Abstract
Nicotinate dehydrogenase (NDH) from Eubacterium barkeri is a molybdoenzyme catalyzing the hydroxylation of nicotinate to 6-hydroxynicotinate. Reactivity of NDH critically depends on the presence of labile (nonselenocysteine) selenium with an as-yet-unidentified form and function. We have determined the crystal structure of NDH and analyzed its active site by multiple wavelengths anomalous dispersion methods. We show that selenium is bound as a terminal Mo Se ligand to molybdenum and that it occupies the position of the terminal sulfido ligand in other molybdenum hydroxylases. The role of selenium in catalysis has been assessed by model calculations, which indicate an acceleration of the critical hydride transfer from the substrate to the selenido ligand in the course of substrate hydroxylation when compared with an active site containing a sulfido ligand. The MoO(OH)Se active site of NDH shows a novel type of utilization and reactivity of selenium in nature.
Se ligand to molybdenum and that it occupies the position of the terminal sulfido ligand in other molybdenum hydroxylases. The role of selenium in catalysis has been assessed by model calculations, which indicate an acceleration of the critical hydride transfer from the substrate to the selenido ligand in the course of substrate hydroxylation when compared with an active site containing a sulfido ligand. The MoO(OH)Se active site of NDH shows a novel type of utilization and reactivity of selenium in nature.
Keywords: Eubacterium barkeri, hydroxylase, molybdoprotein, molybdopterin, selenium
Selenium is an essential component of several enzymes and has a key role in various biological redox processes. Usually selenium occurs in proteins as selenocysteine, which is cotranslationally inserted as the 21st amino acid (1) and is found in a variety of proteins in all 3 kingdoms of life (2). Selenium also finds a natural use as 5-methylaminomethyl-2-selenouridine in the “wobble” position of some tRNAs (3). The ionic radii and electronegativities of selenium and sulfur are similar, but selenide is a stronger reducing agent than sulfide. Because of the lower pKa value of selenols compared with thiols selenocysteine is deprotonated under physiological conditions, whereas cysteine is mostly protonated (4). The ionization state together with the better polarizability of selenium makes selenocysteine a good nucleophile. Several molybdenum- and tungsten-containing enzymes have been shown to contain selenium (5), and a recent comprehensive genomic analysis has revealed a clear relationship between selenium and molybdenum utilization across all 3 domains of life (6). Selenium is found as selenocysteine in some prokaryotic molybdenum-containing oxotransferases like formate dehydrogenase H from Escherichia coli, where it coordinates molybdenum in the oxidized state of the enzyme (7–9). In other enzymes, notably members of the molybdenum hydroxylase family (10, 11), like nicotinate dehydrogenase (NDH) from Eubacterium barkeri (12–14) and the xanthine oxidoreductases (XORs) of Clostridium purinolyticum (15), Clostridium acidiurici (16) and Eubacterium barkeri (17) and the purine hydroxylase from C. purinolyticum (15) contain a labile (nonselenocysteine) selenium in an unidentified form essential for their reactivity (18).
Molybdenum hydroxylases catalyze the hydroxylation of various organic molecules following the general scheme:
This hydroxylation reaction is unique in biology as it uses water as the source for the hydroxyl oxygen and not dioxygen (10). The active site of molybdenum hydroxylases contains molybdenum coordinated by the enedithiolate group of a pyranopterin cofactor, commonly referred to as molybdopterin. The crystal structures of different molybdenum hydroxylases in complex with substrates and inhibitors are known (19–27). These structures demonstrate that the molybdenum coordination sphere typically consists of 1 oxo, 1 hydroxo, and 1 labile sulfido ligand in a square-pyramidal coordination sphere, with the Mo O occupying the apical position. The sulfido ligand is prone to cyanolysis, and its replacement with a second oxo ligand in the process leads to inactivation of the enzymes (10). An extension of the active site is found in aerobic carbon monoxide dehydrogenases. While in an earlier report on the structure the presence of selenium in the active site was suggested (25), a latter study using analytical multiple wavelength anomalous dispersion methods showed that the active site does not contain selenium but a linearly-coordinated Cu(I) ion bridged by a μ-sulfido ligand to the molybdenum (28). This μ-sulfido bridge occupies the position of the Mo
O occupying the apical position. The sulfido ligand is prone to cyanolysis, and its replacement with a second oxo ligand in the process leads to inactivation of the enzymes (10). An extension of the active site is found in aerobic carbon monoxide dehydrogenases. While in an earlier report on the structure the presence of selenium in the active site was suggested (25), a latter study using analytical multiple wavelength anomalous dispersion methods showed that the active site does not contain selenium but a linearly-coordinated Cu(I) ion bridged by a μ-sulfido ligand to the molybdenum (28). This μ-sulfido bridge occupies the position of the Mo S seen in other molybdenum hydroxylases.
S seen in other molybdenum hydroxylases.
The anaerobic soil bacterium E. barkeri is able to ferment nicotinate to propionate, acetate, carbon dioxide, and ammonia with the gain of 1 mol of ATP per mol of nicotinate. The fermentation of nicotinate is initiated by its hydroxylation to 6-hydroxynicotinate catalyzed by NDH (12). NDH has a (αβγδ)2 subunit structure and contains [2Fe-2S] clusters, FAD and a molybdenum center with a pyranopterin cofactor that has been elaborated as the dinucleotide of cytosine and termed molybdopterin cytosine dinucleotide (MCD) (18, 29). The genes encoding NDH occur in the transcriptional order ndhFSLM and are part of a 23.3-kb gene cluster dedicated to the fermentation of nicotinate (30). The NdhF subunit (33 kDa) carries 1 FAD molecule and the NdhS subunit (23 kDa) contains 2 [2Fe-2S] clusters. Contrary to all structurally characterized hydroxylases the molybdenum cofactor appeared to be contained not in 1 but 2 subunits: the NdhL subunit (50 kDa) and NdhM subunit (37 kDa). The most remarkable feature of NDHs is the presence of labile (nonselenocysteine) selenium (14), which is essential to catalyze the hydroxylation reaction (18). Here, we report the X-ray crystal structure of NDH and its Se-containing active site, together with computational studies demonstrating the catalytic profit of the natural selection of Se over its congeners S and O.
Results
Overall Structure.
