Abstract
SET domain proteins are present in chromosomal proteins involved in epigenetic control of transcription. The yeast SET domain protein Set1p regulates chromatin structure, DNA repair, and telomeric functions. We investigated the mechanism by which the absence of Set1p increases DNA repair capacities of checkpoint mutants. We show that deletion of SET1 induces a response relayed by the signaling kinase Rad53p that leads to the MEC1/TEL1-independent hyperphosphorylation of replication protein A middle subunit (Rfa2p). Consequently, the binding of Rfa2p to upstream repressing sequences (URS) of repair genes is decreased, thereby leading to their derepression. Our results correlate the set1Δ-dependent phosphorylation of Rfa2p with the transcriptional induction of repair genes. Moreover, we show that the deletion of the amino-terminal region of Rfa2p suppresses the sensitivity to ultraviolet radiation of a mec3Δ checkpoint mutant, abolishes the URS-mediated repression, and increases the expression of repair genes. This work provides an additional link for the role of Rfa2p in the regulation of the repair capacity of the cell and reveals a role for the phosphorylation of Rfa2p and unveils unsuspected connections between chromatin, signaling pathways, telomeres, and DNA repair.
Keywords: SET domain, RPA, phosphorylation, checkpoint, DNA repair, telomere, gene silencing
Several chromatin regulators involved in epigenetic control of transcription contain a characteristic motif of unknown function called the SET domain that takes its name from Su(v)ar 3–9, E(z) and trithorax (for review, see Jenuwein et al. 1998). The SET domain has been now identified in >40 proteins from fungi, plant, and mammalian species.
The 1081-residue Set1p protein of Saccharomyces cerevisiae contains a carboxy-terminal SET domain of 130 residues. Inactivation of the full-length Set1p or of its SET domain alleviates telomeric position effect (TPE) (Nislow et al. 1997; Corda et al. 1999). The SET domain of Set1p interacts with the carboxy-terminal region of the checkpoint protein Mec3p, which is required for efficient DNA damage-dependent checkpoints at G1/S, intra S, and G2/M (Longhese et al. 1996; Corda et al. 1999). Mec3p belongs to the sensor class of the DNA checkpoint machinery, which along with Mec3p includes Rad9p, Rad17p, Rad24p, and Ddc1p (for review, see Lowndes and Murguia 2000). The sensor proteins are thought to recognize DNA damage and generate a signal relayed by the central transducer Mec1p to the protein kinases Rad53p and Dun1p and the metaphase–anaphase regulator Pds1p (Allen et al. 1994; Sanchez et al. 1996; Sun et al. 1996; Paciotti et al. 1998; Gardner et al. 1999; Pellicoli et al. 1999). The checkpoint genes required for cell cycle delay also regulate the transcriptional induction of DNA repair and DNA metabolism genes, thereby increasing the DNA repair capacities of the cell (Aboussekhra et al. 1996; Huang et al. 1998). We have shown previously that the deletion of SET1 increases the viability after DNA damage of a mec3 null mutant and also of other checkpoint mutants (rad9, rad17, and rad24), but not of cells devoid of the signaling kinase Rad53p. SET1 deletion does not restore any of the three DNA damage checkpoints regulated by Mec3p. Finally, Set1p and Mec3p were found to modulate in opposite directions TPE and telomere length (Corda et al. 1999).
Replication protein A (RPA) is an eukaryotic single-stranded DNA-binding protein essential for replication, recombination, and DNA repair (Wold 1997). Human as well as yeast RPA is composed of three subunits of 70, 32/34, and 14 kD (Brill and Stillman 1991; Philipova et al. 1996). It has been proposed that RPA plays a role in DNA damage sensing mainly on the basis of the fact that RPA 32/34 (in yeast Rfa2p) becomes phosphorylated in response to DNA damage and in the presence of ssDNA (for review, see Iftode et al. 1999). Brush et al. (1996) showed that the yeast checkpoint Mec1p, but not Rad53p, was required for the phosphorylation of Rfa2p during normal cell cycle and after DNA damage. It has been shown recently that Rfa1p is also phosphorylated in response to various forms of genotoxic stress, including radiation and hydroxyurea exposure (Brush and Kelly 2000). Interestingly, phosphorylation of Rfa1p after radiation depends on both checkpoint regulators, Mec1p and Rad53p (Brush and Kelly 2000). RPA has been shown to bind specifically to double-stranded oligonucleotides containing upstream repressing sequences (URS) found upstream of several DNA repair genes (Singh and Samson 1995). It has been hypothesized from these results that RPA may be involved in the transcriptional regulation of repair genes. Finally, it has been shown that in vivo hyperphosphorylation of RPA 32/34 after irradiation was concordant with a decreased ability of RPA to bind ssDNA in mouse cell extracts (Fried et al. 1996).
In this study, we document the set1Δ-induced cellular response. We show that inactivation of Set1p leads to the constitutive MEC1/TEL1-independent hyperphosphorylation of Rfa2p in a process that requires Rad53p. The Set1-dependent phosphorylation of Rfa2p leads to the transcriptional activation of repair genes but not to a cell cycle arrest. This pathway explains why Set1p inactivation increases the repair capacities of the cell when checkpoints fail. This response differs from the Mec1-dependent DNA damage induction pathway.
Results
Rad53-dependent hyperphosphorylation of Rfa2p in set1Δ mutants
Phosphorylation of Rfa2p appears during the G1/S transition and persists through the S phase and in response to genotoxic insult (Din et al. 1990; Brush et al. 1996). Because the deletion of SET1 results in decrease of sensitivity to ionizing radiation in checkpoint mutants, we asked whether the set1Δ mutation was associated with the phosphorylation of Rfa2p. Rfa2p phosphorylation was determined by immunoblot analysis with anti-Rfa2p polyclonal antibodies (Din et al. 1990) of protein extracts prepared from synchronized cells that have been released from α-factor at the indicated times (Fig. 1). Phosphorylated forms of Rfa2p exhibited reduced mobility in SDS/PAGE (Fig. 1A). The two bands that appear in Figure 1A were converted into a single species upon treatment of crude extracts with phosphatase, indicating that the reduced mobility of Rfa2p was due to phosphorylation (data not shown). Flow cytometry analysis was carried out in parallel to monitor cell progression through S phase (Fig. 1A). As expected, cell cycle-regulated Rfa2p phosphorylation was compromised in sml1 mec1-1 cells (Brush et al. 1996) (see Materials and Methods for the sml1 mec1-1 allele). In contrast, in set1Δ cells, the level of Rfa2p phosphorylation was enhanced. The ratio of phosphorylated to unphosphorylated Rfa2p increased as compared with wild type in set1Δ exponentially growing cells. Interestingly, in exponential growing sml1 mec1-1 cells, the absence of SET1 maintained a high level of Rfa2p phosphorylation; >80% of Rfa2p was found under its phosphorylated form in exponentially growing cells and there was only a slight increase of Rfa2p phosphorylation during S phase. We concluded that Rfa2p is hyperphosphorylated in set1Δ cells in a manner independent of MEC1.
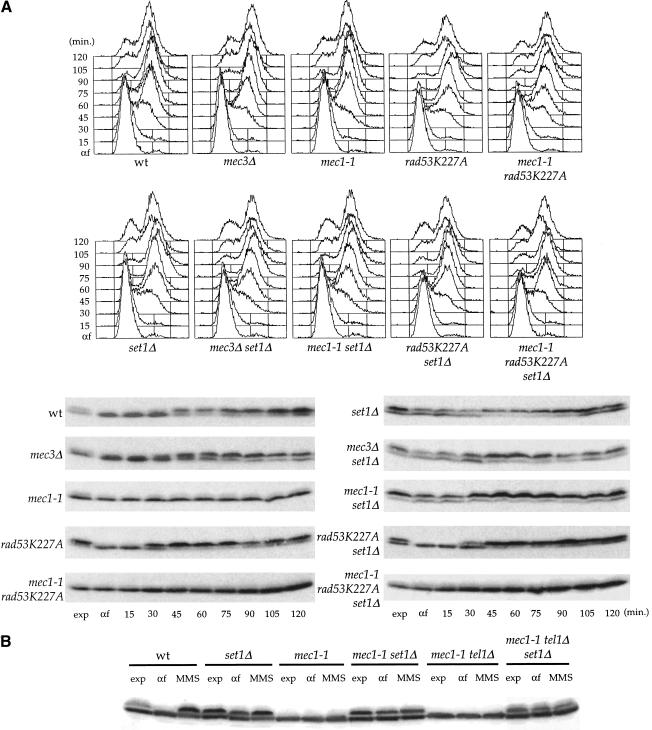
Figure 1.
set1Δ-induced hyperphosphorylation of Rfa2p. (A, top) Exponentially growing cells (exp) of K699 derivative strains with the indicated relevant genotypes were synchronized with α-factor and released at time zero. FACS analysis of the synchronized cultures at the indicated times after α-factor release. (Bottom) Protein extracts prepared at the indicated times after release from α-factor were analyzed by SDS-PAGE and immunoblotting with anti-pRfa2p polyclonal antibodies. The upper band corresponds to the phosphorylated form of Rfa2p. (B) set1Δ-dependent phosphorylation of Rfa2p after MMS treatment in sml1 mec1-1 and sml1 mec1-1 tel1 mutants. Protein extracts from exponentially growing cultures (exp), from G1-arrested cultures (α-F), and MMS treated G1-arrested cultures (MMS), were analyzed by Western blot with anti-pRfa2p polyclonal antibodies.
