Abstract
AvrRxv is a member of a family of pathogen effectors present in pathogens of both plant and mammalian species. Xanthomonas campestris pv. vesicatoria strains carrying AvrRxv induce a hypersensitive response (HR) in the tomato cultivar Hawaii 7998. Using a yeast two-hybrid screen, we identified a 14-3-3 protein from tomato that interacts with AvrRxv called AvrRxv Interactor 1 (ARI1). The interaction was confirmed in vitro with affinity chromatography. Using mutagenesis, we identified a 14-3-3-binding domain in AvrRxv and demonstrated that a mutant in that domain showed concomitant loss of interaction with ARI1 and HR-inducing activity in tomato. These results demonstrate that the AvrRxv bacterial effector recruits 14-3-3 proteins for its function within host cells. AvrRxv homologues YopP and YopJ from Yersinia do not have AvrRxv-specific HR-inducing activity when delivered into tomato host cells by Agrobacterium. Although YopP itself cannot induce HR, its C-terminal domain containing the catalytic residues can replace that of AvrRxv in an AvrRxv-YopP chimera for HR-inducing activity. Phylogenetic analysis indicates that the sequences encoding the C-termini of family members are evolving independently from those encoding the N-termini. Our results support a model in which there are three functional domains in proteins of the family, translocation, interaction, and catalytic.
Keywords: avirulence protein, yeast two hybrid interaction screen, effector, hypersensitive response (HR), Xanthomonas campestris pv. vesicatoria, Yersinia enterocolitica, Solanum esculentum
1. Introduction
Xanthomonas campestris pv vesicatoria (Xcv) (alternatively called X. vesicatoria, X. axonopodis pv. vescicatoria or X. euvesicatoria) is the causal agent of bacterial spot disease in both pepper and tomato [1,2]. Xcv strains carrying the avirulence gene avrRxv elicit resistance, characterized by localized necrosis referred to as the hypersensitive response (HR), in the tomato cultivar Hawaii 7998 [3]. AvrRxv-specific resistance in Hawaii 7998 is genetically complex [3,4]. It was found that avrRxv induces HR on a variety of non-host plants when expressed in Xanthomonas pathogens, including bean, soybean, cowpea, alfalfa, and cotton [5]. In contrast to AvrRxv-specific host resistance, the non-host resistance in bean is simply inherited. Although the plant HR response to avrRxv appear to be highly conserved, the genetic control of the resistance is not.
AvrRxv is a member of a family of pathogen effectors [6-8], that are translocated into host cells by the Type III secretion system (TTSS). AvrRxv exerts its effect intracellularly [9]. In addition to AvrRxv, there are three other AvrRxv family members present in Xcv. These are AvrBsT, XopJ and AvrXv4 [6,10,11], all of which are translocated to the host cell [8,12,13]. Genes with high levels of sequence similarity are also present in other plant pathogens [14-18], and in the plant symbiont Rhizobium [19]. Furthermore, AvrRxv-like sequences are found in mammalian pathogens, including YopJ and YopP from Yersinia spp., AvrA from Salmonella, and VopP from Vibrio parahemeolyticus [7,20-22]. Although there are nuclear localization signal sequences in many of these family members and PopP2 has been shown to localize to the nucleus, most AvrRxv family members, including AvrRxv, PopP1 and AvrXv4, apparently do not [6,13,17,23]. XopJ was shown to localize to the plasma membrane by virtue of a N-myristolylation motif, also essential for ability to induce HR on Nicotiana [24].
Recent studies suggest that YopJ and VopP are acetyltransferases, which regulate signaling by interfering with phosphorylation [25]. Previously, it was hypothesized that family members were similar to cysteine proteases [8,26,27]. Apparently, the catalytic triad that is conserved in all members of the family, may have acetyltransferase rather than protease activity [25]. Mutation of any amino acid in the triad results in loss of function in YopJ, AvrBsT, AvrRxv and AvrXv4 [8,13,23]. It is not clear how to interpret analyses indicating that YopJ and AvrXv4 disrupt SUMO post-translation modifications in host cells [8,13], except possibly by affecting expression levels [25]. In mammalian hosts, YopJ exerts its effect on several known signal transduction pathways, thereby preventing an immune response [28-33]. Mukherjee and coworkers [29] showed that YopJ prevents activation of the mitogen-activated protein kinase (MAPK) and nuclear factor κB (NFκB) signaling pathways through its function as an acetyltransferase. Acetylation of MKKs by YopJ prevent activation. Similarly, VopA has also been shown to be an acetyltransferase that inhibits MAPK signaling by inactivating kinase activity [22]. This family of effectors directly manipulates host cell signaling.
Molecular genetic control of plant host responses specific to these effectors is under study. Resistance to AvrXv4, AvrBsT and PopP2 is controlled by single cognate plant genes [10,34,35]. It has been shown PopP2 physically interacts with the Arabidopsis resistance protein RRS1-R in yeast and co-localizes with RRS1 proteins in the nucleus [17]. The association between the putative transcription factor function of the TIR-NBS-LRR-WRKY RRS1 proteins and sumoylation or acetylation, if any, is not known. Resistance associated with AvrBsT in Arabidopsis is conferred by the lack of a serine hydrolase with carboxylesterase activity called SOBER1 [34]. SOBER1 preferentially works on molecules containing short chain acyl groups. It is possible that in the absence of SOBER1, AvrBsT modulates posttranslational modification of a common substrate by affecting acylation [34]. The interactions of YopJ with members of mammalian signal transduction pathways is direct as demonstrated by Orth and coworkers who isolated several MAPK kinases using YopJ as bait in a yeast two-hybrid screen [30]. Because AvrRxv shares significant sequence identity with other members of the family, it is plausible that AvrRxv also directly interacts with and disrupts signaling pathways in host plants.
14-3-3 proteins are ubiquitous acidic proteins that are involved in a great variety of cellular events mediated by protein-protein interactions [36-38]. Originally named for brain proteins based on electrophoretic mobility and column fractions, 14-3-3 proteins form homo- and/or heterodimers that serve as regulatory adapter proteins in diverse cellular functions, including metabolism, protein targeting, and signal transduction [39-41]. Most 14-3-3 proteins bind specifically to phospho-serine proteins. Analysis of known 14-3-3 binding sites and peptide libraries identified motifs that are bound by 14-3-3 proteins. The first motif discovered is RSXpSXP (X is any amino acid), where the serine is phosphorylated [36,38]. This motif is called the Raf motif because it was first identified in the Raf-1 protein kinase. In addition, there is a 14-3-3 binding motif that does not require phosphorylation called the R18 motif, found in an inhibitory peptide (DI/L/VE) [42]. Although the vast majority of 14-3-3 interactions require phosphorylation, it is not required for binding of 14-3-3 proteins to several cellular proteins in mammals, most importantly, the ADP-ribosyltransferase effector ExoS of Pseudomonas aeruginosa [43-46]. Mutation of the key residues in ExoS for binding to the 14-3-3 inhibits virulence of P. aeruginosa in a cell infection model [47,48].
The involvement of some 14-3-3 proteins in plant host responses to pathogens is inferred by their transcriptional upregulation [49-52]. In addition, the ankyrin repeat-containing protein AKR2 that was shown to bind to a 14-3-3 protein controls level of resistance to a bacterial pathogen in Arabidopsis [53]. In mammalian cells, 14-3-3 proteins are involved in regulating apoptosis [54]. In addition, 14-3-3 proteins have been observed to be associated with kinases, often in large complexes [37]. Many of the mammalian protein kinases that associate with 14-3-3s lie in the same pathways (like the MAPK and ERK pathways) that are affected by the Yersinia effector proteins YopJ and YopP.
In this study we report the identification of a 14-3-3 protein, called ARI1, that interacts with AvrRxv. Xcv strains carrying AvrRxv mutants disrupted for 14-3-3-binding activity can no longer elicit an HR in tomato. We also analyzed the HR-inducing activity of AvrRxv homologues from Yersinia, when delivered by Xcv or expressed by the host. We show that although YopP itself cannot induce HR, its C-terminal domain containing the catalytic residues can replace that of AvrRxv for HR-inducing activity. These results demonstrate that the AvrRxv bacterial effector recruits 14-3-3 proteins for its function within host cells and that the catalytic residues can be swapped between diverse AvrRxv family members.
2. Materials and Methods
2.1. Plant growth and inoculations
All tomatoes (Solanum esculentum) were grown from seed in 4 inch diameter plastic pots with either UC or Sunshine potting mix in a greenhouse with ambient light. All plant inoculations were performed with either a plastic Pasteur pipet or a 1 cc syringe. Inoculated plants were maintained in Conviron E8 growth chambers (Controlled Environment, Inc., Winnipeg, Manitoba, Canada) with 16 hour photoperiods. Plant inoculations of Xanthomonas stains at 5 × 108 CFU/ml in H2O were performed on three to four different plants on the fourth to seventh leaves and experiments were done in triplicate. To assess the effect of gene expression in planta, Agrobacterium-mediated gene transfer was performed in tomato leaves as described [55,56].
2.2. Bacterial strains and molecular biology techniques
Bacterial strains used in this study were Xcv strains 89-1, 92-14, and 92-14 avrRxv:Ω [3,6]; E. coli strains DH5α and TOP10 (Invitrogen, Carlsbad, CA, USA); Agrobacterium tumefaciens C5C81 ± vir [57]. Bacterial strains were grown as described in Morales et al. [55].
Standard molecular biology techniques were used for manipulating plasmids, ligation, transformation, and performing polymerase chain reaction (PCR) [58]. DNA for sequencing was isolated using Qiagen plasmid isolation kits (Valencia, CA, USA). Amplification products were cloned using Topo TA Cloning Kit according to instructions (Invitrogen) and sequenced. For RNA isolations, Xanthomonas cells were lysed at 100°C in 50 mM EDTA, 0.3 M Na-Acetate, 0.625% SDS, 50 mM Tris HCl, pH 8.0. After two extractions each with phenol at 65°C and chloroform, the nucleic acids were precipitated with isopropanol and then 4 M lithium acetate. The integrity of the RNA was determined by denaturing gel electrophoresis.