NDH was crystallized by vapor diffusion methods under anoxic conditions in an atmosphere of 95% N2/5% H2. Crystals belonged to the space group P21 and contained the complete dimer of heterotetramers in 1 asymmetric unit. The structure of NDH was determined with Patterson search techniques by using a substructure of 4-hydroxybenzoyl-CoA reductase (23) as search model. The final model contains all residues and was refined to 2.2-Å resolution (Table 1).
Table 1.
Statistics on diffraction data and structure refinement
| Statistic | Dataset |
||
|---|---|---|---|
| Above Se edge | Below Se edge | Native | |
| Data collection | |||
| Wavelength, Å | 0.9780 | 0.9803 | 1.5418 |
| Space group | P21 | P21 | |
| Cell dimensions (a,b,c in Å, b in °) | 100.00, 72.15, 217.10, 90.56 | 97.08, 71.70, 214.49, 90.23 | |
| Total/unique reflections | 396,539/208,277 | 360,921/196,213 | 519,267/148,165 |
| Rs, % | 7.8 (31.0) | 8.7 (44.8) | 9.7 (53.6) |
| Resolution, Å | 30–2.5 (2.6–2.5) | 30–2.5 (2.6–2.5) | 30–2.2 (2.3–2.2) |
| Completeness, % | 99.2 (97.6) | 93.5 (74.9) | 98.9 (96.8) |
| (I)/(σI) | 6.8 (2.4) | 6.5 (1.5) | 10.8 (2.3) |
| Refinement | |||
| Model Rwork/Rfree factor, % | 21.2/25.0 | ||
| No. atoms | |||
| Protein | 17,180 | ||
| Ligand | 249 | ||
| Water | 954 | ||
| B factors | |||
| Protein | 36.3 | ||
| Ligand | 26.1 | ||
| Water | 29.3 | ||
| rmsd | |||
| Bond lengths, Å | 0.006 | ||
| Bond angles, ° | 1.3 | ||
In the datasets at the Se edge the values given are for unmerged Friedel mates. The values in parentheses indicate the highest resolution.
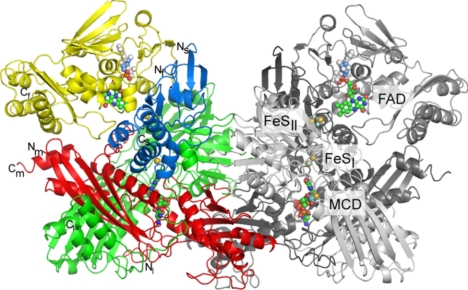
NDH is a dimer of heterotetramers with (FSLM)2 subunit composition in solution (12) and all subunits are present in the crystal structure (Fig. 1). The dimer has overall dimensions of 148 × 100 × 70 Å3. The F subunit (296 residues) harbors the FAD cofactor, which interacts with the N terminal (residues 1–57) and middle domain (residues 58–178) through its adenosin and ribityl moiety. The C-terminal domain (residues 179–291) only interacts with FAD through K187F, whose amino group is in hydrogen-bonding distance to O4 of the isoalloxazine ring. As in other molybdenum hydroxylases, the C-terminal domain contributes to shield the N5 position of the isoalloxazine ring from the solvent. In molybdenum hydroxylases unable to reduce NAD+/NADP+ access to the N5 position is usually blocked by a tyrosine (carbon monoxide dehydrogenase from Oligotropha carboxydovorans, quinoline 2-oxidoreductase) or tryptophan side chain (carbon monoxide dehydrogenase from Hydrogenophaga pseudoflava). In contrast, the less bulky side chain of isoleucine is found in the NAD+/NADP+-reducing NDH and bovine and bacterial XOR (21, 22). All 3 domains of the F subunit show a mixed α/β-fold. The S subunit (157 residues) comprises 2 domains, each coordinating 1 [2Fe-2S] cluster. The N-terminal domain (residues 1–79) is similar to “plant-type ferredoxins” and binds the [2Fe-2S] cluster close to the FAD, which is designated as Fe/S II or type II on the basis of its EPR characteristics in homologous molybdenum hydroxylases (31–33). The C-terminal domain (residues 80–157) displays a 4-helix bundle with 2-fold symmetry unique to molybdenum hydroxylases (19) and coordinates the [2Fe-2S] cluster (type I) closest to the molybdopterin. The L subunit (425 residues) and M subunit (330 residues) harbor the Mo-bound pyranopterin cofactor. Both subunits have an extended structure and lie approximately perpendicular on top of each other. The L subunit interacts with the M and S subunits and has an N-terminal extension (residues 1–28L) that wraps around the C-terminal domain of the S subunit. A middle domain (residues 29–129L and 181–277L) with 2 antiparallel β-sheets of 2 and 7 strands and 3 α-helices follows the N-terminal extension. The C-terminal domain of the L subunit is dominated by a mixed 5-stranded β-sheet flanked on 1 side by 2 α-helices that continue into a 2-stranded antiparallel β-sheet and a C-terminal α-helix. The M subunit can be divided into 2 domains, both containing mixed 4-stranded β-sheets and 3 α-helices (Fig. 1).
Fig. 1.
Overall structure of NDH. Ribbon plot representation of the NDH dimer. In the left monomer each subunit has its own color with green and red for the MCD coordinating L and M subunits, blue for the [2Fe-2S] clusters containing S subunit, and yellow for the FAD containing F subunit. The right monomer is colored in different shades of gray. Cofactors are labeled, and FeSI and FeSII indicate the position of the type I and type II [2Fe-2S] clusters of the S subunit. The shortest distances between the cofactors are 5.4 Å (MCD-FeSI), 11.5 Å (FeSI-FeSII), and 6.4 Å (FeSII-FAD), with the shortest metal-to-metal distances of 14.7 Å (MCD-FeSI) and 12.4 Å (FeSI-FeSII). All pictures were prepared by using PyMol (48).