Noteworthy, we found that the deletion of MEC3 did not modify the phosphorylation pattern of Rfa2p in wild-type and in set1Δ cells.
To identify a kinase involved in the phosphorylation of Rfa2p in set1Δcells, we analyzed the phosphorylation of Rfa2p in rad53 mutants. We introduced in our mutant strains a rad53 allele (rad53K227A) carrying a substitution within the conserved kinase domain of Rad53p (Fay et al. 1997; Pellicioli et al. 1999). We found that inactivation of the kinase activity of Rad53p did not alter the phosphorylation of Rfa2p in wild-type cells. In contrast, introduction of the rad53K227A allele in set1Δ cells abolished the set1Δ-dependent hyperphosphorylation of Rfa2p (Fig. 1A). This observation was confirmed by the analysis of Rfa2p phosphorylation in sml1 mec1-1 rad53K227A and sml1 mec1-1 rad53K227A set1Δ mutants. We failed to detect any significant phosphorylation of Rfa2p when the rad53K227A allele was introduced in sml1 mec1-1 and sml1 mec1-1 set1Δ mutants.
We observed that G1 cells, which normally do not exhibit phosphorylation of Rfa2p, showed such a modification in set1Δ cells. This set1Δ-dependent extra phosphorylation of Rfa2p, which seems to be cell cycle independent, disappeared in set1Δ rad53K227A cells, whereas the cell cycle-dependent phosphorylation of Rfa2p, which is apparently Mec1p dependent, remained unaffected (Fig. 1A).
We concluded that Rfa2p is hyperphosphorylated in set1Δ cells by both Rad53p- and Mec1p-dependent mechanisms. The Rad53p-dependent phosphorylation of Rfa2p, evidenced in set1Δ cells, seems to be cell cycle independent.
Hyperphosphorylation of Rfa2p in set1Δ mutants occurs in the absence of both ATM homologs, Mec1p and Tel1p
The response resulting from the inactivation of SET1 appears different from the one resulting from DNA damage. We therefore asked whether the high level of phosphorylation of Rfa2p in set1Δ cells can be further increased after DNA damage (Brush et al. 1996). As overexpression of TEL1 can restore some responses in checkpoint mutant cells, we also asked whether hyperphosphorylation of Rfa2p in set1Δ mutants could occur in the absence of both ATM homologs, Mec1p and Tel1p. To distinguish DNA damage-induced Rfa2p phosphorylation from the cell cycle-regulated reaction, we blocked cells in G1 with α-factor and compared methyl methane sulfonate (MMS)-treated and untreated cells. Phosphorylation of Rfa2p was analyzed 30 min after MMS addition. The level of Rfa2p phosphorylation was increased by roughly the same level in wild-type and set1Δ cells (Fig. 1B). This result indicates that the level of phosphorylation of Rfa2p in set1Δ cells is not maximal and can still be increased by DNA damage. We then investigated whether this DNA damage-induced phosphorylation in set1Δ cells was depending on both ATM homologs MEC1 and TEL1. In a sml1 mec1-1 mutant, DNA damage-induced phosphorylation of Rfa2p was weak and totally absent in a sml1 mec1-1 tel1 mutant in agreement with the study of Brush et al. (1996). As above, we observed that Rfa2p was hyperphosphorylated in set1Δ cells even in the absence of MEC1 and TEL1. However, no significant increase in Rfa2p phosphorylation after MMS treatment was detected in sml1 mec1-1 set1Δ and sml1 mec1-1 tel1 set1Δ (Fig. 1B). Therefore, the MMS-induced Rfa2p phosphorylation is impaired in the absence of MEC1 and TEL1 in set1Δ cells. Our findings indicate that Rfa2p phosphorylation occurs in the absence of MEC1 and TEL1 in set1Δ cells and can still be increased by DNA damage in set1Δ cells in the presence of MEC1 and TEL1.
set1Δ-dependent phosphorylation of Rfa2p requires the amino terminus of Rfa2p
To further characterize the Set1-depending Rfa2p phosphorylation, we expressed a chromosomally encoded truncated form of Rfa2p, lacking 38 amino acids from the amino terminus (Rfa2p Δ40). In the human counterpart of the yeast Rfa2p, this region has been shown to contain the major phosphorylation sites (Niu et al. 1997; Zernik-Kobak 1997). In yeast, the rfa2 Δ40 allele corresponds to the largest viable amino-terminal truncation of Rfa2p (Philipova et al. 1996).
Interestingly, the yeast-made Rfa2p Δ40 protein is phosphorylated, indicating the presence of phosphorylation sites outside of the amino-terminal region (Fig. 2A). When we treated native extracts from the exponentially growing cells producing Rfa2p Δ40 with phosphatase, we observed that the upper band was converted to the lower one, indicating that the reduced mobility of Rfa2p Δ40 is due to phosphorylation (Fig. 2B). We also observed that Rfa2p Δ40 was neither phosphorylated in sml1 mec1-1 nor in sml1 mec1-1set1Δ mutants (Fig. 2A). This latter result suggests that the Mec1-dependent phosphorylation of Rfa2p may occur outside of the conserved amino-terminal region of Rfa2p, whereas the set1Δ-dependent phosphorylation of Rfa2p, revealed in set1Δ cells appears to depend on the presence of the amino-terminal region of Rfa2p.
Figure 2.
Phosphorylation of Rfa2p Δ40. (A) Exponentially growing cells of K699 derivative strains with the indicated relevant genotypes were synchronized with α-factor and released at time zero. Protein extracts prepared at the indicated times after release from α-factor were analyzed by Western blot with anti-pRfa2p polyclonal antibodies. The upper band corresponds to the phosphorylated form of Rfa2p Δ40. (B) Native extracts of exponentially growing rfa2 Δ40 and rfa2 Δ40 set1Δ. Cells were divided in two samples and treated (+) or not (−) with λ-phosphatase.
The set1Δ mutation does not induce a DNA damage checkpoint signal
We report above that the deletion of SET1 induces the constitutive phosphorylation of Rfa2p. Because phosphorylation of Rfa2p occurs after DNA damage, we investigated whether the set1Δ mutation induces a DNA damage checkpoint signal. Thus, we examined the phosphorylation of proteins involved in DNA damage checkpoints in set1Δ cells. As Rfa2p, the checkpoint protein Ddc1p is phosphorylated periodically during the cell cycle and in response to DNA damage (Longhese et al. 1997). Moreover, Ddc1p phosphorylation during an unpertubated cell cycle and after DNA damage depends on Mec1p (Paciotti et al. 1998). Ddc1p phosphorylation was analyzed under the same conditions as for Rfa2p phosphorylation in Figure 1A. We found that in set1Δ cells, Ddc1p is phosphorylated during S phase and its phosphorylation is very similar to the one of wild-type cells (data not shown). We also observed by immunoblot analysis that the effector kinase Rad53p was apparently not phosphorylated in set1Δ cells (data not shown). We concluded that set1Δ does not induce a constitutive hyperphosphorylation of checkpoint proteins.
set1Δ-dependent hyperphosphorylation of Rfa2p correlates with the enhancing of viability of checkpoint mutants after DNA damage
We have reported previously that the deletion of SET1 increases the viability of rad9, rad17, rad24, and mec3 mutants but not the viability of cells carrying the sad1-1 allele of RAD53 (Corda et al. 1999). In agreement with our previous work, we also observed that the set1Δ mutation did not increase the viability after DNA damage of the rad53K227A mutant (Fig. 3A). Because Rfa2p is not phosphorylated in set1Δ rad53K227A cells, we considered the possibility that the set1Δ-dependent phosphorylation of Rfa2p was linked to the increase of the DNA repair capacities of the cell.
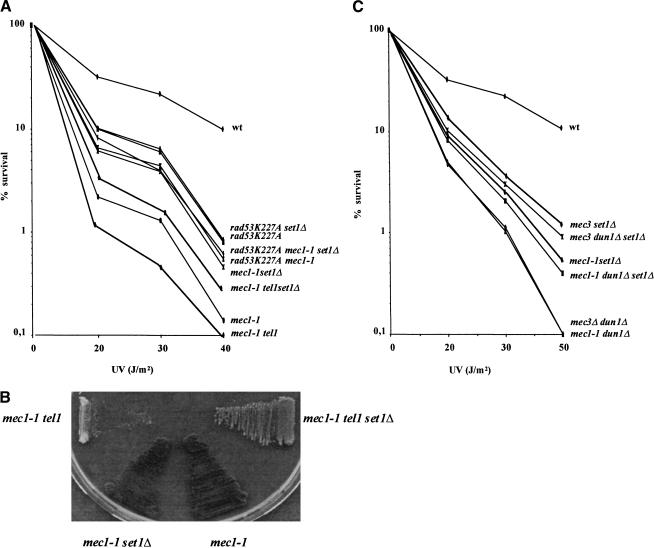
Figure 3.
