Specific nucleoporins and nuclear pore complex–dependent transport events directly influence aging in yeast.
Abstract
The eukaryotic nuclear permeability barrier and selective nucleocytoplasmic transport are maintained by nuclear pore complexes (NPCs), large structures composed of ∼30 proteins (nucleoporins [Nups]). NPC structure and function are disrupted in aged nondividing metazoan cells, although it is unclear whether these changes are a cause or consequence of aging. Using the replicative life span (RLS) of Saccharomyces cerevisiae as a model, we find that specific Nups and transport events regulate longevity independent of changes in NPC permeability. Mutants lacking the GLFG domain of Nup116 displayed decreased RLSs, whereas longevity was increased in nup100-null mutants. We show that Nup116 mediates nuclear import of the karyopherin Kap121, and each protein is required for mitochondrial function. Both Kap121-dependent transport and Nup116 levels decrease in replicatively aged yeast. Overexpression of GSP1, the small GTPase that powers karyopherin-mediated transport, rescued mitochondrial and RLS defects in nup116 mutants and increased longevity in wild-type cells. Together, these studies reveal that specific NPC nuclear transport events directly influence aging.
Introduction
Aging is a degenerative process caused by an accumulation of damage that leads to reduced viability or survival. Because age is a risk factor for a variety of human disorders including cancer, heart attack, and neurodegenerative disease (López-Otín et al., 2013), defining the molecular and cellular causes of aging is important for understanding approaches to improve health. Nuclear pore complexes (NPCs) are large macromolecular assemblies embedded in the nuclear envelope that regulate transport between the nucleus and cytoplasm (Wente and Rout, 2010) and change as metazoans age (D’Angelo et al., 2009; Toyama et al., 2013). Moreover, in the premature aging Hutchinson–Gilford progeria syndrome (HGPS), patient fibroblasts show nuclear import defects and reduced nuclear levels of the transport factor Ran (Kelley et al., 2011; Snow et al., 2013). Investigating the mechanisms by which NPCs and nuclear transport impact aging will require directly manipulating NPC proteins (nucleoporins [Nups]) and transport factors and assessing the impact on longevity.
NPCs maintain a selective permeability barrier that allows ions, metabolites, and macromolecules to passively diffuse in and out of the nucleus (Wente and Rout, 2010). The rate of diffusion is inversely proportional to the size of the molecule and in the absence of their specific transport, those larger than roughly 50 kD are generally excluded from the nuclei of metazoan cells (Keminer and Peters, 1999). Large proteins and cargoes that accumulate against a concentration gradient are actively and directionally transported, which requires transport receptor interactions with NPCs and energy-dependent events on the nuclear or cytoplasmic face of NPCs (Moore and Blobel, 1993; Rexach and Blobel, 1995; Tran et al., 2007). Others report that NPC selectivity decreases with age, as isolated nuclei from aged rats are permeable to larger dextrans excluded from young nuclei (D’Angelo et al., 2009). Moreover, tubulin aggregates, which have been linked to some neurodegenerative diseases, are detected in these “leaky” aged nuclei. It has therefore been suggested that changes in NPC permeability might contribute to aging.
Inhibited NPC function in aged metazoan cells also correlates with age-linked changes in NPC composition. NPCs are composed of multiple copies of ∼30 distinct Nups, many of which stably associate with minimal protein turnover after their assembly into NPCs. For example, in terminally differentiated rat cells, several scaffolding Nups are still present in the NPCs over a year after being produced (Savas et al., 2012). This low exchange rate for several Nups will inherently limit replacement of subunits damaged during the aging process. Indeed, oxidized forms of the scaffolding Nup93 are detectable in aged rat brains (D’Angelo et al., 2009).
Interestingly, aged nuclei with increased NPC permeability have lower levels of FG Nups (D’Angelo et al., 2009). The FG Nups harbor domains enriched in phenylalanine-glycine (FG) repeats that facilitate nuclear transport (Strawn et al., 2004; Terry and Wente, 2007; Stelter et al., 2011) by transiently interacting with Karyopherin (Kap) transport receptors. The Kaps are responsible for binding cargo with specific NLSs or nuclear export signals. GLFG Nups are a subset of FG proteins that are required for both maintaining the NPC passive permeability barrier and facilitating Kap transport of cargo–adaptor complexes (Terry and Wente, 2007; Laurell et al., 2011; Hülsmann et al., 2012). Thus, both structural Nups and FG Nups are potentially key to the aging mechanism.
Together, much progress has been made in defining important correlations between longevity, NPCs, and nuclear transport. Thus far, however, there is no direct causal evidence that altering NPC function can regulate organismal or cellular life span. To address this question, we have analyzed the replicative life span (RLS) of Saccharomyces cerevisiae as a model to test whether NPCs directly affect longevity. S. cerevisiae cells divide asymmetrically, producing a mother and daughter with different properties; RLS is defined as the number of daughters that bud from a mother before senescence or death (Park et al., 2002). RLS is a robust model for metazoan aging as a result of the conservation of several longevity pathways (Steinkraus et al., 2008; Schleit et al., 2012). By directly manipulating NPCs in S. cerevisiae in this study, we reveal that specific FG Nups and nuclear transport events regulate aging.
Results
Specific GLFG domains regulate RLS
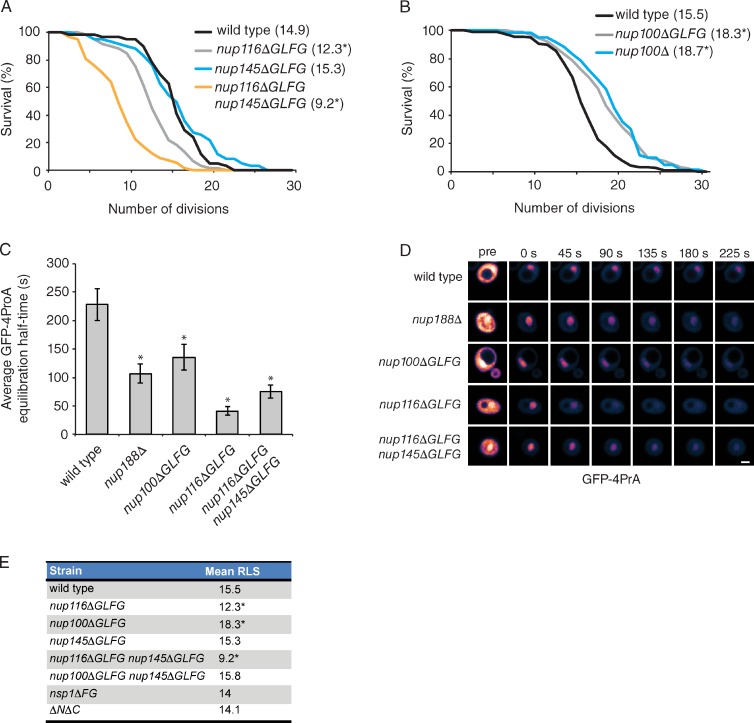
Based on previous studies showing the composition and functions of NPCs change as cells and organisms age (D’Angelo et al., 2009; Kelley et al., 2011), we hypothesized altering NPC function might affect RLS in S. cerevisiae. To test this, we focused on measuring the RLSs of a subset of NPC mutant strains lacking specific FG domains (ΔFG; Strawn et al., 2004; Terry and Wente, 2007). The mean RLS of nup116ΔGLFG mutants was decreased ∼20% when compared with wild-type W303 cells (Fig. 1 A). Although RLS was unaffected in nup145ΔGLFG cells, mean RLS was further reduced 20% in nup116ΔGLFG nup145ΔGLFG double mutants (Fig. 1 A). Surprisingly, deletion of NUP100 (nup100Δ) or its GLFG domain (nup100ΔGLFG) significantly increased mean RLS 17% in W303 (Fig. 1 B) and 23% in BY4741 cells (Fig. S1 A). In contrast, RLS was not significantly altered in nsp1ΔFG, nup145ΔGLFG nup100ΔGLFG, or ΔN ΔC (a strain lacking the five FG domains from Nup42, Nup159, Nup1, Nup2, and Nup60) mutants (Fig. 1 E). Immunoblotting of total cell lysates showed the levels of Nup116 and Nup100 were not significantly altered in nup116ΔGLFG nup145ΔGLFG and nup100ΔGLFG cells (Fig. S1, C and D). Together, these results suggested there are specific defects associated with nup116ΔGLFG, nup116ΔGLFG nup145ΔGLFG, and nup100ΔGLFG cells that alter RLS, and demonstrated longevity is regulated by NPC structure and function in S. cerevisiae.
Figure 1.