Both monomers of NDH show the same arrangement of subunits and cofactors building 2 independently working electron transfer chains (Fig. 1), similar to other structurally characterized molybdenum hydroxylases. However, NDH is unusual in containing 4 subunits per monomer, whereas other molybdenum hydroxylases of known structure, contain 1, 2, or 3 subunits. Compared with all structurally characterized molybdenum hydroxylases, the Mo-pyranopterin binding subunit/domain has been split into 2 polypeptides in NDH: the L subunit corresponds to the N-terminal domain and the M subunit is homologous to the C-terminal domain. Bioinformatic analysis has revealed split molybdopterin subunits in 1 of 3 other bacterial NDHs (SI Text and Fig. S1). Moorella thermoacetica, Carboxydothermus hydrogenoformans, Petrotoga mobilis, Clostridium asparagiforme, 5 α-proteobacterial species, Alkaliphilus oremlandii, and Alkaliphilus metalliredigens have split subunits in NDH homologs of hitherto unknown catalytic activity (SI Text and Fig. S2).
Active-Site Structure and Selenium Detection.
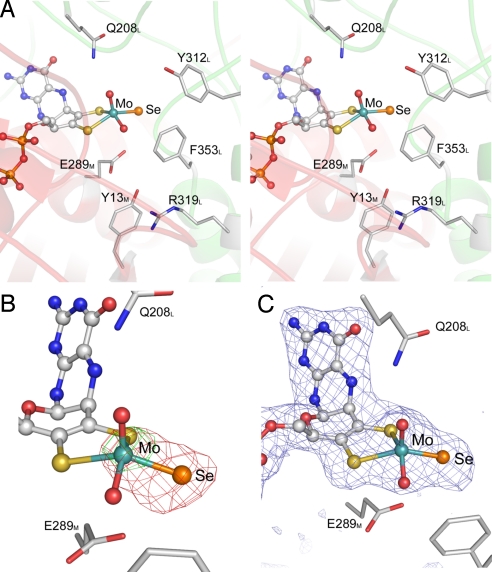
The substrate channel is formed by residues from the M and L subunits, which create a funnel leading to the active site. The active site contains the molybdenum ion with a distorted square pyramidal coordination (Fig. 2A). Two enedithiolate sulfurs and 1 hydroxo-ligand occupy the equatorial positions at the Mo-ion and an oxo-ligand is found in the apical position. The activity of NDH is known to depend on the presence of selenium in the active site, and different Se occupancies depending on preparation and specific activity have been observed (14, 18, 34). Therefore, detection of selenium could not solely rely on the strength of the observed electron densities, which depend on the occupancy of the ligands. To locate Se in the structure of NDH we collected complete crystallographic datasets at 2 different X-ray wavelengths, one at the low-energy side (λ = 0.9803 Å) and one at the high-energy side (λ = 0.9780 Å) of the X-ray absorption K-edge of selenium (Table 1). Whereas Bijvoet difference maps of the low-energy dataset show only a signal at the Mo ion, Bijvoet difference maps calculated from the high-energy dataset reveal strong additional anomalous scattering from only 1 of the equatorial ligands of the Mo ion (Fig. 2B). That this position is indeed occupied by selenium is supported by the stronger electron density observed at this position and from a Mo ligand bond length of 2.3 Å (Fig. 2C), which is longer than the typical Mo–S bond (Table S1). Se is a direct ligand and coordinates Mo in the equatorial plane. The lack of additional density around Se indicates that it is a terminal ligand. We estimated the occupancy of the Se ligand to be ≈80% by adjusting the Se occupancy of the model such that the Se atom refines to a similar B factor as the neighboring atoms including the other Mo ligands.
Fig. 2.
The Mo-Se active site of NDH. (A) Stereoview of the active-site environment. Transparent ribbons are colored as in Fig. 1 (left monomer). (B) Selenium identification by anomalous scattering detected at energies higher (λ = 0.9780 Å in red) and smaller (λ = 0.9803 Å in green) than the energy of the Se K edge. Both Bijvoet difference maps are contoured at the +5.0-σ level. (C) Fobs − Fcalc map for the Mo and pyranopterin cofactor at a contour level of +5.0 σ. For the calculation of the map all atoms of the Mo ion and the pyranopterin cofactor were omitted.
The second coordination sphere around the Mo-ion is formed by E289M in trans to the apical oxo-ligand, Q208L, in hydrogen-bonding distance to the apical oxo-ligand and residues that may contribute to the binding and stabilization of the substrate, like Y312L, R319L, F353L, and Y13M (Fig. 2A). An additional small ligand bound near the active site has been modeled as nitrate, which is present in the crystallization buffer. Nitrate interacts with the main chain of residues forming the substrate-binding pocket and is 8 Å away from the Mo ion and 6 Å away from the Se. Nitrate is found at the same position where an acetate molecule has been modeled in the crystal structure of bXOR (21) and both show the same pattern of interactions with the surrounding amino acids. A physiological role for this conserved anion-binding site near the active site has not been demonstrated.
Structure-Based Reaction Mechanism of NDH.
Increasing evidence on the chemistry of XORs supports a mechanism of substrate hydroxylation involving a base-assisted nucleophilic attack of the equatorial Mo-OH group on the substrate, with a concomitant hydride transfer from the substrate to the Mo(+VI) S group to give Mo(+IV)-SH. Two residues in the direct environment of molybdenum, Q208L in NDH (Q767 in bXOR) and E289M (E1261 in bXOR), are conserved among the molybdenum hydroxylases. E1261 serves as general base catalyst in accepting the proton from Mo-OH upon reaction in bXOR (11) (Fig. S3) and, accordingly, replacement of this residue in bacterial XOR by alanine profoundly compromises the reactivity of the enzyme (35). Additionally, NDH and bXOR have common residues in the active site like R319L (R880 in bXOR) and F353L (F914 in bXOR) (Fig. S3). R880 of bXOR was suggested to stabilize the developing negative charge on the substrate during the hydroxylation step, and its mutation results in an increase of the dissociation constant, KD, for substrate binding and a decrease in the rate constant of enzyme reduction, kred (36). The conservation of amino acids essential for catalysis indicates common ways of substrate binding and transition state stabilization in NDH and XORs. Based on these similarities (Fig. S3), we constructed a structural model for nicotinate binding in the active site of NDH in analogy to the structure of bXOR in complex with 2-hydroxy-6-methylpurine (27). It has recently been shown that 2-hydroxy-6-methylpurine binds between phenylalanine residues in the active site of bXOR, with the carbon atom to be hydroxylated in close proximity to the equatorial hydroxyl ligand. To allow binding of nicotinate in the active site of NDH the side chain of F353L has to rotate to assume a similar conformation as found for substrate/ligand-bound bXOR (21, 27) (Fig. S3). Modeling of substrate binding in analogy to bXOR produces a mechanistically reasonable complex. The nitrogen atom and carboxylate of nicotinate are at hydrogen-bonding distance to Y13M and R319M, respectively. C6, the carbon atom to be hydroxylated, is in a distance of ≈2 Å from the hydroxyl ligand and 2.5 Å from the Se ligand. This binding mode would facilitate the nucleophilic attack of the hydroxyl ligand on C6 and the concomitant hydride transfer from C6 to the Se ligand (Fig. S4).