To determine the 5′-terminus of the transcription product of avrRxv, primer extension was performed [58]. Ten pmol each of four primers (MLP1, 5′-TCTAGGTTGCACTCTTATGGAGTC-3′; MLP3, 5′-TGAGCCGCCCACTCCTAATGAT-3′; MLP4, 5′-TGATGTTGAAAACGACTGCTGCTT-3′; DA1, 5′-CCGCCCACTCCTAATGATCTGA-3′) were labeled using T4 Polynucleotide Kinase (Promega, Madison, WI, USA) and (γ-32P)ATP in T4 PKN buffer according to manufacturer's instructions. Labeled primers were purified using Pharmacia Nick Column Sephadex G-50 (Pharmacia Biotech, Piscataway, NJ, USA), annealed to 15 μg denatured RNA at 65°C for 3 hours in first strand buffer (Promega), and extended at 42°C with Superscript II Reverse Transcriptase (Invitrogen) and 0.1M DTT, 7 μg actinomycin D (Sigma-Aldrich, St. Louis, MO, USA), 35 U recombinant RNasin ribonuclease inhibitor (Promega) and 10 mM dNTPs. After extension, the samples were treated with RNAse A at 37°C for 15 minutes, extracted with phenol, chloroform and iso-amylalcohol (25:24:1) and then precipitated with ethanol. A 35S-dATP-labeled dideoxy sequence ladder was generated on pUC118RXV3 [3]. After fractionation, samples were visualized either by autoradiography or with a phosphoimager (Molecular Dynamics, Inc., Sunnyvale, CA, USA).
2.3. Interaction screen and confirmation of interaction
A tomato cDNA library was made from the tomato cultivar Hawaii 7998, and cloned into the phagemid vector Hybrizap (Stratagene, La Jolla, CA, USA), containing the activation domain. The coding sequence of AvrRxv was cloned into the yeast two hybrid vector pBD as a gene fusion with Gal4 DNA binding domain (Stratagene), creating pBD-avrRxv. The pBD-avrRxv construct was transformed into the yeast strain PJ69-4a and tested for autoactivation of reporter genes. The yeast two-hybrid screen was carried out according to manufacturer's instructions (Stratagene).
Full length ARI1 was cloned with 5′ RACE. The ARI1 open reading frame was tagged at its 3′-end with the c-myc epitope tag sequence using Tft9-Eco-For, 5′-GAATTCATGGCTTCTTCCAAAGAACG-3′ and 14-3-3-c-myc-Bam-Rev, 5′-GGATCCTCACAAATCTTCTTCAGAAATCAACTTTTGCTCTGCATCTTCACCTCCACCAGC-3′ into pCRII and then cloned into pAD and pRSETB on an EcoRI fragment from a partial digest, creating pAD-AriI-myc, and p6XHis-Ari1-myc, respectively.
The interaction in the yeast two-hybrid system of tagged versions of the proteins was confirmed using pBD-avrRxv-flag and pAD-AriI-myc. The interaction of 6XHis-Ari1-Myc and AvrRxv-Flag was verified in vitro by co-immunoprecipitation. The MEGAscript T7 Transcription Kit (Ambion, Austin, TX, USA) and the Flexi Rabbit Reticulocyte Lysate System (Promega) were used according to manufacturer's instructions. In vitro translated 35S-methionine labeled AvrRxv-Flag lysates were incubated alone and with excess unlabeled 6XHis-ARI1-Myc overnight, with Ni-NTA agarose according to instructions (Qiagen, Valencia, CA, USA). Eluted proteins were size fractionated on a 4-20% tris-glycine polyacrylamide gradient gel and immunoblotted as described below.
2.4. Cloning of AvrRxv variants, homologues and chimeras
PCR-mediated cloning was used to construct clones of avrRxv subclones, variants, homologues and chimeras incorporating restriction enzyme sites. After sequence verification of PCR products cloned into pCRII (Invitrogen), inserts were cloned behind the lac promoter in pVSP61 (DNA Plant Technology, Oakland, CA, USA) or pDSK519 [59] for testing in Xanthomonas, behind the CmR promoter in pTM100 for testing in Xanthomonas [60], or behind the 35S promoter in binary vectors pCB302 [61] or pMD1 [62] for testing in Agrobacterium.
The Yersinia homologue yopJ was cloned from pJ [28] into pVSP61 on a 1.8 kb SalI/EcoRI fragment, creating pGPB3. The yopJ gene was cloned into pCRII and pMD1 using PCR primers yopJ-U34XbaI, 5′-TGCGTCTAGATGTCATACCGCTGTTAATTCCCTG-3′ and yopJ-L34XhoI, 5′-ATTCTCGAGGGGGTATTCCCATACTGGAGCAAGA-3′, incorporating XbaI and XhoI sites. yopJ was removed from pCRII on an EcoRI digest and cloned into pBD.
The yopP gene was cloned from Y. enterolitica plasmid JB580v using PCR and the following primers, yopP-fwd, 5′-GAATTCATGATTGGGCCAATATCAC-3′ and yopP-rev, 5′-GTCGACTTATACTTTGAGAAGTGT-3′, incorporating EcoRI and SalI sites. yopP was removed from pCRII on an EcoRI fragment and cloned into pTM100 and pCB302, and on an EcoRI/SalI fragment into pBD.
Chimeras were made between yopP and avrRxv by sequence overlap extension (SOE) as described in Ciesiolka et al. [6]. The chimeric coding sequence for N-AvrRxv1-172-YopP104-289-C was made with PCR using primers avrRxv/yopP-fwd, 5′-GAGCGGTAGTGCGGCTGGGGGAGGGTGGAATACATTTC-3′ and avrRxv/yopP-rev, 5′-GAAATGTATTCCACCCTCCCCCAGCCGCACTACCGCTC-3′, along with F1-avrRxv, 5′-ATGTGCGACTCCATAAGAGTG-3′, and yopP-rev to make pCRII-RY4. For N-YopP1-103-AvrRxv173-373-C, primers yopP/avrRxv-fwd, 5′-CGCTTCATAATTAACATGGATGAAGACCCTAGGAG-3′ and yopP/avrRxv-rev, 5′-CTCCTAGGGTCTTCATCCATGTTAATTATGAAGCG-3′ were used with yopP-fwd and 3′-avrRxv-Sal, 5′-GTCGACATTGTCTCAGGATTGTAAGGC-3′ to make pCRII-YR3. The resulting chimeras were cloned into pTM100 and pCB302 on EcoRI fragments.
Mutations in two putative 14-3-3 binding sites in AvrRxv, were made using SOE. For the AvrRxv-Raf mutant, primers AvrRXV-D147A-forward, 5′-CTGCGCAGCTTCGCCACTCCG-3′, and AvrRXV-D147A-reverse, 5′-CTGCCCCGGAGTGGCGAAGCT-3′ incorporated the D147A mutant codon, and were used along with 5′avrRxv-Nco, 5′-GCCCATGGATGTGCGACTCCATAAG-3′, and 3′avrRxv-BamHI, 3′-CGGATCCATTGTCTCAGGATTGTAAGGC-3′ and ligated into pCRII, to make pCRII-avrRxv-Raf. An EcoRI fragment was cloned from that into pTM100, pCB302 and pBD. For the AvrRxv-R18 mutant, SOE primers MNIE-F, 5′-GAGTCACTTCGGCTGATGAACATTGAAAATCTC-3′, MNIE-R, 5′-CAGATGGGGGAGATTTTCAATGTTCATCAGCCG-3′, 5′avrRxv-Nco and 3′avrRxv-BamHI were used to incorporate the D123N mutation, and ligated into pCRII to make pCRII-avrRxv-D123N. pCRII-avrRxv-D123N was used as a template for SOE for the second mutation E125Q, and the product ligated into pCRII to make pCRII-avrRxv-R18. The insert was cloned into pTM100, pCB302 and pBD on an EcoRI fragment. To make a mutant in both the Raf and R18 domains, pCRII-avrRxv-R18 was used as a template for SOE with the Raf mutagenic primer sets, and cloned into pCRII to make pCRII-avrRxv- Raf/R18. The insert was cloned into pBD on an EcoRI fragment to make pBD-avrRxv-Raf/R18.
For interaction tests, the avrRxv open reading frame and was tagged at the 3′-end using the FLAG epitope coding sequence [9] using primers avrRxv-BamHI, 5′-GGATCCATGTGCGACTCCATAAG-3′ and Xba-Flag-Rev, 5′-TCTAGATCACTTATCATCATCATCCTTGTAATCGGATTCTAAGGCGTGACGGATCTTTCG -3′ and cloned into pCRII. The avrRxv-flag insert was cloned into pCB302 and pBD on an EcoRI fragment.
Two avrRxv deletion constructs were made in the pBD bait vector. An N-terminal deletion AvrRxv142-373, was constructed using PCR primers 5′M/avrRxv-EcoRI, 5′-GGGAATTCCTGCGCAGCTTCGACACTCCG-3′ and 3′-avrRxv-Sal. The PCR product was ligated into pCRII and then cloned on an EcoRI/SalI fragment to make pBD-avrRxv-5′Δ. The C-terminal deletion AvrRxv1-315 in pBD-avrRxv-3′Δ was constructed by cloning an EcoRI/XhoI fragment into pBD cut with EcoRI/SalI.
