Abstract
Italian genetic history was profoundly shaped by the Romans. While the Iron Age Central Italian gene pool was comparable to that of other coeval Central/Western European regions, during the Imperial age, it was significantly influenced by Eastern Mediterranean ancestries. To explain this genetic shift, it has been proposed that people from eastern provinces of the Empire migrated toward its political center: Rome and its surroundings. In this study, by presenting a new Roman Republic individual (1.25×) and comparing it to other published Republican samples, we propose a novel perspective for the presence of the Eastern Mediterranean ancestry in the Imperial gene pool. We show that the spread of this genetic ancestry may have taken place earlier than previously thought, during the Late Republican period, therefore predating the onset of the Empire by ∼200 years. The diffusion of this ancestry occurred due to early East-to-West movements, either because Eastern Mediterranean regions were under Roman political influence since the Late Republican period or even as a result of internal movements from Southern Italy where several Greek and Phoenician settlements were established during the 1st millennium BCE.
Keywords: Eastern Mediterranean, Central Italy, ancient DNA, archaeogenetics, Roman Republic
Significance.
Compared with the previous period, the Imperial Roman gene pool of Central Italy is characterized by a sharp shift toward Eastern Mediterranean regions, usually explained as the genetic consequence of migrations from the eastern provinces mainly driven by the attractive power of Rome. In this article, we present a high-quality genome of a new Central Italian Roman individual, dated before the onset of the Empire, in the Late Republican period. Interestingly, this sample already shows signs of the Eastern Mediterranean genetic ancestry. By analyzing it with other published Central Italian Republican samples with a similar genetic makeup and framing our results in the historical and archaeological context, we suggest that the genetic shift toward the Eastern Mediterranean ancestry was an ongoing process during the Late Republican period, predating the onset of the Empire by ∼200 years.
Introduction
The Italian Peninsula, due to its central position, has been a melting pot for the people living across the whole Mediterranean Sea (Abulafia 2011; Broodbank 2013). The study of ancient DNA (aDNA) greatly enhanced our comprehension of the movements of individuals within Italy and their impact on the local gene pool (Aneli et al. 2021). Overall, all these migrations resulted in frequent shifts in the genetic landscape of the peninsula, starting from the Paleolithic (Posth et al. 2023) to modern times (Antonio et al. 2019; Raveane et al. 2019; Posth et al. 2021).
A pivotal role in the cultural and genetic evolution of Italy has been played by the Romans. The people that inhabited the Italian Peninsula during the previous Iron Age (IA) were characterized by a great cultural diversity (Pallottino 1991; Bietti Sestieri 2018) and a relatively low genetic diversity (Serventi et al. 2018; Antonio et al. 2019; Posth et al. 2021; Aneli et al. 2022; Ravasini et al. 2024). On the contrary, with the conquest of the peninsula, the Romans imposed their culture, language, institutions, and social aspects, eventually influencing even the evolution of the gene pool (Antonio et al. 2019; Posth et al. 2021; Scorrano et al. 2022; Coia et al. 2023; Moots et al. 2023; Antonio et al. 2024; Pilli et al. 2024; Ravasini et al. 2024).
The spread of the Roman domination began during the Republican period (509 to 27 BCE), a fundamental stage of the history of Rome in which the small city-state became a hegemonic power, not only in Italy but also across the whole Mediterranean Sea. During their conquest of the peninsula, the Romans asserted their power over Magna Graecia, the Greek colonies established from the 8th century BCE in Southern Italy. These colonies reached elevated levels of political, cultural, and societal development, even competing with their motherland (Brizzi 1997; van Dommelen 2012; De Angelis 2016; Hodos 2020). The political integration of Magna Graecia greatly influenced Roman society, eventually resulting in major cultural transformations. In the next centuries, the territories conquered and embedded in the Roman state steadily increased, coming into close contact with different people, which, like the Southern Italian Greeks, contributed to the enrichment of the social substrate of Roman society. At the onset of the Empire (27 BCE), Rome was a powerful and cosmopolitan state encompassing the Mediterranean area.
Besides the political and cultural influence, the Roman Empire (27 BCE to 476 CE in Italy) had a great impact also on the genomic landscape of the conquered territories, especially in Italy. Indeed, in the Imperial time, a sharp shift toward Eastern Mediterranean ancestry can be observed, compared to the previous Italian IA populations (Antonio et al. 2019; Posth et al. 2021; Aneli et al. 2022; Scorrano et al. 2022; Coia et al. 2023; Pilli et al. 2024; Ravasini et al. 2024). This great Eastern Mediterranean genetic influence has been mostly explained as the result of massive migrations during the Imperial time from Eastern regions into Rome (Antonio et al. 2019; Lazaridis et al. 2022) and other regions of the peninsula (Posth et al. 2021), possibly triggered by the high population density of Eastern Imperial provinces and/or the attractiveness of a power center such as Rome (Scheidel et al. 2007; Antonio et al. 2019). It is important to note that in this article, we use the term Eastern Mediterranean ancestry/genetic influence with its broadest geographic connotation, encompassing all the Mediterranean territories from present-day Greece eastward. Indeed, unfortunately, with the data currently available, it is not always possible to clearly distinguish more specific contributing regions (such as the Aegean or the Levant), considering that their differences are nuanced, and possibly movements from the entire macroregion may have contributed to this genetic influence in Central Italy.
Despite the current interpretation, historical and genetic considerations may point to an earlier arrival of the Eastern Mediterranean genetic influence in Central Italy, before the onset of the Roman Empire. First, Rome annexed some of the rich and densely populated Eastern territories decades, if not centuries, before the onset of the Empire (Macedonia and Greece between 168 and 146 BCE; Western part of Anatolia in 133 BCE and soon afterward Cilicia, the Southern part of Anatolia, from 100 BCE; and Syria in 64 BCE (Piganiol 1927; Rinaldi Tufi et al. 1971; Brizzi 1997; Fig. 1). This indicates that migrations from those regions might have started much earlier. Second, the individuals genetically analyzed so far and dated to the early years of the Empire already show a nonnegligible amount of Eastern Mediterranean ancestry. For example, the genomes retrieved from the archaeological site of Pompeii (Scorrano et al. 2022; Pilli et al. 2024) (which, due to the eruption that destroyed the city, are precisely dated to the 79 CE) cluster with other later Imperial individuals. They also show an excess of Iran Neolithic and/or Caucasus hunter-gatherer genetic components, usually good proxies for Eastern Mediterranean ancestries in Central and Southern Europe because they are found at high proportions in post-Chalcolithic Eastern Mediterranean groups (Skourtanioti et al. 2020; Lazaridis et al. 2022; Koptekin et al. 2023; Antonio et al. 2024). Although not impossible, it seems unlikely that migrations bringing the Eastern Mediterranean ancestry had an impact on the Italian gene pool in just a few generations since the onset of the Empire. In this context, it is important to note that the Eastern Mediterranean genetic component may have arrived in Central-Northern Italy as a result of internal migrations after the conquest of genetically understudied Southern Italy and Sicily IA. Indeed, while some studies highlighted the role of Greek and Punic settlements in Western Mediterranean islands in shaping the local gene pool (Fernandes et al. 2020; Marcus et al. 2020; Reitsema et al. 2022; Ringbauer et al. 2025), we are still lacking an extensive genomic characterization of Southern Italian Magna Graecia individuals. These groups probably carried Eastern Mediterranean ancestry since they originally came from the aforementioned regions. Finally, among the few analyzed individuals dated to the latest period of the Roman Republic (the last two centuries of the 1st millennium BCE), there are several ones interpreted as “genetic outliers” with an Eastern Mediterranean putative origin (Antonio et al. 2019; Posth et al. 2021; Moots et al. 2023). To better shed light on this fundamental shift of the Italian gene pool, we performed a medium coverage sequencing (1.25×) of a Late Republican individual from the site of Villa Falgari, located in Tarquinia, Viterbo (only 70 km away from Rome; Fig. 1). By analyzing our new sample in the context of coeval individuals with similar genetic makeup, we show that the gene flow bringing Eastern Mediterranean ancestry in Central Italy was probably already occurring during the Late Republican phase, possibly indicating these “genetic outliers” as the representatives of the ongoing arrival of Eastern Mediterranean ancestry, which later characterized the genomic landscape of the Imperial period.
