg22, EF-Tu receptor (EFR) recognizing elf18, and PEP1 receptor 1 (PEPR1)/PEPR2 recognizing AtPEPs (51, 75, 166, 177). Crystal structure analyses indicate that both N- and C-terminal segments of flg22 bind to the inner surface of the FLS2 LRR solenoid (149). Tomato LRR-RLK SlFLS3 perceives flgII-28, a flagellin fragment distinct from flg22 (61), indicating that different regions of flagellin are recognized by different LRR-RLK receptors, likely to ensure the specificity of recognition. Arabidopsis RLK7 recognizes DAMPs PIP1 and PIP2 (62). Rice XA21 is the first cloned RLK that confers resistance against different X00 strains (144). The tyrosine-sulfated protein RaxX from X00 is a potential ligand of XA21 (124).
LRR-RLPs function in PTI signaling either as receptors or as signaling components. RLP23 is required for nlp20-triggered responses in Arabidopsis and specifically binds to nlp20 in vitro. Ec-topic expression of RLP23 causes potato plant sensitivity to nlp20 and enhances resistance against oomycete and fungal pathogens (3). Thus, RLP23 is a bona fide receptor of nlp20. Tobacco LRRRLP CSP receptor (NbCSPR) associates with csp22 and is required for csp22-triggered defense responses (134). Recognition of elicitin INF1 from Phytophthora infestans is mediated by the potato LRR-RLP elicitin response (StELR) (34). Tomato LRR-RLPs SlEIX1 and SlEIX2 associate with EIX and heterodimerize in a ligand-dependent manner. SlEIX2 activates defense responses that are suppressed by SlEIX1 (130). In addition, Arabidopsis RLP30, RLP1, and RLP42 and tomato LRR-RLP cuscuta receptor 1 (CuRe1) are genetically required for the defense responses triggered by SCFE1, eMAX, and PG and the parasitic plant Cuscuta peptide ligand, respectively (58, 65, 169, 170). Tomato LRR-RLPs, Cfs and Ve1, confer resistance against fungal pathogens Cladosporium fulvum and Verticillium dahliae carrying the cognate avirulence proteins, respectively (33, 133). This resistance occurs in a race-specific manner; thus, Cfs and Ve1 are considered as R proteins.
LysM Domain–Containing Pattern Recognition Receptors
Plant LysM domain–containing PRRs, including LysM-RLKs and LysM-RLPs, function in GlcNAc-containing microbial pattern perception and signaling (Figure 3b). Rice chitin elicitor-binding protein (OsCEBiP), a LysM-RLP, homodimerizes upon chitin binding and forms a sandwich-type receptor complex with two OsCEBiP molecules simultaneously binding to one chitin oligosaccharide (57). The second LysM domain (LysM2) of OsCEBiP directly binds to a chitin oligomer (94). Rice LysM-RLK chitin elicitor receptor kinase 1 (OsCERK1), which lacks detectable chitin binding ability, complexes with OsCEBiP and is involved in signaling activation (140). The Arabidopsis genome encodes 5 LysM-RLKs and 3 LysM-RLPs. In contrast, Arabidopsis CERK1 directly binds to chitin and homodimerizes for chitin-induced signaling activation (95). Interestingly, Arabidopsis LysM-RLK LYK5 without a detectable kinase activity possesses a significantly higher chitin-binding affinity than CERK1. Additionally, it heterodimerizes with CERK1 and is required for chitin-induced CERK1 phosphorylation, suggesting that LYK5 is a major chitin receptor in Arabidopsis (Figure 3b) (20). LysM-RLPs appear to not be involved in chitin-induced signaling in Arabidopsis, implicating that distinct chitin perception and signaling systems exist in different plant species.
Although lacking an apparent role in chitin-induced signaling, Arabidopsis LysM-RLPs LYM1 and LYM3 directly bind to PGNs and are required for PGN-mediated responses (160). Similarly, the rice LysM-RLPs OsLYP4 and OsLYP6 physically bind to PGN and chitin and mediate PGN-and chitin-triggered responses (92). Interestingly, CERK1 is also required in PGN-triggered signaling in Arabidopsis and rice, yet direct binding is not evident (92, 160). It is plausible that CERK1 complexes with LYM1 and LYM3 to mediate PGN-induced signaling. Apparently, CERK1 plays dual roles in chitin- and PGN-induced signaling in both monocots and dicots.
Lectin Domain–Containing and Epidermal Growth Factor Domain–Containing Receptor-Like Kinases
The Arabidopsis bulb-type lectin S-domain RLK lipooligosaccharide-specific reduced elicitation (LORE) mediates the sensitivity to the lipid A moiety of LPS from Pseudomonas and Xanthomonas (128). LPS-LORE recognition seems to be restricted to Brassicaceae. Transient expression of LORE in tobacco gains sensitivity to LPS, supporting the idea that LORE is the LPS receptor. The Arabidopsis legume-type lectin domain-containing RLK does not respond to nucleotides 1 (DORN1), binds to ATP and possibly other purine nucleotides, and is a receptor of eATP (26). In addition, Arabidopsis EGF motif-containing wall-associated kinase 1 (WAK1) was suggested to function as the receptor of DAMP OG (18). It is likely that other types of RLKs or RLPs with distinct ectodomains function as receptors of other MAMPs and DAMPs.
Pattern Recognition Receptor Complex Formation and Regulations
Ligand-induced dimerization of PRRs and their cognate coreceptors is a common and immediate event in initiating intracellular signaling (Figure 2). BRI1-associated receptor kinase 1 (BAK1) and other family members of somatic embryogenesis receptor-like kinases (SERKs) function as coreceptors of multiple LRR-RLKs involved in plant immunity and development (88, 103). BAK1 interacts with and acts as a coreceptor of FLS2, EFR, and PEPR1 (Figure 3a). The LRR-containing ectodomain of BAK1 directly contacts the C-terminus of FLS2-bound flg22, and flg22 appears to stabilize the FLS2-BAK1 complex as a molecular glue (149). Tomato FLS3 also complexes with BAK1 upon flgII-28 perception (61). In addition, upon the cognate ligand perception, BAK1 is recruited into multiple LRR-RLP complexes, including tomato Cfs, Arabidopsis RLP23, and tobacco NbCSPR (3, 123, 134). However, BAK1 does not seem to be involved in the signaling pathway mediated by LysM- or lectin-domain containing RLKs, including LYK5, LYM1/LYM3, and LORE, which perceive chitin, PGN, and LPS, respectively (128). Suppressor of BIR1 (SOBIR1), another LRR-RLK with a short LRR ectodomain, constitutively complexes with various LRR-RLPs, including tomato Ve1 and Cfs and Arabidopsis RLP23 but not LRRRLKs, in a ligand-independent manner (3, 87). The aforementioned CERK1 heterodimerizes with LYK5 or OsCEBiP in chitin-triggered signaling (Figure 3b) and possibly with LYM1 and LYM3 in PGN-triggered signaling (20, 140, 160). It is possible that CERK1 functions as a shared coreceptor or signaling partner for LysM-domain containing PRRs.
PRR complexes associate with different regulatory proteins that orchestrate the timing, duration, and intensity of defense responses (Figure 3a). The pseudokinase BAK1-interacting receptor-like kinase 2 (BIR2) associates with BAK1 and likely prevents the FLS2-BAK1 interaction in the resting state (55). Perception of flg22 by FLS2 leads to the release of BIR2 from BAK1, enabling the formation of FLS2-BAK1 complex (55). Specific subunits of protein serine/threonine phosphatase type 2A (PP2A) associate with BAK1 and negatively regulate the BAK1 basal phosphorylation (137). The phosphatase activity of PP2A is attenuated upon elf18 perception, thereby resulting in enhanced BAK1 kinase activity. Rice protein phosphatase 2C (PP2C) XB15 negatively regulates the XA21-mediated immune responses by dephosphorylating XA21 (118). However, rice ATPase XB24 promotes XA21 autophosphorylation via its ATPase activity but inhibits XA21-mediated resistance (21). The Arabidopsis E3 ubiquitin ligases PUB12 and PUB13 are recruited to FLS2 in a BAK1-dependent manner and directly ubiquitinate FLS2 for FLS2 degradation, leading to the attenuation of FLS2 signaling (99, 175). Ubiquitination of membrane receptors also leads to receptor endocytosis and lysosome/vacuole targeting. The ligand-activated PRRs, such as FLS2 and PEPR1, undergo clathrin-dependent internalization and enter an endocytic trafficking route from PM to vacuoles, which is likely required for the full activation of defense responses (109, 117). It remains an open question whether any E3 ligase–mediated ubiquitination is involved in PRR endocytosis.
