ABSTRACT
Kaposi's sarcoma herpesvirus (KSHV) (or human herpesvirus 8) is the cause of Kaposi's sarcoma, primary effusion lymphoma (PEL), and the plasma cell variant of multicentric Castleman's disease (MCD). The transmembrane K15 protein, encoded by KSHV, has been shown to activate NF-κB and the mitogen-activated protein kinases (MAPKs) c-jun-N-terminal kinase (JNK) and extracellular signal-regulated kinase (Erk) as well as phospholipase C gamma (PLCγ) and to contribute to KSHV-induced angiogenesis. Here we investigate how the K15 protein activates the NF-κB pathway. We show that activation of NF-κB involves the recruitment of NF-κB-inducing kinase (NIK) and IKK α/β to result in the phosphorylation of p65/RelA on Ser536. A K15 mutant devoid in NIK/IKK recruitment fails to activate NF-κB but remains proficient in the stimulation of both NFAT- and AP1-dependent promoters, showing that the structural integrity of the mutant K15 protein has not been altered dramatically. Direct recruitment of NIK represents a novel way for a viral protein to activate and manipulate the NF-κB pathway.
IMPORTANCE KSHV K15 is a viral protein involved in the activation of proinflammatory and angiogenic pathways. Previous studies reported that K15 can activate the NF-κB pathway. Here we show the molecular mechanism underlying the activation of this signaling pathway by K15, which involves direct recruitment of the NF-κB-inducing kinase NIK to K15 as well as NIK-mediated NF-κB p65 phosphorylation on Ser536. K15 is the first viral protein shown to activate NF-κB through direct recruitment of NIK. These results indicate a new mechanism whereby a viral protein can manipulate the NF-κB pathway.
INTRODUCTION
Kaposi's sarcoma (KS) herpesvirus (KSHV) or human herpesvirus 8 (HHV-8) (1), a γ2-herpesvirus (rhadinovirus), causes three human neoplastic diseases: Kaposi's sarcoma, a tumor derived from endothelial cells (2, 3), as well as two B cell tumors, i.e., body cavity-based lymphoma (BCBL)/primary effusion lymphoma (PEL) (4) and the plasma cell variant of multicentric Castleman's disease (5). Endothelial and spindle cells of KS tumors as well as PEL cells and perifollicular B cells of multicentric Castleman's disease are infected with KSHV (6–10). Like other herpesviral infections, most of the KSHV-associated infections are latent and are associated with limited viral gene expression. Only a small fraction of infected cells undergo spontaneous lytic replication (8, 10–12). The open reading frame (ORF) K15 of KSHV is located between the terminal repeat region and the right end of the long unique coding region of the genome. Several divergent alleles of K15 have been identified, among them the predominant K15-P type and the minor K15-M type, which share only 33% of their amino acids (13–16) (Fig. 1A). The K15 gene is expressed in latently infected endothelial cells and B cells, and expression increases during the lytic cycle (13–15). It consists of eight exons, which encode 12 transmembrane domains and a C-terminal cytoplasmic domain (amino acids [aa] 355 to 489 in K15-P) (13, 14, 17). This predicted structure shows some resemblance to the latent membrane proteins LMP1 and LMP2 of Epstein-Barr virus (EBV), which are essential for the ability of EBV to transform primary B cells (14). While LMP1 combines 6 transmembrane domains with a C-terminal cytoplasmic domain, LMP2 features an N-terminal domain followed by 12 transmembrane domains. The C-terminal cytoplasmic domain of both K15 alleles contain several putative signaling motifs underlined in Fig. 1A, right panel (listed here for K15-P): two SH2-binding sites (VFGY431ASI and DDLY481EEV), a proline-rich SH3-binding site, and a TRAF-binding site (A473TQPTDD). K15-P activates the mitogen-activated protein (MAP) kinases c-jun-N-terminal kinase 1 (JNK 1) and extracellular signal-regulated kinase (Erk2), and the transcription factors NF-κB and AP-1 (14, 17, 18). Moreover, K15, by recruiting and activating PLCγ1, contributes to KSHV-mediated angiogenesis (16). The phosphorylation of Y481 in the C-terminal cytoplasmic domain of K15-P by Src kinases (17) is required for most of the K15-P-induced gene expression (17). Mutation of tyrosine 481 to phenylalanine (K15-P Y481F) leads to impaired downstream signaling by K15-P (16–18).
FIG 1.
K15-P induces the binding of NF-κB to target DNA sequence. (A) Left, schematic diagram of K15 and its functional domains. Right, alignment of the protein sequence of the K15-P and M cytoplasmic domains. The conserved motifs are underlined, and the motif in K15-P (aa 359 to 364), shown to be important in this report for the recruitment of NIK and IKKs, is shaded in gray. (B) HEK 293-T cells were cotransfected in duplicate with a vector control or increasing amounts (200 ng, 500 ng, 1 μg) of expression constructs for K15-P wt or K15-P Y481F and an NF-κB-responsive reporter vector. Forty hours after transfection, cells were lysed and the luciferase activity was measured. Equal expression levels of K15 were analyzed by immunoblotting. (C) Electrophoretic mobility shift assays were carried out with 32P-end-labeled double-stranded oligonucleotides corresponding to the consensus binding site for NF-κB (wt NF-κB) or with oligonucleotides containing a mutated NF-κB consensus binding site (mut NF-κB). Reactions were performed either without nuclear extracts (−) or with lysates from HeLa cells transfected with a vector control, K15-P wt, K15-P Y481F, NIK, or vFLIP expression constructs. The NF-κB-oligonucleotide complex is indicated as NF-κB-C. (D) Supershift analyses (SS) were performed with K15-P wt-transfected cells using antibodies specific for different NF-κB proteins (5 μg each), as well as 5 μg anti-IgG as an isotype control. Competition experiments were carried out with 200-fold molar excess of unlabeled wild-type (wt NF-κB) or mutated (mut NF-κB) oligonucleotides. All the experiments were performed three times. **, P < 0.01; ***, P < 0.001.
The NF-κB family of dimeric transcription factors consists of five members, of which three, RelA (p65), RelB, and c-Rel, have transactivating functions, and two, NF-κB1 (p100/p52) and NF-κB2 (p105/p50), have DNA-binding activity (19). NF-κB molecules are sequestered in the cytoplasm by inhibitory proteins, including IκBα, IκBβ, IκBε, p100, and p105 (20). Upon stimulation by cytokines, mitogens, DNA-damaging agents, or microbial infections (21), the inhibitory proteins become phosphorylated by distinct IκB kinase (IKK) complexes and are consequently degraded in a ubiquitin/proteasome-dependent manner (22, 23). The classical IKK complex consists of two catalytic subunits, IKKα and IKKβ, and the regulatory subunit IKKγ (NEMO, IKKAP1, or FIP-3) (21). Currently, two NF-κB pathways can be distinguished: the canonical (or classical) and the noncanonical (or alternative) pathway. The classical pathway plays an important role in immune and inflammatory responses, cell growth, and apoptosis (19, 24, 25) and is activated, among others, by tumor necrosis factor alpha (TNF-α) or interleukin-1 (IL-1). Its activation involves activation of the IKK complex, in particular IKKβ, leading to the phosphorylation and degradation of inhibitory IκB proteins and resulting in the generation of free NF-κB heterodimers (most commonly p50/RelA) and their translocation into the nucleus. The alternative pathway is activated by the B cell-activating factor (BAFF), CD40L, and lymphotoxin β (26–29) and is important for the development of lymphoid organs and the maturation of B cells (30–33). Here, the NF-κB-inducing kinase (NIK) activates IKKα by phosphorylation of Ser176 (34) followed by the processing of p100 to p52 and liberation of p52/RelB dimers (30, 35). Induction of the alternative pathway is independent of IKKβ and IKKγ, which are, however, essential for the classical NF-κB pathway (30, 35). In addition to NF-κB activation through the liberation of NF-κB subunits from cytoplasmic retention, these subunits are also regulated in their transactivational potential. For example, p65 can be phosphorylated at two independent transactivation (TA) domains (TA1 and TA2) within its C-terminal 120 amino acids (36, 37). TNF-α-mediated signaling involves p65 Ser529 phosphorylation (38, 39), whereas p65 Ser536 can be phosphorylated by multiple kinases, including IKKα and IKKβ, depending on the stimuli (40, 41). For example, in LTβ-stimulated cells, IKKα phosphorylates p65 at Ser536 in a NIK-dependent manner (42). Deregulated activation of NF-κB plays a role in several diseases (43–45), and several viral proteins are implicated in the active induction of NF-κB signaling. Examples are the latent membrane protein 1 (LMP1) of KSHV's close relative Epstein-Barr virus, which induces the classical and alternative NF-κB pathway via two signaling domains, termed C-terminal-activating region 1 (CTAR1) 1 and CTAR2 (46–48). While EBV LMP1 CTAR1 is responsible for the recruitment of TRAF1, -2, -3, and -5 with subsequent activation of the noncanonical pathway through the NIK-IKKα-p100 axis, CTAR2 activates the canonical NF-κB pathway. The latter event requires the recruitment of TNIK as well as TRAF6 activation, thereby leading to IKK kinase activation and p65 nuclear translocation (49–51).
Among other γ2 herpesviruses, both herpesvirus saimiri and herpesvirus ateles usurp the NF-κB pathway. The former expresses the saimiri transforming protein C (STPC) responsible for the induction of NF-κB via its association with TRAF1, -2, and -3 and by activating NIK (52, 53). The latter encodes the TiO (two in one) oncoprotein, which directly recruits TRAF6 and stabilizes NIK, thus triggering NF-κB activation via IKKγ-IKKβ (54, 55).
