Abstract
Proteolysis-targeting chimeras (PROTAC) are emerging and promising molecules for targeted protein degradation which have the potential to overcome critical bottlenecks in traditional small molecule drug development. However, the scarcity of publicly available data on molecular compound structures has significantly hindered computational drug discovery and AI-aided drug discovery/design (AIDD) in this field. Patents are an important but underutilized source of novel chemical structures in medicinal chemistry. In this study, we collected PROTAC patents published in 2013–2023 and the associated chemical structures disclosed therein. Through manual screening and expert curation, we identified 63,136 unique PROTAC compounds under 590 patent families, along with 252 targets. Additionally, we employed the ADMETlab 3.0 platform to predict 120 physicochemical properties for all compounds. The dataset is publicly available on the Figshare platform, and an online webserver (http://protacpatentdb.com) has also been established. Given the rapid growth of PROTAC patent literature, this dataset can be further expanded as new patents are continuously published.
Subject terms: Chemical libraries, Chemical libraries, Drug screening
Background & Summary
Proteolysis-targeting chimeras (PROTACs) are emerging small molecules for targeted protein degradation in drug development. PROTACs facilitate the design of bifunctional molecules capable of simultaneously engaging disease-associated proteins of interest (POIs) and E3 ubiquitin ligases. This dual engagement efficiently induces ubiquitination of POIs, facilitating their subsequent recognition and degradation by the endogenous ubiquitin–proteasome system (UPS)1. Since traditional small molecules occupy the active sites of target proteins, the event-driven mechanism endows PROTACs with three key advantages: overcoming the limitations of traditionally “undruggable” targets, circumventing target protein resistance, and potentially achieving therapeutic effects at lower doses2.
Continuous advances in PROTAC drug design, including the use of computer-aided drug discovery/design (CADD) and artificial intelligence-aided drug discovery/design (AIDD) methods, have enhanced and accelerated the discovery and development of PROTAC molecules3–5. However, a notable challenge in PROTAC drug discovery is the scarcity of high-quality molecular data, which are essential to provide a robust foundation for artificial intelligence (AI)-assisted drug design. Generally, within the constraints of model architecture and computational resources, increasing the amount of high-quality data can enhance the predictive accuracy, robustness, and reliability of AI models6.
To address this gap, Weng et al.7 constructed the PROTAC-DB database (http://cadd.zju.edu.cn/protacdb/) by manually collecting molecular structure, target protein, and activity information for PROTACs from PubMed-indexed publications. This database has undergone three iterations to date, with PROTAC-DB 3.08 being the latest version. PROTAC-DB 3.0 is the largest and most comprehensive database of PROTAC compounds available, comprising data on 6,111 PROTAC molecules. Further resources include the PROTACpedia database (https://protacpedia.weizmann.ac.il/ptcb/main), which includes 1,190 PROTAC compounds identified in the published literature, and the PROTAC-Databank (https://bailab.siais.shanghaitech.edu.cn/services/deepprotac-db), which contains data on 3,645 PROTAC molecules identified in scientific publications. However, PROTAC-DB, PROTACpedia, and PROTAC-Databank cover a limited chemical space, with substantial data overlap between PROTAC-DB and PROTACpedia4. Multiple reviews have pointed out that the scarcity of available high-quality PROTAC compound data remains a major challenge3–5,9,10, highlighting the need for such data.
The patent literature represents an important but underutilized source of chemical structure information. Compared to journal articles, medicinal chemistry patents generally disclose novel compounds and their structural details earlier and more comprehensively11. Importantly, many innovative molecules are exclusively disclosed in patents and are never reported in scientific publications12. Therefore, we decided to extract PROTAC compound structural data from medicinal chemistry patents.
In this study, we retrieved PROTAC patents from Derwent Innovation patent database and, through rigorous manual screening, refined our dataset to 590 distinct patent families. Subsequently, by leveraging the SciFinder database, we extracted and annotated 63,136 unique PROTAC compound structures. Furthermore, we employed the ADMETlab 3.0 platform13 to predict absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties for these compounds, thereby constructing a comprehensive and high-quality structural dataset for PROTAC research.
Table 1 shows a comparative overview of the scale of the dataset developed in this study compared to existing PROTAC compound databases. To our knowledge, this study presents the first patent-based PROTAC compound dataset, and it comprises the most extensive collection of PROTAC chemical structures. Overall, the dataset substantially expands the chemical range of existing PROTAC databases and effectively addresses the inherent gaps in the literature-derived PROTAC resources. The integration of these complementary datasets established a comprehensive foundation to support future PROTAC research and drug discovery endeavors.
Table 1.
