Abstract
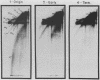
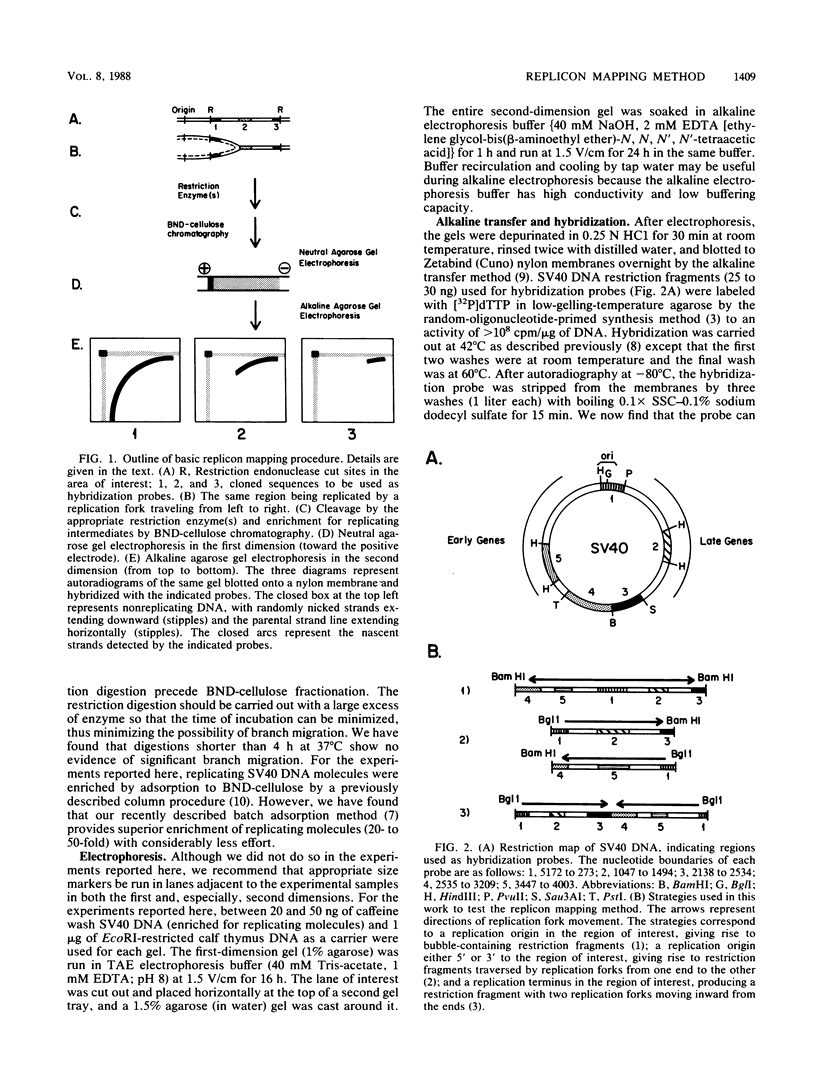
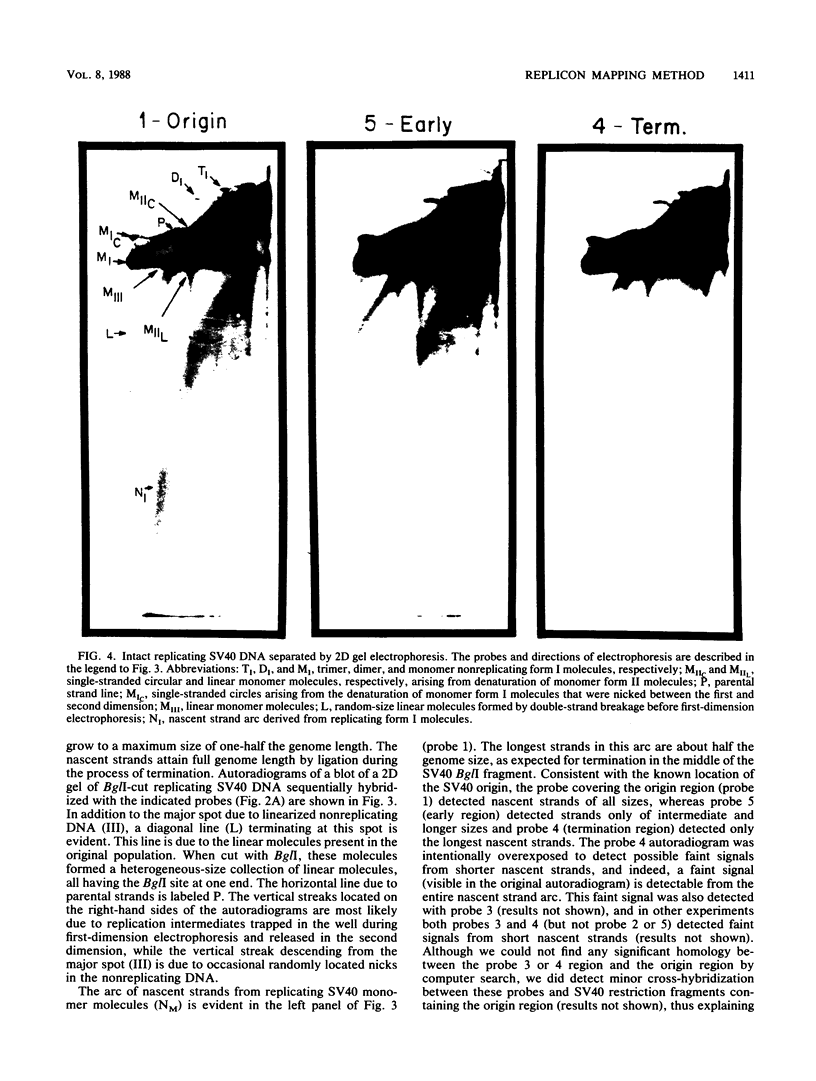
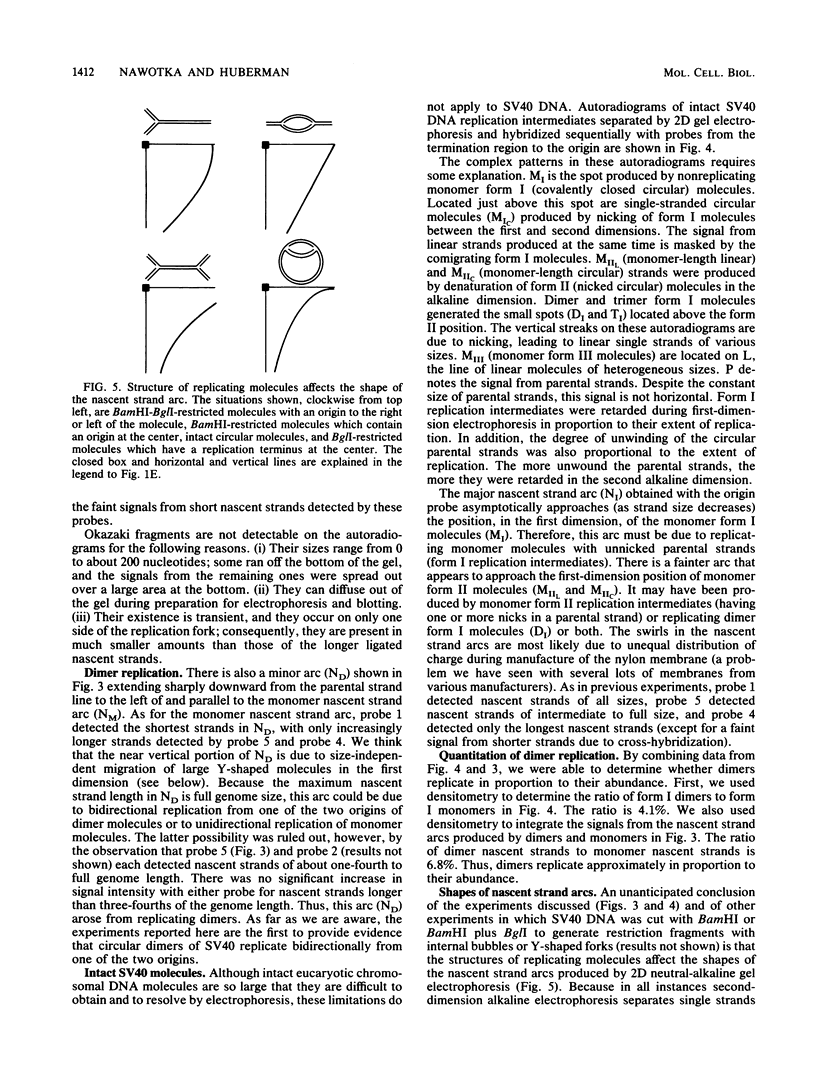
We describe in detail a method which allows determination of the directions of replication fork movement through segments of DNA for which cloned probes are available. The method uses two-dimensional neutral-alkaline agarose gel electrophoresis followed by hybridization with short probe sequences. The nascent strands of replicating molecules form an arc separated from parental and nonreplicating strands. The closer a probe is to its replication origin or to the origin-proximal end of its restriction fragment, the shorter the nascent strands that are detected by the probe. The use of multiple probes allows determination of directions of replication fork movement, as well as locations of origins and termini. In this study, we used simian virus 40 as a model to demonstrate the feasibility of the method, and we discuss its applicability to other systems.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brewer B. J., Fangman W. L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987 Nov 6;51(3):463–471. doi: 10.1016/0092-8674(87)90642-8. [DOI] [PubMed] [Google Scholar]