In other virus families, the respiratory syncytial virus (RSV) rapidly activates the alternative pathway prior to activating the classical pathway (56, 57). Furthermore, Tax of human T cell leukemia virus type 1 (HTLV-1) (58) and Tat of human immunodeficiency virus (HIV) are able to induce NF-κB (59). Among the proteins expressed by KSHV, the viral FLICE-inhibitory protein (vFLIP) and the tegument protein ORF75 (60) as well as K15 (17, 18, 61) can activate the NF-κB pathway.
Here we investigate the molecular mechanisms involved in the activation of NF-κB by K15. We show that this involves the recruitment of a NIK/IKK complex by K15 and phosphorylation of p65 on Ser536, leading to an increased binding of the p65/p50 complex to cognate DNA sequences. Since KSHV K15 is the first viral protein known to induce NF-κB and especially p65 Ser536 phosphorylation by directly recruiting NIK, these findings highlight a new mechanism whereby a viral protein can manipulate the NF-κB pathway.
MATERIALS AND METHODS
Cell cultures and transfections.
HEK 293, HEK 293-T, and HeLa cells were grown in Dulbecco's modified Eagle medium (DMEM; Gibco) supplemented with 10% fetal calf serum (FCS), 50 IU/ml penicillin, and 50 μg/ml streptomycin at 37°C in a 5% CO2 incubator. For transfection, cells were grown to subconfluence in six-well plates and transfected with the indicated expression constructs with FuGENE transfection reagent (Roche) according to the manufacturer's instructions. As indicated, cells were stimulated with 20 ng/ml of TNF-α (PeproTech) for 5 min prior to cell lysis. Wedelolactone (EMD Millipore) was added at a concentration of 80 μM to cells.
DNA constructs.
The alanine scanning mutants of GST-K15355–489 (where GST is glutathione S-transferase), GST-K15355–374, and K15-P wt (wild type) were constructed using site-directed mutagenesis with the following primers: pGEXK15-Pdelta4/359–361/for (5′-GTTACGCAGCGGCAAGGCGGCGCATATACACGCGGGACCAG-3′) and pGEXK15-P delta4/359–361/rev (5′-CTGGTCCCGCGTGTATATGCGCCGCCTTCGCTGCGTAAC-3′) for GST-K15 RQR359–361/AAA; pGEXK15-Pdelta4/362–364/for (5′-CCGACAGAGAGCGGCGGCGATATACACGCGGGACCAGAACTTAC-3′) and pGEXK15-Pdelta4/362–364/rev (5′-GTAAGTTCTGGTCCCGCGTGTATATCGCCGCCGCTCTCTGTC GG-3′) for GST-K15 RRR362–364/AAA; pGEXK15-Pdelta4/365–367/for (5′-GGCGGCGCGCAGCCGCGCGGGACCAGAACTTACACC-3′) and pGEXK15-Pdelta4/365–367/rev (5′-GGTGTAAGTTCTGGTCCCGCGCGGCTGCGCGCCGCC-3′) for GST-K15 IYT365–367/AAA; pGEXK15-Pdelta4 368–370 for (5′-GGCGGCGCATATACACGGCGGCCGCGAACTTACACCAC-3′) and pGEXK15-Pdelta4 368–370 rev (5′-GTGGTGTAAGTTCGCGGCCGCCGTGTATATGCGCCGCC-3′) for GST-K15 RDQ 368–370/AAA; pGEXK15-Pdelta4/371–373/for (5′-CGGCGCATATACACGCGGGACCAGGCCGCAGCCCACACAAAGGG-3′) and pGEXK15-Pdelta4/371–373/rev (5′-CCCTTTGTGTGGGCTGCGGCCTGGTCCCGCGTGTATATGCGCCG-3′) for GST-K15 NLH371–373/AAA; and pGEXK15-Pdelta4/374/for (5′-CGCGGGACCAGAACTTACACGCCACAAAGGGAATTC-3′) and pGEXK15-Pdelta4/374/rev (5′-GAATTCCCTTTGTGGCGTGTAAGTTCTGGTCCCGCG-3′) for GST-K15 H374/A.
The construction of other GST-K15 fusion proteins and the K15 mutant K15-P Y481F have been described previously (17), as have the K15-P wt construct clone 35 (13) and the vFLIP-expressing lentiviral vector (62). The expression vectors for NIK (pSC3MT-NIK wt and pSC3MT-NIK [KK429–430AA {dnNIK}]), which lacks the kinase activity due to a double replacement of lysine with alanine in positions 429 and 430, as well as IKKβ were kind gifts of M. Kracht (63) and A. Kieser, respectively. The LMP1 and IKKα constructs have been described previously (64). Reporter vectors were obtained and have been described previously as follows: the NF-κB reporter p3EnhκBconA-Luc (65) was provided by A. Eliopoulos, and the AP-1 reporter pRTU14 and the control vector pRTU1 (29) and the pNFAT-TA-Luc and pTA-Luc vector were purchased from Clontech (18).
siRNA transfections.
Small inhibitory RNAs (siRNAs) were purchased from Dharmacon Research Inc. (Lafayette, CO). HEK 293 or HeLa cells were transfected with 160 pmol siRNA using Lipofectamine 2000 (Invitrogen) at 30 to 50% confluence according to the manufacturer's instructions. Six hours posttransfection, the medium was replaced and cells were transfected with the indicated expression constructs.
Luciferase-based reporter assays.
To assay NF-κB, AP-1, or NFAT activity, HEK 293 and HEK 293-T cells were transiently cotransfected in duplicate with 50 ng of reporter plasmid and expression constructs as indicated in the figure legends. Forty hours after transfection, cells were washed once with phosphate-buffered saline (PBS) and lysed in 300 μl reporter lysis buffer (Promega) per well of a six-well plate. Luciferase activity was measured in cleared lysates with a luciferase system in accordance with the manufacturer's instructions (Promega). Each experiment was performed at least three times in duplicate; the graphs represent a merge of three independent experiments, the error bars indicate the standard deviations, and to assess whether there was a significant difference between the different conditions, a Kruskal-Wallis test with Dunn's posttest was performed.
Immunoblotting.
For the detection of K15 protein by Western blotting, cleared lysates were not boiled prior to SDS-polyacrylamide gel electrophoresis. As indicated, the following primary antibodies were used for immunostaining of immunoblots: mouse anti-actin (Chemicon), rabbit anti-K15 (17), mouse anti-IKKα (BD Biosciences), rabbit anti-IKKβ, mouse anti-IKKγ, rabbit anti-NIK, rabbit anti-NF-κB p105/p50, rabbit anti-phospho p65 (Ser536), rabbit anti-NF-κB2 p100/p52, rabbit phospho IKKα/β (Ser176/180) (Cell Signaling Technology), mouse anti-NF-κB p65 (Santa Cruz), mouse monoclonal anti-hemagglutinin (HA) tag (Roche), and rabbit anti-FLAG (Sigma).
Coimmunoprecipitation.
Forty-eight hours after cotransfection of HEK 293-T cells with the indicated expression constructs, cells were lysed with 250 μl radioimmunoprecipitation assay medium (RIPA-100; 20 mM Tris-HCl [pH 7.5], 1 mM EDTA, 100 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate [DOC], 0.1% SDS) per well of a six-well plate. Cleared lysates (200 μl) were incubated with 0.9 μg of anti-FLAG M2 antibody bound to protein G Sepharose beads, or anti-c-myc monoclonal AB-agarose beads (Clontech), or mouse immunoglobulin (Dako) protein G Sepharose beads at 4°C with gentle shaking overnight. Beads were washed three times with TBST (20 mM Tris-HCl [pH 7.4], 1 mM EDTA, 100 mM NaCl, 1% Triton X-100) and analyzed by SDS-PAGE and Western blotting.
In vitro translation.
For in vitro translation of NIK, IKKα, and IKKβ, the TNT coupled reticulocyte lysate system (Promega) was used according to the manufacturer's instructions. The translation product was incubated with GST fusion proteins or GST only as described below.
GST fusion protein binding assays.
For GST pulldown experiments, Escherichia coli Rosetta cultures transformed with GST expression plasmids or GST only were grown at 37°C in LB medium plus ampicillin and chloramphenicol. Cultures were induced at an optical density at 600 nm of 0.4 to 0.6 with 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) and harvested by centrifugation 5 h after induction. Pelleted cells were resuspended in 500 μl PBS with protease inhibitors, sonicated for 1 min on ice, supplemented with 1% Triton X-100, and incubated for 1 h at 4°C. After centrifugation, the supernatant was incubated with 100 μl glutathione Sepharose beads (Amersham Biosciences) overnight at 4°C. Beads were washed twice with TBST plus protease inhibitors and visualized on a Coomassie blue-stained gel to estimate equal amounts of GST fusion proteins. HEK 293-T cells were transfected as described above with the indicated expression constructs or left untransfected for experiments with endogenous IKKα or IKKβ. Forty-eight hours after transfection, cells were washed once with PBS and lysed with TBS-T lysis buffer (20 mM Tris HCl [pH 7.4], 150 mM NaCl, 1 mM EDTA, 1% Triton X-100) with protease inhibitors for 10 min on ice. Cleared lysates (100 μl) were incubated with precalculated amounts of either GST fusion proteins or GST control protein overnight at 4°C. Beads were washed three times with TBST with protease inhibitors and analyzed by SDS-PAGE and Western blotting.
EMSA.