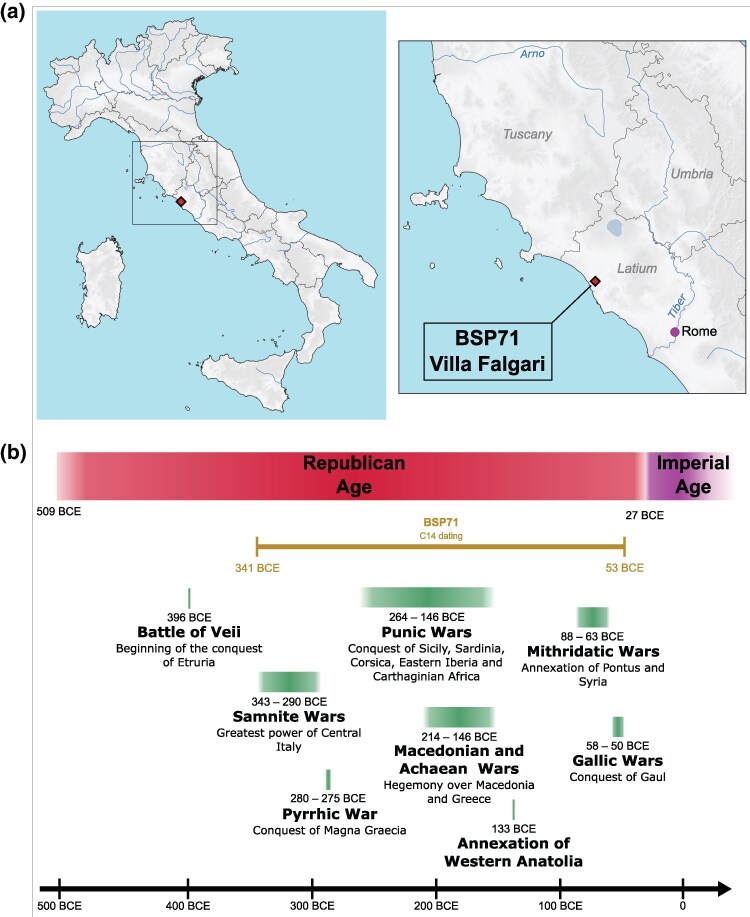
Fig. 1.
a) Geographic location of Villa Falgari, where the individual BSP71 was discovered. b) Timeline of the Roman Republic and the first years of the Empire. Two sigma-calibrated radiocarbon dating of the sample BSP71 from Villa Falgari and the time period of some of the main wars and annexations of the Republic are indicated below the timeline.
Results
Villa Falgari in the Central Italian Genetic Context
We extracted DNA from an individual (BSP71) found in the site of Villa Falgari, Tarquinia (Fig. 1a; Table S1) (Barich et al. 1968). Radiocarbon dating places this individual in the Republican period (341 to 53 2σ calBCE, 2126 ± 24 BP), likely in the latest phase of the Republic, after the beginning of Roman expansionism in the Eastern Mediterranean (Fig. 1; Table S2). Shotgun sequencing was performed resulting in a depth of coverage of 1.25×. The sequences were confirmed to be ancient with negligible amount of contamination, and a male genetic sex was assigned for this individual (Table S1). Genotype data were merged with ancient and modern individuals included in the AADR v54.1 (Mallick et al. 2024; Mallick and Reich 2024) and in Reitsema et al. (2022) (Table S3).
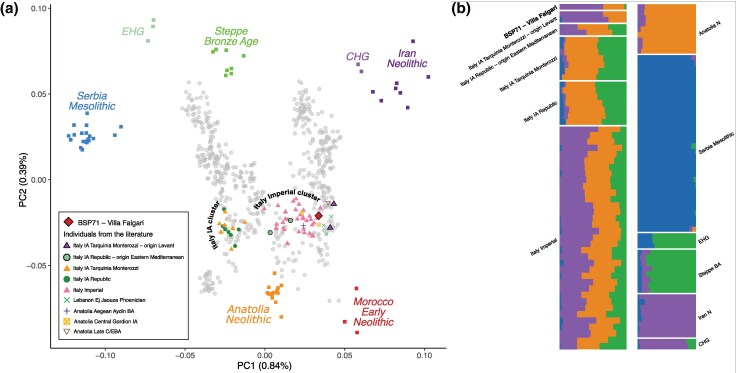
Principal component analysis (PCA) was performed to contextualize the Villa Falgari site in the genetic variability of Western Eurasia and North Africa. BSP71 lies outside the main cluster of Italian IA/Republican individuals, being strongly shifted toward Eastern Mediterranean ancient and modern individuals and overlapping with the genetic variability of the subsequent Imperial period (Fig. 2a; Fig. S1). Interestingly, other IA/Republican individuals reported in previous studies show a similar genetic shift (Fig. 2a; Fig. S1) (Antonio et al. 2019; Moots et al. 2023). In particular, two individuals from Tarquinia (only a few kilometers far from Villa Falgari and likewise dated to the Late Republican phase labeled as Italy_TarquiniaMonterozzi_IA_oLevant) are placed in the most Eastern extreme of the distribution of the PCA, similar to IA Levant groups like Phoenicians (Moots et al. 2023) (Fig. 2a; Fig. S1). Similarly, multidimensional scaling (MDS) analysis based on 1 − f3 statistics matrix between relevant populations highlights close genetic proximity between Villa Falgari, Italy_TarquiniaMonterozzi_IA_oLevant, Bronze Age individuals from Aegean Anatolia and Phoenicians (Fig. S2). Admixture analysis (Alexander et al. 2009) reflects these findings, showing a high proportion of Iran Neolithic/Caucasus hunter-gatherer (Iran_N/CHG) genetic component in aforementioned individuals and in Villa Falgari, while it has negligible proportion in the main IA/Republic cluster (Fig. 2b; Fig. S3). Similarly, Imperial time individuals show a high proportion of Iran_N/CHG genetic component, suggesting some kind of genetic connections between this period and the IA/Republican samples with Eastern Mediterranean ancestry (hereafter indicated as IA/Republic oEM, as for “outlier Eastern Mediterranean”) (Fig. 2b).
