Abstract
Childhood stunting is a persistent public health challenge in Ethiopia, significantly impacting children’s physical growth, cognitive development, and overall well-being. This study overcame a key limitation in previous stunting prediction models by developing a multi-class classification model that predicts stunting severity (severe, moderate, normal) using Ethiopia’s nationally representative EDHS data from 2011 to 2016. Secondary data from the 2011 and 2016 Ethiopian Demographic and Health Surveys (EDHS) were analyzed, comprising 18,451 instances with 28 features. Data preprocessing included handling missing values, duplicate removal, feature selection, and synthetic minority over-sampling technique (SMOTE) for class balancing, resulting in 33,495 instances with 18 selected features. Four ensemble machine learning algorithms Random Forest, AdaBoost, XGBoost, and CatBoost were implemented and evaluated based on accuracy, precision, recall, F1-score, and ROC-AUC. Among the models, Random Forest achieved the highest performance with an accuracy of 97.985%, precision of 97.986%, recall of 97.985%, F1-score of 97.954%, and ROC-AUC of 99.995%. The top risk factors contributing to stunting included child’s age, maternal education level, birth order, household wealth index, mother’s BMI, breastfeeding duration, and access to clean water and sanitation. This study demonstrates the effectiveness of machine learning in accurately predicting childhood stunting in Ethiopia. The findings provide critical insights for healthcare professionals and policymakers to implement targeted intervention strategies, ultimately reducing childhood stunting prevalence.
Keywords: Childhood stunting, Risk factors of stunting, Ensemble machine learning
Subject terms: Diseases, Health care, Risk factors
Introduction
Childhood stunting, marked by low height-for-age, reflects chronic malnutrition and harms physical and cognitive development1. Stunting, according to the World Health Organization (WHO), is a height-for-age Z-score (HAZ) below − 2 standard deviations, reflecting chronic undernutrition. A HAZ ≥ -2 SD indicates normal growth, between − 2 and − 3 SD indicates moderate stunting, and below − 3 SD indicates severe stunting2.
The widespread prevalence of stunting, affecting nearly 154.8 million children globally, underscores its status as a major public health issue that demands urgent attention and evidence-based intervention3.
The consequences of stunting are multi-faceted and long-lasting, impacting not only physical growth but also cognitive abilities, school performance, and future productivity4,5. Specifically, children who experience stunting are more susceptible to chronic diseases later in life, have increased morbidity and mortality rates, and are often trapped in cycles of poverty due to diminished earning potential as adults4.
Consequently, the social and economic ramifications of childhood stunting extend beyond individuals and families, posing a substantial impediment to national development, especially for low-income countries5.
In Ethiopia, stunting affects over one-third of children under the age of five5, placing it among the most severely affected nations globally. This high prevalence is the result of a complex combination of factors, including inadequate access to healthcare, poor prenatal care, food insecurity, socio-economic challenges, and detrimental environmental influences3,4,6.
These challenges contribute to Ethiopia’s economic losses, reducing the nation’s growth by an estimated 8% due to lost productivity. A significant percentage of all child deaths are linked to malnutrition and stunting. As a result, the Ethiopian government and many international agencies are attempting to combat stunting through various intervention programs and policies7,8.
Previous studies, primarily using statistical methods and cross-sectional data, have identified various factors associated with stunting in Ethiopia4,6,9–12. However, these studies were often limited by their geographical scope, sample sizes, and inability to capture complex relationships between risk factors10–12.
Furthermore, they have not fully utilized machine learning approaches to develop robust predictive models or actionable artifacts to guide intervention strategies. Studies conducted in Zambia, Papua New Guinea, and Pakistan using machine learning have also had some limitations, particularly in binary classification, limiting age ranges, and incomplete consideration of determinant factors like birth interval and antenatal care visits13–15.
Recent research has demonstrated the growing importance of machine learning, deep learning, and optimization techniques in improving predictive accuracy in public health and biomedical domains16–18. These approaches, including hybrid and ensemble strategies, have shown effectiveness in uncovering complex relationships between health risk factors and outcomes16,18.
Hybrid machine learning architectures have shown promise for early detection tasks across domains. The CNN-LSTM framework proposed for crop disease surveillance demonstrates how ensemble methods can capture complex temporal patterns19. This reinforces the applicability of machine learning-based models for predicting childhood stunting, a similarly multifactorial health issue.
While machine learning has been applied to stunting prediction in other contexts13–15., this study advances the field by: (1) utilizing Ethiopia’s largest nationally representative dataset (EDHS 2011/2016), (2) introducing a multi-class severity classification (WHO HAZ thresholds: <-3SD severe, -3SD to -2SD moderate), and (3) quantifying specific risk factors (e.g., partner’s occupation, birth interval) previously overlooked in binary models. These contributions enable policymakers to prioritize interventions based on localized severity and determinants. Specifically, we aimed to answer the following questions:
What are the key determinant factors that contribute to stunting among children under five in Ethiopia?
Which ensemble machine learning algorithms demonstrate the highest predictive performance for classifying stunting status?
How effectively does the developed model perform in predicting the stunting status of children under five?
