Abstract
Empirical observations of how labs conduct research indicate that the adoption rate of open practices for transparent, reproducible, and collaborative science remains in its infancy. This is at odds with the overwhelming evidence for the necessity of these practices and their benefits for individual researchers, scientific progress, and society in general. To date, information required for implementing open science practices throughout the different steps of a research project is scattered among many different sources. Even experienced researchers in the topic find it hard to navigate the ecosystem of tools and to make sustainable choices. Here, we provide an integrated overview of community-developed resources that can support collaborative, open, reproducible, replicable, robust and generalizable neuroimaging throughout the entire research cycle from inception to publication and across different neuroimaging modalities. We review tools and practices supporting study inception and planning, data acquisition, research data management, data processing and analysis, and research dissemination. An online version of this resource can be found at https://oreoni.github.io. We believe it will prove helpful for researchers and institutions to make a successful and sustainable move towards open and reproducible science and to eventually take an active role in its future development.
Keywords: Open science, Reproducibility, MRI, PET, MEG, EEG
1. Introduction
Science is an incremental progress towards creating and organizing knowledge through theories and testable predictions. Reproducibility is a core part of science: being able to repeat or recreate scientific results is essential for the complex process of knowledge accumulation. Due to its relevance, different terms have been introduced to describe specific aspects of the process, including “reproducibility” when the same data and methods are used, “replicability” when new data but same methods are used, “robustness” when the same data but different methods are used, and “generalizability” when new data and methods are used (Whitaker, 2019). Here, we use “reproducibility” as an umbrella term, encompassing all aspects of recreating scientific results (Poldrack et al., 2020). Open science tools and practices have been developed to advance reproducibility, as well as accessibility and transparency at all stages of the research cycle and across all levels of society. Together, they remove barriers to sharing and facilitate collaboration, with the goal of improving reproducibility and, ultimately, accelerating scientific discoveries. Importantly, such practices facilitate, but do not guarantee, higher quality.
Empirical observations of how labs conduct research indicate that the adoption rate of open practices and tools for reproducible and collaborative science, unfortunately, remains in its infancy. Even when members of a specific scientific community have taken a central role in open science advocacy and tool development, like in the neuroimaging community, the impact on the rest of the very same community is limited. A recent survey (Paret et al., 2022) including researchers who are senior and likely to hold a positive attitude towards open science, indicated that 42% have never pre-registered a neuroimaging study and 34% have never shared their raw neuroimaging data. Many of those who indicated that they pre-registered or shared their data at least once likely did not do so in all their studies, and thus, the actual rate of pre-registration and data sharing in neuroimaging is likely much lower.
The limited adoption of open science practices is at odds with the overwhelming evidence that a lack of open practices in general can hinder reproducibility with costs for scientific progress and for society. Indeed, reproducibility issues have been undermining the foundation of scientific research in several fields, such as psychology (Open Science Collaboration, 2015; Klein et al., 2018), social sciences (Camerer et al., 2016, 2018), neuroimaging (Munafò et al., 2017; Botvinik-Nezer et al., 2020; Li et al., 2021), preclinical cancer biology research (Errington et al., 2021; Errington et al., 2021), and more (Hutson, 2018; Nissen et al., 2016; Serra-Garcia and Gneezy, 2021). As a response, there has been a rise in the development of tools and approaches to facilitate reproducibility and open science, in the spirit of Findability, Accessibility, Interoperability, and Reusability principles (FAIR) (Wilkinson et al., 2016; Gorgolewski and Poldrack, 2016; Nosek et al., 2018; Nosek et al., 2012; Nosek and Lakens, 2014; Poldrack et al., 2017; Poldrack et al., 2020; Poldrack et al., 2019; Clayson et al., 2022). Beyond their potential to mitigate transparency and reproducibility issues, these practices provide important benefits for individual researchers by increasing exposure, reputation, chances of publication, number of citations, media attention, potential collaborations, and position and funding opportunities (Allen and Mehler, 2019; McKiernan et al., 2016; Nosek et al., 2022; Markowetz, 2015; Hunt, 2019). Hence, one could have expected a higher uptake for such beneficial practices and tools.
Recently, a parallel top-down change of policies started to further support the adoption of open science practices and tools. For example, funding agencies are now enforcing the implementation of certain open data practices for publicly funded research (e.g., the NIH in the U.S. and the ERC in Europe; de San Román 2021; de Jonge et al., 2021), and some require a plan for research data storage and sharing, openly accessible publication formats and dissemination plans beyond the classical journal publication. Additionally, they provide funding for the development of necessary software, hardware, and collaborative infrastructure to support the transition to open and reproducible neuroscience (e.g., the NIH BRAIN Initiative, NIH ReproNim project (Kennedy et al., 2019), NSF CRCNS, EU Human Brain Project, German NFDI). These efforts by funding agencies are complemented by stakeholder institutions like the OHBM, the International Neuroinformatics Coordinating Facility (INCF), the Chinese Open Science Network (COSN), and the Open Science Framework (OSF), who provide platforms for the development of standards and best practices of open and FAIR neuroscience research, assemble training material, and promote open science practices. More-over, journals have started changing their policies with regard to open access options and data sharing. Together, these institutional measures aim at fostering the benefits of open science practices, and the adoption of open and reproducible science standards will be increasingly required for labs and individual researchers.
Nevertheless, multiple barriers of entry to open science practices are driving the modest rate of adoption in the general research community. Among them are lack of knowledge or training and lack of skills or resources. A survey by Borghi and Van Gulick (2018) found that 65% of researchers reported openness and reproducibility as motivation for implementing research data management in MRI, but between 40-50% pointed to the lack of best practices/tools and knowledge/training as main obstacles for embracing these practices. Likewise, a more recent survey indicated that similar percentages of researchers in neuroimaging have never learned how to pre-register or share their data online and that they know too little about pre-registration platforms and suitable data repositories (Paret et al., 2022). These later challenges could be alleviated by a simplified overview of the open resources available. However, information required for implementing open science practices over the full research cycle is currently scattered among many different sources. Even experienced researchers in the topic often find it hard to navigate the ecosystem of community-developed tools and to make sustainable choices.
This manuscript provides an integrated overview of community-developed resources critical to support open and reproducible neuroimaging throughout the entire research cycle and across different neuroimaging modalities (particularly MRI, MEG, EEG, and PET). Instead of detailing, as others before, why one should adopt open and reproducible practices (Munafò et al., 2017; Nosek et al., 2012; Poldrack et al., 2017; McKiernan et al., 2016), we focus on providing a resource overview. Our goal is to make it easier for scientists to select the most valuable instruments for their practice at every step of the research workflow, and consequently accelerate the broader adoption of open science tools and practices, increasing scientific reproducibility and openness. We provide justification on why each implements good practices, as well as how to integrate them into the research workflow.
In this review we do not aim to recommend particular tools over others, as the ideal ones may depend on many factors that vary between researchers. However, we highlight some points to consider at the time of selection. Typically it is advisable to choose tools that integrate with other tools and practices already established in the lab, have a relatively fast learning curve, and a long-term benefit. In order to increase sustainability, tools should be relatively mature, well maintained, and supported by an active community. Another indicator is whether the tools and practices are integrated in already established toolboxes or supported by large open science organizations. If still multiple tools meet these criteria, then it might be advantageous to choose one that is used by peers and collaborating partners. When we recommend practices, we state the problems they are supposed to address. We also encourage the readers to join the development teams and leadership of those tools, becoming an active part of the open neuroimaging community. Contributions from individuals who are experiencing barriers to the uptake of specific practices are particularly encouraged, since they can help mitigate these barriers for the benefit of everyone.
The manuscript is organized following the different steps of the research cycle: study inception and planning, data acquisition, research data management, data processing and analysis, and research dissemination. For each step we provide a figure with subgoals (subsections in the text) in the headings, some recommendations on how to achieve them in a bullet list, and supporting tools indicated by icons (see Figs. 1–5). To further guide the readers, the manuscript is accompanied by a detailed table containing links and pointers to the resources featured in the text of each section (see Table S1). In addition, the content is available online as a Jupyter Book at https://oreoni.github.io (https://doi.org/10.5281/zenodo.7083031).
Fig. 1.

Study inception and planning
For each step, the figure contains the main goals (headings), specific recommendations (bullet list), and useful tools (icons).
Sources: Icons from the Noun Project: Registration by WEBTECHOPS LLP; Share by arjuazka; Computer warranty by Thuy Nguyen; Logos: used with permission by the respective copyright holders.
Fig. 5.
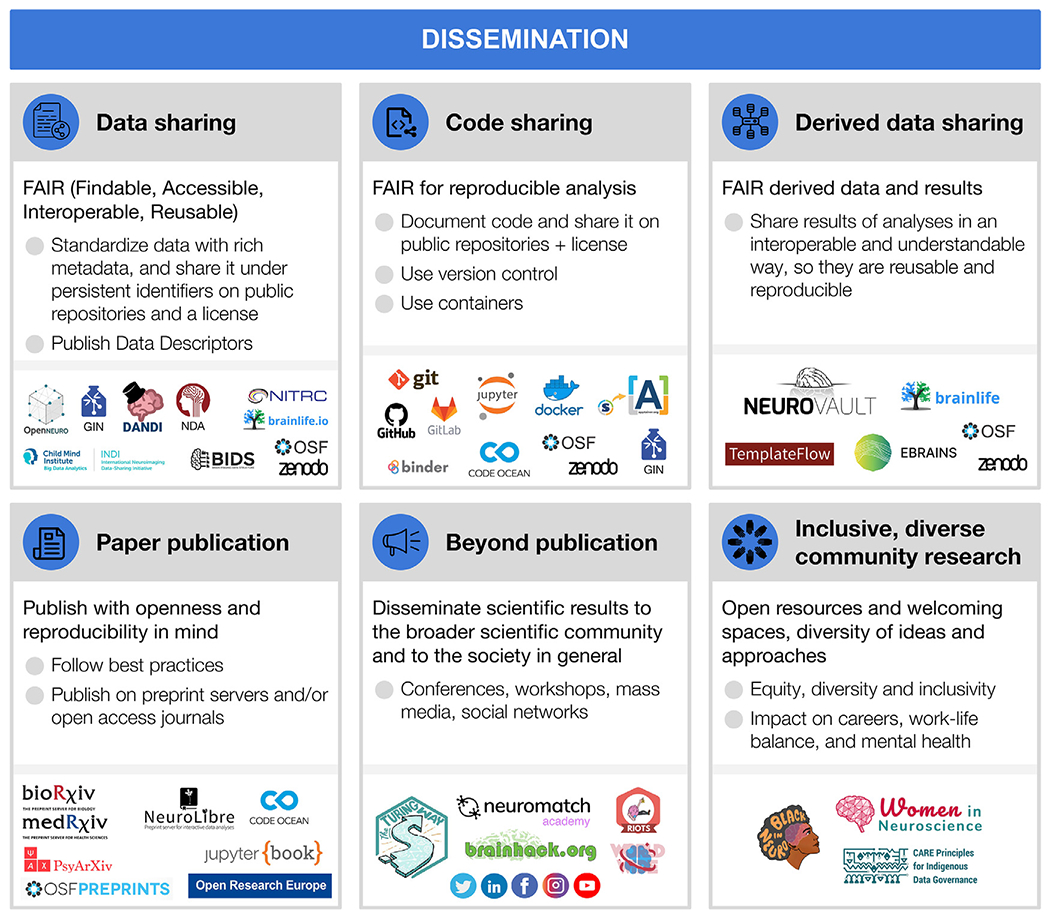

Dissemination
For each step, the figure contains the main goals (headings), specific recommendations (bullet list), and useful tools (icons).
Sources: Icons from the Noun Project: Data Sharing by Design Circle; Share Code by Danil Polshin; Data by Nirbhay; Publication by Adrien Coquet; Broadcast by Amy Chiang; Logos: used with permission by the copyright holders.
2. Study inception, planning, and ethics
Each individual decision from the beginning of the study will contribute to facilitate or hamper reproducibility. In the current section, we will describe practices and tools for preparation, piloting, preregistration, obtaining participants’ consent and ongoing quality control and assessment (see Fig. 1).
2.1. Study preparation and piloting
Research projects usually begin with descriptions of general, theoretical questions in documents such as grants or thesis proposals. Such foundations are essential but necessarily broad. When the project moves from proposal to implementation, these descriptions are translated into concrete protocols and stimuli, a process that can be streamlined by the incorporation of open procedures and comprehensive piloting. The promise is that the more preparation and piloting is conducted prior to data collection, the more likely it is that the project will be successful: that analyses of its data can contribute to answering its motivating ideas and questions (Strand, 2021).
Standard Operating Procedures (SOPs) can take different forms, and are powerful tools for planning, conducting, recording, and sharing projects. Ideally, SOPs describe the entire data collection procedure (e.g., recruitment, performing the experiment, data storage, preprocessing, quality control analyses), in sufficient detail for a reader to conduct the experiment themselves with minimal supervision, thereby contributing to reproducibility. SOPs may begin as an outline with vague descriptions, preferably during the pilot stage, and then become more detailed over time. For example, if a session is lost due to a button box signal failure, an image of its correct settings could be added to the SOPs. At the end of the project, its SOPs should be released along with its publications and datasets, to provide a source of answers for the detailed procedural information that may be needed for experiment reproducibility or dataset reuse, but are not included in typical publications.
Many resources can assist with experiment planning and SOP creation. Documents and experiences from similar studies conducted locally are valuable, but should not be the only source of information during planning, since, for example, a procedure may be considered standard in one institution but not in another. Public SOPs can serve as examples, as can protocols published on specialized sites (e.g., Protocol Exchange, protocols.io, Nature Protocols; see Table S1). Best practices guides are now available for many imaging modalities (MRI: Nichols et al. 2017; MEG/EEG: Pernet et al. 2020; fNIRS: Yucel et al. 2021; PET: Knudsen et al. 2020). Open resources for stimulus presentations and behavioral data acquisition are also recommended to increase reproducibility (see Section 3.2).
Piloting should be considered an integral part of the planning process. By “piloting” we mean the acquisition and evaluation of data prior to the collection of the actual experimental data, verifying the feasibility of the whole research workflow. While it is not a necessary prerequisite for reproducibility, it is a good scientific practice to produce higher quality research, and facilitates reproducibility via better documentation and SOPs. A piloting stage before starting data collection is important, not only for ensuring that the protocol will go smoothly when the first participant arrives, but also that the SOPs are complete and, critically, that the planned analyses can be carried out with the actual experimental data recorded. For example, pilot tests may be set up to confirm that the task produces the expected behavioral patterns (e.g., faster reaction time in condition A than B), that the log files are complete, and that image acquisition can be synchronized with stimulus presentation. Piloting should also include testing the data storage and retrieval strategies, which may include storing consent documents (Section 2.3), questionnaire responses, imaging data, and quality control reports. SOPs also prescribe how data will be organized, preferably according to a schema (e.g., the Brain Imaging Data Structure, Section 4.1). Data organization largely determines the efficient implementation of analysis pipelines and improved reproducibility, reusability, and shareability of the data and results.
Analyses of the pilot data are very important and can take several forms. One is to test for the effects of interest: establishing that the desired analyses can be performed and that the data quality is sufficient to produce valid and reproducible results (Sections 2.2. and 2.4; for power estimation tools see Table S1). A second type of pilot analysis is to establish tests for effects not of direct interest, but suitable for controls. As discussed further in Section 2.4, positive control analyses involve strong, well-understood effects that must be present in a valid dataset. It is worth mentioning that well structured and documented openly available datasets (see Section 4) could also serve for analysis piloting though they would lack the test for potential site specific technical issues. Simulations could also be used to ensure that the planned analysis is doable and valid.
2.2. Pre-registration
Pre-registration is the specification of the research plan in advance, prior to data collection or at least prior to data analysis (Nosek and Stephen Lindsay, 2018). Pre-registration usually includes the study design, the hypotheses and the analysis plan. It is submitted to a registry, resulting in a frozen time-stamped version of the research plan. Its main aim is to distinguish between hypothesis-testing (confirmatory) research and hypothesis-generating (exploratory) research. While both are necessary for scientific progress, they require different tests and the conclusions that can be inferred based on them are different (Nosek et al., 2018)
Registered reports is a relatively novel publishing format that can be seen as advanced pre-registration. This format is becoming very common, with a growing number of hundreds of participating journals (Hardwicke and Ioannidis, 2018; Chambers, 2019). In a registered report, a detailed pre-registration is submitted to a specific journal, including introduction, planned methods and potentially preliminary data. Then, it goes through peer review prior to data collection (or prior to data analysis in certain cases, for example for studies that rely on large-scale publicly shared data). If the proposed plan is approved following peer review, it receives an “in-principle acceptance”, indicating that if the researchers follow their accepted plan, and their conclusions fit their findings, their paper will be published. An in-principle accepted registered report is sometimes required to be additionally pre-registered. Recently, a platform for peer review of registered reports preprints was launched, named “Peer Community in registered reports” (see Table S1).
There are many benefits to pre-registration, from the field to the individual level. Transparency with regard to the research plan, and whether an analysis is confirmatory or exploratory, increases the credibility of scientific findings. It helps to mitigate some of the effects of human biases on the scientific process, and reduces analytical flexibility, p-hacking (Simmons et al., 2011) and hypothesizing after the results are known (Nosek et al., 2019; Munafò et al., 2017; Nature, 2015; Kerr, 1998). There is initial evidence that the quality of pre-registered research is judged higher than in conventional publications (Soderberg et al., 2021). Nonetheless, it should be noted that pre-registration and registered reports are not sufficient to fully protect against questionable research practices (Paul et al., 2021; Devezer et al., 2021; Rubin, 2020) and their general impact will depend on the extent journals will implement them. Registered reports also mitigate publication bias by accepting papers based on hypothesis and methods, independently of the findings. Indeed, it has been shown that pre-registered studies and registered reports include more null findings (Allen and Mehler, 2019; Kaplan and Irvin, 2015; Scheel, 2020) and report lower effect sizes (Schäfer and Schwarz, 2019) compared to other studies. For the individual researcher, registered reports with a two-stage review are an excellent example in which authors benefit from feedback on their methods before even starting data collection. They can help improve the research plan and spot mistakes in time, and provide assurance that the study will be published (Wagenmakers and Dutilh, 2016; Kidwell et al., 2016). It should be noted, though, that registered reports can require a significant time commitment, that is likely to pay off in the long-term but is not easily accommodated in many traditional project funding models.