S group to give Mo(+IV)-SH. Two residues in the direct environment of molybdenum, Q208L in NDH (Q767 in bXOR) and E289M (E1261 in bXOR), are conserved among the molybdenum hydroxylases. E1261 serves as general base catalyst in accepting the proton from Mo-OH upon reaction in bXOR (11) (Fig. S3) and, accordingly, replacement of this residue in bacterial XOR by alanine profoundly compromises the reactivity of the enzyme (35). Additionally, NDH and bXOR have common residues in the active site like R319L (R880 in bXOR) and F353L (F914 in bXOR) (Fig. S3). R880 of bXOR was suggested to stabilize the developing negative charge on the substrate during the hydroxylation step, and its mutation results in an increase of the dissociation constant, KD, for substrate binding and a decrease in the rate constant of enzyme reduction, kred (36). The conservation of amino acids essential for catalysis indicates common ways of substrate binding and transition state stabilization in NDH and XORs. Based on these similarities (Fig. S3), we constructed a structural model for nicotinate binding in the active site of NDH in analogy to the structure of bXOR in complex with 2-hydroxy-6-methylpurine (27). It has recently been shown that 2-hydroxy-6-methylpurine binds between phenylalanine residues in the active site of bXOR, with the carbon atom to be hydroxylated in close proximity to the equatorial hydroxyl ligand. To allow binding of nicotinate in the active site of NDH the side chain of F353L has to rotate to assume a similar conformation as found for substrate/ligand-bound bXOR (21, 27) (Fig. S3). Modeling of substrate binding in analogy to bXOR produces a mechanistically reasonable complex. The nitrogen atom and carboxylate of nicotinate are at hydrogen-bonding distance to Y13M and R319M, respectively. C6, the carbon atom to be hydroxylated, is in a distance of ≈2 Å from the hydroxyl ligand and 2.5 Å from the Se ligand. This binding mode would facilitate the nucleophilic attack of the hydroxyl ligand on C6 and the concomitant hydride transfer from C6 to the Se ligand (Fig. S4).
Computational Study Comparing Mo Se and Mo
Se and Mo S.
S.
The functional advantage of selenium over sulfur as a ligand is not immediately evident. The molybdenum sulfido ligand found in the active site of molybdenum hydroxylases like XOR (Mo S) plays an important role in the catalytic activity of these enzymes. Its replacement by a (second) Mo
S) plays an important role in the catalytic activity of these enzymes. Its replacement by a (second) Mo O group to give the so-called desulfo form of the enzyme leads to its complete inactivation. The hydride transfer taking place in the initial step of the reaction can be considered a nucleophilic attack of the hydride on the Mo
O group to give the so-called desulfo form of the enzyme leads to its complete inactivation. The hydride transfer taking place in the initial step of the reaction can be considered a nucleophilic attack of the hydride on the Mo E antibonding orbital (π*) (E = O, S). The stronger π interaction leads to higher π* anti-bonding energy, which raises the barrier of the reaction. The strength of the Mo
E antibonding orbital (π*) (E = O, S). The stronger π interaction leads to higher π* anti-bonding energy, which raises the barrier of the reaction. The strength of the Mo O π bond relative to that of Mo
O π bond relative to that of Mo S leads to a very high π* antibonding orbital and is presumably the basis for the lack of activity seen in the desulfo form of the enzymes (Fig. 3).
S leads to a very high π* antibonding orbital and is presumably the basis for the lack of activity seen in the desulfo form of the enzymes (Fig. 3).
Fig. 3.
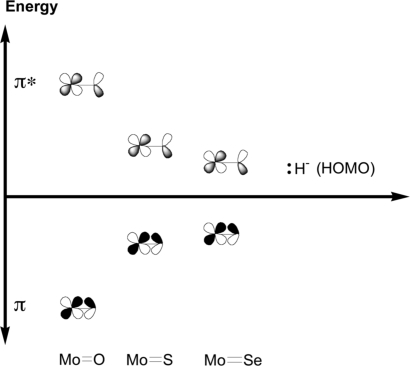
Molecular orbital view and the relative energy of Mo E π and π* (E = O, S, Se). Depicted are the bonding interactions between the dxy orbital of the molybdenum and the px orbital of the coordinated chalcogen in bonding (lower) and antibonding configurations, with the relative energies for Mo
E π and π* (E = O, S, Se). Depicted are the bonding interactions between the dxy orbital of the molybdenum and the px orbital of the coordinated chalcogen in bonding (lower) and antibonding configurations, with the relative energies for Mo O, Mo
O, Mo S, and Mo
S, and Mo Se species indicated.
Se species indicated.