Related work
The problem of childhood stunting is a complex one, deeply intertwined with nutritional, socioeconomic, and environmental factors. As such, research on this topic has taken diverse paths, ranging from purely statistical analyses to the application of more advanced machine learning techniques. Within the specific context of Ethiopia, much of the early research adopted a primarily descriptive and statistical approach, focusing on identifying the most influential risk factors4,6,9–12.
These studies4,6,9–12 often relied on data extracted from the Ethiopian demographic and health surveys (EDHS), a valuable resource for understanding various health indicators across the nation. However, the focus of many of these studies was constrained by the available data at the time and the chosen methodology, which, while informative, presented certain limitations that the current study seeks to address.
A significant body of the existing Ethiopian research primarily utilized cross-sectional data, examining stunting prevalence and associated factors at a single point in time. Techniques like bivariate and multivariate logistic regression were frequently employed to quantify the relationship between various predictor variables and the presence of stunting.
These investigations4,6,9–12 identified a variety of determinants, such as maternal height, educational attainment, and breastfeeding duration. Additionally, household factors, like food security, sanitation, access to clean water, and overall socioeconomic status, were repeatedly highlighted.
Studies10–12exemplify this trend, focusing on geographically limited areas and exploring the impact of factors such as nutritional diversity and parental education. However, the scope of these studies was narrow, with datasets often limited to single regions or specific zones, rendering it challenging to make generalizations at the national level and implement policy changes for the whole country.
Furthermore, the cross-sectional nature of these studies was limiting, because it made it hard to determine causal relationships. For instance, while a study may indicate a correlation between stunting and lower educational attainment in mothers, it could not conclusively determine if the lack of education was the cause of the stunting or if other confounding factors were at play. Also, none of these studies developed a predictive model.
Recent studies highlight the increasing role of machine learning, deep learning, and optimization in enhancing prediction accuracy in public health and biomedical fields. Hybrid and ensemble methods effectively uncover complex links between health risk factors and outcomes16–18. This supports the use of machine learning models for predicting multifactorial issues like childhood stunting.
Advanced computational methods are increasingly vital in biomedical research. For instance, machine learning algorithms predict cardiovascular risk using electronic health records20, enabling timely interventions and improved patient outcomes. These technologies contribute to better understanding and management of complex diseases, reinforcing their value in modern healthcare.
Expanding beyond Ethiopia, researchers in Zambia, Papua New Guinea, and Pakistan have employed machine learning (ML) techniques in an effort to develop predictive models for identifying children at risk of stunting13–15. The methods used included: decision trees, support vector machines (SVMs), Random Forest, Extreme Gradient Boosting (XGBoost), and Naïve Bayes.
The study15used machine learning methods, including Random Forest and SVM to predict the probability of stunting. The study identified several significant risk factors for stunting among children under five in Zambia such as mother’s education and age of the child, and determined that Random Forest had the highest accuracy.
Similar efforts, reported by Children and attempted to classify the level of stunting by using a different machine learning technique13,14, like logistic regression, Random Forest, XGBoost, and neural networks. However, even these machine learning-based studies were not without limitations. A key shortcoming was that they frequently focused on binary classification, only predicting whether a child was “stunted” or “not stunted”, which didn’t fully address the range of stunting severity.
The study conducted by Asad13 did not include children under 6 months, which is not representative because children can be stunted at an early age and the study used a different classification for stunting prediction that includes moderate, marginal and severely stunted. Additionally, some studies did not incorporate determinant factors that are known to be influential, like birth interval and antenatal care visits. While these studies made strides in applying advanced modeling techniques, there were gaps in the modeling approach that did not account for all important factors.
This study, as outlined in the document, is poised to address the aforementioned limitations and offer a more comprehensive understanding and prediction capability for childhood stunting in Ethiopia. The authors have identified key gaps including the need for multi-class models, the lack of nationally representative data and the omission of specific determinant factors.
The study aims to directly address these gaps. Firstly, by moving beyond simple binary classification, the current study endeavors to develop a robust, multi-class predictive model capable of classifying stunting into three distinct categories: severe, moderate, and normal. We have used ensemble machine learning algorithms, which have not been widely used in prior Ethiopian studies on stunting prediction. This granular classification would facilitate more targeted interventions and help provide a more detailed understanding of the extent of the problem.
Secondly, the research utilizes a large dataset combining data from 2011 to 2016 EDHS surveys, which would provide national-level insights and improve the generalizability of the findings. This approach would make the findings more useful for policy formulation and intervention planning at a broader level. Thirdly, the current study makes a point of incorporating more specific determinants such as birth interval and antenatal care visits into the model that have been shown to be influential factors, but omitted in several studies and feature selection techniques.
In essence, the current study positions itself as a crucial step forward in understanding and combating childhood stunting by leveraging advanced machine learning techniques, utilizing comprehensive datasets, and developing practical applications to support health professionals.
Methods
Data source and Preparation
This study used secondary data from the 2011 and 2016 Ethiopian demographic and health surveys (EDHS), collected by the Ethiopian central statistical agency. The surveys included anthropometric, sociodemographic, and health-related data for women and children. We used responses that were related to under five children.
