Abstract
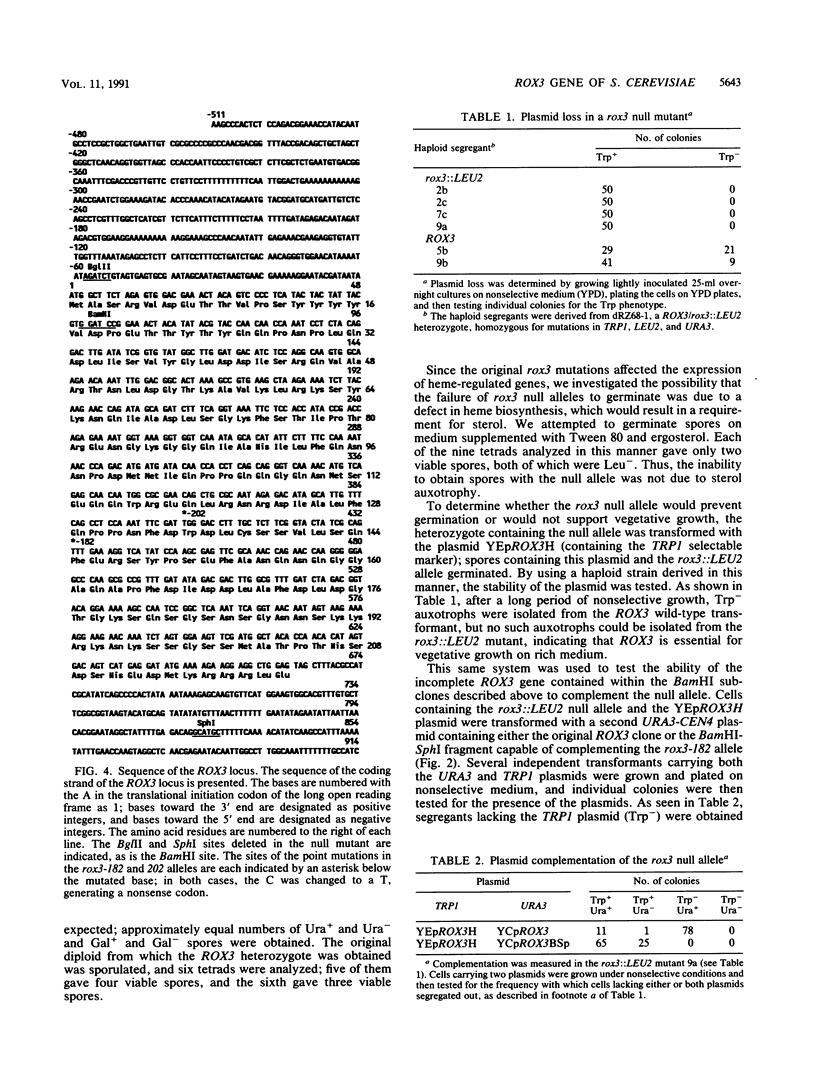
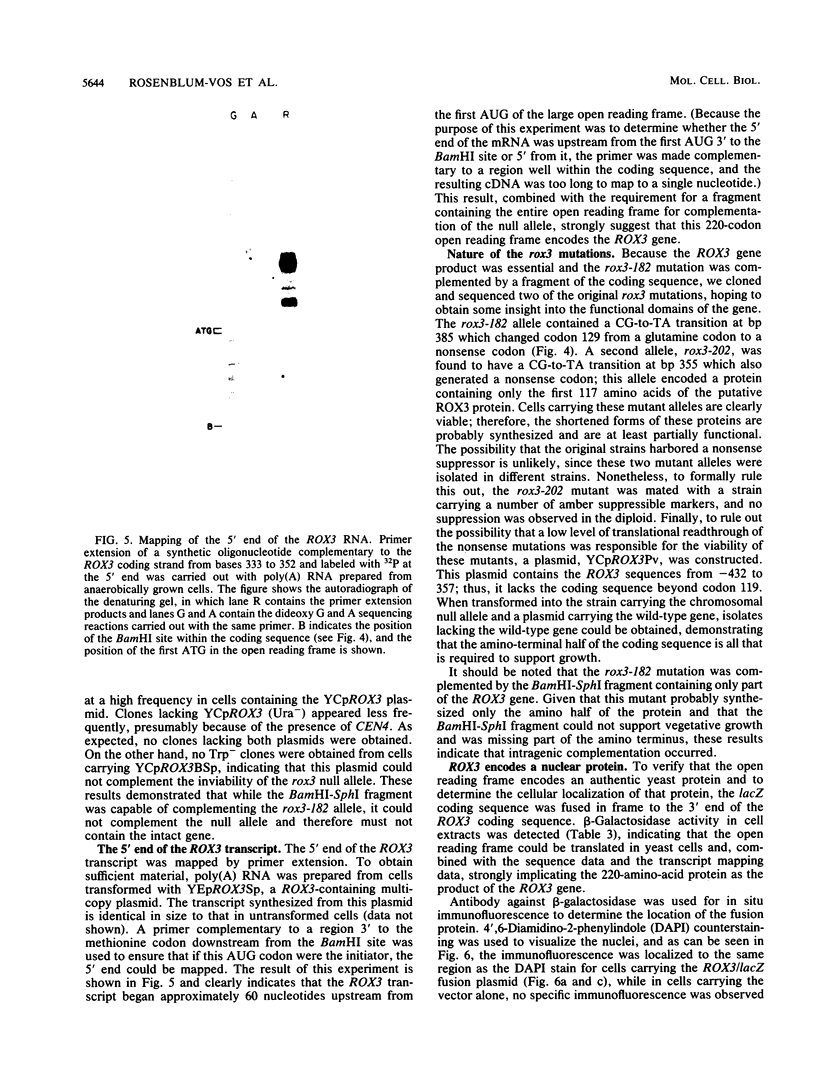
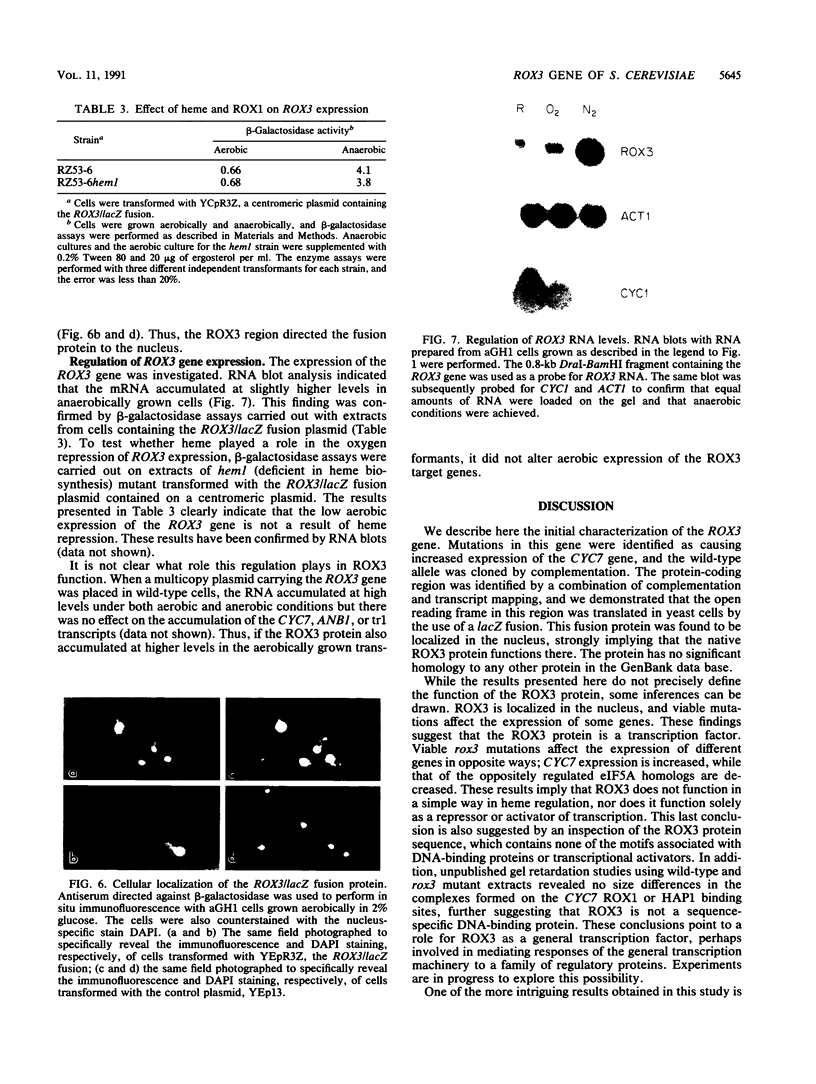
The ROX3 gene was identified during a hunt for mutants with increased expression of the heme-regulated CYC7 gene, which encodes the minor species of cytochrome c in the yeast Saccharomyces cerevisiae. The rox3 mutants caused a 10-fold increase in CYC7 expression both in the presence and absence of heme, had slightly increased anaerobic expression of the heme-activated CYC1 gene, and caused decreases in the anaerobic expression of the heme-repressed ANB1 gene and the aerobic expression of its heme-induced homolog. The wild-type ROX3 gene was cloned, and the sequence indicated that it encodes a 220-amino-acid protein. This protein is essential; deletion of the coding sequence was lethal. The coding sequence for beta-galactosidase was fused to the 3' end of the ROX3 coding sequence, and the fusion product was found to be localized in the nucleus, strongly suggesting that the wild-type protein carries out a nuclear function. Mutations in the rox3 gene showed an interesting pattern of intragenic complementation. A deletion of the 5' coding region complemented a nonsense mutation at codon 128 but could not prevent the lethality of the null mutation. These results suggest that the amino-terminal domain is required for an essential function, while the carboxy-terminal domain can be supplied in trans to achieve the wild-type expression of CYC7. Finally, RNA blots demonstrated that the ROX3 mRNA was expressed at higher levels anaerobically but was not subject to heme repression. The nuclear localization and the lack of viability of null mutants suggest that the ROX3 protein is a general regulatory factor.
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Broach J. R., Strathern J. N., Hicks J. B. Transformation in yeast: development of a hybrid cloning vector and isolation of the CAN1 gene. Gene. 1979 Dec;8(1):121–133. doi: 10.1016/0378-1119(79)90012-x. [DOI] [PubMed] [Google Scholar]
- Chattoo B. B., Sherman F., Azubalis D. A., Fjellstedt T. A., Mehnert D., Ogur M. Selection of lys2 Mutants of the Yeast SACCHAROMYCES CEREVISIAE by the Utilization of alpha-AMINOADIPATE. Genetics. 1979 Sep;93(1):51–65. doi: 10.1093/genetics/93.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clavilier L., Péré G., Slonimski P. P. Mise en évidence de plusieurs loci indépendants impliqués dans la synthèse de l'iso-2-cytochrome c chez la levure. Mol Gen Genet. 1969;104(2):195–218. doi: 10.1007/BF00272801. [DOI] [PubMed] [Google Scholar]