- Campbell J. L. Eukaryotic DNA replication. Annu Rev Biochem. 1986;55:733–771. doi: 10.1146/annurev.bi.55.070186.003505. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Hay R. T., DePamphilis M. L. Initiation of SV40 DNA replication in vivo: location and structure of 5' ends of DNA synthesized in the ori region. Cell. 1982 Apr;28(4):767–779. doi: 10.1016/0092-8674(82)90056-3. [DOI] [PubMed] [Google Scholar]
- Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967 Jun 14;26(2):365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- Huberman J. A., Spotila L. D., Nawotka K. A., el-Assouli S. M., Davis L. R. The in vivo replication origin of the yeast 2 microns plasmid. Cell. 1987 Nov 6;51(3):473–481. doi: 10.1016/0092-8674(87)90643-x. [DOI] [PubMed] [Google Scholar]
- Pruitt S. C., Reeder R. H. Effect of topological constraint on transcription of ribosomal DNA in Xenopus oocytes. Comparison of plasmid and endogenous genes. J Mol Biol. 1984 Mar 25;174(1):121–139. doi: 10.1016/0022-2836(84)90368-1. [DOI] [PubMed] [Google Scholar]
- Reed K. C., Mann D. A. Rapid transfer of DNA from agarose gels to nylon membranes. Nucleic Acids Res. 1985 Oct 25;13(20):7207–7221. doi: 10.1093/nar/13.20.7207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spotila L. D., Huberman J. A. Method of mapping DNA replication origins. Mol Cell Biol. 1985 Jan;5(1):85–92. doi: 10.1128/mcb.5.1.85. [DOI] [PMC free article] [PubMed] [Google Scholar]