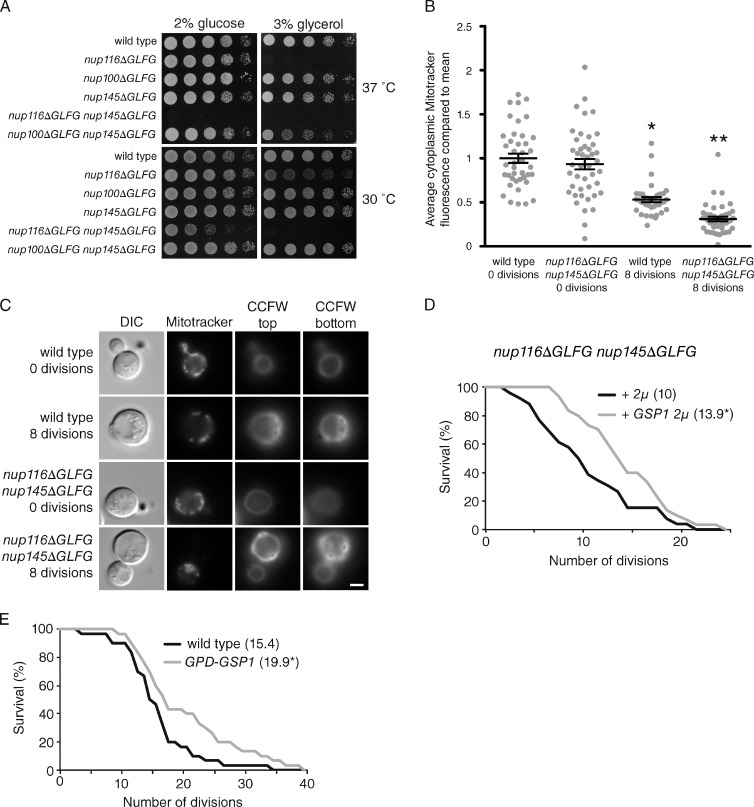
Specific GLFG domains regulate RLS independently of NPC permeability. (A and B) Survival curves of wild-type and mutant cells (n ≥ 50) grown at 30°C. Mean RLSs are listed in parentheses next to genotypes. *, P < 0.001 when the curve is compared with wild type using a log-rank test. (C) Mean half-times for GFP-4PrA equilibration after cytoplasmic bleaching in the listed strains. *, P < 0.01 using Tukey’s post-hoc test when compared with wild-type cells after a one-way analysis of variance (ANOVA; n ≥ 10 cells per strain). Error bars represent SEM. (D) Representative time-lapse FRAP images of single cells expressing GFP-4PrA at the listed times relative to the bleach. Bar, 2.5 µm. (E) A list of mutant strains and their mean RLSs. *, P < 0.05 when the survival curve is compared with wild type using a log-rank test with n ≥ 50 cells.
Because the permeability of NPCs increases as a function of age in metazoans (D’Angelo et al., 2009), a novel live-cell FRAP assay was used to determine whether changes in passive NPC permeability correlate with the RLSs of ΔFG mutants. Mutant strains were transformed with a GFP-4PrA reporter, which localizes to both the nucleoplasm and cytoplasm because its size (molecular mass of 53.5 kD) and conformation allow passive diffusion through NPCs. The cytoplasmic signal was bleached, and subsequent images were used to visualize the diffusion of unbleached GFP-4PrA out of the nucleus until equilibration was reached. The amount of time required for equilibration is inversely proportional to the permeability of NPCs. nup188Δ cells, which have a reported NPC permeability defect (Shulga et al., 2000), were used as a positive control. The mean half-time of GFP-4PrA equilibration in wild-type cells was 228 s and was significantly decreased to 107 s in nup188Δ cells (Fig. 1, C and D). nup116ΔGLFG cells displayed the most severe permeability defect with a mean half-time of 41 s, whereas nup100ΔGLFG and nup116ΔGLFG nup145ΔGLFG cells also had significant increases in NPC permeability (Fig. 1, C and D). These results were consistent with previous studies showing that GLFG domains are essential to maintain the passive permeability barrier of NPCs (Laurell et al., 2011; Hülsmann et al., 2012). Strikingly, there was only a poor correlation (Pearson’s r correlation of 0.58) between RLS and NPC permeability. Thus, changes in NPC permeability were not causative for aging and permeability could not be the sole NPC-related factor influencing RLS.
KAP121 is required for RLS
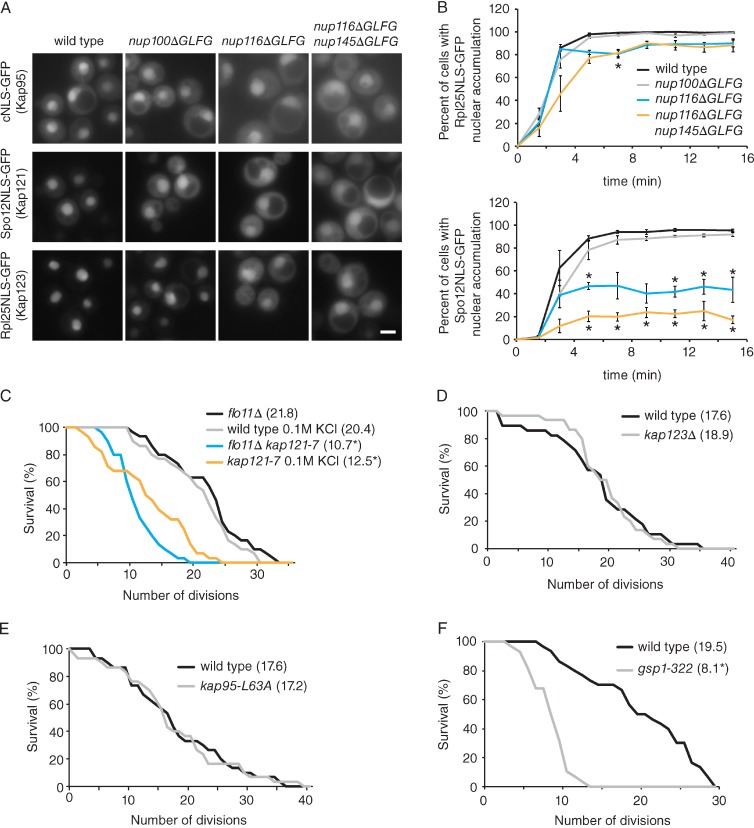
In addition to maintaining the NPC permeability barrier, different GLFG domains are required for specific nuclear transport pathways (Strawn et al., 2004; Terry and Wente, 2007; Stelter et al., 2011). Because RLS did not correlate strongly with NPC permeability, we investigated whether the GLFG domains of Nup116 and Nup100 might mediate specific nuclear transport events that affect longevity. Fluorescence microscopy was used to determine the steady-state localization of different NLS-GFP reporters expressed in wild-type, nup100ΔGLFG, nup116ΔGLFG, and nup116ΔGLFG nup145ΔGLFG cells. Reporters tested include the Srp1-Kap95 substrate classical NLS (cNLS)–GFP (Enenkel et al., 1995; Shulga et al., 1996), the Kap123 substrate Rpl25NLS-GFP (Rout et al., 1997; Schlenstedt et al., 1997; Timney et al., 2006), and the Kap121 substrate Spo12NLS-GFP (Chaves and Blobel, 2001). nup100ΔGLFG cells displayed minor, though detectible amounts of each NLS reporter mislocalized in the cytoplasm compared with wild-type cells (Fig. 2 A), potentially caused by increased NPC permeability. Other than cNLS-GFP, nup116ΔGLFG and nup116ΔGLFG nup145ΔGLFG cells exhibited high levels of each reporter in the cytoplasm (Fig. 2 A), indicating the GLFG domain of Nup116 is required for Kap121- and Kap123-mediated import.
Figure 2.
KAP121 is required for full RLS. (A) Steady-state localization of NLS-GFP reporters in the listed mutant strains at 25°C. Bar, 2.5 µm. (B) Import kinetics of NLS-GFP reporters in wild-type and mutant cells at 25°C. Rates were calculated by measuring the percentage of cells with nuclear accumulation of the reporter at each time point. Error bars represent SEM from at least three independent experiments. *, P < 0.05 when compared with wild type using Bonferri’s post-hoc test after a two-way ANOVA. (C) Survival curves of flo11Δ and flo11Δ kap121-7 (n = 30) cells at 30°C, as well as wild-type and kap121-7 cells supplemented with 0.1 M KCl (n ≥ 25) at 30°C. *, P < 0.001 when compared with wild type/flo11Δ cells using a log-rank test. (D and E) Survival curves of the listed strains (n = 30) grown at 30°C. (F) Survival curves of wild-type and gsp1-322 cells grown at 30°C. *, P < 0.0001 using a log-rank test when the curve is compared with wild-type cells with n = 30 cells. Mean RLSs are listed in parentheses.
Based on their altered steady-state localizations, the relative import rates of Rpl25NLS- and Spo12NLS-GFP were measured in wild-type and ΔFG cells. These assays were especially important given the fact the NPCs of ΔFG strains were also more permeable (Fig. 1, C and D), which might alter the steady-state localization of each reporter. Cells were incubated in 10 mM sodium azide and 10 mM deoxyglucose at 25°C for 30 min to inhibit active transport and equilibrate the reporters between the nucleus and cytoplasm and then transferred to glucose media to initiate transport (Shulga et al., 1996). Relative import rates were calculated by measuring the percentage of cells with GFP accumulated in the nucleus at different time points after recovery. Wild-type and nup100ΔGLFG cells had similar import rates for both reporters (Fig. 2 B). The rate of Kap123-mediated import in nup116ΔGLFG cells was largely comparable to wild type; however, it was delayed in nup116ΔGLFG nup145ΔGLFG cells. Kap121-mediated import was significantly impaired in both nup116ΔGLFG and nup116ΔGLFG nup145ΔGLFG mutants; many of these cells did not accumulate Spo12NLS-GFP in their nucleus even after 15 min of recovery. There was a strong correlation (Pearson’s r correlation of 0.86) between disrupted Kap121-mediated import and decreased RLS. Consistent with our previous study (Terry and Wente, 2007), these results showed the GLFG domain of Nup116 is required for Kap121-mediated import, and it also plays a role in Kap123-mediated import along with Nup145.
