Abstract
During HIV-1 entry, the highly conserved gp41 N-trimer pocket region becomes transiently exposed and vulnerable to inhibition. Using mirror-image phage display and structure-assisted design, we have discovered protease-resistant D-amino acid peptides (D-peptides) that bind the N-trimer pocket with high affinity and potently inhibit viral entry. We also report high-resolution crystal structures of two of these D-peptides in complex with a pocket mimic that suggest sources of their high potency. A trimeric version of one of these peptides is the most potent pocket-specific entry inhibitor yet reported by three orders of magnitude (IC50 = 250 pM). These results are the first demonstration that D-peptides can form specific and high-affinity interactions with natural protein targets and strengthen their promise as therapeutic agents. The D-peptides described here address limitations associated with current L-peptide entry inhibitors and are promising leads for the prevention and treatment of HIV/AIDS.
Keywords: microbicide, phage display, protein design
HIV entry is mediated by the viral envelope glycoprotein, which comprises noncovalently associated surface (gp120) and transmembrane (gp41) subunits. gp120 is primarily involved in recognition of cellular receptors, whereas gp41 directly mediates membrane fusion. When peptides isolated from the gp41 N- and C-peptide regions (N- and C-peptides) are mixed in solution, they form a six-helix bundle, which represents the postfusion gp41 structure (1–3). Three N-peptides form a central parallel trimeric coiled coil (N-trimer) surrounded by three antiparallel helical C-peptides that nestle into long grooves between neighboring N-peptides. The importance of this structure is indicated by the dominant negative inhibition of HIV entry by N- and C-peptides (4).
The available inhibitory and structural data support a working model of HIV membrane fusion (Fig. 1) (4). Initially, gp120 interacts with cellular CD4 and a chemokine coreceptor (typically CXCR4 or CCR5), causing large conformational changes in gp120 that propagate to gp41 via the gp120–gp41 interface. gp41 then undergoes a dramatic structural rearrangement that exposes its N-terminal fusion peptide, which embeds in the target cell membrane. At this stage of fusion, gp41 adopts an extended “prehairpin intermediate” conformation that bridges both viral and cellular membranes and exposes its N-trimer region. This intermediate is relatively long-lived (minutes) (4–6) but ultimately collapses as the N- and C-peptide regions of each gp41 monomer associate to form a hairpin structure. Three such hairpins (trimer-of-hairpins) form the six-helix bundle, which forces the viral and cellular membranes into tight apposition inducing membrane fusion.
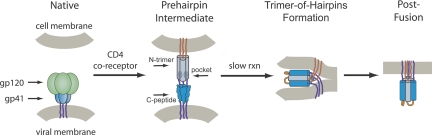
Fig. 1.
HIV entry pathway. Upon cellular receptor recognition, gp120 and gp41 undergo conformational changes resulting in exposure of the N-trimer and its hydrophobic pocket in the prehairpin intermediate. Formation of the trimer-of-hairpins structure juxtaposes cellular and viral membranes and causes fusion. The gp41 fusion peptide (red) and transmembrane domain (purple) are also shown. For clarity, gp120 is omitted from the prehairpin intermediate. Adapted from ref. 35.
According to this model, any inhibitor that binds to the N-trimer and prevents hairpin formation will inhibit viral entry. This prediction has been well supported by the discovery of numerous peptide, protein, and small-molecule inhibitors that bind the N-trimer (7). A particularly interesting feature of the N-trimer is the deep hydrophobic “pocket” formed by the N-peptide's 17 C-terminal residues. This pocket has several enticing features as an inhibitory target including (i) a very highly conserved sequence (1, 8, 9), (ii) an essential role in viral entry (10), (iii) a compact binding site that is vulnerable to inhibition by small molecules or short peptides, and (iv) the availability of several designed peptides that authentically mimic the pocket structure (e.g., IQN17, IZN17, 5-helix, and NCCGN13) (8, 9, 11, 12).
The first direct proof that pocket-specific binding is sufficient to block HIV-1 entry was provided by D-peptides that bind to the N-trimer pocket and inhibit HIV-1 entry with modest potency (IC50 ≈ 10 μM) (8). Numerous other attempts have been made to develop potent, pocket-specific entry inhibitors including minimized C-peptides (13–15), helical mimics (16, 17), antibodies (18), and small molecules (19–23), but all of these inhibitors suffer from limited potency and/or toxicity in standard viral infectivity or cell–cell fusion assays.