For electrophoretic mobility shift assays (EMSAs), oligonucleotide probes were prepared as follows: 2 μl NF-κB wild-type or mutant consensus oligonucleotides (Santa Cruz), 3 μl of 10× PNK buffer, 2 μl T4-PNK (NEB), 4 μl γ32P-ATP (222 TBq/mmol), and 19 μl distilled water were mixed and incubated for 1 h at 37°C.
The labeled double-stranded (ds) oligonucleotides were purified using the ProbeQuant G-50 Micro Columns (GE Healthcare) according to the manufacturer's instructions and diluted to 20,000 cpm/μl.
For EMSAs, 8 μg of nuclear extracts (nuclear/cytosol fractionation kit; BioVision; filled up to a total volume of 2.86 μl with 0.42 M NaCl) was added to a 12.14-μl band shift mixture containing 2 μl poly(dI-dC) (1 mg/ml), 1.5 μl 10× band shift buffer (100 mM Tris-HCl [pH 7.5], 100 mM dithiothreitol [DTT], 50% glycerol, 10 mM EDTA), 2 μl labeled ds-oligonucleotide probe, and distilled water and incubated on ice for 30 min. For supershift or cold-competition reactions, the reaction mixtures were pipetted without labeled probe and 5 μg of supershift antibody or 200× excess of unlabeled probe was added to the band shift mixture and incubated on ice for 1 h prior to adding the labeled probe. The native acrylamide gel was prerun for 1 h at 160 V in 1× TBE (890 mM Tris base, 890 mM boric acid, 20 mM EDTA, pH 8). Afterwards, the band shift reaction mixtures were loaded onto the gel and separated for 2 h at 160 V. The gel was dried using the gel drying frame (Roth) according to the manufacturer's instructions. Dried gels were analyzed by autoradiography.
RESULTS
K15 induces the binding of NF-κB proteins to their target DNA sequences.
Using luciferase-based reporter assays, we have previously shown that both the P and M alleles of the KSHV K15 protein activate the NF-κB pathway and that this is at least partially dependent on the phosphorylation of tyrosine 481 in the SH2-B motif (Y481EEV) of the cytoplasmic domain of K15 (Fig. 1A and B and reference 17). The aim of this study was to investigate how K15 activates the NF-κB pathway. First, we checked if the binding of NF-κB proteins to target DNA sequences is induced upon expression of K15. We therefore performed electrophoretic mobility shift assays with radioactively labeled DNA oligonucleotides containing the consensus binding site for NF-κB and nuclear extracts of cells that had been transiently transfected with K15 expression vectors. Nuclear lysates of K15 wt-transfected HeLa cells showed enhanced binding of NF-κB dimers to target DNA sequences (wt NF-κB) compared to lysates of control vector-transfected cells (Fig. 1C). Moreover, when we used lysates of cells transfected with the K15 Y481F mutant, we observed decreased NF-κB/DNA-binding activity compared to lysates of K15 wt-transfected cells. Experiments carried out with lysates of KSHV vFLIP-transfected cells led to the strongest shift of oligonucleotides containing the NF-κB consensus sequence (Fig. 1C). This is in accordance with the observation that vFLIP is the strongest NF-κB inducer among all KSHV proteins (60). Overexpression of the NF-κB-inducing kinase, NIK, also increases interaction of NF-κB with DNA (Fig. 1C). To investigate which NF-κB proteins bind to DNA in response to K15 stimulation, we performed supershift assays (Fig. 1D). We observed the strongest supershift with an antibody to p65/RelA and a less pronounced effect with an antibody to p50. In contrast, we saw no effect with antibodies to p52, c-Rel, and RelB (Fig. 1D). The specificity of supershifts was confirmed by using an isotype control antibody (IgG), and the specificity of the NF-κB/oligonucleotide complex was confirmed by competition experiments with a 200-fold molar excess of unlabeled wt NF-κB and mutant (mut) NF-κB oligonucleotides (cold competition). In summary, these results demonstrate that primarily the NF-κB subunit p65/RelA is bound to cognate DNA upon K15 expression.
siRNA-mediated silencing of IKK proteins, NIK, and p65 impairs K15-mediated NF-κB activation.
In order to elucidate which components of the NF-κB signaling pathway are engaged during the activation by K15, we used small inhibitory RNAs (siRNAs) to downregulate the expression of IKK proteins and NIK, important upstream mediators of NF-κB activation, as well as p65 and p100, prior to the transfection of HEK 293 cells with an NF-κB-responsive luciferase-based reporter vector and an expression vector for the P allele of K15. As can be seen in Fig. 2A, reduced expression of IKKα, IKKβ, and IKKγ led to a moderate decrease of K15-mediated NF-κB activation of about 30%. Of note, siRNA-mediated silencing of NF-κB p100, the precursor of p52, did not result in a reduction of K15-mediated NF-κB activation (Fig. 2A). However, treatment of cells with siRNAs directed against p65 and NIK (Fig. 2A and B) reduced the K15-mediated NF-κB activation to background levels. This is in accordance with our observation that p65 is the most prominent NF-κB protein recruited to its target DNA upon K15 transfection (Fig. 1). In addition, a dominant negative NIK mutant, which lacks the kinase activity due to a double replacement of lysines with alanine residues in positions 429 and 430, strongly reduced the K15-induced activation of the NF-κB-responsive reporter (Fig. 2C).
FIG 2.
Silencing/inhibition of NIK and p65 inhibits K15-dependent NF-κB activation. (A and B) HEK 293 cells were transfected with the indicated siRNAs. Six hours later, the medium was replaced and cells were cotransfected with the indicated expression constructs and an NF-κB-responsive reporter vector. Forty hours after transfection, cells were lysed and the luciferase activity was measured. Shown are relative light units (RLUs) based on duplicate samples. Expression levels of different proteins were analyzed by immunoblotting. (C) HEK 293-T cells were transiently transfected with an NF-κB-responsive reporter vector, an empty vector, or a K15 expression vector and increasing amounts of dnNIK (500 ng and 1 μg). Forty hours after transfection, cells were lysed and luciferase activity was measured. Shown are relative light units based on duplicate samples. Expression levels of K15-P and dnNIK proteins were analyzed by immunoblotting. The experiment was performed three times in duplicate. **, P < 0.01; ***, P < 0.001.
K15 induces p65/RelA phosphorylation in a NIK-dependent manner.
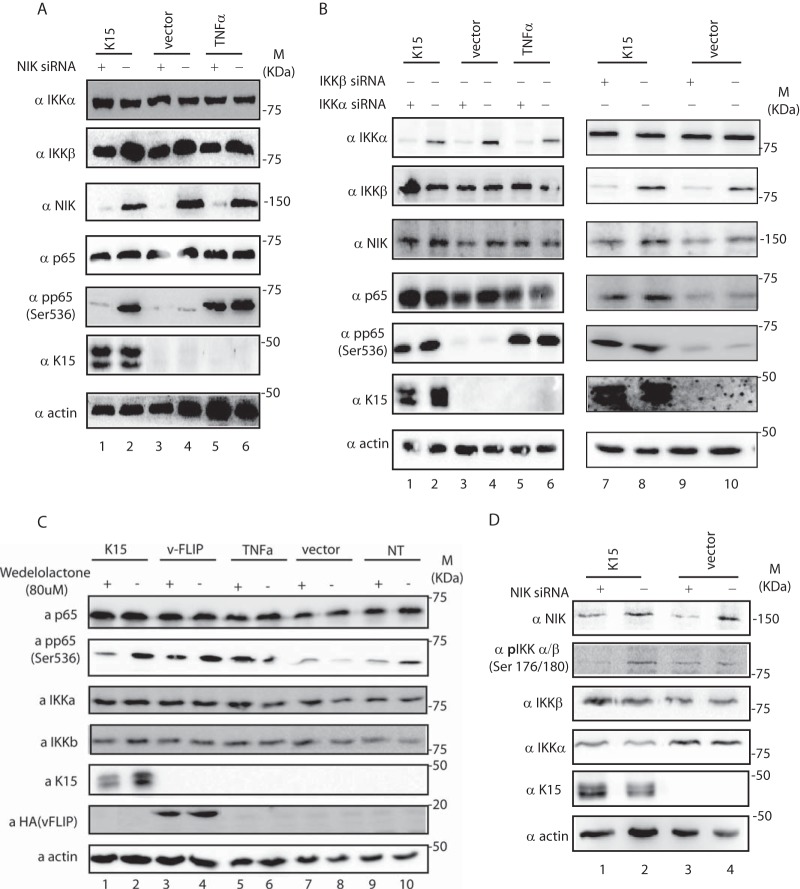
Our results so far suggested an involvement of NIK in the K15-mediated activation of p65/RelA (Fig. 1 and 2). NIK has previously been shown to contribute to the phosphorylation of p65 on Ser536 in response to lymphotoxin β stimulation (42). We therefore explored if K15 expression leads to the phosphorylation of p65/RelA and if this involves NIK. We transfected HeLa cells, which had previously been treated with siRNAs against NIK, IKKα, IKKβ, or control siRNA, with a K15-expressing or a control vector. As a positive control, we stimulated HeLa cells with TNF-α for 5 min before lysis. Figure 3A shows that expression of K15 induces phosphorylation of p65 at Ser536 (compare lanes 2 and 4), which could also be seen in cells stimulated with TNF-α (lanes 5 and 6). Interestingly, siRNA-mediated downregulation of NIK impairs K15- but not TNF-α-mediated phosphorylation of p65. In contrast, individual downregulation of IKKα and IKKβ by siRNA had no effect on the K15-mediated phosphorylation of p65 on Ser536 (Fig. 3B). In an alternative approach to investigate whether K15-induced p65 phosphorylation is dependent on the combined actions of IKKα and IKKβ, we used wedelolactone, a selective and irreversible inhibitor of IKKα and IKKβ enzymatic activity (66) (Fig. 3C). K15, vFLIP, or control vector-transfected HeLa cells were treated 16 h after transfection with 80 μM wedelolactone and then lysed 28 h later, while, as a positive control, cells were treated with TNF-α 5 min prior to lysis. Wedelolactone treatment reduced K15-, as well as vFLIP-induced p65 phosphorylation on Ser536. To further confirm that NIK is responsible for the activation of IKKs by K15 and subsequent p65 phosphorylation, we transfected HeLa cells with either NIK or control siRNA and measured by Western blotting the level of IKKα/β phosphorylation in the presence or absence of K15 (Fig. 3D). We observed that K15 induces IKKα/β phosphorylation in a NIK-dependent manner (compare lanes 1 and 2).