Viability after UV irradiation. Approximately 1000 cells of K699 derivative strains with the indicated relevant genotypes were spread on YPD plates and exposed to ultraviolet irradiation. Cells were grown at 30°C for 5 d before we counted the number of colonies. (A) The set1Δ mutation partially rescues ultraviolet sensitivity of sml1 mec1-1 but not of rad53K227A and sml1 mec1-1 rad53K227A mutants; (B) the deletion of SET1 suppresses the growth defect of a sml1 mec1-1 tel1 double mutant; (C) the set1Δ mutation increases the viability after ultraviolet of mec3Δ dun1Δ and sml1 mec1-1 dun1Δ cells.
We first asked whether inactivation of Set1p partially rescued the ultraviolet sensitivity of a sml1 mec1-1 mutant (Fig. 3A). We showed that the viability after irradiation of a sml1 mec1-1 mutant was increased in the absence of SET1 (Fig. 3A). Similarly, the set1Δ mutation increased the viability after UV of a sml1 mec1-1 tel1 mutant (Fig. 3A).
To examine whether the increase of viability of sml1 mec1-1 set1Δ cells with respect to sml1 mec1-1 cells after ultraviolet treatment requires the kinase activity of Rad53p, we compared the ultraviolet sensitivity of a sml1 mec1-1 rad53K227A with the isogenic sml1 mec1-1 rad53K227A set1Δ. Both mutants had the same sensitivity to UV irradiation (Fig. 3A). Moreover, the deletion of SET1 did not increase the viability after UV irradiation of a mec3Δ rad53Δ double mutant (data not shown). These results indicate that the set1Δ-dependent increase of viability of sml1 mec1-1 and mec3Δ mutants after DNA damage is mediated by kinase activity of Rad53p.
Surprisingly and yet unexplained, we observed repeatedly that the sml1 mec1-1 rad53K227A (or rad53Δ) mutant was less sensitive to DNA damage than the sml1 mec1-1 mutant. Thus, the inactivation of RAD53 increased the viability after DNA damage of a sml1 mec1-1 mutant. This explained the fact that the sml1 mec1-1 rad53K227A set1Δ exhibited increased survival after UV irradiation with respect to the sml1 mec1-1 set1Δ, contrary to what one could have expected, as Rfa2p is not hyperphosphorylated in the sml1 mec1-1 rad53K227A set1Δ strain.
We also observed that the deletion of SET1 suppressed the growth defect of the sml1 mec1-1tel1 mutant (Fig. 3B), strengthening evidence for the existence of pathways activated by SET1 inactivation, but independent of MEC1 and TEL1. Interestingly, we noticed that the deletion of SET1 in a sml1 mec1Δ tel1Δ strain increased telomere length (data not shown). This could account for the rescue of the poor viability of the sml1 mec1Δ tel1Δ mutant by the deletion of SET1. From these data and the data in Figure 1B, we ruled out that the set1Δ mutation exerts its action via MEC1 and TEL1.
The protein kinase Dun1p has been shown to act downstream of Rad53p and to be involved in the induction of the expression of many repair genes after DNA damage (Zhou and Elledge 1993). We asked whether Dun1p relays the increase of the DNA repair capacities in set1Δ cells. We introduced the set1Δ mutation in dun1Δ mec3Δ and dun1 mec1-1 sml1 mutants. We found that the deletion of SET1 increased the viability of both mutants, dun1Δ mec3Δ and dun1 mec1-1 sml1 after ultraviolet treatment (Fig. 3C).
These results suggest that in set1Δ cells, the Rad53 kinase-dependent phosphorylation of Rfa2p induces a Dun1-independent increase of the DNA repair capacities of the cell.
set1Δ-dependent phosphorylation of Rfa2p decreases its binding to URS elements of repair genes, providing a mechanism for transcriptional induction
Our results indicate that phosphorylation of Rfa2p correlates with the increase of the DNA repair capacity of the cell. We therefore investigated whether the set1Δ-dependent phosphorylation of Rfa2p could modulate the induction of repair genes. It has been reported by Singh and Samson (1995) that RPA binds in vitro to the URS of 3-methyladenine DNA glycosylase repair gene (MAG) and to MAG URS1-like sequences of other repair genes, some of them being inducible by DNA-damaging agents such as RAD51 and RNR2.
We first investigated the association of Rfa2p to URS of RNR2 and RAD51 in vivo by performing chromatin immunoprecipitation experiments (ChIP) (Strahl-Bolsinger et al. 1997). We fused a 18-mer Myc epitope to the caboxyl terminus of the genomic copy of RFA2 and also of HDF2 (yKU80). We used yKu80 as a control for the specific association of Rfa2p with the URS. Chromatin from cells (UCC1001) expressing either Rfa2p–Myc18 or yKu80–Myc18 was cross-linked, immunoprecipitated, and analyzed by PCR. We performed PCR on input and on immunoprecipitated DNA with primers flanking the URS elements of RAD51 and RNR2 identified by Singh and Samson (1995) (Fig. 4A). Immunoprecipitation of Rfa2p–Myc18 resulted in the retention of PCR products corresponding to a region flanking the URS elements of RAD51 and RNR2, whereas the URS PCR fragments were not retained in untagged or yKu80–Myc18 strains (Fig. 4B). To show that the interaction between Rfa2p and URS2 is specific, we examined, as an additional control, the binding of Rfa2p to DNA regions adjacent to the URS2 of RAD51 and RNR2. These regions start in the coding sequences of RAD51 and RNR2 and are localized at about 1400 bp of the URS2 sequences of RNR2 and RAD51. As shown in Figure 4B, Rfa2p does not bind to these regions.
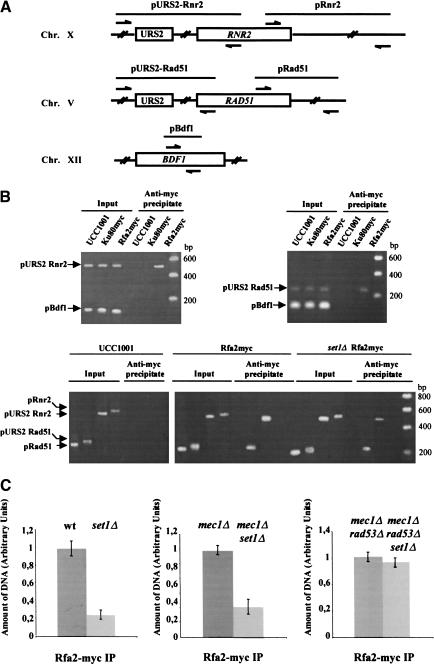
Figure 4.
set1Δ-dependent phosphorylation of Rfa2p decreases its binding to URS elements and leads to transcriptional induction of repair genes. (A) Scheme of the DNA regions flanking the URS elements of RNR2 (pURS2–Rnr2) and RAD51 (pURS2–Rad51) that were used to analyze the binding of Rfa2p to the URS elements. The pairs of primers used to amplify pURS2–Rnr2, pURS2–Rad51, and the control fragments pRnr2, pRad51, and pBdf1 are indicated. (B, top) Rfa2p binds to URS elements in vivo. The ChIP assay was performed as described in Materials and Methods. Rpa2myc and Ku80myc indicate that the parental strain UCC1001 bears a 18-mer Myc epitope fused to the carboxyl terminus of the genomic copy of RFA2 and HDF2, respectively. Immunoprecipitation was performed with anti-myc antibodies (9 E10). After DNA purification, PCR was performed with primers for the sequences flanking the URS2 of RNR2 (pURS2–Rnr2) and RAD51 (pURS2–Rad51), and for the control sequence in BDF1 (pBdf1). PCR reactions were performed on immunoprecipitated DNA and on the corresponding imput DNA. PCR products (indicated by arrows) were resolved on 2% agarose gels. (Bottom) Rfa2p binds to URS2–Rnr2 and URS2–Rad51 but not to the adjacent sequences, pRnr2 and pRad51. DNA regions pRnr2 and pRad51 are localized at ∼1400 bp of the URS2 sequences of RNR2 and RAD51, respectively. (C) In vivo association of Rfa2p with the URS2 of RAD51 is diminished in set1Δ cells. Chromatin solutions from wild-type and set1Δ cells expressing the Rfa2-myc were subjected to ChIP. The amount of precipitated DNA was analyzed by quantitative PCR (see Materials and Methods). The signals obtained by PCR in the Rfa2-myc immunoprecipitates were quantitated by referring them to a standard curve. This standard curve was obtained by plotting the signals obtained from PCRs performed with the same primers on serial dilutions of the corresponding input DNA. The effect of the set1Δ mutation was analyzed in different parental strains. For each couple of strains (UCC1001 (wt), UCC1001 set1Δ), (UCC1001 mec1Δ sml1Δ, UCC1001 mec1Δ sml1Δ set1Δ), and (UCC1001 mec1Δ rad53Δ sml1Δ UCC1001, mec1Δ rad53Δ sml1Δ set1Δ), amplification of the input DNA samples with the specific and control primers produced similar amount of pBdf1 and pURS2–Rad51 PCR DNA fragments. For each graph, the amount of DNA corresponding to pURS2–Rad51 in the precipitated DNA from the parental strain is arbitrary plotted as 1. The experiments shown were done in triplicate
These cross-linking experiments indicate that Rfa2p binds to URS of RNR2 and RAD51 in vivo.