After data cleaning steps including the handling of missing values, and removing duplicates, the original data consisted of 18,451 instances with 28 features. The target variable “stunting status” was categorized into three classes: severely stunted (height-for-age Z-score <-3SD), moderately stunted (-3SD ≤ height-for-age Z-score < -2SD), and normal (height-for-age Z-score ≥-2SD), according to the WHO standards.
Data preprocessing
The initial raw datasets contained inconsistencies, missing values, and duplicates, therefore, an extensive data preprocessing was carried out to ensure data quality21. First, we addressed the missing values in various features using different approaches depending on the type of data.
To evaluate the impact of imputation, we compared model performance with and without imputation using Random Forest. Results showed negligible differences (accuracy difference < 9%), justifying our approach. Missing categorical values were filled using the mode22, while numerical values were imputed with the mean to maintain consistency across all ensemble models22.
Next, we removed any duplicate data instances, using drop_duplicates() pandas function, to avoid bias from similar data instances23. After handling missing values and duplicates, we checked for quasi-constant features to reduce the dimensionality of the data. We used variance threshold function for this purpose. Quasi constant features are features with the same value for a significant majority of the observations in the data set. Such features do not provide valuable information to the model, as they lack variability.
To make it suitable for machine learning, we discretized some numerical features. This procedure transforms continuous variables into discrete variables by grouping the values in to categories, and it helps to enhance the model performance and reduce complexity for model. The “child size at birth” attribute was re-binned into three categories: ‘small’, ‘average’, and ‘large’ by combining very small and smaller than average as small category, average as average category and very large and larger than average to large category; the wealth index was re-binned into three categories: ‘poor’, ‘middle’, and ‘rich’. This was achieved by grouping poorer and poorest as poor category, middle as middle category, and richer and richest as rich category; and toilet facilities were transformed into ‘improved’ and ‘unimproved’ categories.
Although CatBoost handles categorical features directly, we applied discretization to ensure consistency across all models, including Random Forest and AdaBoost, which require numerical inputs. This step also enhanced interpretability, reduced noise, and aligned with domain knowledge to improve performance and comparability across models.
Ordinal encoding was also applied to encode categorical variables into numerical representations suitable for machine learning algorithms, which ensure that each unique category has a unique corresponding numerical value. This encoding was used for variables such as religion and birth order.
Finally, the target variable “stunting status,” which was initially represented using standard deviations (<-3SD, -3SD to -2SD, >=2SD), was converted to categorical variables. For model training purposes, these categories were numerically coded: 0 for severely stunted, 1 for moderately stunted, and 2 for normal. These steps ensured that all data, including numerical and categorical attributes are suitable for machine learning models.
Class imbalance handling
The initial data set of 18,451 instances had an imbalanced class distribution with the majority of data categorized into “normal” stunting and smaller counts in the “moderately stunted,” and “severely stunted” categories. Such class imbalance can negatively impact machine learning models24, particularly for detecting minority class which is important in this study.
Before applying SMOTE, the class distribution 3415(18.5%), 3871(21%), and 11,165(60.5%) severe stunting, moderate stunting, and normal respectively. After SMOTE, each class contained approximately 11,165 records, achieving balance.
To address the class imbalance in the dataset, we applied the Synthetic Minority Over-sampling Technique (SMOTE). SMOTE was selected over random oversampling because it generates synthetic examples rather than duplicating existing ones, thereby reducing overfitting. Additionally, under sampling was not used, as it would remove valuable data from an already limited dataset25.
The over sampling of SMOTE was applied by setting the number of desired observations for each minority class. After applying SMOTE to the dataset, we expanded the dataset to a total of 33,495 instances. The balanced dataset with even distributions of cases in each stunting categories helps to produce more accurate model prediction outcomes without skew.
Feature selection
Feature selection was performed using filter and wrapper methods to select the most relevant features. Because Filter methods quickly rank features using statistical measures without involving models and wrapper methods use models to evaluate feature subsets, capturing feature interactions for better accuracy. As a filter, mutual information, chi-square test, Analysis of Variance F-test methods were applied, and a wrapper method, sequential forward selection (SFS), and sequential backward selection (SBS) techniques were applied using a Random Forest classifier.
Our approach aligns with bio-inspired optimization strategies in clinical research. They used snake optimization to streamline cardiovascular risk predictors while preserving model performance, mirroring our goal of identifying parsimonious stunting determinants20. Our feature selection strategy is conceptually aligned with nature-inspired optimization techniques proven effective in clinical data analysis. The Greylag Goose Optimization algorithm developed for lung cancer biomarker selection demonstrates how bio-inspired methods can enhance identification of critical health predictors26. We extend through our statistical feature selection framework for stunting determinants.
Based on step backward feature selection, a set of features that scored high accuracy were selected. In addition, we also incorporated essential features recommended by domain experts for the construction of the final model. Finally 18 features were selected.