EARLY PATTERN RECOGNITION RECEPTOR SIGNALING EVENTS
In contrast to the high recognition specificity of PRRs to the cognate MAMPs or DAMPs, the downstream signaling events triggered by PRRs converge at multiple modules, including the activation of receptor-like cytoplasmic kinases (RLCKs), G proteins, MAPK cascades, and calcium-dependent protein kinases (CDPKs/CPKs) (Figures 1 and 3). Apparently, the transcriptional reprogramming triggered by different MAMPs or DAMPs also largely overlaps (82).
Receptor-Like Cytoplasmic Kinases Complex with Pattern Recognition Receptors to Relay Pattern-Triggered Immunity Signaling
RLCKs belong to the same superfamily as RLKs but lack extracellular and transmembrane domains (143). Tomato ACIK1, which is induced upon Cf9 activation, is among the first RLCKs to be shown to have a role in disease resistance (133). Recent evidence indicates that RLCKs associate with PRR complexes to relay diverse intracellular signaling (Figure 3). The Arabidopsis RLCK, Botrytis-induced kinase 1 (BIK1), together with its close family member PBL1, interacts with multiple PRRs, including FLS2, EFR, PEPR1, and coreceptor BAK1 (98, 100, 168). BAK1 directly phosphorylates BIK1 at multiple serine, threonine, and tyrosine residues (89). Another RLCK, BR-signaling kinase 1 (BSK1), also associates with FLS2 and positively regulates certain PTI responses (139). In addition, PTI-compromised RLCK 1 (PCRK1) and PCRK2 associate with FLS2, and possibly other PRRs, and positively regulate multiple PTI responses, disease resistance, and SA biosynthesis (74, 146). Rice OsRLCK176 and OsRLCK185 interact with OsCERK1 and regulate both chitin- and PGN-mediated MAPK activation and defense gene induction (163). PBL27, the Arabidopsis ortholog of OsRLCK185, is required for chitin-triggered responses via direct interaction with and phosphorylation by CERK1 in Arabidopsis (Figure 3b) (142). Interestingly, CERK1 preferentially phosphorylates PBL27, whereas BAK1 preferentially phosphorylates BIK1. It is likely that different RLCKs transduce signaling from specific PRRs via distinct phosphorylation by different coreceptors or receptors.
RLCK-mediated phosphorylation of different substrates appears to further relay PTI signaling to trigger different downstream events. BIK1 directly interacts with and phosphorylates RBOHD to positively regulate flg22- or elf18-triggered ROS bursts (68, 85). Interestingly, another BIK1 family member PBL13 also interacts with RBOHD but negatively regulates PTI responses (90). It is possible that distinct phosphorylation by BIK1 and PBL13 results in opposing activities of RBOHD or that the different interacting partners of BIK1 and PBL13 contribute to the differential regulations of RBOHD. Although loss-of-function of BIK1 and PBL1 does not significantly compromise flg22- or elf18-triggered MAPK activation, CERK1-activated PBL27 directly phosphorylates and activates a MAPK cascade consisting of MAPKKK5-MKK4/5-MPK3/6 in chitin-triggered signaling in Arabidopsis (162). Additional RLCK targets are likely involved in transducing signals into diverse intracellular outputs. RLCKs are also subjected to layered regulations. A phosphatase PP2C38 negatively regulates PTI responses by modulating BIK1 phosphorylation and activity toward RBOHD (28). BIK1 is phosphorylated by CPK28, which likely leads to 26S proteasome-dependent BIK1 turnover at the resting state (114). In addition, a heterotrimeric G-protein complex regulates BIK1 protein stability using an elusive mechanism (86). The trigger and mechanisms for RLCKs turnover require further investigations.
G Proteins Relay Signaling from Pattern Recognition Receptor Complexes
Despite the lack of G protein–coupled receptors (GPCRs) found in animals, plants do possess both small G proteins and heterotrimeric G-protein complexes that play critical roles in PTI signaling via direct association with PRR complexes and downstream components. Chitin treatment activates rice small GTPase OsRac1, which interacts with the guanine nucleotide exchange factor OsRacGEF1 (2). Chitin-activated OsCERK1 directly phosphorylates and activates OsRacGEF1. Both OsRac1 and OsRacGEF1 positively regulate chitin-triggered defense responses and resistance against the rice blast fungus M. oryzae. In addition, OsRac1 interacts with the N-terminus of NADPH oxidase to regulate ROS bursts (Figure 3b) (161). Notably, OsRacGEF1 also interacts with OsFLS2, suggesting its broad involvement in different PRR complexes.
The canonical heterotrimeric G-protein complex includes one Gα (GPA1), one Gβ (AGB1), and three Gγ (AGG1, AGG2, and AGG3) in Arabidopsis. AGB1 connects a MAPK cascade activated by the serine protease PrpL secreted from P. aeruginosa or ArgC from X. campestris via the scaffold protein RACK1 (22). Although RACK1 and Gα do not appear to be involved in flg22-or elf18-triggered signaling, Gβ and Gγ are required for flg22- or elf18-induced ROS bursts and defense gene expression but not for MAPK activation (93). The Arabidopsis genome also contains three Gα-like extra-large G proteins (XLG1, XLG2, and XLG3), of which XLG2 is important in pathogen resistance and flg22- or elf18-triggered ROS bursts (108). Apparently, XLG2-Gβ-Gγ forms a noncanonical heterotrimeric G-protein complex to transduce flg22- or elf18-triggered signaling independent of MAPK activation. In addition, XLG2 interacts with FLS2 and BIK1 and attenuates BIK1 degradation for potentiating immune activation. Flg22 perception leads to BIK1-mediated phosphorylation of XLG2, which is subsequently released from Gβ to activate downstream proteins, such as RBOHD (86). Furthermore, BAK1 directly phosphorylates regulator of G-protein signaling 1 (RGS1), which keeps the G-protein complex at the resting state (156). Therefore, heterotrimeric G-protein complex emerges as an additional component downstream of PRR complexes to transduce PTI signaling.
Mitogen-Activated Protein Kinases and Calcium-Dependent Protein Kinases Are Convergent Components Downstream of Pattern Recognition Receptor Complexes
Rapid and transient activation of MAPKs is a hallmark of PTI responses (Figure 2) (6, 111). A MAPK cascade typically contains three sequentially activated kinases of a MAPK kinase kinase (MAPKKK or MEKK), a MAPK kinase (MAPKK or MKK), and a MAPK (MPK). At least two canonical MAPK cascades consisting of MEKK1/MEKK-MKK4/MKK5-MPK3/MPK6 and MEKK1-MKK1/MKK2-MPK4 in Arabidopsis have been indicated to exert opposing roles in plant defense, with the MPK3/MPK6 cascade having a positive role and the MPK4 cascade having a negative role (6, 73, 111). Chitin-activated CERK1-PBL27 phosphorylation relay activates another MAPK cascade consisting of MAPKKK5-MKK4/MKK5-MPK3/MPK6 (162). Interestingly, in contrast to the compromised MPK3/MPK6 activation triggered by chitin, the mapkkk5 mutant shows elevated MPK3/MPK6 activation stimulated by flg22, suggesting that MAPKKK5 inversely regulates chitin- and flg22-triggered signaling (162). In contrast, the MAPKKK MKKK7, associated with FLS2 and phosphorylated upon flg22 treatment, negatively regulates flg22-triggered MPK6 activation and the ROS burst through unknown mechanisms (113).