A Mo Se rather than Mo
Se rather than Mo S is expected to yield an even weaker π bond with molybdenum than sulfur and oxygen (O > S > Se) (Fig. 3) and is thus expected to increase the reactivity of the enzyme in any mechanism involving hydride transfer. In the case of less reactive carbon centers, the increased reactivity may be important in catalyzing the hydroxylation. Computational studies have been carried out after the reaction of a molybdenum center such as that found in bXOR and NDH, but with an equatorial Mo
S is expected to yield an even weaker π bond with molybdenum than sulfur and oxygen (O > S > Se) (Fig. 3) and is thus expected to increase the reactivity of the enzyme in any mechanism involving hydride transfer. In the case of less reactive carbon centers, the increased reactivity may be important in catalyzing the hydroxylation. Computational studies have been carried out after the reaction of a molybdenum center such as that found in bXOR and NDH, but with an equatorial Mo Se rather than Mo
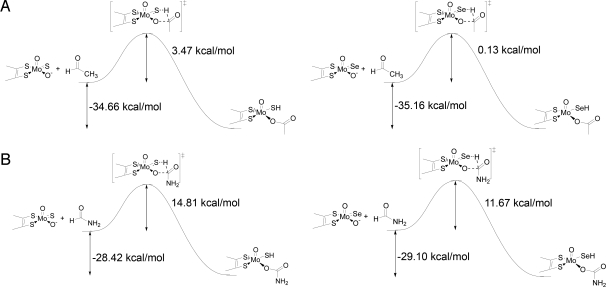
Se rather than Mo S group, with ethylaldehyde and formamide as simple substrates (Fig. 4 A and B, respectively). Ethylaldehyde and formamide were used as simple analogs for xanthine and nicotinate to reduce the necessary computational resources. Additionally, it is not necessary to include alternative protonation states with these 2 substrates. The calculations indicate that the transition state for the first step of the reaction is stabilized by 3.1–3.4 kcal/mol upon substitution of selenium for sulfur in the reaction with both ethylaldehyde (ΔH‡ is 3.47 kcal/mol for Mo
S group, with ethylaldehyde and formamide as simple substrates (Fig. 4 A and B, respectively). Ethylaldehyde and formamide were used as simple analogs for xanthine and nicotinate to reduce the necessary computational resources. Additionally, it is not necessary to include alternative protonation states with these 2 substrates. The calculations indicate that the transition state for the first step of the reaction is stabilized by 3.1–3.4 kcal/mol upon substitution of selenium for sulfur in the reaction with both ethylaldehyde (ΔH‡ is 3.47 kcal/mol for Mo S and 0.13 kcal/mol for Mo
S and 0.13 kcal/mol for Mo Se) and formamide (ΔH‡ is 14.81 kcal/mol for Mo
Se) and formamide (ΔH‡ is 14.81 kcal/mol for Mo S and 11.67 kcal/mol for Mo
S and 11.67 kcal/mol for Mo Se). This amounts to a factor of 200–300 in rate acceleration for this step.
Se). This amounts to a factor of 200–300 in rate acceleration for this step.
Fig. 4.
Calculated reaction coordinates. (A) Reaction of ethylaldehyde with the MoO(OH)S (Left) and the MoO(OH)Se analogue (Right). (B) Reaction of formamide with the MoO(OH)S (Left) and the MoO(OH)Se analogue (Right).
The geometry of the optimized Mo Se-containing structure is square pyramidal, with the Mo
Se-containing structure is square pyramidal, with the Mo E (E = S, Se), the hydroxyl group, and the enedithiolate ligand of the pyranopterin cofactor in the equatorial plane (Fig. S5). Table S1 gives the metric parameters for both Mo
E (E = S, Se), the hydroxyl group, and the enedithiolate ligand of the pyranopterin cofactor in the equatorial plane (Fig. S5). Table S1 gives the metric parameters for both Mo S and Mo
S and Mo Se structures. It can be seen that substitution of selenium for sulfur does not have a significant influence on the bond angles made by the ligands, although the Mo
Se structures. It can be seen that substitution of selenium for sulfur does not have a significant influence on the bond angles made by the ligands, although the Mo Se is longer than the Mo
Se is longer than the Mo S (2.31 vs. 2.18 Å), as expected given the larger selenium and weaker Mo
S (2.31 vs. 2.18 Å), as expected given the larger selenium and weaker Mo Se bond. The calculated geometry of the selenium-containing model is very similar to what has been found in the structure of NDH and both show a Mo-Se distance of ≈2.3 Å.
Se bond. The calculated geometry of the selenium-containing model is very similar to what has been found in the structure of NDH and both show a Mo-Se distance of ≈2.3 Å.
Discussion
The nature of the selenium moiety of NDH has been investigated over the last decade by using various approaches. After the discovery that selenium is essential for hydroxylase activity (13), Dilworth (14) showed that selenium is a component of NDH, which could be released by heat treatment or the addition of chaotrophic agents like urea. That selenium in NDH is not part of a stable organic molecule but that is instead present as a neutral selenol or an inorganic selenide has been indicated by the liberation of selenium from NDH by incubation with alkylating agents, resulting in the formation of dialkylselenides (14). The approximate ratio of 1 mol of selenium per mol of NDH was established by EPR studies of NDH with the 77Se isotope that showed the nuclear spin of 77Se couples to the Mo(V) electron spin, suggesting that the selenium could be a ligand to molybdenum (18). This left the possibilities that selenium replaced the sulfido ligand at the molybdenum or that it could be weakly bound to a heteroatom of a separate cofactor adjacent to molybdenum. The crystal structure of NDH presented here shows that selenium indeed replaces sulfur as a molybdenum ligand, present as Mo Se. The refined bond length of 2.3 Å agrees well with the expected bond length for such a terminal selenido ligand and specifically is too short for a selenol ligand (37). No further stabilization of the selenium by other interactions is observed, nor do we find residues such as cysteines in the vicinity of the Mo ion, which might form a bond to the selenido ligand. Access to the selenido ligand is partly blocked by F353L (Fig. 2A), which could explain why incubation of active NDH with potassium cyanide does not lead to a rapid inactivation of the enzyme and inactive enzyme could not be reactivated by the addition of sodium selenide or selenophosphate (18, 34).
Se. The refined bond length of 2.3 Å agrees well with the expected bond length for such a terminal selenido ligand and specifically is too short for a selenol ligand (37). No further stabilization of the selenium by other interactions is observed, nor do we find residues such as cysteines in the vicinity of the Mo ion, which might form a bond to the selenido ligand. Access to the selenido ligand is partly blocked by F353L (Fig. 2A), which could explain why incubation of active NDH with potassium cyanide does not lead to a rapid inactivation of the enzyme and inactive enzyme could not be reactivated by the addition of sodium selenide or selenophosphate (18, 34).