The only currently approved HIV-1 entry inhibitor is Trimeris' Fuzeon (also known as T-20 or enfuvirtide), a 36-residue C-peptide that binds to the N-trimer, but not the pocket region (24, 25). Although highly effective, Fuzeon has several serious limitations that have hampered its widespread clinical adoption, including delivery by injection, high dosing requirements (90 mg, twice daily), cost (approximately $25,000 per year), and the emergence of resistant strains both in vitro (25) and in patients (26). As a result, Fuzeon's use has been limited to salvage therapy for patients with multidrug-resistant HIV.
Several of Fuzeon's limitations stem from protease sensitivity, a problem common to all unstructured L-peptides. In contrast, D-peptides have several theoretical advantages: (i) they are resistant to proteases (27), which can dramatically increase serum half-life (28), (ii) short D-peptides can be absorbed systemically when taken orally (29, 30), whereas L-peptides must be injected to avoid digestion, and (iii) D-peptides are a rich unexplored source of structural diversity because they can bind to targets with unique interface geometries not available to L-peptides. Despite these potential advantages, however, the promise of D-peptides has thus far been largely unfulfilled.
In this study we used modified mirror-image phage display screens and structure-assisted design to discover D-peptide pocket-specific inhibitors of entry (PIE) with up to 40,000-fold improved antiviral potency over previously reported D-peptides (8). These D-peptides are promising leads for the prevention of HIV-1 infection (microbicides) and will allow the therapeutic potential of D-peptides to be tested.
Results and Discussion
Previously, Eckert et al. (8) used mirror-image phage display to discover a first generation of D-peptides that bind specifically to the hydrophobic pocket of the gp41 N-trimer and inhibit HIV-1 entry (IC50 = 11–270 μM, HXB2 strain). Briefly, in mirror-image phage display (31), the desired natural target is made synthetically with D-amino acids and is used to screen for binding of L-peptides displayed on phage. By symmetry, D-peptide versions of the phage peptides will bind to the natural L-target. This phage library contained 10 randomized residues (10-mer library) flanked by cysteines (CX10C). Because of the vast possible sequence diversity of this library, only one in ≈3 × 106 possible sequences was screened, and we therefore reasoned that more potent D-peptide inhibitors likely remained to be discovered.
Importantly, a consensus sequence (CX5EWXWLC) was identified from the original phage screen that allowed us to develop a constrained library in which the consensus residues (underlined) were fixed while the other six positions were randomized. This constraint allowed us to construct a comprehensive library that comprised all possible sequences. As expected, phage display screening of this library identified a family of D-peptides with improved average potency over the original D-peptides (≈4-fold; data not shown). Surprisingly, one of the most potent D-peptides identified (2K-PIE1) was an 8-mer (i.e., missing two of the randomized residues, CX3EWXWLC). This phage clone (PIE1-ϕ) was not intentionally part of the library and likely arose from a very rare replication error. The selection of this sequence despite its very low prevalence in the initial library suggested that the 8-mer family might be a richer source of tight binders than the 10-mers.
Crystal Structure of the IQN17:2K-PIE1 Complex.
To more fully understand the interaction of 2K-PIE1 with its target we determined the crystal structure of its complex with the gp41 N-trimer pocket mimic IQN17 (8) (Fig. 2). The structure was solved at 1.7 Å by molecular replacement and contains two IQN17 subunits and two 2K-PIE1 inhibitors in the asymmetric unit. A crystallographic threefold axis generates two trimers from the two independent subunit–inhibitor complexes [see supporting information (SI) Table 3 and SI Text for a description of data collection and refinement statistics]. Electron density clearly shows a number of important features of the inhibitor, including the main pocket-binding residues (dTrp10, dTrp12, and dLeu13) and the disulfide bond between dCys5 and dCys14 (Fig. 2B).
Fig. 2.
Structural analysis of the IQN17:2K-PIE1 inhibitor complex. (A) IQN17, consisting of IQ (orange) and gp41 (N17, gray) segments, with inhibitors (green, yellow, and purple) located in the canonical gp41 binding pockets. The purple inhibitor is mostly occluded in this view. (B) Omit map for 2K-PIE1 contoured at 3.0 × rmsd. Five of the eight pocket residues (gray, HXB2 numbering) that make hydrophobic contacts with 2K-PIE1 (green) are shown. Two hydrogen bonds (black) at the binding interface are also shown. (C) Overlay of D10-p1 (slate) and 2K-PIE1 (green) superposed by alignment of the IQN17 trimers. Intramolecular disulfide bonds (solid yellow) are also shown. (D) A slab view through the center of 2K-PIE1 (green) reveals an intact hydrophobic core (black) that excludes water. (E) A similar view of D10-p1 (slate) reveals the presence of several water molecules (red) in its core that nearly form a water channel. (F) End-on view of the complex (same color scheme as A) in which the surface from the last three residues of IQN17 have been removed. This view illustrates the packing of the inhibitor into the deep hydrophobic pocket. dK2 (blue), equivalent to the N-terminal Lys in PIE7 used for cross-linking, is highlighted.