In parallel, activation of CDPKs, which are unique Ca2+ sensor protein kinases, has been observed upon MAMP perception (15). Arabidopsis CPK4, CPK5, CPK6, and CPK11 are transiently activated in response to flg22 treatment and redundantly regulate expression of a subset of MAMP-responsive genes distinct from or overlapping with those controlled by MAPKs (15). In addition, CPK5 positively regulates flg22-induced ROS bursts via direct phosphorylation of RBOHD at specific residues different from those phosphorylated by BIK1 (35, 68). In contrast, CPK28 negatively regulates PTI signaling by modulating the BIK1 protein level through phosphorylation (114). The mechanism by which CDPKs regulate MAMP-responsive gene expression remains poorly understood.
Dynamic phosphorylation of different substrates by MAPKs and CDPKs further bifurcates PTI signaling. For instance, MPK3 and MPK6 phosphorylate and activate 1-aminocyclopropane-1-carboxylic acid synthase 2 (ACS2) and ACS6, the rate-limiting enzymes in ET biosynthesis, to regulate flg22-triggered ET production (97). In addition, flg22-activated MPK3 and MPK6 phosphorylate tandem zinc finger protein 9 (TZF9), residing in cytoplasmic processing (P) bodies, the sites for mRNA storage and decay, for full PTI responses (107). Flg22-activated MPK4 phosphorylates PAT1, a key component in mRNA decay, leading to its accumulation in P-bodies and contributing to posttranscriptional regulation of gene expression (132). MAPKs also regulate the activation of specific transcription factors in PTI signaling (see below). Compared to MAPKs, the CDPK substrates controlling defense gene expression are less understood.
Transcriptional Reprograming Leads to Pattern-Triggered Immunity–Associated Gene Expression
PTI elicitation induces rapid, dynamic, and global transcriptional changes of plant genes involved in a broad range of biological functions (82). Specific transcription factors directly phosphorylated by MAPKs or acting downstream of Ca2+ signaling have been shown to play a role in regulating MAMP-responsive gene expression. The Arabidopsis transcription factors ERF104 and BES1 phosphorylated by MPK6 play positive roles in MAMP-responsive gene expression (10, 70), whereas the trihelix transcription factor ASR3 phosphorylated by MPK4 negatively regulates expression of a subset of MAMP-responsive genes (81). In other cases, transcription factors are not the direct targets of MAPKs, but instead their activation is monitored by MAPK phosphorylation substrates. MPK3 and MPK6 phosphorylate a subset of MPK3/MPK6-targeted VQ-motif-containing proteins (MVQs), each of which interacts with a specific subclass of WRKYs via the VQ motif (119). Phosphorylation by MPK3/MPK6 destabilizes MVQ1 and subsequently activates WRKYs (119). The transcription factors CaM-binding protein 60g (CBP60g) and its homolog SARD1 bind to the promoter of the SA biosynthetic gene ICS1 and regulate MAMP-induced SA accumulation (159, 173). Interestingly, CBP60g and SARD1 directly bind to the promoters of genes encoding PRR complexes or signaling components, including BAK1, BIK1, and MPK3, and modulate their expression (147).
Specific modulation of the general transcription machinery also plays an essential role in warranting rapid activation of MAMP-responsive genes. MAMP treatments induce rapid and transient phosphorylation of RNA polymerase II (RNAPII) C-terminal domain (CTD) via cyclin-dependent kinase Cs (CDKCs), which are phosphorylated and activated by MPK3 and MPK6(83). CTD phosphatase-like 3 (CPL3) dephosphorylates MAMP-activated CTD phosphorylation and negatively regulates expression of a large portion of MAMP-responsive genes (83). The Mediator complex is also implicated in plant defense, yet its precise connection with PTI signaling is not established (82). Using a forward genetic screen based on flg22-induced gene transcriptional change, protein poly(ADP-ribosyl)ation (PARylation), which is reversibly regulated by poly(ADP-ribose) polymerases (PARPs) and poly(ADP-ribose) glycohydrolases (PARGs), was indicated to play a role in regulating MAMP-responsive gene expression (41). flg22 treatment induces the PARylation of the forkhead-associated domain protein DAWDLE, a substrate of Arabidopsis PARP2, to regulate some of late PTI responses (42). It is postulated that PARylation of DDL may induce chromatin remodeling that creates a permissive chromatin environment for RNAPII-mediated transcription of MAMP-responsive genes. Thus, the coordinated actions of the general transcriptional machinery, gene-specific transcription factors, and chromatin remodeling contribute to the rapid and accurate transcriptional reprogramming of MAMP-responsive genes.
CONCLUSIONS AND PERSPECTIVE
Tremendous progress has been achieved over the past decades in the identification of microbial patterns, characterization of pattern-triggered plant responses, and elucidation of plant PRR receptors and signaling networks. Despite differences in certain details, a common theme is that conserved microbial patterns are perceived by plant PRRs and trigger a series of physiological and cellular responses and transcriptional reprogramming, collectively contributing to plant defense against a broad spectrum of pathogens. Activation of PRRs, which are often cell surface–resident RLKs or RLPs, is regulated by dynamic association with coreceptors and various regulatory components as well as by the posttranslational modifications of the receptor complexes. The activated receptor complexes further relay the signaling to different intracellular modules, including RLCKs, G proteins, MAPKs, and CDPKs, leading to complex and intertwined physiological and cellular responses and transcriptional reprogramming to mount effective PTI.
Perception of MAMPs or DAMPs by PRRs instantaneously emanates layered and intertwined molecular, cellular, and physiological responses. Although the relationship and epistasis of these responses remain ambiguous in most cases, increase of [Ca2+]cyt appears to be a prerequisite for other responses to occur, such as the ROS burst and PM depolarization. Considering the difference between cytosolic and extracellular Ca2+ concentrations in plant cells, a rapid [Ca2+]cyt burst likely ensures the sensitivity of defense elicitation. The mechanism by which [Ca2+]cyt increase is sensed to exert multifaceted cellular functions remains elusive. A ROS burst is a robust and common response upon MAMP perception, but it is a potential double-edged sword for plants. Not surprisingly, NADPH oxidases, the key enzymes mediating a ROS burst, are regulated at multiple levels, including phosphorylation by BIK1 and CDPKs, G proteins, and signaling molecules, such as PA, with both positive and negative effects. It will be interesting to dissect how NADPH oxidase activity is orchestrated by these components upon MAMP perception. MAPK activation is another robust and universal PTI response. In chitin-triggered signaling, MAPK cascades are directly activated by the upstream RLCK in the PRR complex (162). How MAPKs are activated remains enigmatic in flg22 and elf18 signaling. Different PRRs perceive distinct MAMPs/DAMPs but recruit the same group of coreceptors, such as BAK1. Notably, BAK1 and related RLKs are also coreceptors of LRR-RLKs involved in plant growth and cell differentiation and elongation (88, 103). How these coreceptors function as shared signaling modules yet maintain a high degree of signaling specificity in different biological processes remains an open question.
ACKNOWLEDGMENTS
We apologize to colleagues whose work was not discussed here because of space limitations. We thank Dr. Mike Kolomiets, Dr. Jen Sheen, Mr. Kevin Cox, and Mr. Pierce Jamieson for critical reading and editing of the manuscript. The work was supported by National Science Foundation (IOS-1252539) to P.H., and National Institutes of Health (R01GM097247) and the Robert A. Welch foundation (A-1795) to L.S.
Footnotes
DISCLOSURE STATEMENT
The authors are not aware of any affiliations, memberships, funding, or financial holdings that might be perceived as affecting the objectivity of this review.