Mutant avrRxv genes in the catalytic domain were made by SOE and are named by the amino acid codon that changed. Primers (Table 1) were used in the following combinations for wildtype-untagged, EG16/EG13; for wildtype-HA, EG4/EG10; for H180A-untagged, EG16/EG15, EG14/EG13, EG16/EG13; for H180A-HA, EG6/EG12, EG11/EG10, EG6/EG10; for E200A-untagged, EG16/IL, IR/EG13, EG16/EG13; for E200A-HA, EG4/IL, IR/EG10, EG4/EG10; for C244A-untagged, EG6/EG9, EG8/EG7, EG6/EG7; for C244A-HA, EG4/EG5, EG2/EG1, EG4/EG1; for E258A-untagged, EG16/EG18, EG17/EG13, EG16/EG13; and for E258-HA, EG6/EG18, EG17/EG10, EG6/EG10; to clone into pCRII and then into pDSK519 and pMD1 on XbaI/BamHI fragments. An EcoRI sublcone from pCRII-avrRxv was cloned into pTM100, pCB302 and pBD. An EcoRI/HindIII fragment containing avrRxv from pUC118RXV3 [3] was cloned into pDSK519, creating pXVSC910, and into pVSP61, creating pLLM1.
Table 1.
Primers used to construct mutant variants of avrRxv by sequence overlap extension
| Primer Name | Primer sequence, 5′ to 3′a |
|---|---|
| EG1 | TGGATCCTCACGCATAGTCAGGCACATCGTAAGGGTAGGATTCTAAGGCGTGACG |
| EG2 | GCGGAAGCACTTAAGTCAATCGGTGGGGCTGTCATATTTTCTCTTGATTATGC |
| EG4 | TTCTAGATGTGCGACTCCATAAGAGTGCAATTCAGATCCATACAAAAAATGGT |
| EG5 | GCATAATCAAGAGAAAATATGACAGCCCCACCGATTGACTTAAGTGCTTCC |
| EG6 | TTCTAGATGTGCGACTCCATAAGAGTGCAATTCAG |
| EG7 | TGGATCCTCAGGATTCTAAGGCGTGACG |
| EG8 | TCAATCGGTGGGGCTGTCATATTTTCTCTTGAT |
| EG9 | AAAATATGACAGCCCCACCGATTGACTTAAG |
| EG10 | TGGATCCTCACGCATAGTCAGGCACATCGTAAGGGTAGGATTCTAAGGCGTGACGG |
| EG11 | GAAGACCCTAGGAGATGGGCTCGCGTCGCGTTCGACGTGCGCAAC |
| EG12 | TCGAACGCGACGCGAGCCCATCTCCTAGGGTCTTC |
| EG13 | TGGATCCTCAGGATTCTAAGGCGTGACGGATCTTTCGAGCTCT |
| EG14 | ACCCTAGGAGATGGGCTCGCGTCGCGTTCGACGTG |
| EG15 | CACGTCGAACGCGACGCGAGCCCATCTCCTAGGGT |
| EG16 | TGCCCTCTAGACGGCTGTGCGACTCCATAAGAGTGCAATTCAG |
| E200A-IL | CAATGCGATAATCGTCGTGT |
| E200A-IR | GATTATCGCATTGGCTCCTG |
TCTAGA XbaI site; GGATCC BamHI site; bold, altered nucleotide to change codon to alanine
2.5. Immunoblotting
To assess the stability of mutant proteins, immunoblots were performed on the catalytic domain mutants. Xcv was grown on NYG medium and resuspended to an OD600 of 2.0 with 100 mM MgCl2. A 20 μL aliquot was mixed 1:1 with 3× Laemmli buffer, boiled for 5 minutes and centrifuged at 14,000 × g for 5 minutes. Fifteen μL of the supernatant was fractionated by 8-12.5% SDS-PAGE and transferred onto a nitrocellulose membrane (BioRad, Hercules, CA, USA). Immunoblotted membranes were incubated with mouse monoclonal antibodies that recognize HA (Covance, Berkeley, CA, USA) according to manufacturer's instructions. Detection was performed using the EZ-ECL Chemiluminescence Detection Kit for HRP (Biological Industries, Kibbutz Beit Haemek, Isreal) or SuperSignal West Pico (Pierce Biotechnology, Rockford, IL, USA), according to manufacturer's protocols.
2.6. AvrRxv family phylogeny
Homologues of AvrRxv, PopP1 and YopP were identified with BLASTP [63]. Amino acid and nucleotide sequences were aligned using ClustalX version 1.83.1 [64]. To optimize the nucleotide sequence alignments, nucleotide triplets were manually aligned using the amino acid alignments as a guide using McClade 4 (Sinauer Associates, Inc., Sunderland, MA, USA). Gaps were maintained. The 5′- and 3′-coding sequences were split at the gap in the amino acid alignment of family members that precedes D173 in AvrRxv. Phylogenetic relationships of coding sequences were inferred by using PAUP*Ver.4.0b10 (Sinauer Associated, Inc.) to implement likelihood analyses. Maximum likelihood models were generated using ModelTest version 3.6 [65]. A Jukes-Cantor neighbor-joining tree [66] was created to estimate its parameters based on the best-fit model from ModelTest. These initial parameter estimates were used to begin an iterative maximum likelihood analysis using a neighbor-joining heuristic search until the tree parameters converged [67]. To evaluate statistical support for the tree branches, a bootstrap analysis using a heuristic search of 100 random stepwise addition replicates was executed [68].
In addition, the two tree topologies based on either end of the gene were compared using the Shimodaira–Hasegawa test [69] to test for significant divergence in tree lengths. This test was performed using RELL with 1000 bootstrap replicates and the results evaluated as a one-tailed test. The Likelihood Ratio Test (LRT) indicated that HKY + Γ was the most appropriate model for the 5′ dataset. This model allows for rate variation between transitions and transversions (Ti/Tv=0.922796), unequal base frequencies (A = 0.248985, C = 0.289080, G = 0.271095, T = 0.190841), and Γ distribution (α = 2.148966 across four categories). The maximum likelihood search using this model resulted in one tree (-lnL=13033.63456). For the 3′ dataset, the Likelihood Ratio Test (LRT) indicated that HKY + Γ was again the most appropriate model. This model allows for rate variation between transitions and transversions (Ti/Tv=0.985939), unequal base frequencies (A = 0.250419, C = 0.259215, G = 0.275177, T = 0.215188), and Γ distribution (α = 1.470186 across four categories). The maximum likelihood search using this model resulted in one tree (-lnL=15691.99289).]. All of the effector's N-termini and C-termini were also compared individually with BlastP to those of AvrRxv and YopP to further analyze the general evolution of the two termini. Results are expressed as % similarity, which takes into account conservative amino acid substitutions according to the BLOSUM62 algorithm [63].
3. Results
3.1. AvrRxv interacts with a 14-3-3 protein in the yeast two-hybrid system
To identify potential host factors that interact with AvrRxv, a cDNA library from tomato Hawaii 7998 was co-transformed with avrRxv in the yeast two-hybrid system system. The pBD-avrRxv construct did not autoactivate in yeast strain PJ69-4a, and the fusion protein was detected in yeast extracts by immunoblot analysis. The pBD-avrRxv bait construct was used to screen approximately 5×106 yeast cfu expressing clones of the Hawaii 7998 cDNA library for potential interactors. Several potential positives were identified in the primary screen and sequenced. More than half were ribosomal proteins that were not further characterized. The remaining potential interactors were re-transformed into yeast strain PJ69-4a containing the bait plasmid pBD-avrRxv and re-tested for interaction. One protein, ARI1 (AvrRxv interactor 1), continued to interact with AvrRxv in the secondary screens (Table 2). The ARI1 sequence is a 14-3-3 family member and is nearly identical to the previously cloned 14-3-3 protein from tomato, TFT9 (Genbank accession no. X98865) [51]. The pAD-Ari1 was tested for autoactivation by co-transforming it with either pBD or pBD-Lamin C. Neither the pBD and pAD-Ari1, nor the pBD-Lamin C and pAD-Ari1 plasmid pairs allowed growth of the yeast strain on selective media or induced β-galactosidase (Table 2). YopJ and YopP cloned as pBD fusions did not interact with ARI1. ARI1 was also tested for promiscuous interaction with the plant disease resistance gene product PTO, and a constitutively active mutant PTO Y207D [56]. No interaction was observed using PTO or PTO Y207D.
Table 2.
Yeast two-hybrid screen for proteins that interact with AvrRxv.
| Interactors | Interaction Analysis | |||
|---|---|---|---|---|
| Growth on mediumc | ||||
| Baita | Targetb | Adenine minus | Histidine minus | β-gal activityd |
| pBD-p53 | SV40 | + | + | 4.38 ± 0.02 |
| pBD-Lamin C | SV40 | - | - | 0.83 ± 0.20 |
| pBD-Lamin C | ARI1 | - | - | 0.00 ± 0.00 |
| pBD | ARI1 | - | - | 0.84 ± 0.12 |
| pBD-avrRxv | ARI1 | ++ | ++ | 61.44 ± 3 |
| pBD-avrRxv 3′Δ | ARI1 | ++ | ++ | 122.33 ± 9 |
| pBD-avrRxv 5′Δ | ARI1 | - | - | 0.94 ± 0.40 |
| pBD-avrRxv:Raf | ARI1 | ++ | ++ | 55.82 ± 7 |
| pBD-avrRxv:R18 | ARI1 | ± | ± | 8.35 ± 4 |
| pBD-avrRxv:Raf/R18 | ARI1 | ± | ± | 17.57 ± 5 |
Vector pBD contains the Gal4 DNA binding domain.
Target coding sequence constructs are in pAD, containing the activation domain.
Bait and target constructs co-transformed into yeast strain pJ69-4a; no growth, -; some slow growth, ±; good growth, +; very good growth, ++.
β-galactosidase activity in Miller units.
To localize the ARI1 interaction domain within the AvrRxv sequence, two avrRxv deletion constructs were tested for interaction. An N-terminal deletion of AvrRxv142-373, contained in pBD-avrRxv 5′Δ, did not interact with ARI1 (Table 2). The first 141 amino acids of AvrRxv are essential for interaction with ARI1. On the other hand, the C-terminal deletion of AvrRxv1-315 did interact. In fact, the β-galactosidase activity associated with the interaction of ARI1 with the C-terminal deletion AvrRxv1-315 in pBD-avrRxv 3′Δ was twice that of the full-length construct (Table 2). The C-terminal 58 amino acids of AvrRxv, which are C-terminal to the protease domain, are not only dispensable for interaction with ARII, but may normally inhibit the interaction.