Comparison of our 2K-PIE1 and the previously reported D10-p1 (8) structure, both of which were determined in complex with IQN17, reveals a striking similarity in the pocket-binding interface (Fig. 2C). The inhibitors' pocket-binding residues dTrp10, dTrp12, and dLeu13 (2K-PIE1 numbering), which contribute ≈60% of the binding surface, are nearly superposable (Fig. 2C). The essentially identical binding interfaces and buried solvent-accessible surface areas (475 Å2 for 2K-PIE1 vs. 469 Å2 for D10-p1) are surprising in light of 2K-PIE1's significantly improved potency over D10-p1 and suggest that binding of these inhibitors depends significantly on factors remote from the direct contact surface.
Overall, the comparison suggests that the improved potency and binding (see below) of 2K-PIE1 is a consequence of its reduced size (10-mer to 8-mer), which creates a more compact D-peptide with better packing while maintaining the pocket-binding interface. One major difference between the inhibitors is the path of the backbone distal to the pocket interface (Fig. 2C). dPro8 in 2K-PIE1 appears to facilitate the turn required for circularization, possibly allowing other residues to adopt more relaxed conformations. In support of this idea, a Pro in this position appears to be a better solution for 8-mers than other residues (see below). The more compact structure of 2K-PIE1 vs. D10-p1 (volume is 1,556 vs. 1,858 Å3, excluding N-terminal Lys residues) allows it to form a better-packed hydrophobic core (Fig. 2 D and E) that excludes the water molecules seen in the core of D10-p1 (Fig. 2E).
Phage Display of an 8-mer Library.
The surprising emergence of 2K-PIE1 from a 10-mer library and its apparent structural advantages motivated us to perform a dedicated screen of 8-mer sequences. We generated a comprehensive 1.5 × 108 member 8-mer phage library of the form CX4WXWLC (3.4 × 107 possible sequences). Our mirror-image target was the second-generation trimeric pocket mimic IZN17 (12).
For this screen, we used solution-phase phage display (32) combined with a soluble competitor to increase selection pressure (see SI Text for additional details). Several sequences were identified after six rounds of phage display and characterized in a phage clone binding assay (SI Fig. 5).
Potency of D-Peptides Against HXB2 Entry.
D-peptide versions of the best phage clones (PIE2, PIE7, and PIE8-ϕ) were synthesized and tested against the standard HIV-1 laboratory strain HXB2 in a single-cycle viral infectivity assay (Table 1 and Fig. 3A). As expected from the phage binding data, PIE7 is the most potent inhibitor (IC50 = 620 nM) and is ≈15-fold more potent than the best first-generation D-peptide (D10-p5).
Table 1.
D-peptide binding and neutralization
| Sample | Sequence | HXB2 IC50,* μM | HXB2 KD,* μM | JRFL IC50, μM | BaL IC50, μM |
|---|---|---|---|---|---|
| D10-p5 | KKGACELLGWEWAWLCAA | 9.5† | 7.0 | >25† | 12† |
| 2K-PIE1 | KKGACESPEWRWLCAA | 2.2 | ND | NI | — |
| 2K-PIE2 | KKGACDYPEWRWLCAA | 2.6 | 0.17 | — | — |
| PIE2 | KGACDYPEWRWLCAA | 1.3 | 0.13 | 66 | — |
| PIE7 | KGACDYPEWQWLCAA | 0.62 | 0.08 | 24 | 2.2 |
| PIE8 | KGACDYKEWQWLCAA | 1.7 | 0.17 | ≈110 | — |
| PEG-PIE7 | PEG–KGACDYPEWQWLCAA | 0.94 | 0.12 | ≈90 | — |
| (PIE7)2 | PEG–(KGACDYPEWQWLCAA)2 | 0.0019 | 0.0025 | 2.3 | 0.0073 |
| (PIE7)3 | PEG–(KGACDYPEWQWLCAA)3 | 0.00025 | ≈0.00007 | 0.22 | 0.00065 |
| C37 | — | 0.0014 | 0.00093 | 0.013 | 0.0026 |
| Fuzeon | — | 0.0037 | — | 0.005 | 0.0063 |
To aid comparisons, all D-peptides are numbered relative to PIE1 (i.e., PIE7 starts at Lys-2). ND, not determined because of complex binding behavior; NI, no inhibition seen at 100 μM.
*IC50 SEM is <25% and KD SEM is < 5% for duplicate assays for all values.
†Toxicity was observed at 100 μM for D10-p5, and this point was excluded. No other toxicity was observed (see Materials and Methods).
Fig. 3.