LITERATURE CITED
- 1.Ahuja I, Kissen R, Bones AM. 2012. Phytoalexins in defense against pathogens. Trends Plant Sci 17:73–90 [DOI] [PubMed] [Google Scholar]
- 2.Akamatsu A, Wong HL, Fujiwara M, Okuda J, Nishide K, et al. 2013. An OsCEBiP/OsCERK1-OsRacGEF1-OsRac1 module is an essential early component of chitin-induced rice immunity. Cell Host Microbe 13:465–76 [DOI] [PubMed] [Google Scholar]
- 3.Albert I, Bohm H, Albert M, Feiler CE, Imkampe J, et al. 2015. An RLP23-SOBIR1-BAK1 complex mediates NLP-triggered immunity. Nat. Plants 1:15140. [DOI] [PubMed] [Google Scholar]
- 4.Ali R, Ma W, Lemtiri-Chlieh F, Tsaltas D, Leng Q, et al. 2007. Death don’t have no mercy and neither does calcium: Arabidopsis CYCLIC NUCLEOTIDE GATED CHANNEL2 and innate immunity. Plant Cell 19:1081–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Arnaud D, Hwang I. 2014. A sophisticated network of signaling pathways regulates stomatal defenses to bacterial pathogens. Mol. Plant 8:566–81 [DOI] [PubMed] [Google Scholar]
- 6.Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL, et al. 2002. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415:977–83 [DOI] [PubMed] [Google Scholar]
- 7.Ausubel FM. 2005. Are innate immune signaling pathways in plants and animals conserved? Nat. Immunol 6:973–79 [DOI] [PubMed] [Google Scholar]
- 8.Avni A, Bailey BA, Mattoo AK, Anderson JD. 1994. Induction of ethylene biosynthesis in Nicotiana tabacum by a Trichoderma viride xylanase is correlated to the accumulation of 1-aminocyclopropane-1-carboxylic acid (Acc) synthase and Acc oxidase transcripts. Plant Physiol 106:1049–55 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bent AF, Mackey D. 2007. Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu. Rev. Phytopathol 45:399–436 [DOI] [PubMed] [Google Scholar]
- 10.Bethke G, Unthan T, Uhrig JF, Poschl Y, Gust AA, et al. 2009. Flg22 regulates the release of an ethylene response factor substrate from MAP kinase 6 in Arabidopsis thaliana via ethylene signaling. PNAS 106:8067–72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blein JP, Milat ML, Ricci P. 1991. Responses of cultured tobacco cells to cryptogein, a proteinaceous elicitor from Phytophthora cryptogea: possible plasmalemma involvement. Plant Physiol 95:486–91 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bohm H, Albert I, Fan L, Reinhard A, Nurnberger T. 2014. Immune receptor complexes at the plant cell surface. Curr. Opin. Plant Biol 20:47–54 [DOI] [PubMed] [Google Scholar]
- 13.Bohm H, Albert I, Oome S, Raaymakers TM, Van den Ackerveken G, Nurnberger T. 2014. A conserved peptide pattern from a widespread microbial virulence factor triggers pattern-induced immunity in Arabidopsis. PLOS Pathog 10:e1004491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Boller T, Felix G. 2009. A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol 60:379–406 [DOI] [PubMed] [Google Scholar]
- 15.Boudsocq M, Willmann MR, McCormack M, Lee H, Shan LB, et al. 2010. Differential innate immune signalling via Ca2+ sensor protein kinases. Nature 464:418–22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Brauer EK, Ahsan N, Dale R, Kato N, Coluccio AE, et al. 2016. The Raf-like kinase ILK1 and the high affinity K+ transporter HAK5 are required for innate immunity and abiotic stress response. Plant Physiol 171:1470–84 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brunner F, Rosahl S, Lee J, Rudd JJ, Geiler C, et al. 2002. Pep-13, a plant defense-inducing pathogen-associated pattern from Phytophthora transglutaminases. EMBO J 21:6681–88 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Brutus A, Sicilia F, Macone A, Cervone F, De Lorenzo G. 2010. A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. PNAS 107:9452–57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cai R, Lewis J, Yan S, Liu H, Clarke CR, et al. 2011. The plant pathogen Pseudomonas syringae pv. tomato is genetically monomorphic and under strong selection to evade tomato immunity. PLOS Pathog 7:e1002130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cao YR, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, et al. 2014. The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. eLife 3:e03766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chen X, Chern M, Canlas PE, Ruan D, Jiang C, Ronald PC. 2010. An ATPase promotes autophosphorylation of the pattern recognition receptor XA21 and inhibits XA21-mediated immunity. PNAS 107:8029–34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cheng Z, Li JF, Niu Y, Zhang XC, Woody OZ, et al. 2015. Pathogen-secreted proteases activate a novel plant immune pathway. Nature 521:213–16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nurnberger T, et al. 2007. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448:497–500 [DOI] [PubMed] [Google Scholar]
- 24.Cho MH, Lee SW. 2015. Phenolic phytoalexins in rice: biological functions and biosynthesis. Int. J. Mol. Sci 16:29120–33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Choi HW, Manohar M, Manosalva P, Tian M, Moreau M, Klessig DF. 2016. Activation of plant innate immunity by extracellular high mobility group box 3 and its inhibition by salicylic acid. PLOS Pathog 12:e1005518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Choi J, Tanaka K, Cao YR, Qi Y, Qiu J, et al. 2014. Identification of a plant receptor for extracellular ATP. Science 343:290–94 [DOI] [PubMed] [Google Scholar]
- 27.Clay NK, Adio AM, Denoux C, Jander G, Ausubel FM. 2009. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323:95–101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Couto D, Niebergall R, Liang X, Bucherl CA, Sklenar J, et al. 2016. The Arabidopsis protein phosphatase PP2C38 negatively regulates the central immune kinase BIK1. PLOS Pathog 12:e1005811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Couto D, Zipfel C. 2016. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol 16537–52 [DOI] [PubMed] [Google Scholar]
- 30.Cui W, Lee JY. 2016. Arabidopsis callose synthases CalS1/8 regulate plasmodesmal permeability during stress. Nat. Plants 2:16034. [DOI] [PubMed] [Google Scholar]
- 31.Daudi A, Cheng Z, O’Brien JA, Mammarella N, Khan S, et al. 2012. The apoplastic oxidative burst peroxidase in Arabidopsis is a major component of pattern-triggered immunity. Plant Cell 24:275–87 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Day B, Henty JL, Porter KJ, Staiger CJ. 2011. The pathogen-actin connection: a platform for defense signaling in plants. Annu. Rev. Phytopathol 49:483–506 [DOI] [PubMed] [Google Scholar]
- 33.de Jonge R, van Esse HP, Maruthachalam K, Bolton MD, Santhanam P, et al. 2012. Tomato immune receptor Ve1 recognizes effector of multiple fungal pathogens uncovered by genome and RNA sequencing. PNAS 109:5110–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Du J, Verzaux E, Chaparro-Garcia A, Bijsterbosch G, Keizer LC, et al. 2015. Elicitin recognition confers enhanced resistance to Phytophthora infestans in potato. Nat. Plants 1:15034. [DOI] [PubMed] [Google Scholar]
- 35.Dubiella U, Seybold H, Durian G, Komander E, Lassig R, et al. 2013. Calcium-dependent protein kinase/NADPH oxidase activation circuit is required for rapid defense signal propagation. PNAS 110:8744–49 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Elmore JM, Coaker G. 2011. The role of the plasma membrane H+-ATPase in plant-microbe interactions. Mol. Plant 4:416–27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Erbs G, Silipo A, Aslam S, De Castro C, Liparoti V, et al. 2008. Peptidoglycan and muropeptides from pathogens Agrobacterium and Xanthomonas elicit plant innate immunity: structure and activity. Chem. Biol 15:438–48 [DOI] [PubMed] [Google Scholar]