The RLSs of kap121-7 (Seedorf and Silver, 1997) and kap123Δ (Giaever et al., 2002) cells were assessed to directly test whether inhibited Kap121- or Kap123-mediated import decreased longevity. Deletion of the nonessential KAP123 produced no obvious changes in RLS at 30°C (Fig. 2 D), indicating that Kap123 does not significantly regulate longevity. A significant portion of kap121-7 cells entered a filamentous growth state at 30°C, causing the cells to stick together and preventing accurate measurement of their RLS. Deletion of FLO11, a nonessential gene required for filamentous growth (Lo and Dranginis, 1998), suppressed this phenotype in kap121-7 cells. Compared with flo11Δ cells, which had a mean RLS of 21.8 divisions, flo11Δ kap121-7 cells had a significantly decreased mean RLS of 10.7 divisions (Fig. 2 C). Similar results were also observed for wild-type and kap121-7 cells analyzed on 0.1 M KCl, which also largely suppressed filamentous growth (Fig. 2 C). Using a different kap121-21 mutant (Seedorf and Silver, 1997), we observed even more pronounced replicative aging defects when grown on 0.1 M KCl (unpublished data). We found there was no significant change in RLS in kap95-L63A mutants at 30°C (Fig. 2 E), suggesting specific transport pathways regulate life span in yeast. Together, these experiments revealed that KAP121 is required for yeast longevity and suggested that the Kap121-mediated transport defects of nup116ΔGLFG and nup116ΔGLFG nup145ΔGLFG mutants contribute to their decreased RLSs.
Because Ran-GTP dissociates Kap121–cargo (and other Kap–cargo) complexes once they enter the nucleus (Rexach and Blobel, 1995; Izaurralde et al., 1997), we predicted that inhibiting Ran would also decrease RLS. The RLSs of wild-type and gsp1-322 (Oki et al., 1998) cells were measured at 30°C to test this prediction. GSP1 is an essential gene that codes for one of two Ran orthologues found in S. cerevisiae (Belhumeur et al., 1993); the gsp1-322 allele is a temperature-sensitive mutant with a growth defect at 30°C. Wild-type cells had a mean RLS of 19.5 divisions (Fig. 2 F), whereas the mean RLS of gsp1-322 cells was significantly decreased to 8.1 divisions (Fig. 2 F). This indicated that Ran function is required for RLS and is consistent with NPC-mediated transport regulating life span.
NPCs are altered in replicatively aged cells
Given our experiments that demonstrated altering NPC function can affect RLS, we next tested whether NPCs and GLFG Nups are affected during the aging process of wild-type S. cerevisiae cells. Because there are lower amounts of FG Nups at the nuclear rim in aged rat cells, a phenotype that correlates with increased NPC permeability (D’Angelo et al., 2009), we hypothesized FG and GLFG Nup levels might be decreased in replicatively aged yeast. This possibility seemed particularly likely because NPCs with decreased levels of Nup82 appear to be selectively retained in mother yeast cells (Makio et al., 2013). A biochemical enrichment technique (Park et al., 2002) was used to isolate populations of young (∼0–1 divisions) and aged (∼6–9 divisions) cells. Replicative age was assessed by counting the number of bud scars visualized by calcofluor white (CCFW) staining, and immunoblotting was used to measure relative protein levels.
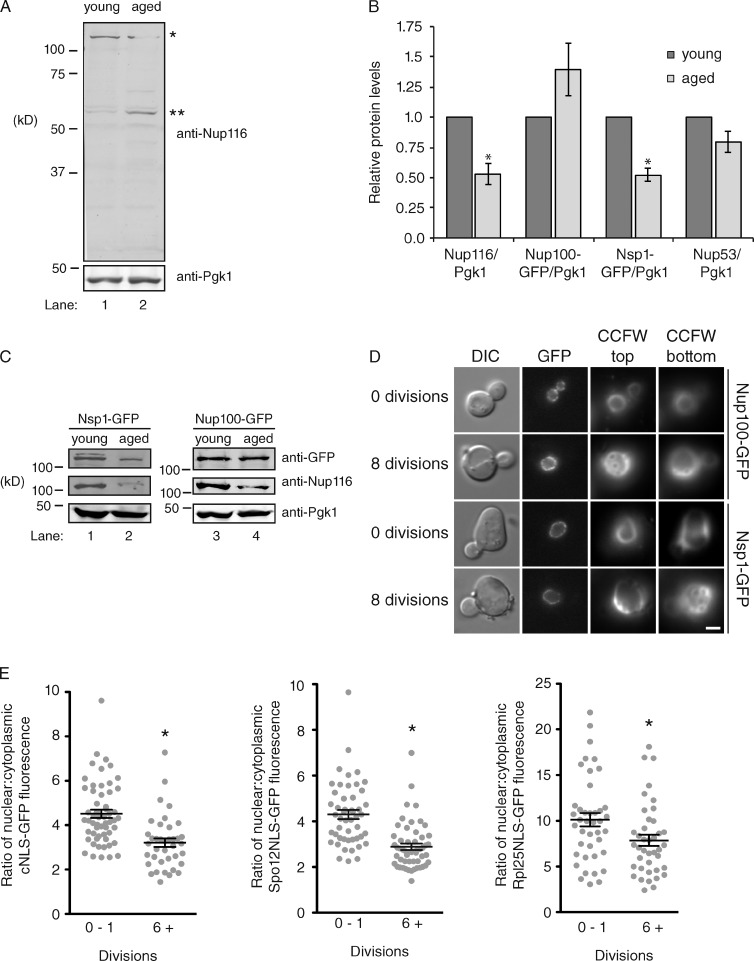
Significantly reduced amounts of full-length Nup116 were detected in aged cells (Fig. 3, A–C) using an antibody that recognizes the C-terminal region of Nup116. The amount of Pgk1 did not significantly decrease in aged cells (Fig. 3, A and B), indicating all cellular proteins were not similarly affected. We also detected a lower migrating band in aged cells in the blot in Fig. 3 A, likely corresponding to a degradation product of Nup116 lacking a portion of its N-terminal region (which includes FG and GLFG domains). Full-length Nsp1-GFP levels were also decreased in aged yeast, whereas the amounts of Nup100-GFP and the scaffolding Nup53 were unaffected (Fig. 3, B and C; and Fig. S2 A). Quantitative RT-PCR demonstrated that the amounts of Nup100, Nup116, Nup53, and Nsp1 transcripts were not significantly altered in aged cells (Fig. S2 B), suggesting decreased Nup116 and Nsp1 protein levels are not caused by reduced mRNA expression. Although levels of full-length Nsp1-GFP decreased in aged cells, the localization of Nsp1-GFP was largely unaffected, with the majority of Nsp1-GFP signal present at the nuclear rim (Fig. 3 D). The localization of Nup100-GFP was also unchanged in replicatively aged yeast (Fig. 3 D). A ratiometric analysis of Nup100-GFP and Nsp1-mCherry levels in young and aged cells was also performed, which showed a decrease in the amount of Nsp1-mCherry relative to Nup100-GFP when comparing young and aged cells (Fig. S2 C). We therefore concluded that full-length levels of Nup116 and Nsp1 decrease in replicatively aged mothers.
Figure 3.
NPCs and nuclear transport deteriorate in replicatively aged cells. (A) Enriched mother (∼6–9 divisions) and young (∼0–1 divisions) wild-type cells were lysed, and immunoblotting was conducted using anti-Pgk1 and anti-Nup116 C-terminal antibodies. One asterisk corresponds to full-length Nup116, whereas two asterisks correspond to a degradation product. (B) Quantification of Nup116, Nup53, Nup100-GFP, and Nsp1-GFP levels in young (∼0–1 divisions) and aged (∼6–9) cells compared with Pgk1. The Nup/Pgk1 ratio was set as 100% in young cells. Error bars represent SEM from at least three independent experiments. *, P < 0.05 when compared with young cells using a two-tailed Student’s t test. (C) Enriched aged (∼6–9 divisions) and young (∼0–1 divisions) Nup100-GFP or Nsp1-GFP cells were lysed and immunoblotting conducted using anti-GFP, anti-Pgk1, and anti-Nup116 C-terminal antibodies. (D) Zero or eight division cells from the listed strains were stained with CCFW and visualized using fluorescence microscopy. CCFW images are from the cortical regions of the cell to show most bud scars. Bar, 2.5 µm. (E) The nuclear versus cytoplasmic ratios of the listed NLS-GFP reporters were calculated in young (zero divisions) or aged (six or more divisions) cells using ImageJ. Error bars represent SEM, and *, P < 0.03 using a two-tailed Student’s t test when compared with young cells. At least 10 cells from each condition were measured in three independent experiments.
We predicted that nuclear transport would be inhibited in aged cells caused by decreased levels of full-length Nsp1 and Nup116. NPC function was assessed by measuring the ratio of GFP fluorescence in the nucleus versus cytoplasm for GFP-NLS reporters in wild-type cells that had divided zero or more than five times. Cytoplasmic levels of Spo12NLS-GFP, Rpl25NLS-GFP, and cNLS-GFP increased in aged cells (Fig. 3 E). The fact that Spo12NLS-GFP and Rpl25NLS-GFP reporters were mislocalized when there were decreased levels of full-length Nup116 in aged cells was consistent with our observation that the import of these GFP reporters was delayed in nup116ΔGLFG nup145ΔGLFG mutants (Fig. 2 B). Mislocalization of cNLS-GFP suggested that other NPC functions are compromised in aged cells because cNLS-GFP import was not significantly dependent on the GLFG region of Nup116 (Fig. 2 A). Together, these experiments indicated nuclear transport is inhibited in replicatively aged yeast, which is likely caused by decreased levels of full-length and/or functional Nup116 and Nsp1.