Fig. 2.
a) PCA plot with the newly reported individual and a subset of the used relevant ancient (colored) and modern (gray) samples from the literature (see Fig. S1 for the plot with all the samples included in the analysis). The main Italian IA and Imperial clusters are indicated, together with the IA individuals with Eastern Mediterranean ancestry. b) Unsupervised admixture analysis (K = 4), on the left, Italian IA and Imperial individuals are represented; on the right, populations representative of the main European genetic ancestries.
D-statistics was used to confirm the greater affinity of the Imperial and the IA/Republic oEM individuals to Iran_N ancestry compared to the main IA/Republic cluster (Fig. S4; Table S4a). IA/Republic oEM and Imperial groups tend to share significantly more derived alleles with Iran_N compared to the IA/Republic main cluster. On the other hand, when Imperial and IA/Republic oEM groups are compared, no significant difference can be observed, indicating a similar influence of the Iran_N ancestry. These results suggest genetic continuity between these groups and indicate that they form a homogeneous genetic cluster in their affinity to Iran_N when they are compared to the IA/Republican main cluster. Although it was not possible to finely identify the source of the Eastern Mediterranean genetic component in Villa Falgari using only D-statistics, some tests performed point to a greater affinity to Anatolian Bronze and IA groups with respect to IA/Republican main cluster (see for examples tests where |Z| ≥ 2 in Table S4b).
Interestingly, also the Y chromosome analysis comparing ancient individuals with modern ones shows clues of an Eastern Mediterranean ancestry in BSP71 and the other four IA/Republic oEM (R437, R10337, R10341, and R850) individuals (Fig. S5; Table S5) (Karmin et al. 2015; Mallick et al. 2016; Poznik et al. 2016; Martiniano et al. 2022). BSP71 belongs to the R1-M269/Z2105 haplogroup and clusters with several Eastern individuals, mostly from Late Bronze Age/IA Armenia. The R437 individual belongs to a sister clade (R1-M269/P312), now frequent in Western Europe but also observed in Bronze Age Croatians. R10337 belongs to the R2-M479 clade, observed in present-day Western Asian groups and in ancient Middle Eastern people. Finally, R10341 and R850 belong to the J1-M267/P58 and T-M70 haplogroups, respectively. These lineages are now observed mostly in Middle Eastern populations, but they were also present among ancient Eastern Mediterranean individuals, mostly from Greece and Anatolia. The mitochondrial DNA (mtDNA) haplogroup of BSP71 (T1a1; Table S1) is found in an individual from the Greek colony of Himera in Sicily, possibly indicating a genetic connection on the maternal side (Reitsema et al. 2022). However, the wide distribution of this haplogroup all over Eurasia in ancient and modern individuals does not allow more precise reconstructions.
The Spread of Eastern Mediterranean Ancestry in Central Italy
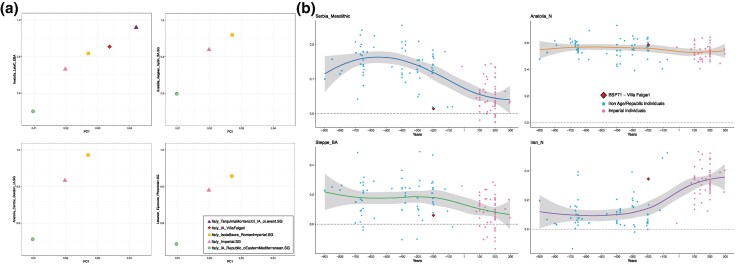
To shed light on the origin of the Eastern Mediterranean ancestry in Central Italy, we explored more in detail these results by performing different tests of qpAdm modeling (Harney et al. 2021). In particular, we tested if the Imperial gene pool and the IA/Republic oEM individuals can be modeled as a mixture between local IA gene pool and Eastern Mediterranean populations or whether they are genetically originating entirely from the Eastern Mediterranean regions. The results indicate that the IA/Republic oEM and the Imperial groups can be modeled either with a two-source model including the main cluster IA/Republican (i.e. Italy_IA_Republic) and an Eastern Mediterranean group (Fig. 3a; Table S6) or as deriving their ancestry directly from these Eastern groups (Table S7). In particular, the feasibility and the admixture proportions of these models correlate with the distribution of targets and sources along the first component of the PCA, which reflects their distance from the Iran_N individuals (Fig. 3a; Table S6). When using as Eastern source the group that shows the greater affinity to Iran_N among the ones tested (i.e. Anatolia_LateC_EBA, a Late Chalcolithic/Early Bronze Age group from Şırnak in Southeastern modern Turkey), the model fits for all the targets analyzed, showing a descending gradient for the value of this ancestry going from right to left along the PC1: Italy_TarquiniaMonterozzi_IA_oLevant (0.959 ± 0.08); Villa Falgari (0.854 ± 0.09); Italy_IsolaSacra_RomanImperial (0.817 ± 0.059); Italy_Imperial (0.731 ± 0.049); and Italy_IA_Republic_oEasternMediterranean (0.501 ± 0.082) (Fig. 3a; Table S6).
Fig. 3.
a) Comparisons of qpAdm results (Table S6) and the position of targets along the PC1 of the PCA (Fig. 2; Fig. S1). The four plots show different two-source qpAdm models where one of the sources is always Italy_IA_Republic.SG as a proxy of the typical IA/Republican gene pool, while the other is represented by different Eastern Mediterranean populations (shown on the y axes). When the sources Anatolia_Aegean_Aydin_BA.SG, Anatolia_Central_Gordion_IA.SG, and Lebanon_EjJaouze_Phoenician.SG are used, Italy_TarquiniaMonterozzi_IA_oLevant.SG and Italy_IA_VillaFalgari cannot be modeled and so they are not reported in the respective plots (Table S6). However, they can be modeled as deriving their ancestry entirely from these Eastern Mediterranean sources (see the one source qpAdm models in Table S7). b) Contribution of the four main sources that contributed to the peopling of Europe (Serbia Mesolithic, Anatolia Neolithic, Steppe Bronze Age, and Iran Neolithic) in Central Italian individuals from 900 BCE to 300 CE, modeled with qpAdm analysis. Component proportions are on the y axes and time on the x axes of the single plots. LOESS smooth lines indicate the trend of the component proportions over time (0.95 confidence interval represented by gray area). A sharp increase of the Iran Neolithic source can be observed starting from ∼200 BCE.