FIG 3.
K15 induces a NIK-dependent phosphorylation of p65 on Ser536. (A and B) HeLa cells were transfected with the indicated siRNAs. Six hours later, the medium was replaced and cells were transfected with the indicated expression constructs for K15, vFLIP, or vector control or left untransfected. Forty hours after transfection, cells were treated as indicated with TNF-α for 5 min and lysed in TBST buffer. Expression levels of different proteins were analyzed by immunoblotting. (C) HeLa cells were either transfected with 1 μg of the indicated expression construct or left untransfected, and 16 h later cells were treated with either wedelolactone or dimethyl sulfoxide (DMSO). Forty hours after transfection, cells were treated with TNF-α for 5 min as indicated and then lysed as described for panels A and B. Protein expression levels were analyzed by immunoblotting. (D) HeLa cells were treated as described for panel A and lysed in TBST buffer. Expression levels of the indicated proteins were measured by Western blotting. With the exception of the experiment shown in panel D, all other experiments were performed three times.
The cytoplasmic domain of K15 directly interacts with NIK.
NIK is a cytoplasmic and nuclear kinase, which is rapidly turned over by the E3 ubiquitin ligase TRAF3. Upon activation of the lymphotoxin β receptor, TRAF3 levels are reduced, resulting in a stabilization of NIK (67). As we had previously observed that the cytoplasmic domain of K15 interacts with TRAF3 (14), we explored if K15 has a direct effect on NIK. We did not observe an increase in NIK levels in K15-transfected cells (Fig. 3 and data not shown). However, we found that K15 and NIK interact directly. We first carried out coimmunoprecipitation experiments with full-length K15 and NIK. Lysates of HEK 293T cells transiently transfected with expression constructs of a Flag-tagged K15 P allele and NIK were incubated with Sepharose beads coated with either anti-Flag or mouse IgG antibody and analyzed by immunoblotting after washing. Figure 4A shows the coprecipitation of NIK with K15, demonstrating an interaction of NIK and K15-P. Using a set of GST fusion proteins containing either the complete K15 cytoplasmic domain or truncated versions thereof (Fig. 4B), we were able to show, in GST pulldown assays, that NIK interacts with the K15 cytoplasmic domain and, in particular, with a short membrane-proximal region (aa 355 to 374) of K15 (Fig. 4C). We were able to confirm the interaction of NIK with the K15 cytoplasmic domain using in vitro-translated NIK, indicating a direct interaction between the two proteins (Fig. 4D). In addition to NIK, its target IKKα, as well as IKKβ, appears to be recruited to the same region (aa 355 to 374), as indicated by GST pulldown assays with lysates from transfected HEK 293-T cells (Fig. 4C, lower panels). In vitro-translated IKKα also showed a strong binding to the cytoplasmic domain of K15 in a GST pulldown assay, whereas the binding of in vitro-translated IKKβ appeared to be weak and inconsistent (Fig. 4D, lower panels). As a next step, we sought to map the interaction site for NIK, IKKα, and IKKβ in the cytoplasmic domain of K15 more precisely. Using site-directed mutagenesis, we generated alanine scanning mutants of the shortest K15 region (aa 355 to 374) that we had found to interact with NIK, IKKα, and IKKβ in a GST pulldown assay (Fig. 4B and C). In these alanine scanning mutants, we substituted triplets of neighboring amino acids in the GST K15 aa 355 to 374 region. These mutated GST-K15 fusion proteins were used in a pulldown experiment with lysates of NIK-transfected or untransfected (for IKKα and IKKβ) HEK 293-T cells. Figure 4E (upper panel) shows that two alanine scanning mutants, GST-K15355–374 RQR359–361/AAA and GST-K15355–374 RRR362–364/AAA were no longer able to interact with NIK, pointing to an essential role of these six amino acids for the interaction of the two proteins. Similar results could be seen for IKKα and IKKβ (Fig. 4E, lower panels). We then introduced the two triple alanine mutations RQR359–361/AAA and RRR362–364/AAA into the GST construct containing the whole cytoplasmic domain of K15 (GST K15355–489). Figure 4F shows that replacement of amino acids 359 to 364 with alanines in the background of the whole cytoplasmic domain of K15 still abolishes the interaction with NIK. In contrast, the recruitment of IKKα and IKKβ was abolished only when amino acids 362 to 364 were mutated, whereas the interaction with IKKβ was preserved when amino acids 359 to 561 were replaced with alanines (Fig. 4F, lower panels). IKKα showed reduced, but still detectable, binding to RQR359–361/AAA.
FIG 4.
K15-P interacts directly with NIK, IKKα, and IKKβ via six amino acids near the last transmembrane domain of K15-P. (A) HEK 293-T cells were cotransfected with the indicated expression constructs or control vector and lysed 48 h later as described in Materials and Methods. Cleared cell lysates were used in a coimmunoprecipitation assay with either α-FLAG or mouse immunoglobulin protein G Sepharose beads. The beads were washed and analyzed by SDS-PAGE and Western blotting. (B) Schematic diagram of GST-K15 fusion proteins used in panels C to F. (C) GST fusion constructs, as depicted in panel B, were used in a GST pulldown assay with transiently expressed NIK or endogenous IKKα and IKKβ. GST proteins were incubated overnight with eukaryotic cell lysates. After extensive washing, beads were analyzed by SDS-PAGE and immunoblotting with an antibody against NIK, IKKα, or IKKβ. (D) NIK, IKKα, and IKKβ proteins were in vitro translated using the TNT coupled reticulocyte lysate system (Promega) and incubated overnight with K15-P fusion proteins or GST only, with which Sepharose beads were coated, at 4°C. The beads were washed, heated with sample buffer, and analyzed by SDS-PAGE and Western blotting. (E) K15-P alanine scanning mutants (see the text) in the background of GST-K15355–374 were used in a GST pulldown assay with transiently expressed NIK or endogenous IKKα and IKKβ. GST proteins were incubated overnight with eukaryotic cell lysates. After extensive washing, beads were analyzed by SDS-PAGE and immunoblotting with antibodies against NIK, IKKα, or IKKβ. (F) GST fusion proteins containing the entire K15 cytoplasmic domain for K15 wt and the indicated mutants were used in a GST pulldown assay with transiently expressed NIK or endogenous IKKα and IKKβ. GST proteins and eukaryotic cell lysates were incubated overnight. After extensive washing, beads were analyzed by SDS-PAGE and immunoblotting with antibodies against NIK, IKKα, or IKKβ. The experiments were performed three times.
A NIK-/IKKα-/IKKβ binding-deficient K15 mutant fails to activate NF-κB activation but retains the ability to stimulate other K15-driven promoters.
To investigate if mutant K15 RQR359–361/AAA or K15 RRR362–364/AAA can still activate NF-κB, we performed NF-κB reporter experiments in HEK 293-T cells. Cells were cotransfected with an NF-κB-responsive reporter vector and increasing amounts of a eukaryotic expression vector for full-length K15 wt, K15 RQR359–361/AAA, or K15 RRR362–364/AAA. As a negative control, we cotransfected increasing amounts of K15 Y481F, a K15 mutant impaired in downstream signaling (17, 18). K15 wt activated NF-κB up to 6-fold in comparison to vector-transfected cells (Fig. 5A). As expected, the tyrosine mutant K15 Y481F showed only weak NF-κB activation. Interestingly, only the K15 RRR362–364/AAA mutant, but not the K15 RQR359–361/AAA mutant, was impaired in NF-κB activation in this reporter assay. Furthermore, we performed EMSAs with nuclear extracts of cells transfected with K15 wt and K15 mutants (Fig. 5B). As already shown above (Fig. 1B), K15 wt induced a bandshift of a radioactively labeled oligonucleotide representing the NF-κB binding site. In contrast, the two K15 mutants were significantly compromised in their ability to induce the binding of NF-κB proteins to cognate DNA sequences in this assay. The K15 RRR362–364/AAA mutant showed a more pronounced defect in inducing the binding of NF-κB to DNA than the K15 RQR359–361/AAA mutant, reflecting to some extent the results of the NF-κB-responsive reporter assay (Fig. 5A).
FIG 5.