Representative viral entry inhibition data. Each point represents the average of quadruplicate measurements normalized to uninhibited control. Error bars represent the SEM. (A) IC50 curves for various inhibitors against HXB2. (B) IC50 curves for PIE7 and (PIE7)3 against JRFL, BaL, and HXB2.
The importance of optimizing residues that do not directly contact the pocket is highlighted by several pairwise comparisons (using 2K-PIE1 numbering) between peptides in Table 1 and SI Fig. 5. For example, PIE7 differs from PIE2 only at residue 11, for which Gln is preferred over Arg. Similarly, PIE7 differs from PIE8 only at residue 8, where Pro is preferred.
It was previously noted that introduction of Lys residues at the N terminus of D-peptides, required for solubility, adversely affects potency (8). 2K-PIE2 is ≈2-fold less potent than PIE2 (Table 1). Because 1K versions of our second-generation D-peptides have good solubility and improved potency, we decided to make 1K the standard N terminus of our second-generation D-peptides (all second-generation peptides have a single N-terminal Lys unless otherwise labeled).
Crystal Structure of the IQN17:PIE7 Complex.
In an attempt to understand the source of PIE7's improved affinity compared with 2K-PIE1, we determined two independent crystal structures of PIE7 in complex with IQN17 at 2.0-Å and 1.66-Å resolution (see SI Table 3 and SI Text for a description of data collection and refinement statistics). A comparison of 2K-PIE1 and PIE7 reveals several interesting differences (Fig. 4). First, an intramolecular polar contact between the hydroxyl of dSer7 and the carbonyl of dGly3 in 2K-PIE1 is lost in PIE7 but is replaced with a new interaction between the side chain carboxylate of dAsp6 and the amide of dGly3. Second, new hydrophobic interactions are created in PIE7 between the ring carbons of dTyr7 and the pocket residue Trp-571 (SI Fig. 6A). Third, the carbonyl of dLys2 of PIE7, although somewhat flexible in orientation, forms a direct hydrogen bond with the ε nitrogen of Trp-571 in some of the structures. This interaction is water-mediated in 2K-PIE1. Fourth, in some of the structures the hydroxyl of dTyr7 in PIE7 forms a new water-mediated hydrogen bond with the pocket residue Gln-575. This interaction cannot be formed in the 2K-PIE1 structure. These subtle changes expand PIE7's pocket binding interface and may account for the enhanced potency of PIE7.
Fig. 4.
Structural analysis of the IQN17:2K-PIE1 and IQN17:PIE7 inhibitor complexes. Shown is a comparison of unique polar contacts observed in the 2K-PIE1 (A) and PIE7 (B) costructures (described in the text).
Multimeric D-Peptides.
Based on the trimeric nature of gp41, we predicted that multimeric D-peptides would have significantly improved affinity for the N-trimer and enhanced antiviral potency. To test this idea, we used a bis(NHS ester)PEG cross-linker to dimerize PIE7 via its unique primary amine (N-terminal Lys) (Fig. 2F). The length of the PEG spacer (35 Å) was chosen to cover, with slack, the distance between the N-termini of neighboring D-peptides in the crystal structures. To construct the PIE7 trimer, we used two of the same cross-linker to connect a central 2K-PIE7 to two flanking PIE7s. PEG is an ideal material for cross-linking because it is highly flexible, very soluble, nonimmunogenic, and has been used in several approved therapeutic peptides and proteins (33).
The resulting dimeric and trimeric inhibitors, (PIE7)2 and (PIE7)3, have IC50 values of 1.9 nM and 250 pM (Table 1 and Fig. 3A) against HXB2, respectively. This dramatic gain in potency is a 325- and 2,500-fold improvement over the PIE7 monomer, respectively. To control for possible nonspecific effects of the PEG moiety, we reacted mono(NHS ester)PEG with PIE7 to generate PEG-PIE7. Addition of this PEG group caused a modest ≈1.5-fold reduction in potency against HXB2. Therefore, the improved potency of the oligomers cannot be attributed to an interaction of the PEG with virus, cells, or the D-peptide but is a genuine avidity effect caused by multiple D-peptides binding to the N-trimer.
Surface Plasmon Resonance (SPR) Characterization.
To determine whether the improved potency of our second-generation D-peptides stems from optimization of affinity for the pocket, we characterized the binding properties of the D-peptides to an immobilized N-trimer mimic (IZN36) using SPR (Table 1). The rank order of measured KD values correlated well with antiviral IC50 values, indicating that D-peptide binding to a pocket mimic in vitro is a good predictor of antiviral potency.