- Downie J. A., Stewart J. W., Brockman N., Schweingruber A. M., Sherman F. Structural gene for yeast iso-2-cytochrome c. J Mol Biol. 1977 Jun 25;113(2):369–384. doi: 10.1016/0022-2836(77)90147-4. [DOI] [PubMed] [Google Scholar]
- Forsburg S. L., Guarente L. Identification and characterization of HAP4: a third component of the CCAAT-bound HAP2/HAP3 heteromer. Genes Dev. 1989 Aug;3(8):1166–1178. doi: 10.1101/gad.3.8.1166. [DOI] [PubMed] [Google Scholar]
- Gietz R. D., Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene. 1988 Dec 30;74(2):527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- Guarente L., Mason T. Heme regulates transcription of the CYC1 gene of S. cerevisiae via an upstream activation site. Cell. 1983 Apr;32(4):1279–1286. doi: 10.1016/0092-8674(83)90309-4. [DOI] [PubMed] [Google Scholar]
- Hahn S., Guarente L. Yeast HAP2 and HAP3: transcriptional activators in a heteromeric complex. Science. 1988 Apr 15;240(4850):317–321. doi: 10.1126/science.2832951. [DOI] [PubMed] [Google Scholar]
- Healy A. M., Helser T. L., Zitomer R. S. Sequences required for transcriptional initiation of the Saccharomyces cerevisiae CYC7 genes. Mol Cell Biol. 1987 Oct;7(10):3785–3791. doi: 10.1128/mcb.7.10.3785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodge M. R., Kim G., Singh K., Cumsky M. G. Inverse regulation of the yeast COX5 genes by oxygen and heme. Mol Cell Biol. 1989 May;9(5):1958–1964. doi: 10.1128/mcb.9.5.1958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klebe R. J., Harriss J. V., Sharp Z. D., Douglas M. G. A general method for polyethylene-glycol-induced genetic transformation of bacteria and yeast. Gene. 1983 Nov;25(2-3):333–341. doi: 10.1016/0378-1119(83)90238-x. [DOI] [PubMed] [Google Scholar]
- Kunkel T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc Natl Acad Sci U S A. 1985 Jan;82(2):488–492. doi: 10.1073/pnas.82.2.488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laz T. M., Pietras D. F., Sherman F. Differential regulation of the duplicated isocytochrome c genes in yeast. Proc Natl Acad Sci U S A. 1984 Jul;81(14):4475–4479. doi: 10.1073/pnas.81.14.4475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemontt J. F., Fugit D. R., Mackay V. L. Pleiotropic Mutations at the TUP1 Locus That Affect the Expression of Mating-Type-Dependent Functions in SACCHAROMYCES CEREVISIAE. Genetics. 