Comparative overview of current PROTAC compound databases.
| Data Category | PROTAC-DB 3.0 | PROTAC-Databank | PROTACpedia | PROTAC-PatentDB |
|---|---|---|---|---|
| Number of PROTACs | 6,111 | 3,645 | 1,190 | 63,136 |
| Number of target proteins | 442 | 337 | 82 | 252 |
| Data source | Articles | Articles | Articles | Patents |
To demonstrate the broader coverage of chemical space, we performed a UMAP-based dimensionality reduction analysis comparing compounds from PROTAC-PatentDB with those in two representative public databases, PROTAC-DB and PROTACpedia (Figure S1). The visualization clearly shows that compounds in PROTAC-PatentDB distribute across a wider and more diverse chemical space compared with the other two databases. This indicates that our database not only expands the accessible chemical diversity but also provides a richer resource for machine learning applications in the field of targeted protein degradation.
We publicly deposited the complete PROTAC compound dataset described in this study on the Figshare platform14. Additionally, to further enhance accessibility and facilitate dataset usage by researchers, we established an online webserver (http://protacpatentdb.com) that allows users to download and explore the data conveniently.
Methods
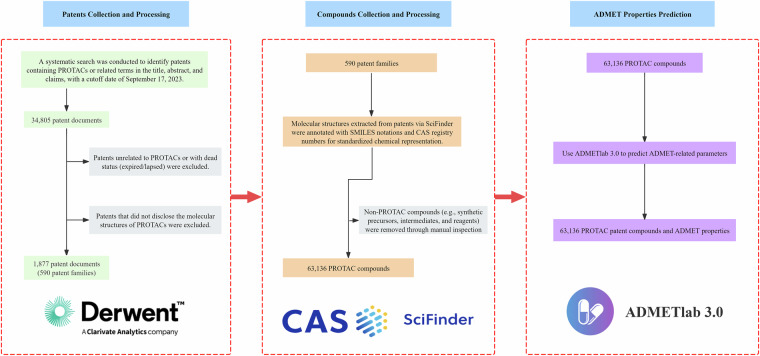
We employed a structured workflow (Fig. 1) to construct a comprehensive PROTAC compound dataset. Initially, we retrieved patents from the Derwent Innovation patent database (https://derwentinnovation.clarivate.com) using a detailed keyword search based on PROTAC-related terms which yielded 34,805 documents, the full list of specific search terms and query syntax used in this study are provided in the supplementary material. We applied stringent filtering to these records to exclude inactive patents (“dead” status), irrelevant entries, and patents lacking explicit PROTAC structure disclosure, which ultimately resulted in 590 distinct patent families. For each patent family, we extracted the chemical structures using the SciFinder database (https://scifinder.cas.org) and then performed expert-curated annotation to remove intermediates, catalysts, and structural redundancies. Consequently, we compiled 63,136 unique PROTAC chemical structures. Finally, we comprehensively predicted the ADMET properties of these compounds using ADMETlab 3.0.
Fig. 1.
Flowchart for data collection and processing.
Patent collection and processing
We obtained patents from the Derwent Innovation patent database (https://derwentinnovation.clarivate.com) because it provides global patent information. To ensure a comprehensive search, we searched for terms related to PROTAC in the title, abstract, claim, and Derwent World Patent Index (DWPI) fields. The search conducted on September 17, 2023, returned 34,805 patent documents.
We refined the dataset through several filtering steps. First, we identified patents with “dead” legal statuses and excluded them. A “dead” status indicates that a patent application is no longer active, encompassing situations such as abandonment, lapsing, expiration, revocation, or rejection. Subsequently, we manually screened the titles, abstracts, and claims to eliminate patents that were irrelevant to PROTACs. We then conducted further filtering by inspecting the patent specifications to verify the explicit disclosure of PROTAC molecular structures. Additionally, we consolidated patent families—groups of related patents derived from the same initial priority application and subsequently filed across multiple countries or regions. This study adopts the Derwent World Patents Index (DWPI) patent family classification standard. Based on common priority and consistency of technical content, the same invention patents worldwide are classified into a unified family to improve the accuracy and consistency of patent analysis. Ultimately, this yielded a refined dataset of 1,877 patent documents relating to 590 distinct patent families. Through a manual examination of abstracts and specifications, we further identified and confirmed the specific molecular targets addressed by patented compounds.
Figure 2 provides a comprehensive overview of PROTAC-related patent activities, including application trends, geographic distributions, leading patent assignees, and key molecular targets. Since 2015, the number of patent filings in the PROTAC field has increased rapidly, particularly between 2019 and 2022, reflecting significant global interest and intensive research and development (R&D) investment. However, we observed a slight decrease in 2023 attributable to the data cutoff date (September 2023); hence, overall, the patent activity remained robust. Geographically, the United States (US, 23.02%) and China (CN, 20.94%) were the most active jurisdictions for PROTAC patent disclosures, indicating that these two countries enjoy competitive advantages in PROTAC research. Additionally, notable contributions from the European Patent Office (EPO, 16.78%) and the World Intellectual Property Organization (WIPO, 15.62%) highlighted the global nature of patent filings and strategic efforts to establish worldwide intellectual property protection in the PROTAC field. Regarding leading patent assignees, institutions such as the Dana-Farber Cancer Institute, Kymera Therapeutics, Yale University, and the University of Michigan ranked highly, emphasizing that innovation in the PROTAC field is largely driven by top-tier academic institutions and specialized biotechnology companies. In terms of targeted proteins, androgen receptor (AR), Bruton’s tyrosine kinase (BTK), bromodomain-containing protein 4 (BRD4), estrogen receptor (ER), and epidermal growth factor receptor (EGFR) emerged as the principal targets within the PROTAC patent literature, underscoring an R&D focus primarily oriented toward oncology-related targets. In particular, AR and BTK exhibited significantly higher patent activity than other targets, highlighting their pivotal role and commercial potential in the development of PROTAC-based therapeutics.