While pre-registration is not the common practice yet, it is becoming more common over time (Nosek and Stephen Lindsay, 2018) and requirements by journals and funding agencies are already changing. There are many available templates and forms for pre-registration, organized by discipline or study types, for example for fMRI and EEG (see Table S1), and published guidelines for pre-registration in EEG are also available (Govaart and Schettino, 2022; Paul et al., 2021). There are different approaches as to what should be pre-registered. For instance, some believe it should be an exhaustive description of the study, including the background and justification for the research plan, while others believe it should be a short and concise document, including only the necessary details to reduce the likelihood of p-hacking and allowing reviewers to review it properly during the peer review process (Simmons et al., 2021). Pre-registration can also be flexible and adaptive by pre-registering contingency plans or complex decision trees (Benning et al., 2019).
Once researchers develop an idea and design a study, they can write and pre-register their research plan (Nosek and Stephen Lindsay, 2018; Paul et al., 2021). Pre-registration could be performed following the piloting stage (Section 2.1), but studies can be pre-registered irrespective of whether they include a piloting stage or not. There are many online registries where researchers can pre-register their study. The three most frequently used platforms are: (1) OSF, a platform that can also be used to share additional information about the study/project (such as data and code), with multiple templates and forms for different types of pre-registration, in addition to extensive resources about pre-registration and other open science practices; (2) aspredicted.org, a simplified form for pre-registration (Simmons et al., 2021); and (3) clinicaltrials.gov, which is used for registration of clinical trials in the U.S. (see Table S1).
Once the pre-registration is submitted, it can remain private or become public immediately, depending on the platform and the re-searcher’s preferences. Then, the researcher collects the data and executes the research plan. When writing the manuscript to report the study, the researcher is advised to include a link to the pre-registration, clearly and transparently describe and justify any deviation from the pre-registered plan and also report all registered analyses. Additional analyses to deepen some results or looking into unexpected effects are encouraged, and are part of the routine scientific investigation. The added benefit of pre-registration is that such analyses do not need to reach pre-specified levels of significance because they are reported as exploratory.
2.3. Ethical review and data sharing plan
The optimism of the scientific community about improving science by making all research assets open and transparent has to take into account privacy, ethics and the associated legal and regulatory needs for each institution and country. Whereas on the one hand sharing data (most often collected with public funds) is critical to advance science, on the other hand, sharing data can in some situations become infeasible to safeguard privacy. Data governance concerns the regulatory and ethical aspects of data management and sharing of data files, metadata and data-processing software (see Sections 6.1–3). When data sharing crosses national borders, data governance is called International Data Governance (IDG). IDG depends on ethical, cultural and international laws.
Data sharing is beneficial for both reproducibility and the exploration and formulation of new hypotheses. Therefore, it is important to ensure, prior to data collection, that the collected data could be later shared. Open and reproducible neuroimaging thus starts by (1) planning which data would be collected; (2) planning how these data would later be shared; (3) having ethical and legal clearance to share data; but also (4) the infrastructural means for this sharing (for more information about data sharing and available platforms, see Section 6; for data governance see Section 4).
Since 2014, the Open Brain Consent project (see Table S1), which was founded under the ReproNim project umbrella, provides examples and templates translated to multiple languages to help researchers prepare consent forms for data sharing, including the recent development of an EU GDPR-compliant template (The Open Brain Consent working group, 2021). Such consent should include a statement about how the data will be shared, with whom, potential risks, and that the consent to share can be withdrawn (which is separate from consent to participate and withdraw from the study). Data sharing forms should also make explicit how these factors determine to what extent a later withdrawal or editing of the data on the repository is possible. Given the international nature of the majority of neuroscience projects, IDG has become a priority (Eiss, 2020). Further work will be needed to implement an IDG approach that can facilitate research while protecting privacy and ethics. Specific recommendations on how to implement IDG have been proposed (Eke et al., 2022).
It should be noted that ethical review comprises more than data sharing procedures. Its goals are safety, self determination, and the protection of rights of study participants. A central element is informed consent to participate in the study, which requires that technical and scientific aspects of the study as well as regulations for participation in the study are transparently communicated to the participants. Clinical research may require adherence to additional, country-specific regulations. Finally, when planning the recruitment procedures, it is important to aim for equity, diversity and inclusivity (Henrich et al., 2010; Forbes et al., 2021), avoiding obtaining results that may not generalize to larger populations and improving the quality of research (e.g., Baggio et al. 2013).
2.4. Looking at the data early and often: monitoring quality
Inevitably, unexpected events and errors will occur during every experiment and in every part of the research workflow. These can take many forms, including dozing participants, hardware malfunction, data transfer errors, and mislabeled results files. As data progresses through the workflow, issues are likely to cascade and amplify, perhaps masking or mediating experimental effects, thereby damaging the reliability of the results. The impact of such surprises can range from the trivial and easily corrected to the catastrophic, rendering the collected data unusable or conclusions drawn from it invalid. Identifying issues and errors as early as possible is important to enable adding corrective measures to the protocol, but also because some issues are much easier to detect when the data are in a less-processed form. For example, a number of typical artifacts in anatomical MRI are known to be easier to identify in the background of the image and regions of no-interest (Mortamet et al., 2009), and can easily remain undetected if the first quality control check is set up after, e.g., brain extraction, which masks out non brain tissue. Thus, it is fundamental to pre-establish within the SOPs (Section 2.1) the mechanisms set in place to ensure the quality of the study. There are several mechanisms available that help to ensure that all required data are being recorded with sufficient quality and in a way that makes them analyzable.
Quality control checkpoints.
Establishing quality control (QC) checkpoints (Strother, 2006) is necessary for every project: which data are usable for analyses, and which are not? At these key points in the pre-processing or analysis workflow the data’s quality is checked, and if insufficient, it does not move on to the next stage. Results from low quality data are much less likely to be reproducible with new data or methods. Critically, the exclusion criteria of each checkpoint must be defined in advance (preferably stated in the SOPs and the pre-registration document, see Sections 2.1 and 2.2) to preempt unintentional cherry-picking (i.e., excluding data points to reinforce the results), which is a major contributor to irreproducibility via undisclosed flexibility. Some criteria are widely accepted and applicable, for example, that all neuroimaging data should be screened to eliminate clear artifacts, such as data corrupted by incidental electromagnetic interference or participants movements. A similarly well-established checkpoint of the workflow is visualizing and inspecting the outputs of surface reconstruction methods in MRI, checking time activity curves in high binding regions for PET or power spectral content in MEG and EEG; these fundamental QC checkpoints and their implementation are heavily dependent on the immediately previous processing step. Such QC may be conducted manually by experts using software aids, like visual summary reports or visualization software such as MRIQC (Esteban et al., 2017). More objective, automatic exclusion criteria, are currently an open and active line of work in neuroimaging (e.g., Ding et al. 2019; Kollada et al. 2021; Esteban, Blair, et al. 2019). Some QC checkpoints, such as for acceptable task performance or participant movement, are often defined for individual tasks, experiments and hypotheses.
Quality assurance (QA).
Tracking QC decisions will also enable identifying structured failures and artifacts that require not just excluding affected datasets, but rather taking corrective actions to preempt propagation to additional datasets. When a mishap occurs, the experimenters should investigate its cause, and if possible, change the SOPs (Section 2.1) and related materials to reduce the chance of it happening again. For example, if many participants report confusion about task instructions, the training procedure and experimenter script could be altered. Automated checks and reports can be very effective, such as real-time monitoring of participant motion during data collection (Heunis et al., 2020), or validating that image parameters are as expected before storage (e.g., with XNAT Marcus et al. 2007 or ReproNim tools Kennedy et al. 2019).
Positive control analyses.
A final aspect of quality assurance is the incorporation of positive control analyses: analyses included not because they are of interest for the scientific questions, but because they provide evidence that the dataset is of sufficient quality to conduct the analyses of interest, and that the analysis is valid. Ideally, positive control analyses focus on strong, well-established effects that must be present if the dataset is valid. For example, with task fMRI designs, button pressing, which should be associated with contralateral motor activation, is often a convenient target for positive control analysis. In MEG and EEG, participants can be asked to blink their eyes, open their mouths, or clench their jaws, and the recordings checked for the associated artifacts. Positive control analyses should also be carried out during piloting, when changes to the protocol are still possible (see Section 2.1). For example, if button presses are not clearly detectable during piloting, the acquisition sequence may not have sufficient SNR for the planned analyses and thus should be modified. Positive controls can further serve for analysis pipeline optimization prior to conducting the optimized analysis on the outcome of interest, thus preventing legitimate optimization from turning into p-hacking.
3. Data acquisition
Data acquisition is largely carried out with vendored systems. Manufacturers typically keep their software and hardware closed or semi-open at most. As a result, researchers often receive highly processed (e.g., reconstructed) data as ‘raw’ data from the devices. The lack of transparency in the acquisition details and downstream proprietary processing prevents end-to-end reproducible neuroimaging workflows. Reproducibility is endangered, for instance, by heterogeneity in data formats, definition of critical experimental parameters, and technological differences that are translated into the data as spurious, non-biological differences between acquisition devices.
A central issue is the proprietary nature of acquisition protocols. Many imaging device manufacturers require developers to use building blocks from vendor-exclusive toolboxes. This closes the door on open-source development and hampers multi-center consensus for modern imaging methods, especially in research. These shortcomings of mostly closed solutions have triggered a growing interest in open-source acquisition hardware and software (Winter et al., 2016), Here, we provide a brief review of these developments and accompanying solutions aimed at fostering open and collaborative acquisition method development across imaging modalities (see Fig. 2).
Fig. 2.
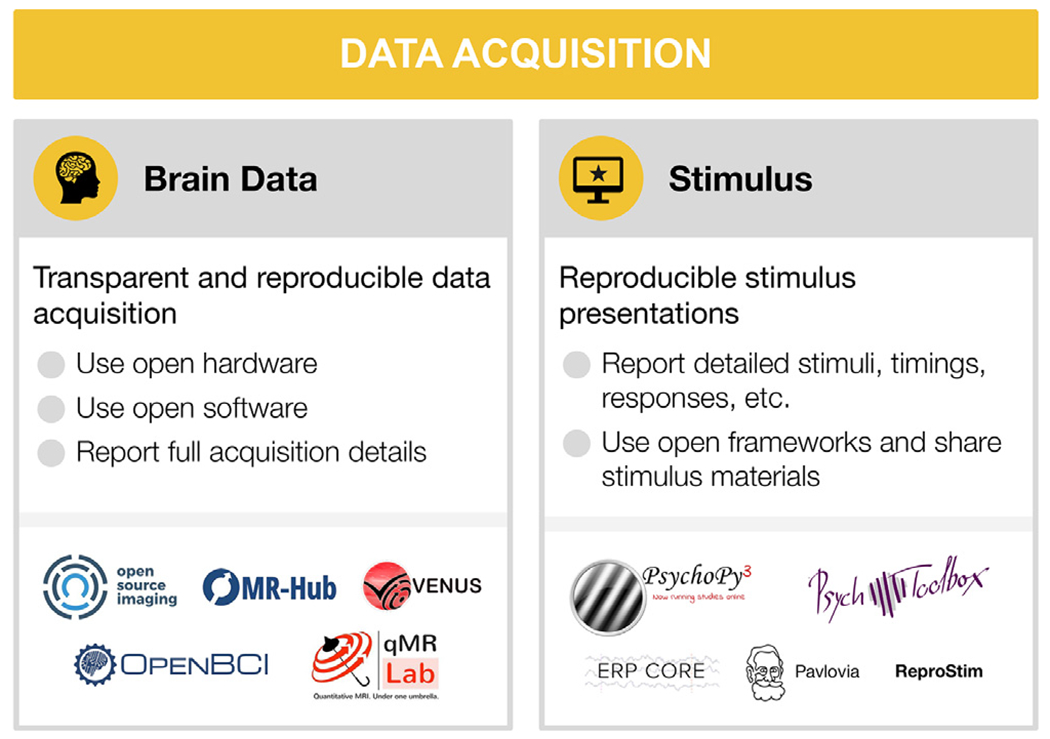
Data acquisition
For each step, the figure contains the main goals (headings), specific recommendations (bullet list), and useful tools (icons).
Sources: Icons from the Noun Project: Brain by parkjisun; Computer Screen by Icon Solid (adapted with a star); Logos: used with permission by the copyright holders.
3.1. Brain data acquisition
A common approach advocated by MRI researchers is establishing consensus protocols to standardize data acquisition. One of the flagship applications of this strategy is the Human Connectome Project (HCP) protocol, which achieved this within the confines of a single vendor (Smith et al., 2013). The HCP acquisition sequences and reconstruction software are compiled for different MRI scanner versions of a single vendor, openly distributed and maintained for fMRI applications (Uğurbil et al., 2013). However, it is generally difficult to achieve good inter vendor agreement using off the shelf software even for widely used protocols, such as apparent diffusion coefficient and longitudinal relaxation time (Sasaki et al., 2008; Lee et al., 2019). In addition, not all software options are available from all vendors (for example, compressed sensing (Lustig et al., 2008) and frequency-domain based parallel imaging methods (Breuer et al., 2005; Griswold et al., 2002). More over, even seemingly simple image enhancement protocols, such as image inhomogeneity corrections, are often scarcely documented and validated but can affect inferences drawn from an experiment (Schmitt and Rieger, 2021; Jellús and Kannengiesser 2014). Users typically have access to key parameters of pulse sequences, which are at the center of data acquisition. The exact pulse sequence descriptions are vendor-specific and may even change between software upgrades of a single vendor. This makes it difficult to evaluate multi-center replicability of new acquisition methods or to acquire longitudinal data with confidence.
Fortunately, in the last decade, several vendor-neutral data acquisition pulse sequences and reconstruction frameworks have been developed to mitigate this problem: Pulseq (Layton et al., 2017), PyPulseq (Ravi et al., 2019), GammaStar (Cordes et al., 2020), TOPPE (Nielsen and Noll, 2018), ODIN (Jochimsen and von Mengershausen, 2004), and SequenceTree (Magland et al., 2016) (see Table S1). Although these tools vary in vendor compatibility and the flexibility of their acquisition runtime, they enable vendor-neutral deployment of pulse sequences with transparent access to all the details needed. Nevertheless, vendor-neutral raw data (k-space, i.e. the 2D or 3D Fourier space representation of the image) collection is half the battle.
To complete the puzzle of MRI acquisition, interoperable and open-source reconstruction frameworks are essential. Thanks to ISMRM-RD (Inati et al., 2017), a k-space data standard, community-developed reconstruction tools can have a unified way to run advanced reconstruction algorithms against undersampled raw data (Maier et al., 2021). Some of these tools include Gadgetron (Hansen and Sørensen, 2013), BART (Uecker et al., 2015), MRIReco.Jl (Knopp and Grosser, 2021) (see Table S1 for further tools and details). By streamlining these acquisition and reconstruction tools using data standards at multiple levels (Karakuzu et al., 2021; Inati et al., 2017) on a data-driven and container mediated workflow engine (Di Tommaso et al., 2017), end-to-end reproducible MRI workflows can be developed. A recent study has shown that this approach can significantly reduce inter-vendor variability of quantitative MRI measurements (Karakuzu et al., 2020; Karakuzu et al., 2022). Given the growing open-source MRI acquisition ecosystem, a variety of end-to-end workflows are possible. Therefore, community-driven validation frameworks have a key importance for interoperable solutions (Tong et al., 2021). Facilitated by these standards, effective and open communication methods development sets the future direction for reproducible MRI research (Stikov et al., 2019).
In PET, the variety between different scanners is even larger than in MRI. An overview over different scanner types based on their usage for a specific radiotracer targeting the serotonin transporter, namely [11 C]DASB, is given in (Nørgaard et al., 2019). Different PET scanners export images in slightly different data formats with little overlap in the Digital Imaging and Communications in Medicine (DICOM) PET specific tags. As with MRI, reconstruction is vendor/machine specific but open source solutions to image reconstruction are being developed, for instance the OMEGA toolbox (Wettenhovi et al., 2021). Data acquisition for PET is further complicated by the use of different PET tracers, injection methods, scan duration and scan framing or injected radioactivity dose.
In MEG and EEG, the problem of standardized data acquisition starts even earlier: unlike the common DICOM data format used across vendors in MRI or PET, MEG and EEG manufacturers do not use a common data format, and format specifications are rarely made public. More importantly, equipment implementation significantly differs between vendors, for example with respect to MEG sensor types, software noise suppression techniques, and EEG amplifiers and electrodes. There have been some efforts on developing open versions of some proprietary tools, for example, the Maxwell filtering for signal space separation by the MNE-python team (Gramfort et al., 2014). Additionally, initiatives, such as the OpenBCI, offer open EEG hardware and tools for biosensing and brain computer interfacing through continuous community driven development. As we have mentioned, very little is known on how the variability of data acquisition parameters affect downstream comparability of results. The EEGManyLabs project (Pavlov et al., 2021) will provide a comprehensive dataset in this regard, as many labs with different equipment try to replicate the same studies.
Given the large variations across different vendors for all neuroimaging modalities, which often cannot be overcome, it is crucial to report all data acquisition parameters in a comprehensive and standardized manner to make potential differences in data acquisition across studies and sites transparent (for a discussion of reporting guidelines see Section 6.4).
3.2. Stimulus presentation and behavior
Several actively maintained programs for stimulus presentation and response logging are available. Open source software include PsychoPy (Peirce et al., 2019) in Python and Psychtoolbox (Brainard, 1997; Pelli, 1997; Kleiner et al., 2007) in MATLAB. Both have many users, making it possible to get assistance and perhaps find an already-implemented task protocol (e.g., on Pavlovia for Psychopy). Modality specific resources also exist, for instance the ERP CORE (Compendium of Open Resources and Experiments; Kappenman et al. 2021) openly provides optimized paradigms for several widely used ERP components, along with scripts, data processing pipelines, and sample data.
Using open stimuli and presentation software generally increases the likelihood a dataset will be useful to others, and its results reproducible (Strand and Brown, n.d.2022). Although desirable, it is not always possible to use fully open stimuli, particularly in the case of commercial movies, audio plays, and image databases. Stimuli, presentation scripts, behavioral tests and related material should be shared whenever possible (see DuPre et al. 2019 for a list of datasets sharing naturalistic stimuli and Section 6). Researchers should always check the licenses on the stimulus materials they plan to use or share. To facilitate stimuli feature analysis and exact reproducibility of the experimental paradigms, such projects as ReproNim’s ReproStim (Connolly and Halchenko, 2022) could automate recording and archival of audio-visual stimuli. When specific stimuli or material can not be released, they should be described as unambiguously as possible and, if possible, providing the source, such as identification number (e.g., a GTIN), and scripts to (re)produce used stimuli from the commercial media.