The PIE7 monomer and multimers had similar rapid association rates, but the dimer and trimer (data not shown) showed dramatically slowed dissociation rates compared with the monomer (SI Fig. 7). The trimer's binding to the pocket was too tight (low to mid pM) to measure accurately by SPR (the value reported in Table 1 is approximate and likely underestimates the trimer's true affinity). Interestingly, the trimer's potency against HXB2 did not improve as much as expected from its KD, suggesting that trimer potency against HXB2 may have reached a potency limit imposed by association kinetics, as has been reported for another entry inhibitor, 5-helix (34). This kinetic limitation is expected because the exposed N-trimer has an ≈10- to 20-min lifetime in the gp41 prehairpin intermediate (4–6), similar to the time required for binding of our peptides at mid to high pM concentrations.
D-Peptide Inhibitors Are Also Active Against Primary HIV-1 Strains.
HXB2 is a widely used laboratory-adapted HIV-1 strain that is typically more sensitive to entry inhibitors than primary (clinical) HIV-1 strains. Here we report D-peptide inhibitory data against primary strains (standard clade B strains BaL and JRFL). The most potent first-generation D-peptide, D10-p5, showed little or no inhibitory activity against JRFL and modest activity against BaL (Table 1). In contrast, PIE7 inhibits both JRFL (IC50 = 24 μM) and BaL (IC50 = 2.2 μM) entry, although ≈40- and ≈4-fold less potently than HXB2 entry, respectively (Table 1). Similar differences in potency between JRFL and HXB2 have been reported for other entry inhibitors that target the pocket region (18) (e.g., the C-peptide inhibitor C37) (Table 1). Interestingly, Fuzeon, which does not bind to the pocket, shows only a small loss of activity against JRFL (Table 1). Despite the relative insensitivity of JRFL to the PIE7 monomer, the PIE7 trimer is a moderately potent inhibitor of this strain (IC50 = 220 nM) and an extremely potent inhibitor against BaL (IC50 = 650 pM).
Possible Sources of JRFL's Relative Insensitivity to Inhibition by PIE7.
Compared with the sequence of the BaL and HXB2 pocket region (N17), JRFL has the conservative L565M substitution (Table 2, highlighted). All other pocket residues that contact our D-peptides are >97% identical in the >5,000 clade A, B, and C HIV-1 strains from the Los Alamos National Laboratory HIV sequence database (www.hiv.lanl.gov). Residue 565 is Leu or Met in ≈99% of these strains. Our crystal structures show that the D-peptide C-terminal Ala interacts with the L565 equivalent position of IQN17 in each of the available crystal structures (e.g., Fig. 2B). Residue 580 (Table 2, highlighted) does not contact the pocket.
Table 2.
Pocket region (N17) alignment
| Strain | Sequence |
|---|---|
| HXB2 | LLQLTVWGIKQLQARIL |
| JRFL | MLQLTVWGIKQLQARVL |
| BaL | LLQLTVWGIKQLQARVL |
The L565M substitution might affect binding of our D-peptides to the JRFL pocket. To test this possibility, we measured binding of PIE7 to JRFL and HXB2 versions of IZN36 by SPR and observed an ≈4-fold increase in KD for binding to the JRFL pocket (data not shown). Another possible contributing factor to JRFL's relative insensitivity is the reduced steric accessibility of JRFL's N-trimer region compared with HXB2 (35), which may explain why PEG-PIE7 has ≈4-fold lower potency than PIE7 against JRFL (vs. a 1.5-fold difference against HXB2) (Table 1). These differences in inhibitor binding caused by the L565M substitution or reduced steric accessibility do not appear to fully account for the ≈40-fold reduction in potency against JRFL. Therefore, an unknown complex factor (e.g., kinetics) is also likely to be involved.
Third Generation D-Peptides.
Although our second-generation multimeric D-peptides are sufficiently potent to begin preclinical studies, the ideal D-peptide for clinical use will require further optimization to improve potency against challenging strains like JRFL. Our current results suggest several straightforward strategies to further improve D-peptide potency and achieve this goal. First, we predict that optimization of the cross-linker length and connectivity in our multimeric D-peptides will strengthen avidity. In our initial strategy, we connected the N termini of monomers (N–N) via the existing N-terminal Lys in PIE7. From the crystal structures, it is apparent that C–C or C–N linkages could be significantly shorter than our current N–N linkage. Second, our structures show that flanking residues (those outside the disulfide bond) present on the phage (and peptides) make important interactions with the pocket. Optimization of these residues in the context of the PIE7 core sequence by phage display will likely provide further improvements in potency.