1980 Apr;94(4):899–920. doi: 10.1093/genetics/94.4.899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry C. V., Lieber R. H. Negative regulation of the Saccharomyces cerevisiae ANB1 gene by heme, as mediated by the ROX1 gene product. Mol Cell Biol. 1986 Dec;6(12):4145–4148. doi: 10.1128/mcb.6.12.4145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry C. V., Weiss J. L., Walthall D. A., Zitomer R. S. Modulator sequences mediate oxygen regulation of CYC1 and a neighboring gene in yeast. Proc Natl Acad Sci U S A. 1983 Jan;80(1):151–155. doi: 10.1073/pnas.80.1.151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry C. V., Zitomer R. S. Oxygen regulation of anaerobic and aerobic genes mediated by a common factor in yeast. Proc Natl Acad Sci U S A. 1984 Oct;81(19):6129–6133. doi: 10.1073/pnas.81.19.6129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowry C. V., Zitomer R. S. ROX1 encodes a heme-induced repression factor regulating ANB1 and CYC7 of Saccharomyces cerevisiae. Mol Cell Biol. 1988 Nov;8(11):4651–4658. doi: 10.1128/mcb.8.11.4651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNeil J. B., Smith M. Transcription initiation of the Saccharomyces cerevisiae iso-1-cytochrome c gene. Multiple, independent T-A-T-A sequences. J Mol Biol. 1986 Feb 5;187(3):363–378. doi: 10.1016/0022-2836(86)90439-0. [DOI] [PubMed] [Google Scholar]
- Mehta K. D., Leung D., Lefebvre L., Smith M. The ANB1 locus of Saccharomyces cerevisiae encodes the protein synthesis initiation factor eIF-4D. J Biol Chem. 1990 May 25;265(15):8802–8807. [PubMed] [Google Scholar]
- Mukai Y., Harashima S., Oshima Y. AAR1/TUP1 protein, with a structure similar to that of the beta subunit of G proteins, is required for a1-alpha 2 and alpha 2 repression in cell type control of Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jul;11(7):3773–3779. doi: 10.1128/mcb.11.7.3773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng R., Abelson J. Isolation and sequence of the gene for actin in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3912–3916. doi: 10.1073/pnas.77.7.3912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfeifer K., Arcangioli B., Guarente L. Yeast HAP1 activator competes with the factor RC2 for binding to the upstream activation site UAS1 of the CYC1 gene. Cell. 1987 Apr 10;49(1):9–18. doi: 10.1016/0092-8674(87)90750-1. [DOI] [PubMed] [Google Scholar]