K15 mutants deficient in binding to NIK, IKKα, and IKKβ show a decreased ability to activate NF-κB and induce p65 phosphorylation but retain the ability to activate other K15-P-driven promoters. (A) HEK 293-T cells were transiently cotransfected with reporter vectors responsive to NF-κB and increasing amounts (200 ng, 500 ng, 1 μg) of expression constructs of either K15-P wt, K15-P Y481F, K15-P 359–361 RQR/AAA, or K15-P362–364 RRR/AAA. Forty hours after transfection, cells were lysed and luciferase activity was measured. Shown are relative light units based on duplicate samples. Expression levels of K15-P wt and mutants were analyzed by immunoblotting. (B) Electrophoretic mobility shift assays were carried out with 32P-end-labeled double-stranded oligonucleotides corresponding to the consensus binding site for NF-κB. Reactions were performed either without nuclear extracts (−) or with lysates from HeLa cells transfected with empty vector, K15-P wt, K15-P359–361RQR/AAA, or K15-P362–364 RRR/AAA expression constructs. The NF-κB-oligonucleotide complex is indicated as NF-κB-C. (C) HeLa cells were transfected with the indicated expression constructs or left untransfected. Forty hours after transfection, cells were treated with TNF-α for 5 min, where indicated, and lysed. Expression levels of K15 were measured with antibody to K15, of vFLIP with an antibody against HA tag cloned into the vFLIP expression vector, and of p65 or phosphorylated (Ser536) p65 with commercial antibodies (see Materials and Methods). (D and E) HEK 293T cells were transfected with an AP1 reporter plasmid (D) or an NFAT reporter plasmid (E) and increasing concentrations of the indicated K15 expression vectors. Luciferase assays were carried out as described for panel A. The experiments were performed three times in duplicate. n.s., not significant; ***, P < 0.001.
We also wanted to explore if the NIK-, IKKα-, and/or IKKβ-binding-deficient K15 mutants are impaired in inducing the phosphorylation of p65 on Ser536. HeLa cells were transfected with different K15 or vFLIP expression constructs or were treated with TNF-α as a positive control. As can be seen in Fig. 5C, expression of K15 wt or K15 Y481F induces the phosphorylation of p65, as does KSHV vFLIP transfection and TNF-α stimulation. In contrast, both K15 triple alanine mutants (K15 RQR359–361/AAA and K15 RRR362–364/AAA) are impaired in Ser536 phosphorylation of p65. To ascertain if the K15 triple alanine mutants also lead to a reduced induction of other K15-driven signaling pathways, we performed luciferase-based reporter assays with AP-1- and NFAT-responsive promoters (Fig. 5D and E), which we have previously shown to be activated by the expression of K15 (14, 17, 18). Interestingly, the NIK-binding-deficient mutants of K15 retained the ability to activate AP-1- and NFAT-responsive promoters, while, as expected (17, 18), the SH2-binding site mutant K15 Y481F was compromised in this regard. These data suggest that the binding motif for NIK, IKKα, and IKKβ in the cytoplasmic tail of K15 is involved in the activation of NF-κB but not of other K15-P-driven signaling pathways.
DISCUSSION
Previous studies showed that the K15 protein of KSHV induces several signal transduction pathways, including the activation of NF-κB (16–18). Induction of NF-κB via the classical or the alternative pathway is accompanied by the translocation of specific NF-κB dimers from the cytoplasm to the nucleus. Activation of the classical pathway leads in most cases to the release of p50/p65 dimers from cytoplasmic IκB-mediated retention, whereas the alternative pathway induces the processing of p100 and the accumulation of p52/RelB dimers in the nucleus. However, the composition of these dimers is flexible and cannot always be assigned to just one of the pathways mentioned above. We found that expression of K15 wt induces the binding of NF-κB subunits p65 and p50 to cognate target DNA sequences (Fig. 1C and D). We also found the activation of the NF-κB pathway by K15 to be dependent on the presence of a functionally active NIK, and expression of K15 together with a dominant negative NIK (dnNIK) led to a reduced activation of an NF-κB-responsive promoter (Fig. 2C). Suppression of NIK and p65 with siRNAs strongly reduced the activation of NF-κB (Fig. 2A and B), whereas reduced expression of individual components of the IKK complex (IKKα, IKKβ, and IKKγ) led to only a minor effect on K15-mediated NF-κB signaling (Fig. 2A). The K15 Y481F mutant was impaired in the induction of NF-κB/DNA complex formation (Fig. 1C), which, in addition to previously reported experiments (16–18), provides further evidence for the importance of the Y481EEV motif for K15-mediated downstream signaling.
NIK has previously been implicated in LTβR-induced phosphorylation of p65 at position Ser536, a process that also involves IKKα (42). In other settings, IKKβ overexpression stimulates Ser536 phosphorylation of p65 (41). We show here that K15 expression stimulates Ser536 phosphorylation of p65 and IKKα/β in a NIK-dependent manner (Fig. 3A to D). In contrast, silencing the expression of IKKα or IKKβ individually did not reduce the level of K15-dependent NF-κB activation (Fig. 2A) or p65 Ser536 phosphorylation (Fig. 3B). However, treatment with wedelolactone, an inhibitor of both IKKα and IKKβ, reduced the levels of K15-induced p65 Ser536 phosphorylation. As the suppression of IKKα and IKKβ by siRNA was not complete (Fig. 2A and 3B), this suggests the possibility that K15-induced activation of NIK, and subsequent phosphorylation of p65, may involve IKKα and/or IKKβ, which could be able to compensate for each other when only one of them is suppressed by siRNA. Moreover, NIK seems to be responsible for the K15-mediated IKKα/β activation, since silencing of NIK by siRNA (Fig. 4D) abolished IKKα/β phosphorylation. The NF-κB-inducing kinase (NIK) has been identified as the kinase responsible for phosphorylation of IKKα and the consequent activation of alternative NF-κB signaling (34). NIK has also been implicated in the activation of the classical NF-κB pathway by receptors that induce both the classical and the alternative pathways. In contrast to receptors that activate only the classical pathway, these receptors require the recruitment of NIK in addition to the IKK complex to activate both NF-κB pathways (68). Although the well-known activators of the alternative NF-κB pathway, including CD40, LTβR, BAFF, and viral proteins (26–28, 69, 70), utilize NIK for NF-κB activation, only a few of them directly interact with NIK. The present study provides evidence for a direct interaction between K15 and NIK. Coimmunoprecipitation experiments revealed an interaction of the P allele of K15 and NIK in vivo (Fig. 4A), and GST pulldown assays with lysates of NIK-transfected cells and in vitro-translated NIK showed that this interaction occurs directly (Fig. 4C and D). Using alanine scanning mutagenesis, we defined six amino acids (aa 359 to 364) responsible for NIK binding (Fig. 4E and F). Functional assays confirmed that K15 mutants RQR359–361/AAA and RRR362–364/AAA showed a decreased ability to activate NF-κB (Fig. 5A, B, and C). However, while both mutants showed a decreased ability to induce an NF-κB bandshift (Fig. 5B) and p65 Ser536 phosphorylation (Fig. 5C), only mutant RRR362–364/AAA failed to activate an NF-κB promoter (Fig. 5A). Since this mutant retained residual activity in the NF-κB bandshift assay (Fig. 5B), we assume that this residual activity is sufficient to activate the NF-κB reporter vector. Alternatively, another explanation could be that K15 does not need NIK binding for NF-κB activation, as measured in the reporter assay (Fig. 5A) and in the NF-κB bandshift assay (Fig. 5B), but that NIK binding to K15 could be required for p65 phosphorylation and a subsequent modification of NF-κB dimers and, thereby, the spectrum of NF-κB target genes that are activated by K15.
We have also shown that the targets of NIK, IKKα, and IKKβ, are directly recruited to the same region of the K15 cytoplasmic domain (Fig. 4C to F). In contrast to what occurs with NIK, substitution of amino acids 359 to 361 of K15 did not completely abolish the interaction with IKKα and IKKβ, whereas mutation of amino acids 362 to 364 entirely prevented the recruitment of IKKα and IKKβ to K15 (Fig. 4F). Thus, K15 appears to utilize a mechanism of activating the NF-κB pathway that is different from that of of KSHV vFLIP and HTLV-1 Tax, which both directly recruit IKKγ/NEMO (74–77). It is thus possible that K15 functionally resembles IKKγ/NEMO, working as a “regulatory platform” able to recruit IKKs and modulate the NF-κB signal. While IKKγ/NEMO functions are normally regulated by ubiquitination, K15 might work as a constitutively active NF-κB inducer. Further investigation would be needed to address this question.
Recruitment of NF-κB pathway components has also been shown for other viruses. The hepatitis C virus core protein suppresses NF-κB activation by directly recruiting IKKβ (71), and the Epstein-Barr virus LMP1 protein activates the NF-κB pathway in a NIK-dependent manner, but the mechanism of its recruitment is not clear (72). The Tio protein of herpesvirus saimiri activates the alternative NF-κB pathway by stabilizing NIK (55).
While the K15 mutant RRR362–364RRR/AAA shows a pronounced defect in NF-κB activation in NF-κB reporter and NF-κB bandshift assays and is defective in phosphorylating Ser536 of p65, it is capable of activating AP-1 and NFAT-responsive promoter activity (Fig. 5D and E). This observation indicates that the K15362–364RRR/AAA mutant likely does not suffer from a structural defect that would affect all currently known K15 functions (14, 17, 18, 44) but is selectively deficient in NF-κB activation. Thus, the region located in the most proximal region of the cytoplasmic tail of K15 (comprising amino acids 362 to 364) seems to selectively contribute to activation of the transcription factor NF-κB but not of others, such as NFAT or AP-1. It thereby differs from the YEEV SH2 binding site, which appears to be involved in all K15-activated pathways reported so far (14, 16–18, 73). Interestingly, while a mutation of this SH2 binding site decreases K15-dependent activation of an NF-κB reporter (Fig. 1B) and recruitment of p65/p50 to a cognate NF-κB binding site (Fig. 1C), phosphorylation of p65 at Ser536 is not affected (Fig. 5C). Since the two K15 mutants K15359–361RQR/AAA and K15362–364RRR/AAA fail to induce p65 Ser536 phosphorylation, this observation suggests that two K15-dependent signals contribute to NF-κB activation, one of which, recruitment of the NIK/IKKα/IKKβ complex to aa 359 to 364 of K15, leads to the phosphorylation of p65 Ser536.