Fig. 2.
Overview of PROTAC patents. (a) annual trends in PROTAC patent applications, represented by patent documents and patent families, (b) geographic distribution of PROTAC patent documents by publication authority, (c) top 15 patent holders ranked by number of patent families, and (d) top 15 molecular targets ranked by number of patent families.
Compound collection and processing
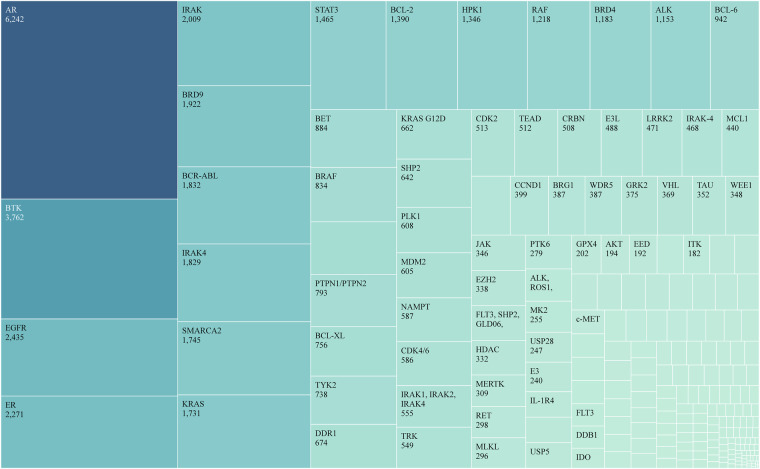
For the 590 patent families mentioned previously, we retrieved specific patent numbers through the “references” module of the SciFinder database (https://scifinder.cas.org). Subsequently, we accessed the “substances” category to obtain the corresponding compound lists. The exported data included structural identifiers, such as CAS registry numbers, canonical SMILES, and InChI keys. After merging and consolidating compound tables from all 590 patent families, we manually curated and validated the PROTAC compound patent families to ensure the exclusion of reaction intermediates, catalytic ligands, and other nonterminal product noise molecules. Considering potential compound structure redundancies arising from citation relationships among patent families, we retained only the patent with the earliest priority year for each chemical structure, thus ensuring the uniqueness and originality of each compound entry in the dataset. Finally, we obtained 63,136 unique PROTAC chemical structures. Figure 3 shows the distribution of molecular targets for compounds in PROTAC-PatentDB. By examining patent specifications, we identified 252 specific targets addressed by the 63,136 PROTAC compounds, among which the top five targets with the highest number of compounds were AR, BTK, EGFR, ER, and IRAK.
Fig. 3.
Distribution of specific molecular targets among 63,136 PROTAC compounds.
ADMET property prediction
We employed ADMETlab 3.013 (https://admetlab3.scbdd.com/) to predict the ADMET properties of the 63,136 unique PROTAC compounds identified. ADMETlab 3.0 is an online platform designed for the evaluation and prediction of key ADMET properties, and it provides predictions across 120 individual parameters. However, it is worth noting that ADMETlab 3.0 was originally designed and validated for small organic compounds. Its application to PROTACs should be interpreted with caution due to the challenges of property prediction for PROTAC compounds caused by their relatively large molecular weight, flexible linkers, bifunctional structures and “beyond the Rule of 5” characteristics15.
Table 2 shows a comparative analysis of Lipinski’s rule of five16 (Ro5) properties across various PROTAC databases. Notably, PROTAC-PatentDB, which we constructed in this study, includes significantly more compounds than other databases, allowing for a comprehensive overview of the PROTAC chemical space. On average, the molecular weights (MW, 920.7 Da), cLogPs (3.8), hydrogen bond acceptors (HBAs, 16.0), hydrogen bond donors (HBDs, 3.6), and topological polar surface areas (TPSAs, 192.5 Ų) given in PROTAC-PatentDB were consistent with the known characteristics of PROTAC molecules, typically exceeding traditional Ro5 criteria. Compared to ARVINAS-DB17, PROTAC-DB 1.018, and PROTACpedia (https://protacpedia.weizmann.ac.il/), our dataset displays intermediate and representative physicochemical parameters, reflecting balanced chemical diversity and providing a valuable complementary resource for PROTAC-based drug design and further research efforts.
Table 2.