3.2. Mutational analysis of 14-3-3 protein-binding sequence motifs in AvrRxv
Analysis of the AvrRxv sequence indicated that there are two domains in AvrRxv that resemble known 14-3-3 binding motifs, RSFDTP (called Raf domain) and MDIE (called R18 domain) [42,45]. To test if either the Raf or R18 domain is important for the AvrRxv and ARI1 interaction, mutant versions of AvrRxv were assayed for interaction with ARI1 in yeast. The altered AvrRxv-Raf in which the aspartic acid in the putative Raf domain (RSFDTP) was changed to an alanine (D147A), behaved no differently than the full length AvrRxv (Table 2). Whereas, AvrRxv-R18 in which the two negatively charged amino acids aspartic acid and glutamic acid in the R18 domain MDIE were changed to the positively charged residues asparagine and glutamine (D123N/E125Q), interacted significantly less based on growth phenotype and β-galactosidase activity (14% of wildtype activity). These results are consistent with results from the N-terminal deletion mutant AvrRxv142-373, which contains the Raf-like motif but lacks the R18 motif (Table 2). The AvrRxv Raf/R18 mutant containing all three mutations interacted less than wildtype, but more than the R18 mutant. Yeast strains containing the AvrRxv Raf/R18 mutant, compared to wildtype AvrRxv exhibited less growth on selective media and β-galactosidase activity (29% wildtype activity). Accordingly, neither AvrRxv-R18 nor AvrRxv–Raf/R18 induced HR when carried by Xcv or delivered by Agrobacterium (Table 3). However, AvrRxv-Raf was no different from wildtype AvrRxv. These results suggest that the ARI1 interaction, mediated by the R18 interaction domain in AvrRxv, is essential for AvrRxv's function in planta.
Table 3.
Response of tomato lines to inoculation with Xanthomonas and Agrobacterium containing plasmid-borne AvrRxv and variants thereof.
| Tomato Host Responsea | ||||||
|---|---|---|---|---|---|---|
| Xcv 89-1 | Agrobacterium | |||||
| AvrRxvb | Xcv plasmidc | Hawaii 7998 | Bonny Best/Money-maker | Agro plasmidd | Hawaii 7998 | Bonny Best/Money-maker |
| Wildtype | pTM100-avrRxv | HR | WS | pCB302-avrRxv | HR | NS |
| Rafe | pTM100-Raf | HR | WS | pCB302-Raf | HR | NS |
| R18f | pTM100-R18 | WS | WS | pCB302-R18 | NS | NS |
| Raf/R18 | pTM100-Raf/R18 | WS | WS | pCB302-Raf/R18 | NS | NS |
| Wildtype | pXVSC910 | HR | WS | pMD1-avrRxv | HR | NS |
| H180A | pDSK519-H180A | WS | WS | pMD1-H180A | NS | NS |
| E200A | pDSK519-E200A | WS | WS | pMD1-E200A | NS | NS |
| C244A | pDSK519-C244A | WS | WS | pMD1-C244A | NS | NS |
| E258A | pDSK519-E258A | HR | WS | pMD1-E258A | HR | NS |
Host responses are HR, hypersensitive response; WS, watersoaking; NS, no symptom.
avrRxv wildtype gene and mutants, with an HA tag, induced same host response as untagged gene.
Xcv carrying pTM100 or pDSK519 induced WS on Hawaii 7998 and Bonny Best.
Agrobacterium carrying pCB302 or pMD1 induced NS on Hawaii 7998 and Bonny Best.
Raf mutation in AvrRxv, D147A.
R18 mutation in AvrRxv, D123N/E125Q
3.3. AvrRxv and ARI1 physically interact in vitro
To confirm the yeast two-hybrid results, the interaction of ARI1 and AvrRxv was tested using affinity chromatography and immunoprecipitation. The c-myc epitope-tagged and 6XHis-tagged ARI1 in the pAD vector maintained their interaction in yeast with AvrRxv and AvrRxv-Flag, encoded in pBD-avrRxv and pBD-avrRxv-flag, respectively. The cloned avrRxv-flag was shown to maintain HR-inducing activity in Xcv as shown previously [9]. In vitro translated 35S-methionine labeled AvrRxv-Flag lysates were incubated with excess unlabeled 6XHis-ARI1-Myc and the mixture was affinity purified on Ni-NTA columns. AvrRxv-Flag proteins co-eluted with the 6XHis-ARI1-Myc fraction (Fig. 1). No AvrRxv-Flag was found in the eluant of the control column that lacked 6XHis-ARI1-Myc, although it was detected in the flow-through fraction of the control column. These results confirm the interaction of AvrRxv and ARI1.
Figure 1.
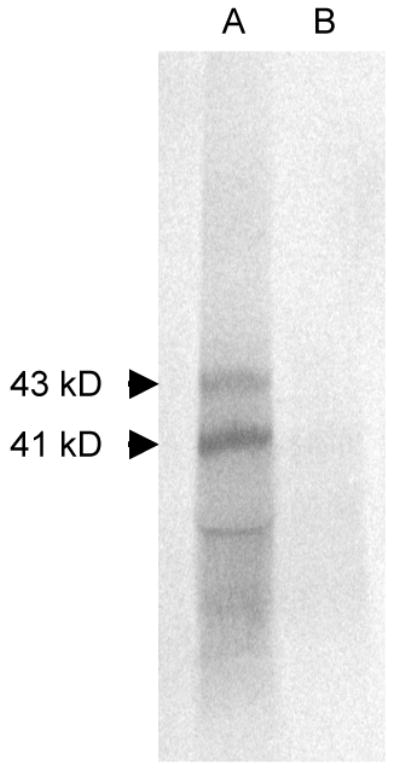
Affinity purification of AvrRxv with ARI1. AvrRxv labeled with 35S-methionine by in vitro translatation was affinity purified by interaction with ARI1-Myc containing a N-terminal 6X His tag. Lane A, elution of 35S-methionine labeled AvrRxv-Flag bound to 6XHis-ARI1-Myc from Ni-NTA column; Lane B, elution of 35S-methionine labeled AvrRxv-Flag alone from Ni-NTA column. Arrows mark two AvrRxv translation products.
An alternative approach to verify the interaction in vitro was used as well. Recombinant AvrRxv-Flag labeled with 35S-methionine was mixed with equal amounts of recombinant ARI1-Myc or negative control luciferase and immunoprecipitated with anti-Myc antibody and Dynabeads Protein G. Although there was cross reactivity of the AvrRxv-Flag with the Dynabeads Protein G, the intensity of the labeled protein band was four times greater in the AvrRxv-Flag and ARI1-Myc reaction, than in the negative controls AvrRxv-Flag alone or AvrRxv-Flag and luciferase reactions (data not shown). AvrRxv and ARI1 interact in vitro.
3.4. AvrRxv and YopP chimera has HR-inducing activity
AvrRxv homologues from Yersina were tested for activity in leaves of tomato lines Hawaii 7998 and Bonny Best (Table 4). AvrRxv induces a resistance, hypersensitive response (HR) on Hawaii and a susceptible, watersoaking (WS) response on Bonny Best, when carried by Xcv. Accordingly, when delivered by Agrobacterium, avrRxv induced HR on Hawaii, but not on Bonny Best. Both Yersinia homologues, YopJ and YopP were not recognized by either plant line with both methods of delivery. However, when a chimera was made between the N-terminal 173 amino acids of AvrRxv and the C-terminal 185 amino acids of YopP, an HR was induced on Hawaii 7998 (Table 4; Fig. 2). In contrast, a chimera between the coding sequence for the N-terminal 103 amino acids of YopP and the C-terminal 200 amino acids of AvrRxv, did not induce HR. It appears that the catalytic domain of AvrRxv family member YopP functions to induce the host response associated with AvrRxv, when the appropriate AvrRxv N-terminal domain is present.
Table 4.
Host responses to inoculation with Xanthomonas and Agrobacterium strains containing genes for AvrRxv, AvrRxv homologues and chimeras thereof.
| Tomato Host Responsea | |||||||
|---|---|---|---|---|---|---|---|
| Protein termini | Xcv 89-1 | Agrobacterium | |||||
| N | C | Xcv plasmid b | Hawaii 7998 | Bonny Best | Agro plasmidc | Hawaii 7998 | Bonny Best |
| AvrRxv | pTM100-avrRxv | HR | WS | pCB302-avrRxv | HR | NS | |
| YopP | pTM100-yopP | WS | WS | pCB302-yopP | NS | NS | |
| AvrRxv | YopP | pTM100-RY4 | HR | WS | pCB302-RY4 | HR | NS |
| YopP | AvrRxv | pTM100-YR3 | WS | WS | pCB302-YR3 | NS | NS |
Host responses are HR, hypersensitive response; WS, watersoaking; NS, no symptom.
Xcv 89-1 carrying pTM100 induced WS on Hawaii 7998 and Bonny Best.
Agrobacterium carrying pCB302 induced NS on Hawaii 7998 and Bonny Best.
Figure 2.

Response of tomato line Hawaii 7998 to Xcv 89-1 carrying genes encoding (A) AvrRxv; (B) N-AvrRxv1-172-YopP104-289-C; (C) N-YopP1-103-AvrRxv173-373-C; and (D) vector alone, at 6 days post-inoculation.
3.5. Phylogeny of coding sequence of C-terminus differs from that of N-terminus in the AvrRxv-family
Multiple alignments of AvrRxv family members indicate that the C-termini of the proteins have more overall similarity than the N-termini. When compared to AvrRxv, the plant pathogen effector C-termini are more similar (55% average similarity) than the N-termini (41% average similarity). Likewise the animal pathogen effector C-termini are also more similar to that of YopP (74% average similarity) than the N-termini (69% average similarity). Moreover, although the C-terminus of the plant pathogen effector AvrRxv has similarity to the C-termini of the animal effectors (43% average similarity) and the C-terminus of YopP has similarity to the C-termini of the plant effectors (30% average similarity), the N-termini in the cross-comparison do not (0-10% average similarity). The C-termini are more conserved than the N-termini.