- 38.Faulkner C, Petutschnig E, Benitez-Alfonso Y, Beck M, Robatzek S, et al. 2013. LYM2-dependent chitin perception limits molecular flux via plasmodesmata. PNAS 110:9166–70 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Felix G, Boller T. 2003. Molecular sensing of bacteria in plants. The highly conserved RNA-binding motif RNP-1 of bacterial cold shock proteins is recognized as an elicitor signal in tobacco. J. Biol. Chem 278:6201–8 [DOI] [PubMed] [Google Scholar]
- 40.Felix G, Duran JD, Volko S, Boller T. 1999. Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J 18:265–76 [DOI] [PubMed] [Google Scholar]
- 41.Feng B, Liu C, de Oliveira MV, Intorne AC, Li B, et al. 2015. Protein poly(ADP-ribosyl)ation regulates Arabidopsis immune gene expression and defense responses. PLOS Genet 11:e1004936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Feng B, Ma S, Chen S, Zhu N, Zhang S, et al. 2016. PARylation of the forkhead-associated domain protein DAWDLE regulates plant immunity. EMBO Rep 17:1799–813 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ferrari S, Galletti R, Denoux C, De Lorenzo G, Ausubel FM, Dewdney J. 2007. Resistance to Botrytis cinerea induced in Arabidopsis by elicitors is independent of salicylic acid, ethylene, or jasmonate signaling but requires PHYTOALEXIN DEFICIENT3. Plant Physiol 144:367–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Fliegmann J, Mithofer A, Wanner G, Ebel J. 2004. An ancient enzyme domain hidden in the putative β-glucan elicitor receptor of soybean may play an active part in the perception of pathogen-associated molecular patterns during broad host resistance. J. Biol. Chem 279:1132–40 [DOI] [PubMed] [Google Scholar]
- 45.Foresi NP, Laxalt AM, Tonon CV, Casalongue CA, Lamattina L. 2007. Extracellular ATP induces nitric oxide production in tomato cell suspensions. Plant Physiol 145:589–92 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Frei dit Frey N, Mbengue M, Kwaaitaal M, Nitsch L, Altenbach D, et al. 2012. Plasma membrane calcium ATPases are important components of receptor-mediated signaling in plant immune responses and development. Plant Physiol 159:798–809 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Frias M, Gonzalez C, Brito N. 2011. BcSpl1, a cerato-platanin family protein, contributes to Botrytis cinerea virulence and elicits the hypersensitive response in the host. New Phytol 192:483–95 [DOI] [PubMed] [Google Scholar]
- 48.Furukawa T, Inagaki H, Takai R, Hirai H, Che FS. 2014. Two distinct EF-Tu epitopes induce immune responses in rice and Arabidopsis. Mol. Plant-Microbe Interact 27:113–24 [DOI] [PubMed] [Google Scholar]
- 49.Galletti R, Denoux C, Gambetta S, Dewdney J, Ausubel FM, et al. 2008. The AtrbohD-mediated oxidative burst elicited by oligogalacturonides in Arabidopsis is dispensable for the activation of defense responses effective against Botrytis cinerea. Plant Physiol 148:1695–706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Glazebrook J. 2005. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol 43:205–27 [DOI] [PubMed] [Google Scholar]
- 51.Gomez-Gomez L, Boller T. 2000. FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol. Cell 5:1003–11 [DOI] [PubMed] [Google Scholar]
- 52.Gust AA, Biswas R, Lenz HD, Rauhut T, Ranf S, et al. 2007. Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J. Biol. Chem 282:32338–48 [DOI] [PubMed] [Google Scholar]
- 53.Guzel Deger A, Scherzer S, Nuhkat M, Kedzierska J, Kollist H, et al. 2015. Guard cell SLAC1-type anion channels mediate flagellin-induced stomatal closure. New Phytol 208:162–73 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Halim VA, Altmann S, Ellinger D, Eschen-Lippold L, Miersch O, et al. 2009. PAMP-induced defense responses in potato require both salicylic acid and jasmonic acid. Plant J 57:230–42 [DOI] [PubMed] [Google Scholar]
- 55.Halter T, Imkampe J, Mazzotta S, Wierzba M, Postel S, et al. 2014. The leucine-rich repeat receptor kinase BIR2 is a negative regulator of BAK1 in plant immunity. Curr. Biol 24:134–43 [DOI] [PubMed] [Google Scholar]
- 56.Hamada H, Kurusu T, Okuma E, Nokajima H, Kiyoduka M, et al. 2012. Regulation of a proteinaceous elicitor-induced Ca2+ influx and production of phytoalexins by a putative voltage-gated cation channel, OsTPC1, in cultured rice cells. J. Biol. Chem 287:9931–39 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hayafune M, Berisio R, Marchetti R, Silipo A, Kayama M, et al. 2014. Chitin-induced activation of immune signaling by the rice receptor CEBiP relies on a unique sandwich-type dimerization. PNAS 111:E404–13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hegenauer V, Furst U, Kaiser B, Smoker M, Zipfel C, et al. 2016. Detection of the plant parasite Cuscuta reflexa by a tomato cell surface receptor. Science 353:478–81 [DOI] [PubMed] [Google Scholar]
- 59.Henty-Ridilla JL, Li J, Day B, Staiger CJ. 2014. ACTIN DEPOLYMERIZING FACTOR4 regulates actin dynamics during innate immune signaling in Arabidopsis. Plant Cell 26:340–52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Henty-Ridilla JL, Shimono M, Li J, Chang JH, Day B, Staiger CJ. 2013. The plant actin cytoskeleton responds to signals from microbe-associated molecular patterns. PLOS Pathog 9:e1003290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Hind SR, Strickler SR, Boyle PC, Dunham DM, Bao Z, et al. 2016. Tomato receptor FLAGELLIN-SENSING 3 binds flgII-28 and activates the plant immune system. Nat. Plants 2:16128. [DOI] [PubMed] [Google Scholar]
- 62.Hou S, Wang X, Chen D, Yang X, Wang M, et al. 2014. The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLOS Pathog 10:e1004331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Huffaker A, Pearce G, Ryan CA. 2006. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. PNAS 103:10098–103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Jabs T, Tschope M, Colling C, Hahlbrock K, Scheel D. 1997. Elicitor-stimulated ion fluxes and O2− from the oxidative burst are essential components in triggering defense gene activation and phytoalexin synthesis in parsley. PNAS 94:4800–5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Jehle AK, Lipschis M, Albert M, Fallahzadeh-Mamaghani V, Furst U, et al. 2013. The receptor-like protein ReMAX of Arabidopsis detects the microbe-associated molecular pattern eMax from Xanthomonas. Plant Cell 25:2330–40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Jeworutzki E, Roelfsema MR, Anschutz U, Krol E, Elzenga JT, et al. 2010. Early signaling through the Arabidopsis pattern recognition receptors FLS2 and EFR involves Ca-associated opening of plasma membrane anion channels. Plant J 62:367–78 [DOI] [PubMed] [Google Scholar]
- 67.Jones JD, Dangl JL. 2006. The plant immune system. Nature 444:323–29 [DOI] [PubMed] [Google Scholar]
- 68.Kadota Y, Sklenar J, Derbyshire P, Stransfeld L, Asai S, et al. 2014. Direct regulation of the NADPH oxidase RBOHD by the PRR-associated kinase BIK1 during plant immunity. Mol. Cell 54:43–55 [DOI] [PubMed] [Google Scholar]
- 69.Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, et al. 2006. Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. PNAS 103:11086–91 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Kang S, Yang F, Li L, Chen H, Chen S, Zhang J. 2015. The Arabidopsis transcription factor BRASSINOSTEROID INSENSITIVE1-ETHYL METHANESULFONATE-SUPPRESSOR1 is a direct substrate of MITOGEN-ACTIVATED PROTEIN KINASE6 and regulates immunity. Plant Physiol 167:1076–86 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Khatib M, Lafitte C, Esquerre-Tugaye M-T, Bottin A, Rickauer M. 2004. The CBEL elicitor of Phytophthora parasitica var. nicotianae activates defence in Arabidopsis thaliana via three different signalling pathways. New Phytol 162:501–10 [Google Scholar]