As set1Δ-dependent phosphorylation of Rfa2p correlates with the enhancing of the DNA repair capacities, we compared the binding of Rfa2p with the URS of RAD51 in wild-type and set1Δ cells. We carried out ChIP experiments on chromatin from wild-type and set1Δ cells expressing Rfa2p–Myc18. We performed quantitative PCR with a Light Cycler on input DNA and on immunoprecipitates with URS-specific primers allowing the amplification of a PCR fragment flanking the URS of RAD51. PCR amounts in the immunoprecipitates were calculated by comparing the signal in the immunoprecipitates with a standard curve obtained by performing PCRs with the same primers on corresponding serially diluted input DNAs. The PCR quantitation analysis indicated that the amount of DNA corresponding to URS of RAD51 bound to Rfa2p was decreased by fourfold in a set1Δ mutant (Fig. 4C). As above, no URS was retained in wild-type or yKu80–Myc18 strains. These results may reflect the fact that the set1Δ-dependent hyperphosphorylation of Rfa2p decreases the binding of RPA to URS of RAD51. We further tested the effect of the SET1 deletion on the Rfa2p URS-binding activity by analyzing the in vivo association of Rfa2p to RAD51 URS in set1Δ cells in the presence or absence of MEC1 and RAD53. The ChIP assays indicate that the amount of Rfa2p-associated URS in sml1Δ mec1Δ set1Δ cells (hyperphosphorylation of Rfa2p) was 3.7-fold lower than the one in sml1Δ mec1Δ cells (no phosphorylation) (Fig. 4C). In contrast, the amount of Rfa2p associated with the URS remains approximately the same in sml1Δ rad53Δ mec1Δ cells (no phosphorylation of Rfa2p) and in sml1Δ rad53Δ mec1Δ set1Δ cells (no phosphorylation of Rfa2p) (Fig. 4C). Together these results indicate that the URS-binding activity of Rfa2p is reduced in set1Δ cells in a Rad53-dependent manner. The binding activity of Rfa2p to URS is thus likely to depend on its phosphorylation.
Such regulation may account for the transcriptional derepression of RNR2 and RAD51 genes in set1Δ cells. To confirm this hypothesis, we used a CYC1–lacZ vector (pNG22) containing the MAG URS1 region cloned 3′ of the CYC1 UAS (pNG22+URS1) (Singh and Samson 1995). We transformed pNG22 and pNG22+URS1 in wild-type and set1Δ strains and measured the β-galactosidase activity of equivalent amounts of cells. As reported by Singh and Samson (1995), we observed a threefold decrease of CYC1 UAS function in the presence of the MAG URS1 in a wild-type strain (Fig. 5A). In contrast, we observed that the repressive effect of the MAG URS1 region was abolished in set1Δ cells (Fig. 5A). Moreover, we found that the repressing activity of the MAG URS1 region was restored in set1Δ cells in the absence of the Rad53p kinase (Fig. 5A), which links the phosphorylation of Rfa2p with transcriptional induction. To further investigate the effect of the set1Δ mutation on the induction of repair genes, wild-type and set1Δ cells containing a chromosomally encoded RAD51–lacZ gene fusion were assayed for β-galactosidase activity. We observed that RAD51–lacZ expression was increased threefold in exponentially growing set1Δ cells relative to wild-type cells (Fig. 5B, left). This result suggests that the level of Rad51p and probably of other repair proteins is increased in set1Δ cells. Finally, results of Figure 3C indicate that Dun1p was not required for the effect of the set1Δ mutation on the increase of the DNA repair capacities of the cell. We further investigated this point by measuring β-galactosidase activities of cells bearing the β-galactosidase-reporter plasmid pZZ20. This plasmid contains a 42-bp element (D1) from the RNR2 promoter inserted between the α1 gene UAS and the CYC1 basal promoter. The D1 element has been shown to be critical for the Mec1–Rad53–Dun1-dependent transcriptional response to DNA damage (Huang et al. 1998). We found that wild-type and set1Δ cells bearing this plasmid had a similar lacZ activity suggesting that the absence of SET1 did not mediate any changes in the expression of RNR2 through sequences within D1 (Fig. 5B, right).
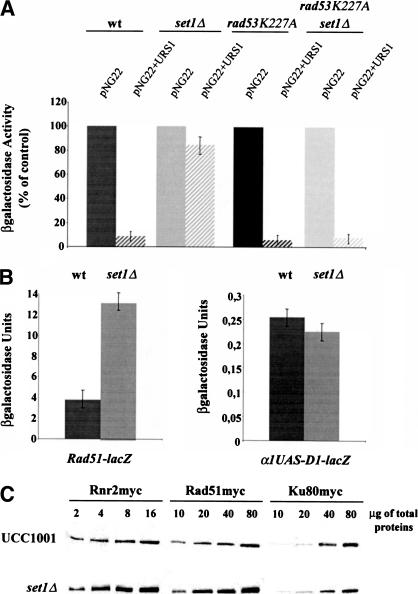
Figure 5.
(A) The set1Δ mutation abolishes the repression mediated by the MAG URS1 element. We compared for each strain the β-galactosidase activity associated with the CYC1–lacZ vector (pNG22) and with pNG22 containing the MAG URS1 element (pNG22+URS1). The enzymatic activity associated with pNG22 was taken as control. (B, left) The set1Δ mutation increases the RAD51–lacZ expression; (right) the set1Δ mutation does not act via the D1 element of the RNR2 promoter. (C) protein levels of Rad51p and Rnr2p increase in set1Δ cells. A 13-mer Myc epitope was fused to the carboxyl terminus of the genomic copy of RAD51 and RNR2. Ku80myc was used as an internal control. Identical amounts of total protein extracts were analyzed by SDS-10% PAGE and immunoblotting. The presence of Rnr2p, Rad51p, and Ku80p was revealed with 9 E10 (anti-myc) antibodies. Relative amounts of proteins were determined by analyzing the chemiluminescence signals.
The use of lacZ reporter constructs suggests that RAD51 expression is increased threefold in exponentially growing set1Δ cells relative to wild-type cells. To confirm that the protein levels of Rad51p and Rnr2p increase in set1Δ cells, we compared the amounts of Rnr2p, Rad51p, and Ku80p in set1Δ cells with wild-type cells. We analyzed by immunoblotting in wild-type and set1Δ cells identical amounts of total protein extracts for the presence of Myc-tagged Rnr2p, Rad51p, and Ku80p (Fig. 5C). Quantification analysis indicates that there is a three- to fourfold induction of the expression of Rnr2p and Rad51p in set1Δ cells compared with wild-type, whereas the amount of Ku80p remains unchanged.
Taken together, these results indicate that the set1Δ-dependent phosphorylation of Rfa2p decreases its binding to URS, resulting in a derepression of repair genes.
Rfa2 regulates the transcriptional induction of repair genes
To directly test the role of Rfa2p in the induction of the expression of repair genes, we analyzed the ultraviolet sensitivity of a mec3Δ strain carrying the rfa2 Δ40 allele. The mec3Δ rfa2 Δ40 strain produces Rfa2p Δ40 as a sole form of Rfa2p. Strikingly, the rfa2 Δ40 allele strongly suppressed the sensitivity to ultraviolet of the checkpoint mec3Δ mutant strain (Fig. 6A). These data indicate that the sensitivity to DNA damage of the checkpoint mutant mec3Δ is suppressed by the deletion of the amino-terminal region of Rfa2p. Moreover, we found that the set1Δ mutation had no additive effect on the ultraviolet sensitivity of the mec3Δ rfa2 Δ40 (Fig. 6A), indicating that the deletion of the amino-terminal region of Rfa2p abolishes the effect of the set1Δ mutation. Overall, these data are in full agreement with the idea that the amino terminus of Rfa2p is a target of a phosphorylation regulated by SET1.
Figure 6.