Whether selenium is also a ligand to molybdenum in other selenium-dependent molybdenum hydroxylases remains to be established in future structural studies. For the selenium-containing XOR from E. barkeri it has been shown that the enzyme can be inactivated with potassium cyanide and enzyme thus inactivated can be reactivated by incubation with selenide under reducing conditions (17), observations that could be explained by the presence of a selenido ligand bound to molybdenum such as is seen here with NDH. However, with the purine hydroxylase from C. purinolyticum no magnetic interaction between the nuclear spin of 77Se and the Mo(V) electron spin could be detected, raising the possibility that the labile selenium is not coordinated to Mo in all forms of this enzyme (38). The catalytic advantage of the incorporation of selenium in a molybdenum hydroxylase is evident from comparing XORs of different organisms. While for bXORs with a sulfido ligand at the molybdenum turnover number of 2–25 s−1 has been reported (39), the selenium containing XOR from E. barkeri achieves turnover rates >400 s−1 (17). If the XOR from E. barkeri has a selenido ligand like NDH, the higher rates are likely caused by an acceleration of the hydride transfer step, as suggested by our model calculations.
For the molybdenum hydroxylase family we now see that at least 3 variations for the common theme MoO(OH)X coordination sphere exist, where X is: (i) S for XORs (40), quinoline oxidoreductase (24), and 4-hydroxybenzoyl-CoA reductase (23); (ii) SCu for carbon monoxide dehydrogenase (28); and, as detailed above (iii) Se for NDH. All active sites can be converted to an inactive state in which X is O, to give a second Mo O ligand. These variations demonstrate how nature adapts a protein bound ligand-metal motif for different substrates and reactivities by the exchange of 1 ligand and indicate a flexible biological chemistry of molybdenum enzymes by ligand tuning.
O ligand. These variations demonstrate how nature adapts a protein bound ligand-metal motif for different substrates and reactivities by the exchange of 1 ligand and indicate a flexible biological chemistry of molybdenum enzymes by ligand tuning.
Materials and Methods
Purification of NDH.
As NDH is instable and looses activity with time (34) our experimental strategy was to take <1 week from breaking the E. barkeri cells used to purify the protein until freezing the protein crystals used for structure determination. For all steps, including crystallization of NDH, buffer conditions were used for which NDH was reported to be most stable and active (34). Frozen E. barkeri cells were resuspended in 50 mM Tris·HCl (pH 7.8), 10 mM NaCl, 1 mM EDTA, and 2 mM DTT (buffer A) and broken by 5 cycles of sonication. After centrifugation for 30 min the cell-free extract was loaded on a Source 30Q column. NDH was eluted from the anionic exchange column by a linear gradient of buffer A containing 0.5 M NaCl. Pooled fractions were loaded onto a hydroxyapatite column without preceding buffer exchange. Active NDH eluted with 120 mM KPO4, pH 7.3. Active fractions were collected, rebuffered in 50 mM Tris·HCl (pH 7.8), 0.2 M KCl, 1 mM EDTA, and 2 mM DTT and concentrated to 20 mg/mL. The protein was stored at +4 °C and used within 5 days. Approximately 1.5 mg NDH could be obtained from 10 g of cells. To obtain protein with higher purity, an additional gelfiltration step (Superdex 200) was performed in 50 mM Tris·HCl (pH 7.8), 0.2 M KCl, 1 mM EDTA, and 2 mM DTT. All purification steps were carried out under anoxic conditions in a glove box containing 95% N2 and 5% H2.
Activity Measurement.
Enzyme assays were conducted according to Gladyshev et al. (34) with some modifications. Hydroxylase activity was measured under anoxic conditions in 100 mM KPO4 (pH 7.0), 50 mM nicotinate (pH 7.5), 1 mM NADP+, and 5 mM DTT. The reaction was started by addition of NADP+ and followed by absorption increase at 340 nm.
Crystallographic Methods.
NDH was crystallized by the vapor diffusion method in a hanging drop. The enzyme preparations used for crystallization had specific activities of 11–20 units·mg−1. The drop contained a 1:1 mixture of protein solution (10 mg/mL−1) with reservoir solution containing 18–20% PEG 3350, 0.1 M Tris·HCl (pH 7.5), 75 mM NaNO3, 5% glycerol, or 1% 2,4-methylpentandiol. Crystals were harvested in the corresponding soaking solutions containing 15% (vol/vol) (2R,3R)-butanediol (Sigma), shock-frozen, and stored in liquid nitrogen. Diffraction data were collected at −180 °C on a rotating anode X-ray generator (Nonius FR591; Bruker) equipped with an image plate detector (mar345dtb; Marresearch) (native, Table 1) and at the BM-14, European Synchrotron Radiation Facility, Grenoble (above and below Se-edge, Table 1). The protein structure model was build with Coot (41) and MAIN 2000 (42). Positional and temperature refinement was carried out with CNS (43). Anomalous difference Fourier maps were calculated by using CNS (43). The final refinement statistics and stereochemical analyses using PROCHECK (44) are shown in Table 1. Ramachandran statistics revealed 86% in the most favored, 13% in the additionally allowed, 1% in the generously allowed, and 0% in the disallowed regions. Simulated annealing omit maps were used to validate the active-site structure reported.
Model Calculations.
Calculations were carried out by using hybrid density functional theory (B3LYP) as implemented in Gaussian 03. The structures were fully optimized and confirmed as minima or transition state by calculating the vibrational frequency. The structures were optimized with B3LYP using LANL2DZ ECP (45, 46) augmented with f polarization functions (47) for Mo and 6–31G(d) for the rest of atoms. For the hydrogen that performs the hydride transfer 6–31++G(d,p) was used. The zero point energies were calculated with same basis set and level of theory. All quoted energies are corrected to zero point energy and are without temperature correction.
Supplementary Material
Acknowledgments.
H.D. was supported by Deutsche Forschungsgemeinschaft Grant DO 785/2–2 and the Fonds der Chemischen Industrie. R.H. and H.D. received travel funds from the Bavaria California Technology Center to initiate the collaboration.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: The atomic coordinates have been deposited in the Protein Data Bank, www.pdb.org (PDB ID codes 3HRD).
This article contains supporting information online at www.pnas.org/cgi/content/full/0902210106/DCSupplemental.