Comparison of Ro5 properties across PROTAC databases.
| Property | PROTAC-PatentDB | ARVINAS-DB | PROTAC-DB 1.0 | PROTACpedia | |
|---|---|---|---|---|---|
| Count | 63,136 | 1,573 | 9,380 | 1,202 | |
| MW | Mean | 920.7 | <950 | 939.7 | 1,000.1 |
| Max | 3,487.5 | / | 2,750.3 | 1,963.2 | |
| Min | 444.3 | / | 409.4 | 382.3 | |
| cLogP | Mean | 3.8 | 1–7 | 5.0 | 5.7 |
| Max | 15.0 | / | 19.2 | 14.4 | |
| Min | −5.0 | / | −18.0 | −3.0 | |
| HBA | Mean | 16.0 | <15 | 17.7 | 14.3 |
| Max | RDKIT | / | / | / | |
| Min | RDKIT | / | / | / | |
| HBD | Mean | 3.6 | <2 | 4.5 | 4.4 |
| Max | RDKIT | / | / | / | |
| Min | RDKIT | / | / | / | |
| TPSA | Mean | 192.5 | <200 | 240.0 | 227.2 |
| Max | 1,072.9 | / | / | 460.0 | |
| Min | 34.3 | / | / | 98.9 | |
Data Record
We have uploaded14 the complete datasets and accompanying supporting files to Figshare19 and provided the complete compound dataset in the file “PROTAC_Patent_Compounds.xlsx.” The file includes the structures of 63,136 unique compounds (covering 252 distinct targets) along with their basic attributes (e.g., SMILES, targets, CAS registry numbers, InChI keys, patent numbers, patent publication years, patent family numbers, and patent assignees), as well as 120 physicochemical properties predicted using ADMETlab 3.0. The file “PROTAC_ADMET_Properties_Overview.xlsx” provides detailed definitions of these 120 physicochemical properties. The complete patent dataset is available in “PROTAC_Patent_Information.xlsx,” which provides comprehensive patent information (e.g., publication number, assignee, inventor, publication date, application date, title, abstract, and claim), target classifications, patent family details, legal statuses, and citation information. Thus, it enables researchers to investigate the background, application strategy, and technological scope of each PROTAC-related patent.
It is worth noting that the current dataset is in its preliminary stage, primarily serving to validate the feasibility of data collection and curation, and therefore does not include compound activity data yet. In future work, scholars may expand the dataset by continuously tracking newly disclosed patent molecules, incorporating corresponding bioactivity data, and performing a more granular analysis of structural components within these molecules to ensure data completeness and representativeness. Furthermore, it is worth to develop and refine ADMET prediction specifically tailored for PROTAC molecules, leveraging advanced computational approaches such as machine learning and molecular modeling to enhance predictive accuracy and practical applicability. These efforts will not only enrich the PROTAC-related data resources but also provide a robust foundation for subsequent molecular design and drug discovery endeavors.
Technical Validation
To ensure the reliability and technical quality of the dataset, we constructed the PROTAC compound dataset in this study using a series of rigorous technical validation procedures. First, during the patent data collection and cleaning processes, we strictly adhered to the guidelines outlined in the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA20) and the Reporting Items for Patent Landscapes (RIPL) statement21, thus ensuring a transparent research approach. Specifically, we individually and manually reviewed the title, abstract, claims, and specifications of each patent to ensure that all included patents accurately pertained to PROTACs.
Second, all PROTAC compounds included in this dataset were manually validated and verified. Specifically, medicinal chemistry experts independently conducted three rounds of review and annotation for each chemical structure to ensure that all entries explicitly fell within the PROTAC structural category. This manual review effectively eliminated potential errors and noise introduced by the SciFinder database’s automated recognition process, including structural fragments, synthetic intermediates, catalysts, and unrelated impurities. These noisy compounds are frequently misidentified as final products by automated extraction tools, which is a common issue when automatically extracting chemical structures from the patent literature22. Therefore, integrating multiple rounds of expert reviews and annotations significantly improved the dataset’s accuracy and reliability, ensuring its scientific validity.
Finally, we conducted further comparative analyses based on the widely recognized Ro5 drug-likeness parameters to validate the chemical rationality of our dataset. The results showed that the Ro5 properties of the compounds in our dataset were within reasonable ranges compared to other publicly available PROTAC databases, indicating their structural diversity and suitability for drug design. Additionally, we predicted the ADMET properties of all 63,136 compounds using the ADMETlab 3.0 platform, which is well-known for its high accuracy, efficiency, and reliability.