Although this study focused mainly on K15-P-mediated NF-κB signaling, previous reports (17, 18) showed that the M allele of K15 can also activate NF-κB. Interestingly, of the two regions found to be important for K15-P-mediated NF-κB activation, one, the YEEVL motif, is conserved in K15M, while the other one (aa 359 to 364), shown here to be involved in the recruitment of NIK, IKKα/β, is not (Fig. 1A). This observation is compatible with two possible interpretations: (i) K15M does not recruit NIK or IKKα/β and activates NF-κB in a different manner, which however still relies on the conserved YEEVL motif; (ii) other regions in the K15M cytoplasmic tail are required for the recruitment of NIK and/or IKKα/β. Further experiments are therefore required to elucidate the exact mechanism by which K15M mediates NF-κB activation.
Several KSHV proteins activate the NF-κB pathway and may utilize the NF-κB pathway to regulate different aspects of the viral life cycle. A KSHV mutant from which the K15 gene has been deleted induces a slightly different cellular gene expression pattern in infected primary endothelial cells from that brought about by a KSHV wt virus, and some of these differentially regulated cellular genes, such as that encoding IL-8, are known to be regulated by NF-κB (16). It is thus possible that the recruitment of NIK by K15 described in this report contributes to the regulation of NF-κB-dependent cellular inflammatory genes in KSHV-infected cells. A detailed analysis of this question would require a recombinant KSHV carrying the K15362–364RRR/AAA mutant instead of wt K15, which we plan to construct and test in future experiments.
ACKNOWLEDGMENTS
This study was supported by SFB 566 (TP11) and SFB 796 (TP B9) of the Deutsche Forschungsgemeinschaft (DFG), by DFG project grant Schu1688/3-1, by the European Union Integrated Project INCA (LSHC-CT-2005-018704), and by the Interdisciplinary Center for Clinical Research (IZKF) of the Clinical Center Erlangen.
Footnotes
Published ahead of print 3 September 2014
REFERENCES
- 1.Chang Y, Cesarman E, Pessin MS, Lee F, Culpepper J, Knowles DM, Moore PS. 1994. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi's sarcoma. Science 266:1865–1869. 10.1126/science.7997879. [DOI] [PubMed] [Google Scholar]
- 2.Moore PS, Chang Y. 1995. Detection of herpesvirus-like DNA sequences in Kaposi's sarcoma in patients with and without HIV infection. N. Engl. J. Med. 332:1181–1185. 10.1056/NEJM199505043321801. [DOI] [PubMed] [Google Scholar]
- 3.Schulz TF. 2001. KSHV/HHV8-associated lymphoproliferations in the AIDS setting. Eur. J. Cancer 37:1217–1226. 10.1016/S0959-8049(01)00115-0. [DOI] [PubMed] [Google Scholar]
- 4.Cesarman E, Chang Y, Moore PS, Said JW, Knowles DM. 1995. Kaposi's sarcoma-associated herpesvirus-like DNA sequences in AIDS-related body-cavity-based lymphomas. N. Engl. J. Med. 332:1186–1191. 10.1056/NEJM199505043321802. [DOI] [PubMed] [Google Scholar]
- 5.Soulier J, Grollet L, Oksenhendler E, Cacoub P, Cazals-Hatem D, Babinet P, d'Agay MF, Clauvel JP, Raphael M, Degos L, Sigaux F. 1995. Kaposi's sarcoma-associated herpesvirus-like DNA sequences in multicentric Castleman's disease. Blood 86:1276–1280. [PubMed] [Google Scholar]
- 6.Boshoff C, Schulz TF, Kennedy MM, Graham AK, Fisher C, Thomas A, McGee JO, Weiss RA, O'Leary JJ. 1995. Kaposi's sarcoma-associated herpesvirus infects endothelial and spindle cells. Nat. Med. 1:1274–1278. 10.1038/nm1295-1274. [DOI] [PubMed] [Google Scholar]
- 7.Dupin N, Fisher C, Kellam P, Ariad S, Tulliez M, Franck N, van Marck E, Salmon D, Gorin I, Escande JP, Weiss RA, Alitalo K, Boshoff C. 1999. Distribution of human herpesvirus-8 latently infected cells in Kaposi's sarcoma, multicentric Castleman's disease, and primary effusion lymphoma. Proc. Natl. Acad. Sci. U. S. A. 96:4546–4551. 10.1073/pnas.96.8.4546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Katano H, Sato Y, Kurata T, Mori S, Sata T. 2000. Expression and localization of human herpesvirus 8-encoded proteins in primary effusion lymphoma, Kaposi's sarcoma, and multicentric Castleman's disease. Virology 269:335–344. 10.1006/viro.2000.0196. [DOI] [PubMed] [Google Scholar]
- 9.Parravicini C, Chandran B, Corbellino M, Berti E, Paulli M, Moore PS, Chang Y. 2000. Differential viral protein expression in Kaposi's sarcoma-associated herpesvirus-infected diseases: Kaposi's sarcoma, primary effusion lymphoma, and multicentric Castleman's disease. Am. J. Pathol. 156:743–749. 10.1016/S0002-9440(10)64940-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rainbow L, Platt GM, Simpson GR, Sarid R, Gao SJ, Stoiber H, Herrington CS, Moore PS, Schulz TF. 1997. The 222- to 234-kilodalton latent nuclear protein (LNA) of Kaposi's sarcoma-associated herpesvirus (human herpesvirus 8) is encoded by orf73 and is a component of the latency-associated nuclear antigen. J. Virol. 71:5915–5921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blasig C, Zietz C, Haar B, Neipel F, Esser S, Brockmeyer NH, Tschachler E, Colombini S, Ensoli B, Sturzl M. 1997. Monocytes in Kaposi's sarcoma lesions are productively infected by human herpesvirus 8. J. Virol. 71:7963–7968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chiou CJ, Poole LJ, Kim PS, Ciufo DM, Cannon JS, ap Rhys CM, Alcendor DJ, Zong JC, Ambinder RF, Hayward GS. 2002. Patterns of gene expression and a transactivation function exhibited by the vGCR (ORF74) chemokine receptor protein of Kaposi's sarcoma-associated herpesvirus. J. Virol. 76:3421–3439. 10.1128/JVI.76.7.3421-3439.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Choi JK, Lee BS, Shim SN, Li M, Jung JU. 2000. Identification of the novel K15 gene at the rightmost end of the Kaposi's sarcoma-associated herpesvirus genome. J. Virol. 74:436–446. 10.1128/JVI.74.1.436-446.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Glenn M, Rainbow L, Aurade F, Davison A, Schulz TF. 1999. Identification of a spliced gene from Kaposi's sarcoma-associated herpesvirus encoding a protein with similarities to latent membrane proteins 1 and 2A of Epstein-Barr virus. J. Virol. 73:6953–6963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Poole LJ, Zong JC, Ciufo DM, Alcendor DJ, Cannon JS, Ambinder R, Orenstein JM, Reitz MS, Hayward GS. 1999. Comparison of genetic variability at multiple loci across the genomes of the major subtypes of Kaposi's sarcoma-associated herpesvirus reveals evidence for recombination and for two distinct types of open reading frame K15 alleles at the right-hand end. J. Virol. 73:6646–6660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bala K, Bosco R, Gramolelli S, Haas DA, Kati S, Pietrek M, Havemeier A, Yakushko Y, Singh VV, Dittrich-Breiholz O, Kracht M, Schulz TF. 2012. Kaposi's sarcoma herpesvirus K15 protein contributes to virus-induced angiogenesis by recruiting PLCgamma1 and activating NFAT1-dependent RCAN1 expression. PLoS Pathog. 8:e1002927. 10.1371/journal.ppat.1002927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brinkmann MM, Glenn M, Rainbow L, Kieser A, Henke-Gendo C, Schulz TF. 2003. Activation of mitogen-activated protein kinase and NF-kappaB pathways by a Kaposi's sarcoma-associated herpesvirus K15 membrane protein. J. Virol. 77:9346–9358. 10.1128/JVI.77.17.9346-9358.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Brinkmann MM, Pietrek M, Dittrich-Breiholz O, Kracht M, Schulz TF. 2007. Modulation of host gene expression by the K15 protein of Kaposi's sarcoma-associated herpesvirus. J. Virol. 81:42–58. 10.1128/JVI.00648-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Siebenlist U, Franzoso G, Brown K. 1994. Structure, regulation and function of NF-kappa B. Annu. Rev. Cell Biol. 10:405–455. 10.1146/annurev.cb.10.110194.002201. [DOI] [PubMed] [Google Scholar]
- 20.Baldwin AS., Jr 1996. The NF-kappa B and I kappa B proteins: new discoveries and insights. Annu. Rev. Immunol. 14:649–683. 10.1146/annurev.immunol.14.1.649. [DOI] [PubMed] [Google Scholar]