The feature importance scores in Fig. 1 were computed using the Gini Importance (Mean Decrease in Impurity) metric. This method measures the contribution of each feature by evaluating how much the Gini impurity is reduced when a split is made using that feature. Higher values indicate more important features in predicting stunting status.
 |
1 |
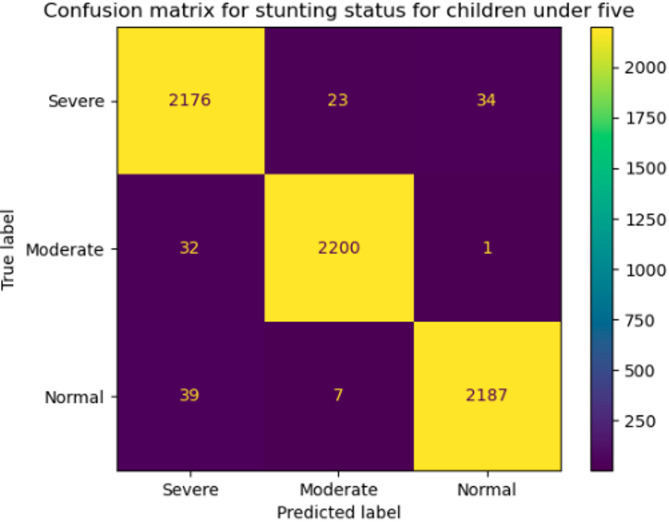
Fig. 1.
Confusion matrix.
where ΔGiniΔGini is the decrease in impurity.
Ensemble machine learning model development
Ensemble machine learning techniques have shown promise in health-related predictive modeling due to their ability to handle complex patterns and reduce overfitting. Recent studies have successfully applied such techniques to disease prediction16–18,27–29.
For instance, an explainable ensemble model for Parkinson’s diagnosis using optimized features18. Similarly, Parkinson’s Disease Explainable integrated boosting with selective features to improve diagnostic transparency16, while a multimodal framework explored feature optimization for AI-driven health insights17. These studies validate the effectiveness of ensemble models in biomedical contexts, reinforcing their use in our stunting prediction framework.
Four ensemble machine learning algorithms were used for model development: Random Forest, Adaptive Boosting(AdaBoost), Extreme Gradient Boosting (XGBoost), and CatBoost. These algorithms were chosen based on their effectiveness in handling complex datasets, capturing non-linear patterns, and strong performance for classification tasks27–29.
Model optimization follows best practices in epidemiological machine learning. BPSO framework for COVID-19 spread prediction illustrates how evolutionary algorithms can enhance prediction efficiency a strategy30 we adapt for parameter tuning in our ensemble models. Our hyperparameter optimization strategy aligns with metaheuristic approaches proven effective for biological data analysis. While modified-BER algorithm was designed for EEG signal classification, their success in optimizing model parameters for noisy physiological data reinforces the value of systematic tuning in health prediction tasks31.
A systematic hyperparameter optimization was performed for Random Forest, AdaBoost, XGBoost, and CatBoost using Scikit-learn’s RandomizedSearchCV with 10-fold cross-validation to ensure robust evaluation and reduce overfitting. For each model, 100 hyperparameter combinations were randomly sampled from predefined search spaces, focusing on key parameters like n_estimators, max_depth, and learning_rate due to their impact on model complexity, learning capacity, and generalization. Model performance was assessed using validation accuracy, and the best configurations for each model are presented in Table 1.
Table 1.
Hyper parameter tuning.
| Model | Best parameters |
|---|---|
| Random Forest | bootstrap = False, criterion=’entropy’, max_features=’log2’, min_samples_split = 2, n_estimators = 650, max_depth = 23, max_leaf_nodes = 7450 |
| AdaBoost | base_estimator = DecisionTreeClassifier(max_depth = None, max_leaf_nodes = 400,criterion=’gini’, splitter=’best’, min_samples_split = 10,min_samples_leaf = 2,max_features = None, n_estimators = 321,learning_rate = 0.5 |
| XGBOOST |
colsample_bytree = 0.6, learning_rate = 0.02, max_depth = 27, n_estimators = 326, reg_lambda = 0.005, reg_alpha = 0.0 |
| CatBoost | depth = 16, iterations = 290,l2_leaf_reg = 10,learning_rate = 1.0 |
The dataset was stratified into 80% training (26,796 instances) and 20% testing (6,699 instances) to ensure proportional representation of all stunting classes. All reported metrics (accuracy, precision, recall, f1-score.) are based on the testing set.
Evaluation metrics
The performance of the models was assessed using several evaluation metrics. These included: Accuracy, precision, recall, F1-score and receiver operating characteristic curve (ROC-AUC) were calculated to assess the models ability in correct classification of cases. And also Confusion matrix is used to analyze the number of correctly and incorrectly classified cases.
Here are the mathematical formulas for recall, precision, accuracy, and F1-score, all crucial metrics in evaluating machine learning models, particularly in classification tasks:
Formulas:
 |
2 |
 |
3 |
 |
4 |
 |
5 |
Where: TP = True Positives, TN = True Negatives, FP = False Positives, FN = False Negatives.
To complement the evaluation metrics, we computed 95% confidence intervals (CIs) for accuracy, precision, recall, and F1-score using bootstrapping with 1000 resamples on the test set. Additionally, we conducted pairwise statistical comparisons of model performance using McNemar’s test to assess whether differences in classification outcomes between models (Random Forest vs. other model) were statistically significant.