References
- 1.Böck A, et al. Selenocysteine: The 21st amino acid. Mol Microbiol. 1991;5:515–520. doi: 10.1111/j.1365-2958.1991.tb00722.x. [DOI] [PubMed] [Google Scholar]
- 2.Stadtman TC. Selenocysteine. Annu Rev Biochem. 1996;65:83–100. doi: 10.1146/annurev.bi.65.070196.000503. [DOI] [PubMed] [Google Scholar]
- 3.Stadtman TC. Selenium biochemistry. Annu Rev Biochem. 1990;59:111–127. doi: 10.1146/annurev.bi.59.070190.000551. [DOI] [PubMed] [Google Scholar]
- 4.Wessjohann LA, Schneider A, Abbas M, Brandt W. Selenium in chemistry and biochemistry in comparison to sulfur. Biol Chem. 2007;388:997–1006. doi: 10.1515/BC.2007.138. [DOI] [PubMed] [Google Scholar]
- 5.Gladyshev VN. Comparison of selenium-containing molybdoenzymes. Molybdenum and tungsten: Their roles in biological processes. Metal Ions Biol Syst. 2002;39:655–672. [PubMed] [Google Scholar]
- 6.Zhang Y, Gladyshev VN. Molybdoproteomes and evolution of molybdenum utilization. J Mol Biol. 2008;379:881–899. doi: 10.1016/j.jmb.2008.03.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gladyshev VN, Khangulov SV, Axley MJ, Stadtman TC. Coordination of selenium to molybdenum in formate dehydrogenase H from Escherichia coli. Proc Natl Acad Sci USA. 1994;91:7708–7711. doi: 10.1073/pnas.91.16.7708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Boyington JC, et al. Crystal structure of formate dehydrogenase H: Catalysis involving Mo, molybdopterin, selenocysteine, and an Fe4S4 cluster. Science. 1997;275:1305–1308. doi: 10.1126/science.275.5304.1305. [DOI] [PubMed] [Google Scholar]
- 9.Raaijmakers HC, Romao MJ. Formate-reduced Escherichia coli formate dehydrogenase H: The reinterpretation of the crystal structure suggests a new reaction mechanism. J Biol Inorg Chem. 2006;11:849–854. doi: 10.1007/s00775-006-0129-2. [DOI] [PubMed] [Google Scholar]
- 10.Hille R. The mononuclear molybdenum enzymes. Chem Rev. 1996;96:2757–2816. doi: 10.1021/cr950061t. [DOI] [PubMed] [Google Scholar]
- 11.Hille R. Molybdenum-containing hydroxylases. Arch Biochem Biophys. 2005;433:107–116. doi: 10.1016/j.abb.2004.08.012. [DOI] [PubMed] [Google Scholar]
- 12.Holcenberg JS, Stadtman ER. Nicotinic acid metabolism. 3. Purification and properties of a nicotinic acid hydroxylase. J Biol Chem. 1969;244:1194–1203. [PubMed] [Google Scholar]
- 13.Imhoff D, Andreesen JR. Nicotinic acid hydroxylase from Clostridium barkeri: Selenium-dependent formation of active enzyme. FEMS Microbiol Lett. 1979;5:155–158. [Google Scholar]
- 14.Dilworth GL. Properties of the selenium-containing moiety of nicotinic acid hydroxylase from Clostridium barkeri. Arch Biochem Biophys. 1982;219:30–38. doi: 10.1016/0003-9861(82)90130-8. [DOI] [PubMed] [Google Scholar]
- 15.Self WT, Stadtman TC. Selenium-dependent metabolism of purines: A selenium-dependent purine hydroxylase and xanthine dehydrogenase were purified from Clostridium purinolyticum and characterized. Proc Natl Acad Sci USA. 2000;97:7208–7213. doi: 10.1073/pnas.97.13.7208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wagner R, Cammack R, Andreesen JR. Purification and characterization of xanthine dehydrogenase from Clostridium acidiurici grown in the presence of selenium. Biochim Biophys Acta. 1984;791:63–74. [Google Scholar]
- 17.Schrader T, Rienhofer A, Andreesen JR. Selenium-containing xanthine dehydrogenase from Eubacterium barkeri. Eur J Biochem. 1999;264:862–871. doi: 10.1046/j.1432-1327.1999.00678.x. [DOI] [PubMed] [Google Scholar]
- 18.Gladyshev VN, Khangulov SV, Stadtman TC. Nicotinic acid hydroxylase from Clostridium barkeri: Electron paramagnetic resonance studies show that selenium is coordinated with molybdenum in the catalytically active selenium-dependent enzyme. Proc Natl Acad Sci USA. 1994;91:232–236. doi: 10.1073/pnas.91.1.232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Romao MJ, et al. Crystal structure of the xanthine oxidase-related aldehyde oxido-reductase from Desulfovibrio gigas. Science. 1995;270:1170–1176. doi: 10.1126/science.270.5239.1170. [DOI] [PubMed] [Google Scholar]
- 20.Rebelo J, et al. Gene sequence and crystal structure of the aldehyde oxidoreductase from Desulfovibrio desulfuricans ATCC 27774. J Mol Biol. 2000;297:135–146. doi: 10.1006/jmbi.2000.3552. [DOI] [PubMed] [Google Scholar]
- 21.Enroth C, et al. Crystal structures of bovine milk xanthine dehydrogenase and xanthine oxidase: Structure-based mechanism of conversion. Proc Natl Acad Sci USA. 2000;97:10723–10728. doi: 10.1073/pnas.97.20.10723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Truglio JJ, et al. Crystal structures of the active and alloxanthine-inhibited forms of xanthine dehydrogenase from Rhodobacter capsulatus. Structure (London) 2002;10:115–125. doi: 10.1016/s0969-2126(01)00697-9. [DOI] [PubMed] [Google Scholar]
- 23.Unciuleac M, et al. Structure of a xanthine oxidase-related 4-hydroxybenzoyl-CoA reductase with an additional [4Fe-4S] cluster and an inverted electron flow. Structure (London) 2004;12:2249–2256. doi: 10.1016/j.str.2004.10.008. [DOI] [PubMed] [Google Scholar]
- 24.Bonin I, et al. Active site geometry and substrate recognition of the molybdenum hydroxylase quinoline 2-oxidoreductase. Structure (London) 2004;12:1425–1435. doi: 10.1016/j.str.2004.05.014. [DOI] [PubMed] [Google Scholar]