- 72.Klemptner RL, Sherwood JS, Tugizimana F, Dubery IA, Piater LA. 2014. Ergosterol, an orphan fungal microbe-associated molecular pattern (MAMP). Mol. Plant Pathol 15:747–61 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Kong Q, Qu N, Gao M, Zhang Z, Ding X, et al. 2012. The MEKK1-MKK1/MKK2-MPK4 kinase cascade negatively regulates immunity mediated by a mitogen-activated protein kinase kinase kinase in Arabidopsis. Plant Cell 24:2225–36 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kong Q, Sun T, Qu N, Ma J, Li M, et al. 2016. Two redundant receptor-like cytoplasmic kinases function downstream of pattern recognition receptors to regulate activation of SA biosynthesis. Plant Physiol 171:1344–54 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Krol E, Mentzel T, Chinchilla D, Boller T, Felix G, et al. 2010. Perception of the Arabidopsis danger signal peptide 1 involves the pattern recognition receptor AtPEPR1 and its close homologue AtPEPR2. J. Biol. Chem 285:13471–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Kunze G, Zipfel C, Robatzek S, Niehaus K, Boller T, Felix G. 2004. The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16:3496–507 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Laxalt AM, Raho N, Have AT, Lamattina L. 2007. Nitric oxide is critical for inducing phosphatidic acid accumulation in xylanase-elicited tomato cells. J. Biol. Chem 282:21160–68 [DOI] [PubMed] [Google Scholar]
- 78.Lecourieux D, Mazars C, Pauly N, Ranjeva R, Pugin A. 2002. Analysis and effects of cytosolic free calcium increases in response to elicitors in Nicotiana plumbaginifolia cells. Plant Cell 14:2627–41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Lee JY, Lu H. 2011. Plasmodesmata: the battleground against intruders. Trends Plant Sci 16:201–10 [DOI] [PubMed] [Google Scholar]
- 80.Lee JY, Wang X, Cui W, Sager R, Modla S, et al. 2011. A plasmodesmata-localized protein mediates crosstalk between cell-to-cell communication and innate immunity in Arabidopsis. Plant Cell 23:3353–73 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Li B, Jiang S, Yu X, Cheng C, Chen S, et al. 2015. Phosphorylation of trihelix transcriptional repressor ASR3 by MAP KINASE4 negatively regulates Arabidopsis immunity. Plant Cell 27:839–56 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Li B, Meng X, Shan L, He P. 2016. Transcriptional regulation of pattern-triggered immunity in plants. Cell Host Microbe 19:641–50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Li F, Cheng C, Cui F, de Oliveira MV, Yu X, et al. 2014. Modulation of RNA polymerase II phosphor-ylation downstream of pathogen perception orchestrates plant immunity. Cell Host Microbe 16:748–58 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Li J, Henty-Ridilla JL, Staiger BH, Day B, Staiger CJ. 2015. Capping protein integrates multiple MAMP signalling pathways to modulate actin dynamics during plant innate immunity. Nat. Commun 6:7206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Li L, Li M, Yu LP, Zhou ZY, Liang XX, et al. 2014. The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe 15:329–38 [DOI] [PubMed] [Google Scholar]
- 86.Liang XX, Ding PT, Liang KH, Wang JL, Ma MM, et al. 2016. Arabidopsis heterotrimeric G proteins regulate immunity by directly coupling to the FLS2 receptor. eLife 5:e13568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Liebrand TWH, van den Berg GCM, Zhang Z, Smit P, Cordewener JHG, et al. 2013. Receptor-like kinase SOBIR1/EVR interacts with receptor-like proteins in plant immunity against fungal infection. PNAS 110:10010–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Liebrand TWH, van den Burg HA, Joosten MHAJ. 2014. Two for all: receptor-associated kinases SOBIR1 and BAK1. Trends Plant Sci 19:123–32 [DOI] [PubMed] [Google Scholar]
- 89.Lin WW, Li B, Lu DP, Chen SX, Zhu N, et al. 2014. Tyrosine phosphorylation of protein kinase complex BAK1/BIK1 mediates Arabidopsis innate immunity. PNAS 111:3632–37 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Lin ZJ, Liebrand TW, Yadeta KA, Coaker G. 2015. PBL13 is a serine/threonine protein kinase that negatively regulates Arabidopsis immune responses. Plant Physiol 169:2950–62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Liu B, Li JF, Ao Y, Li Z, Liu J, et al. 2013. OsLYP4 and OsLYP6 play critical roles in rice defense signal transduction. Plant Signal. Behav 8:e22980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Liu B, Li JF, Ao Y, Qu J, Li Z, et al. 2012. Lysin motif–containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 24:3406–19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Liu J, Ding P, Sun T, Nitta Y, Dong O, et al. 2013. Heterotrimeric G proteins serve as a converging point in plant defense signaling activated by multiple receptor-like kinases. Plant Physiol 161:2146–58 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Liu S, Wang J, Han Z, Gong X, Zhang H, Chai J. 2016. Molecular mechanism for fungal cell wall recognition by rice chitin receptor OsCEBiP. Structure 24:1192–200 [DOI] [PubMed] [Google Scholar]
- 95.Liu T, Liu Z, Song C, Hu Y, Han Z, et al. 2012. Chitin-induced dimerization activates a plant immune receptor. Science 336:1160–64 [DOI] [PubMed] [Google Scholar]
- 96.Liu X, Grabherr HM, Willmann R, Kolb D, Brunner F, et al. 2014. Host-induced bacterial cell wall decomposition mediates pattern-triggered immunity in Arabidopsis. eLife 3:e01990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Liu Y, Zhang S. 2004. Phosphorylation of 1-aminocyclopropane-1-carboxylic acid synthase by MPK6, a stress-responsive mitogen-activated protein kinase, induces ethylene biosynthesis in Arabidopsis. Plant Cell 16:3386–99 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Liu Z, Wu Y, Yang F, Zhang Y, Chen S, et al. 2013. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. PNAS 110:6205–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Lu D, Lin W, Gao X, Wu S, Cheng C, et al. 2011. Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science 332:1439–42 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Lu D, Wu S, Gao X, Zhang Y, Shan L, He P. 2010. A receptor-like cytoplasmic kinase, BIK1, associates with a flagellin receptor complex to initiate plant innate immunity. PNAS 107:496–501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Luna E, Pastor V, Robert J, Flors V, Mauch-Mani B, Ton J. 2011. Callose deposition: a multifaceted plant defense response. Mol. Plant-Microbe Interact 24:183–93 [DOI] [PubMed] [Google Scholar]
- 102.Ma W, Smigel A, Tsai YC, Braam J, Berkowitz GA. 2008. Innate immunity signaling: Cytosolic Ca2+ elevation is linked to downstream nitric oxide generation through the action of calmodulin or a calmodulin-like protein. Plant Physiol 148:818–28 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ma X, Xu G, He P, Shan L. 2016. SERKing coreceptors for receptors. Trends Plant Sci 21:1017–33 [DOI] [PubMed] [Google Scholar]
- 104.Ma Y, Walker RK, Zhao YC, Berkowitz GA. 2012. Linking ligand perception by PEPR pattern recognition receptors to cytosolic Ca2+ elevation and downstream immune signaling in plants. PNAS 109:19852–57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Ma Y, Zhao Y, Walker RK, Berkowitz GA. 2013. Molecular steps in the immune signaling pathway evoked by plant elicitor peptides: Ca2+-dependent protein kinases, nitric oxide, and reactive oxygen species are downstream from the early Ca2+ signal. Plant Physiol 163:1459–71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Ma Z, Song T, Zhu L, Ye W, Wang Y, et al. 2015. A Phytophthora sojae glycoside hydrolase 12 protein is a major virulence factor during soybean infection and is recognized as a PAMP. Plant Cell 27:2057–72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Maldonado-Bonilla LD, Eschen-Lippold L, Gago-Zachert S, Tabassum N, Bauer N, et al. 2014. The Arabidopsis tandem zinc finger 9 protein binds RNA and mediates pathogen-associated molecular pattern-triggered immune responses. Plant Cell Physiol 55:412–25 [DOI] [PubMed] [Google Scholar]