Case studies
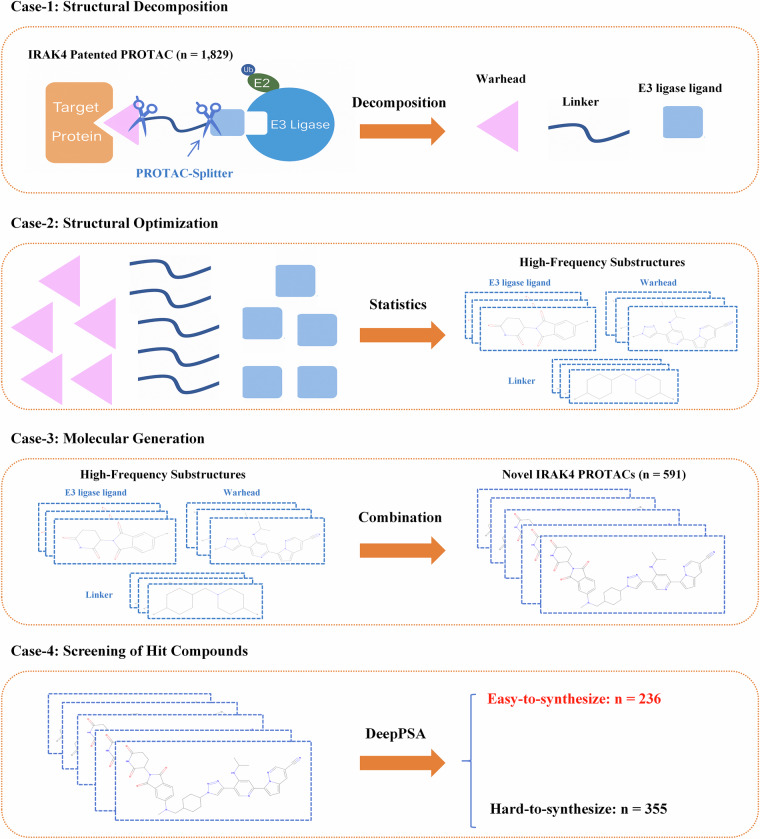
Taking IRAK4-targeting PROTAC molecules as an example, we present four case studies to demonstrate the utility and potential of the PROTAC-PatentDB dataset in facilitating the key stages of PROTAC drug discovery (Fig. 4). Specifically, these application scenarios include: structural deconstruction (Case 1), structural optimization (Case 2), molecular generation (Case 3), and screening of hit compounds (Case 4).
Fig. 4.
Workflow of case studies.
Case 1: Structural Decomposition
PROTAC molecules typically comprise three core structural components: a ligand targeting the protein of interest (warhead), a linker, and a ligand binding to the E3 ubiquitin ligase (E3 ligand). The structural characteristics and precise combinations of these three components directly determine the biological activity, selectivity, and drug-likeness of PROTAC molecules23–25. Therefore, accurately decomposing PROTAC molecules into these three distinct substructures is essential for facilitating subsequent PROTAC drug discovery efforts, such as novel PROTAC molecule design and synthetic route development, highlighting the significant research implications of this process.
Traditionally, structural decomposition of PROTACs relies predominantly on manual identification and annotation. Although manual annotation generally provides high accuracy, it is time-consuming and inefficient for processing large-scale datasets. Recently, rapid accumulation of PROTAC research data has created an urgent demand for automated and efficient tools capable of facilitating structural decomposition. In response to this challenge, researchers developed PROTAC-Splitter (https://huggingface.co/spaces/ailab-bio/PROTAC-Splitter-App), a machine learning-based automatic structure decomposition tool. PROTAC-Splitter integrates a hybrid modeling approach combining advanced graph-based XGBoost and sequence-based Transformer models. This combination allows for rapid and efficient structural decomposition of large-scale PROTAC datasets, accurately recognizing and annotating complex molecular structures through models trained on more than one million simulated PROTAC data points.
In this case study, we utilized the PROTAC-Splitter to automate structural decomposition of 1,829 IRAK4-targeted PROTAC molecules derived from PROTAC-PatentDB. Specifically, we first standardized the SMILES structural representations for each PROTAC molecule, then uploaded the entire CSV file to the PROTAC-Splitter web server (https://huggingface.co/spaces/ailab-bio/PROTAC-Splitter-App). Subsequently, the model automatically identified and delineated the boundaries of warheads, linkers, and E3 ligands within each molecule, explicitly labeling the connection sites using dummy atoms. This automated process significantly enhanced data processing efficiency, enabling rapid structural decomposition of large-scale datasets within a short timeframe.
Despite the demonstrated efficiency and accuracy of the PROTAC-Splitter tool for large-scale data processing, certain inherent limitations of automated structural decomposition must be acknowledged. Machine-learning models may encounter difficulties in accurately recognizing substructures or defining precise boundaries between warheads and linkers. Therefore, we strongly recommend that researchers incorporate manual review and validation in practical applications to ensure the accuracy and reliability of structural annotations.
Case 2: Structural Optimization
Systematic statistical analysis of substructure fragments derived from reported PROTAC molecules can effectively reveal structural preferences and trends present in current research and patent protection. Such structural feature analysis is valuable for identifying common chemical patterns and structural characteristics shared among PROTAC compounds targeting specific proteins, thus providing clear reference information for researchers in future PROTAC molecule design and optimization efforts.