On the other hand, models including Eastern sources less shifted toward Iran_N in the PCA do not work for all the groups under study. Indeed, the models including Lebanon_EjJaouze_Phoenician, Anatolia_Aegean_Aydin_BA, or Anatolia_Central_Gordion_IA are feasible exclusively for the Imperial groups and Italy_IA_Republic_oEasternMediterranean with different proportions of the Eastern sources. Italy_IA_Republic_oEasternMediterranean, i.e. the group farther from Eastern Mediterranean groups in the PCA, has the lowest proportion of the Eastern Mediterranean components (0.486 ± 0,072, 0.598 ± 0.113, and 0.512 ± 0.094 for Lebanon_EjJaouze_Phoenician, Anatolia_Aegean_Aydin_BA, and Anatolia_Central_Gordion_IA, respectively) (Fig. 3a; Table S6), while Imperial groups (Italy_Imperial and Italy_IsolaSacra_RomanImperial) show higher proportions (0.780 ± 0.051 and 0.857 ± 0.059, respectively, for the Lebanon_EjJaouze_Phoenician component; 0.839 ± 0.073 and 0.919 ± 0.085, respectively, for the Anatolia_Aegean_Aydin_BA; and 0.833 ± 0.078 and 0.972 ± 0.104, respectively, for the Anatolia_Central_Gordion_IA) (Fig. 3a; Table S6). On the contrary, these models are not feasible for the more Eastern-shifted Republican individuals (BSP71 and Italy_TarquiniaMonterozzi_IA_oLevant). Indeed, they can be modeled as deriving their entire ancestry from Eastern Mediterranean populations (Table S7), consistent with their position along the PC1 (Fig. 2a) and their genetic proximity to Anatolian Aegean BA individuals in the 1 − f3 MDS analysis (Fig. S2). Overall, these findings confirm the Eastern Mediterranean genetic influence in all the analyzed groups and suggest that the gene pool of the Late Republican phase and Imperial period emerged thanks to the gene flow from Eastern regions into Central Italy, with some Late Republican individuals (like the Villa Falgari and the Italy_TarquiniaMonterozzi_IA_oLevant ones) that may have genetically originated entirely from Eastern Mediterranean regions.
Moreover, the Imperial groups (Italy_Imperial and Italy_IsolaSacra_RomanImperial) can be mostly modeled as a mixture between groups of the main IA/Republican cluster (i.e. Italy_IA_Republic) and Villa Falgari or Italy_TarquiniaMonterozzi_IA_oLevant (Fig. S6; Table S8), with a high proportion (∼70% to 80%) of the latter component. This indicates that the Eastern Mediterranean ancestry in the Imperial gene pool may have arrived thanks to admixture events between local IA people and individuals similar to the more Eastern-like IA/Republic oEM ones arrived during the Late Republican phase. This would eventually indicate that the Eastern Mediterranean genetic influence in Central Italy began to spread during the Late Republic rather than at the onset of the Empire.
We further investigated this possibility by performing qpAdm on single individuals from Central Italy dated between 900 BCE and 300 CE (therefore encompassing the early IA, the Republican and first Imperial period) with a more general model using four source populations, which contributed to the peopling of Europe: Serbia_Mesolithic, Anatolia Neolithic (Anatolia_N), Steppe Bronze Age (Steppe_BA), and Iran_N (Fig. 3; Table S9). While the Anatolia_N component (present in Italy as soon as the early stages of the Neolithic) remains stable over the centuries, the Iran_N one sharply increases starting from 200 BCE at the expense of the Steppe_BA and the Serbia_Mesolithic components (Fig. 3). This result is mirrored by the position of these samples along the first component of the PCA, showing a strong increase in their affinity toward Eastern Mediterranean populations as approaching to the Late Republican and Imperial period (Fig. S7). Despite the small number of samples belonging to the first two centuries BCE, this result may suggest that it is possible to frame the beginning of the spread of Eastern Mediterranean genetic influence in Central Italy during the Late Republican phase rather than in the Imperial period.
Finally, we investigated the genetic similarity of BSP71 to ancient and modern populations from Europe and the Mediterranean basin and tested the putative legacy of this individual (or similar ones) on the modern gene pool. We performed an outgroup f3 statistic in the form f3(BSP71, Test; Onge) where Tests are Eurasian groups comprising a 1,000-year range centered around the average date of Villa Falgari individual (therefore including Imperial individuals) or modern populations (Fig. S8; Table S10). This analysis shows that for ancient populations, the area with greater genetic similarity to BSP71 is the Central-Eastern Mediterranean, while for modern populations, this area shifts toward Central-Southern Europe, suggesting that individuals like BSP71 may represent the first steps of the formation of the modern European gene pool.
Discussion and Conclusions
The main purpose of this article is to provide a new framework for the studying of population dynamics in Central Italy at the turning point between the IA/Republican and the Imperial period, by combining archaeogenetic and historical data. In this study, we analyzed a Late Roman Republican individual from the site of Villa Falgari, and, reanalyzing published individuals from the same area with similar genetic makeup, we suggest a new perspective on the diffusion of the Eastern Mediterranean genetic influence in Central Italy. Given the presence of a high proportion of this ancestry in Late Republican individuals, like Villa Falgari, here, we propose that the observed shift in Central Italian gene pool toward Eastern Mediterranean genetic components was not a consequence of the establishment of the Roman Empire, but rather an ongoing process began probably 200 years before. Movements of people from Eastern regions may have already occurred after the end of the Punic Wars, when Rome became the hegemonic power of the Mediterranean Sea, and they probably increased when Macedonia, Greece, and Anatolia were annexed to the Republic in the 2nd century BCE. Moreover, it is even plausible that part of the Eastern Mediterranean genetic influence arrived earlier in Central Italy as a consequence of internal migrations along the peninsula. Indeed, the Roman conquest of Southern Italy, before divided between Magna Graecia and Phoenicians, may have favored the spread of individuals who already carried Eastern Mediterranean ancestry.
Although at the moment few IA samples from Southern Italy have been analyzed, some hints suggest that their gene pool was characterized by Eastern Mediterranean ancestries. Middle and Late Bronze Age individuals from Sicily already have the Iran Neolithic genetic component (Fernandes et al. 2020), although not to the extent of the Central Italian Imperial period, and the subsequent IA autochthonous people of the Sicani seem to be in genetic continuity with Bronze Age populations (Fig. S1) (Reitsema et al. 2022). It is, however, among the few samples available from the Greek colony of Himera that it is possible to observe individuals with a genetic makeup more similar to Villa Falgari and the Imperial individuals (Table S6) (Reitsema et al. 2022). It is therefore possible that Magna Graecia did not influence only the cultural aspects of the Roman world but also contributed to its population dynamics. With the extensive analysis of new Bronze and IA individuals from Southern Italy, especially from a Greek cultural background, it will be possible to finely place in space and time the origin of the Eastern Mediterranean ancestry in Central Italy.