4. Research data management
Good research data management (RDM), i.e. how data are organized, maintained, annotated, tracked, stored, and accessed throughout a research project, forms the basic foundations of result reproducibility, data reusability, and research efficiency (Wilkinson et al., 2016; Gorgolewski and Poldrack, 2016; Nosek et al., 2018; Nosek et al., 2012; Nosek and Lakens, 2014; Poldrack et al., 2017; Poldrack et al., 2020; Poldrack et al., 2019; Borghi and Van Gulick, 2021a; Poline et al., 2022). Consequently, Data Management Plans (DMPs) are widely required by funders even at the application phase (e.g., NIH and NSF in the U.S., ERC in Europe), increasingly expected by scientific peers, and holds considerable benefits for individual researchers. It is good practice to develop, review and execute DMPs for every experiment, whether or not it is required by the funding agency. While specific RDM requirements vary across subdisciplines, this section highlights RDM standards and tools applicable across neuroimaging, ranging from data organization to annotation and publication (see Fig. 3).
Fig. 3.
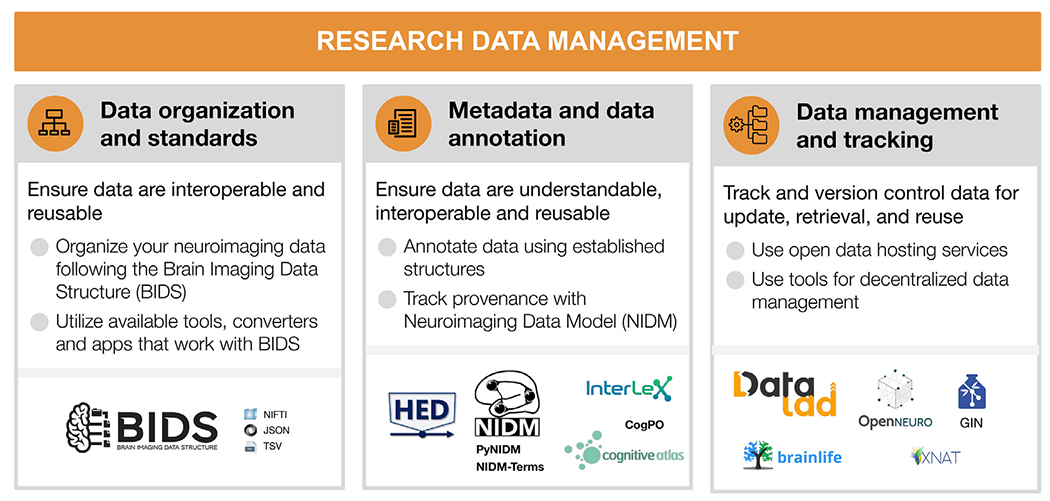
Research data management
For each step, the figure contains the main goals (headings), specific recommendations (bullet list), and useful tools (icons).
Sources: Icons from the Noun Project: Structure by Adam Baihaqi from NounProject.com; Metadata by M. Oki Orlando; Data Management by ProSymbols; Logos: used with permission by the copyright holders.
4.1. Data organization and standards
Neuroimaging experiments result in complicated data that can be arranged in many different ways. Historically, data were organized differently between institutions and within labs. This lack of consensus (or a standard) could lead to misunderstandings and suboptimal usage of various resources: human (e.g., time wasted on rearranging data or rewriting scripts expecting certain structure), infrastructure (e.g., data storage space, duplicates), and financial (e.g., disorganized data have limited longevity and value after first publication, because it is hard or even impossible for other researchers to understand and use them). Finally, and most importantly, it produces poor reproducibility of results, even within the lab where data were collected, because it is more likely to include errors and less likely to be accessible to future lab members (or even to the original researcher who obtained the dataset, months or years after they worked on it). Therefore, the need for a data standard in the neuroimaging community became essential.
The Brain Imaging Data Structure (BIDS) is a community-led standard for organizing, describing, and sharing neuroimaging data [RRID:SCR_016124]. BIDS is an evolving standard, which supports multiple neuroimaging modalities including MRI (Gorgolewski et al., 2016), quantitative MRI (Karakuzu et al., 2021), MEG (Niso et al., 2018), EEG (Pernet et al., 2019), intracranial EEG (Holdgraf et al., 2019), PET (Norgaard et al., 2022), Microscopy (Bourget et al., 2022), and imaging genetics (Moreau et al., 2020). Many more extensions are under active development, for example, fNIRS, motion capture, and animal neurophysiology. The BIDS specification documents how to organize the data, generally based on simple file formats (such as NIfTI for tomographic data (Cox et al., 2004), and JSON for metadata) and folder structures. This specification can be extended through community-driven processes to incorporate new neuroimaging modalities or sets of data types.
Multiple applications and tools have been released to make it easy for researchers to incorporate BIDS into their current workflows, maximizing reproducibility, enabling effective data sharing, and supporting good data management practices. For example, BIDS converters make it easier to convert data into BIDS format (e.g., MNE-BIDS (Appelhoff et al., 2019) for MEG and EEG, dcm2bids, ReproNim’s HeuDiConv (Halchenko et al., 2021) and ReproIn (Visconti di Oleggio Castello et al., 2020) for MRI and PET2BIDS for PET; see many more on Table S1). The BIDS validator can help researchers make sure their dataset is BIDS-valid following conversion.
Once data are in BIDS, tools are available to ease interaction with the data. Two commonly used software packages are PyBIDS (Yarkoni et al., 2019), and BIDS-Matlab (Gau et al., 2022). These tools facilitate useful dataset queries—such as how many participants are part of a dataset or what tasks were performed— as well as programmatically retrieving specific files—such as all functional runs for a specific subject. Finally, BIDS apps are containerized analysis pipelines that use full BIDS datasets as their input and produce derivative data (Gorgolewski et al., 2017). Examples of BIDS apps include MRIQC (Esteban et al., 2017) for MRI quality control, fMRIPrep (Esteban et al., 2019) for fMRI preprocessing, and PyMVPA (Hanke et al., 2009) for statistical learning analyses of large datasets (see more at Table S1).
BIDS is a community-led standard and strives to be open and inclusive. The BIDS specification is the result of the ongoing collaboration, shared knowledge, discussion, and consensus through the email discussion list, shared Google docs, and GitHub. Questions are also answered on the Neurostars forum and the Brainhack Mattermost channel. BIDS has a well-specified governance structure where everybody is welcome to participate (see BIDS Code of Conduct, Table S1), and the BIDS Starter Kit is a growing resource intended to simplify the learning process for newcomers.
4.2. Metadata and data annotation
Metadata and data annotation induces consistency and facilitates data replication and reuse. It improves the clarity of the dataset, the ability for collaborators to understand the conditions in which the data were collected, and the ability to effectively share and reuse them. Commonly, metadata files are data dictionaries that map key terms from an agreed-upon vocabulary to data values that contain detailed and standardized information about the key terms. For example, a key called “SampleFrequency” might map to a numerical value, or a key “TaskDescription” might map to a free-form text that describes the task used in a specific experiment. The BIDS standard has proposed a consistent metadata structure in its specification along with a set of specification terms and tags.
Data annotation is also crucial for most data analyses in neuroimaging. For example, when analyzing task-based data, an experiment’s reproducibility is largely determined by the extent to which events are clearly documented. Beyond reproducing previous findings, exhaustively annotated events can allow researchers to re-use the data for means that were originally not thought of during data collection (Bigdely-Shamlo et al., 2020). However, even if each study is fully annotated, without a standard to consistently describe facets of events, all annotations will remain cumbersome and error-prone to work with, and achieving a state of machine readability will require effortful labor.
To address this problem, the Hierarchical Event Descriptor (HED) standard has been continuously developed over the past years (Robbins et al., 2021; Robbins et al., 2021). Drawing on a set of hierarchical vocabulary structures (the HED base schema) and application rules, the HED standard allows for both human- and machine readability, validation, and search of annotations across studies. HED is also fully integrated with the BIDS standard (see Section 4.1), and can be extended by researcher supplied schemas.
Additionally, the Neuroimaging Data Model (NIDM; Maumet et al., 2016; Keator et al., 2013) effort aims to build a core structure for neuroscience datasets to improve searching across publicly-available datasets. The initiative also provides tools to create and use NIDM documents from BIDS datasets (Appelhoff et al., 2019). To effectively describe neuroscience data, well-developed community-driven vocabularies are needed. NIDM is built using semantic web techniques and builds off the PROV (provenance) vocabulary (Moreau et al., 2015). Moreover, the NIDM-Terms effort has begun to collect and extend sets of community-developed controlled vocabularies and techniques for associating concepts with selected study variables of publicly-available neuroimaging datasets (e.g., OpenNeuro, ABIDE, ADHD200, and CoRR). This keeps a registry of the domain-relevant vocabularies and concepts used to annotate datasets, further facilitating concept reuse, and improved inter-dataset search. The NIDM team has developed a JavaScript web application, as well as Python-based command line annotation tools, that allow researchers to annotate their BIDS structured datasets and single tabular files (e.g., csv and tsv spreadsheets), and export BIDS JSON-formatted data dictionaries, NIDM JSON-LD data dictionaries, and NIDM semantic web documents, into sidecar files that accompany the data files. Currently, the NIDM-Terms annotation tools allow researchers to associate their study variables with concepts available in the Cognitive Atlas (Poldrack et al., 2011), the InterLex information resource, and those in the canonical NIDM terminology/ontology as well as encourage them to add descriptive information to improve the clarity of their variables. Such an effort harmonizes and improves the consistency of neuroimaging data and thus makes querying across neuroimaging datasets more efficient.
4.3. Data management and tracking
Raw data and derivatives (outputs from processed data) form the basis for scientific analyses and insights. Being able to efficiently store, retrieve, and update data, derivatives, and metadata across a variety of available storage options is crucial to enable further research (Borghi and Van Gulick, 2021b). As files change and evolve over the course of a project, there is a need to identify which data have been used in the generation of a result, and, in case the data were subject to change or updates, which exact version of the data has been used. The ability to manage data and metadata and track the data-analysis process provides a basis for rigor and reproducibility.
DataLad (Halchenko et al., 2021) is an open-source, community-developed, general purpose tool for managing and version controlling digital files in a decentralized manner. It tracks data of any type or size in a scalable, Git-repository-based overlay structure, called the dataset (practically, a structure of folders and files). DataLad allows tracking data and metadata files stored on local devices as well as remote or cloud infrastructure. DataLad can retrieve public data from major providers such as OpenNeuro, the Canadian Open Neuroscience Platform, the International Neuroimaging Data-sharing Initiative, the Healthy Brain Network Serial Scanning Initiative, Data sharing for Collaborative Research in Computational Neuroscience, the Human Connectome Project’s open access dataset (Van Essen et al., 2013), and many more. Beyond public data, with appropriate permissions or authentication, it can retrieve data from web-based storage providers including major cloud storage services, and local and remote paths (Halchenko et al., 2021; Hanke et al., 2021). DataLad implements this decentralized data management functionality in order to ensure streamlined access to tracked data regardless of hosting service, and to expose datasets for easy access on repository hosting structure. It separates management of file content from lean metadata management by tracking pointers to the services that host managed files (i.e., local infrastructure, remote hosting services, or multiple storage solutions at once). Using these pointers, it enables streamlined on-demand file retrieval in uniquely identified versions from the registered source. Importantly, data retrieval works via streamlined commands regardless of where the data are hosted. Information about DataLad can be found in the DataLad Handbook (Wagner et al. 2021, see Table S1). Entire computing environments could be efficiently managed in DataLad using datalad-container extension (Meyer et al., 2021) developed in collaboration between DataLad and ReproNim projects.
Brainlife is another open science project that allows data management. Brainlife is a free and open community-oriented, non-commercial cloud platform that provides web services to support reproducible data management and analysis. Brainlife tracks data provenance automatically for the users. As data are analyzed using the Graphical User Interfaces (GUI) and the platform’s data processing applications, provenance metadata information is automatically generated and associated with the data derivatives. The users do not have to manually save data versions, the platform does that automatically and it allows visualizing data provenance graphs.
DataLad and Brainlife are synergistic but not overlapping projects that address different user bases and needs. Indeed, DataLad and Brain-life interact nicely with one another and all published datasets retrieved by DataLad are readily accessible at brainlife.io/datasets.
5. Data processing and analysis
Researchers typically execute a set of signal pre-processing steps prior to advanced data analysis, to, for instance, identify and remove noise, align data spatially and temporally, segment spatio-temporal regions of interest, identify patterns and latent signal structures (e.g., clustering), integrate the information from several modalities, introduce prior knowledge about the device or the physiology of the specimen, etc. The combination of the operations that take the unprocessed data as the input, prepare the data for analysis, and finally, perform advanced analysis, comprise a full analysis pipeline or workflow. In implementing such analysis workflows, software has emerged as a critical research instrument greatly relevant to ensure the reproducibility of studies (see Fig. 4).
Fig. 4.
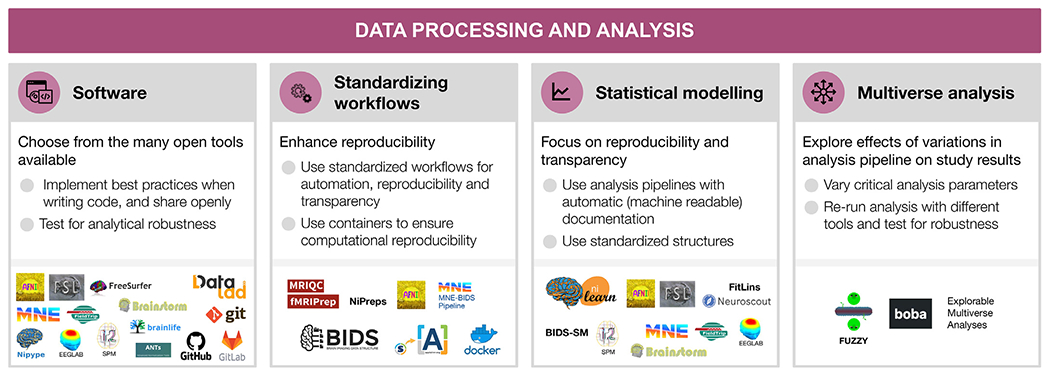
Data processing and analysis
For each step, the figure contains the main goals (headings), specific recommendations (bullet list), and useful tools (icons).
Sources: Icons from the Noun Project: Software by Adrien Coquet; Workflow by D. Sahua; Statistics by Creative Stall; Chaos Sigil by Avana Vana; Logos: used with permission by the copyright holders.
5.1. Software as a research instrument
The digital nature of neuroimaging data along with the large, and constantly increasing, net amounts of daily acquired data, place software as a central instrument of the neuroimaging research workflow. As a result, many toolboxes containing utilities ranging from early steps of preprocessing to statistical analysis and visualization of results have emerged, and some have largely shaped the software development in the field, e.g., AFNI (Cox, 1996; Cox and Hyde, 1997), FSL (Jenkinson et al., 2012), SPM (Penny et al., 2011; Litvak et al., 2011; Flandin and Friston, 2008), FreeSurfer (Dale et al., 1999; Dale and Sereno, 1993), Brainstorm (Tadel et al., 2011, 2019), EEGLAB (Delorme and Makeig, 2004; Delorme et al., 2021), MNE-Python (Gramfort et al., 2013, 2014), Field-Trip (Oostenveld et al., 2011) (see Table S1). More recently, some software packages have been developed to cover additional aspects of the neuroimaging workflow. For instance, nibabel (Brett et al., 2020) to read and write images in many formats, the Advanced Normalization Tools (ANTs) for image registration and segmentation, or Nilearn (Abraham et al., 2014) for statistical analysis and visualization. Workflow engines conveniently connect between the building blocks and determine how the steps are executed in the computational environment. Solutions range from general-purpose scripting (e.g., Bash or Python) to neuroimaging-specific libraries (e.g., NiPype; Gorgolewski et al. 2011). Researchers have all these tools (and others) at their disposal to “mix-and-match” in their workflow. Therefore, ensuring the proper development and operation of the software engine is critical to ensure the reproducibility of results (Tustison et al., 2013).
Relatedly, the variety of software implementations is an additional motive of concern. As remarked by Carp (2012a, 2012b) based on the analysis of thousands of fMRI pipelines, analytical flexibility in combination with incomplete reporting precludes the reproducibility of the results. A recent comprehensive investigation, the Neuroimaging Analysis Replication and Prediction Study (NARPS; Botvinik-Nezer et al. 2020), found that when 70 different teams were asked to analyze the same fMRI data to test the same hypotheses, each team chose a distinct pipeline and results were highly variable. Other studies suggest similar problems in EEG (Šoškić et al., 2021; Clayson et al., 2021), PET (Nørgaard et al., 2020)and diffusion MRI (Schilling et al., 2021).
There are two crucial aspects of the high analytical variability and its effect on results in neuroimaging. First, when high analytical variability (that potentially affects results) is combined with partial reporting or with incentives to find significant effects, it can alarmingly undermine the reliability and reproducibility of results. Second, even in the apparently ideal scenario in which the researcher performs a single pre-registered valid analysis and reports it fully and transparently, it is still likely that the results are not robust to arbitrary analytical choices. Therefore, new tools are needed to allow researchers to perform a “multiverse analysis” (Section 5.4), where multiple data workflows are used on the same dataset and all the results are reported and their agreement or convergence discussed. Community-led efforts to develop high-quality “gold standard” workflows may also reduce researchers’ degrees of freedom as well as accelerate data analysis, although different pipelines may be optimal for different research questions and data.
Nevertheless, neuroimaging researchers frequently encounter gaps that readily available toolboxes do not cover. These gaps, amongst a number of other reasons (e.g., deploying a data workflow on a high-performance computer), pushes researchers into creating their own software implementations. However, most neuroimaging researchers are not formally trained in related fields of computer science, data science, or software engineering, and formal software development practices are often not included in undergraduate or graduate level neuroimaging training. This mismatch often results in undocumented, hard to maintain, and disorganized code; largely as a consequence of unawareness of software development practices. It also increases the likelihood of undetected errors that may remain even after running tests on the code.
The first and foremost strategy available to maximize the transparency of research methods is openly sharing the code with the minimal restrictions possible (see Section 6.2; Barnes, 2010; Gorgolewski and Poldrack, 2016). Complementarily, version control systems, such as Git (Blischak et al., 2016, see Table S1), are the most basic and effective tool to track how software is developed, and to collaboratively produce code. Beyond making the code available to others, software tools can implement further transparency strategies by thoroughly documenting their tools and by supporting implementations with scientific publications (Barnes, 2010; Gorgolewski and Poldrack, 2016).