Third, it is possible that 8-mers are not the most optimal length for these D-peptides. Modeling one or two residue deletions from the 2K-PIE1 structure indicates that 7-mers are still long enough to present the WXWL binding motif to the pocket and maintain the disulfide bond, whereas 6-mers are not (data not shown). Screening of a naïve 7-mer library (CX7C) again identified the WXWL consensus motif and confirmed that 7-mers can bind the pocket and inhibit HIV entry (B.D.W., Y. Shi, and M.S.K., unpublished results). Future phage display screening will ultimately determine which geometry is most optimal for high-affinity pocket binding. Fourth, the crystal structure of PIE7 (our best current monomer) is a valuable platform for rational design using nonnatural amino acid derivatives. For example, PIE7's dTyr7 hydroxyl is not optimally positioned to make direct hydrogen bonds with the pocket. It may be possible to stabilize the complex by extending the position of the dTyr7 hydroxyl by one or two carbons.
Finally, it is important to note that the avidity of our multimers predicts that small improvements in the potency of monomers will result in geometric improvements in the corresponding dimers and trimers, up to the potency limit imposed by association kinetics. We also predict that it will be beneficial to “overengineer” future D-peptides to improve affinity even after reaching this potency limit. Such inhibitors will not show improved potency, but will have a reserve of binding energy that acts as a “resistance capacitor” to defend against potential resistance mutations [i.e., resistance mutations that moderately affect binding would have no effect on potency, as has been reported for the entry inhibitor 5-helix (34)]. Of particular importance, this property will discourage the stepwise accumulation of multiple subtle mutations that combine to confer resistance. Individual mutations would have no effect on inhibitor potency and would not confer a growth advantage in the presence of inhibitors. This resistance capacitor would be especially beneficial for trimeric inhibitors, because resistance mutations would simultaneously affect all three pockets. As a further defense against the development of resistance, our trimeric D-peptides could also be constructed by using three different D-peptide sequences, each with a distinct resistance profile. Such a heterotrimer would present a significant additional barrier to the development of resistance.
Potential Uses of D-Peptides.
Our D-peptides target the highly conserved gp41 hydrophobic pocket region and will likely have an improved resistance profile compared with Fuzeon (25), which targets a less well conserved region of gp41. Further studies of our D-peptides against panels of viruses from diverse HIV-1 clades and in vitro selections for resistance mutations will be required to determine the breadth of their activity and predict susceptibility to resistance mutations. Because the hydrophobic pocket is not targeted by Fuzeon or other entry inhibitors currently in advanced clinical trials (e.g., BMS-378806, PRO 542, Vicriviroc, and Maraviroc), our D-peptides should be additive (or possibly synergistic) with these inhibitors and could form part of a emerging entry inhibitor “cocktail,” similar to the mixtures of HIV-1 protease and reverse transcriptase inhibitors currently used in highly active antiretroviral therapy.
D-peptides represent a new class of drugs that have not been extensively tested in vivo. Because D-peptides are not degraded by proteases they have the potential for oral bioavailability (29, 30), extended persistence in circulation (28), reduced immunogenicity (36), long shelf life, and use in harsh mucosal environments as a topical prophylactic microbicide. The D-peptides reported here are now sufficiently potent for preclinical studies, which will ultimately determine whether these theoretical advantages translate into a valuable new class of agents for the prevention and treatment of HIV/AIDS. These results also suggest that D-peptides may be useful for diverse applications against other therapeutic targets.
Materials and Methods
Peptide and Protein Production.
All peptides and proteins were produced and purified as described in SI Text.
Phage Display.
Phage display was performed by using the M13KE plasmid from New England Biolabs as described in SI Text.
Viral Infectivity Assay.
Pseudovirions were produced and viral infectivity was measured essentially as previously described (35), using HOS-CD4-CXCR4 (for HXB2) or HOS-CD4-CCR5 (for BaL and JRFL) target cells, with the following modifications: 8 μg/ml DEAE-Dextran was used as a fusion enhancer, and media were changed ≈24 h after infection. Samples dissolved in DMSO for solubility [D10-p5, PEG-PIE7, and (PIE7)2] were tested at 1% final DMSO concentration and normalized to an uninhibited control containing 1% DMSO. The following reagents were obtained through the AIDS Research and Reference Reagent Program, Division of AIDS, National Institute of Allergy and Infectious Diseases, National Institutes of Health: T-20 (Fuzeon) and BaL.01 from Roche and John Mascola, respectively.
The cellular toxicity of all D-peptides was assessed by (i) microscopic inspection of cells ≈24 h after infection and (ii) measurement of luciferase levels in inhibitor-treated cells infected with VSV-G pseudotyped virions (same protocol as above). Because the D-peptides are not expected to inhibit VSV-G-mediated entry, a >10% reduction in luciferase signal was interpreted as cellular toxicity.
Crystallography.
Details of crystallographic methods and data analysis are described in SI Text.
SPR Analysis.