Phylogenetic analyses were performed on the 5′- and 3′-coding sequences of family members. The Shimodaira-Hasegawa test showed that the maximum likelihood trees were very different (P=0.0001) between the 5′- and the 3′-ends of the coding sequences of AvrRxv family members. These results show that the two parts of the gene are evolving independently and in very different ways (Fig. 3).
Figure 3.
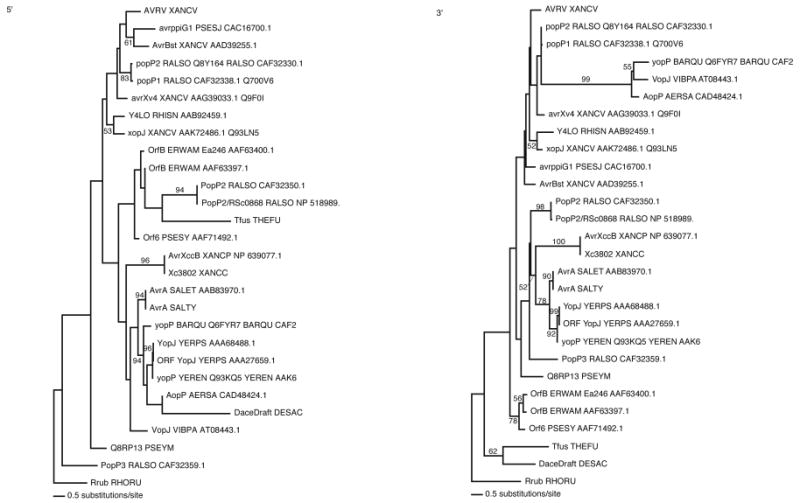
Phylogenetic analysis of the 5′- and 3′-ends of the coding sequences of AvrRxv family members. Phylogenetic relationships were inferred and maximum likelihood models were generated with neighbor-joining tree estimations and heuristic searching until the tree parameters converged. Statistical support for the tree branches was evaluated by bootstrap analysis using a heuristic search of 100 random stepwise addition replicates. The two tree topologies were shown to be statistically different using the Shimodaira–Hasegawa test for significant divergence in tree lengths.
3.6. Catalytic triad mutants are inactive
Wildtype avrRxv induces an HR in leaves of tomato line Hawaii 7998 when carried by Xcv 89-1, or when delivered by Agrobacterium-mediated transformation (Table 3) [3,9]. When any one of the three active site amino acid residues is mutated as in H180A, E200A, and C244A, HR inducing activity is lost when carried by Xcv 89-1 or delivered by Agrobacterium (Table 3). A mutation in an amino acid outside the active site has no effect on activity (E258A). The catalytic triad is essential for HR-inducing activity in our system.
3.7. Identification of the transcription initiation site
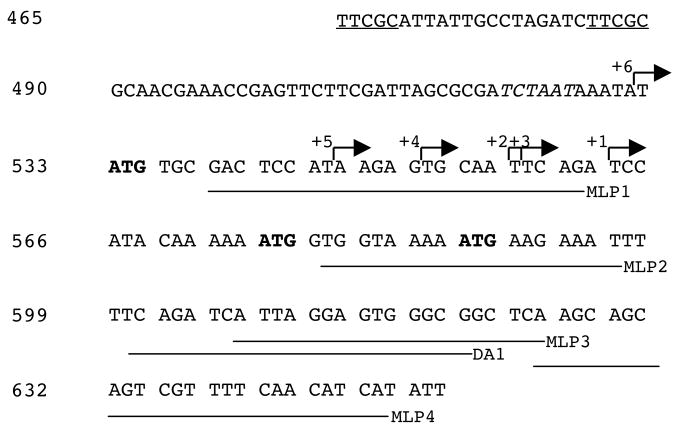
To identify the 5′-end of the avrRxv transcript, primer extension on RNA isolated from Xcv 89-1 (pXVSC910) was performed. The major extension products using MLP3, MLP4 and DA1 aligned with base 563 of the sequencing reaction (Fig. 4). There was no detectable extension product with primer MLP1. Minor start sites were also observed upstream of base 563 at 558, 557, 552, 546, and 530. No extension products were observed with RNA from the negative control strain Xcv 89-1 (pDSK519) or from Xcv 92-14 ± avrRxv. This transcription initiation site is 12 and 24 bases upstream of two start codons at 575 and 587, respectively, and 36 bases downstream of a -10 promoter motif (Fig. 4) [70].
Figure 4.
Sequence of region upstream of avrRxv with transcription initiation sites labeled. Primer extension was performed with primers MLP1, MLP2, DA1, MLP3 and MLP4 (indicated by horzontal lines). The 5′-end of extension products is indicated with arrows, with +1 being the major initiation site, and +2 through +6, progressively more minor. Potential translation start codons are indicated in bold. PIP-box sequence underlined; promoter -10 sequence italicized.
Discussion
Xcv uses the TTSS to translocate effector proteins into host cells, where they affect cellular physiology, causing disease in tomato and pepper. Seventeen effectors have been confirmed in Xcv [71], with only a few characterized for molecular or biochemical function. Several members of the YopJ/AvrRxv family of effectors have been shown to use a catalytic triad to target host proteins [8,13,22,25]. In this study, we used the yeast two-hybrid interaction screen to identify the protein(s) with which AvrRxv physically interacts in tomato. We have shown that AvrRxv interacts with a 14-3-3 protein, which we call ARI1 for AvrRxv interactor (Table 2). This result was supported by binding of recombinant AvrRxv to ARI1 via in vitro affinity chromatography (Fig. 1). ARI1 is nearly identical to the previously cloned 14-3-3 from tomato, TFT9 [51]. TFT9 is one of at least ten 14-3-3 proteins in tomato. Tft9 expression is induced in response to treatment with fusicoccin, a membrane depolarizing fungal toxin, but not in a resistance response to Avr9 [51]. Similar to ARI1 in this study, TFT9 is thought to play a role in defense signaling.
14-3-3 proteins bind their ligands in a conserved amphipathic groove [36]. Ligands are thought to bind to 14-3-3 proteins using three basic mechanisms [47], including the phosphorylation-independent manner via negatively charged residues coordinated by a basic cluster. The aspartate and glutamate residues of R18 peptide substitute for the phosphorylated residues in the phosphorylation-dependent mechanism. By mutagenesis of the coding sequence of AvrRxv, we demonstrated that although a putative Raf domain was not essential for both ARI1 binding or HR-inducing activity, a domain similar to the R18 domain was essential (Table 2). The AvrRxv-R18 mutant (D123N/E125Q), in which the acidic aspartate and glutamate residues were replaced by neutral amino acids, asparagines and glutamine, lost the ability to interact with ARI1 in yeast and lost HR-inducing activity, both when delivered by Xcv or Agrobacterium (Table 2; Table 3). Accordingly, an N-terminal deletion that removed the R18 domain (AvrRxv142-373) did not interact with ARI1 in yeast (Table 2). These results further support the assertion that the interaction with ARI1 is specific to its function in the signaling pathway in tomato in response to AvrRxv. As mentioned above, studies of the structure of particular 14-3-3 proteins and their ligands have identified different binding mechanisms [47]. The fact that mutation of the acidic residues in the R18 domain of AvrRxv resulted in loss in ARI1 binding in vitro and loss of HR-inducing activity, suggests that ARI1 may interact with AvrRxv by a basic cluster of amino acids within the amphipathic groove of ARI1. Similar to Zhang et al. [72], future experiments will include analysis of the impact on AvrRxv binding by mutagenesis of these basic residues in ARI1.
14-3-3 proteins are involved in a wide variety of protein interaction-mediated functions in plants [38,40,41]. In Arabidopsis, there are up to 15 members of the 14-3-3 family [73]. They are categorized by sequence into groups and isoforms. Based on its phylogenetic proximity to Arabidopsis protein general regulatory factor 9 (GRF9), ARI1 is a member of the μ isoform subfamily of the ε group (D group) of the phylogenetic tree [38]. Our study of ARI1's phylogeny, indicates that there are 18 members in ARI1's clade. The expression of five clade relatives is up-regulated in host responses to pathogens in tomato [51] and chickpea (Genbank accession no. ABQ95991) or to elicitors in poplar [50]. In barley, 14-3-3 proteins are upregulated by powdery mildew [74]. Affinity purification of barley 14-3-3 binding partners indicated that the second largest class of interactors was involved in defense, including three different disease NBS-LRR resistance proteins [75]. This suggests that there is a direct role of 14-3-3 in resistance gene-mediated resistance signaling. To assess the impact of ARI1 on the AvrRxv-specific HR in tomato, ARI1 will be silenced. Interaction partners downstream of ARI1 will be identified.
Although there are quite a few close relatives of ARI1, not much is known about their specific function. ARI1's second nearest relative is a 14-3-3 protein D75 in tobacco that binds to and potentially regulates the RSG transcriptional activator involved in GA responses [76]. The closest Arabidopsis relative GRF9 (G14μ) binds to the signal peptide of a thylakoid-targeted chloroplast precursor protein and is found in the chloroplast stroma, possibly suggesting a role in protein import [77]. A soybean 14-3-3 was up-regulated upon infection by P. syringae pv. glycinea [52]. The Arabidopsis relatives are localized in the chloroplast stroma and in the cytoplasm [77], and if that is generally true for all of ARI1's relatives, it rules out exclusive subcellular localization as a regulatory factor [38]. ARI1's precise role in the signaling response of tomato to the presence of AvrRxv remains to be discovered.
Interestingly, the C-terminal 58 amino acids of AvrRxv, which are C-terminal to the protease domain, are not only dispensable for interaction with ARII, but may normally inhibit the interaction (Table 2). It is possible that the C-terminus plays a guarding or regulatory function. Additional AvrRxv mutagenesis experiments will allow dissection of this possibility.