- 25.Dobbek H, Gremer L, Meyer O, Huber R. Crystal structure and mechanism of CO dehydrogenase, a molybdo iron-sulfur flavoprotein containing S-selanylcysteine. Proc Natl Acad Sci USA. 1999;96:8884–8889. doi: 10.1073/pnas.96.16.8884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hänzelmann P, et al. The effect of intracellular molybdenum in Hydrogenophaga pseudoflava on the crystallographic structure of the seleno-molybdo-iron-sulfur flavoenzyme carbon monoxide dehydrogenase. J Mol Biol. 2000;301:1221–1235. doi: 10.1006/jmbi.2000.4023. [DOI] [PubMed] [Google Scholar]
- 27.Pauff JM, Zhang J, Bell CE, Hille R. Substrate orientation in xanthine oxidase: Crystal structure of enzyme in reaction with 2-hydroxy-6-methylpurine. J Biol Chem. 2008;283:4818–4824. doi: 10.1074/jbc.M707918200. [DOI] [PubMed] [Google Scholar]
-
28.Dobbek H, et al. Catalysis at a dinuclear [CuSMo(

 O)OH] cluster in a CO dehydrogenase resolved at 1.1-Å resolution. Proc Natl Acad Sci USA. 2002;99:15971–15976. doi: 10.1073/pnas.212640899. [DOI] [PMC free article] [PubMed] [Google Scholar]
O)OH] cluster in a CO dehydrogenase resolved at 1.1-Å resolution. Proc Natl Acad Sci USA. 2002;99:15971–15976. doi: 10.1073/pnas.212640899. [DOI] [PMC free article] [PubMed] [Google Scholar] - 29.Gladyshev VN, Lecchi P. Identification of molybdopterins in molybdenum- and selenium-containing enzymes. Biofactors. 1995;5:93–97. [PubMed] [Google Scholar]
- 30.Alhapel A, et al. Molecular and functional analysis of nicotinate catabolism in Eubacterium barkeri. Proc Natl Acad Sci USA. 2006;103:12341–12346. doi: 10.1073/pnas.0601635103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Caldeira J, et al. Analysis of the electron paramagnetic resonance properties of the [2Fe-2S]1+ centers in molybdenum enzymes of the xanthine oxidase family: Assignment of signals I and II. Biochemistry. 2000;39:2700–2707. doi: 10.1021/bi9921485. [DOI] [PubMed] [Google Scholar]
- 32.Canne C, et al. Comparative EPR and redox studies of three prokaryotic enzymes of the xanthine oxidase family: Quinoline 2-oxidoreductase, quinaldine 4-oxidase, and isoquinoline 1-oxidoreductase. Biochemistry. 1997;36:9780–9790. doi: 10.1021/bi970581d. [DOI] [PubMed] [Google Scholar]
- 33.Gremer L, et al. Binding of flavin adenine dinucleotide to molybdenum-containing carbon monoxide dehydrogenase from Oligotropha carboxidovorans. J Biol Chem. 2000;275:1864–1872. doi: 10.1074/jbc.275.3.1864. [DOI] [PubMed] [Google Scholar]
- 34.Gladyshev VN, Khangulov SV, Stadtman TC. Properties of the selenium- and molybdenum-containing nicotinic acid hydroxylase from Clostridium barkeri. Biochemistry. 1996;35:212–223. doi: 10.1021/bi951793i. [DOI] [PubMed] [Google Scholar]
- 35.Leimkühler S, et al. The role of active site glutamate residues in catalysis of Rhodobacter capsulatus xanthine dehydrogenase. J Biol Chem. 2004;279:40437–40444. doi: 10.1074/jbc.M405778200. [DOI] [PubMed] [Google Scholar]
- 36.Pauff JM, et al. The role of arginine 310 in catalysis and substrate specificity in xanthine dehydrogenase from Rhodobacter capsulatus. J Biol Chem. 2007;282:12785–12790. doi: 10.1074/jbc.M700364200. [DOI] [PubMed] [Google Scholar]
- 37.Ma X, Schulzke C, Schmidt HG, Noltemeyer M. Structural, electrochemical, and oxygen atom transfer properties of a molybdenum selenoether complex [Mo2O4(OC3H6SeC3H6O)2] and its thioether analogue [Mo2O4(OC3H6SC3H6O)2] Dalton Trans. 2007;2007:1773–1780. doi: 10.1039/b617652f. [DOI] [PubMed] [Google Scholar]
- 38.Self WT, Wolfe MD, Stadtman TC. Cofactor determination and spectroscopic characterization of the selenium-dependent purine hydroxylase from Clostridium purinolyticum. Biochemistry. 2003;42:11382–11390. doi: 10.1021/bi030136k. [DOI] [PubMed] [Google Scholar]
- 39.Pai EF, Nishino T. The molybdenum-containing xanthine oxidoreductases and picolinate dehydrogenases. Molybdenum and tungsten: Their roles in biological processes. Metal Ions Biol Syst. 2002;39:431–454. [PubMed] [Google Scholar]
- 40.Kuwabara Y, et al. Unique amino acids cluster for switching from the dehydrogenase to oxidase form of xanthine oxidoreductase. Proc Natl Acad Sci USA. 2003;100:8170–8175. doi: 10.1073/pnas.1431485100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Emsley P, Cowtan K. COOT: Model-building tools for molecular graphics. Acta Crystallogr D. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 42.Turk D. Münich: Technische Universität; 1992. PhD thesis. [Google Scholar]
- 43.Brünger AT, et al. Crystallography and NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D. 1998;54:905–921. doi: 10.1107/s0907444998003254. [DOI] [PubMed] [Google Scholar]
- 44.Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: A program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26:283–291. [Google Scholar]
- 45.Hay PJ, Wadt WR. Ab initio effective core potentials for molecular calculations: Potentials for the transition-metal atoms Sc to Hg. J Chem Phys. 1985;82:270–283. [Google Scholar]
- 46.Hay PJ, Wadt WR. Ab initio effective core potentials for molecular calculations: Potentials for K to Au including the outermost core orbitals. J Chem Phys. 1985;82:299–310. [Google Scholar]
- 47.Ehlers AW, et al. A set of F-polarization functions for pseudo-potential basis sets of the transition metals Sc-Cu, Y-Ag and La-Au. Chem Phys Lett. 1993;208:111–114. [Google Scholar]
- 48.DeLano WL. PyMol. San Carlos, CA: DeLano Scientific; 2002. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.