5.2. Standardizing preprocessing and workflows
Although the diversity in methodological alternatives has been key to extracting scientific insights from neuroimaging data, appropriately combining heterogeneous tools into complete workflows requires substantial expertise. Traditionally, researchers used default workflows distributed along with individual software packages, or alternatively, individual laboratories have developed in-house analysis workflows that resulted in highly specialized pipelines. Such pipelines are often not thoroughly validated and difficult to reuse due to lack of documentation or accessibility to outside labs. In response, several community-led efforts have spearheaded the development of robust, standardized workflows.
An early effort towards workflow standardization was the Configurable Pipeline for the Analysis of Connectomes (C-PAC; Craddock et al. 2013), which is a “nose-to-tail” preprocessing and analysis pipeline for resting state fMRI. C-PAC offers a comprehensive configuration file, editable directly with a text editor or through C-PAC’s graphical user-interface, prescribing all the tools and parameters to be executed, and thereby making strides towards keeping methodological decisions closely traced. Similarly, large-scale acquisition initiatives released workflows tailored for their official imaging protocols (e.g., the HCP Pipelines Glasser et al. 2013 and the UK Biobank Alfaro-Almagro et al. 2016).
Conversely, fMRIPrep (Esteban et al., 2019) proposed the alternative approach of restricting the pipeline goals to the preprocessing step, while accepting the maximum diversity possible of the input data (i.e., not tailored to a particular experimental design or analysis-agnosticity). This approach has recently been proposed for additional modalities (e.g., dMRI, ASL, PET) and population/species of interest (e.g., fMRIPrep-rodents, fMRIPrep-infants) under a common framework called NiPreps (Neuroimaging PREProcessing toolS). NiPreps is a community-led endeavor with the goal of ensuring the generalization of the building blocks of preprocessing across modalities (e.g., the alignment of fMRI and dMRI with the same participant / animal’s anatomical image) and specimens (e.g., using the same brain extraction from anatomical data using the same algorithm and implementation on both human adults and rodents). Similar standardization efforts are starting to be adopted for EEG (Desjardins et al., 2021) and MEG (e.g., MNE-BIDS pipeline; Jas et al., 2018). Further examples of standardized workflows are found in Table S1.
An additional and relevant premise of standardized workflows is transparency — tools must be transparent not only in their implementation, but also in their reporting. For example, fMRIPrep produces visual reports with the double goal of assessing the quality of results, and also providing the researcher with a resource to comprehensively understand every step of the workflow. In addition, the report includes a text description which comprehensively describes each major step in the pipeline, including the exact software version and principle citation. This text, referred to as the “citation boilerplate”, is released under a public domain license, and therefore can be included verbatim in researcher’s manuscripts, facilitating accurate reporting and proper referencing of academic software. A final relevant aspect towards transparency is the comprehensive documentation of pipelines.
In most cases, standardized workflows preprocess datasets in a fully automated manner, taking a BIDS dataset as input and outputting data that is ready for subsequent analysis with little manual intervention. Importantly, such workflows are typically designed to be as robust as possible to diverse input data (e.g., with varying parameters or sampling distant populations), a challenge that is facilitated by data standardization (i.e., BIDS). Additionally, workflows must be portable, enabling users to execute them in a wide variety of environments. A key technology in this endeavor is containers—such as Docker and Apptainer/Singularity—which facilitate packaging specific versions of heterogeneous dependencies while ensuring cross-platform compatibility (e.g., high-performance computing clusters, desktop, or cloud services). The BIDS apps framework (Section 4.1) leverages containers by standardizing input parameters to make it trivially easy to execute a wide variety of standardized workflows on BIDS datasets. An example of a higher-level combination of workflows is found in Esteban et al. (2020), which describes an MRI research protocol using MRIQC and fMRIPrep. Finally, recent efforts to standardize the outputs of workflows (BIDS Derivatives), further enhances the interoperability of workflows, by ensuring their outputs are compatible with subsequent analysis.
5.3. Statistical modeling and advanced analysis
Analysis of neuroimaging data is particularly heterogeneous and prone to excessive analytical flexibility and underspecified reporting (Carp 2012a,2012b). Whereas preprocessing is ideally performed once per dataset, there is often a large number of types of analyses that may be used with the preprocessed data. In MRI and fNIRS, for example, analyses range from multi-stage general linear models (GLMs), multivariable decoding analyses, to anatomical and functional connectivity, and more. In PET, analyses consist of region-wise averaging, although voxel-wise approaches are gaining popularity, followed by kinetic modeling and subsequent statistical analyses, which can be GLM or more advanced, such as latent variable models. In MEG and EEG, the broad variety includes analyses such as evoked response potentials, power spectral density, source reconstructions, time-frequency, connectivity, advanced statistics and more. Each type of analysis also has a wide variety of subtypes, parameters, and statistical models that can be specified, and the form of that specification varies across the dozens of analysis packages that implement each type of analysis.
Data analysis reporting may be made more transparent by sharing code that relies on open-source software. A prime example is SPM (Flandin and Friston, 2008), which has been open source since its inception in 1991. Additional widely used open-source tools for data analysis are FSL and AFNI for MRI, and some examples of reproducible pipelines for MEG and EEG developed based on each of the following software: EEGLAB (Pernet et al. 2020) Fieldtrip (Andersen, 2018b; Meyer et al., 2021; Popov et al., 2018), Brainstorm (Niso et al., 2019; Tadel et al., 2019), SPM (Henson et al., 2019) and MNE-Python (Andersen 2018a; van Vliet et al., 2018; Jas et al., 2018) (see Niso et al. 2022 for a detailed review on main EEG and MEG open toolboxes and reproducible pipelines). Reproducibility is also improved when relying on modular and well-documented software such as Nilearn, which offers versatile methods to perform advanced analyses of fMRI data, from GLM to connectomic and machine learning (Abraham et al., 2014). Ideally, a single analysis script is created, from signal extraction, data analysis, and reproducing all figures.
An additional challenge for the reproducibility of analysis workflows is the representation of statistical models across distinct implementations of analysis software. For example, GLM approaches to analyze fMRI time series are prevalent and supported by all of the major statistical packages (e.g., AFNI, SPM, FSL, Nilearn). However, specifying equivalent models across packages is non-trivial and requires time consuming package specific model specification (Pernet, 2014), which obfuscates details of the statistical model, exacerbates variability across pipelines, and makes it difficult to perform multiverse analyses (see Section 5.4). The BIDS Stats Model (BIDS-SM, see Table S1) specification has been proposed as a implementation-independent representation of fMRI GLM models. BIDS-SM describes the inputs, steps, and specification details of GLM-type analyses, and encodes them in a machine readable JSON format. The PyBIDS library provides tooling to facilitate reading BIDS-SM, and FitLins (Markiewicz et al., 2021) is a reference workflow that fits BIDS-SM using AFNI or Nilearn. The transformative potential of BIDS-SM is showcased by Neuroscout (de la Vega et al., 2022), a turnkey platform for fast and flexible neuroimaging analysis. Neuroscout provides a user-friendly web application for creating BIDS-SM on a curated set of public neuroimaging datasets, and leverages FitLins to fit statistical models in a fully reproducible and portable workflow. By standardizing the entire process of statistical modeling, users can formally specify a hypothesis and produce statistical results in a matter of minutes, while simultaneously ensuring a fully reproducible and transparent analysis that can be readily disseminated to the scientific community.
5.4. Multiverse analysis
The variety of data workflows reflects the enormous interest and the need for novel software instruments, but it also poses an important risk to reproducibility. The multitude of possible combinations of methods and parameters in each of the analysis steps creates an extremely large number of combinations to select from. This problem is often referred to as “researcher degrees of freedom” or “the garden of forking paths” (Gelman and Loken, 2013). Importantly, analytical choices affect results. This has been shown for preprocessing of fMRI data (Strother et al. 2004; Churchill et al., 2012; Churchill et al., 2012). While this work focused mainly on the aspect of tailoring preprocessing to e.g. maximize predictive models, recent efforts in fMRI (task fMRI: Botvinik-Nezer et al., 2020; Carp, 2012a; preprocessing of resting-state fMRI: Li et al. 2021) and PET (specifically for preprocessing: Nørgaard et al. 2020) focused more on the variability of outcomes in general when analysis pipelines were varied. In addition, recent studies showed high variability in diffusion-based tractography dissection (Schilling et al., 2021) and event-related potentials in EEG preprocessing (Šoškić et al., 2021; Clayson et al., 2021). Another large-scale attempt to estimate the analytical variability for EEG, EEGManyPipelines (see Table S1), is currently ongoing.
The converging findings of these studies across modalities suggest that it is crucial to test the robustness of reported results to specific analytical choices. One proposed solution to tackle the analytical variability, where many different analytical approaches are compared, is multiverse analysis (Hall et al., 2022). There are two broad types of multiverse tools. In a “numerical instabilities” approach, different setups and numerical errors or uncertainties in computational tools are evaluated, analyses are rerun several times, and variability, robustness, and “mean answer” are estimated (Kiar et al., 2020). One tool of this type that is being developed is “Fuzzy” (Kiar et al., 2021). Alternatively, in a “classic multiverse analysis”, multiple pipelines are used with the same data and the results are compared across pipelines. Such an analysis could be conducted by a single or by multiple researchers (Aczel et al. 2021). Although multiverse analysis was suggested before in other fields (Simonsohn et al., 2020; Steegen et al., 2016; Simonsohn et al., 2015; Patel et al., 2015), there are not yet mature “classic multiverse analysis” tools for high-dimensional data like in neuroimaging. Explorable Multiverse Analyses is an R-tool that allows the readers to explore different statistical approaches in a paper (Dragicevic et al., 2019). Other tools, such as the Python-based Boba (Liu et al., 2021), aim to facilitate multiverse analyses by allowing users to specify the shared and the varying parts of the code only once and by providing useful visualizations of the pipelines and results. However, these tools currently fit simpler analyses and datasets compared to the ones common in neuroimaging.
In neuroimaging, recent progress has been made in creating infrastructure for multiverse analysis in fMRI, based on the C-PAC tool (see Section 5.2; Li et al. 2021). Ongoing efforts to formalize machine-readable standards for statistical models (BIDS-SM) and pipelines to estimate them (e.g., FitLins; Markiewicz et al., 2021), and their integration with datasets using platforms such as Brainlife (Avesani et al., 2019), could facilitate the development of multiverse tools. In order to make sense of a multiverse analysis, one needs methods to test for convergence across results of diverse analysis pipelines with the same data. Such a method for fMRI image-based meta-analysis was recently used in NARPS (Botvinik-Nezer et al., 2020) as well as in subsequent projects (Bowring et al., 2021). Another simple statistical approach to a multiverse analysis was presented with PET data (Nørgaard et al., 2019), although it lacks statistical power, due to the use of a very conservative statistic. A different approach is to use active learning to approximate the whole multiverse space (Dafflon et al., 2020). Moreover, Boos et al. (2021) provided an online application to explore the effects of the choice of parameters on the results (data-driven auditory encoding, see Table S1). Progress is still needed until such tools are mature enough to allow scalable multiverse analysis in neuroimaging.
6. Research dissemination
Through the whole research cycle a range of outputs far beyond publications are produced, and each of them can have different levels of reproducibility and openness (see Fig. 5). For shared resources to be useful, they need to follow the FAIR principles (Wilkinson et al., 2016), to ensure they are: Findable (e.g., using persistent identifiers, such as Digital Object Identifiers (DOI) or Research Resource Identifiers (RRIDs), and described with rich metadata indexed in a searchable resource), Accessible (e.g., shared in public repositories, under open access or controlled access depending on regulations, so they can be retrievable by their identifier using standardized communication protocols), Interoperable (e.g., following a common standard for organization and vocabulary), and Reusable (e.g., richly described, with detailed provenance and an appropriate license). Indeed, without a license, materials (data, code, etc.) become unusable by the community due to the lack of permission and conditions for reuse, copy, modification, or distribution. Therefore, consenting through a license is essential for any material to be publicly shared.
A useful generalpurpose resource, beyond neuroimaging, including practical guidelines on reproducible research, project design, communication, collaboration, and ethics is The Turing Way (TTW, The Turing Way Community et al. 2019, see Table S1). TTW is an open collaborative community-driven project, aiming to make data science accessible and comprehensible to ensure more reproducible and reusable projects.
6.1. Data sharing
Making data available to the community is important for reproducibility, allows more scientific knowledge to be obtained from the same number of participants (animal or human), and also enables scientists to learn and teach others to reuse data, develop new analysis techniques, advance scientific hypotheses, and combine data in mega- or meta-analyses (Poldrack and Gorgolewski, 2014; Laird, 2021; Madan, 2021). Moreover, the willingness to share has been shown to be positively related to the quality of the study (Wicherts et al., 2011). Because of the many advantages data sharing brings to the scientific community (Milham et al. 2018), more and more journals and funding agencies are requiring scientists to make their data public (curated and archived with a public record, but controlled access) or open (public data with uncontrolled access) upon the completion of the study, as long as it does not compromise participants’ privacy, legal regulations, or the ethical agreement between the researcher and participants (see Sections 2.3 and 7).
For data to be interoperable and reusable, it should be organized following an accepted standard, such as BIDS (Section 4.1) and with at least a minimal set of metadata. Free data-sharing platforms are available for publicly sharing neuroimaging data, such as OpenNeuro (Markiewicz et al., 2021), Brainlife (Avesani et al., 2019), GIN (G-Node Infrastructure), Ebrains, Distributed Archives for Neurophysiology Data Integration (DANDI), International Neuroimaging Data-Sharing Initiative (INDI), Neuroimaging Tools & Resources Collaborator (NITRC), etc. (see Table S1). Data could also be shared on institutional and funder archives such as the National Institute of Mental Health Data Archive (NDA); on dedicated repositories, such as the the Cambridge Centre for Ageing and Neuroscience, Cam-CAN (Shafto et al., 2014; Taylor et al., 2017) or The Open MEG Archive, OMEGA (Niso et al., 2016); or on generic archives that are not neuroscience or neuroimaging specific, such as figshare, GitHub, the Open Science Framework, and Zenodo. If allowed by the law and participants’ consent (see Section 2.3), data sharing can be made open, or at least public.
Once curated and archived, data can further benefit the individual researcher, for example by adding them to the scientific literature in the form of data descriptors. Such an article type is not intended to communicate findings or interpretations but rather to provide detailed information about how the dataset was collected, what it includes and how it could be used, along with the shared data. In addition, an “open science badge” for data sharing is available in an increasing number of scientific journals (Kidwell et al., 2016) and some prizes are also available as a recognition for such efforts (e.g., OHBM’s Data Sharing prize).
It is important to note that there are unresolved issues with international data sharing that some researchers should consider before sharing their neuroimaging data. First, privacy regulations can differ tremendously between cultural, legal, and ethical regions, and these differences have an impact on whether certain data can be shared (e.g., unprocessed MRI images) and if so, under which restrictions (e.g. openly or after signing a data user agreement). Data sharing platforms vary in physical location and access policies, adding complexity to the choice of site. There is an ongoing discussion of these issues (see e.g., Jwa and Poldrack, 2022; Eke et al., 2022) and solutions are under development, for instance via the EBRAINs infrastructure (Amunts et al., 2019, 2016). It can be expected that data sharing procedures will undergo further transformations as privacy laws in some jurisdictions shift towards GDPR-type laws, and more adequacy decisions will be made by the EU Commission (e.g. the Consumer Privacy Protection Act in Canada, or the California Consumer Privacy Act). Second, it is unclear how researchers are properly credited for data they collected and shared. Sharing data with a DOI, or as a data paper when appropriate, allows the researchers to receive some academic credit via citations.
6.2. Methodological transparency
Documenting performed analysis steps is key for reproducing studies’ results. For studies containing a small number of procedures, the methods section of an article could detail them in full length. This is, however, often not the case in current neuroimaging studies, where authors may need to summarize content in order to fit into the designated space, likely omitting relevant details. Therefore, the programming code itself becomes the most accurate source of the exact analysis steps performed on the data, and is anyway needed for reproducibility. Thus, it needs to be organized and clear (see recommendations suggested by Sandve et al. 2013; van Vliet 2020; Wilson et al. 2017), otherwise, results may not be reproducible, or even correct (Casadevall and Fang, 2014; Pavlov et al., 2021). It should be noted that sharing an imperfect code is still much better than not sharing at all (Barnes, 2010; Gorgolewski and Poldrack, 2016).
While GUIs are a great and interactive way to learn to analyze data, one has to pay special attention to properly report the steps followed, as manual operations are more difficult to be reported and reproduced than automated code (Pernet and Poline, 2015). Some toolboxes using GUIs keep track of data ‘history’ and ‘provenance’ (e.g., Brainlife, Brainstorm, EEGLAB, FSL’s Feat tool) facilitating this task. Efforts are being put to improve these features.
To ensure long-term preservation of the shared code, we suggest using version control systems such as Git, and social coding platforms such as GitHub in combination with an archival database for assigning permanent DOI to code served for research, for instance, Zenodo (Troupin et al., 2018), brainlife.io/apps (Avesani et al., 2019) or Software Heritage (Di Cosmo, 2018). These platforms help keep a snapshot of the version of the code used for the paper published, allowing exact reproduction in case of later code updates.
6.3. Derived data sharing
Sharing data derivatives is perhaps the most critical yet most challenging aspect of data sharing in support of reproducible science. The BIDS standard provides a general description of common derivative data (e.g., preprocessed data and statistical maps) and is actively working towards extending advanced specific derivatives for the different neuroimaging modalities. Yet, standards for the description of advanced derivatives (such as activation or connectivity maps, or diffusion measures) are currently not available or mature for wide use. As a result, to date, the community lacks clear guidance and tools on how derived data should be organized to maximize its reuse and to encompass its provenance, and where such data could be shared.
Solutions for sharing derivatives comprise a mixture of in-house and semi-standardized data-tracking and representation methods. Examples of data-derivative sharing are the high-profile, centralized projects, such as the Human Connectome Projects (Van Essen et al., 2012), Ebrains of the Human Brain Project (Amunts et al., 2019, 2016), the UK-Biobank (Alfaro-Almagro et al., 2016), the NKI-Rockland sample (Nooner et al., 2012), and the Adolescent Brain Cognitive Development (ABCD; Feldstein Ewing and Luciana, 2018) to name a few (see Table S1). These projects have developed project-specific solutions and in doing so also have provided a first-level implementation of what could be considered a data derivative standard. Yet, these projects are far from being open or community-developed as they must be centrally governed and mandated by the directives of the research plan.