Our result demonstrating that ARI1 directly interacts with AvrRxv is similar to the interaction of the effector ExoS from P. aeruginosa with a 14-3-3 protein [45]. ExoS requires the host factor 14-3-3 for activation and binds to it in a phosphorylation-independent manner [72]. It is possible that ARI1 also enhances the enzymatic activity of AvrRxv. In addition to the 14-3-3/ExoS interaction, other bacterial pathogens encode proteins that complex with mammalian 14-3-3s [78]. Our work provides additional evidence that bacterial effectors and proteins recruit 14-3-3 proteins for their function within host cells. Interestingly, a YopJ mutant (DVE53-55NAQ) that turns out to be in the equivalent R18 domain is inactive [79]. Perhaps, deeper analysis of YopJ host interactions may uncover a 14-3-3 protein for full level of activation. The elucidation of the role of these interactions may allow the rational design of inhibitory molecules that interfere with widely conserved mechanisms of virulence.
A chimera of AvrRxv and YopP has HR-inducing activity in leaves of Hawaii 7998 (Table 4; Fig. 2). That along with the results that both Yersinia homologues, YopJ and YopP and a YopP-AvrRxv chimera were not able to induce an HR, have important implications for our understanding of structure and function relationships in the family. Despite their divergent origins, the catalytic domain of YopP is interchangeable with that of AvrRxv. This suggests a strong conservation of function in the C-terminal domains of YopJ family members as was demonstrated by YopJ function in inhibition of yeast MAPK pathway [80,81]. The functional conservation of catalytic triad residues supports this as well (Table 3) [8,13,23]. We find this to be the case in our phylogenetic analyses of the family, where the C-termini containing the catalytic triad are significantly more conserved than the N-termini. Remarkably, the coding sequences for the two termini appear to be evolving independently (Fig. 3). Even when the YopP effector is delivered directly to the host cell via Agrobacterium-mediated expression, without the AvrRxv N-terminus, it cannot function (Table 4). Along these lines, the family member AvrA from Salmonella, fails to complement YopJ activity [82]. In Xcv family members, TTSS signals are present in about the first 50 amino acids, comprising a translocation domain [12,34,83,84]. Based on our results, we predict that there is a third essential domain, the host-specific interaction domain that is C-terminal to the translocation domain and N-terminal to the protease domain. In AvrRxv the host-specific interaction domain contains the R18 motif, required for full level interaction with host factor ARI1 and HR-inducing activity. Analysis of the activity of other Xcv family members in tomato, and delineation of the translocation and host-specific interactions domains in them will allow testing of this prediction. The host-specific interaction domain may be highly specific to not only the host and particular target cells, but also to the protein target.
We have identified the transcription initiation site in avrRxv (Fig. 4), allowing more accurate prediction of the translation start codon. Based on alignment of the amino acid sequence of AvrRxv with that of the closely related family member AvrBsT [6], and the distance between other Xanthomonas transcription initiation sites and start codons [70], it is most likely the ATG at base 587 that serves as the start codon. The PIP-box that is 31 bases upstream of the -10 promoter motif has not yet been shown to function in HrpX control of expression of avrRxv [6]. Analysis of the genome of Xcv revealed that there are PIP-boxes upstream of coding sequences for genes that have no obvious association with pathogenicity and also, HrpX controlled genes that do not contain this motif [11,12,85]. More work has to be done to understand the role of the PIP-box.
In conclusion, we have shown that the AvrRxv bacterial effector recruits a 14-3-3 protein for its function within host cells. An interaction domain in AvrRxv was essential for this recruitment and associated function in host cells. Different domains in this family of effector proteins were shown to have evolved differently, suggesting host-specific adaptations are ongoing.
Acknowledgments
We thank Edward Moe for expert assistance in seed production. This work was supported by a NSF Cooperative Agreement BIR-8920216 to CEPRAP. T.R., M.Y., C.Q.M., M.C.W. were supported by NIH MBRS SCORE SO6 GM52533 (M.C.W.) and NIH AREA 5 R21 GM059022-01 (M.C.W.); T.R. and V.K. and research by NSF Cooperative Agreement BIR-8920216 (U.C.D. CEPRAP); A.A. by NIH NIGMS PREP 5 R25-GM64078 (S.F.S.U.); C.T. by NIH NIGMS MBRS-RISE MS/PhD 5 R25-GM59298 (S.F.S.U.); M.L. and E.E. by NIH NIGMS Bridges to the PhD 5 R25 GM48972 (S.F.S.U.); G.P-.B., E.G., L.L.M. by NIH NIGMS MARC 5 T34-GM08574 (S.F.S.U.); C.S.T. by NIH MORE CCSF/SFSU Bridges to the Baccalaureate (M.C.W.); the work by NIH RIMI P20 RR11805.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Jones JB, Lacy GH, Bouzar H, Stall RE, Schaad NW. Reclassification of the xanthomonads associated with bacterial spot disease of tomato and pepper. Syst Appl Microbiol. 2004;27:755–62. doi: 10.1078/0723202042369884. [DOI] [PubMed] [Google Scholar]
- 2.Vauterin L, Hoste B, Kersters K, Swings J. Reclassification of Xanthomonas. Int J Syst Bacteriol. 1995;45:472–89. [Google Scholar]
- 3.Whalen MC, Wang JF, Carland FM, Heiskell ME, Dahlbeck D, Minsavage GV, et al. Avirulence gene avrRxv from Xanthomonas campestris pv. vesicatoria specifies resistance on tomato line Hawaii 7998. Mol Plant-Microbe Interact. 1993;6:616–27. doi: 10.1094/mpmi-6-616. [DOI] [PubMed] [Google Scholar]
- 4.Yu ZH, Wang JF, Stall RE, Vallejos CE. Genomic localization of tomato genes that control a hypersensitive reaction to Xanthomonas campestris pv. vesicatoria (Doidge) Dye. Genetics. 1995;141:675–82. doi: 10.1093/genetics/141.2.675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Whalen MC, Stall RE, Staskawicz BJ. Characterization of a gene from a tomato pathogen determining hypersensitive resistance in non-host species and genetic analysis of this resistance in bean. Proc Natl Acad Sci USA. 1988;85:6743–7. doi: 10.1073/pnas.85.18.6743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ciesiolka LD, Hwin T, Gearlds JD, Minsavage GV, Saenz R, Bravo M, et al. Regulation of expression of avirulence gene avrRxv and identification of a family of host interaction factors by sequence analysis of avrBsT. Mol Plant-Microbe Interact. 1999;12:35–44. doi: 10.1094/MPMI.1999.12.1.35. [DOI] [PubMed] [Google Scholar]
- 7.Hardt WD, Galan JE. A secreted Samonella protein with homology to an avirulence determinant of plant pathogenic bacteria. Proc Natl Acad Sci USA. 1997;94:9887–92. doi: 10.1073/pnas.94.18.9887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Orth K, Xu Z, Mudgett MB, Bao ZQ, Palmer LE, Bliska JB, et al. Disruption of signaling by Yersinia effector YopJ, a ubiquitin-like protein protease. Science. 2000;290:1594–7. doi: 10.1126/science.290.5496.1594. [DOI] [PubMed] [Google Scholar]
- 9.Rossier O, Wengelnik K, Hahn K, Bonas U. The Xanthomonas Hrp type III system secretes proteins from plant and mammalian bacterial pathogens. Proc Natl Acad Sci USA. 1999;96:9368–73. doi: 10.1073/pnas.96.16.9368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Astua-Monge G, Minsavage GV, Stall RE, Vallejos CE, Davis MJ, Jones JB. Xv4-Avrxv4: a new gene-for-gene interaction identified between Xanthomonas campestris pv. vesicatoria race T3 and wild tomato relative Lycopersicon pennellii. Mol Plant Microbe Interact. 2000;1312:1346–55. doi: 10.1094/MPMI.2000.13.12.1346. [DOI] [PubMed] [Google Scholar]
- 11.Noël L, Thieme F, Nennstiel D, Bonas U. Two novel type III-secreted proteins of Xanthomonas campestris pv. vesicatoria are encoded within the hrp pathogenicity island. J Bacteriol. 2002;184:1340–8. doi: 10.1128/JB.184.5.1340-1348.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Noël L, Thieme F, Gäbler J, Büttner D, Bonas U. XopC and XopJ, two novel type III effector proteins from Xanthomonas campestris pv. vesicatoria. J Bacteriol. 2003;185:7092–102. doi: 10.1128/JB.185.24.7092-7102.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Roden J, Eardley L, Hotson A, Cao Y, Mudgett MB. Characterization of the Xanthomonas AvrXv4 effector, a SUMO protease translocated into plant cells. Mol Plant Microbe Interact. 2004;17:633–43. doi: 10.1094/MPMI.2004.17.6.633. [DOI] [PubMed] [Google Scholar]