- Pfeifer K., Prezant T., Guarente L. Yeast HAP1 activator binds to two upstream activation sites of different sequence. Cell. 1987 Apr 10;49(1):19–27. doi: 10.1016/0092-8674(87)90751-3. [DOI] [PubMed] [Google Scholar]
- Pinkham J. L., Olesen J. T., Guarente L. P. Sequence and nuclear localization of the Saccharomyces cerevisiae HAP2 protein, a transcriptional activator. Mol Cell Biol. 1987 Feb;7(2):578–585. doi: 10.1128/mcb.7.2.578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose M. D., Novick P., Thomas J. H., Botstein D., Fink G. R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60(2-3):237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- Rose M., Botstein D. Construction and use of gene fusions to lacZ (beta-galactosidase) that are expressed in yeast. Methods Enzymol. 1983;101:167–180. doi: 10.1016/0076-6879(83)01012-5. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J., Sherman F. Genes affecting the expression of cytochrome c in yeast: genetic mapping and genetic interactions. Genetics. 1980 Apr;94(4):871–889. doi: 10.1093/genetics/94.4.871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnier J., Schwelberger H. G., Smit-McBride Z., Kang H. A., Hershey J. W. Translation initiation factor 5A and its hypusine modification are essential for cell viability in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jun;11(6):3105–3114. doi: 10.1128/mcb.11.6.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz J., Carlson M. Molecular analysis of SSN6, a gene functionally related to the SNF1 protein kinase of Saccharomyces cerevisiae. Mol Cell Biol. 1987 Oct;7(10):3637–3645. doi: 10.1128/mcb.7.10.3637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapira S. K., Chou J., Richaud F. V., Casadaban M. J. New versatile plasmid vectors for expression of hybrid proteins coded by a cloned gene fused to lacZ gene sequences encoding an enzymatically active carboxy-terminal portion of beta-galactosidase. Gene. 1983 Nov;25(1):71–82. doi: 10.1016/0378-1119(83)90169-5. [DOI] [PubMed] [Google Scholar]
- Sherman F., Stewart J. W. Genetics and biosynthesis of cytochrome c. Annu Rev Genet. 1971;5:257–296. doi: 10.1146/annurev.ge.05.120171.001353. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trueblood C. E., Wright R. M., Poyton R. O. Differential regulation of the two genes encoding Saccharomyces cerevisiae cytochrome c oxidase subunit V by heme and the HAP2 and REO1 genes. Mol Cell Biol. 1988 Oct;8(10):4537–4540. doi: 10.1128/mcb.8.10.4537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trumbly R. J. Cloning and characterization of the CYC8 gene mediating glucose repression in yeast. Gene. 1988 Dec 15;73(1):97–111. doi: 10.1016/0378-1119(88)90316-2. [DOI] [PubMed] [Google Scholar]