(A) The sensitivity to ultraviolet radiation of the mec3Δ mutant is suppressed by the deletion of the amino-terminal region of Rfa2p. Approximately 1000 cells of K699 derivative strains with the indicated relevant genotypes were spread on YPD plates and exposed to ultraviolet irradiation. Cells were grown at 30°C for 5 d before we counted the number of colonies. (B) Analysis of intraS checkpoint of the rfa2 Δ40 mutants. Exponentially growing (exp) wild-type, mec3Δ rfa2 Δ40 and mec3Δ rfa2 Δ40 cells were synchronized with α-factor and released in YPD medium with (+MMS) or without 0.02% MMS. (Left) Untreated or MMS-treated samples were taken at the indicated times after α-factor release and analyzed. (Right) Aliquots were removed from the MMS-treated cultures to determine cell number and to score for colony-forming units on YPD plates at 25°C. (C) The rfa2 Δ40 mutation abolishes the repression mediated by the MAG URS2 element. We measured the β-galactosidase activity associated with the CYC1–lacZ vector (pNG22) and with pNG22 containing the MAG URS1 element (pNG22+URS1). (D) The rfa2 Δ40 mutation increases the RAD51–lacZ expression.
Because the sensitivity to ultraviolet radiation of a mec3Δ checkpoint mutant was suppressed by the rfa2 Δ40 mutation, we determined whether the rfa2 Δ40 mutation modified cell-cycle progression after DNA damage and whether this mutation restored the ability of mec3Δ cells to arrest cell cycle progression. We released cells synchronized in G1 into the cell cycle in either the presence or absence of a sublethal dose of MMS and measured the DNA content of cell cultures every 15 min. (Fig. 6B). In the absence of DNA damage, both rfa2 Δ40 and rfa2 Δ40 mec3Δ mutants appear to replicate their DNA slightly more slowly than wild-type or mec3Δ cells (Fig. 6B). In the presence of MMS, wild-type and rfa2 Δ40 cells have still not completed replication 150 min after release from α-factor, whereas most mec3Δ cells had a 2C DNA content 75 min after α-factor release. This indicates that the rfa2 Δ40 mutation does not alter the ability of cells to delay replication after DNA damage. During MMS treatment, DNA replication in mec3Δ rfa2 Δ40 cells is completed 120 min after release, whereas rfa2 Δ40 cells still have a 1C DNA content 150 min after release. Thus, the rfa2 Δ40 mutation does not seem to suppress the checkpoint defect of mec3Δ cells. We also measured cell lethality during the MMS treatment (Fig. 6B). In agreement with the results described in Figure 6A, the deletion of the amino-terminal domain of Rfa2p suppressed the sensitivity to DNA damage of the mec3Δ mutant.
We then asked whether enhancing of the DNA repair capacities in the rfa2 Δ40 mutant was associated with transcriptional derepression of repair genes as is the case in set1Δ cells. We first transformed pNG22 (CYC1–lacZ vector) and pNG22+URS1 in the rfa2 Δ40 mutant and measured the β-galactosidase activity. As for the set1Δ mutation, we observed that the repressive effect of the MAG URS1 region was abolished in rfa2 Δ40 cells (Fig. 6C). We further investigated the effect of the rfa2 Δ40 mutation on the induction of repair genes by fusing lacZ to RAD51 in rfa2 Δ40 cells. We observed that expression of the chromosomally encoded RAD51–lacZ was increased about threefold in exponentially growing rfa2 Δ40 cells relative to wild-type cells (Fig. 6D). This result suggests that the level of Rad51p and probably of other repair proteins is increased in rfa2 Δ40 cells as is the case in set1Δ cells.
Discussion
The absence of SET1 activates a Rad53p-signaling pathway
We have shown previously that the deletion of SET1 abolishes TPE, reduces telomere length, and increases the viability after DNA damage of a mec3 null mutant and also of other checkpoint mutants (rad9, rad17, rad24). The major contribution of this work is the finding that, in response to the loss of SET1, a novel Rad53-dependent pathway leading to Rfa2p hyperphosphorylation is activated and this results in activation of repair genes.
The proteins containing a SET domain are considered to behave as global chromatin regulators, a concept strengthened by the recent finding of a SET-dependent histone H3-specific methyltransferase activity (Rea et al. 2000). The first interpretation of the Rad53-dependent pathway induction upon the loss of SET1 is that this response results from an alteration in the chromatin organization. It is tempting to speculate that this pathway is initiated by changes in the posttranslational modification of chromatin components. This suggests the existence of chromatin damages that can be relayed by signaling pathways.
Another, not exclusive, interpretation for the set1Δ-induced cellular response is that SET1 regulates phosphorylation pathways by modulating the activity of kinases or phosphatases. For instance, phosphatases that dephosphorylate Rfa2p could be potential targets for Set1p. Cui et al. (1998) proposed that interaction between the SET domain and SET interacting domains (SID) in mammalian cells might regulate the activity of phosphatases or so-called antiphosphatases, thereby engaging cells through growth or differentiation. Interestingly, sml1 mec1-1 set1Δ cells exhibit elongated buds (V. Schramke and V. Gelí, unpubl.) similar to those displayed by hsl1 and hsl7 (histone synthetic lethal) or cdc55 mutants (Ma et al. 1996; Jiang and Broach 1999). hsl1 and hsl7 were identified in a screen for mutations that are lethal in combination with a deletion of the amino terminus of histone H3, whereas Cdc55p is a regulatory subunit of phosphatase 2A. Strikingly, we have also observed that the deletion of SET1 suppresses the growth defect of a sml1 mec1-1 tel1 double mutant. sml1 mec1-1 tel1 double-mutant strains have very short telomeres and a senescent phenotype in S. cerevisiae and lose all telomeric sequences in Schizosaccharomyces pombe (Naito et al. 1998; Ritchie et al. 1999). As a model for the functions of Tel1p and Mec1p, it has been proposed that Tel1p and Mec1p affect the accessibility of telomeres to telomerase and exonucleases by phosphorylation of target proteins located at telomeres (Ritchie et al. 1999). Taken together, these observations suggest further links between SET1, chromatin, telomeres, and signaling phosphorylation.
The set1Δ pathway overlaps with but is distinct from the DNA damage pathway
Our work establishes that the set1Δ-induced response is different from the response produced by a DNA damage, although some components are shared and although both pathways result in an increased repair potency. In particular, we showed that (1) the hyperphosphorylation of Rfa2p is ATM homolog independent; (2) although the Rad53 kinase domain is required, no apparent phosphorylation of Rad53p can be visualized in set1Δ cells; (3) the activation of the repair genes in set1Δ cells does not require DUN1; (4) the checkpoint functions and cell cycle progression are not altered in set1Δ cells. On the basis of these findings, a model for the signal transduction pathway in response to set1Δ can be suggested (Fig. 7; see below). We do not think that the effects of set1Δ are simply due to the set1Δ-induced loss of TPE or to the alteration in expression of genes near telomeres, because a sas2 mutant, which also has a clear TPE defect (Reifsnyder et al. 1996), does not exhibit the phenotypes associated with the set1Δ mutant.
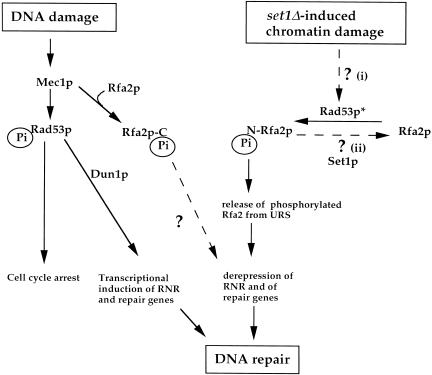
Figure 7.
A schematic representation of the set1Δ-induced cellular response that leads to the transcriptional activation of repair genes. We propose that phosphorylation of Rfa2p can result either from (1) the activation of Rad53p (Rad53*) in response to a set1Δ-induced chromatin damage, or (2) from the deregulation of a phosphorylation pathway controlled by Set1p. Both models are not exclusive. The set1Δ cellular response is shown in parallel with the DNA damage response. N-Pi and C-Pi indicated phosphorylation in the amino and carboxyl terminus of Rfa2p, respectively.
Rfa2p hyperphosphorylation in response to the loss of SET1
Phosphorylation of yeast Rfa2p has been shown to depend on Mec1p and Tel1p (Brush et al. 1996). Nevertheless, it has not been determined whether MEC1 and TEL1 encode RPA kinases, a role that has been mainly suggested for DNA–PK, ATM, and cyclin-dependent kinases (CDK) in human cells (Liu and Weaver 1993; Brush et al. 1996; Henricksen et al. 1996; Niu et al. 1997). Our results suggest that the amino-terminal region of Rfa2p is required for the Set1p-regulated phosphorylation of Rfa2p in a Mec1-independent manner. Importantly, we show that Rad53p is required for this set1Δ-dependent phosphorylation. Our data also indicate that either Mec1p is involved in the phosphorylation of a region outside of the first 40 residues of Rfa2p, or that truncation of the amino-terminal region of Rfa2p unveils a cryptic site phosphorylated in a Mec1-dependent way. Additional work will be required to identify the Mec1- and Set1-dependent phosphorylation sites of Rfa2p.
set1Δ alleviates the repression of repair genes
We observed previously that the deletion of SET1 increased the viability after DNA damage of DNA damage sensor mutants (rad9, rad17, rad24, mec3) without restoring any of the three DNA damage checkpoints (Corda et al. 1999). We now establish that this holds true for sml1 mec1-1 and sml1 mec1-1 tel1 double mutants and that the increased repair capacity of set1Δ cells can be, at least to some extent, explained by an enhanced expression of repair genes. The transcription of many repair genes is under the negative control of URS elements that are recognized in vitro by RPA (Singh and Samson 1995). We show here that these sequences are efficiently bound by Rfa2p in vivo, but are no longer associated with Rfa2p in set1Δcells. Furthermore, the loss of SET1 relieves the repressive effect of an URS element located upstream of a heterologous promoter in a Rad53-dependent fashion. Altogether, these results suggest that the loss of SET1 alleviates the repressive effect of RPA at the promoter of some repair genes. Whether the Mec1-dependent phosphorylation of Rfa2p contributes to the derepression of the repair genes remains to be determined.