As a result of the paucity of community-oriented standards, archives and software methods, sharing highly-processed neuroimaging data is still the frontier of reproducible science. One community-open archive for highly processed neuroimaging data derivatives is NeuroVault (Gorgolewski et al., 2015). The archive accepts brain statistical maps derived from fMRI with the goal of being reused for meta-analytic studies. Data upload is open to researchers world-wide and the archive can accept brain maps submitted using most major formats but preferably using the NeuroImaging Data Model (NIDM; see Section 4.2). Another interesting example for automated and standardized composition and sharing of derived data is TemplateFlow (Ciric et al., 2021), which provides an open and distributed framework for establishment, access, management, and vetting of reference anatomies and atlases.
Another general neuroimaging platform, which allows lowering the barrier to sharing highly processed data derivatives, is Brainlife. Brainlife provides methods for publishing derivatives integrated with the data-processing applications used for generating results from the data via easy-to-use web interfaces for data-upload, processing, and publishing. Different licenses can be selected when publishing a record, allowing reuse of data and derivatives to other researchers (see sources that support the selection of an appropriate license listed in Table S1).
Sharing lighter-weight data products such as tables and figures is easier using generic repositories (e.g., OSF, Figshare, Github or Zenodo) under, for example, CC-BY license. This allows authors to retain rights on the figures they created, and others to re-use their figures, either in other publications or for educational purposes, while giving credit to the originating team. Additional material, such as slide presentations, or supporting content, should be shared using accessible formats (e.g., image files, pdf, PowerPoint slides, Markdown, jupyter notebooks, etc) via online repositories or institutional platforms, with appropriate licenses to indicate how the work could be reused. Whenever possible, using platforms that ensure long-term preservation is recommended (e.g., Zenodo, FigShare, OSF). Using platforms that provide a DOI is particularly encouraged, because it ensures that the shared data would be identifiable in the future.
6.4. Publication of scientific results
Scientific papers are currently the most important means for disseminating research results. However, they should also be written with reproducibility in mind. Guidelines to improve reproducibility can support the writing. The OHBM Committee on Best Practices in Data Analysis and Sharing (COBIDAS), has been promoting best practices, including open science. Recommendations from the committees for MRI (Nichols et al., 2017) and MEG and EEG (Pernet et al., 2020) provide guidance on what to report. Other recent community efforts also led to guidelines for PET (Knudsen et al., 2020) and EEG reporting (e.g., Agreed Reporting Template for EEG Methodology - International Standard (ARTEM-IS) Styles et al. 2021). One tool that could help authors follow these guidelines while writing their report are their web-based apps (see Table S1). For the data description and preprocessing aspects, some tools (pyBIDS, bids-matlab) or pipelines (fMRIPrep) can also generate reports automatically, and/or method templates are provided (see Section 5.2). Exact description of methods is mandatory for reproducibility alongside detailed reporting of results.
In recent years, it has become very common in neuroimaging to publish papers as preprints, on servers such as bioRxiv, medRxiv, PsyArXiv or OSF (see Table S1), prior to peer review in scientific journals. Preprints are publicly available, expedite the process of releasing new findings, and also, importantly, allow authors to get feedback on their paper from a broader audience prior to final publication (Moshontz et al., 2021). There are also initiatives for open community reviews of preprints, such as PREreview. Other initiatives have emerged to adapt to this paradigm shift, such as the recently launched Open Europe Research platform for publication and open peer review, which also includes the different outputs obtained throughout the research cycle (e.g., study protocol, data, methods and brief reports through the process) for research stemming from Horizon 2020 and Horizon Europe funding. In addition, novel publication formats have been developed, like NeuroLibre (Karakuzu et al., 2022), a preprint server to publish hybrid research objects including text, code, data, and runtime environment (DuPre et al., 2022). More traditional publishers have successfully partnered with companies such as CodeOcean (Cheifet, 2021) to provide similar services.
Crucially, papers should be accessible to others, preferably to everyone. Many scientific publications are hidden behind paywalls, practically denying access from many people who could have gained from them. This is now slowly changing (Piwowar et al., 2018), with both researchers and funding agencies pushing towards open access, meaning that papers are fully open to all. Although the adoption of the concept of open access by major publishers is in itself a positive development, the way it was adopted could be considered arguable. For example, several journals considerably increased article processing charges (Khoo, 2019; Budzinski et al., 2020), increasingly excluding research produced in labs with lower levels of resources, particularly in Low and Middle Income Countries, from being published open access (Nabyonga-Orem et al., 2020). Additionally, some publishers implement massive tracking technology with the argument to protect their rights and offer the data or derivatives of them for sale, as a recent report published by the German Research Foundation criticizes (DFG, 2021). This raises many questions, related, for example, to the influence publishers and their algorithms will have in the future on strategic decisions of science institutions and freedom of science.
6.5. Beyond publication
The research lifecycle continues beyond paper publication. Disseminating scientific results to the broader scientific community and to the society in general is of utmost importance, to translate the newly acquired knowledge and give back to society. Oral and poster presentations at conferences contribute to the dissemination of results (including preliminary or intermediate results) within the scientific community, and also provide opportunities for feedback prior to publication. Workshops and other educational events contribute to expand the knowledge further and induce new communities. Popular and social media (e.g., press releases, interviews, podcasts, blog posts, twitter, facebook, linkedin, youtube, etc.) may reach an even wider and more heterogeneous audience. Different types of audiences may have different degrees of expertise and scientific knowledge, hence, for an effective communication, each of the outreach events should adapt accordingly (e.g., avoiding jargon and over interpretation, identifying your audience, promoting accessibility in content and language (Amano et al. 2021), and considering disabilities). See the TTW Guide for Communication for recommendations (The Turing Way Community et al., 2019). Slides presentations, and further outreach content should be shared FAIRly for higher impact (see Section 6.3).
Hackathons - such as the Brainhacks in the neuroimaging community (Gau et al., 2021) - typically offer times for “unconferences” in which attendees can propose a short talk to present some work-in-progress, an open question, or any other topic they wish to discuss with other participants. This deviates from more typical conferences in which only well-polished, finalized results can be presented. Other initiatives, such as Neuromatch Academy, Neurohackademy, OHBM Open Science Room, and Brainhack school MLT facilitate open science and provide opportunities for researchers to learn and get hands-on experience with open science practices, and also to engage with other researchers in the community. Those hackathons and the related online communities are also well-known as kick-starters for the development of community tools and standards in which researchers and engineers from different labs join forces. As those tools and standards get shaped, typically in multi-lab collaborations, researchers get the chance to exchange their views and practices. Overall, such events, slowly but surely, help shape a research culture that is driven by open collaborative communities rather than single groups of researchers.
6.6. Towards inclusive, diverse and community driven research
Taken to the next level, the described developments and introduced tools provide an opportunity for a paradigm shift: rather than carrying out a study from inception to results and only then disseminating the findings to the community, researchers now get multiple opportunities to share their ideas and results as they are being developed. Scientific research can now become more transparent, inclusive and collaborative throughout the research cycle.
Inclusivity, in particular, has the potential to increase reproducibility (and more specifically, the generalizability and robustness) and research results quality, by diversifying neuroimaging research, from the participants included in the samples to the views and ideas of the researchers (Henrich et al., 2010; Laird, 2021; Forbes et al., 2021; Hofstra et al., 2020). Publishing Code of Conduct for collaborative projects is one practice that helps ensure a more welcoming and inclusive space for everyone regardless of background or identity. Initiatives such as TTW (The Turing Way Community et al., 2019) or the OHBM Conference have detailed Code of Conducts that can be of inspiration to adapt and use in new collaborative projects. Over the past years, the awareness and number of initiatives to mitigate bias and inequity at individual and institutional levels are growing (Llorens et al., 2021; Levitis et al., 2021; Malkinson et al., 2021; Schreiweis et al., 2019). These aim to produce better research powered by a broader range of perspectives and ideas and to reduce the negative impact on the careers, work-life balance, and mental health of underrepresented groups. By providing open resources and promoting welcoming and inclusive spaces, we are also improving access to the tools, training and infrastructure which can facilitate reproducible research, which will accelerate discoveries, and ultimately, advance science.
7. Conclusions
Recent years have marked the rise of “open science”, producing numerous tools and practices that enhance the reproducibility, transparency, inclusivity and diversity of research in general and in neuroimaging specifically. These tools and practices yield benefits at multiple levels ranging from the individual researcher to the society. At the societal level, they can increase transparency and credibility of research, foster the public understanding of scientific findings, and promote participation. Higher public credibility in research results can support decision-makers in basing their decisions on scientific knowledge. For the scientific community, such practices can increase the quality and generalizability of scientific products. They also increase the cost-effectiveness of invested resources (money, time, personnel, etc.), by, for example, enabling reuse of collected data and developed methods and tools (Milham et al., 2018). Acquiring and analyzing data also has a substantial environmental cost, which can be minimized when research data and products are shared and reused1 https://ohbm-environment.org/. For individual researchers, the application of open science practices can improve their chances for funding and recognition in the community by meeting related requirements from funding institutions, agencies, and scientific journals. Furthermore, the use of open science tools and practices can ease the use of novel analysis techniques and open the researcher new opportunities for collaborations and contributions, which in turn transform the research culture.
In this review we have attempted to comprehensively summarize a broad range of open and reproducible science practices. However, to maximize their impact it is important to position these efforts in the broader scientific reform debate. In particular in psychology, many have argued that the very theories that guide the design of experiments lack rigor (Oberauer and Lewandowsky, 2019), and overreach due to improper use of inferential statistics (Yarkoni, 2022). It has been suggested that a formalization of theoretical models and claims (Lee et al., 2019; Guest and Martin, 2021; Devezer et al., 2021) — including claims made in favor of the open and reproducible practices and tools reviewed here — is critical to truly advance the field. Although these issues require ongoing deep introspection and cannot be solved solely by adopting the practices we reviewed, these practices increase our field’s rigor by helping scientists ensure they achieve their stated standards and constitute first steps towards better formalization of designs, models, and conclusions (e.g. with SOPs, formalized statistical models, and pre-registrations which could lead to better formalization of theory-based experimental designs and predictions).
The abundance of tools and practices for open and reproducible neuroimaging is both promising and challenging. They should support scientific practices rather than setting up new hurdles, for example with exceedingly rigid rules, particularly time consuming processes, or requiring highly developed programming skills. This review was written to assist neuroimaging researchers in making informed and sustainable implementation choices in their own research, by means of understanding the purpose of each tool, how they interact together, how to use them, and where to look for further information. We believe it will prove helpful for researchers and institutions to make a successful and sustainable move towards open and reproducible science, contributing to improving scientific research and, ultimately, accelerating scientific discoveries.
Supplementary Material
Acknowledgments
G.N. was supported by the AXA Research Fund and by NIH CR-CNS: US-France Data Sharing Proposal (NIBIB (USA) R01 EB030896 and ANR-20-NEUC-0004-01). R.B-N. is an Awardee of the Weizmann Institute of Science -Israel National Postdoctoral Award Program for Advancing Women in Science. A.d.l.V was supported by NIH grant 5R01MH109682. O.E. was supported by the Swiss National Science Foundation (SNSF) Project 185872 and NIMH grant (RF1MH121867, O.E.). J.A.E was supported by NIH grant R37MH066078 to Todd S. Braver. K.F. was supported by the Polish National Agency for Academic Exchange (the Bekker programme; PPN/BEK/2020/1/00279/U/00001). M.G. was supported by the Elsass foundation (18-3-0147). Y.O.H. was supported by NIH grant 1P41EB019936-01A1. P.H. was supported in parts by funding from the Canada First Research Excellence Fund, awarded to McGill University for the Healthy Brains for Healthy Lives initiative, the National Institutes of Health (NIH) NIH-NIBIB P41 EB019936 (ReproNim), the National Institute Of Mental Health of the NIH under Award Number R01MH096906, a research scholar award from Brain Canada, in partnership with Health Canada, for the Canadian Open Neuroscience Platform initiative, as well as an Excellence Scholarship from Unifying Neuroscience and Artificial Intelligence - Québec. A.K. was supported by the TransMedTech Institute Postdoc Fellowship. D.B.K. was supported by the National Institute of Mental Health under grant RF1 MH120021 (PI: Keator), and acknowledges the International Neuroinformatics Coordinating Facility (INCF). C.R.P. was supported by the Novo Nordisk Fonden NNF20OC0063277. F.P. is supported by NSF grants IIS 1636893, IIS 1912270, and BCS 1734853, NIH National Institute of Biomedical Imaging and Bioengineering (NIBIB) grant 1R01EB029272, NIH NIMH 1R01MH126699, and a Microsoft Investigator Fellowship. N.Q. acknowledges the NIDM-Terms grant (National Institutes of Mental Health (NIMH) grant 1RF1MH120021 PI: David B Keator). K.J.W. was supported by The UKRI Strategic Priorities Fund under EPSRC Grant EP/T001569/1, particularly the “Tools, Practices and Systems” theme within that grant, and by The Alan Turing Institute under the EPSRC grant EP/N510129/1. J.W.R. was supported by the DFG Device Center Grant INST 184/216-1 “Tools and infrastructure for open and reproducible neuroimaging”, and the DFG grant 390895286 of the Excellence Strategy (EXC 2177/1, Hearing4All).
Footnotes
Declaration of Competing Interest
The authors declare no conflict of interest related to this work.
Credit authorship contribution statement
Guiomar Niso: Conceptualization, Project administration, Supervision, Writing – original draft, Writing – review & editing. Rotem Botvinik-Nezer: Conceptualization, Project administration, Supervision, Writing – original draft, Writing – review & editing. Stefan Appelhoff: Conceptualization, Writing – original draft, Writing – review & editing. Alejandro De La Vega: Conceptualization, Writing – original draft, Writing – review & editing. Oscar Esteban: Conceptualization, Writing – original draft, Writing – review & editing. Joset A. Etzel: Conceptualization, Writing – original draft, Writing – review & editing. Karolina Finc: Writing – original draft, Writing – review & editing. Melanie Ganz: Writing – original draft, Writing – review & editing. Rémi Gau: Writing – original draft, Writing – review & editing. Yaroslav O. Halchenko: Conceptualization, Writing – original draft, Writing – review & editing. Peer Herholz: Conceptualization, Writing – original draft, Writing – review & editing. Agah Karakuzu: Conceptualization, Writing – original draft, Writing – review & editing. David B. Keator: Writing – original draft, Writing – review & editing. Christopher J. Markiewicz: Writing – review & editing. Camille Maumet: Conceptualization, Writing – original draft, Writing – review & editing. Cyril R. Pernet: Conceptualization, Writing – original draft, Writing – review & editing. Franco Pestilli: Conceptualization, Writing – original draft, Writing – review & editing. Nazek Queder: Conceptualization, Writing – original draft, Writing – review & editing. Tina Schmitt: Writing – original draft, Writing – review & editing. Weronika Sójka: Writing – review & editing. Adina S. Wagner: Conceptualization, Writing – original draft, Writing – review & editing. Kirstie J. Whitaker: Writing – review & editing. Jochem W. Rieger: Conceptualization, Project administration, Supervision, Writing – original draft, Writing – review & editing.
Supplementary materials
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.neuroimage.2022.119623.
Data and code availability statement
Not applicable
Data availability
No data was used for the research described in the article.
References
- Abraham A, Pedregosa F, Eickenberg M, Gervais P, Mueller A, Kossaifi J, Gram-fort A, Thirion B, Varoquaux G, 2014. Machine learning for neuroimaging with scikit-learn. Front. Neuroinf 8 (February), 14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aczel B, Szaszi B, Nilsonne G, Akker OR, Albers CJ, Assen MA, Bastiaansen JA, et al. , 2021. Consensus-based guidance for conducting and reporting multi-analyst studies. eLife 10 (November). doi: 10.7554/eLife.72185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alfaro-Almagro F, Jenkinson M, Bangerter N, Andersson J, Griffanti L, and Douaud G. 2016. “UK biobank brain imaging: automated processing pipeline and quality control for 100,000 subjects.” In, 1877.
- Allen C, Mehler DMA, 2019. Open science challenges, benefits and tips in early career and beyond. PLoS Biol. 17 (5), e3000246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amano T, Rojas CR, Boum Ii Y, Calvo M, Misra BB, 2021. Ten tips for overcoming language barriers in science. Nat. Hum. Behav 5 (9), 1119–1122. [DOI] [PubMed] [Google Scholar]
- Amunts K, Ebell C, Muller J, Telefont M, Knoll A, Lippert T, 2016. The human brain project: creating a european research infrastructure to decode the human brain. Neuron 92 (3), 574–581. [DOI] [PubMed] [Google Scholar]
- Amunts K, Knoll AC, Lippert T, Pennartz CMA, Ryvlin P, Destexhe A, Jirsa VK, D’Angelo E, Bjaalie JG, 2019. The human brain project: synergy between neuroscience, computing, informatics, and brain-inspired technologies. PLoS Biol. 17 (7), e3000344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen LM, 2018a. Group analysis in MNE-python of evoked responses from a tactile stimulation paradigm: a pipeline for reproducibility at every step of processing, going from individual sensor space representations to an across-group source space representation. Front. Neurosci doi: 10.3389/fnins.2018.00006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen LM, 2018b. Group analysis in fieldtrip of time-frequency responses: a pipeline for reproducibility at every step of processing, going from individual sensor space representations to an across-group source space representation. Front. Neurosci 12 (May), 261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Appelhoff S, Bates JF, Ghosh S, Keator DB, Kennedy DN, Poldrack R, Poline JB, et al. , 2019. BIDS and the neuroimaging data model (NIDM). F1000Research 8 (1924), 1924. [Google Scholar]
- Appelhoff S, Sanderson M, Brooks T, Vliet M, Quentin R, Holdgraf C, Chaumon M, et al. , 2019. MNE-BIDS: organizing electrophysiological data into the BIDS format and facilitating their analysis. J. Open Source Softw 4 (44), 1896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avesani P, McPherson B, Hayashi S, Caiafa CF., Henschel R, Garyfallidis E, Kitchell L, et al. , 2019. The open diffusion data derivatives, brain data upcycling via integrated publishing of derivatives and reproducible open cloud services. Sci. Data 6 (1), 69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baggio G, Corsini A, Floreani A, Giannini S, Zagonel V, 2013. Gender medicine: a task for the third millennium. Clin. Chem. Lab. Med 51 (4), 713–727 CCLM/FESCC. [DOI] [PubMed] [Google Scholar]
- Barnes N, 2010. Publish your computer code: it is good enough. Nature 467 (7317), 753. [DOI] [PubMed] [Google Scholar]
- Benning SD, Bachrach RL, Smith EA, Freeman AJ, Wright AGC, 2019. The registration continuum in clinical science: a guide toward transparent practices. J. Abnorm. Psychol 128 (6), 528–540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bigdely-Shamlo N, Touryan J, Ojeda A, Kothe C, Mullen T, Robbins K, 2020. Automated EEG mega-analysis I: spectral and amplitude characteristics across studies. Neuroimage 207 (February), 116361. [DOI] [PubMed] [Google Scholar]
- Blischak JD, Davenport ER, Wilson G, 2016. A quick introduction to version control with Git and GitHub. PLoS Comput. Biol 12 (1), e1004668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boos M, Lücke J, Rieger JW, 2021. Generalizable dimensions of human cortical auditory processing of speech in natural soundscapes: a data-driven ultra high field fMRI approach. Neuroimage 237 (August), 118106. [DOI] [PubMed] [Google Scholar]
- Borghi JA, and Van Gulick AE. 2021. “Promoting open science through research data management. ”arXiv. https://arxiv.org/abs/2110.00888.