ΔFG mutants regulate RLS independently of characterized longevity factors
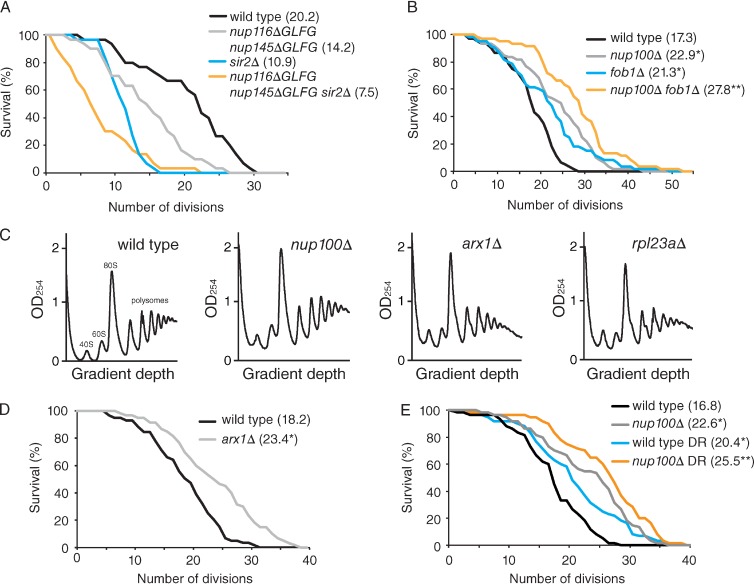
We next investigated whether signaling pathways known to regulate RLS were affected by ΔFG mutants. Extrachromosomal ribosomal DNA circles (ERCs) are a cause of replicative aging in yeast (Sinclair and Guarente, 1997), and factors that influence ERC formation significantly alter RLS. Sir2 inhibits ERC production and RLS is accordingly decreased in sir2Δ cells (Kaeberlein et al., 1999), whereas fob1Δ cells have increased RLSs because Fob1 promotes ERC formation (Defossez et al., 1999). Because Sir2 and Fob1 both localize to the nucleolus (Gotta et al., 1997; Defossez et al., 1999), their nuclear import could be disrupted in ΔFG strains. The RLSs of sir2Δ, nup116ΔGLFG nup145ΔGLFG, and sir2Δ nup116ΔGLFG nup145ΔGLFG mutants were measured to ascertain whether the decreased RLSs of nup116ΔGLFG nup145ΔGLFG cells were solely caused by inhibition of Sir2. If Sir2 is significantly impaired in nup116ΔGLFG nup145ΔGLFG cells, sir2Δ nup116ΔGLFG nup145ΔGLFG cells should have similar RLSs compared with nup116ΔGLFG nup145ΔGLFG cells. We found that the mean RLS of sir2Δ nup116ΔGLFG nup145ΔGLFG cells was significantly decreased compared with both sir2Δ (31%) and nup116ΔGLFG nup145ΔGLFG cells (48%; Fig. 4 A), indicating the GLFG domains of Nup116 and Nup145 regulate RLS independently of Sir2. Similar experiments were performed to determine whether Fob1 activity is inhibited in nup100Δ cells, which could explain why deletion of NUP100 increased RLS. Both fob1Δ and nup100Δ mutants had increased RLSs relative to wild-type cells (Fig. 4 B). Because mean RLS was further increased ∼20% in fob1Δ nup100Δ cells (Fig. 4 B), Nup100 inhibited RLS independently of Fob1. SIR2 is not required for the extended life span of nup100Δ mutants, as nup100Δ sir2Δ cells have increased RLSs compared with sir2Δ cells (Fig. S3 B). Similarly, RLSs of nup116ΔGLFG cells are extended when FOB1 is deleted (Fig. S3 A). These experiments suggested the GLFG regions of Nup100 and Nup116 do not significantly affect Sir2 or Fob1 function.
Figure 4.
ΔFG mutants regulate RLS independently of SIR2, FOB1, and DR. (A) Survival curves for the listed strains grown at 30°C. P < 0.02 when comparing all curves to one another using a log-rank test (n = 30 cells). (B) Survival curves for the listed strains grown at 30°C. *, P < 0.02 when the curve is compared with wild type using a log-rank test; **, P < 0.01 when the curve is compared with fob1Δ or nup100Δ (n ≥ 50 cells). (C) Representative polysome profiles of the listed strains; at least three separate profiles were performed for each strain. 40S, 60S, 80S, and polysome peaks are labeled in the wild-type profile. (D) Survival curves for wild-type and arx1Δ cells grown at 30°C. *, P < 0.001 when the curve is compared with wild type using a log-rank test with n ≥ 50 cells. (E) Survival curves for wild-type and nup100Δ cells grown on 2% or 0.05% glucose (DR) at 30°C, with mean RLSs listed in parentheses. *, P < 0.01 when compared with wild type using a log-rank test; **, P < 0.04 when compared with nup100Δ or wild-type DR (n ≥ 50 cells).
Inhibition of GSP1 function using the gsp1-322 mutant largely suppressed the extended RLSs of nup100Δ cells (compare with nup100Δ kap121-7 cells; Fig. S3, C and D), suggesting that Nup100 mediates one or more nuclear transport pathways that affect longevity. RLS increases in S. cerevisiae when 60S ribosomal subunit levels are decreased by genetic or chemical means (Steffen et al., 2008); we therefore speculated that the increased RLSs of nup100Δ and nup100ΔGLFG cells could be explained by inhibited preribosomal subunit export, which might decrease 60S subunit levels. We initially tested whether inhibiting ribosome export affected RLS using an arx1Δ mutant, which is partially defective in nuclear export of pre-60S subunits (Bradatsch et al., 2007; Hung et al., 2008). The mean RLS of arx1Δ cells was increased roughly 28% compared with wild-type cells (Fig. 4 D), showing that inhibited ribosome export increases longevity in yeast. Polysomes were next analyzed to determine whether 60S subunit levels were decreased in nup100Δ and arx1Δ cells. rpl23aΔ cells, which have a reported decrease in 60S subunit levels (Steffen et al., 2008), were used as a positive control. Polysome profiles of wild-type cells showed increased amounts of 60S subunits compared with 40S subunits (Fig. 4 C). The profiles of rpl23aΔ- and arx1Δ-null cells displayed roughly similar amounts of 40S and 60S subunits as well as half-mers (Fig. 4 C), both of which are indicative of decreased 60S subunit levels (Rotenberg et al., 1988) and are consistent with previous profiles of arx1Δ cells (Hung et al., 2008). The polysome profiles of nup100Δ cells appeared similar to wild type (Fig. 4 C), suggesting that Nup100 is not required for export of pre-60S subunits. These results indicated that decreased 60S export can affect longevity, although Nup100 does not regulate longevity by altering this process.
Others have also shown that decreasing levels of 60S ribosome subunits extends RLS through activation of dietary restriction (DR) pathways (Steffen et al., 2008). DR refers to reducing the amount of caloric intake of an organism (typically 75–95% in yeast, and 20–40% in mammals) and increases RLS in most yeast backgrounds as well as longevity in worms, flies, and mice (Schleit et al., 2012). We analyzed the RLSs of wild-type and nup100Δ BY4741 cells grown on 0.05% glucose to determine whether Nup100 regulates another aspect of DR. This particular yeast background was used because the RLSs of wild-type BY4741 cells have consistently been shown to increase when grown on low glucose (Schleit et al., 2012). As expected, the mean RLS of wild-type cells increased when grown on 0.05% glucose (Fig. 4 E). The mean RLS of nup100Δ cells (Fig. 4 E) grown on 0.05% glucose was greater than the mean RLS of wild-type DR cells (25%) and nup100Δ cells grown on 2% glucose (13%). Thus, Nup100 inhibited RLS through a mechanism independent of DR.
Nup116 and Gsp1 influence life span through regulation of mitochondrial function
Alterations in mitochondrial function influence longevity in a variety of model organisms including S. cerevisiae. For example, mitochondria tend to become more fragmented and less functional as yeast replicatively age; mutations that inhibit this deterioration significantly increase RLS (Scheckhuber et al., 2007; Hughes and Gottschling, 2012). We speculated that strains with decreased life spans might display reduced mitochondrial function. This was tested by serially diluting different ΔFG strains on glucose or glycerol. S. cerevisiae mutants with compromised mitochondrial activity can grow on fermentable carbon sources such as glucose because respiration is largely not used for energy production but display specific growth defects on nonfermentable carbon sources that require respiration such as glycerol; this is commonly referred to as a petite phenotype.
With the exception of nup116ΔGLFG nup145ΔGLFG cells, which do not grow at temperatures >30°C, other ΔFG mutants displayed normal growth on glucose at 30 and 37°C (Fig. 5 A). At 30°C, nup116ΔGLFG nup145ΔGLFG cells did not grow on glycerol, whereas the growth of nup116ΔGLFG mutants was decreased compared with wild-type cells (Fig. 5 A). Growth of nup116ΔGLFG cells was completely inhibited on glycerol at 37°C. Transformation of nup116ΔGLFG cells with a vector containing full-length NUP116, but not nup116ΔGLFG, rescued glycerol sensitivity (Fig. S4 A). Importantly, this glycerol growth defect was specific to ΔFG mutants with decreased life spans, as nup100ΔGLFG, nup145ΔGLFG, and nup100ΔGLFG nup145ΔGLFG cells grew normally on glycerol at all temperatures tested (Fig. 5 A). Ran is also required for mitochondrial activity, as gsp1-322 mutants also display a petite phenotype (Fig. S4 C). Other ΔFG mutants with wild-type RLSs also grew on glucose and glycerol at 37°C (Fig. S4 B). Thus, mutants lacking the GLFG region of Nup116 displayed reduced mitochondrial function.