Results
Dataset characteristics and preprocessing outcome
The initial raw dataset comprised 18,451 instances with 28 features directly from the 2011 and 2016 rounds of the Ethiopian demographic and health surveys (EDHS) before applying preprocessing techniques including synthetic minority over-sampling (SMOTE). Data cleaning steps, such as imputation of missing values, removal of duplicates, and handling quasi-constant features, transformed the data, to ensure data quality for model training. The application of synthetic minority over-sampling technique (SMOTE) resulted in a balanced dataset comprising 33,495 instances with 18 features, and that was used for model development and evaluation.
The discretization of numerical features was a notable step in the preprocessing phase. Child size at birth was successfully re-binned into three categories (small, average, and large), wealth index was re-binned into three categories (poor, middle, and rich), and toilet facilities were reduced to two categories (improved and unimproved). The application of feature selection techniques led to a refined subset of 18 determinant features from the initial 28 features.
Predictive performance of ensemble machine learning models
The study evaluated the performance of four ensemble machine learning algorithms using data from the EDHS dataset: Random Forest, AdaBoost, XGBoost, and CatBoost. Table 1 presents the performance metrics, including accuracy, precision, recall, F1-score, and area under the receiver operating characteristic curve (ROC-AUC) of the four models based on the initial dataset before any class imbalance handling. The Random Forest and CatBoost models demonstrated better performance compared to other models.
Table 2 presents the performance metrics of the four models after applying SMOTE. SMOTE provided more balanced data and improved the overall model performance. The Random Forest algorithm achieved the highest accuracy of 97.746%, precision of 97.756%, recall of 97.748%, F1-score of 97.751%, and ROC-AUC of 99.99%. The performance increase, post class imbalance handling using SMOTE, is an indication of the effectiveness of re-sampling in improving model performance when the dataset is imbalanced Table 3.
Table 2.
Performance metrics of ensemble ML models before SMOTE sampling.
| Evaluation Metrics | Random Forest | AdaBoosting | XGboost | Catboost |
|---|---|---|---|---|
| Accuracy | 73.772% | 65.944% | 74.101% | 74.482% |
| Precision | 66.443% | 56.590% | 66.672% | 67.202% |
| Recall | 60.524% | 57.024% | 62.894% | 63.013% |
| F1_score | 62.023% | 56.793% | 64.303% | 64.542% |
| Roc curve | 98.711% | 97.432% | 95.184% | 95.533% |
Table 3.
Performance metrics of ensemble ML models after SMOTE sampling.
| Evaluation Metrics | Random Forest | AdaBoosting | XGboost | Catboost |
|---|---|---|---|---|
| Accuracy | 97.985% | 94.925% | 97.627% | 97.119% |
| Precision | 97.986% | 95.054% | 97.625% | 97.119% |
| Recall | 97.985% | 94.925% | 97.627% | 97.119% |
| F1_score | 97.954% | 94.946% | 97.626% | 97.119% |
| Roc curve | 99.995% | 99.673% | 99.984% | 99.977% |
| Cross validation | 99.998% | 99.900% | 98.410% | 97.200% |
The 95% confidence intervals for the performance metrics of Random Forest were: Accuracy: 97.985% [97.764%, 98.206%], Precision: 97.986% [97.745%, 98.227%], Recall: 97.985% [97.752%, 98.217%], F1-score: 97.954% [97.703%, 98.205%]. McNemar’s test comparing Random Forest and other model yielded a p-value < 0.05, indicating that the difference in classification performance was statistically significant.
Comparative analysis of model performance
A detailed comparative analysis of the four models highlights the superiority of Random Forest, not only in classification accuracy but also in computational efficiency. As shown in Table 2, Random Forest achieves the highest accuracy (97.985%), followed by XGBoost (97.627%), CatBoost (97.119%), and AdaBoost (94.925%).
One of the key advantages of Random Forest is its robust ensemble learning approach, which reduces overfitting by training multiple decision trees independently and averaging their predictions. Unlike boosting methods, which iteratively adjust weights and may overfit noisy data, Random Forest maintains high generalization performance with minimal hyper parameter tuning.
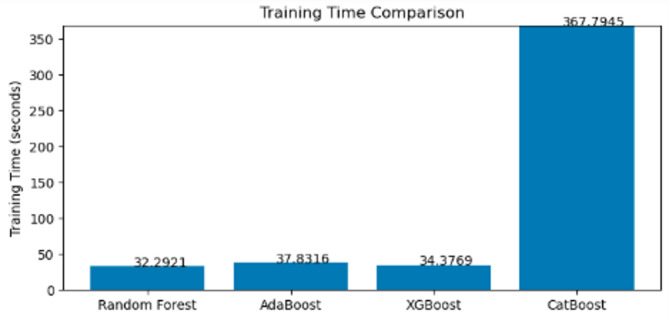
Additionally, Fig. 2 illustrates that Random Forest has the lowest training time (32 s), while CatBoost requires significantly more computation (367.79 s).
Fig. 2.
Training time comparison.