- 14.Alfano JR, Charkowski AO, Deng WL, Badel JL, Petnicki-Ocwieja T, van Dijk K, et al. The Pseudomonas syringae Hrp pathogenicity island has a tripartite mosaic structure composed of a cluster of type III secretion genes bounded by exchangeable effector and conserved effector loci that contribute to parasitic fitness and pathogenicity in plants. Proc Natl Acad Sci USA. 2000;97:4856–61. doi: 10.1073/pnas.97.9.4856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Arnold DL, Jackson RW, Fillingham AJ, Goss SC, Taylor JD, Mansfield JW, Vivian A. Highly conserved sequences flank avirulence genes: isolation of novel avirulence genes from Pseudomonas syringae pv. pisi. Microbiology. 2001;147:1171–82. doi: 10.1099/00221287-147-5-1171. [DOI] [PubMed] [Google Scholar]
- 16.Deng WL, Rehm AH, Charkowski AO, Rojas CM, Collmer A. Pseudomonas syringae exchangeable effector loci: sequence diversity in representative pathovars and virulence function in P. syringae pv. syringae B728a. J Bacteriol. 2003;185:2592–602. doi: 10.1128/JB.185.8.2592-2602.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Deslandes L, Olivier J, Peeters N, Feng DX, Khounlotham M, Boucher C, et al. Physical interaction between RRS1-R, a protein conferring resistance to bacterial wilt, and PopP2, a type III effector targeted to the plant nucleus. Proc Natl Acad Sci USA. 2003;100:8024–9. doi: 10.1073/pnas.1230660100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lavie M, Shillington E, Eguiluz C, Grimsley N, Boucher C. PopP1, a new member of the YopJ/AvrRxv family of type III effector proteins, acts as a host-specificity factor and modulates aggressiveness of Ralstonia solanacearum. Mol Plant-Microbe Interact. 2002;15:1058–68. doi: 10.1094/MPMI.2002.15.10.1058. [DOI] [PubMed] [Google Scholar]
- 19.Freiberg C, Fellay R, Bairoch A, Broughton WJ, Rosenthal A, Perret X. Molecular basis of symbiosis between Rhizobium and legumes. Nature. 1997;387:394–401. doi: 10.1038/387394a0. [DOI] [PubMed] [Google Scholar]
- 20.Galyov EE, Håkansson S, Forsberg A, Wolf-Watz H. A secreted protein kinase of Yersinia pseudotuberculosis is an indispensable virulence determinant. Nature. 1993;361:730–2. doi: 10.1038/361730a0. [DOI] [PubMed] [Google Scholar]
- 21.Mills SD, Boland A, Sory MP, van der Smissen P, Kerbourch C, Finlay BB, et al. Yersinia enterocolitica induces apoptosis in macrophages by a process requiring functional type III secretion and translocation mechanisms and involving YopP, presumably acting as an effector protein. Proc Natl Acad Sci USA. 1997;94:12638–43. doi: 10.1073/pnas.94.23.12638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Trosky JE, Li Y, Mukherjee S, Keitany G, Ball H, Orth K. VopA inhibits ATP binding by acetylating the catalytic loop of MAPK kinases. J Biol Chem. 2007;282:34299–305. doi: 10.1074/jbc.M706970200. [DOI] [PubMed] [Google Scholar]
- 23.Bonshtien A, Lev A, Gibly A, Debbie P, Avni A, Sessa G. Molecular properties of the Xanthomonas AvrRxv effector and global transcriptional changes determined by its expression in resistant tomato plants. Mol Plant Microbe Interact. 2005;18:300–10. doi: 10.1094/MPMI-18-0300. [DOI] [PubMed] [Google Scholar]
- 24.Thieme F, Szczesny R, Urban A, Kirchner O, Hause G, Bonas U. New type III effectors from Xanthomonas campestris pv. vesicatoria trigger plant reactions dependent on a conserved N-myristoylation motif. Mol Plant Microbe Interact. 2007;20:1250–61. doi: 10.1094/MPMI-20-10-1250. [DOI] [PubMed] [Google Scholar]
- 25.Mukherjee S, Keitany G, Li Y, Wang Y, Ball HL, Goldsmith EJ, Orth K. Yersinia YopJ acetylates and inhibits kinase activation by blocking phosphorylation. Science. 2006;312:1211–4. doi: 10.1126/science.1126867. [DOI] [PubMed] [Google Scholar]
- 26.Hotson A, Mudgett MB. Cysteine proteases in phytopathogenic bacteria: identification of plant targets and activation of innate immunity. Curr Opin Plant Biol. 2004;7:384–90. doi: 10.1016/j.pbi.2004.05.003. [DOI] [PubMed] [Google Scholar]
- 27.Orth K. Function of the Yersinia effector YopJ. Curr Opin Microbiol. 2002;5:38–43. doi: 10.1016/s1369-5274(02)00283-7. [DOI] [PubMed] [Google Scholar]
- 28.Monack DM, Mecsas J, Ghori N, Falkow S. Yersinia signals macrophages to undergo apoptosis and YopJ is necessary for this cell death. Proc Natl Acad Sci USA. 1997;94:10385–90. doi: 10.1073/pnas.94.19.10385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mukherjee S, Hao YH, Orth K. A newly discovered post-translational modification--the acetylation of serine and threonine residues. Trends Biochem Sci. 2007;32:210–6. doi: 10.1016/j.tibs.2007.03.007. [DOI] [PubMed] [Google Scholar]
- 30.Orth K, Palmer LE, Bao ZQ, Stewart S, Rudolph AE, Bliska JB, et al. Inhibition of the mitogen-activated protein kinase kinase superfamily by a Yersinia effector. Science. 1999;285:1920–3. doi: 10.1126/science.285.5435.1920. [DOI] [PubMed] [Google Scholar]
- 31.Palmer LE, Hobbie S, Galán JE, Bliska JB. YopJ of Yersinia pseudotuberculosis is required for the inhibition of macrophage TNF-alpha production and downregulation of the MAP kinases p38 and JNK. Mol Microbiol. 1998;27:953–65. doi: 10.1046/j.1365-2958.1998.00740.x. [DOI] [PubMed] [Google Scholar]
- 32.Palmer LE, Pancetti AR, Greenberg S, Bliska JB. YopJ of Yersinia spp. is sufficient to cause downregulation of multiple mitogen-activated protein kinases in eukaryotic cells. Infect Immun. 1999;67:708–16. doi: 10.1128/iai.67.2.708-716.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhou H, Monack DM, Kayagaki N, Wertz I, Yin J, Wolf B, et al. Yersinia virulence factor YopJ acts as a deubiquitinase to inhibit NF-kappa B activation. J Exp Med. 2005;202:1327–32. doi: 10.1084/jem.20051194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Cunnac S, Wilson A, Nuwer J, Kirik A, Baranage G, Mudgett MB. A conserved carboxylesterase is a suppressor of avrBsT-elicited resistance in Arabidopsis. Plant Cell. 2007;19:688–705. doi: 10.1105/tpc.106.048710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Deslandes L, Pileur F, Liaubet L, Camut S, Can C, Williams K, et al. Genetic characterization of RRS1, a recessive locus in Arabidopsis thaliana that confers resistance to the bacterial soilborne pathogen Ralstonia solanacearum. Mol Plant Microbe Interact. 1998;11:659–67. doi: 10.1094/MPMI.1998.11.7.659. [DOI] [PubMed] [Google Scholar]
- 36.Dougherty MK, Morrison DK. Unlocking the code of 14-3-3. J Cell Sci. 2004;117:1875–84. doi: 10.1242/jcs.01171. [DOI] [PubMed] [Google Scholar]
- 37.Ferl RJ. 14-3-3 proteins and signal transduction. Annu Rev Plant Physiol Plant Mol Biol. 1996;47:49–73. doi: 10.1146/annurev.arplant.47.1.49. [DOI] [PubMed] [Google Scholar]
- 38.Sehnke PC, Rosenquist M, Alsterfjord M, DeLille J, Sommarin M, Larsson C, et al. Evolution and isoform specificity of plant 14-3-3 proteins. Plant Mol Biol. 2002;50:1011–8. doi: 10.1023/a:1021289127519. [DOI] [PubMed] [Google Scholar]
- 39.Milne FC, Moorhead G, Pozuelo Rubio M, Wong B, Kulma A, Harthill JE, et al. Affinity purification of diverse plant and human 14-3-3-binding partners. Biochem Soc Trans. 2002;30:379–81. doi: 10.1042/bst0300379. [DOI] [PubMed] [Google Scholar]
- 40.Roberts MR. 14-3-3 proteins find new partners in plant cell signaling. Trends Plant Sci. 2003;8:218–23. doi: 10.1016/S1360-1385(03)00056-6. [DOI] [PubMed] [Google Scholar]
- 41.Schoonheim PJ, Veiga H, Pereira Dda C, Friso G, van Wijk KJ, de Boer AH. A comprehensive analysis of the 14-3-3 interactome in barley leaves using a complementary proteomics and two-hybrid approach. Plant Physiol. 2007;143:670–83. doi: 10.1104/pp.106.090159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang B, Yang H, Liu YC, Jelinek T, Zhang L, Ruoslahti E, Fu H. Isolation of high-affinity peptide antagonists of 14-3-3 proteins by phage display. Biochem. 1999;38:12499–504. doi: 10.1021/bi991353h. [DOI] [PubMed] [Google Scholar]
- 43.Hallberg B. Exoenzyme S binds its cofactor 14-3-3 through a non-phosphorylated motif. Biochem Soc Trans. 2002;30:401–5. doi: 10.1042/bst0300401. [DOI] [PubMed] [Google Scholar]
- 44.Henriksson ML, Francis MS, Peden A, Aili M, Stefansson K, Palmer R, Aitken A, Hallberg B. A nonphosphorylated 14-3-3 binding motif on exoenzyme S that is functional in vivo. Eur J Biochem. 2002;269:4921–9. doi: 10.1046/j.1432-1033.2002.03191.x. [DOI] [PubMed] [Google Scholar]
- 45.Masters SC, Pederson KJ, Zhang L, Barbieri JT, Fu H. Interaction of 14-3-3 with a nonphosphorylated protein ligand, exoenzyme S of Pseudomonas aeruginosa. Biochem. 1999;38:5216–21. doi: 10.1021/bi982492m. [DOI] [PubMed] [Google Scholar]
- 46.Zhai J, Lin H, Shamim M, Schlaepfer WW, Cañete-Soler R. Identification of a novel interaction of 14-3-3 with p190RhoGEF. J Biol Chem. 2001;276:41318–24. doi: 10.1074/jbc.M107709200. [DOI] [PubMed] [Google Scholar]