- Verdière J., Creusot F., Guarente L., Slonimski P. P. The overproducing CYP1 and the underproducing hap1 mutations are alleles of the same gene which regulates in trans the expression of the structural genes encoding iso-cytochromes c. Curr Genet. 1986;10(5):339–342. doi: 10.1007/BF00418404. [DOI] [PubMed] [Google Scholar]
- Verdière J., Creusot F., Guérineau M. Regulation of the expression of iso 2-cytochrome c gene in S. cerevisiae: cloning of the positive regulatory gene CYP1 and identification of the region of its target sequence on the structural gene CYP3. Mol Gen Genet. 1985;199(3):524–533. doi: 10.1007/BF00330769. [DOI] [PubMed] [Google Scholar]
- Williams F. E., Trumbly R. J. Characterization of TUP1, a mediator of glucose repression in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Dec;10(12):6500–6511. doi: 10.1128/mcb.10.12.6500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wright C. F., Zitomer R. S. Point mutations implicate repeated sequences as essential elements of the CYC7 negative upstream site in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Nov;5(11):2951–2958. doi: 10.1128/mcb.5.11.2951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wright R. M., Poyton R. O. Release of two Saccharomyces cerevisiae cytochrome genes, COX6 and CYC1, from glucose repression requires the SNF1 and SSN6 gene products. Mol Cell Biol. 1990 Mar;10(3):1297–1300. doi: 10.1128/mcb.10.3.1297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zagorec M., Buhler J. M., Treich I., Keng T., Guarente L., Labbe-Bois R. Isolation, sequence, and regulation by oxygen of the yeast HEM13 gene coding for coproporphyrinogen oxidase. J Biol Chem. 1988 Jul 15;263(20):9718–9724. [PubMed] [Google Scholar]
- Zagorec M., Labbe-Bois R. Negative control of yeast coproporphyrinogen oxidase synthesis by heme and oxygen. J Biol Chem. 1986 Feb 25;261(6):2506–2509. [PubMed] [Google Scholar]
- Zitomer R. S., Sellers J. W., McCarter D. W., Hastings G. A., Wick P., Lowry C. V. Elements involved in oxygen regulation of the Saccharomyces cerevisiae CYC7 gene. Mol Cell Biol. 1987 Jun;7(6):2212–2220. doi: 10.1128/mcb.7.6.2212. [DOI] [PMC free article] [PubMed] [Google Scholar]