In this study, guided by the aforementioned objectives, we conducted detailed statistical analyses of the three substructure fragments (warhead, linker, and E3 ligand) derived from the structural decomposition of 1,829 IRAK4-targeted PROTAC molecules obtained from patent literature. The purpose was to clearly elucidate the distribution patterns and frequencies of these substructures within existing patents. Specifically, all substructure fragments obtained from the decomposition step were initially subjected to structural standardization to ensure consistency and comparability of their SMILES representations. Subsequently, RDKit was employed to extract structural scaffolds and calculate chemical descriptors for each fragment, facilitating a clear statistical evaluation of their frequency distributions. Finally, we selected the top 10 most frequently occurring representative fragments for each structural category (warhead, linker, and E3 ligand) and performed a more comprehensive analysis of their physicochemical properties and structural characteristics to achieve a thorough understanding of the chemical attributes of these high-frequency substructures.
In this study, guided by the objectives mentioned above, we conducted detailed statistical analyses on the three structural fragments—warheads, linkers, and E3 ligands—derived from the decomposition of 1,829 IRAK4-targeted PROTAC molecules obtained from patent literature. The purpose of these analyses was to elucidate the distribution patterns and frequencies of these substructures within existing patents. Specifically, all extracted substructure fragments were first standardized to ensure consistency and comparability of their SMILES representations. Subsequently, we employed RDKit to extract structural scaffolds and calculate chemical descriptors for each fragment, clearly defining and quantifying the frequency distributions of these structural fragments. Ultimately, we selected the top 10 most frequently occurring representative fragments for each structural category (warhead, linker, and E3 ligand) (Table S1), and conducted more detailed physicochemical property and structural characteristic analyses to comprehensively understand the chemical attributes of these high-frequency structures.
In the statistical analysis of warhead structures, we observed that the top-ranking fragments generally contained distinct aromatic or heterocyclic scaffolds, such as pyrazole, pyrimidine, and benzimidazole. The analysis of linker structures primarily focused on their length, flexibility, and structural types. Among the 1,829 IRAK4-targeted PROTACs analyzed, the most frequent linker fragments were predominantly flexible, with chain lengths typically ranging from approximately 5 to 12 atoms. For the structural analysis of E3 ligands, we found a clear predominance of CRBN (Cereblon)-related ligands, likely due to their clinical validation, well-defined structural features, and relative maturity in chemical synthesis.
Case 3: Molecular Generation
Systematically and efficiently combining existing structural fragments to generate novel PROTAC candidates remains a significant challenge. Data-driven combinatorial construction can rapidly expand the chemical space accessible to PROTAC molecules, producing extensive candidate libraries with potential for innovation and optimization26. Such combinatorial approaches fully exploit the design potential inherent in existing structural fragments, reduce the complexity and cost associated with discovering new compounds, and quickly generate diverse molecular design options.
In this study, based on the previously described statistical analyses of substructures, we selected the ten most frequently occurring warhead, linker, and E3 ligand fragments identified from patented IRAK4-targeted PROTACs to generate new candidate compounds. Specifically, by exhaustively combining these top-ranked structural fragments—comprising ten E3 ligands, ten warheads, and ten linkers—we generated 1,000 candidate PROTAC molecules. After comparing these newly generated molecules with those already documented in our PROTAC-PatentDB dataset, we identified and removed 409 duplicates, ultimately yielding 591 novel IRAK4-targeted PROTAC candidates. This combinatorial case study underscores the practical value and potential of the PROTAC-PatentDB dataset in supporting and accelerating innovation in PROTAC drug discovery.
Case 4: Screening of Hit Compounds
After generating novel PROTAC candidate molecules, the next critical step in drug discovery involves selecting high-quality “hit compounds” that warrant experimental validation and further optimization. Typically, initial molecular libraries derived from computational approaches contain a large proportion of compounds with varying degrees of synthetic complexity and feasibility. Therefore, before initiating costly and resource-intensive laboratory synthesis and biological testing, it is essential to perform preliminary screening to rapidly identify compounds with favorable synthetic accessibility and practical development potential. Synthetic accessibility evaluation serves as an effective computational screening method to prioritize promising candidate molecules, significantly streamlining the selection of viable hit compounds27.
In this study, we employed DeepPSA28 (https://bailab.siais.shanghaitech.edu.cn/psa), a deep learning-based SA prediction tool to evaluate the synthetic accessibility of PROTAC molecules. DeepPSA utilizes deep learning algorithms trained on extensive synthetic reaction datasets comprising millions of data points, providing rapid and robust predictions of synthetic feasibility for novel molecular structures. Specifically, we submitted the SMILES structural information of the 591 newly designed IRAK4-targeted PROTAC candidate molecules, derived from Case 3, to the DeepPSA online server. DeepPSA analyzes each molecular structure and returns a synthetic accessibility score between 0 and 1. Scores closer to 1 indicate a lower predicted synthetic difficulty, suggesting higher synthetic feasibility under laboratory conditions, whereas scores approaching 0 suggest greater synthetic complexity. To clarify the practical significance of these results, we established 0.5 as the threshold for synthetic feasibility; molecules scoring ≥ 0.5 were classified as “predicted synthetically accessible.”