Figure 5.
The mitochondria of nup116ΔGLFG nup145ΔGLFG cells exhibit morphological and functional defects. (A) Strains were serially diluted onto glucose or glycerol and incubated at 30 or 37°C until grown as shown. (B) Enriched mother (eight divisions) and young cells (zero divisions) were stained with MitoTracker red CMXRos. ImageJ was used to quantify the mean fluorescence intensity of each cell; ≥10 cells per condition were quantified from three independent experiments. *, P < 0.01 when compared with wild-type zero divisions using Tukey’s post-hoc test after a one-way ANOVA. **, P < 0.01 when compared with wild-type eight divisions using Tukey’s post-hoc test. Error bars represent SEM. (C) Representative single cells from B stained using CCFW and MitoTracker red CMXRos. CCFW images show the top and bottom of cells for visualization of most bud scars. DIC, differential interference contrast. Bar, 2.5 µm. (D) Survival curves of nup116ΔGLFG nup145ΔGLFG cells transformed with an empty or GSP1 2µ vector grown at 30°C. *, P < 0.04 when the curves are compared using a log-rank test (n ≥ 25 cells). (E) Survival curves of wild-type cells or those overexpressing GSP1 under a TDH3 promoter grown at 30°C. *, P < 0.05 when compared with wild type using a log-rank test with n ≥ 30 cells. Mean RLSs are listed in parentheses.
ΔFG strains were stained with the mitochondrial membrane potential (ΔΨ)–dependent dye MitoTracker red CMXRos to detect obvious changes in morphology or membrane potential. The mitochondria of wild-type cells were largely localized to the periphery of the cell, although there were often large regions lacking any detectable MitoTracker signal (Fig. S4 C). About 25% of nup116ΔGLFG nup145ΔGLFG cells exhibited a morphological defect where their mitochondria were distributed uniformly throughout the cortical region of the cell at 25°C. This phenotype was not observed in wild-type or other ΔGLFG strains (Fig. S4, D and E). As a control, localization was also analyzed using nup116ΔGLFG nup145ΔGLFG cells in which the chromosomal copy of the mitochondrial ribosomal protein Mrpl11 (Kitakawa et al., 1997; Tkach et al., 2012; Breker et al., 2013) was GFP tagged (Fig. S4 C). The morphological defect was similar to that observed with MitoTracker staining.
Young and aged wild-type and nup116ΔGLFG nup145ΔGLFG cells were stained with MitoTracker red CMXRos to determine whether age-dependent changes in ΔΨ are affected by nuclear transport events. ΔΨ was quantified by measuring the mean intensity of MitoTracker signal throughout a cell that had divided zero or eight times. Similar to previous analyses (Hughes and Gottschling, 2012), we observed a statistically significant decrease in MitoTracker intensity as wild-type cells replicatively aged (Fig. 5, B and C). Although the mean MitoTracker intensity of wild-type and nup116ΔGLFG nup145ΔGLFG cells was similar in young cells, it was significantly more decreased in nup116ΔGLFG nup145ΔGLFG cells after eight divisions (Fig. 5, B and C). There was no noticeable change in the intensity of Mrpl11-GFP fluorescence when comparing aged wild-type and mutant cells (Fig. S4 F), suggesting membrane potential is specifically affected. This result indicated the GLFG domains of Nup116 and Nup145 influence the function of mitochondria during the aging process.
Overexpression of GSP1 specifically rescued the glycerol sensitivity of nup116ΔGLFG cells without significantly affecting their growth on glucose (Fig. S5, A and B), suggesting that mitochondrial defects in nup116ΔGLFG mutants are caused by disrupted nuclear transport. We assessed whether GSP1 overexpression increased longevity in nup116ΔGLFG nup145ΔGLFG cells to determine whether their decreased RLSs could be caused by inhibited mitochondrial function. GSP1 overexpression significantly extended the RLS of nup116ΔGLFG nup145ΔGLFG mutants (Fig. 5 D), linking mitochondrial activity to life span. Because we observed age-dependent changes in NPCs (Fig. 3), we also tested whether overexpressing GSP1 could also influence the RLS of wild-type cells. Overexpression (Fig. S5 D) increased RLS when cells were grown on either glucose (Fig. 5 E) or glycerol (Fig. S5 C). Importantly, this confirms that nuclear transport can influence RLS and suggests longevity can be increased by limiting the functional deterioration of NPCs during the aging process.
Mitochondrial dysfunction in kap121 cells
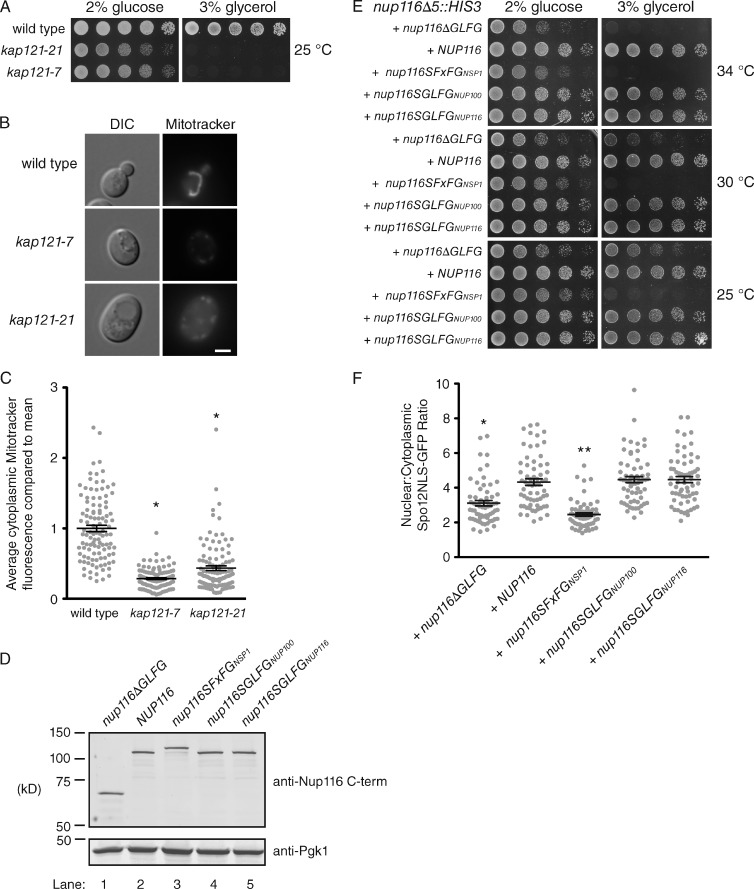
Because Kap121 transport was mediated by Nup116’s GLFG domain (Fig. 2), kap121-7 and kap121-21 mutants were analyzed to determine whether they also displayed inhibited mitochondrial activity. This possibility seemed likely as KAP121 was isolated in an overexpression screen for genes that enhance the import of hydrophobic membrane proteins into the mitochondria (Corral-Debrinski et al., 1999). Both kap121-7 and kap121-21 cells were not viable on glycerol at 25°C (Fig. 6 A), indicative of strong defects in mitochondrial function. Quantification of MitoTracker red CMXRos staining showed the ΔΨ of kap121-7 and kap121-21 mutants was significantly decreased compared with wild-type cells (Fig. 6, B and C). When the brightness was adjusted to help visualize MitoTracker signal in kap121 mutants, the mitochondria also often appeared more fragmented compared with wild-type cells (Fig. S5 E).
Figure 6.
Kap121-mediated transport is required for mitochondrial function. (A) Wild-type and mutant strains were serially diluted onto glucose or glycerol and incubated at 25°C until grown as shown. (B) Single representative wild-type or kap121 mutant cells stained with MitoTracker red CMXRos. DIC, differential interference contrast. Bar, 2.5 µm. (C) Quantification of the mean MitoTracker red CMXRos cytoplasmic intensity for strains listed in B. *, P < 0.01 using Tukey’s post-hoc test when compared with wild type after a one-way ANOVA (n ≥ 90 with 30–40 cells quantified in three separate experiments). (D) Immunoblots of lysates derived from nup116Δ5::HIS3 cells transformed with pRS314 vectors containing the constructs listed above each lane. Lysates were immunoblotted with anti-Pgk1 and anti-Nup116 C-terminal (C-term) antibodies. (E) nup116Δ5::HIS3 cells transformed with pRS314 vectors containing the listed constructs were serially diluted onto glucose or glycerol and incubated at 25, 30, or 34°C until grown as shown. (F) nup116Δ5::HIS3 cells transformed with pRS314 vectors containing the listed constructs as well as Spo12NLS-GFP were visualized using fluorescence microscopy. The mean nuclear/cytoplasmic GFP intensity ratios of ≥33 cells from three different experiments were calculated. *, P < 0.01 when compared with NUP116 using Tukey’s post-hoc test following a one-way ANOVA; **, P < 0.05 when compared with nup116ΔGLFG. Error bars represent SEM.