- Borghi JA, Van Gulick AE, 2018. Data management and sharing in neuroimaging: practices and perceptions of MRI researchers. PLoS One 13 (7), e0200562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borghi JA, Van Gulick AE, 2021b. Data management and sharing: practices and perceptions of psychology researchers. PLoS One 16 (5), e0252047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botvinik-Nezer R, Holzmeister F, Camerer CF, Dreber A, Huber J, Johannesson M, Kirchler M, et al. , 2020. Variability in the analysis of a single neuroimaging dataset by many teams. Nature 582 (7810), 84–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourget MH, Kamentsky L, Ghosh SS, Mazzamuto G, Lazari A, Markiewicz CJ, Oostenveld R, et al. , 2022. Microscopy-BIDS: an extension to the brain imaging data structure for microscopy data. Front. Neurosci 16 (April), 871228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowring A, Nichols TE, and Maumet C. 2021. “Isolating the sources of pipeline-variability in group-level task-fMRI results.”bioRxiv. bioRxiv. doi: 10.1101/2021.07.27.453994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brainard DH, 1997. The psychophysics toolbox. Spat. Vis 10 (4), 433–436. [PubMed] [Google Scholar]
- Brett M, Markiewicz CJ, Hanke M, Côté MA, Cipollini B, McCarthy P, Jarecka D, et al. 2020. “Nipy/nibabel: 3.2.1. ” Zenodo. doi: 10.5281/zenodo.4295521. [DOI] [Google Scholar]
- Breuer FA, Blaimer M, Heidemann RM, Mueller MF, Griswold MA, Jakob PM, 2005. Controlled Aliasing in Parallel Imaging Results in Higher Acceleration (CAIPIRINHA) for multi-slice imaging. Mag. Reson. Med 53 (3), 684–691 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Budzinski O, Grebel T, Wolling J, Zhang X, 2020. Drivers of article processing charges in open access. SSRN Electron. J 124, 2185–2206. [Google Scholar]
- Camerer CF, Dreber A, Forsell E, Ho TH, Huber J, Johannesson M, Kirchler M, et al. , 2016. Evaluating replicability of laboratory experiments in economics. Science 351 (6280), 1433–1436. [DOI] [PubMed] [Google Scholar]
- Camerer CF, Dreber A, Holzmeister F, Ho TH, Huber J, Johannesson M, Kirchler M, et al. , 2018. Evaluating the replicability of social science experiments in nature and science between 2010 and 2015. Nat. Hum. Behav 2 (9), 637–644. [DOI] [PubMed] [Google Scholar]
- Carp J, 2012a. On the plurality of (methodological) worlds: estimating the analytic flexibility of fmri experiments. Front. Neurosci 6 (OCT), 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carp J, 2012b. The secret lives of experiments: methods reporting in the fMRI literature. Neuroimage 63 (1), 289–300. [DOI] [PubMed] [Google Scholar]
- Casadevall A, Fang FC, 2014. Causes for the persistence of impact factor mania. mBio 5 (2), e00064–e00114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers C, 2019. What’s next for registered reports? Nature 573 (7773), 187–189. [DOI] [PubMed] [Google Scholar]
- Cheifet B, 2021. Promoting reproducibility with code ocean. Genom. Biol 22 (1), 65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Churchill NW, Oder A, Abdi H, Tam F, Lee W, Thomas C, Ween JE, Graham SJ, Strother SC, 2012. Optimizing preprocessing and analysis pipelines for single-subject fMRI. I. Standard temporal motion and physiological noise correction methods. Hum. Brain Mapp 33 (3), 609–627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Churchill NW, Yourganov G, Oder A, Tam F, Graham SJ, Strother SC, 2012. Optimizing preprocessing and analysis pipelines for single-subject fMRI: 2. interactions with ICA, PCA, task contrast and inter-subject heterogeneity. PLoS One 7 (2), e31147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciric R, Thompson WH, Lorenz R, Goncalves M, MacNicol E, Markiewicz CJ, Halchenko YO, et al. 2021. “TemplateFlow: FAIR-sharing of multi-scale, multi-species brain models.”bioRxiv. doi: 10.1101/2021.02.10.430678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clayson PE, Baldwin SA, Rocha HA, Larson MJ, 2021. The data-processing multiverse of Event-Related Potentials (ERPs): a roadmap for the optimization and standardization of ERP processing and reduction pipelines. Neuroimage (November), 118712. [DOI] [PubMed] [Google Scholar]
- Clayson PE, Keil A, Larson MJ, 2022. Open science in human electrophysiology. Int. J. Psychophysiol 174 (April), 43–46 Official Journal of the International Organization of Psychophysiology. [DOI] [PubMed] [Google Scholar]
- Connolly A, and Halchenko Y. 2022. ReproNim/reprostim: doi: 10.5281/zenodo.6354036. [DOI]
- Cordes C, Konstandin S, Porter D, Günther M, 2020. Portable and platform-independent MR pulse sequence programs. Mag. Reson. Med 83 (4), 1277–1290 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Cox RW, 1996. AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput. Biomed. Res 29 (29), 162–173. [DOI] [PubMed] [Google Scholar]
- Cox RW, Ashburner J, Breman H, Fissell K, Haselgrove C, Holmes CJ, Lancaster JL, et al. 2004. “A (sort Of) new image data format standard: NiFTI-1.” In. https://nifti.nimh.nih.gov/nifti-1/documentation/hbm_nifti_2004.pdf.
- Cox RW, Hyde JS, 1997. Software tools for analysis and visualization of fMRI data. NMR Biomed. 10 (4-5), 171–178. [DOI] [PubMed] [Google Scholar]
- Craddock C, Sikka S, Cheung B, Khanuja R, Ghosh SS, Yan C, Li Q, et al. , 2013. Towards automated analysis of connectomes: the configurable pipeline for the analysis of connectomes (c-Pac). Front. Neuroinf 42, 10–3389. [Google Scholar]
- Dafflon J, Da Costa PF, Váša F, Monti RP, and Bzdok D. 2020. “Neuroimaging: into the multiverse.”bioRxiv. 10.1101/2020.10.29.359778v1.abstract. [DOI] [Google Scholar]
- Dale AM, Fischl B, Sereno MI, 1999. Cortical surface-based analysis. I. Segmentation and surface reconstruction. Neuroimage 9 (2), 179–194. [DOI] [PubMed] [Google Scholar]
- Dale AM, Sereno MI, 1993. Improved localizadon of cortical activity by combining EEG and MEG with MRI cortical surface reconstruction: a linear approach. J. Cogn. Neurosci 5 (2), 162–176. [DOI] [PubMed] [Google Scholar]
- Delorme A, Makeig S, 2004. EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J. Neurosci. Methods 134 (1), 9–21. [DOI] [PubMed] [Google Scholar]
- Delorme A, Truong D, Martinez-Cancino R, Pernet CR, Sivagnanam S, Yoshimoto K, Poldrack R, Majumdar A, and Makeig S. 2021. “Tools for importing and evaluating BIDS-EEG formatted data.” In doi: 10.1109/ner49283.2021.9441399. [DOI] [Google Scholar]
- Desjardins JA, van Noordt S, Huberty S, Segalowitz SJ, Elsabbagh M, 2021. EEG Integrated Platform Lossless (EEG-IP-L) pre-processing pipeline for objective signal quality assessment incorporating data annotation and blind source separation. J. Neurosci. Methods 347 (January), 108961. [DOI] [PubMed] [Google Scholar]
- Devezer B, Navarro DJ, Vandekerckhove J, Buzbas EO, 2021. The case for formal methodology in scientific reform. R. Soc. Open Sci 8 (3), 200805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DFG, 2021. Data tracking in research: aggregation and use or sale of usage data by academic publishers. DFG. https://www.dfg.de/download/pdf/foerderung/programme/lis/datentracking_papier_en.pdf. [Google Scholar]
- Di Cosmo R. 2018. “Software heritage: why and how we collect, preserve and share all the software source code.” In, 2–2.
- Ding Y, Suffren S, Bellec P, Lodygensky GA, 2019. Supervised machine learning quality control for magnetic resonance artifacts in neonatal data sets. Hum. Brain Mapp 40 (4), 1290–1297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Tommaso P, Chatzou M, Floden EW, Barja PP, Palumbo E, Notredame C, 2017. Nextflow enables reproducible computational workflows. Nat. Biotechnol 35 (4), 316–319. [DOI] [PubMed] [Google Scholar]
- Dragicevic P, Jansen Y, Sarma A, Kay M, Chevalier F, 2019. Increasing the Transparency of Research Papers with Explorable Multiverse Analyses. Association for Computing Machinery, New York, NY, USA, pp. 1–15. [Google Scholar]
- DuPre E, Hanke M, and Poline JB. 2019. “Nature abhors a paywall: how open science can realize the potential of naturalistic stimuli.” doi: 10.31234/osf.io/sdbqv. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DuPre E, Holdgraf C, Karakuzu A, Tetrel L, Bellec P, Stikov N, Poline JB, 2022. Beyond advertising: new infrastructures for publishing integrated research objects. PLoS Comput. Biol 18 (1), e1009651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eke D, Bernard A, Bjaalie JG, Chavarriaga R, Hanakawa T, Hannan A, Hill S, et al. , 2022. International data governance for neuroscience. Neuron 110 (4), 600–612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errington TM, Denis A, Perfito N, Iorns E, Nosek BA, 2021. Challenges for assessing replicability in preclinical cancer biology. eLife 10. doi: 10.7554/elife.67995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errington TM, Mathur M, Soderberg CK, Denis A, Perfito N, Iorns E, Nosek BA, 2021. Investigating the replicability of preclinical cancer biology. eLife 10 (December). doi: 10.7554/eLife.71601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esteban O, Birman D, Schaer M, Koyejo OO, Poldrack RA, Gorgolewski KJ, 2017. MRIQC: advancing the automatic prediction of image quality in MRI from unseen sites. PLoS One 12 (9), e0184661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esteban O, Blair RW, Nielson DM, Varada JC, Marrett S, Thomas AG, Poldrack RA, Gorgolewski KJ, 2019. Crowdsourced MRI quality metrics and expert quality annotations for training of humans and machines. Sci. Data 6 (1), 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esteban O, Ciric R, Finc K, Blair RW, Markiewicz CJ, Moodie CA, Kent JD, et al. , 2020. Analysis of task-based functional MRI data preprocessed with fMRIPrep. Nat. Protoc 15 (7), 2186–2202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esteban O, Markiewicz CJ, Blair RW, Moodie CA, Isik AI, Erramuzpe A, Kent JD, et al. , 2019. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16 (1), 111–116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eiss Robert, 2020. Confusion over Europe’s data-protection law is stalling scientific progress. Nature 584, 498. doi: 10.1038/d41586-020-02454-7. [DOI] [PubMed] [Google Scholar]
- Feldstein Ewing S, Luciana M, 2018. The Adolescent Brain Cognitive Development (ABCD) consortium: rationale, aims, and assessment strategy [Special Issue]. Dev. Cogn. Neurosci 32, 1–164.29496476 [Google Scholar]
- Flandin G, Friston K, 2008. Statistical parametric mapping (SPM). Scholarpedia J. 3 (4), 6232. [Google Scholar]
- Forbes SH, Aneja P, and Guest O. 2021. “The myth of (A)typical development.” doi: 10.31234/osf.io/ajynp. [DOI] [Google Scholar]
- Gau R, Flandin G, Janke A, tanguyduval R Oostenveld C Madan GN Galán, et al. 2022. Bids-Matlab. doi: 10.5281/zenodo.5910585. [DOI] [Google Scholar]
- Gau R, Noble S, Heuer K, Bottenhorn KL, Bilgin IP, Yang YF, Huntenburg JM, et al. , 2021. Brainhack: developing a culture of open, inclusive, community-driven neuroscience. Neuron 109 (11), 1769–1775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gelman A, and Loken E. 2013. “The garden of forking paths: why multiple comparisons can be a problem, even when there is no ‘fishing expedition’ or ‘p-Hacking’ and the research hypothesis was posited ahead of time.” 2013. http://www.stat.columbia.edu/~gelman/research/unpublished/p_hacking.pdf.
- Glasser MF, Sotiropoulos SN, Anthony Wilson J, Coalson TS, Fischl B, Andersson JL, Xu J, et al. , 2013. The minimal preprocessing pipelines for the human connectome project. Neuroimage 80 (October), 105–124 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorgolewski KJ, Alfaro-Almagro F, Auer T, Bellec P, Capotă M, Chakravarty MM, Churchill NW, et al. , 2017. BIDS apps: improving ease of use, accessibility, and reproducibility of neuroimaging data analysis methods. PLoS Comput. Biol 13 (3), e1005209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorgolewski KJ, Auer T, Calhoun VD, Craddock RC, Das S, Duff EP, Flandin G, et al. , 2016. The brain imaging data structure: a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 3, 1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorgolewski KJ, Burns CD, Madison C, Clark D, Halchenko YO, Waskom ML, Ghosh SS, 2011. Nipype: a flexible, lightweight and extensible neuroimaging data processing framework in python. Front. Neuroinf 5 (August). doi: 10.3389/fn-inf.2011.00013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorgolewski KJ, Poldrack RA, 2016. A practical guide for improving transparency and reproducibility in neuroimaging research. PLoS Biol. 14 (7), e1002506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorgolewski KJ, Varoquaux G, Rivera G, Schwarz Y, Ghosh SS, Maumet C, Sochat VV, et al. , 2015. NeuroVault.org: a web-based repository for collecting and sharing unthresholded statistical maps of the human brain. Front. Neuroinf 9 (April), 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Govaart H, Gisela, Schettino, Antonio, et al. , 2022. EEG ERP Preregistration Template. MetaArXiv doi: 10.31222/osf.io/4nvpt. [DOI] [Google Scholar]
- Gramfort A, Luessi M, Larson E, Engemann DA, Strohmeier D, Brodbeck C, Goj R, et al. , 2013. MEG and EEG data analysis with MNE-Python. Front. Neurosci 7 (December), 267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gramfort A, Luessi M, Larson E, Engemann DA., Strohmeier D, Brodbeck C, Parkkonen L, Hämäläinen MS, 2014. MNE software for processing MEG and EEG data. Neuroimage 86 (February), 446–460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griswold MA, Jakob PM, Heidemann RM, Nittka M, Jellus V, Wang J, Kiefer B, Haase A, 2002. Generalized Autocalibrating Partially Parallel Acquisitions (GRAPPA). Magn. Reson. Med 47 (6), 1202–1210. [DOI] [PubMed] [Google Scholar]
- Guest O, Martin AE, 2021. How computational modeling can force theory building in psychological science. Perspect. Psychol. Sci. J. Assoc. Psychol. Sci 16 (4), 789–802. [DOI] [PubMed] [Google Scholar]
- Halchenko Y, Goncalves M, di Oleggio Castello M. Visconti, Ghosh S, Salo T, Hanke M, Velasco P, et al. 2021. Nipy/heudiconv: doi: 10.5281/zenodo.5557588. [DOI] [Google Scholar]
- Halchenko Y, Meyer K, Poldrack B, Solanky D, Wagner A, Gors J, MacFarlane D, et al. , 2021. DataLad: distributed system for joint management of code, data, and their relationship. J. Open Source Softw 6 (63), 3262. [Google Scholar]
- Hall BD, Liu Y, Jansen Y, Dragicevic P, Chevalier F, Kay M, 2022. A survey of tasks and visualizations in multiverse analysis reports. Comput. Graph. Forum (February) doi: 10.1111/cgf.14443, Journal of the European Association for Computer Graphics. [DOI] [Google Scholar]
- Hanke M, Halchenko YO, Sederberg PB, Hanson SJ, Haxby JV, Pollmann S, 2009. PyMVPA: a python toolbox for multivariate pattern analysis of fMRI data. Neuroinformatics 7 (1), 37–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanke M, Pestilli F, Wagner AS, Markiewicz CJ, Poline JB, Halchenko YO, 2021. In defense of decentralized research data management. Neuroforum 27 (1), 17–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen MS, Sørensen TS, 2013. Gadgetron: an open source framework for medical image reconstruction. Mag. Reson. Med 69 (6), 1768–1776 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Hardwicke TE, Ioannidis JPA, 2018. Mapping the universe of registered reports. Nat. Hum. Behav 2 (11), 793–796. [DOI] [PubMed] [Google Scholar]
- Henrich J, Heine SJ, Norenzayan A, 2010. The weirdest people in the world? Behav. Brain Sci 33 (2-3), 61–83. [DOI] [PubMed] [Google Scholar]
- Henson RN, Abdulrahman H, Flandin G, Litvak V, 2019. Multimodal integration of M/EEG and f/MRI data in SPM12. Front. Neurosci 13 (April), 300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heunis S, Lamerichs R, Zinger S, Caballero-Gaudes C, Jansen JFA, Aldenkamp B, Breeuwer M, 2020. Quality and denoising in real-time functional magnetic resonance imaging neurofeedback: a methods review. Hum. Brain Mapp 41 (12), 3439–3467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holdgraf C, Appelhoff S, Bickel S, Bouchard K, D’Ambrosio S, David O, Devinsky O, et al. , 2019. iEEG-BIDS: extending the brain imaging data structure specification to human intracranial electrophysiology. Sci. Data 6 (1), 102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofstra Bas, Kulkarni Vivek V., Galvez Sebastian Munoz-Najar, McFarland Daniel A., 2020. The Diversity–Innovation Paradox in Science. PNAS 117 (17), 9284–9291. doi: 10.1073/pnas.1915378117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunt LT, 2019. The life-changing magic of sharing your data. Nat. Hum. Behav 3 (4), 312–315. [DOI] [PubMed] [Google Scholar]
- Hutson M, 2018. Artificial intelligence faces reproducibility crisis. Science 359 (6377), 725–726. [DOI] [PubMed] [Google Scholar]
- Inati SJ, Naegele JD, Zwart NR, Roopchansingh V, Lizak MJ, Hansen DC, Liu CY, et al. , 2017. ISMRM raw data format: a proposed standard for MRI raw datasets. Mag. Reson. Med 77 (1), 411–421 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jas M, Larson E, Engemann DA, Leppäkangas J, Taulu S, Hämäläinen M, Gramfort A, 2018. A reproducible MEG/EEG group study with the MNE software: recommendations, quality assessments, and good practices. Front. Neurosci 12 (August), 530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jellús V, and Kannengiesser SAR. 2014. “Adaptive coil combination using a body coil scan as phase reference.” In, 4406.