Following the DeepPSA evaluation, among the 591 newly generated PROTAC candidate molecules submitted, 236 achieved scores exceeding the 0.5 threshold, indicating that these molecules are more likely to be practically synthesized in the laboratory. These predicted synthetically accessible molecules provide reliable and experimentally feasible candidates for subsequent medicinal chemistry studies and drug development efforts.
Usage Notes
Users can obtain the PROTAC-PatentDB dataset through two primary channels: (1) The dataset has been publicly deposited on the Figshare platform16, providing standardized download formats for all compounds; (2) An interactive webserver (http://protacpatentdb.com) has been established to enable customized data exploration and bulk downloads.
The PROTAC compound dataset developed in this study can be applied to underpin computational drug discovery and AI-driven modeling. Chemical structures within the dataset are presented in standard cheminformatics formats, including SMILES representations, InChI keys, CAS registry numbers, and predicted ADMET properties. Researchers can readily access and process this dataset using common cheminformatics tools. A distinct advantage of this dataset is that all included compounds originate exclusively from the patent literature, thus providing extensive chemical diversity and facilitating exploration of the novel chemical space disclosed in patents. In contrast, existing publicly available databases (e.g., PROTAC-DB, PROTACpedia, and PROTAC-Databank) comprise predominantly compound information drawn from high-impact academic publications, offering only experimentally validated biological and pharmacological data. Therefore, we recommend that researchers integrate this patent-derived dataset with literature-based PROTAC databases to leverage the complementary strengths of both data sources, enhancing the breadth and depth of subsequent research.
Supplementary information
Acknowledgements
We gratefully acknowledge the funding support from the Science and Technology Development Fund of Macau SAR and the University of Macau for this work. This research was funded by the Science and Technology Development Fund of Macau SAR (No.: 005/2023/SKL, SKL-QRCM(UM)-2023-2025, and 0049/2024/AGJ) and by the University of Macau (No.: MYRG-CRG2023-00007-ICMS-IAS, and MYRG-GRG2024-00268-ICMS-UMDF).
Author contributions
Data collection: Hong Cai and Tianyi Zhang. Data processing and analysis: Hong Cai, Gengyuan Yao, Yulong Shi and Tianyi Zhang. Technical validation support: Yulong Shi, Yuanjia Hu and Hong Cai. Writing and reviewing the manuscript: Hong Cai, Gengyuan Yao, Yulong Shi and Yuanjia Hu. Research supervision and manuscript revision: Yuanjia Hu. Funding support: Yuanjia Hu.
Data availability
The dataset has been deposited in the repository Figshare and is available for unrestricted access via the following link: 10.6084/m9.figshare.2935132114. It can also be accessed through an interactive webserver: PROTAC-PatentDB (http://protacpatentdb.com).
Code availability
No custom code was used to generate or process the data described in the manuscript.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Hong Cai, Gengyuan Yao, Yulong Shi.
Supplementary information
The online version contains supplementary material available at 10.1038/s41597-025-06136-9.
References
- 1.Li, X. & Song, Y. C. Proteolysis-targeting chimera (PROTAC) for targeted protein degradation and cancer therapy. J Hematol Oncol13, 50, 10.1186/s13045-020-00885-3 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Békés, M., Langley, D. R. & Crews, C. M. PROTAC targeted protein degraders: the past is prologue. Nat Rev Drug Discov21, 181–200, 10.1038/s41573-021-00371-6 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ge, J., Hsieh, C. Y., Fang, M., Sun, H. Y. & Hou, T. Development of PROTACs using computational approaches. Trends Pharmacol Sci45, 1162–1174, 10.1016/j.tips.2024.10.006 (2024). [DOI] [PubMed] [Google Scholar]
- 4.Gharbi, Y. & Mercado, R. A comprehensive review of emerging approaches in machine learning for de novo PROTAC design. Digital Discovery3, 2158–2176, 10.1039/D4DD00177J (2024). [Google Scholar]
- 5.Tan, S. Y., Chen, Z. L., Lu, R. Q., Liu, H. X. & Yao, X. J. Rational Proteolysis Targeting Chimera Design Driven by Molecular Modeling and Machine Learning. WIREs: Computational Molecular Science15, 10.1002/wcms.70013 (2025).