- 47.Ottmann C, Yasmin L, Weyand M, Veesenmeyer JL, Diaz MH, Palmer RH, et al. Phosphorylation-independent interaction between 14-3-3 and exoenzyme S: from structure to pathogenesis. EMBO J. 2007;26:902–13. doi: 10.1038/sj.emboj.7601530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yasmin L, Jansson AL, Panahandeh T, Palmer RH, Francis MS, Hallberg B. Delineation of exoenzyme S residues that mediate the interaction with 14-3-3 and its biological activity. FEBS J. 2006;273:638–46. doi: 10.1111/j.1742-4658.2005.05100.x. [DOI] [PubMed] [Google Scholar]
- 49.Brandt J, Thordal-Christensen H, Vad K, Gregersen PL, Collinge DB. A pathogen-induced gene of barley encodes a protein showing high similarity to a protein kinase regulator. Plant J. 1992;2:815–20. [PubMed] [Google Scholar]
- 50.Lapointe G, Luckevich MD, Cloutier M, Séguin A. 14-3-3 gene family in hybrid poplar and its involvement in tree defence against pathogens. J Exp Bot. 2001;52:1331–8. [PubMed] [Google Scholar]
- 51.Roberts MR, Bowles DJ. Fusicoccin, 14-3-3 proteins, and defense responses in tomato plants. Plant Physiol. 1999;119:1243–50. doi: 10.1104/pp.119.4.1243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Seehaus K, Tenhaken R. Cloning of genes by mRNA differential display induced during the hypersensitive reaction of soybean after inoculation with Pseudomonas syringae pv. glycinea. Plant Mol Biol. 1998;38:1225–34. doi: 10.1023/a:1006036827841. [DOI] [PubMed] [Google Scholar]
- 53.Yan J, Wang J, Zhang H. An ankyrin repeat-containing protein plays a role in both disease resistance and antioxidation metabolism. Plant J. 2002;29:193–202. doi: 10.1046/j.0960-7412.2001.01205.x. [DOI] [PubMed] [Google Scholar]
- 54.Rosenquist M. 14-3-3 proteins in apoptosis. Braz J Med Biol Res. 2003;36:403–8. doi: 10.1590/s0100-879x2003000400001. [DOI] [PubMed] [Google Scholar]
- 55.Morales CQ, Posada J, Macneale E, Franklin D, Rivas I, Bravo M, et al. Functional analysis of the early chlorosis factor gene. Mol Plant Microbe Interact. 2005;18:477–86. doi: 10.1094/MPMI-18-0477. [DOI] [PubMed] [Google Scholar]
- 56.Rathjen JP, Chang JH, Staskawicz BJ, Michelmore RW. Constitutively active Pto induces a Prf-dependent hypersensitive response in the absence of avrPto. EMBO J. 1999;18:3232–40. doi: 10.1093/emboj/18.12.3232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.VanLarebeke N, Engler G, Holsters M, Van den Elsacker S, Zaenen I, Schilperoort RA, et al. Large plasmid in Agrobacterium tumefaciens essential for crown gall inducing ability. Nature. 1984;252:169–70. doi: 10.1038/252169a0. [DOI] [PubMed] [Google Scholar]
- 58.Ausubel F, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, et al. Short protocols in molecular biology. 5th. New Jersey: Wiley; 2002. [Google Scholar]
- 59.Keen NT, Tamaki S, Kobayashi D, Trollinger D. Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene. 1988;70:191–7. doi: 10.1016/0378-1119(88)90117-5. [DOI] [PubMed] [Google Scholar]
- 60.Michiels T, Cornelis GR. Secretion of hybrid proteins by the Yersinia Yop export system. J Bacteriol. 1991;173:1677–85. doi: 10.1128/jb.173.5.1677-1685.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Xiang C, Han P, Lutziger I, Wang K, Oliver DJ. A mini binary vector series for plant transformation. Plant Mol Biol. 1999;40:711–7. doi: 10.1023/a:1006201910593. [DOI] [PubMed] [Google Scholar]
- 62.Tai TH, Dahlbeck D, Clark ET, Gajiwala P, Pasion R, Whalen MC, et al. Expression of the Bs2 pepper gene confers resistance to bacterial spot disease in tomato. Proc Natl Acad Sci USA. 199;96:14153–8. doi: 10.1073/pnas.96.24.14153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 64.Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research. 1997;24:4876–82. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Posada D, Crandall KA. Modeltest: testing the model of DNA substitution. Bioinformatics. 1998;14:817–8. doi: 10.1093/bioinformatics/14.9.817. [DOI] [PubMed] [Google Scholar]
- 66.Jukes TH, Cantor DR. Evolution of protein molecules. In: Nunro HN, editor. Mammalian protein metabolism. New York: Academic Press; 1969. pp. 21–132. [Google Scholar]
- 67.Swofford DL, Sullivan J. Phylogenetic inference using parsimony and maximum likelihood using PAUP*. In: Salemi M, Vandamme AM, editors. The Phylogenetic Handbook: A Practical Approach to DNA and Protein Phylogeny. Cambridge: Cambridge Univ. Press; 2003. pp. 160–96. [Google Scholar]
- 68.Felenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–91. doi: 10.1111/j.1558-5646.1985.tb00420.x. [DOI] [PubMed] [Google Scholar]
- 69.Shimodaira H, Hasegawa M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol. 1999;16:1114–6. [Google Scholar]
- 70.Koebnik K, Kruger A, Thieme F, Urban A, Bonas U. Specific binding of the Xanthomonas campestris pv. vesicatoria AraC-type transcriptional activator HrpX to plant-inducible promoter boxes. J Bacteriol. 2006;188:7653–60. doi: 10.1128/JB.00795-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Gürlebeck D, Thieme F, Bonas U. Type III effector proteins from the plant pathogen Xanthomonas and their role in the interaction with the host plant. J Plant Physiol. 2006;163:233–55. doi: 10.1016/j.jplph.2005.11.011. [DOI] [PubMed] [Google Scholar]
- 72.Zhang L, Wang H, Masters SC, Wang B, Barbieri JT, Fu H. Residues of 14-3-3 zeta required for activation of exoenzyme S of Pseudomonas aeruginosa. Biochem. 1999;38:12159–64. doi: 10.1021/bi991019l. [DOI] [PubMed] [Google Scholar]
- 73.DeLille JM, Sehnke PC, Ferl RJ. The Arabidopsis 14-3-3 family of signaling regulators. Plant Physiol. 2001;126:35–8. doi: 10.1104/pp.126.1.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Finni C, Andersen CH, Borch J, Gjetting S, Christensen AB, de Boer AH, et al. Do 14-3-3 proteins and plasma membrane H+-ATPases interact in the barley epidermis in response to the barley powdery mildew fungus? Plant Mol Biol. 2002;49:137–47. doi: 10.1023/a:1014938417267. [DOI] [PubMed] [Google Scholar]
- 75.Alexander RD, Morris PC. A proteomic analysis of 14-3-3 binding proteins from developing barley grains. Proteomics. 2006;6:1886–96. doi: 10.1002/pmic.200500548. [DOI] [PubMed] [Google Scholar]
- 76.Igarashi D, Ishida S, Fukazawa J, Takahashi Y. 14-3-3 proteins regulate intracellular localization of the bZIP transcriptional activator RSG. Plant Cell. 2001;13:2483–97. doi: 10.1105/tpc.010188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Sehnke PC, Henry R, Cline K, Ferl RJ. Interaction of a plant 14-3-3 protein with the signal peptide of a thylakoid-targeted chloroplast precursor protein and the presense of 14-3-3 isoforms in the chloroplast stroma. Plant Physiol. 2000;122:235–41. doi: 10.1104/pp.122.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Scidmore MA, Hackstadt T. Mammalian 14-3-3beta associates with the Chlamydia trachomatis inclusion membrane via its interaction with IncG. Mol Microbiol. 2001;39:1638–50. doi: 10.1046/j.1365-2958.2001.02355.x. [DOI] [PubMed] [Google Scholar]
- 79.Schesser K, Spiik AK, Dukuzumuremyi JM, Neurath MF, Pettersson S, Wolf-Watz H. The yopJ locus is required for Yersinia-mediated inhibition of NF-kappaB activation and cytokine expression: YopJ contains a eukaryotic SH2-like domain that is essential for its repressive activity. Mol Microbiol. 1998;28:1067–79. doi: 10.1046/j.1365-2958.1998.00851.x. [DOI] [PubMed] [Google Scholar]
- 80.Hao YH, Wang Y, Burdette D, Mukherjee S, Keitany G, Goldsmith E, et al. Structural Requirements for Yersinia YopJ Inhibition of MAP Kinase Pathways. PLoS ONE. 2008;3:e1375. doi: 10.1371/journal.pone.0001375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Yoon S, Liu Z, Eyobo Y, Orth K. Yersinia effector YopJ inhibits yeast MAPK signaling pathways by an evolutionarily conserved mechanism. J Biol Chem. 2003;278:2131–5. doi: 10.1074/jbc.M209905200. [DOI] [PubMed] [Google Scholar]
- 82.Schesser K, Dukuzumuremyi JM, Cilio C, Borg S, Wallis TS, Pettersson S, et al. The Salmonella YopJ-homologue AvrA does not possess YopJ-like activity. Microbial Pathogenesis. 2000;28:59–70. doi: 10.1006/mpat.1999.0324. [DOI] [PubMed] [Google Scholar]
- 83.Mudgett MB, Chesnokova O, Dahlbeck D, Clark ET, Rossier O, Bonas U, et al. Molecular signals required for type III secretion and translocation of the Xanthomonas campestris AvrBs2 protein to pepper plants. Proc Natl Acad Sci USA. 2000;97:13324–9. doi: 10.1073/pnas.230450797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Roden JA, Belt B, Ross JB, Tachibana T, Vargas J, Mudgett MB. A genetic screen to isolate type III effectors translocated into pepper cells during Xanthomonas infection. Proc Natl Acad Sci USA. 2004;101:16624–9. doi: 10.1073/pnas.0407383101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Noël L, Thieme F, Nennstiel D, Bonas U. cDNA-AFLP analysis unravels a genome-wide hrpG-regulon in the plant pathogen Xanthomonas campestris pv. vesicatoria. Mol Microbiol. 2001;41:1271–81. doi: 10.1046/j.1365-2958.2001.02567.x. [DOI] [PubMed] [Google Scholar]