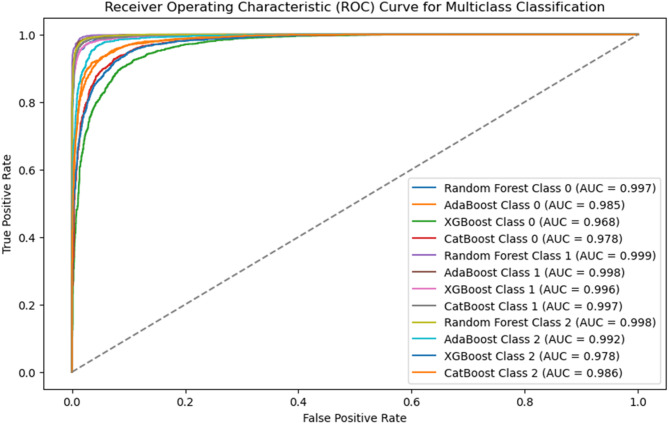
This efficiency is attributed to Random Forest’s parallelized tree construction, making it ideal for large datasets. Figure 3 further supports its superiority by presenting the ROC curves and AUC scores, where Random Forest was consistently outperforms the other models. The Random Forest model shows strong performance in classifying the Severe class. Out of 2,233 actual Severe cases, the model correctly predicted 2,176 as Severe, while misclassifying 23 as Moderate and 34 as Normal. This indicates a high level of accuracy in identifying Severe cases, with only a small proportion of errors. The performance for the Moderate and Normal classes is also impressive, as shown in Fig. 1.
Fig. 3.
ROC curve comparison.
The ability of Random Forest to handle high-dimensional data, and maintain stability across different dataset distributions makes it a highly effective and novel approach in this classification task.
Identification of key risk factors
While Random Forest itself is not novel, its application to this context provided unique interpretive insights into childhood stunting in Ethiopia. As shown in Fig. 4, which ranks the 18 features according to their relative importance, child age emerged as the most important factor, followed by child height and child weight, which represent basic anthropometrics that directly contribute to the stunting determination. Other highly important factors includes Children anemia level and birth interval indicating the importance of the general health of the child and its mothers.
Fig. 4.
Feature importance ranking.
Other factors like, partner occupation, and education are also among the important factors. Also the wealth index, region, religion, and water sources have a moderate impact on predicting the likelihood of stunting. The lower importance of birth order, breastfeeding duration, and place of residence, while still relevant, suggests that they are less influential compared to the other risk factors that we have considered.
The details of feature importance score is presented in Fig. 4. Importantly, the classification was based on the WHO definition of stunting, using height-for-age Z-scores: children with HAZ < -3 SD are severely stunted, between − 3 and − 2 SD are moderately stunted, and ≥ -2 SD are normal. By incorporating this clinical categorization into a machine learning framework, the model can accurately assign stunting levels based on multi-dimensional input features, going beyond binary classification used in most prior studies.
Moreover, the model’s interpretability measured through feature importance using Gini decrease allows for actionable conclusions. For example, the high importance of maternal education and birth spacing aligns with known intervention levers. This supports the use of such models not just for prediction, but for policy prioritization and health strategy design.
Discussion
This research sought to develop an effective predictive model for stunting status among Ethiopian children under five using ensemble machine learning algorithms, with a focus on practical applicability. The study is the first to apply multi-class classification using national EDHS data from two rounds (2011 and 2016), enabling context-specific policy guidance and overcoming prior limitations of geographically narrow studies in Ethiopia.
The Random Forest model, trained on a balanced dataset after applying SMOTE, achieved the highest predictive accuracy and demonstrated lower training time compared to other models. The results also revealed significant determinant factors of stunting and highlighted the potential for building intelligent systems to improve child health interventions.
The multi-class classification approach adopted in this study categorizing children as severely stunted, moderately stunted, or normal offers a more granular understanding of stunting severity. Contrasting with the binary approaches used in prior research13–15. This nuanced view allows for more targeted interventions. Compared to previous studies from Pakistan, Zambia, and Papua New Guinea13–15, our model outperforms in accuracy, achieving 97.98% with Random Forest, while those earlier binary classification approaches reported accuracies ranging from 72.8 to 98.5% using various ML algorithms.
The superior performance of the Random Forest algorithm can be attributed to its capacity for handling both numerical and categorical variables, resistance to overfitting, and support for parallel processing, which reduces training time. These attributes make Random Forest particularly effective for public health datasets with diverse variable types.
In addition to high performance, the model identified several key risk factors for stunting, including child age, maternal education, household wealth index, child anemia level, and access to clean water and sanitation.
These findings align with previous studies but also provide new evidence on the relative importance of less commonly addressed factors such as partner education, birth interval, and breastfeeding duration. These insights reinforce the need for holistic intervention strategies that go beyond nutritional support to address socioeconomic and environmental determinants.
Despite its strengths, this study also faces limitations. The use of secondary data restricts the inclusion of more specific variables such as feeding practices, local food security metrics, and sanitation quality. Furthermore, reliance on cross-sectional data prevents the analysis of causal relationships and long-term outcomes. The model has yet to be externally validated on data from.
Real-World deployment, policy implications, and future directions
While the Random Forest model shows strong predictive performance, real-world deployment presents challenges, including data privacy, ethical considerations, and limited infrastructure in rural areas. Ensuring secure data handling and compliance with protection regulations is crucial, especially in settings dealing with sensitive health data. Additionally, interpretability remains a barrier to adoption. Incorporating explainable artificial intelligence methods, such as SHAP values or visual dashboards, could enhance transparency and build trust among healthcare professionals and policymakers.