- 6.Schneider, P. et al. Rethinking drug design in the artificial intelligence era. Nat Rev Drug Discov19, 353–364, 10.1038/s41573-019-0050-3 (2020). [DOI] [PubMed] [Google Scholar]
- 7.Weng, G. Q. et al. PROTAC-DB: an online database of PROTACs. Nucleic Acids Res49, D1381–d1387, 10.1093/nar/gkaa807 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ge, J. X. et al. PROTAC-DB 3.0: an updated database of PROTACs with extended pharmacokinetic parameters. Nucleic Acids Res53, D1510–d1515, 10.1093/nar/gkae768 (2025). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lv, W. X. et al. In silico modeling of targeted protein degradation. Eur J Med Chem289, 117432, 10.1016/j.ejmech.2025.117432 (2025). [DOI] [PubMed] [Google Scholar]
- 10.Zattoni, J. et al. A comprehensive primer and review of PROTACs and their In Silico design. Comput Methods Programs Biomed264, 108687, 10.1016/j.cmpb.2025.108687 (2025). [DOI] [PubMed] [Google Scholar]
- 11.Southan, C., Varkonyi, P., Boppana, K., Jagarlapudi, S. & Muresan, S. Tracking 20 years of compound-to-target output from literature and patents. PLoS One8, e77142, 10.1371/journal.pone.0077142 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bregonje, M. Patents: A unique source for scientific technical information in chemistry related industry? World Patent Information27, 309–315 (2005). [Google Scholar]
- 13.Fu, L. et al. ADMETlab 3.0: an updated comprehensive online ADMET prediction platform enhanced with broader coverage, improved performance, API functionality and decision support. Nucleic Acids Res52, W422–w431, 10.1093/nar/gkae236 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cai, H., Yao, G., Shi, Y., Zhang, T. & Hu, Y. PROTAC-PatentDB: A PROTAC patent compound dataset. Figshare Dataset10.6084/m9.figshare.29351321 (2025). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cantrill, C. et al. Fundamental aspects of DMPK optimization of targeted protein degraders. Drug Discov Today25, 969–982, 10.1016/j.drudis.2020.03.012 (2020). [DOI] [PubMed] [Google Scholar]
- 16.Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev46, 3–26, 10.1016/s0169-409x(00)00129-0 (2001). [DOI] [PubMed] [Google Scholar]
- 17.Hornberger, K. R. & Araujo, E. M. V. Physicochemical Property Determinants of Oral Absorption for PROTAC Protein Degraders. J Med Chem66, 8281–8287, 10.1021/acs.jmedchem.3c00740 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jimenez, D. G., Sebastiano, M. R., Caron, G. & Ermondi, G. Are we ready to design oral PROTACs®? Admet dmpk9, 243–254, 10.5599/admet.1037 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Thelwall, M. & Kousha, K. Figshare: a universal repository for academic resource sharing? Online Information Review40, 333–346, 10.1108/OIR-06-2015-0190 (2016). [Google Scholar]
- 20.Knobloch, K., Yoon, U. & Vogt, P. M. Preferred reporting items for systematic reviews and meta-analyses (PRISMA) statement and publication bias. J Craniomaxillofac Surg39, 91–92, 10.1016/j.jcms.2010.11.001 (2011). [DOI] [PubMed] [Google Scholar]
- 21.Smith, J. A. et al. The Reporting Items for Patent Landscapes statement. Nat Biotechnol36, 1043–1047, 10.1038/nbt.4291 (2018). [DOI] [PubMed] [Google Scholar]
- 22.Ohms, J. Validity of PubChem compounds supplied by Patentscope or SureChEMBL. World Patent Information70, 102134, 10.1016/j.wpi.2022.102134 (2022). [Google Scholar]
- 23.Paiva, S. L. & Crews, C. M. Targeted protein degradation: elements of PROTAC design. Curr Opin Chem Biol50, 111–119, 10.1016/j.cbpa.2019.02.022 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bondeson, D. P. et al. Lessons in PROTAC Design from Selective Degradation with a Promiscuous Warhead. Cell Chem Biol25, 78–87.e75, 10.1016/j.chembiol.2017.09.010 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kumar, H. & Sobhia, M. E. Interplay of PROTAC Complex Dynamics for Undruggable Targets: Insights into Ternary Complex Behavior and Linker Design. ACS Med Chem Lett15, 1306–1318, 10.1021/acsmedchemlett.4c00189 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Osman, J., Thompson, P. E., Jörg, M. & Scanlon, M. J. Methods to accelerate PROTAC drug discovery. Biochem J482, 921–937, 10.1042/bcj20243018 (2025). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Skoraczyński, G., Kitlas, M., Miasojedow, B. & Gambin, A. Critical assessment of synthetic accessibility scores in computer-assisted synthesis planning. J Cheminform15, 6, 10.1186/s13321-023-00678-z (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang, R. et al. DeepPSA: A Geometric Deep Learning Model for PROTAC Synthetic Accessibility Prediction. J Chem Inf Model65, 6861–6873, 10.1021/acs.jcim.5c00366 (2025). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The dataset has been deposited in the repository Figshare and is available for unrestricted access via the following link: 10.6084/m9.figshare.2935132114. It can also be accessed through an interactive webserver: PROTAC-PatentDB (http://protacpatentdb.com).
No custom code was used to generate or process the data described in the manuscript.