- 108.Maruta N, Trusov Y, Brenya E, Parekh U, Botella JR. 2015. Membrane-localized extra-large G proteins and Gβγ of the heterotrimeric G proteins form functional complexes engaged in plant immunity in Arabidopsis. Plant Physiol 167:1004–16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Mbengue M, Bourdais G, Gervasi F, Beck M, Zhou J, et al. 2016. Clathrin-dependent endocytosis is required for immunity mediated by pattern recognition receptor kinases. PNAS 113:11034–39 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Melotto M, Underwood W, Koczan J, Nomura K, He SY. 2006. Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–80 [DOI] [PubMed] [Google Scholar]
- 111.Meng X, Zhang S. 2013. MAPK cascades in plant disease resistance signaling. Annu. Rev. Phytopathol 51:245–66 [DOI] [PubMed] [Google Scholar]
- 112.Millet YA, Danna CH, Clay NK, Songnuan W, Simon MD, et al. 2010. Innate immune responses activated in Arabidopsis roots by microbe-associated molecular patterns. Plant Cell 22:973–90 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Mithoe SC, Ludwig C, Pel MJ, Cucinotta M, Casartelli A, et al. 2016. Attenuation of pattern recognition receptor signaling is mediated by a MAP kinase kinase kinase. EMBO Rep 17:441–54 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Monaghan J, Matschi S, Shorinola O, Rovenich H, Matei A, et al. 2014. The calcium-dependent protein kinase CPK28 buffers plant immunity and regulates BIK1 turnover. Cell Host Microbe 16:605–15 [DOI] [PubMed] [Google Scholar]
- 115.Nuhse TS, Bottrill AR, Jones AM, Peck SC. 2007. Quantitative phosphoproteomic analysis of plasma membrane proteins reveals regulatory mechanisms of plant innate immune responses. Plant J 51:931–40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Nurnberger T, Brunner F, Kemmerling B, Piater L. 2004. Innate immunity in plants and animals: striking similarities and obvious differences. Immunol. Rev 198:249–66 [DOI] [PubMed] [Google Scholar]
- 117.Ortiz-Morea FA, Savatin DV, Dejonghe W, Kumar R, Luo Y, et al. 2016. Danger-associated peptide signaling in Arabidopsis requires clathrin. PNAS 113:11028–33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Park CJ, Peng Y, Chen XW, Dardick C, Ruan DL, et al. 2008. Rice XB15, a protein phosphatase 2C, negatively regulates cell death and XA21-mediated innate immunity. PLOS Biol 6:1910–26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Pecher P, Eschen-Lippold L, Herklotz S, Kuhle K, Naumann K, et al. 2014. The Arabidopsis thaliana mitogen-activated protein kinases MPK3 and MPK6 target a subclass of “VQ-motif “-containing proteins to regulate immune responses. New Phytol 203:592–606 [DOI] [PubMed] [Google Scholar]
- 120.Pel MJC, van Dijken AJH, Bardoel BW, Seidl MF, van der Ent S, et al. 2014. Pseudomonas syringae evades host immunity by degrading flagellin monomers with alkaline protease AprA. Mol. Plant-Microbe Interact 27:603–10 [DOI] [PubMed] [Google Scholar]
- 121.Pinosa F, Buhot N, Kwaaitaal M, Fahlberg P, Thordal-Christensen H, et al. 2013. Arabidopsis phospholipase dδ is involved in basal defense and nonhost resistance to powdery mildew fungi. Plant Physiol 163:896–906 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Poinssot B, Vandelle E, Bentejac M, Adrian M, Levis C, et al. 2003. The endopolygalacturonase 1 from Botrytis cinerea activates grapevine defense reactions unrelated to its enzymatic activity. Mol. Plant-Microbe Interact 16:553–64 [DOI] [PubMed] [Google Scholar]
- 123.Postma J, Liebrand TW, Bi G, Evrard A, Bye RR, et al. 2016. Avr4 promotes Cf-4 receptor-like protein association with the BAK1/SERK3 receptor-like kinase to initiate receptor endocytosis and plant immunity. New Phytol 210:627–42 [DOI] [PubMed] [Google Scholar]
- 124.Pruitt RN, Schwessinger B, Joe A, Thomas N, Liu F, et al. 2015. The rice immune receptor XA21 recognizes a tyrosine-sulfated protein from a Gram-negative bacterium. Sci. Adv 1:e1500245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Pugin A, Frachisse JM, Tavernier E, Bligny R, Gout E, et al. 1997. Early events induced by the elicitor cryptogein in tobacco cells: involvement of a plasma membrane NADPH oxidase and activation of glycolysis and the pentose phosphate pathway. Plant Cell 9:2077–91 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Rajniak J, Barco B, Clay NK, Sattely ES. 2015. A new cyanogenic metabolite in Arabidopsis required for inducible pathogen defence. Nature 525:376–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Ranf S, Eschen-Lippold L, Pecher P, Lee J, Scheel D. 2011. Interplay between calcium signalling and early signalling elements during defence responses to microbe- or damage-associated molecular patterns. Plant J 68:100–13 [DOI] [PubMed] [Google Scholar]
- 128.Ranf S, Gisch N, Schaffer M, Illig T, Westphal L, et al. 2015. A lectin S-domain receptor kinase mediates lipopolysaccharide sensing in Arabidopsis thaliana. Nat. Immunol 16:426–33 [DOI] [PubMed] [Google Scholar]
- 129.Robinson SM, Bostock RM. 2015. β-Glucans and eicosapolyenoic acids as MAMPs in plant-oomycete interactions: past and present. Front. Plant Sci 5:797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Ron M, Avni A. 2004. The receptor for the fungal elicitor ethylene-inducing xylanase is a member of a resistance-like gene family in tomato. Plant Cell 16:1604–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Rotblat B, Enshell-Seijffers D, Gershoni JM, Schuster S, Avni A. 2002. Identification of an essential component of the elicitation active site of the EIX protein elicitor. Plant J 32:1049–55 [DOI] [PubMed] [Google Scholar]
- 132.Roux ME, Rasmussen MW, Palma K, Lolle S, Regue AM, et al. 2015. The mRNA decay factor PAT1 functions in a pathway including MAP kinase 4 and immune receptor SUMM2. EMBO J 34:593–608 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Rowland O, Ludwig AA, Merrick CJ, Baillieul F, Tracy FE, et al. 2005. Functional analysis of Avr9/Cf-9 rapidly elicited genes identifies a protein kinase, ACIK1, that is essential for full Cf-9-dependent disease resistance in tomato. Plant Cell 17:295–310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Saur IM, Kadota Y, Sklenar J, Holton NJ, Smakowska E, et al. 2016. NbCSPR underlies age-dependent immune responses to bacterial cold shock protein in Nicotiana benthamiana. PNAS 113:3389–94 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Scheler C, Durner J, Astier J. 2013. Nitric oxide and reactive oxygen species in plant biotic interactions. Curr. Opin. Plant Biol 16:534–39 [DOI] [PubMed] [Google Scholar]
- 136.Schmelz EA, Huffaker A, Sims JW, Christensen SA, Lu X, et al. 2014. Biosynthesis, elicitation and roles of monocot terpenoid phytoalexins. Plant J 79:659–78 [DOI] [PubMed] [Google Scholar]
- 137.Segonzac C, Macho AP, Sanmartin M, Ntoukakis V, Sanchez-Serrano JJ, Zipfel C. 2014. Negative control of BAK1 by protein phosphatase 2A during plant innate immunity. EMBO J 33:2069–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Seybold H, Trempel F, Ranf S, Scheel D, Romeis T, Lee J. 2014. Ca2+ signalling in plant immune response: from pattern recognition receptors to Ca2+ decoding mechanisms. New Phytol 204:782–90 [DOI] [PubMed] [Google Scholar]
- 139.Shi H, Shen Q, Qi Y, Yan H, Nie H, et al. 2013. BR-SIGNALING KINASE1 physically associates with FLAGELLIN SENSING2 and regulates plant innate immunity in Arabidopsis. Plant Cell 25:1143–57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Shimizu T, Nakano T, Takamizawa D, Desaki Y, Ishii-Minami N, et al. 2010. Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice. Plant J 64:204–14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Shinya T, Nakagawa T, Kaku H, Shibuya N. 2015. Chitin-mediated plant-fungal interactions: catching, hiding and handshaking. Curr. Opin. Plant Biol 26:64–71 [DOI] [PubMed] [Google Scholar]
- 142.Shinya T, Yamaguchi K, Desaki Y, Yamada K, Narisawa T, et al. 2014. Selective regulation of the chitin-induced defense response by the Arabidopsis receptor-like cytoplasmic kinase PBL27. Plant J 79:56–66 [DOI] [PubMed] [Google Scholar]