Italy has always been a place of great genetic variability in the European context (Raveane et al. 2019; Aneli et al. 2021), and the IA/Republican period does not represent an exception (Antonio et al. 2019; Posth et al. 2021; Aneli et al. 2022; Moots et al. 2023; Ravasini et al. 2024). However, it is clear that the genetic variability observed is also shaped over time rather than by geography only. In Central Italy, the gene pool varies from being more similar to Central Europe, in the first centuries of the IA, to a higher diversity shifting toward the Eastern Mediterranean in the Late Republican phase, possibly resulting more similar to IA Southern Italy. Therefore, for this genetically heterogeneous region, thinking in broad chronological terms (e.g. IA and Bronze Age) may be simplistic, not allowing for a thorough study of all its complexity. In this particular case and given the results of this study, Villa Falgari and the other coeval individuals with a strong Eastern Mediterranean genetic component should be considered an integrating part of the Late Republican gene pool rather than “genetic outliers,” since they may represent the first step of the genetic shift typical of the subsequent periods.
Materials and Methods
aDNA processing, including DNA extraction, library preparation, quality control, and sequencing was performed at the aDNA laboratory of the Estonian Biocenter, Institute of Genomics, University of Tartu, Estonia. Radiocarbon dating was performed at the Chrono Centre, Queen's University Belfast, UK.
Archaeological Context
The individual analyzed in this study comes from Villa Falgari, Tarquinia (Italy), and the archaeological context of this finding is really scarce (Barich et al. 1968). In 1960, a well-preserved almost complete skull was found in this locality without grave goods or any other artifacts allowing for an archaeological dating. Originally, it has been placed in an Eneolithic context because of other findings discovered in the same years in the nearby locality of Bandita San Pantaleo (Barich et al. 1968). Nevertheless, radiocarbon dating allowed to precisely date this individual to the Late Roman Republican phase, suggesting that it might be in some way connected to one of the many other Republican and/or Etruscan necropolises in the Tarquinia area. This was one of the major Etruscan cities and remained an important commercial center also during the Roman Republican period, with strong connections with the Greek area (Gianni 2016).
DNA Extraction, Library Preparation, and Sequencing
For the BSP71, the final portion of the molar root was sampled for analysis. All the steps related to DNA extraction, library preparation, and sequencing were performed as detailed in Saupe et al. (2021) and Ravasini et al. (2024). In summary, after decontamination steps with 6% (w/v) bleach and UV light, DNA was extracted from the root portion with EDTA and Proteinase K and incubated at room temperature for 72 h. DNA was then purified with the MinElute PCR Purification Kit (QIAGEN). Sequencing library was built with NEBNext DNA library Prep Master Mix Set for 454 (E6070, New England Biolabs) and with Illumina-specific adaptors (Meyer and Kircher 2010) following established protocols (Meyer and Kircher 2010; Orlando et al. 2013; Malaspinas et al. 2014; Saupe et al. 2021). Quality control procedures were carried out to check that library preparation was successful and quantify the concentration of DNA/sequencing library. In particular, fluorometric quantification (Qubit, Thermo Fisher Scientific), parallel capillary electrophoresis (Fragment Analyzer, Agilent Technologies), and qPCR. The sequencing was performed using the Illumina NextSeq500/550 High-Output single-end 75-cycle kit.
Mapping
Adapters were removed with cutadapt-2.1 (Martin 2011) together with indexes, poly-G tails, and sequences shorter than 28 bp (parameter --minimum-length). The mapping step to the human reference sequence GRCh37 (hs37d5) was performed with BWA-0.7.17 (Li and Durbin 2009) with the command bwa aln and relaxed alignment parameters (-n 0.01 -o 2) in combination with disabling seeding (parameter -l 1024) (Kircher 2012; Schubert et al. 2012; Martiniano et al. 2020; Ravasini et al. 2024). Mapped sequences were converted in BAM format file with samtools-1.9 (Li et al. 2009). picard-2.20.8 (http://broadinstitute.github.io/picard/index.html) was used to remove duplicates (parameter REMOVE_DUPLICATES = true). Indels were realigned using GATK-3.5 (McKenna et al. 2010) with the command IndelRealigner. Reads with mapping quality lower than 25 were removed with samtools-1.9 as suggested in Martiniano et al. (2020). Reported statistics of the final BAM file are in Table S1.
aDNA Authentication, Contamination Rate and Genetic Sex Estimation
MapDamage-2.0 (Jónsson et al. 2013) was used to control for aDNA damage, in particular to estimate the frequency 5′ ends of sequences C > T transitions (Table S1). The contamination rate was estimated with two different methods: (i) on the mtDNA as detailed in Jones et al. (2017) and (ii) on the X chromosome with the method described in Rasmussen et al. (2011) using ANGSD (Korneliussen et al. 2014) with default parameters. In both cases, contamination estimates were <1% (Table S1). The presence of aDNA postmortem damage and the negligible amount of contamination at both mtDNA and nuclear level allowed us to include BSP71 in further analyses. Genetic sex of BSP71 was assigned with the method described in Skoglund et al. (2013), setting the R_y limit for males to 0.074.
mtDNA Haplogroup Assignment
mtDNA haplogroup was assigned with Haplogrep2 (Weissensteiner et al. 2016). As described in Ravasini et al. (2024), a VCF file was obtained by performing a pileup with bcftools-1.14 (Danecek et al. 2021), with the command bcftools mpileup and the flag --ignore-RG. Then, only the variant positions were called with the command bcftools call -m --ploidy 1 -v. This VCF file was used as input for the online version of Haplogrep2 (https://haplogrep.i-med.ac.at/haplogrep2/)
Y Chromosome Haplogroup Assignment and Phylogeny
Phylogenetic relationships among the Y chromosome of BSP71 and other ancient individuals (Antonio et al. 2019; Lazaridis et al. 2022; Moots et al. 2023) were reconstructed with pathPhynder (Martiniano et al. 2022) using default parameters, BAM files as input and the phylogenetic tree in Martiniano et al. (2022) as reference tree. To visualize the resulting tree, we used the R packages treeio and ggtree (Yu et al. 2017). The pathPhynder haplogroup assignment has been refined reporting the defining markers for main haplogroups/subhaplogroups, following the nomenclature proposed by the PhyloTree Y (van Oven et al. 2014) and checking the ISOGG tree v. 15.73.
Variant Calling on Autosomes
For the autosomes, haploid genotypes were called with ANGSD-0.917 (Korneliussen et al. 2014) sampling a random base for each position of the 1,240K SNP panel and considering major and minor alleles as they are indicated in the panel (parameters -doHaploCall 1, -doCounts 1, -doMajorMinor 3 and -sites providing the list of the 1,240K SNPs). The function haploToPlink was used to convert the resulting .haplo file into PLINK (Purcell et al. 2007) format files.