- 21.Karin M, Delhase M. 2000. The I kappa B kinase (IKK) and NF-kappa B: key elements of proinflammatory signalling. Semin. Immunol. 12:85–98. 10.1006/smim.2000.0210. [DOI] [PubMed] [Google Scholar]
- 22.Bonizzi G, Karin M. 2004. The two NF-kappaB activation pathways and their role in innate and adaptive immunity. Trends Immunol. 25:280–288. 10.1016/j.it.2004.03.008. [DOI] [PubMed] [Google Scholar]
- 23.Hayden MS, Ghosh S. 2004. Signaling to NF-kappaB. Genes Dev. 18:2195–2224. 10.1101/gad.1228704. [DOI] [PubMed] [Google Scholar]
- 24.Karin M, Lin A. 2002. NF-kappaB at the crossroads of life and death. Nat. Immunol. 3:221–227. 10.1038/ni0302-221. [DOI] [PubMed] [Google Scholar]
- 25.Silverman N, Maniatis T. 2001. NF-kappaB signaling pathways in mammalian and insect innate immunity. Genes Dev. 15:2321–2342. 10.1101/gad.909001. [DOI] [PubMed] [Google Scholar]
- 26.Claudio E, Brown K, Park S, Wang H, Siebenlist U. 2002. BAFF-induced NEMO-independent processing of NF-kappa B2 in maturing B cells. Nat. Immunol. 3:958–965. 10.1038/ni842. [DOI] [PubMed] [Google Scholar]
- 27.Coope HJ, Atkinson PG, Huhse B, Belich M, Janzen J, Holman MJ, Klaus GG, Johnston LH, Ley SC. 2002. CD40 regulates the processing of NF-kappaB2 p100 to p52. EMBO J. 21:5375–5385. 10.1093/emboj/cdf542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Dejardin E, Droin NM, Delhase M, Haas E, Cao Y, Makris C, Li ZW, Karin M, Ware CF, Green DR. 2002. The lymphotoxin-beta receptor induces different patterns of gene expression via two NF-kappaB pathways. Immunity 17:525–535. 10.1016/S1074-7613(02)00423-5. [DOI] [PubMed] [Google Scholar]
- 29.Kayagaki N, Yan M, Seshasayee D, Wang H, Lee W, French DM, Grewal IS, Cochran AG, Gordon NC, Yin J, Starovasnik MA, Dixit VM. 2002. BAFF/BLyS receptor 3 binds the B cell survival factor BAFF ligand through a discrete surface loop and promotes processing of NF-kappaB2. Immunity 17:515–524. 10.1016/S1074-7613(02)00425-9. [DOI] [PubMed] [Google Scholar]
- 30.Senftleben U, Cao Y, Xiao G, Greten FR, Krahn G, Bonizzi G, Chen Y, Hu Y, Fong A, Sun SC, Karin M. 2001. Activation by IKKalpha of a second, evolutionary conserved, NF-kappa B signaling pathway. Science 293:1495–1499. 10.1126/science.1062677. [DOI] [PubMed] [Google Scholar]
- 31.Shinkura R, Kitada K, Matsuda F, Tashiro K, Ikuta K, Suzuki M, Kogishi K, Serikawa T, Honjo T. 1999. Alymphoplasia is caused by a point mutation in the mouse gene encoding Nf-kappa b-inducing kinase. Nat. Genet. 22:74–77. 10.1038/8780. [DOI] [PubMed] [Google Scholar]
- 32.Yilmaz ZB, Weih DS, Sivakumar V, Weih F. 2003. RelB is required for Peyer's patch development: differential regulation of p52-RelB by lymphotoxin and TNF. EMBO J. 22:121–130. 10.1093/emboj/cdg004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yin L, Wu L, Wesche H, Arthur CD, White JM, Goeddel DV, Schreiber RD. 2001. Defective lymphotoxin-beta receptor-induced NF-kappaB transcriptional activity in NIK-deficient mice. Science 291:2162–2165. 10.1126/science.1058453. [DOI] [PubMed] [Google Scholar]
- 34.Ling L, Cao Z, Goeddel DV. 1998. NF-kappaB-inducing kinase activates IKK-alpha by phosphorylation of Ser-176. Proc. Natl. Acad. Sci. U. S. A. 95:3792–3797. 10.1073/pnas.95.7.3792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Xiao G, Harhaj EW, Sun SC. 2001. NF-kappaB-inducing kinase regulates the processing of NF-kappaB2 p100. Mol. Cell 7:401–409. 10.1016/S1097-2765(01)00187-3. [DOI] [PubMed] [Google Scholar]
- 36.Schmitz ML, dos Santos Silva MA, Baeuerle PA. 1995. Transactivation domain 2 (TA2) of p65 NF-kappa B. Similarity to TA1 and phorbol ester-stimulated activity and phosphorylation in intact cells. J. Biol. Chem. 270:15576–15584. [DOI] [PubMed] [Google Scholar]
- 37.Schmitz ML, Stelzer G, Altmann H, Meisterernst M, Baeuerle PA. 1995. Interaction of the COOH-terminal transactivation domain of p65 NF-kappa B with TATA-binding protein, transcription factor IIB, and coactivators. J. Biol. Chem. 270:7219–7226. 10.1074/jbc.270.13.7219. [DOI] [PubMed] [Google Scholar]
- 38.Wang D, Baldwin AS., Jr 1998. Activation of nuclear factor-kappaB-dependent transcription by tumor necrosis factor-alpha is mediated through phosphorylation of RelA/p65 on serine 529. J. Biol. Chem. 273:29411–29416. 10.1074/jbc.273.45.29411. [DOI] [PubMed] [Google Scholar]
- 39.Wang D, Westerheide SD, Hanson JL, Baldwin AS., Jr 2000. Tumor necrosis factor alpha-induced phosphorylation of RelA/p65 on Ser529 is controlled by casein kinase II. J. Biol. Chem. 275:32592–32597. 10.1074/jbc.M001358200. [DOI] [PubMed] [Google Scholar]
- 40.Mattioli I, Sebald A, Bucher C, Charles RP, Nakano H, Doi T, Kracht M, Schmitz ML. 2004. Transient and selective NF-kappa B p65 serine 536 phosphorylation induced by T cell costimulation is mediated by I kappa B kinase beta and controls the kinetics of p65 nuclear import. J. Immunol. 172:6336-6344. 10.4049/jimmunol.172.10.6336. [DOI] [PubMed] [Google Scholar]
- 41.Sakurai H, Chiba H, Miyoshi H, Sugita T, Toriumi W. 1999. IkappaB kinases phosphorylate NF-kappaB p65 subunit on serine 536 in the transactivation domain. J. Biol. Chem. 274:30353–30356. 10.1074/jbc.274.43.30353. [DOI] [PubMed] [Google Scholar]
- 42.Jiang X, Takahashi N, Matsui N, Tetsuka T, Okamoto T. 2003. The NF-kappa B activation in lymphotoxin beta receptor signaling depends on the phosphorylation of p65 at serine 536. J. Biol. Chem. 278:919–926. 10.1074/jbc.M208696200. [DOI] [PubMed] [Google Scholar]
- 43.Nakshatri H, Bhat-Nakshatri P, Martin DA, Goulet RJ, Jr, Sledge GW., Jr 1997. Constitutive activation of NF-kappaB during progression of breast cancer to hormone-independent growth. Mol. Cell. Biol. 17:3629–3639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Neumann M, Grieshammer T, Chuvpilo S, Kneitz B, Lohoff M, Schimpl A, Franza BR, Jr, Serfling E. 1995. RelA/p65 is a molecular target for the immunosuppressive action of protein kinase A. EMBO J. 14:1991–2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Rayet B, Gelinas C. 1999. Aberrant rel/nfkb genes and activity in human cancer. Oncogene 18:6938–6947. 10.1038/sj.onc.1203221. [DOI] [PubMed] [Google Scholar]
- 46.Eliopoulos AG, Caamano JH, Flavell J, Reynolds GM, Murray PG, Poyet JL, Young LS. 2003. Epstein-Barr virus-encoded latent infection membrane protein 1 regulates the processing of p100 NF-kappaB2 to p52 via an IKKgamma/NEMO-independent signalling pathway. Oncogene 22:7557–7569. 10.1038/sj.onc.1207120. [DOI] [PubMed] [Google Scholar]
- 47.Eliopoulos AG, Young LS. 2001. LMP1 structure and signal transduction. Semin. Cancer Biol. 11:435–444. 10.1006/scbi.2001.0410. [DOI] [PubMed] [Google Scholar]
- 48.Mosialos G. 2001. Cytokine signaling and Epstein-Barr virus-mediated cell transformation. Cytokine Growth Factor Rev. 12:259–270. 10.1016/S1359-6101(00)00035-6. [DOI] [PubMed] [Google Scholar]
- 49.Schultheiss U, Puschner S, Kremmer E, Mak TW, Engelmann H, Hammerschmidt W, Kieser A. 2001. TRAF6 is a critical mediator of signal transduction by the viral oncogene latent membrane protein 1. EMBO J. 20:5678–5691. 10.1093/emboj/20.20.5678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Luftig M, Prinarakis E, Yasui T, Tsichritzis T, Cahir-McFarland E, Inoue J, Nakano H, Mak TW, Yeh WC, Li X, Akira S, Suzuki N, Suzuki S, Mosialos G, Kieff E. 2003. Epstein-Barr virus latent membrane protein 1 activation of NF-kappaB through IRAK1 and TRAF6. Proc. Natl. Acad. Sci. U. S. A. 100:15595–15600. 10.1073/pnas.2136756100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shkoda A, Town JA, Griese J, Romio M, Sarioglu H, Knofel T, Giehler F, Kieser A. 2012. The germinal center kinase TNIK is required for canonical NF-kappaB and JNK signaling in B-cells by the EBV oncoprotein LMP1 and the CD40 receptor. PLoS Biol. 10:e1001376. 10.1371/journal.pbio.1001376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lee H, Choi JK, Li M, Kaye K, Kieff E, Jung JU. 1999. Role of cellular tumor necrosis factor receptor-associated factors in NF-kappaB activation and lymphocyte transformation by herpesvirus Saimiri STP. J. Virol. 73:3913–3919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Sorokina EM, Merlo JJ, Jr, Tsygankov AY. 2004. Molecular mechanisms of the effect of herpesvirus saimiri protein StpC on the signaling pathway leading to NF-kappaB activation. J. Biol. Chem. 279:13469–13477. 10.1074/jbc.M305250200. [DOI] [PubMed] [Google Scholar]