SPR was performed as described (35). Each binding study was performed in duplicate at 20°C by using a 3-fold decreasing concentration series starting from 10 μM for D10-p5, 60 nM for C37, (PIE)2, and (PIE7)3, or 2 μM (all others). Only 2K-PIE1, C37, (PIE7)2, and (PIE7)3 required a specific surface regeneration procedure (one 20-s pulse of 6 M GuHCl or 0.05% SDS). The IZN36 surface was very stable to these regeneration conditions. Data were analyzed by using Scrubber2 (BioLogic Software).
Supplementary Material
Acknowledgments
We thank Debra Eckert and Yu Shi for valuable advice and discussion, John McIntosh for cross-linking assistance, Lucrezia Solano for JRFL-IZN36 cloning assistance, and Jun Aishima for technical assistance and discussion. We also thank the following University of Utah core facilities: DNA and Peptide (Robert Schackmann and Scott Endicott), Protein Interactions (David Myszka), DNA Sequencing, and Mass Spectrometry. Operations of the National Synchrotron Light Source are supported by the Office of Basic Energy Sciences at the U.S. Department of Energy and by the National Institutes of Health. Data collection at the National Synchrotron Light Source was funded by the National Center for Research Resource. This research was funded by the National Institutes of Health (Grant PO1 GM66521 and a Training Grant in Biological Chemistry), the University of Utah Technology Commercialization Project, and the American Cancer Society (Grant PF0304001GMC).
Abbreviations
- SPR
surface plasmon resonance
- PIE
pocket-specific inhibitor of entry.
Footnotes
The authors declare no conflict of interest.
Data deposition: The atomic coordinates have been deposited in the Protein Data Bank, www.pdb.org (PDB ID codes 2R3C, 2R5B, and 2R5D).
This article contains supporting information online at www.pnas.org/cgi/content/full/0708109104/DC1.
References
- 1.Chan DC, Fass D, Berger JM, Kim PS. Cell. 1997;89:263–273. doi: 10.1016/s0092-8674(00)80205-6. [DOI] [PubMed] [Google Scholar]
- 2.Weissenhorn W, Dessen A, Harrison SC, Skehel JJ, Wiley DC. Nature. 1997;387:426–430. doi: 10.1038/387426a0. [DOI] [PubMed] [Google Scholar]
- 3.Tan K, Liu J, Wang J, Shen S, Lu M. Proc Natl Acad Sci USA. 1997;94:12303–12308. doi: 10.1073/pnas.94.23.12303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Eckert DM, Kim PS. Annu Rev Biochem. 2001;70:777–810. doi: 10.1146/annurev.biochem.70.1.777. [DOI] [PubMed] [Google Scholar]
- 5.Chan DC, Kim PS. Cell. 1998;93:681–684. doi: 10.1016/s0092-8674(00)81430-0. [DOI] [PubMed] [Google Scholar]
- 6.Furuta RA, Wild CT, Weng Y, Weiss CD. Nat Struct Biol. 1998;5:276–279. doi: 10.1038/nsb0498-276. [DOI] [PubMed] [Google Scholar]
- 7.Root MJ, Steger HK. Curr Pharm Des. 2004;10:1805–1825. doi: 10.2174/1381612043384448. [DOI] [PubMed] [Google Scholar]
- 8.Eckert DM, Malashkevich VN, Hong LH, Carr PA, Kim PS. Cell. 1999;99:103–115. doi: 10.1016/s0092-8674(00)80066-5. [DOI] [PubMed] [Google Scholar]
- 9.Root MJ, Kay MS, Kim PS. Science. 2001;291:884–888. doi: 10.1126/science.1057453. [DOI] [PubMed] [Google Scholar]
- 10.Chan DC, Chutkowski CT, Kim PS. Proc Natl Acad Sci USA. 1998;95:15613–15617. doi: 10.1073/pnas.95.26.15613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Louis JM, Bewley CA, Clore GM. J Biol Chem. 2001;276:29485–29489. doi: 10.1074/jbc.C100317200. [DOI] [PubMed] [Google Scholar]
- 12.Eckert DM, Kim PS. Proc Natl Acad Sci USA. 2001;98:11187–11192. doi: 10.1073/pnas.201392898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Judice JK, Tom JY, Huang W, Wrin T, Vennari J, Petropoulos CJ, McDowell RS. Proc Natl Acad Sci USA. 1997;94:13426–13430. doi: 10.1073/pnas.94.25.13426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jin BS, Ryu JR, Ahn K, Yu YG. AIDS Res Hum Retroviruses. 2000;16:1797–1804. doi: 10.1089/08892220050195757. [DOI] [PubMed] [Google Scholar]