Design of Datasets for Genomic Analysis
BSP71 was compared to modern and ancient samples from the literature present in the AADR dataset v54.1 (Mallick et al. 2024; Mallick and Reich 2024), both the 1,240K and the 1,240K + HO datasets. The dataset with the 1,240K data was used for analyses like D-statistics and qpAdm, while the dataset with 1,240K + HO data was used for PCA, Admixture and f3 statistics. Additionally, the samples reported in Reitsema et al. (2022) and in Koptekin et al. (2023) were included in both the 1,240K and the 1,240K + HO datasets. The data coming from different datasets were converted into binary PLINK file format and merged with Plink-1.9 (Purcell et al. 2007) with the command “—bmerge.” Later, we excluded all the transitions and retained only transversions from the resulting datasets. This filtering step was performed to minimize possible errors or bias due to postmortem damage of aDNA. Therefore, all the autosomal population genetic analyses were performed using only transversion sites. For published individuals, we maintained the group nomenclature assigned in the Allen Ancient DNA Resource dataset (AADR, v54.1) (Dataverse 7.0; November 16 2022, https://doi.org/10.7910/DVN/FFIDCW; Mallick et al. 2024; Mallick and Reich 2024) to allow easier identification.
PCA
PCA was performed with a dataset of modern and ancient Eurasian and North African samples including a total of 1,108 individuals (Table S3). PLINK format files were converted into EIGENSTRAT format with convertf of the EIGENSOFT-7.2.0 package (Patterson et al. 2006; Price et al. 2006) using the parameter familynames:NO. After conversion, smartpca was used to perform PCA with the parameters lsqproject:YES, autoshrink:YES. Ancient individuals were projected onto the PCs built on modern samples. PCs were visualized in R-4.1.3 (https://www.r-project.org/) using the package ggplot2 (https://ggplot2.tidyverse.org/).
ADMIXTURE Analysis
ADMIXTURE analysis was performed including a dataset of modern and ancient West Eurasian and North African individuals from the literature (Table S3). SNPs were pruned for linkage disequilibrium with Plink-1.9 (Purcell et al. 2007) with the parameter --indep-pairwise 50 5 0.5. Diploid genomes (when present) were converted into pseudo-haploid genomes by discarding a random allele for each variant. Ten independent runs (with different random seed) for the k values from 2 to 6 using the option --haploid= ‘*’ were performed. R package pophelper (Francis 2017) was used to merge and visualize the results from the different runs. As in Ravasini et al. (2024), we represented k = 4 because it discriminates between CHG/Iran N and EHG/Steppe Bronze Age components. The ancestral components in the main text (Anatolia_Neolithic, Serbia Mesolithic, EHG/Steppe Bronze Age, CHG/Iran Neolithic) were named based on the populations considered basal to modern Europeans, which have the highest amount of that component.
D-statistics
D-statistics was performed in the form D(X, Y; Iran_N, Onge), where X and Y are ancient Italian IA/Republic or Imperial and post-Imperial groups. X and Y populations were selected also to reduce possible bias related to different sequencing strategies, therefore not including in the same tests shotgun or capture data. D-statistics was performed with the program qpDstat of the package admixtools-7.0.1. (Patterson et al. 2012) with the parameters “printsd: YES” and “inbreed: YES.”
qpAdm Analysis
qpAdm analysis was performed with admixtools-7.0.1. (Patterson et al. 2012) with the option “allsnps: YES” in different forms, as described below.
Target groups were single individuals from Central Italy of the IA/Republican and Imperial period; source populations were ancient Eurasian groups known to have had a major impact in shaping the European gene pool (Haak et al. 2015), observed also in ADMIXTURE analysis: Anatolia_N, Iran_GanjDareh_N, Serbia_IronGates_Mesolithic, and Steppe_BA. In addition, the feasibility of this four-source model in the analyzed groups is amply demonstrated by previous studies (Antonio et al. 2019; Posth et al. 2021; Ravasini et al. 2024). Given the statistical uncertainty in the estimates when performing the model in single individuals, some of them result in negative ancestry proportions. For the scatterplots regarding this model (Fig. 3), the trend of the component proportions was analyzed with a LOESS (LOcally Estimated Scatterplot Smoothing) smooth line with the function geom_smooth of ggplot2 in R with default parameters. Gray areas around the smooth line indicate the confidence interval set to 0.95;
Targets were ancient Italian (IA/Republic or Imperial) groups and sources were different combinations of ancient Italian and Eastern Mediterranean groups. Sources and targets were selected to reduce as much as possible bias related to different sequencing strategies (Davidson et al. 2023).
“Right” (outgroup) populations were selected based on Lazaridis et al. (2022), excluding the ones that were used as sources in the single individuals test, and reported in Tables S6 to S9. For model (2), Italian Bronze Age/Early Bronze Age individuals were also added as “right” population to pull for local Italian Peninsula ancestry.
Outgroup f3 Statistics
f3 statistics were computed with the program qp3pop of the package admixtools-7.0.1 (Patterson et al. 2012) using the option “inbreed:YES”. We used the outgroup f3 statistics with two different approaches:
To explore the similarities between different ancient groups, f3 statistics was performed in the form f3(X, Y; Onge) where X and Y are all the possible combinations of the following populations: Italy_IA_VillaFalgari, Turkey_Aegean_Aydin_BA.SG, Turkey_Central_Gordion_IA.SG, Iran_GanjDareh_N, Turkey_LateC_EBA, Greece_Delphi_IA, Italy_Imperial.SG, Italy_IA_Republic.SG, Italy_IA_Republic_oEasternMediterranean.SG, Italy_TarquiniaMonterozzi_IA_oLevant.SG, Lebanon_EjJaouze_Phoenician.SG, Italy_TarquiniaMonterozzi_IA.SG, Anatolia_N, Serbia_IronGates_Mesolithic.SG, Steppe_BA, EHG, CHG, and Italy_IsolaSacra_RomanImperial.SG. 1 − f3 values were converted into a pairwise distant matrix, and a MDS analysis was performed on this matrix with cmdscale function of R with parameters eig = TRUE and k = 2. Results were visualized with ggplot2. The goodness of fit of the MDS analysis was obtained by creating a new distance matrix from the original MDS and computing its correlation coefficient with the original matrix. The value of the goodness of fit resulted to be low (r2 = 0.15) probably due to the high genetic similarity of several groups from Italy, Greece, and Anatolia.
To investigate the genetic similarity between Villa Falgari and other ancient and modern populations, outgroup f3 statistics was performed in the form f3(Italy_IA_VillaFalgari, Test; Onge) where test populations are (i) relevant modern Eurasians and North Africans or (ii) ancient groups in a time interval of 1,000 years, with the mid-date set as the one of Villa Falgari. Longitude and latitude of the test groups were computed as the mean of the ones of the individuals. Interpolation maps, with the inverse distance weighted method and using the f3 values, were performed with QGIS-3.26.1 (https://www.qgis.org/en/site/), as described in Ravasini et al. (2024). For the representation of the maps, the option “singleband pseudocolor” was selected, the minimum value was set to 0.18 and the maximum to 0.27, and the option “Interpolation: Discrete” was selected.
Supplementary Material
Acknowledgments
Bioinformatic analyses were performed with the facilities of the High Performance Computing Center of the University of Tartu. The authors are grateful to the aDNA lab at the Institute of Genomics of the University of Tartu including Lehti Saag, Helja Kabral, and Anu Solnik for the support in bioinformatic and wet lab analysis. The EASI-Genomics project has received funding from the European Union's Horizon 2020 research and innovation program under grant agreement No 824110.