- Jenkinson M, Beckmann CF, Behrens TEJ, Woolrich MW, Smith SM, 2012. FSL. Neuroimage 62 (2), 782–790. [DOI] [PubMed] [Google Scholar]
- Jochimsen TH, Mengershausen M, 2004. ODIN: object-oriented development interface for NMR. J. Magn. Reson 170 (1), 67–78. [DOI] [PubMed] [Google Scholar]
- Jonge H, Cruz M, Holst S, 2021. Funders need to credit open science. Nature 599 (7885), 372. [DOI] [PubMed] [Google Scholar]
- Jwa AS, Poldrack RA, 2022. The spectrum of data sharing policies in neuroimaging data repositories. Hum. Brain Mapp 43 (8), 2707–2721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaplan RM, Irvin VL, 2015. Likelihood of null effects of large NHLBI clinical trials has increased over time. PLoS One 10 (8), e0132382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kappenman ES, Farrens JL, Zhang W, Stewart AX, Luck SJ, 2021. ERP CORE: an open resource for human event-related potential research. Neuroimage 225 (January), 117465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karakuzu A, Appelhoff S, Auer T, Boudreau M, Feingold F, Khan AR, Lazari A, et al. 2021. “qMRI-BIDS: an extension to the brain imaging data structure for quantitative magnetic resonance imaging data.” medRxiv. doi: 10.1101/2021.10.22.21265382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karakuzu A, Biswas L, Cohen-Adad J, Stikov N, 2022. Vendor-neutral sequences and fully transparent workflows improve inter-vendor reproducibility of quantitative MRI. Mag. Reson. Med 88 (3), 1212–1228 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Karakuzu A, Boudreau M, Duval T, Boshkovski T, Leppert I, Cabana JF, Gagnon I, et al. , 2020. qMRLab: quantitative MRI analysis, under one umbrella. J. Open Source Softw 5 (53), 2343. [Google Scholar]
- Karakuzu A, DuPre E, Tetrel L, Bermudez P, Boudreau M, Chin M, Poline JB, Das S, Bellec P, and Stikov N. 2022. “NeuroLibre : a preprint server for full-fledged reproducible neuroscience.” doi: 10.31219/osf.io/h89js. [DOI] [Google Scholar]
- Keator DB, Helmer K, Steffener J, Turner JA, Van Erp TGM, Gadde S, Ashish N, Burns GA, Nichols BN, 2013. Towards structured sharing of raw and derived neuroimaging data across existing resources. Neuroimage 82 (November), 647–661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kennedy DN, Abraham SA, Bates JF, Crowley A, Ghosh S, Gillespie T, Goncalves M, et al. , 2019. Everything matters: the ReproNim perspective on reproducible neuroimaging. Front. Neuroinf 13 (February), 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerr NL, 1998. HARKing: hypothesizing after the results are known. Personal. Soc. Psychol. Rev 2 (3), 196–217 An Official Journal of the Society for Personality and Social Psychology, Inc. [DOI] [PubMed] [Google Scholar]
- Khoo SYS, 2019. Article processing charge hyperinflation and price insensitivity: an open access sequel to the serials crisis. LIBER Q. doi: 10.18352/lq.10280, The Journal of the Association of European Research Libraries. [DOI] [Google Scholar]
- Kiar G, Chatelain Y, Glatard T, Salari A, Castro PO, michaelnicht AH, and Vadariya M. 2021. Verificarlo/fuzzy: Fuzzy v0.5.0. doi: 10.5281/zenodo.5027708. [DOI] [Google Scholar]
- Kiar G, Castro PO, Rioux P, Petit E, Brown ST, Evans AC, Glatard T, 2020. Comparing perturbation models for evaluating stability of neuroimaging pipelines. Int. J. High Perform. Comput. Appl 34 (5), 491–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kidwell MC, Lazarević LB, Baranski E, Hardwicke TE, Piechowski S, Falkenberg LS, Kennett C, et al. , 2016. Badges to acknowledge open practices: a simple, low-cost, effective method for increasing transparency. PLoS Biol. 14 (5), e1002456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleiner M, Brainard D, Pelli D, 2007. What’s New in Psychtoolbox-3? Perception 36 (14). [Google Scholar]
- Klein RA, Vianello M, Hasselman F, Adams BG, Adams RB Jr, Alper S, Aveyard M, et al. , 2018. Many Labs 2: investigating variation in replicability across samples and settings. Adv. Methods Pract. Psychol. Sci 1 (4), 443–490. [Google Scholar]
- Knopp T, Grosser M, 2021. MRIReco.jl: an MRI reconstruction framework written in Julia. Mag. Reson. Med 86 (3), 1633–1646 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Knudsen GM, Ganz M, Appelhoff S, Boellaard R, Bormans G, Carson RE, Catana C, et al. , 2020. Guidelines for the content and format of PET brain data in publications and archives: a consensus paper. J. Cereb. Blood Flow Metab 40 (8), 1576–1585 Official Journal of the International Society of Cerebral Blood Flow and Metabolism. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kollada M, Gao Q, Mellem MS, Banerjee T, Martin WJ, 2021. A generalizable method for automated quality control of functional neuroimaging datasets. In: Shaban-Nejad A, Michalowski M, Buckeridge DL (Eds.), Explainable AI in Healthcare and Medicine: Building a Culture of Transparency and Accountability. Springer International Publishing, Cham, pp. 55–68. [Google Scholar]
- Laird AR, 2021. Large, open datasets for human connectomics research: considerations for reproducible and responsible data use. Neuroimage 244 (December), 118579. [DOI] [PubMed] [Google Scholar]
- Layton KJ, Kroboth S, Jia F, Littin S, Yu H, Leupold J, Nielsen JF, Stöcker T, Zaitsev M, 2017. Pulseq: a rapid and hardware-independent pulse sequence prototyping framework. Mag. Reson. Med 77 (4), 1544–1552 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Lee MD, Criss AH, Devezer B, Donkin C, Etz A, Leite FP, Matzke D, et al. , 2019. Robust modeling in cognitive science. Comput. Brain Behav 2 (3), 141–153. [Google Scholar]
- Lee Y, Callaghan MF, Acosta-Cabronero J, Lutti A, Nagy Z, 2019. Establishing intra- and inter-vendor reproducibility of T relaxation time measurements with 3T MRI. Mag. Reson. Med 81 (1), 454–465 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Litvak V, Mattout J, Kiebel S, Phillips C, Henson R, Kilner J, Barnes G, et al. , 2011. EEG and MEG data analysis in SPM8. Comput. Intell. Neurosci 2011 (March), 852961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Llorens A, Tzovara A, Bellier L, Bhaya-Grossman I, Bidet-Caulet A, Chang WK, Cross ZR, Dominguez-Faus R, Flinker A, Fonken Y, Gorenstein MA, Holdgraf C, Hoy CW, Ivanova MV, Jimenez RT, Jun S, Kam JWY, Kidd C, Marcelle E …, Dronkers NF, 2021. Gender bias in academia: a lifetime problem that needs solutions. Neuron 109 (13), 2047–2074. doi: 10.1016/j.neuron.2021.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levitis E, van Praag CDG, Gau R, Heunis S, DuPre E, Kiar G, Bottenhorn KL, Glatard T, Nikolaidis A, Whitaker KJ, Mancini M, Niso G, Afyouni S, Alonso-Ortiz E, Appelhoff S, Arnatkeviciute A, Atay SM, Auer T, Baracchini G, …, Maumet C, 2021. Centering inclusivity in the design of online conferences—An OHBM–open science perspective. GigaScience 10 (8). doi: 10.1093/gi-gascience/giab051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Kale A, Althoff T, Heer J, 2021. Boba: authoring and visualizing multiverse analyses. IEEE Trans. Visual Comput. Graph 27 (2), 1753–1763. [DOI] [PubMed] [Google Scholar]
- Li X, Ai L, Giavasis S, Jin H, Feczko E, Xu T, Clucas J, et al. 2021. “Moving beyond processing and analysis-related variation in neuroscience.”bioRxiv. 10.1101/2021.12.01.470790v1. [DOI] [PubMed] [Google Scholar]
- Lustig M, Donoho DL, Santos JM, Pauly JM, 2008. Compressed sensing MRI. IEEE Signal Process. Mag 25 (2), 72–82. [Google Scholar]
- Madan CR, 2021. Scan once, analyse many: using large open-access neuroimaging datasets to understand the brain. Neuroinformatics doi: 10.1007/s12021-021-09519-6, May. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malkinson TS, Terhune DB, Kollamkulam M, et al. , 2021. Gender Imbalance in the Editorial Activities of a Researcher-led. Journal doi: 10.1101/2021.11.09.467796, bioRxiv. [DOI] [Google Scholar]
- Magland JF, Li C, Langham MC, Wehrli FW, 2016. Pulse sequence programming in a dynamic visual environment: sequencetree. Mag. Reson. Med 75 (1), 257–265 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maier O, Baete SH, Fyrdahl A, Hammernik K, Harrevelt S, Kasper L, Karakuzu A, et al. , 2021. CG-SENSE revisited: results from the first ISMRM reproducibility challenge. Mag. Reson. Med 85 (4), 1821–1839 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcus DS, Olsen TR, Ramaratnam M, Buckner RL, 2007. The extensible neuroimaging archive toolkit: an informatics platform for managing, exploring, and sharing neuroimaging data. Neuroinformatics 5 (1), 11–34. [DOI] [PubMed] [Google Scholar]
- Markiewicz CJ, Vega ADL, Wagner A, Halchenko YO, Finc K, Ciric R, Goncalves M, et al. 2021. “Poldracklab/fitlins: v0.9.2.” Zenodo. 10.5281/zenodo.5120201. [DOI] [Google Scholar]
- Markiewicz CJ, Gorgolewski KJ, Feingold F, Blair R, Halchenko YO, Miller E, Hardcastle N, et al. , 2021. The openneuro resource for sharing of neuroscience data. eLife 10 (October). doi: 10.7554/eLife.71774 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Markowetz F, 2015. Five selfish reasons to work reproducibly. Genom. Biol 16 (December), 274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maumet C, Auer T, Bowring A, Chen G, Das S, Flandin G, Ghosh S, et al. , 2016. Sharing brain mapping statistical results with the neuroimaging data model. Sci. Data 3 (1), 1–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKiernan B, Brown B, Kenall, 2016. & Spies JR (2016). Point of view: how open science helps researchers succeed. eLife. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer K, Hanke M, Halchenko Y, Poldrack B, and Wagner A. 2021. Datalad/datalad-Container: 1.1.4. 10.5281/zenodo.4701527. [DOI] [Google Scholar]
- Meyer M, Lamers D, Kayhan E, Hunnius S, Oostenveld R, 2021. Enhancing reproducibility in developmental EEG research: BIDS, cluster-based permutation tests, and effect sizes. Dev. Cogn. Neurosci 52 (December), 101036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milham MP, Craddock RC, Son JJ, Fleischmann M, Clucas J, Xu H, Koo B, et al. , 2018. Assessment of the impact of shared brain imaging data on the scientific literature. Nat. Commun 9 (1), 2818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreau CA, Jean-Louis M, Blair R, Markiewicz CJ, Turner JA, Calhoun VD, Nichols TE, Pernet CR, 2020. The genetics-BIDS extension: easing the search for genetic data associated with human brain imaging. GigaScience 9 (10). doi: 10.1093/gigascience/giaa104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreau L, Groth P, Cheney J, Lebo T, Miles S, 2015. The rationale of PROV. Web Semant. 35 (December), 235–257. [Google Scholar]
- Mortamet B, Bernstein MA, Jack CR Jr, Gunter JL, Ward C, Britson PJ, Meuli R, Thiran JP, Krueger G Alzheimer’s Disease Neuroimaging Initiative, 2009. Automatic quality assessment in structural brain magnetic resonance imaging. Mag. Reson. Med 62 (2), 365–372 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moshontz H, Binion G, Walton H, Brown BT, Syed M, 2021. A guide to posting and managing preprints. Adv. Methods Pract. Psychol. Sci 4 (2). doi: 10.1177/25152459211019948. [DOI] [Google Scholar]
- Munafò MR, Nosek BA, Bishop DVM, Button KS, Chambers CD, Sert NPD, Simonsohn U, Wagenmakers EJ, Ware JJ, Ioannidis JPA, 2017. A manifesto for reproducible science. Nat. Hum. Behav 1 (1), 1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nabyonga-Orem J, Asamani JA, Nyirenda T, Abimbola S, 2020. Article processing charges are stalling the progress of African researchers: a call for urgent reforms. BMJ Glob. Health 5 (9). doi: 10.1136/bmjgh-2020-003650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nature, 2015. Let’s think about cognitive bias. Nature; 526 (7572), 163. [DOI] [PubMed] [Google Scholar]
- Nichols TE, Das S, Eickhoff SB, Evans AC, Glatard T, Hanke M, Kriegeskorte N, et al. , 2017. Best practices in data analysis and sharing in neuroimaging using MRI. Nat. Neurosci 20 (3), 299–303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen JF, Noll DC, 2018. TOPPE: a framework for rapid prototyping of MR pulse sequences. Mag. Reson. Med 79 (6), 3128–3134 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niso G, Gorgolewski KJ, Bock E, Brooks TL, Flandin G, Gramfort A, Henson RN, et al. , 2018. MEG-BIDS: the brain imaging data structure extended to magnetoencephalography. Scient. Data 5 (June), 180110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niso G, Krol LR, Combrisson E, Dubarry AS, Elliott MA, Franqois C, Héjja-Brichard Y, et al. , 2022. Good scientific practice in MEEG research: progress and perspectives. Neuroimage (March), 119056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niso G, Rogers C, Moreau JT, Chen LY, Madjar C, Das S, Bock E, et al. , 2016. OMEGA: the open MEG archive. Neuroimage 124 (Pt B), 1182–1187. [DOI] [PubMed] [Google Scholar]
- Niso G, Tadel F, Bock E, Cousineau M, Santos A, Baillet S, 2019. Brainstorm pipeline analysis of resting-state data from the open MEG archive. Front. Neurosci 13 (April), 284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nissen SB, Magidson T, Gross K, Bergstrom CT, 2016. Publication bias and the canonization of false facts. eLife 5 (December). doi: 10.7554/eLife.21451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nooner KB, Colcombe S, Tobe R, Mennes M, Benedict M, Moreno A, Panek L, et al. , 2012. The NKI-rockland sample: a model for accelerating the pace of discovery science in psychiatry. Front. Neurosci. 0. doi: 10.3389/fnins.2012.00152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nørgaard M, Ganz M, Svarer C, Feng L, Ichise M, Lanzenberger R, Lubberink M, et al. , 2019. Cerebral serotonin transporter measurements with [11C]DASB: a review on acquisition and preprocessing across 21 PET centres. J. Cereb. Blood Flow Metab 39 (2), 210–222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nørgaard M, Ganz M, Svarer C, Frokjaer VG, Greve DN, Strother SC, Knudsen GM, 2020. Different preprocessing strategies lead to different conclusions: a [11C]DASB-PET reproducibility study. J. Cereb. Blood Flow Metab 40 (9), 1902–1911 Official Journal of the International Society of Cerebral Blood Flow and Metabolism. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nqrgaard M, Ozenne B, Svarer C, Frokjaer VG, Schain M, Strother SC, and Ganz M. 2019. “Preprocessing, prediction and significance: framework and application to brain imaging.” In, 196–204. Springer International Publishing. [Google Scholar]
- Norgaard M, Matheson GJ, Hansen HD, et al. , 2022. PET-BIDS, an extension to the brain imaging data structure for positron emission tomography. Scientific data 9 (1), 1–7. doi: 10.1038/s41597-022-01164-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nosek BA, Beck ED, Campbell L, Flake JK, Hardwicke TE, Mellor DT, Veer A.E.’t, Vazire S, 2019. Preregistration is hard, and worthwhile. Trends Cogn. Sci. 23 (10), 815–818. [DOI] [PubMed] [Google Scholar]
- Nosek BA, Ebersole CR, DeHaven AC, Mellor DT, 2018. The preregistration revolution. Proc. Natl. Acad. Sci 2017 (15), 201708274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nosek BA, Hardwicke TE, Moshontz H, Allard A, Corker KS, Dreber A, Fidler F, et al. , 2022. Replicability, robustness, and reproducibility in psychological science. Annu. Rev. Psychol 73 (January), 719–748. [DOI] [PubMed] [Google Scholar]
- Nosek BA, Lakens D, 2014. Registered reports: a method to increase the credibility of published reports. Soc. Psychol 45 (3), 137–141. [Google Scholar]
- Nosek BA, Spies JR, Motyl M, 2012. Scientific Utopia: II. Restructuring incentives and practices to promote truth over publishability. Perspect. Psychol. Sci 7 (6), 615–631 A Journal of the Association for Psychological Science. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nosek BA, Stephen Lindsay D, 2018. Preregistration becoming the norm in psychological science. APS Observer. 31 (3). https://www.psychologicalscience.org/observer/preregistration-becoming-the-norm-in-psychological-science/comment-page-1. [Google Scholar]
- Oberauer K, Lewandowsky S, 2019. Addressing the theory crisis in psychology. Psychon. Bull. Rev 26 (5), 1596–1618. [DOI] [PubMed] [Google Scholar]
- Oostenveld R, Fries P, Maris E, Schoffelen JM, 2011. FieldTrip: open source software for advanced analysis of MEG, EEG, and invasive electrophysiological data. Comput. Intell. Neurosci 2011, 156869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Open Science Collaboration, 2015. PSYCHOLOGY. Estimating the reproducibility of psychological science. Science 349 (6251), aac4716. [DOI] [PubMed] [Google Scholar]
- Paret C, Unverhau N, Feingold F, Poldrack RA, Stirner M, Schmahl C, and Sicorello M. 2022. “Survey on open science practices in functional neuroimaging.” NeuroImage. 257, 10.1101/2021.11.26.470115. [DOI] [PubMed] [Google Scholar]
- Patel CJ, Burford B, Ioannidis JPA, 2015. Assessment of vibration of effects due to model specification can demonstrate the instability of observational associations. J. Clin. Epidemiol 68 (9), 1046–1058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paul M, Govaart G, Schettino A, 2021. Making ERP research more transparent: guidelines for preregistration. Int. J. Psychophysiol 164, 52–63. [DOI] [PubMed] [Google Scholar]