- 143.Shiu SH, Karlowski WM, Pan RS, Tzeng YH, Mayer KFX, et al. 2004. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16:1220–34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Song WY, Wang GL, Chen LL, Kim HS, Pi LY, et al. 1995. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–6 [DOI] [PubMed] [Google Scholar]
- 145.Spoel SH, Dong X. 2012. How do plants achieve immunity? Defence without specialized immune cells. Nat. Rev. Immunol 12:89–100 [DOI] [PubMed] [Google Scholar]
- 146.Sreekanta S, Bethke G, Hatsugai N, Tsuda K, Thao A, et al. 2015. The receptor-like cytoplasmic kinase PCRK1 contributes to pattern-triggered immunity against Pseudomonas syringae in Arabidopsis thaliana. New Phytol 207:78–90 [DOI] [PubMed] [Google Scholar]
- 147.Sun T, Zhang Y, Li Y, Zhang Q, Ding Y, Zhang Y. 2015. ChIP-seq reveals broad roles of SARD1 and CBP60g in regulating plant immunity. Nat. Commun 6:10159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Sun W, Dunning FM, Pfund C, Weingarten R, Bent AF. 2006. Within-species flagellin polymorphism in Xanthomonas campestris pv campestris and its impact on elicitation of Arabidopsis FLAGELLIN SENSING2-dependent defenses. Plant Cell 18:764–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Sun YD, Li L, Macho AP, Han ZF, Hu ZH, et al. 2013. Structural basis for flg22-induced activation of the Arabidopsis FLS2-BAK1 immune complex. Science 342:624–28 [DOI] [PubMed] [Google Scholar]
- 150.Tanaka K, Choi JM, Cao YR, Stacey G. 2014. Extracellular ATP acts as a damage-associated molecular pattern (DAMP) signal in plants. Front. Plant Sci 5:446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Tanaka K, Swanson SJ, Gilroy S, Stacey G. 2010. Extracellular nucleotides elicit cytosolic free calcium oscillations in Arabidopsis. Plant Physiol 154:705–19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Testerink C, Munnik T. 2011. Molecular, cellular, and physiological responses to phosphatidic acid formation in plants. J. Exp. Bot 62:2349–61 [DOI] [PubMed] [Google Scholar]
- 153.Torres MA, Jones JD, Dangl JL. 2006. Reactive oxygen species signaling in response to pathogens. Plant Physiol 141:373–78 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Tsuda K, Sato M, Glazebrook J, Cohen JD, Katagiri F. 2008. Interplay between MAMP-triggered and SA-mediated defense responses. Plant J 53:763–75 [DOI] [PubMed] [Google Scholar]
- 155.Tsuda K, Sato M, Stoddard T, Glazebrook J, Katagiri F. 2009. Network properties of robust immunity in plants. PLOS Genet 5:e1000772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Tunc-Ozdemir M, Urano D, Jaiswal DK, Clouse SD, Jones AM. 2016. Direct modulation of heterotrimeric G protein–coupled signaling by a receptor kinase complex. J. Biol. Chem 291:13918–25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.van der Luit AH, Piatti T, van Doorn A, Musgrave A, Felix G, et al. 2000. Elicitation of suspension-cultured tomato cells triggers the formation of phosphatidic acid and diacylglycerol pyrophosphate. Plant Physiol 123:1507–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Vleeshouwers VG, Driesprong JD, Kamphuis LG, Torto-Alalibo T, Van’t Slot KA, et al. 2006. Agroinfection-based high-throughput screening reveals specific recognition of INF elicitins in Solanum. Mol. Plant Pathol 7:499–510 [DOI] [PubMed] [Google Scholar]
- 159.Wang L, Tsuda K, Truman W, Sato M, Nguyen LEV, et al. 2011. CBP60g and SARD1 play partially redundant critical roles in salicylic acid signaling. Plant J 67:1029–41 [DOI] [PubMed] [Google Scholar]
- 160.Willmann R, Lajunen HM, Erbs G, Newman MA, Kolb D, et al. 2011. Arabidopsis lysin-motif proteins LYM1 LYM3 CERK1 mediate bacterial peptidoglycan sensing and immunity to bacterial infection. PNAS 108:19824–29 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Wong HL, Pinontoan R, Hayashi K, Tabata R, Yaeno T, et al. 2007. Regulation of rice NADPH oxidase by binding of Rac GTPase to its N-terminal extension. Plant Cell 19:4022–34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Yamada K, Yamaguchi K, Shirakawa T, Nakagami H, Mine A, et al. 2016. The Arabidopsis CERK1-associated kinase PBL27 connects chitin perception to MAPK activation. EMBO J 35:2468–83 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Yamaguchi K, Yamada K, Ishikawa K, Yoshimura S, Hayashi N, et al. 2013. A receptor-like cytoplasmic kinase targeted by a plant pathogen effector is directly phosphorylated by the chitin receptor and mediates rice immunity. Cell Host Microbe 13:347–57 [DOI] [PubMed] [Google Scholar]
- 164.Yamaguchi T, Minami E, Shibuya N. 2003. Activation of phospholipases by N-acetylchitooligosaccharide elicitor in suspension-cultured rice cells mediates reactive oxygen generation. Physiol. Plant 118:361–70 [Google Scholar]
- 165.Yamaguchi T, Yamada A, Hong N, Ogawa T, Ishii T, Shibuya N. 2000. Differences in the recognition of glucan elicitor signals between rice and soybean: β-glucan fragments from the rice blast disease fungus Pyricularia oryzae that elicit phytoalexin biosynthesis in suspension-cultured rice cells. Plant Cell 12:817–26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Yamaguchi Y, Pearce G, Ryan CA. 2006. The cell surface leucine-rich repeat receptor for AtPep1, an endogenous peptide elicitor in Arabidopsis, is functional in transgenic tobacco cells. PNAS 103:10104–9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Zeidler D, Zahringer U, Gerber I, Dubery I, Hartung T, et al. 2004. Innate immunity in Arabidopsis thaliana: Lipopolysaccharides activate nitric oxide synthase (NOS) and induce defense genes. PNAS 101:15811–16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Zhang J, Li W, Xiang T, Liu Z, Laluk K, et al. 2010. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 7:290–301 [DOI] [PubMed] [Google Scholar]
- 169.Zhang L, Kars I, Essenstam B, Liebrand TW, Wagemakers L, et al. 2014. Fungal endopolygalacturonases are recognized as microbe-associated molecular patterns by the Arabidopsis receptor-like protein RESPONSIVENESS TO BOTRYTIS POLYGALACTURONASES1. Plant Physiol 164:352–64 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Zhang W, Fraiture M, Kolb D, Loffelhardt B, Desaki Y, et al. 2013. Arabidopsis receptor-like protein30 and receptor-like kinase suppressor of BIR1–1/EVERSHED mediate innate immunity to necrotrophic fungi. Plant Cell 25:4227–41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Zhang W, He SY, Assmann SM. 2008. The plant innate immunity response in stomatal guard cells invokes G-protein-dependent ion channel regulation. Plant J 56:984–96 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Zhang X, Mou Z. 2009. Extracellular pyridine nucleotides induce PR gene expression and disease resistance in Arabidopsis. Plant J 57:302–12 [DOI] [PubMed] [Google Scholar]
- 173.Zhang Y, Xu S, Ding P, Wang D, Cheng YT, et al. 2010. Control of salicylic acid synthesis and systemic acquired resistance by two members of a plant-specific family of transcription factors. PNAS 107:18220–25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Zhang Y, Zhu H, Zhang Q, Li M, Yan M, et al. 2009. Phospholipase dα1 and phosphatidic acid regulate NADPH oxidase activity and production of reactive oxygen species in ABA-mediated stomatal closure in Arabidopsis. Plant Cell 21:2357–77 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Zhou J, Lu D, Xu G, Finlayson SA, He P, Shan L. 2015. The dominant negative ARM domain uncovers multiple functions of PUB13 in Arabidopsis immunity, flowering, and senescence. J. Exp. Bot 66:3353–66 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Zhu X, Caplan J, Mamillapalli P, Czymmek K, Dinesh-Kumar SP. 2010. Function of endoplasmic reticulum calcium ATPase in innate immunity-mediated programmed cell death. EMBO J 29:1007–18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JD, et al. 2006. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125:749–60 [DOI] [PubMed] [Google Scholar]