In addition to an increased expression of repair genes, set1Δ cells could improve their repair capacity by other means. For instance, the phosphorylated Rfa2p proteins could participate in the processing of DNA lesions. It has been proposed that RAD9, RAD17, RAD24, and MEC3 process some forms of DNA damage to a single strand by DNA degradation (Lydall and Weinert 1995).
Rfa2p regulates in vivo the expression of repair genes
Our results indicate a strong correlation between the Rad53-dependent phosphorylation of Rfa2p in set1Δ cells and the partial suppression of checkpoint mutant sensitivity to radiation. The role of Rfa2p in this process is strengthened by the observation that the deletion of the amino terminus of Rfa2p suppresses the sensitivity of mec3Δ cells to ultraviolet and increases the expression of the Rad51–lacZ reporter protein. We explain the similarity between the phenotypes displayed by the rfa2 Δ40 mutant and those displayed by set1Δ mutant by the fact that both the Rfa2p Δ40 and the hyperphosphorylated Rfa2p have a reduced ability to bind the URS elements. In agreement with this proposal, the deletion of SET1 does not further increase the ultraviolet resistance of rpa2Δ 40 mec3Δ cells. This shows that the forty-first amino acids of Rfa2p are required for set1Δ to rescue the ultraviolet sensitivity of mec3Δ. Because rfa2 Δ40 appears to be epistatic to set1Δ, this further links hyperphosphorylation of Rfa2p with the increased ultraviolet resistance of a mec3Δ set1Δ strain as compared with a mec3Δ strain.
Suppression of UV and MMS sensitivity of mec3Δ cells by the rfa2 Δ40 allele does not appear to imply suppression of mec3Δ checkpoint defect. These data are in agreement with previous observations (Lydall and Weinert 1995; Corda et al. 1999), suggesting that the loss of viability after DNA damage of the checkpoint mutants is not solely due to their inability to arrest cell cycle progression after DNA damage. The delay in DNA replication that we observed in the mec3Δ rfa2 Δ40 MMS-treated cells compared with the mec3Δ single mutant is likely due to the fact that the rfa2 Δ40 mutation causes, per se, a delay in DNA replication even in the absence of DNA damage. The effect of this replication defect might be amplified when DNA synthesis occurs on a damaged template.
New connections between chromatin and repair
A remarkable result of this work is that the same mutation (set1Δ) increased repair potency of cells and lead to chromatin injuries. The set1Δ cellular response may correspond to the derepression of at least 12 DNA repair and metabolism genes carrying MAG URS-like sequences (Xiao et al. 1993), namely MAG and MGT1 (DNA alkylation repair genes), PHR1 (involved in light-dependent repair of pyrimidine dimers), RAD1, RAD2, RAD4, RAD10, and RAD16 (components of the nucleotide excision repairosome), RAD51 (a central component of the recombinational repair), DDR48 (a stress protein induced by DNA damage), and RNR2 and RNR3 (subunits of the ribonucleotide reductase). Understanding the raison d'être of this response is likely to provide new insights into the connections between chromatin and repair. For instance, one can speculate that the increased repair capacity counteracts an enhanced susceptibility of DNA to damage in the context of a dysfunctional chromatin. Also, the repair genes can have a function in chromatin organization, even in the absence of DNA damage. This could occur in wild-type cells when the chromatin has to be transiently reconstructed, for example, during cell cycle or when large chromatin domains are reorganized in response to modifications of the cellular program.
In light of the conservation through evolution of the proteins involved in the set1Δ pathway, the relationships between chromatin-associated SET domain proteins and signaling kinases established here in yeast are likely to be shared by higher eukaryotes.
Materials and methods
Yeast strains and plasmids
Most yeast strains used in this study are isogenic either to K699 (W303 background) or UCC1001 (YPH250 background). Chromosomal deletions of SET1 and MEC3 were performed as described (Corda et al. 1999). The sml1 mec1-1 strain (DMP2394/18B) is virtually isogenic to K699 (Paciotti et al. 1998). The sml1 mec1-1 allele that we used eliminates the essential functions of MEC1 (Weinert et al. 1994; Zhao et al. 1998). To disrupt the 5′ end of TEL1 with URA3, we linearized plasmid pPG47 with SacI and transformed appropriate yeast strains (Greenwell et al. 1995). The rad53K227A mutant strain is isogenic with W303 (Pellicioli et al. 1999). To introduce the null allele of MEC1 and RAD53, we first disrupted SML1 by PCR amplifying a sml1∷TRP1 disruption cassette from plasmid pWJ700 (kindly provided by X. Zhao, Columbia University, New York). The set1Δ∷KAN, mec1Δ∷KAN, mec1Δ∷LYS2 null mutations were introduced as described (Fairhead et al. 1996). RAD53 was disrupted by integration of a PCR product containing the nourseothricin (nat) resistance gene (Goldstein and McCusker 1998). yKu80 and Rfa2p were tagged at their carboxyl termini with 18 MYC epitopes with the help of pVL1001 (from Connie Nugent, Baylor College of Medicine, Houston, TX) and pGG1 (from Jérôme Tonnelier, CNRS, Marseille, France), respectively. To analyze Ddc1p phosphorylation in set1Δ cells, SET1 has been disrupted in YLL334 that carries a chromosomal DDC1–HA2-tagged allele.
lacZ plasmids to assay the URS function, pNG22 and pNG22+URS1, were described by Singh and Samson (1995). The RAD51–lacZ gene fusion has been described previously (Aboussekhra at al. 1992). The RAD51–lacZ gene fusion containing the entire RAD51 gene has been integrated into the RAD51 locus by homologous recombination. Plasmid pZZ20 that contains the α1–UAS-D1–CYC1–lacZ region has been described by Huang et al. (1998).
The deletion removing the RFA2 intron and the region extending from codon 3 to codon 40 of RFA2 was performed by PCR site-directed mutagenesis. By replacing the chromosomally encoded RFA2 with the rfa2 Δ40 mutant allele, we generated an allele encoding a truncated protein extending from residue 1 to 165 (rfa2 16) that is coexpressed with Rfa2p Δ40. To eliminate potential effects of rfa2 165, this nonviable truncated allele was fully disrupted with a kanMX4 marker. The resulting strain produces Rfa2p Δ40 as the sole form of Rfa2p. As a control, we also constructed a strain producing the truncated Rfa2p 165 with the full-length Rfa2p (data not shown). This control was done to ensure that the phenotypes associated with the allele rfa2 Δ40 are due to the deletion of the amino terminus region.
Protein extracts and Western blot analysis
Protein extracts for Western blot analysis were prepared from trichloroacetic acid (TCA)-treated yeast cells. Protein extracts were resolved by electrophoresis on a 15% SDS–polyacrylamide gel (80:1 acrylamide:bis-acrylamide). Immunoblots were developed with rabbit anti-serum directed against Rfa2p (kindly provided by S. Brill, Rutgers University, NJ and B. Stillman, Cold Spring Harbor Laboratory, NY) or with anti-HA monoclonal antibodies (for the analysis of Ddc1p).
Native extract preparation and phosphatase treatment
Exponentially growing cells (15 mL) were harvested by centrifugation at 4°C. The pellet was washed with 10 mL of ice-cold water and the cells were collected by centrifugation. The pellet was resuspended in 0.2 mL of lysis buffer (50 mM HEPES at pH 7.5, 2 mM EDTA, 2 mM DTT, 20% glycerol, 0.2 mM PMSF). An equal volume of acid-washed glass beads was added and the cells were disrupted by vortexing. Native extracts were divided into two samples, and 15 μL of native extract were incubated for 30 min at 30°C with 2 μL of 10× λ-phosphatase buffer (Biolabs) supplemented with 2 mM MnCl2 with or without 150 units of λ-phosphatase (Biolabs). Rfa2p was analyzed by Western blot analysis as described previously.