This model offers valuable tools for public health planning and resource allocation. By identifying high-risk children and the most impactful risk factors, it supports targeted health interventions particularly those related to maternal education, economic support, and access to clean water and antenatal care. These insights can inform policies aimed at reducing childhood stunting and improving health equity in Ethiopia.
Future research should explore real-time deployment through mobile and web applications to assist frontline health workers in early detection and intervention. Incorporating longitudinal data would provide a deeper understanding of growth patterns over time, while cross-country validation would strengthen the model’s generalizability. Integration with intelligent decision-support systems and ethical AI frameworks could further enhance the model’s impact and practical applicability. other countries or regions, which limits its generalizability.
Conclusion
This study demonstrated the effectiveness of ensemble machine learning algorithms, particularly the Random Forest model, in predicting childhood stunting in Ethiopia using EDHS data. The multi-class classification approach and the application of SMOTE to address class imbalance contributed to the model’s high predictive performance and its ability to identify key risk factors.
The model offers practical utility for early detection and targeted intervention, providing a data-driven foundation for policymakers and health professionals to combat childhood stunting more effectively. While the current model is robust and informative, further work is needed to ensure its deployment in real-world settings through interpretable and scalable systems.
Future developments should aim to integrate the model into real-time platforms, validate its use in diverse populations, and enhance transparency through explainable AI like shape and others. These enhancements can transform the model from a research tool into an actionable system for reducing stunting and improving child health outcomes in Ethiopia and beyond.
The findings provide actionable insights for national health authorities to design localized stunting interventions that prioritize key determinants such as maternal education, water access, and household wealth disparities.
Acknowledgements
We would like to acknowledge the Ethiopian central statistics for providing us with the data with a data set description.
Author contributions
Misganaw Ketema Ayele: Conceptualization, methodology, data curation, machine learning model development, software implementation, and corresponding author. Getachew Alemu Baye: Research design, literature review, statistical analysis, writing—original draft, and manuscript review. Seid Hassen yesuf: Supervision, validation, and critical review of methodology and results. Abebaw Agegne Engda: Data preprocessing, feature selection, and technical support for machine learning implementation. Eshetie Teka Mitiku: Review and editing, manuscript formatting, and final approval of the version to be published. All authors read and approved the final manuscript.
Data availability
The datasets analyzed during the current study are available from the Demographic and Health Surveys (DHS) Program repository (https://dhsprogram.com/data/available-datasets.cfm ). Access to the EDHS data requires registration and approval from the DHS Program. Researchers can request access through the DHS Program’s data portal.
Declarations
Competing interests
The authors declare no competing interests.
Ethics declaration
not applicable because of we have used public available data by requesting to use the data.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Solomons, N. W. Vision of research on human linear growth. Food Nutr. Bull.40 (4), 416–431 (2019). [DOI] [PubMed] [Google Scholar]
- 2.Watson, K. M. et al. Height-age as an alternative to height-for-age z-scores to assess the effect of interventions on child linear growth in low-and middle-income countries. Curr. Dev. Nutr.8 (12), 104495 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Organization, W. H. Reducing stunting in children: equity considerations for achieving the Global Nutrition Targets 2025, (2018).
- 4.Fenta, H. M., Workie, D. L., Zike, D. T., Taye, B. W. & Swain, P. K. Determinants of stunting among under-five years children in Ethiopia from the 2016 Ethiopia demographic and health survey: application of ordinal logistic regression model using complex sampling designs. Clin. Epidemiol. Glob Health. 8 (2), 404–413 (2020). [Google Scholar]
- 5.Ephi, I. Ethiopia mini demographic and health survey 2019: key indicators, Rockville, Maryland, USA: EPHI and ICF, (2019).
- 6.Gobena, W. E., Wotale, T. W., Lelisho, M. E. & Gezimu, W. Prevalence and associated factors of stunting among under-five children in Ethiopia: application of marginal models analysis of 2016 Ethiopian demographic and health survey data. PLoS One. 18 (10), e0293364 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Atalell, K. A., Techane, M. A., Terefe, B. & Tamir, T. T. Mapping stunted children in Ethiopia using two decades of data between 2000 and 2019. A Geospatial analysis through the bayesian approach. J. Health Popul. Nutr.42 (1), 113 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Laillou, A. et al. Wasted children and wasted time: a challenge to meeting the nutrition sustainable development goals with a high economic impact to Ethiopia. Nutrients12 (12), 3698 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fantay Gebru, K., Mekonnen Haileselassie, W., Haftom Temesgen, A., Oumer Seid, A. & Afework Mulugeta, B. Determinants of stunting among under-five children in Ethiopia: a multilevel mixed-effects analysis of 2016 Ethiopian demographic and health survey data. BMC Pediatr.19, 1–13 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tola, G., Kassa, A., Getu, M., Dibaba, B. & Neggesse, S. Prevalence of stunting and associated factors among neonates in Shebadino woreda, Sidama region South Ethiopia; a community-based cross-sectional study 2022. BMC Pediatr.23 (1), 276 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gudeta, H. T., Nagari, S. L., Dadi, D. G., Abdulahi, T. & Abose, S. Predictors of Stunting among 6–35 Months Old Children in Assosa Zone, Northwest Ethiopia: Unmatched Case–Control Study, Adv Public Health, vol. no. 1, p. 3491977, 2023. (2023).