- 54.Heinemann S, Biesinger B, Fleckenstein B, Albrecht JC. 2006. NFkappaB signaling is induced by the oncoprotein Tio through direct interaction with TRAF6. J. Biol. Chem. 281:8565–8572. 10.1074/jbc.M510891200. [DOI] [PubMed] [Google Scholar]
- 55.de Jong SJ, Albrecht JC, Schmidt M, Muller-Fleckenstein I, Biesinger B. 2010. Activation of noncanonical NF-kappaB signaling by the oncoprotein Tio. J. Biol. Chem. 285:16495–16503. 10.1074/jbc.M110.102848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Choudhary S, Boldogh S, Garofalo R, Jamaluddin M, Brasier AR. 2005. Respiratory syncytial virus influences NF-kappaB-dependent gene expression through a novel pathway involving MAP3K14/NIK expression and nuclear complex formation with NF-kappaB2. J. Virol. 79:8948–8959. 10.1128/JVI.79.14.8948-8959.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Liu P, Li K, Garofalo RP, Brasier AR. 2008. Respiratory syncytial virus induces RelA release from cytoplasmic 100-kDa NF-kappa B2 complexes via a novel retinoic acid-inducible gene-I{middle dot}NF-kappa B-inducing kinase signaling pathway. J. Biol. Chem. 283:23169–23178. 10.1074/jbc.M802729200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Uhlik M, Good L, Xiao G, Harhaj EW, Zandi E, Karin M, Sun SC. 1998. NF-kappaB-inducing kinase and IkappaB kinase participate in human T-cell leukemia virus I Tax-mediated NF-kappaB activation. J. Biol. Chem. 273:21132–21136. 10.1074/jbc.273.33.21132. [DOI] [PubMed] [Google Scholar]
- 59.Leghmari K, Bennasser Y, Bahraoui E. 2008. HIV-1 Tat protein induces IL-10 production in monocytes by classical and alternative NF-kappaB pathways. Eur. J. Cell Biol. 87:947–962. 10.1016/j.ejcb.2008.06.005. [DOI] [PubMed] [Google Scholar]
- 60.Konrad A, Wies E, Thurau M, Marquardt G, Naschberger E, Hentschel S, Jochmann R, Schulz TF, Erfle H, Brors B, Lausen B, Neipel F, Sturzl M. 2009. A systems biology approach to identify the combination effects of human herpesvirus 8 genes on NF-kappaB activation. J. Virol. 83:2563–2574. 10.1128/JVI.01512-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Matta H, Mazzacurati L, Schamus S, Yang T, Sun Q, Chaudhary PM. 2007. Kaposi's sarcoma-associated herpesvirus (KSHV) oncoprotein K13 bypasses TRAFs and directly interacts with the IkappaB kinase complex to selectively activate NF-kappaB without JNK activation. J. Biol. Chem. 282:24858–24865. 10.1074/jbc.M700118200. [DOI] [PubMed] [Google Scholar]
- 62.Alkharsah KR, Singh VV, Bosco R, Santag S, Grundhoff A, Konrad A, Sturzl M, Wirth D, Dittrich-Breiholz O, Kracht M, Schulz TF. 2011. Deletion of Kaposi's sarcoma-associated herpesvirus FLICE inhibitory protein, vFLIP, from the viral genome compromises the activation of STAT1-responsive cellular genes and spindle cell formation in endothelial cells. J. Virol. 85:10375–10388. 10.1128/JVI.00226-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Holtmann H, Winzen R, Holland P, Eickemeier S, Hoffmann E, Wallach D, Malinin NL, Cooper JA, Resch K, Kracht M. 1999. Induction of interleukin-8 synthesis integrates effects on transcription and mRNA degradation from at least three different cytokine- or stress-activated signal transduction pathways. Mol. Cell. Biol. 19:6742–6753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Huen DS, Henderson SA, Croom-Carter D, Rowe M. 1995. The Epstein-Barr virus latent membrane protein-1 (LMP1) mediates activation of NF-kappa B and cell surface phenotype via two effector regions in its carboxy-terminal cytoplasmic domain. Oncogene 10:549–560. [PubMed] [Google Scholar]
- 65.Eliopoulos AG, Stack M, Dawson CW, Kaye KM, Hodgkin L, Sihota S, Rowe M, Young LS. 1997. Epstein-Barr virus-encoded LMP1 and CD40 mediate IL-6 production in epithelial cells via an NF-kappaB pathway involving TNF receptor-associated factors. Oncogene 14:2899–2916. 10.1038/sj.onc.1201258. [DOI] [PubMed] [Google Scholar]
- 66.Kobori M, Yang Z, Gong D, Heissmeyer V, Zhu H, Jung YK, Gakidis MA, Rao A, Sekine T, Ikegami F, Yuan C, Yuan J. 2004. Wedelolactone suppresses LPS-induced caspase-11 expression by directly inhibiting the IKK complex. Cell Death Differ. 11:123–130. 10.1038/sj.cdd.4401325. [DOI] [PubMed] [Google Scholar]
- 67.Sanjo H, Zajonc DM, Braden R, Norris PS, Ware CF. 2010. Allosteric regulation of the ubiquitin:NIK and ubiquitin:TRAF3 E3 ligases by the lymphotoxin-beta receptor. J. Biol. Chem. 285:17148–17155. 10.1074/jbc.M110.105874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Ramakrishnan P, Wang W, Wallach D. 2004. Receptor-specific signaling for both the alternative and the canonical NF-kappaB activation pathways by NF-kappaB-inducing kinase. Immunity 21:477–489. 10.1016/j.immuni.2004.08.009. [DOI] [PubMed] [Google Scholar]
- 69.Hiscott J, Nguyen TL, Arguello M, Nakhaei P, Paz S. 2006. Manipulation of the nuclear factor-kappaB pathway and the innate immune response by viruses. Oncogene 25:6844–6867. 10.1038/sj.onc.1209941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Matsushima A, Kaisho T, Rennert PD, Nakano H, Kurosawa K, Uchida D, Takeda K, Akira S, Matsumoto M. 2001. Essential role of nuclear factor (NF)-kappaB-inducing kinase and inhibitor of kappaB (IkappaB) kinase alpha in NF-kappaB activation through lymphotoxin beta receptor, but not through tumor necrosis factor receptor I. J. Exp. Med. 193:631–636. 10.1084/jem.193.5.631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Joo M, Hahn YS, Kwon M, Sadikot RT, Blackwell TS, Christman JW. 2005. Hepatitis C virus core protein suppresses NF-kappaB activation and cyclooxygenase-2 expression by direct interaction with IkappaB kinase beta. J. Virol. 79:7648–7657. 10.1128/JVI.79.12.7648-7657.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Luftig M, Yasui T, Soni V, Kang MS, Jacobson N, Cahir-McFarland E, Seed B, Kieff E. 2004. Epstein-Barr virus latent infection membrane protein 1 TRAF-binding site induces NIK/IKK alpha-dependent noncanonical NF-kappaB activation. Proc. Natl. Acad. Sci. U. S. A. 101:141–146. 10.1073/pnas.2237183100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Pietrek M, Brinkmann MM, Glowacka I, Enlund A, Havemeier A, Dittrich-Breiholz O, Kracht M, Lewitzky M, Saksela K, Feller SM, Schulz TF. 2010. Role of the Kaposi's sarcoma-associated herpesvirus K15 SH3 binding site in inflammatory signaling and B-cell activation. J. Virol. 84:8231–8240. 10.1128/JVI.01696-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Chu ZL, Shin YA, Yang JM, DiDonato JA, Ballard DW. 1999. IKKgamma mediates the interaction of cellular IkappaB kinases with the tax transforming protein of human T cell leukemia virus type 1. J. Biol. Chem. 274:15297–15300. 10.1074/jbc.274.22.15297. [DOI] [PubMed] [Google Scholar]
- 75.Field N, Low W, Daniels M, Howell S, Daviet L, Boshoff C, Collins M. 2003. KSHV vFLIP binds to IKK-gamma to activate IKK. J. Cell Sci. 116:3721–3728. 10.1242/jcs.00691. [DOI] [PubMed] [Google Scholar]
- 76.Harhaj EW, Sun SC. 1999. IKKgamma serves as a docking subunit of the IkappaB kinase (IKK) and mediates interaction of IKK with the human T-cell leukemia virus Tax protein. J. Biol. Chem. 274:22911–22914. 10.1074/jbc.274.33.22911. [DOI] [PubMed] [Google Scholar]
- 77.Jin DY, Giordano V, Kibler KV, Nakano H, Jeang KT. 1999. Role of adapter function in oncoprotein-mediated activation of NF-kappaB. Human T-cell leukemia virus type I Tax interacts directly with IkappaB kinase gamma. J. Biol. Chem. 274:17402–17405. [DOI] [PubMed] [Google Scholar]