Contributor Information
Francesco Ravasini, Department of Biology and Biotechnologies ‘Charles Darwin’, Sapienza University of Rome, Rome, Italy; Centre for Palaeogenetics, Stockholm University, Stockholm, Sweden.
Cecilia Conati Barbaro, Department of Sciences of Antiquities, Sapienza University of Rome, Rome, Italy.
Christiana Lyn Scheib, Department of Zoology, St John's College, University of Cambridge, Cambridge, UK.
Kristiina Tambets, Institute of Genomics, University of Tartu, Tartu, Estonia.
Mait Metspalu, Institute of Genomics, University of Tartu, Tartu, Estonia.
Fulvio Cruciani, Department of Biology and Biotechnologies ‘Charles Darwin’, Sapienza University of Rome, Rome, Italy; Department of Biomedical Science, Institute of Molecular Biology and Pathology, CNR, Rome, Italy.
Beniamino Trombetta, Department of Biology and Biotechnologies ‘Charles Darwin’, Sapienza University of Rome, Rome, Italy.
Eugenia D’Atanasio, Department of Biomedical Science, Institute of Molecular Biology and Pathology, CNR, Rome, Italy.
Supplementary Material
Supplementary material is available at Genome Biology and Evolution online.
Funding
This study was supported by EASI-Genomics 3rd Call for Transnational Access grant nr. PID15152 to B.T.; Gerda Henkel Foundation grant nr. 40/V/20 to E.D., B.T., and F.C.; Sapienza University of Rome grants nr. RM12117A81385C5A, RM122181691E0881, and RM123188F697BDAE to B.T. and RM12218167749457 to F.C.; and Italian Ministry of University and Research PRIN2022 2022PC2TSX to E.D. and 2022E8NN2N to B.T., funded by the European Union - Next Generation EU, Mission 4, Component 1, CUP E53D23007590006 and H53D23003260006. F.R. was supported by Sapienza University of Rome grants nr. RM12218167749457 to F.C.
Data Availability
The new sequences generated and analyzed during the current study are available in the European Nucleotide Archive (ENA, https://www.ebi.ac.uk/ena/browser/home) under Accession Number PRJEB81271.
Literature Cited
- Abulafia D. The great sea: a human history of the Mediterranean. Oxford University Press; 2011. [Google Scholar]
- Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009:19:1655–1664. 10.1101/gr.094052.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aneli S, Caldon M, Saupe T, Montinaro F, Pagani L. Through 40,000 years of human presence in Southern Europe: the Italian case study. Hum Genet. 2021:140:1417–1431. 10.1007/s00439-021-02328-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aneli S, et al. The genetic origin of daunians and the pan-Mediterranean Southern Italian Iron Age context. Mol Biol Evol. 2022:39:msac014. 10.1093/molbev/msac014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antonio ML, et al. Ancient Rome: a genetic crossroads of Europe and the Mediterranean. Science. 2019:366:708–714. 10.1126/science.aay6826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antonio ML, et al. Stable population structure in Europe since the Iron Age, despite high mobility. eLife. 2024:13:e79714. 10.7554/eLife.79714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barich B, Bonadonna FP, Borgognini Tarli S, Parenti R. Trovamenti eneolitici presso tarquinia. Orig.-II-Roma; 1968. [Google Scholar]
- Bietti Sestieri AM. L’Italia nell’età del bronzo e del ferro. Dalle palafitte a Romolo (2200-700 a.C.). Carocci; 2018. [Google Scholar]
- Brizzi G. Storia di Roma. Dalle origini ad Azio. Pàtron; 1997. [Google Scholar]
- Broodbank C. The making of the middle sea. Thames & Hudson; 2013. [Google Scholar]
- Coia V, et al. Ancestry and kinship in a Late Antiquity-Early Middle Ages cemetery in the Eastern Italian Alps. iScience. 2023:26:108215. 10.1016/j.isci.2023.108215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Danecek P, et al. Twelve years of SAMtools and BCFtools. GigaScience. 2021:10:giab008. 10.1093/gigascience/giab008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davidson R, et al. Allelic bias when performing in-solution enrichment of ancient human DNA. Mol Ecol Resour. 2023:23:1823–1840. 10.1111/1755-0998.13869. [DOI] [PubMed] [Google Scholar]
- De Angelis F. Archaic and Classical Greek Sicily: a social and economic history. Oxford University Press; 2016. [Google Scholar]
- Fernandes DM, et al. The spread of steppe and Iranian-related ancestry in the islands of the western Mediterranean. Nat Ecol Evol. 2020:4:334–345. 10.1038/s41559-020-1102-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francis RM. Pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour. 2017:17:27–32. 10.1111/1755-0998.12509. [DOI] [PubMed] [Google Scholar]
- Gianni GB. Tarquinii. Oxford Classical Dictionary; 2016. [Google Scholar]
- Haak W, et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature. 2015:522:207–211. 10.1038/nature14317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harney É, Patterson N, Reich D, Wakeley J. Assessing the performance of qpAdm: a statistical tool for studying population admixture. Genetics. 2021:217:iyaa045. 10.1093/genetics/iyaa045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodos T. The archaeology of the Mediterranean Iron Age: a globalising world c.1100–600 BCE. Cambridge University Press; 2020. [Google Scholar]
- Jones ER, et al. The Neolithic transition in the Baltic was not driven by admixture with early European farmers. Curr Biol. 2017:27:576–582. 10.1016/j.cub.2016.12.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jónsson H, Ginolhac A, Schubert M, Johnson PLF, Orlando L. mapDamage2.0: fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinforma Oxf Engl. 2013:29:1682–1684. 10.1093/bioinformatics/btt193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karmin M, et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res. 2015:25:459–466. 10.1101/gr.186684.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kircher M. Analysis of high-throughput ancient DNA sequencing data. In: Shapiro B, Hofreiter M, editors. Ancient DNA: methods and protocols. Methods in molecular biology. Humana Press; 2012. p. 197–228. [DOI] [PubMed] [Google Scholar]
- Koptekin D, et al. Spatial and temporal heterogeneity in human mobility patterns in Holocene Southwest Asia and the East Mediterranean. Curr Biol. 2023:33:41–57.e15. 10.1016/j.cub.2022.11.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korneliussen TS, Albrechtsen A, Nielsen R. ANGSD: analysis of next generation sequencing data. BMC Bioinformatics. 2014:15:356. 10.1186/s12859-014-0356-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazaridis I, et al. A genetic probe into the ancient and medieval history of Southern Europe and West Asia. Science. 2022:377:940–951. 10.1126/science.abq0755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinforma Oxf Engl. 2009:25:1754–1760. 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, et al. The sequence alignment/map format and SAMtools. Bioinforma Oxf Engl. 2009:25:2078–2079. 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malaspinas A-S, et al. Two ancient human genomes reveal Polynesian ancestry among the indigenous Botocudos of Brazil. Curr Biol. 2014:24:R1035–R1037. 10.1016/j.cub.2014.09.078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallick S, Reich D. The Allen Ancient DNA Resource (AADR): a curated compendium of ancient human genomes. Harvard Dataverse, V7.0 data release [Nov 16, 2022]. Dataset. 2024. 10.7910/DVN/FFIDCW. [DOI] [PMC free article] [PubMed]