- 12.Mengesha, A., Hailu, S., Birhane, M. & Belay, M. M. The prevalence of stunting and associated factors among children under five years of age in Southern Ethiopia: community based cross-sectional study. Ann Glob Health, 87, 1, (2021). [DOI] [PMC free article] [PubMed]
- 13.Asad, M. & Zouq, A. A Machine Learning Approach for Predicting Stunting in Under Five Children (The Case of Pakistan Demographic and Health Survey, 2024).
- 14.Shen, H., Zhao, H. & Jiang, Y. Machine learning algorithms for predicting stunting among under-five children in Papua new Guinea. Children10 (10), 1638 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chilyabanyama, O. N. et al. Performance of machine learning classifiers in classifying stunting among under-five children in Zambia. Children9 (7), 1082 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Khanom, F., Uddin, M. S. & Mostafiz, R. PD_EBM: an integrated boosting approach based on selective features for unveiling Parkinson’s disease diagnosis with global and local explanations. Eng. Rep.7 (1), e13091 (2025). [Google Scholar]
- 17.Khanom, F., Mostafiz, R. & Uddin, K. M. M. Exploring multimodal framework of optimized Feature-Based machine learning to revolutionize the diagnosis of Parkinson’s disease: AI-Driven insights. Biomedical Mater. & Devices, pp. 1–20, (2025).
- 18.Khanom, F., Biswas, S., Uddin, M. S. & Mostafiz, R. XEMLPD: an explainable ensemble machine learning approach for Parkinson disease diagnosis with optimized features. Int. J. Speech Technol.27 (4), 1055–1083 (2024). [Google Scholar]
- 19.Alzakari, S. A., Alhussan, A. A., Qenawy, A. S. T. & Elshewey, A. M. Early detection of potato disease using an enhanced convolutional neural network-long short-term memory deep learning model. Potato Res, pp. 1–19, (2024).
- 20.Tarek, Z., Alhussan, A. A., Khafaga, D. S., El-Kenawy, E. S. M. & Elshewey, A. M. A snake optimization algorithm-based feature selection framework for rapid detection of cardiovascular disease in its early stages. Biomed. Signal. Process. Control. 102, 107417 (2025). [Google Scholar]
- 21.Ahmad, T. & Aziz, M. N. Data preprocessing and feature selection for machine learning intrusion detection systems. ICIC Express Lett.13 (2), 93–101 (2019). [Google Scholar]
- 22.Zhang, Z. Missing data imputation: focusing on single imputation. Ann Transl Med, 4, 1, (2016). [DOI] [PMC free article] [PubMed]
- 23.MAITY, S., PATTANAYAK, S., DEY, D., MITRA, A. & INNOVATIONS IN AUTOMATIC QUESTION GENERATION AND ANSWER PREDICTION. : A COMPREHENSIVE REVIEW OF THE LATEST RESEARCH AND TECHNIQUES.
- 24.Tran, N., Chen, H., Jiang, J., Bhuyan, J. & Ding, J. Effect of class imbalance on the performance of machine learning-based network intrusion detection. Int. J. Perform. Eng.17 (9), 741 (2021). [Google Scholar]
- 25.Elreedy, D. & Atiya, A. F. A comprehensive analysis of synthetic minority oversampling technique (SMOTE) for handling class imbalance. Inf. Sci. (N Y). 505, 32–64 (2019). [Google Scholar]
- 26.Elkenawy, E. S. M., Alhussan, A. A., Khafaga, D. S., Tarek, Z. & Elshewey, A. M. Greylag Goose optimization and multilayer perceptron for enhancing lung cancer classification. Sci. Rep.14 (1), 23784 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Fatima, S., Hussain, A., Bin Amir, S., Ahmed, S. H. & Aslam, S. M. H. Xgboost and random forest algorithms: an in depth analysis. Pak. J. Sci. Res.3 (1), 26–31 (2023). [Google Scholar]
- 28.Amirudin, N. & Abdulkadir, S. J. Comparative study of machine learning algorithms using the CICIOV2024 dataset. Platform: J. Sci. Technol.7 (1), 1–8 (2024). [Google Scholar]
- 29.Almahdi, A. et al. Boosting ensemble learning for freeway crash classification under varying traffic conditions: A hyperparameter optimization approach. Sustainability15 (22), 15896 (2023). [Google Scholar]
- 30.Alkhammash, E. H. et al. Application of machine learning to predict COVID-19 spread via an optimized BPSO model. Biomimetics8 (6), 457 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Elshewey, A. M., Alhussan, A. A., Khafaga, D. S., Elkenawy, E. S. M. & Tarek, Z. EEG-based optimization of eye state classification using modified-BER metaheuristic algorithm. Sci. Rep.14 (1), 24489 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets analyzed during the current study are available from the Demographic and Health Surveys (DHS) Program repository (https://dhsprogram.com/data/available-datasets.cfm ). Access to the EDHS data requires registration and approval from the DHS Program. Researchers can request access through the DHS Program’s data portal.