- 15.Sia SK, Carr PA, Cochran AG, Malashkevich VN, Kim PS. Proc Natl Acad Sci USA. 2002;99:14664–14669. doi: 10.1073/pnas.232566599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ernst JT, Kutzki O, Debnath AK, Jiang S, Lu H, Hamilton AD. Angew Chem Int Ed Engl. 2002;41:278–281. doi: 10.1002/1521-3773(20020118)41:2<278::aid-anie278>3.0.co;2-a. [DOI] [PubMed] [Google Scholar]
- 17.Stephens OM, Kim S, Welch BD, Hodsdon ME, Kay MS, Schepartz A. J Am Chem Soc. 2005;127:13126–13127. doi: 10.1021/ja053444+. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Miller MD, Geleziunas R, Bianchi E, Lennard S, Hrin R, Zhang H, Lu M, An Z, Ingallinella P, Finotto M, et al. Proc Natl Acad Sci USA. 2005;102:14759–14764. doi: 10.1073/pnas.0506927102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Debnath AK, Radigan L, Jiang S. J Med Chem. 1999;42:3203–3209. doi: 10.1021/jm990154t. [DOI] [PubMed] [Google Scholar]
- 20.Ferrer M, Kapoor TM, Strassmaier T, Weissenhorn W, Skehel JJ, Oprian D, Schreiber SL, Wiley DC, Harrison SC. Nat Struct Biol. 1999;6:953–960. doi: 10.1038/13324. [DOI] [PubMed] [Google Scholar]
- 21.Zhao Q, Ernst JT, Hamilton AD, Debnath AK, Jiang S. AIDS Res Hum Retroviruses. 2002;18:989–997. doi: 10.1089/08892220260235353. [DOI] [PubMed] [Google Scholar]
- 22.Jiang S, Lu H, Liu S, Zhao Q, He Y, Debnath AK. Antimicrob Agents Chemother. 2004;48:4349–4359. doi: 10.1128/AAC.48.11.4349-4359.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Frey G, Rits-Volloch S, Zhang XQ, Schooley RT, Chen B, Harrison SC. Proc Natl Acad Sci USA. 2006;103:13938–13943. doi: 10.1073/pnas.0601036103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wild CT, Shugars DC, Greenwell TK, McDanal CB, Matthews TJ. Proc Natl Acad Sci USA. 1994;91:9770–9774. doi: 10.1073/pnas.91.21.9770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rimsky LT, Shugars DC, Matthews TJ. J Virol. 1998;72:986–993. doi: 10.1128/jvi.72.2.986-993.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wei X, Decker JM, Liu H, Zhang Z, Arani RB, Kilby JM, Saag MS, Wu X, Shaw GM, Kappes JC. Antimicrob Agents Chemother. 2002;46:1896–1905. doi: 10.1128/AAC.46.6.1896-1905.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Milton RC, Milton SC, Kent SB. Science. 1992;256:1445–1448. doi: 10.1126/science.1604320. [DOI] [PubMed] [Google Scholar]
- 28.Sadowski M, Pankiewicz J, Scholtzova H, Ripellino JA, Li Y, Schmidt SD, Mathews PM, Fryer JD, Holtzman DM, Sigurdsson EM, et al. Am J Pathol. 2004;165:937–948. doi: 10.1016/s0002-9440(10)63355-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Pappenheimer JR, Dahl CE, Karnovsky ML, Maggio JE. Proc Natl Acad Sci USA. 1994;91:1942–1945. doi: 10.1073/pnas.91.5.1942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pappenheimer JR, Karnovsky ML, Maggio JE. J Pharmacol Exp Ther. 1997;280:292–300. [PubMed] [Google Scholar]
- 31.Schumacher TN, Mayr LM, Minor DL, Jr, Milhollen MA, Burgess MW, Kim PS. Science. 1996;271:1854–1857. doi: 10.1126/science.271.5257.1854. [DOI] [PubMed] [Google Scholar]
- 32.Barbas CF. Phage Display: A Laboratory Manual. New York: Cold Spring Harbor Lab Press; 2001. [Google Scholar]
- 33.Harris JM, Chess RB. Nat Rev Drug Discovery. 2003;2:214–221. doi: 10.1038/nrd1033. [DOI] [PubMed] [Google Scholar]
- 34.Steger HK, Root MJ. J Biol Chem. 2006;281:25813–25821. doi: 10.1074/jbc.M601457200. [DOI] [PubMed] [Google Scholar]
- 35.Hamburger AE, Kim S, Welch BD, Kay MS. J Biol Chem. 2005;280:12567–12572. doi: 10.1074/jbc.M412770200. [DOI] [PubMed] [Google Scholar]
- 36.Chong P, Sia C, Tripet B, James O, Klein M. Lett Pept Sci. 1996;3:99–106. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.