- Pavlov YG, Adamian N, Appelhoff S, Arvaneh M, Benwell CSY, Beste C, Bland AR, et al. , 2021. #EEGManyLabs: investigating the replicability of influential EEG experiments. Cortex 144 (April), 213–229. [DOI] [PubMed] [Google Scholar]
- Peirce J, Gray JR, Simpson S, MacAskill M, Höchenberger R, Sogo H, Kastman E, Lindeløv JK, 2019. PsychoPy2: experiments in behavior made easy. Behav. Res. Methods 51 (1), 195–203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelli DG, 1997. The videotoolbox software for visual psychophysics: transforming numbers into movies. Spat. Vis 10 (4), 437–442. [PubMed] [Google Scholar]
- Penny WD, Friston KJ, Ashburner JT, Kiebel SJ, Nichols TE, 2011. Statistical Parametric Mapping: The Analysis of Functional Brain Images. Elsevier. [Google Scholar]
- Pernet CR, 2014. Misconceptions in the use of the general linear model applied to functional MRI: a tutorial for junior neuro-imagers. Front. Neurosci 8 (January), 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pernet CR, Appelhoff S, Gorgolewski KJ, Flandin G, Phillips C, Delorme A, Oostenveld R, 2019. EEG-BIDS: an extension to the brain imaging data structure for electroencephalography. Sci. Data 6 (1), 103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pernet CR, Garrido MI, Gramfort A, Maurits N, Michel CM, Pang E, Salmelin R, Schoffelen JM, Valdes-Sosa PA, Puce A, 2020. Issues and recommendations from the OHBM COBIDAS MEEG committee for reproducible EEG and MEG research. Nat. Neurosci 23 (12), 1473–1483. [DOI] [PubMed] [Google Scholar]
- Pernet CR, Martinez-Cancino R, Truong D, Makeig S, Delorme A, 2020. From BIDS–formatted EEG data to sensor-space group results: a fully reproducible workflow with EEGLAB and LIMO EEG. Front. Neurosci 14, 610388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pernet CR, Poline JB, 2015. Improving functional magnetic resonance imaging reproducibility. GigaScience 4 (1). doi: 10.1186/s13742-015-0055-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piwowar HA, Priem J, Larivière V, Pablo Alperin J, Matthias L, Norlander B, Farley A, West J, Haustein S, 2018. The state of OA: a large-scale analysis of the prevalence and impact of open access articles. PeerJ 6 (February), e4375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poldrack RA, Baker CI, Durnez J, Gorgolewski KJ, Matthews PM, Munafò MR, Nichols TE, Poline JB, Vul E, Yarkoni T, 2017. Scanning the horizon: towards transparent and reproducible neuroimaging research. Nat. Rev. Neurosci 18 (2), 115–126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poldrack RA, Feingold F, Frank MJ, Gleeson P, Hollander G, Huys QJM, Love BC, et al. , 2019. The importance of standards for sharing of computational models and data. Comput. Brain Behav 2 (3-4), 229–232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poldrack RA, Gorgolewski KJ, 2014. Making big data open: data sharing in neuroimaging. Nat. Neurosci 17 (11), 1510–1517. [DOI] [PubMed] [Google Scholar]
- Poldrack RA, Huckins G, Varoquaux G, 2020. Establishment of best practices for evidence for prediction: a review. JAMA Psychiatry 77 (5), 534–540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poldrack RA, Kittur A, Kalar D, Miller E, Seppa C, Gil Y, Parker DS, Sabb FW, Bilder RM, 2011. The cognitive atlas: toward a knowledge foundation for cognitive neuroscience. Front. Neuroinf 5 (September), 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poldrack RA, Whitaker K, Kennedy D, 2020. Introduction to the special issue on reproducibility in neuroimaging. Neuroimage 218 (September), 116357. [DOI] [PubMed] [Google Scholar]
- Poline JB, Kennedy DN, Sommer FT, Ascoli GA, Essen DCV, Ferguson AR, Grethe JS, et al. , 2022. Is neuroscience FAIR? A call for collaborative standardisation of neuroscience data. Neuroinformatics (January) doi: 10.1007/s12021-021-09557-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popov T, Oostenveld R, Schoffelen JM, 2018. FieldTrip made easy: an analysis protocol for group analysis of the auditory steady state brain response in time, frequency, and space. Front. Neurosci 12 (October), 711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravi K, Geethanath S, Vaughan J, 2019. PyPulseq: a python package for MRI pulse sequence design. J. Open Source Softw 4 (42), 1725. [Google Scholar]
- Robbins K, Truong D, Appelhoff S, Delorme A, Makeig S, 2021. Capturing the nature of events and event context using hierarchical event descriptors (HED). Neuroimage (November), 118766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robbins K, Truong D, Jones A, Callanan I, Makeig S, 2021. Building FAIR functionality: annotating events in time series data using hierarchical event descriptors (HED). OSF doi: 10.31219/osf.io/5fg73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin M, 2020. Does preregistration improve the credibility of research findings? Quant. Methods Psychol 16 (4), 376–390. [Google Scholar]
- Sandve GK, Nekrutenko A, Taylor J, Hovig E, 2013. Ten simple rules for reproducible computational research. PLoS Comput. Biol 9 (10), e1003285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- San R, López de A, 2021. Open Science in Horizon Europe. Zenodo doi: 10.5281/zen-odo.4681073. [DOI] [Google Scholar]
- Sasaki M, Yamada K, Watanabe Y, Matsui M, Ida M, Fujiwara S, Shibata E Acute Stroke Imaging Standardization Group-Japan (ASIST-Japan) Investigators, 2008. Variability in absolute apparent diffusion coefficient values across different platforms may be substantial: a multivendor, multi-institutional comparison study. Radiology 249 (2), 624–630. [DOI] [PubMed] [Google Scholar]
- Schäfer T, Schwarz MA, 2019. The meaningfulness of effect sizes in psychological research: differences between sub-disciplines and the impact of potential biases. Front. Psychol 10 (April), 813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scheel AM, 2020. Registered reports: a process to safeguard high-quality evidence. Q. Life Res 29 (12), 3181–3182 An International Journal of Quality of Life Aspects of Treatment, Care and Rehabilitation. [DOI] [PubMed] [Google Scholar]
- Schilling KG, Rheault F, Petit L, Hansen CB, Nath V, Yeh FC, Girard G, et al. , 2021. Tractography dissection variability: what happens when 42 groups dissect 14 white matter bundles on the same dataset? Neuroimage 243 (November), 118502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmitt T, Rieger JW, 2021. Recommendations of choice of head coil and prescan normalize filter depend on region of interest and task. Front. Neurosci 0. doi: 10.3389/fnins.2021.735290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schreiweis C, Volle E, Durr A, Auffret A, Delarasse C, George N, Dumont M, Hassan BA, Renier N, Rosso C, Thiebaut de Schotten M, Burguiere E, Zujovic V, 2019. A neuroscientific approach to increase gender equality. Nat. Hum. Behav 3 (12), 1238–1239. doi: 10.1038/s41562-019-0755-7. [DOI] [PubMed] [Google Scholar]
- Serra-Garcia M, Gneezy U, 2021. Nonreplicable publications are cited more than replicable ones. Sci. Adv 7 (21). doi: 10.1126/sciadv.abd1705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shafto MA, Cam-CAN, Tyler LK, Dixon M, Taylor JR, Rowe JB, Cusack R, et al. , 2014. The Cambridge Centre for Ageing and Neuroscience (Cam-CAN) study protocol: a cross-sectional, lifespan, multidisciplinary examination of healthy cognitive ageing. BMC Neurol. doi: 10.1186/s12883-014-0204-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simmons JP, Nelson LD, Simonsohn U, 2011. False-positive psychology: undisclosed flexibility in data collection and analysis allows presenting anything as significant. Psychol. Sci 22 (11), 1359–1366. [DOI] [PubMed] [Google Scholar]
- Simmons JP, Nelson L, Simonsohn U, 2021. Pre-registration: why and how. J. Consum. Psychol 31 (1), 151–162 The Official Journal of the Society for Consumer Psychology. [Google Scholar]
- Simonsohn U, Simmons JP, Nelson LD, 2015. Specification curve: descriptive and inferential statistics on all reasonable specifications. SSRN Electron. J (no. November) doi: 10.2139/ssrn.2694998. [DOI] [Google Scholar]
- Simonsohn Uri, Simmons Joseph P., Nelson Leif D., 2020. Specification curve analysis. Nat Hum Behav. 4 (11), 1208–1214. doi: 10.1038/s41562-020-0912-z,Epub 2020 Jul 27. [DOI] [PubMed] [Google Scholar]
- Smith SM, Beckmann CF, Andersson J, Auerbach EJ, Bijsterbosch J, Douaud G, Duff E, et al. , 2013. Resting-state fMRI in the human connectome project. Neuroimage 80 (October), 144–168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soderberg CK, Errington TM, Schiavone SR, Bottesini J, Thorn FS, Vazire S, Esterling KM, Nosek BA, 2021. Initial evidence of research quality of registered reports compared with the standard publishing model. Nat. Hum. Behav 5 (8), 990–997. [DOI] [PubMed] [Google Scholar]
- Šoškić A, Jovanović V, Styles SJ, Kappenman ES, Ković V, 2021. How to do better N400 studies: reproducibility, consistency and adherence to research standards in the existing literature. Neuropsychol. Rev doi: 10.1007/s11065-021-09513-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steegen S, Tuerlinckx F, Gelman A, Vanpaemel W, 2016. Increasing transparency through a multiverse analysis. Perspect. Psychol. Sci. J. Assoc. Psychol. Sci 11 (5), 702–712. [DOI] [PubMed] [Google Scholar]
- Stikov N, Trzasko JD, Bernstein MA, 2019. Reproducibility and the future of MRI research. Mag. Reson. Med 82 (6), 1981–1983 Official Journal of the Society of Magnetic Resonance in Medicine /Society of Magnetic Resonance in Medicine. [DOI] [PubMed] [Google Scholar]
- Strand JF, 2021. Error tight: exercises for lab groups to prevent research mistakes. OSF doi: 10.31234/osf.io/rsn5y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strand JF, and Brown VA. n.d. 2022. “Spread the word: enhancing replicability of speech research through stimulus sharing.” 10.31234/osf.io/amevw. [DOI] [PMC free article] [PubMed]
- Strother SC, 2006. Evaluating fMRI preprocessing pipelines. IEEE Eng. Med. Biol. Mag. 25 (2), 27–41 The Quarterly Magazine of the Engineering in Medicine & Biology Society. [DOI] [PubMed] [Google Scholar]
- Strother SC, Conte SLA, Hansen LK, Anderson J, Zhang J, Pulapura S, Rottenberg D, 2004. Optimizing the fMRI data-processing pipeline using prediction and reproducibility performance metrics: I. A preliminary group analysis. Neuroimage 23 (Suppl 1), S196–S207. [DOI] [PubMed] [Google Scholar]
- Styles SJ, Ković V, Ke H, Šoškić A, 2021. Towards ARTEM-IS: design guidelines for evidence-based EEG methodology reporting tools. Neuroimage 245 (December), 118721. [DOI] [PubMed] [Google Scholar]
- Tadel F, Baillet S, Mosher JC, Pantazis D, Leahy RM, 2011. Brainstorm: a user-friendly application for MEG/EEG analysis. Comput. Intell. Neurosci 2011 (April), 879716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tadel F, Bock E, Niso G, Mosher JC, Cousineau M, Pantazis D, Leahy RM, Baillet S, 2019. MEG/EEG group analysis with brainstorm. Front. Neurosci 13 (February), 76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor JR, Williams N, Cusack R, Auer T, Shafto MA, Dixon M, Tyler, Cam–Can LK, Henson RN, 2017. The Cambridge Centre for Ageing and Neuroscience (Cam-CAN) data repository: structural and functional MRI, MEG, and cognitive data from a cross-sectional adult lifespan sample. Neuroimage 144 (Pt B), 262–269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Open Brain Consent Working Group, 2021. The open brain consent: informing research participants and obtaining consent to share brain imaging data. Hum. Brain Mapp 42 (7), 1945–1951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnold B, Bowler L, Gibson S, Herterich P, Higman R, Krystalli A, Morley A, O’Reilly M, Whitaker K The Turing Way Community, 2019. The Turing Way: A Handbook for Reproducible Data Science. Zenodo doi: 10.5281/ZENODO.3233986. [DOI] [Google Scholar]
- Tong G, Gaspar AS, Qian E, Ravi KS, Vaughan JT, Nunes RG, Geethanath S, 2021. A framework for validating open-source pulse sequences. Magn. Reson. Imaging 87 (November), 7–18. [DOI] [PubMed] [Google Scholar]
- Troupin C, Muñoz C, Gabriel Fernández J, and Rújula MÀ. 2018. “Scientific results traceability: software citation using GitHub and Zenodo.” In. Barcelona. https://orbi.uliege.be/bitstream/2268/230513/1/IMDIS2018_Ctroupin_poster66.pdf. [Google Scholar]
- Tustison NJ, Johnson HJ, Rohlfing T, Klein A, Ghosh SS, Ibanez L, Avants BB, 2013. Instrumentation bias in the use and evaluation of scientific software: recommendations for reproducible practices in the computational sciences. Front. Neurosci 7 (September), 162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uecker M, Ong F, Tamir JI, Bahri D, Virtue P, Cheng JY, Zhang T, and Lustig M. 2015. “Berkeley advanced reconstruction toolbox (BART).” In. 10.5281/zen-odo.31907. [DOI] [Google Scholar]
- Uğurbil K, Xu J, Auerbach EJ, Moeller S, Vu An.T., Duarte-Carvajalino JM, Lenglet C, et al. , 2013. Pushing spatial and temporal resolution for functional and diffusion MRI in the human connectome project. Neuroimage 80 (October), 80–104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Essen DC, Smith SM, Barch DM, Behrens TEJ, Yacoub E, Ugurbil K for the WU-Minn HCP Consortium, 2013. The WU-Minn human connectome project: an overview. Neuroimage 80 (October), 62–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Essen DC, Ugurbil K, Auerbach E, Barch D, Behrens TEJ, Bucholz R, Chang A, et al. , 2012. The human connectome project: a data acquisition perspective. Neuroimage 62 (4), 2222–2231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vega A, Rocca R, Blair RW, Markiewicz CJ, Mentch J, Kent JD, Herholz P, Ghosh SS, Poldrack RA, and Yarkoni T. 2022. “Neuroscout, a unified platform for generalizable and reproducible fMRI research.” bioRxiv. 10.1101/2022.04.05.487222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Visconti di Oleggio Castello M, Dobson JE, Sackett T, Kodiweera C, Haxby JV, Goncalves M, Ghosh S, and Halchenko YO. 2020. ReproNim/reproin 0.6.0. doi: 10.5281/zenodo.3625000. [DOI] [Google Scholar]
- Vliet M, 2020. Seven quick tips for analysis scripts in neuroimaging. PLoS Comput. Biol 16 (3), e1007358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vliet M, Liljeström M, Aro S, Salmelin R, Kujala J, 2018. Analysis of functional connectivity and oscillatory power using DICS: from raw MEG data to group-level statistics in python. Front. Neurosci 12 (September), 586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagenmakers EJ, Dutilh G, 2016. Seven selfish reasons for preregistration. APS Observer. 29 (9). https://www.psychologicalscience.org/observer/seven%C3%A2%E2%82%AC%20selfish%C3%A2%E2%82%AC%20reasons%C3%A2%E2%82%AC%20for%C3%A2%E2%82%AC%20preregistration. [Google Scholar]
- Wagner AS, Waite LK, Meyer K, Heckner MK, Kadelka T, Reuter N, Waite AQ, et al. , 2021. The DataLad Handbook. Zenodo doi: 10.5281/ZENODO.4495560. [DOI] [Google Scholar]
- Wettenhovi VV, Vauhkonen M, Kolehmainen V, 2021. OMEGA: open-source emission tomography software. Phys. Med. Biol 66 (6). doi: 10.1088/1361-6560/abe65f. [DOI] [PubMed] [Google Scholar]
- Whitaker K 2019. “Definitions.” Data 100 at UC Berkeley. October 21, 2019. https://web.archive.org/web/20191030093753/https://the-turing-way.netlify.com/reproducibility/03/definitions.html. [Google Scholar]
- Wicherts JM, Bakker M, Molenaar D, 2011. Willingness to share research data is related to the strength of the evidence and the quality of reporting of statistical results. PLoS One 6 (11), e26828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilkinson MD, Dumontier M, Aalbersberg IJJ, Appleton G, Axton M, Baak A, Blomberg N, et al. , 2016. The FAIR guiding principles for scientific data management and stewardship. Sci. Data 3 (March), 160018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson G, Bryan J, Cranston K, Kitzes J, Nederbragt L, Teal TK, 2017. Good enough practices in scientific computing. PLoS Comput. Biol 13 (6), e1005510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winter L, Haopeng H, Barghoorn A, Hoffmann W, Hetzer S, Winkler S, and Others. 2016. “Open source imaging initiative.” In. Vol. 3638. https://wiki.opensourceecology.org/images/a/a5/OSIIabstract.pdf. [Google Scholar]
- Yarkoni T, 2022. The generalizability crisis. Behav. Brain Sci 45. doi: 10.1017/S0140525X20001685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yarkoni T, Markiewicz CJ, Vega A, Gorgolewski KJ, Salo T, Halchenko YO, McNamara Q, et al. , 2019. PyBIDS: python tools for BIDS datasets. J. Open Source Softw 4 (40). doi: 10.21105/joss.01294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yücel MA, Lühmann AV, Scholkmann F, Gervain J, Dan I, Ayaz H, Boas D, et al. , 2021. Best practices for fNIRS publications. Neurophotonics 8 (1), 012101. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Not applicable
No data was used for the research described in the article.