A series of FG swap mutants were used to define the requirements of Nup116’s GLFG domain in mitochondrial function. In these mutants, the deleted chromosomal copy of nup116Δ5::HIS3 was covered with a plasmid-based construct wherein the GLFG domain of Nup116 was either deleted (nup116ΔGLFG), replaced with its own GLFG domain (nup116SGLFGNup116), replaced with the GLFG domain of Nup100 (nup116SGLFGNup100), or replaced with the FxFG domain of Nsp1 (nup116SFxFGNsp1). Immunoblotting showed all constructs were expressed at equal levels compared with one another and full-length plasmid-based Nup116 (Fig. 6 D). Cells expressing nup116ΔGLFG exhibited minor growth defects on glucose at 25 and 30°C, whereas their growth was largely inhibited on glycerol at 25 and 30°C (Fig. 6 E). Cells expressing full-length NUP116, nup116SGLFGNup116, and nup116SGLFGNup100 all grew similarly on glucose and glycerol, indicating the GLFG domain of Nup100 could functionally substitute for Nup116’s GLFG domain. Substitution of Nsp1’s FxFG domain actually appeared to inhibit respiration more than deletion of the GLFG domain alone, as the nup116SFxFGNsp1 mutant did not grow on glycerol at all temperatures tested (Fig. 6 E).
The FG swap mutants were transformed with Spo12NLS-GFP to test whether Kap121 transport was also rescued by nup116SGLFGNup100. Fluorescence microscopy of steady-state nuclear versus cytoplasmic Spo12NLS-GFP levels showed a strong correlation between inhibited glycerol growth and decreased Kap121 transport (Fig. 6 F), consistent with the requirement of Kap121 in mitochondrial function (Fig. 6, A–C). The fact Nup100’s GLFG domain functionally substituted for Nup116’s implied the location and number of GLFG domains in the NPC (as determined by the C-terminal region of Nup116) is more important than the specific sequence of the GLFG domain. The type of FG domain was critical, however, as Nsp1’s FxFG domain was unable to substitute for Nup116’s GLFG region and might actually inhibit its activity with respect to mitochondrial function.
Discussion
We have shown that specific GLFG Nups and transport events directly regulate longevity in S. cerevisiae. Deletion of Nup116’s GLFG domain significantly decreases RLS, whereas deletion of NUP100 increases RLS. The NPCs of nup116ΔGLFG and nup100ΔGLFG mutants are both more permeable than those of wild-type cells, indicating that NPC permeability does not correlate well with longevity. Defects in specific active transport pathways are linked to decreased RLSs. Kap121-mediated import is inhibited in the nup116ΔGLFG mutant, and kap121 mutants have decreased RLSs. KAP121 and the GLFG domain of Nup116 are also both essential for mitochondrial function in young yeast as well as for maintaining mitochondrial activity during the aging process. Overexpression of GSP1 rescues the mitochondrial defects of nup116ΔGLFG cells and also increases RLS in wild-type cells, likely by limiting the deterioration of NPCs that occurs during the replicative aging process.
We specifically observed NPC deterioration in replicatively aged yeast, potentially caused by direct retention of damaged NPCs in mothers or as an indirect consequence of other factors enriched in aged cells. Metazoan scaffolding Nups have low exchange rates and are thus stably associated with NPCs. Recent work shows that once assembled into NPCs the S. cerevisiae Nup188 and GLFG Nup57 do not exchange over a time course of 3 h (Menendez-Benito et al., 2013). If stably associated Nups are proteolytically or oxidatively damaged, these low exchange rates inherently limit replacement and result in accumulation of such changes in aged cells. Nsp1 is required to overcome a selective barrier at the bud neck for transmitting NPCs into daughter S. cerevisiae cells (Colombi et al., 2013; Makio et al., 2013). When cells are depleted of the Nsp1-binding Nup82, the NPCs lacking Nup82 are retained in mother cells (Makio et al., 2013). Thus, NPCs containing damaged or degraded Nsp1 potentially accumulate in mother cells during the aging process. Furthermore, the basis for our observing less full-length Nup116 in aged cells might be caused by Nup116, but not Nup100, forming a subcomplex with Nsp1, Nup159, and Nup82 (Bailer et al., 2000). Alternatively, other detrimental factors enriched or retained in aged cells might affect NPCs. For example, reactive oxygen species are detectable in replicatively aged yeast without any additional oxidative stress (Laun et al., 2001). Moreover, Caenorhabditis elegans Nup93 is carbonylated in aged worms (D’Angelo et al., 2009), indicating Nups can be affected by oxidative damage.
Our analysis also reveals that both the GLFG domain of Nup116 and Kap121 are required for mitochondrial function and RLS. Because proper import of the Kap121 substrate Spo12NLS-GFP is dependent on Nup116’s GLFG domain (Fig. 2 B), it is likely that the RLSs and mitochondrial defects of nup116ΔGLFG and nup116ΔGLFG nup145ΔGLFG cells are at least partially caused by inhibited Kap121-dependent transport. Mitochondrial function and RLS appear to be functionally linked in NPC mutants as overexpression of GSP1specifically rescues growth on glycerol and also increases RLS of nup116ΔGLFG nup145ΔGLFG cells. We hypothesize that this is important during the aging process in wild-type cells, as GSP1 overexpression also increased life span (Figs. 5 E and S5 C). Because this suggests that some nuclear transport events regulate aging, the decreased life spans of kap121 and nup116ΔGLFG mutants are therefore likely caused by accelerated aging. We hypothesize the decreased life spans of kap121 and nup116ΔGLFG mutants are not caused by stochastic cell death because we did not observe a significant portion failing to divide after one or two cell divisions. Because Kap121-mediated import is significantly inhibited in both nup116ΔGLFG and nup116ΔGLFG nup145ΔGLFG cells (Fig. 2 B), other nuclear transport events required for RLS might also be disrupted in nup116ΔGLFG nup145ΔGLFG cells. Alternatively, nup116ΔGLFG nup145ΔGLFG mutants could be more sensitive to NPC degradation in aged cells because they already lack two GLFG domains. Because KAP121 overexpression potentially promotes mitochondrial membrane import (Corral-Debrinski et al., 1999), we propose that Kap121 mediates the nuclear transport of a factor directly or indirectly required for mitochondrial import of membrane proteins. Future work will be required to fully delineate such factors. We cannot exclude the possibility that Kap121 and Gsp1 regulate RLS independent of their roles in nuclear transport; however, the fact that ΔFG mutants also display altered RLSs is consistent with specific transport events affecting life span.
These findings in S. cerevisiae are informative for ongoing studies in multicellular organisms. Our results suggest that permeability-independent NPC defects, including disrupted Kap121-dependent transport, significantly decrease longevity. We predict that the Nup116 orthologue Nup98 and the Kap121 orthologue Ipo5 (or potentially another importin) are also required for mitochondrial function and longevity in higher eukaryotes. Nuclear import of large cargoes is inhibited in HGPS fibroblasts as a result of Ran mislocalization (Snow et al., 2013), suggesting that disrupted nuclear transport causes some of the phenotypes associated with this disease. Interestingly, glycolytic enzymes such as PGK1 are expressed at higher levels in HGPS fibroblasts compared with wild-type controls, whereas levels of proteins involved in oxidative phosphorylation like cytochrome c are reduced (Rivera-Torres et al., 2013). Moreover, mitochondrial function is impaired in HGPS fibroblasts as well as mouse models of HGPS (Rivera-Torres et al., 2013). This suggests that a conserved set of pathways link mitochondrial function and longevity to NPC- and Ran-dependent transport events.
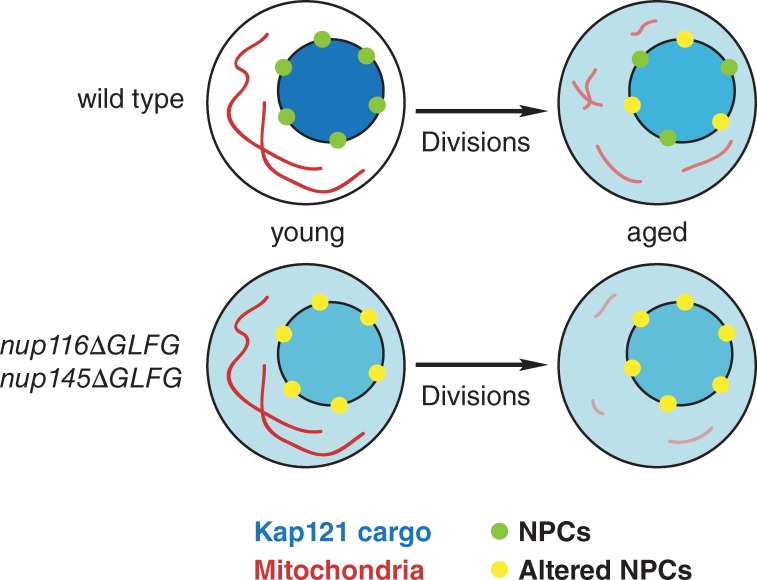
Our results are consistent with a model whereby NPC damage contributes to the aging process in wild-type cells (Fig. 7). Replicatively aged NPCs likely contain partially degraded Nups because full-length Nup116 and Nsp1 levels are decreased in aged cells (Fig. 3 B), even though Nsp1-GFP still localizes to the nuclear rim (Fig. 3 D). Loss of Nup116’s GLFG domain then inhibits Kap121-dependent nuclear import (Fig. 2 B), which compromises mitochondrial function (Figs. 5 and 6) and decreases RLS (Figs. 1 A and 2 D). We speculate that deletion of Nup116’s GLFG domain promotes aging because Kap121-mediated transport is inhibited in young cells, limiting the function of their mitochondria and replicative potential during the aging process. RLS is further reduced in nup116ΔGLFG nup145ΔGLFG cells because of functional redundancy between the GLFG domains of Nup145 and Nup116 (Fig. 1 A). Overexpression of GSP1 suppresses mitochondrial defects in nup116ΔGLFG cells and thus extends the RLS in mutant cells. We hypothesize that GSP1 overexpression increases RLS in wild-type cells by limiting some of the nuclear transport defects observed in aged cells.