- Mallick S, et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature. 2016:538:201–206. 10.1038/nature18964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallick S, et al. The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes. Sci Data. 2024:11:182. 10.1038/s41597-024-03031-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcus JH, et al. Genetic history from the Middle Neolithic to present on the Mediterranean island of Sardinia. Nat Commun. 2020:11:939. 10.1038/s41467-020-14523-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011:17:10–12. 10.14806/ej.17.1.200. [DOI] [Google Scholar]
- Martiniano R, De Sanctis B, Hallast P, Durbin R. Placing ancient DNA sequences into reference phylogenies. Mol Biol Evol. 2022:39:msac017. 10.1093/molbev/msac017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martiniano R, Garrison E, Jones ER, Manica A, Durbin R. Removing reference bias and improving indel calling in ancient DNA data analysis by mapping to a sequence variation graph. Genome Biol. 2020:21:250. 10.1186/s13059-020-02160-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenna A, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010:20:1297–1303. 10.1101/gr.107524.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer M, Kircher M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb Protoc. 2010:pdb.prot5448. 10.1101/pdb.prot5448. [DOI] [PubMed] [Google Scholar]
- Moots HM, et al. A genetic history of continuity and mobility in the Iron Age central Mediterranean. Nat Ecol Evol. 2023:7:1515–1524. 10.1038/s41559-023-02143-4. [DOI] [PubMed] [Google Scholar]
- Orlando L, et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature. 2013:499:74–78. 10.1038/nature12323. [DOI] [PubMed] [Google Scholar]
- Pallottino M. A history of earliest Italy. University of Michigan Press; 1991. [Google Scholar]
- Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS Genet. 2006:2:e190. 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patterson N, et al. Ancient admixture in human history. Genetics. 2012:192:1065–1093. 10.1534/genetics.112.145037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piganiol A. La conquête romaine. F. Alcan; 1927. [Google Scholar]
- Pilli E, et al. Ancient DNA challenges prevailing interpretations of the Pompeii plaster casts. Curr Biol. 2024:34:5307–5318.e7. 10.1016/j.cub.2024.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Posth C, et al. The origin and legacy of the Etruscans through a 2000-year archeogenomic time transect. Sci Adv. 2021:7:eabi7673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Posth C, et al. Palaeogenomics of Upper Palaeolithic to Neolithic European hunter-gatherers. Nature. 2023:615:117–126. 10.1038/s41586-023-05726-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poznik GD, et al. Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences. Nat Genet. 2016:48:593–599. 10.1038/ng.3559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price AL, et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006:38:904–909. 10.1038/ng1847. [DOI] [PubMed] [Google Scholar]
- Purcell S, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007:81:559–575. 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rasmussen M, et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science. 2011:334:94–98. 10.1126/science.1211177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravasini F, et al. The genomic portrait of the Picene culture provides new insights into the Italic Iron Age and the legacy of the Roman Empire in Central Italy. Genome Biol. 2024:25:292. 10.1186/s13059-024-03430-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raveane A, et al. Population structure of modern-day Italians reveals patterns of ancient and archaic ancestries in Southern Europe. Sci Adv. 2019:5:eaaw3492. 10.1126/sciadv.aaw3492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitsema LJ, et al. The diverse genetic origins of a Classical period Greek army. Proc Natl Acad Sci U S A. 2022:119:e2205272119. 10.1073/pnas.2205272119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinaldi Tufi S, et al. Province romane. In: Enciclopedia dell’Arte antica. Istituto dell’Enciclopedia Italiana; 1971. [Google Scholar]
- Ringbauer H, et al. Punic people were genetically diverse with almost no Levantine ancestors. Nature. 2025:643:139–147. 10.1038/s41586-025-08913-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saupe T, et al. Ancient genomes reveal structural shifts after the arrival of Steppe-related ancestry in the Italian Peninsula. Curr Biol. 2021:31:2576–2591.e12. 10.1016/j.cub.2021.04.022. [DOI] [PubMed] [Google Scholar]
- Scheidel W, Morris I, Saller RP. The Cambridge economic history of the Graeco–Roman world. Cambridge university press; 2007. [Google Scholar]
- Schubert M, et al. Improving ancient DNA read mapping against modern reference genomes. BMC Genomics. 2012:13:178. 10.1186/1471-2164-13-178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scorrano G, et al. Bioarchaeological and palaeogenomic portrait of two Pompeians that died during the eruption of Vesuvius in 79 AD. Sci Rep. 2022:12:6468. 10.1038/s41598-022-10899-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Serventi P, et al. Iron Age Italic population genetics: the Piceni from Novilara (8th-7th century BC). Ann Hum Biol. 2018:45:34–43. 10.1080/03014460.2017.1414876. [DOI] [PubMed] [Google Scholar]
- Skoglund P, Storå J, Götherström A, Jakobsson M. Accurate sex identification of ancient human remains using DNA shotgun sequencing. J Archaeol Sci. 2013:40:4477–4482. 10.1016/j.jas.2013.07.004. [DOI] [Google Scholar]
- Skourtanioti E, et al. Genomic history of Neolithic to Bronze Age Anatolia, Northern Levant, and Southern Caucasus. Cell. 2020:181:1158–1175.e28. 10.1016/j.cell.2020.04.044. [DOI] [PubMed] [Google Scholar]
- van Dommelen P. Colonialism and migration in the ancient Mediterranean. Annu Rev Anthropol. 2012:41:393–409. 10.1146/annurev-anthro-081309-145758. [DOI] [Google Scholar]
- van Oven M, Van Geystelen A, Kayser M, Decorte R, Larmuseau MH. Seeing the wood for the trees: a minimal reference phylogeny for the human Y chromosome. Hum Mutat. 2014:35:187–191. 10.1002/humu.22468. [DOI] [PubMed] [Google Scholar]
- Weissensteiner H, et al. HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res. 2016:44:W58–W63. 10.1093/nar/gkw233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Smith DK, Zhu H, Guan Y, Lam TT-Y. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol Evol. 2017:8:28–36. 10.1111/2041-210X.12628. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
- Mallick S, Reich D. The Allen Ancient DNA Resource (AADR): a curated compendium of ancient human genomes. Harvard Dataverse, V7.0 data release [Nov 16, 2022]. Dataset. 2024. 10.7910/DVN/FFIDCW. [DOI] [PMC free article] [PubMed]
Supplementary Materials
Data Availability Statement
The new sequences generated and analyzed during the current study are available in the European Nucleotide Archive (ENA, https://www.ebi.ac.uk/ena/browser/home) under Accession Number PRJEB81271.