Chromatin immunoprecipitation
The extraction and sonication of formaldehyde-fixed yeast cells were done as described (Strahl-Bolsinger et al. 1997). The average size of sonicated DNA was between 500 and 1000 bp. Immunoprecipitation of cross-linked DNA was performed with 9 E10 (anti-myc) agarose-conjugate monoclonal antibodies (Santacruz Biotechnology) for 3 h at 4°C. An aliquot of each sample was not immunoprecipitated (input). PCR reaction parameters were as described (Strahl-Bolsinger et al. 1997) except for the annealing temperature (62°C ). Primers used to amplify the 500-bp RNR2 URS2 (pURS2Rnr2) (5′-GCAACGGGCAACCGTTT GGGG-3′; 5′-GGAAGGGGTCTCTTTAGGC-3′) and the 250-bp RAD51 URS2 (pURS2Rad51) (5′-CGCGTCATTCCGCTAT TTC-3′; 5′-CGAACCGTTCCCGTACTGAAGC-3′) are specific for the upstream regions of RNR2 andRAD51, respectively. Both PCR DNA fragments flank the URS2 sequence. Control primers used to amplify the 520-bp RNR2 (pRnr2) (5′-CATCTCCTTGGCCGGTAAGACC-3′; 5′-CCGCAGATGGG ACCTCTTATG-3′) and the 230-bp RAD51 (pRad51) (5′-CTC ACCTTGCTTACCAGAGGC-3′; 5′-CCGCAGTAGGGTTGC GAGGTA-3′) control sequences are specific for regions starting 120 and 159-bp upstream of the stop codons of RNR2 and RAD51, respectively. Control primers specific for BDF1 (5′-GC CCCGCGCACTTTGTGTGTGC-3′; 5′-GGGAAGGAGCGAC CCAGTAGGC-3′) produce a 100-bp PCR fragment.
For DNA quantitation, amplifications were done by use of a LightCycler (Roche Biochemicals) in a standard PCR reaction containing 0.5 μM of primers specific for the pURS2Rad51 and SYBR Green I Dye (Roche Biochemicals). Template concentration was determined by absorbance at 260 nm. The reactions were cycled 45 times with the following parameters: denaturation at 95°C, annealing at 62°C for 8 sec followed by a 16-sec extension at 72°C. Fluorescence was acquired at the end of the extension phase. The melting curves were obtained at the end of amplification by cooling the sample to 70°C (20°C/sec) and then increasing the temperature to 95°C at 0.1°C/sec. Fluorescence was acquired every 0.1°C. Quantitation data was analyzed by use of the Light Cycler analysis software.
Miscellaneous
Radiation sensitivity and MMS synchrony were carried out as described (Paulovich and Hartwell 1995; Corda et al. 1999). β-galactosidase activity was assayed as described previously (Kovari et al. 1990). Protein concentration was determined with the BCA protein assay reagent. Total protein extracts were TCA precipitated and analyzed by SDS-10% PAGE and immunoblotting. The presence of Rnr2p–myc13, Rad51p–myc13, and Ku80p–myc18 was revealed with 9 E10 (anti-myc) antibodies. Relative amounts of proteins were determined with a Kodak image station 440 CF.
Acknowledgments
We thank B. Stillman and S. Brill for the anti-Rfa2p antibodies. We thank J. Mallory, T. Petes, X. Zhao, and R. Rothstein for their help with TEL1 and SML1 disruptions, respectively; M.B. Vaze and J. Haber for their help in disrupting RAD53; P.A. Goldstein and J.H. McCusker, for pAG25(natMX4); C. Nugent and V. Lundblad for plasmids to tag yKu80p; and M. Foiani for providing the rad53K227A mutant. We thank L. Samson for plasmid pNG22 and pNG22+URS1, F. Fabre for the RAD51–lacZ fusion, and S. Elledge and D. Liu for plasmid pZZ20. We thank C. Mann, C. Marsolier, C. Nugent, J. Haber, F. Fabre, S. Elledge, L. Samson, C. de la Roche Saint Andre, and S. Brill for helpful discussions. The work in the Laboratories of V.G. was supported by l'Association pour la Recherche sur le Cancer, of E.G. by the Ligue Nationale contre le Cancer, of M.P.L and G.L. by the Cofinanziamento 1999 MURST-Università di Milano-Bicocca and CNR Target Project on Biotechnology Grant CT.97.01180.PF49(F).
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
E-MAIL geli@ibsm.cnrs-mrs.fr; FAX 33-4-91-71-21-24.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.193901.
References
- Aboussekhra A, Chanet R, Adjiri A, Fabre F. Semidominant suppressors of Srs2 helicase mutations of Saccharomyces cerevisiae map in the RAD51 gene, whose sequence predicts a protein with similarities to procaryotic RecA proteins. Mol Cell Biol. 1992;12:3224–3234. doi: 10.1128/mcb.12.7.3224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aboussekhra A, Vialard JE, Morrison DE, de la Torre-Ruiz MA, Cernakova L, Fabre F, Lowndes NF. A novel role for the budding yeast RAD9 checkpoint gene in DNA damage-dependent transcription. EMBO J. 1996;15:3912–3922. [PMC free article] [PubMed] [Google Scholar]
- Allen JB, Zhou Z, Siede W, Friedberg EC, Elledge SJ. The SAD1/RAD53 protein kinase controls multiple checkpoints and DNA damage-induced transcription in yeast. Genes & Dev. 1994;8:2416–2428. doi: 10.1101/gad.8.20.2401. [DOI] [PubMed] [Google Scholar]
- Brill SJ, Stillman B. Replication factor-A from Saccharomyces cerevisiae is encoded by three essential genes coordinately expressed at S phase. Genes & Dev. 1991;5:1589–1600. doi: 10.1101/gad.5.9.1589. [DOI] [PubMed] [Google Scholar]
- Brush GS, Kelly TJ. Phosphorylation of the replication protein A large subunit in the Saccharomyces cerevisiae checkpoint response. Nucleic Acids Res. 2000;19:3725–3732. doi: 10.1093/nar/28.19.3725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brush GS, Morrow DM, Hieter P, Kelly TJ. The ATM homologue MEC1 is required for phosphorylation of replication protein A in yeast. Proc Natl Acad Sci. 1996;93:15075–15080. doi: 10.1073/pnas.93.26.15075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corda Y, Schramke V, Longhese MP, Smokvina T, Paciotti V, Brevet V, Gilson E, Geli V. Interaction between Set1p and checkpoint protein Mec3p in DNA repair and telomere functions. Nat Genet. 1999;21:204–208. doi: 10.1038/5991. [DOI] [PubMed] [Google Scholar]
- Cui X, De Vivo I, Slany R, Miyamoto A, Firestein R, Cleary ML. Association of SET domain and myotubularin-related proteins modulates growth control. Nat Genet. 1998;18:331–337. doi: 10.1038/ng0498-331. [DOI] [PubMed] [Google Scholar]
- Din S, Brill SJ, Fairman MP, Stillman B. Cell-cycle-regulated phosphorylation of DNA replication factor A from human and yeast cells. Genes & Dev. 1990;6:968–977. doi: 10.1101/gad.4.6.968. [DOI] [PubMed] [Google Scholar]
- Fairhead C, Llorente B, Denis F, Soler M, Dujon B. New vectors for combinatorial deletions in yeast chromosomes and for gap-repair using split-marker recombination. Yeast. 1996;12:1439–1457. doi: 10.1002/(SICI)1097-0061(199611)12:14%3C1439::AID-YEA37%3E3.0.CO;2-O. [DOI] [PubMed] [Google Scholar]
- Fay DS, Sun Z, Stern DF. Mutations in SPK1/RAD53 that specifically abolish checkpoint but not growth-related functions. Curr Genet. 1997;31:97–105. doi: 10.1007/s002940050181. [DOI] [PubMed] [Google Scholar]
- Fried LM, Koumenis C, Peterson SR, Green SL, van Zijl P, Allalunis-Turner J, Chen DJ, Fishel R, Giaccia AJ, Brown JM, et al. The DNA damage response in DNA-dependent protein kinase-deficient SCID mouse cells: Replication protein A hyperphosphorylation and p53 induction. Proc Natl Acad Sci. 1996;93:13825–13830. doi: 10.1073/pnas.93.24.13825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gardner R, Putman W, Weinert T. RAD53, DUN1 and PDS1 define two paralllel G2/M checkpoint pathways in budding yeast. EMBO J. 1999;18:3173–3185. doi: 10.1093/emboj/18.11.3173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldstein AL, McCusker JH. Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast. 1998;15:1541–1553. doi: 10.1002/(SICI)1097-0061(199910)15:14<1541::AID-YEA476>3.0.CO;2-K. [DOI] [PubMed] [Google Scholar]
- Greenwell PW, Kronmal SL, Porter SE, Gassenhuber J, Obermaier B, Petes TD. TEL1, a gene involved in controlling telomere length in S. cerevisiae, is homologous to the human ataxia telangiectasia gene. Cell. 1995;82:823–829. doi: 10.1016/0092-8674(95)90479-4. [DOI] [PubMed] [Google Scholar]