Figure 7.
NPCs regulate RLS by influencing specific nuclear transport pathways. Functional NPCs in young wild-type cells properly mediate Kap121-dependent nuclear import. Degradation or loss of Nup116 and Nsp1 causes deterioration of NPC function in replicatively aged wild-type cells, which inhibits nucleocytoplasmic transport and likely increases NPC permeability. Along with other signaling pathways that regulate longevity, inhibited Kap121 transport decreases mitochondrial membrane potential and reduces RLS. Mitochondrial membrane potential is similar to wild type in young nup116ΔGLFG nup145ΔGLFG cells but is reduced faster during the aging process because Kap121-dependent transport is inhibited in young cells. Accelerated reduction in membrane potential is likely at least partially responsible for the decreased RLSs of nup116ΔGLFG nup145ΔGLFG cells.
We also discovered that deletion of Nup100 or its GLFG domain increased longevity independently of Kap121 (Fig. 2 B), pre-60S ribosome export, DR, and Fob1 (Fig. 3). This is surprising given the high degree of conservation between Nup100 and Nup116 (Wente et al., 1992). Because the GLFG domain of Nup100 can functionally replace Nup116’s GLFG region (Fig. 6, E and F), we speculate the amounts and/or location of GLFG Nups in the NPC affect how they regulate transport. Because deletion of Nup100’s GLFG domain is sufficient to increase RLS (Fig. 1 B), Nup100 likely mediates one or more nuclear transport pathways that normally inhibit longevity. Consistent with this hypothesis, deletion of NUP100 had only a marginal effect on RLS in a gsp1-322 mutant (Fig. S3 C). Overexpressed Nup100-GFP can form prions that aggregate in cytoplasmic foci (Halfmann et al., 2012); these aggregates could influence RLS. However, we did not observe any Nup100-GFP cytoplasmic foci in young or replicatively aged cells (unpublished data), and additional work will be required to ascertain Nup100’s function in RLS.
In summary, we reveal specific roles for Nups in regulating RLS in S. cerevisiae. This provides key insights into the mechanisms for how NPC degradation potentially accelerates the aging process. Changes in NPC permeability alone are not sufficient, and we identify an important functional link between Kap121, Nup116, and mitochondrial activity in aging. Revealing if and how other NPC functions are involved in longevity is a future goal.
Materials and methods
Yeast strains and plasmids
Strains and plasmids used in this study are listed in Tables S1 and S2. Yeast were grown in synthetic complete (SC) media without appropriate amino acids supplemented with 2% dextrose or YPD (1% yeast extract, 2% peptone, 2% dextrose). Cloning was performed using standard molecular biology strategies as well as the Gibson cloning method (New England Biolabs, Inc.). ΔFG strains are in a W303 background, whereas most individual deletion mutants are in a BY4741/BY4742 background. Gene deletions (SWY5724, 5909, 5912, 6012, 6013, 6014, 6044, 6170, 6236, 6237, and 6239) were constructed by generating a PCR product using primers 50–400 bp upstream and downstream of the gene of interest and genomic DNA from a null allele, typically from the deletion library (Giaever et al., 2002), and then transforming the PCR product into the desired yeast strain and plating on selective media. GFP-tagged strains (SWY5836-40, 5913, 5914, 5915, and 6238) were generated in a similar manner using primers upstream of the C-terminal region and in the 3′ UTR of the gene of interest along with genomic DNA from a GFP-tagged strain, typically from the GFP collection (Huh et al., 2003). Htb2-mCherry–tagged strains (SWY5678, 5680, 5725, 5728, and 5874) were made similarly using genomic DNA from an htb2-mCherry strain. SWY6169 was generated by transforming BY4741 cells with a PCR product to insert a TDH3 promoter upstream of GSP1; the PCR product was made using the Gateway vector pAG306GPD-ccdB (Addgene) as a template. Fluorescence microscopy and/or immunoblotting was used to confirm deletions and tags. Plasmid GFP-4PrA was created by cloning in TPI promoter, GFP, and four Protein A sequences into the BssHI site of pRS415. pSW4070 was generated by cloning GSP1 and ∼500 bp of its 5′ and 3′ UTR in EcoRI-digested pRS426. A. Corbett of Emory University, Atlanta, GA; D. Goldfarb of University of Rochester, Rochester, NY; and J. Aitchison of the Institute for Systems Biology, Seattle, WA provided strains and/or plasmids for some experiments.
Replicative aging assays
RLSs were generally measured as previously described (Park et al., 2002; Steffen et al., 2009). Strains were streaked onto a YPD plate from a frozen glycerol stock and incubated at 25–30°C for 2–3 d. Single colonies were then restreaked as patches near the side of a fresh YPD plate (2–4 patches/plate). After growing overnight, ≥30 single cells were isolated from each patch and moved into rows using a microdissection microscope. Plates were incubated for 2–3 h, and virgin daughter cells were isolated from the 30 individual mother cells; the mother cells were then discarded. These virgin daughter cells then began to divide, becoming mothers. New daughter cells were counted and discarded to measure each mother’s RLS. The RLS was considered complete when a cell hadn’t successfully divided for ≥12 h at 30°C. Plates were stored at 4°C overnight. Assays were performed at 30°C on YPD plates unless otherwise noted. For DR experiments, YP and 0.05% glucose plates were used. Mean RLSs are listed in parentheses next to strain names in figures.
Enrichment of mother cells
Mother cells were enriched as previously described (Park et al., 2002). 1 OD599 unit of yeast grown to mid–log phase was pelleted and washed two times with 500 µl PBS. The cells were resuspended in 500 µl PBS, and 1 µg Sulfa-NHS-LC-biotin was added to the suspension, which was vortexed for 20 s to dissolve the biotin. This was incubated at 25°C while shaking for 20–30 min. The cells were washed two times with PBS and added to 1 liter synthetic media with 2% glucose and 75 µg/ml ampicillin. Cells were grown for 15–16 h at 30°C. Cells were pelleted and washed with 10 ml PBS before resuspending in a 15-ml conical tube with 8 ml PBS and 60 µl of 4-mg/ml streptavidin magnetic beads. The suspension was mixed 30 min at 4°C. The tube was then incubated on a magnetic separation rack at 4°C for 5 min. The supernatant was removed, and 8 ml PBS was added; this process was repeated five times. Some cells in the supernatant were saved as young cells. The aged cells were then resuspended in 1 ml PBS for further analysis.
CCFW staining
Cells were washed twice with 500 µl PBS and then resuspended in 500 µl PBS with 0.5 µg/ml CCFW and mixed 5 min at 4°C. Cells were then washed two times with PBS before fluorescence microscopy analysis. Cells were visualized using a microscope (BX53; Olympus) equipped with a 100×/1.35 NA UPlan oil lens (Olympus) and a camera (Orca-R2; Hamamatsu Photonics) at room temperature to visualize calcofluor dye using a DAPI filter and NIS Elements Advanced Research 3.2 imaging software (Nikon). 13–17 0.65-µm images were acquired for each cell to detect bud scars on the periphery of the cell. ImageJ (National Institutes of Health) was used to analyze images when counting bud scars.
Live-cell NPC permeability assay and confocal microscopy
htb2-mCherry:HIS3 strains transformed with GFP-4PrA cloned in pRS415 were grown to mid–log phase in SC-Leu. Cells were analyzed using an inverted confocal microscope (LSM510 META; Carl Zeiss) equipped with a 63×/1.40 NA Plan Apochromat oil lens (Carl Zeiss) at room temperature and photographed using a charge-coupled device camera (AxioCam; Carl Zeiss). AxioVision version 3.0 SP software (Carl Zeiss) was used to acquire images. GFP signal was detected using a 488-nm excitation signal, whereas Htb2-mCherry was detected using a 543-nm excitation signal. The GFP signal was bleached in a rectangular area away from the nucleus, which was detected using Htb2-mCherry. Subsequent images were taken every 9 s to monitor loss of the GFP signal from the nucleus into the cytoplasm. ImageJ was used to quantify the mean intensity of GFP signal in a nuclear and cytoplasmic region at each time point. NPC permeability was measured by determining the ratio of these nuclear and cytoplasmic GFP intensities over time, producing a graph with an exponential decay pattern. Each plot was analyzed using Prism 5 (GraphPad Software) to generate a best-fit exponential decay line with a calculated half-time. This was repeated to measure the half-times of ≥10 different cells per strain. The calculated half-times for these cells were then averaged to produce a mean strain half-time depicted in the bar graph in Fig. 1 C.
Kinetic import assays
1–2 OD599 units of yeast transformed with NLS-GFP reporters were grown overnight in SC-Leu to mid–log phase and then washed with 500 µl of 10-mM 2-deoxyglucose and 10-mM sodium azide. Cells were resuspended in 500 µl of 10-mM 2-deoxyglucose and 10-mM sodium azide and incubated for 30 min at 25°C while shaking to inhibit active transport of reporters. 5 µl of this suspension was then removed and analyzed using fluorescence microscopy to ensure that GFP reporters were localized pancellularly. Cells were centrifuged, and the supernatant was removed and replaced with 20–40 µl YPD. Cells were immediately transferred to a slide and then visualized at each time point to determine the percentage with an accumulation of NLS-GFP reporter in the nucleus. A microscope (BX53; Olympus) equipped with a 50×/0.9 NA UPlan oil lens (Olympus) and an Orca-R2 camera was used for analysis at room temperature to detect GFP signal with a 488-nm excitation laser using NIS Elements Advanced Research 3.2 acquisition software. ImageJ was subsequently used to analyze the accumulation of GFP reporter in the nucleus. Three independent experiments were performed for each strain and reporter to produce the graphs in Fig. 2 B.