- Henricksen LA, Carter T, Dutta A, Wold MS. Phosphorylation of human replication protein A by the DNA-dependent protein kinase is involved in the modulation of DNA replication. Nucleic Acids Res. 1996;24:3107–3112. doi: 10.1093/nar/24.15.3107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang M, Zhou Z, Elledge SJ. The DNA replication and damage checkpoint pathways induce transcription by inhibition of the Crt1 repressor. Cell. 1998;94:595–605. doi: 10.1016/s0092-8674(00)81601-3. [DOI] [PubMed] [Google Scholar]
- Iftode C, Daniely Y, Borowiec JA. Replication protein A (RPA): The eukaryotic SSB. Crit Rev Biochem Mol Biol. 1999;34:141–180. doi: 10.1080/10409239991209255. [DOI] [PubMed] [Google Scholar]
- Jenuwein T, Laible G, Dorn R, Reuter G. SET domain proteins modulate chromatin domains in eu- and heterochromatin. Cell Mol Life Sci. 1998;54:80–93. doi: 10.1007/s000180050127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang Y, Broach JR. Tor proteins and protein phosphatase 2A reciprocally regulate Tap42 in controlling cell growth in yeast. EMBO J. 1999;18:2782–2792. doi: 10.1093/emboj/18.10.2782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovari L, Sumrada R, Kovari I, Cooper TG. Multiple positive and negative cis-acting elements mediate induced arginase (CAR1) gene expression in Saccharomyces cerevisiae. Mol Cell Biol. 1990;10:5087–5097. doi: 10.1128/mcb.10.10.5087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu VF, Weaver DT. The ionizing radiation-induced replication protein A phosphorylation response differs between ataxia telangiectasia and normal human cells. Mol Cell Biol. 1993;13:2114–2123. doi: 10.1128/mcb.13.12.7222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longhese MP, Fraschini R, Plevani P, Lucchini G. Yeast pip3/mec3 mutants fail to delay entry into S phase and to slow down DNA replication in response to DNA damage and they define a functional link between Mec3 and DNA primase. Mol Cell Biol. 1996;16:3225–3244. doi: 10.1128/mcb.16.7.3235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longhese MP, Paciotti V, Fraschini R, Zaccarini R, Plevani P, Lucchini G. The novel DNA damage checkpoint protein Ddc1p is phosphorylated periodically during the cell cycle and in response to DNA damage in budding yeast. EMBO J. 1997;16:5216–5226. doi: 10.1093/emboj/16.17.5216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowndes NF, Murguia JR. Sensing and responding to DNA damage. Curr Opin Genet Dev. 2000;10:17–25. doi: 10.1016/s0959-437x(99)00050-7. [DOI] [PubMed] [Google Scholar]
- Lydall D, Weinert T. Yeast checkpoint genes in DNA damage processing: Implications for repair and arrest. Science. 1995;270:1488–1491. doi: 10.1126/science.270.5241.1488. [DOI] [PubMed] [Google Scholar]
- Ma XJ, Lu Q, Grunstein M. A search for proteins that interact genetically with histone H3 and H4 amino termini uncovers novel regulators of the Swe1 kinase in Saccharomyces cerevisiae. Genes & Dev. 1996;10:1327–1340. doi: 10.1101/gad.10.11.1327. [DOI] [PubMed] [Google Scholar]
- Naito T, Matsuura A, Ishikawa F. Circular chromosome formation in a fission yeast mutant defective in two ATM homologues. Nat Genet. 1998;20:203–206. doi: 10.1038/2517. [DOI] [PubMed] [Google Scholar]
- Nislow C, Ray E, Pillus L. SET1, a yeast member of the trithorax family, functions in transcriptional silencing and diverse cellular mechanisms. Mol Biol Cell. 1997;8:2421–2436. doi: 10.1091/mbc.8.12.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niu H, Erdjument-Bromage H, Pan ZQ, Lee SH, Tempst P, Hurwitz J. Mapping of amino acid residues in the p34 subunit of human single-stranded DNA-binding protein phosphorylated by DNA-dependent protein kinase and Cdc2 kinase in vitro. J Biol Chem. 1997;272:12634–12641. doi: 10.1074/jbc.272.19.12634. [DOI] [PubMed] [Google Scholar]
- Paciotti V, Lucchini G, Plevani P, Longhese MP. Mec1p is essential for phosphorylation of the yeast DNA damage checkpoint protein Ddc1p, which physically interacts with Mec3p. EMBO J. 1998;17:4199–4209. doi: 10.1093/emboj/17.14.4199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulovich AG, Hartwell LH. A checkpoint regulates the rate of progression through S phase in S. cerevisiae in response to DNA damage. Cell. 1995;82:841–847. doi: 10.1016/0092-8674(95)90481-6. [DOI] [PubMed] [Google Scholar]
- Pellicioli A, Lucca C, Liberi G, Marini F, Lopes M, Plevani P, Romano A, Di Fiore PP, Foiani M. Activation of Rad53 kinase in response to DNA damage and its effect in modulating phosphorylation of the lagging strand DNA polymerase. EMBO J. 1999;18:6561–6672. doi: 10.1093/emboj/18.22.6561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Philipova D, Mullen JR, Maniar HS, Lu J, Gu C, Brill SJ. A hierarchy of SSB protomers in replication protein A. Genes & Dev. 1996;10:2222–2233. doi: 10.1101/gad.10.17.2222. [DOI] [PubMed] [Google Scholar]
- Rea S, Eisenhaber F, O'Carroll D, Strahl BD, Sun ZW, Schmid M, Opravil S, Mechtler K, Ponting CP, Allis CD, et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature. 2000;406:593–599. doi: 10.1038/35020506. [DOI] [PubMed] [Google Scholar]
- Reifsnyder C, Lowell J, Clarke A, Pillus L. Yeast SAS silencing genes and human genes associated with AML and HIV-1 Tat interactions are homologous with acetyltransferases. Nat Genet. 1996;14:42–49. doi: 10.1038/ng0996-42. [DOI] [PubMed] [Google Scholar]
- Ritchie KB, Mallory JC, Petes TD. Interactions of TLC1 (which encodes the RNA subunit of telomerase), TEL1, and MEC1 in regulating telomere length in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1999;19:6065–6075. doi: 10.1128/mcb.19.9.6065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanchez Y, Desany BA, Jones WJ, Liu Q, Elledge SJ. Regulation of RAD53 by the ATM-like kinases MEC1 and TEL1 in yeast cell cycle checkpoint pathway. Science. 1996;271:357–360. doi: 10.1126/science.271.5247.357. [DOI] [PubMed] [Google Scholar]
- Singh KK, Samson L. Replication protein A binds to regulatory elements in yeast DNA repair and DNA metabolism genes. Proc Natl Acad Sci. 1995;92:4907–4911. doi: 10.1073/pnas.92.11.4907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strahl-Bolsinger S, Hecht A, Luo K, Grunstein M. SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes & Dev. 1997;11:83–93. doi: 10.1101/gad.11.1.83. [DOI] [PubMed] [Google Scholar]
- Sun Z, Fay DS, Marini F, Foiani M, Stern DF. Spk1/Rad53 is regulated by Mec1-dependent protein phosphorylation in DNA replication and damage checkpoint pathways. Genes & Dev. 1996;10:395–406. doi: 10.1101/gad.10.4.395. [DOI] [PubMed] [Google Scholar]
- Weinert TA, Kiser GL, Hartwell LH. Mitotic checkpoint genes in budding yeast and the dependence of mitosis on DNA replication and repair. Genes & Dev. 1994;15:652–665. doi: 10.1101/gad.8.6.652. [DOI] [PubMed] [Google Scholar]
- Wold MS. Replication Protein A: A heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu Rev Biochem. 1997;66:61–92. doi: 10.1146/annurev.biochem.66.1.61. [DOI] [PubMed] [Google Scholar]
- Xiao W, Singh KK, Chen B, Samson L. A common element involved in transcriptional regulation of two DNA alkylation repair genes (MAG and MGT1) of Saccharomyces cerevisiae. Mol Cell Biol. 1993;13:7213–7221. doi: 10.1128/mcb.13.12.7213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zernik-Kobak M, Vasunia K, Connelly M, Anderson CW, Dixon K. Sites of UV-induced phosphorylation of the p34 subunit of replication protein A from HeLa cells. J Biol Chem. 1997;272:23896–23904. doi: 10.1074/jbc.272.38.23896. [DOI] [PubMed] [Google Scholar]
- Zhao X, Muller EG, Rothstein R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol Cell. 1998;2:329–340. doi: 10.1016/s1097-2765(00)80277-4. [DOI] [PubMed] [Google Scholar]
- Zhou Z, Elledge SJ. DUN1 encodes a protein kinase that controls the DNA damage response in yeast. Cell. 1993;75:1119–1127. doi: 10.1016/0092-8674(93)90321-g. [DOI] [PubMed] [Google Scholar]