Calculation of nuclear versus cytoplasmic GFP ratios and ratiometric analysis
1–2 OD599 units of yeast transformed with NLS-GFP reporters were grown overnight in SC-Leu to mid–log phase, pelleted, and resuspended in ∼20 µl SC media. Cells were transferred to a slide and visualized using a microscope (BX53) equipped with a 100×/1.35 NA Olympus UPlan oil lens and an Orca-R2 camera at room temperature to visualize GFP signal with a 488-nm excitation laser using NIS Elements AR 3.2 acquisition software. ImageJ was used to quantify the mean GFP intensity in a region of the nucleus and cytoplasm as well as the mean background intensity of the field. Mean background fluorescence was subtracted from the mean nuclear and cytoplasmic fluorescence, and then, the ratio was calculated by dividing the nuclear by the cytoplasmic fluorescence. For ratiometric analysis in Fig. S2 C, ImageJ was used to quantify the intensity of Nup100-GFP and Nsp1-mCherry at the nuclear rim of 20 cells that had divided zero or eight times. The intensity of Nsp1-mCherry was divided by that of Nup100-GFP to get ratios for individual cells.
MitoTracker staining
1–2 OD599 units of yeast grown in synthetic media were pelleted in a light-sensitive 1.5-ml tube and washed once with 500 µl staining buffer (10 mM Hepes, pH 7.4, and 5% glucose). Cells were resuspended in 500 µl staining buffer supplemented with 100 nM MitoTracker red CMXRos (Life Technologies) and shaken for 15–30 min at 25°C. Cells were then washed two times with 500 µl staining buffer and resuspended in 20–30 µl staining buffer for analysis using fluorescence microscopy. For aged cell experiments, cells were stained with CCFW and then washed an additional two times with staining buffer before fluorescence microscopy analysis. Cells were visualized using a microscope (BX53) equipped with a 100×/1.35 NA UPlan oil lens (Olympus) and a camera (Orca-R2) to detect MitoTracker signal using a TRITC excitation laser at 543 nm at room temperature; a DAPI laser was used to detect CCFW. NIS Elements Advanced Research 3.2 was used to acquire images; z stacks were taken as described in the CCFW staining section. For quantification of MitoTracker signals, ImageJ was used to measure the mean cytoplasmic fluorescence of cells and the mean background intensity of the field. Cellular fluorescence was calculated by subtracting the mean background intensity from the mean cytoplasmic fluorescence. The mean intensity was calculated for wild-type zero division cells in Fig. 5 D, and wild-type cells in Fig. 6 C, and then all values in each experiment were divided by this mean to determine relative fluorescence for each condition.
Immunoblotting
Lysates were generated by resuspending 1–3 OD599 units of yeast in 40–50 µl Laemmli sample buffer (SB) and boiling immediately for 5 min. Glass beads were added to fill the volume of the SB, and then, suspensions were vortexed for 2 min. 50–75 µl of SB was then added to samples, which were vortexed and boiled for an additional 30 s before loading. Membranes were stained with 15 ml Ponceau S (0.1% Ponceau S and 5% acetic acid) after each transfer to compare relative protein levels of samples. Membranes were washed 2 min with 20 ml of 50-mM NaOH to remove the Ponceau stain, and then three times with 20 ml PBS. Membranes were incubated in blocking buffer (5% milk, 150 mM NaCl, 50 mM Tris, pH 7.5, and 0.5% Tween 20) for 1 h and then incubated overnight with affinity-purified primary antibody at 4°C in 1–2 ml blocking buffer. Membranes were washed three times for 5–10 min with 10 ml of wash buffer (150 mM NaCl, 50 mM Tris, pH 7.5, and 0.5% Tween 20). Blots were incubated 1 h in 5 ml blocking buffer along with 1:10,000 dilution of Alexa Fluor–conjugated secondary antibody, and then washed three times for 5–10 min with 10 ml of wash buffer. Blots were developed using an imager (Odyssey; LI-COR Biosciences). Primary antibodies were used at the following dilutions: rabbit polyclonal anti-Nup116 C terminus (ASW11.1), 1:1,000; mouse monoclonal anti-Pgk1 (Life Technologies), 1:1,000; rabbit polyclonal anti-GFP (gift from M. Linder, Cornell University, Ithaca, NY), 1:1,000; mouse monoclonal anti-Ran (BD), 1:2,000; and rabbit polyclonal anti-Nup53 (Abcam), 1:2,000. Immunoblots were quantified using ImageJ, and ratios were generated in Fig. 4 C by dividing the protein levels of Nup116/Nup100-GFP/Nsp1-GFP by levels of Pgk1. At least three independent experiments were performed for quantification.
Serial yeast dilutions
3-ml yeast cultures were grown in either YP/SC 2% dextrose medium overnight to an OD599 of 0.1–1.0/ml. 500,000 cells (0.166 OD599 units) were pelleted and resuspended in 250 µl sterile water. 100 µl of this suspension was mixed in a new tube with 400 µl sterile water to dilute the sample fivefold; this was repeated three more times successively to dilute samples further. 5 µl of each dilution was then spotted onto agar plates and then grown for 1–6 d depending on the growth rate of wild-type cells on the media at a given temperature. Plates were scanned, and Photoshop (Adobe) was used to adjust brightness and contrast.
Quantitative RT-PCR
1 OD599 units of yeast were resuspended in 1 ml cold water, pelleted, and then frozen on dry ice in a 1.5-ml tube. The pellets were then resuspended in 400 µl of 10 mM Tris-HCl, pH 7.5, 10 mM EDTA, and 0.5% SDS. 400 µl acid phenol was added, and the tube was incubated for 60 min at 65°C after vortexing 10 s. Tubes were then placed on ice 5 min and then centrifuged 13,000 rpm for 5 min at 4°C. The top aqueous phase was transferred to a new 1.5-ml tube along with 400 µl acid phenol, vortexed, and then centrifuged 13,000 rpm for 5 min at 4°C. The top aqueous phase was again transferred to a new tube, along with 400 µl chloroform, which was vortexed and centrifuged 13,000 rpm for 5 min at 4°C. The aqueous phase was transferred to a fresh 1.5-ml tube along with 1/10 the volume of 3 M sodium acetate, pH 5.3, and 2.5 times the volume of ethanol. The tube was incubated at −70°C for 45 min and then centrifuged 13,000 rpm for 5 min at 4°C. The RNA was washed with 500 µl of ice-cold 70% ethanol and resuspended in 20 µl RNase-free water, and its concentration was determined by measuring the OD260 and OD280. 5 µg RNA was treated with RQ1 DNase according to its manufacturer’s protocol (Promega). cDNA was generated using a reverse transcriptase kit according to the manufacturer’s protocol (Applied Biosystems); a no reverse transcriptase control was included for each sample. The cDNA was added to 2× SYBR green Master Mix (Bio-Rad Laboratories) along with primers to amplify 80–100-nt regions of Nup116, Nup53, Nsp1, Nup100, or Pgk1 according to the manufacturer’s protocol; three replicates were performed for each reaction. Five serial 1:5 dilutions of cDNA were used for each transcript and sample for quantification. A quantitative PCR thermocycler (CFX96; Bio-Rad Laboratories) was used to perform and quantify relative transcript levels in young and aged cells; values for the three replicates/sample were averaged to assess the total level of transcript mRNA for each experiment. The total amount of mRNA was set to 1 for each transcript in young cells to compare with amounts in aged cells. Three independent experiments were performed to produce the graph depicted in Fig. S2.
Polysome profiles
Yeast cells were grown overnight at 30°C in 100 ml YPD to a density of 0.9–1.4 OD599 units/ml. Cycloheximide was added to a concentration of 50 µg/ml, and the cells were incubated on ice for 15 min. Cells were pelleted at 4°C and washed in 5 ml of cold low salt (LS) buffer (20 mM Hepes, pH 7.5, 100 mM KOAc, 5 mM MgOAc, 1 mM EDTA, and 50 µg/ml cycloheximide) before resuspending in 700 µl of LS buffer supplemented with 1 mM PMSF, 1 mM DTT, and protease inhibitor cocktail. Cells were vortexed with an equal amount of glass beads for 5 min at 4°C, and then, the resulting lysate was centrifuged at 500 g for 5 min at 4°C. The supernatant was collected and moved to a fresh tube where the protein concentration was determined using a Bradford assay. 1.5 mg of protein was loaded onto the top of a 10.6 ml of 47–7% continuous sucrose gradient prepared in LS buffer in a 14 × 89–mm ultracentrifuge tube (Beckman Coulter). The gradients were centrifuged at 24,000 g (38,000 rpm) for 2 h at 4°C in a swinging bucket rotor (SW 41 Ti; Beckman Coulter). The absorbance of the gradient at 254 nm was then continuously measured using a wavelength detector to produce graphs depicted in Fig. 3 C.
Online supplemental material
Fig. S1 shows Nup116 and Nup100 levels are unchanged in ΔFG mutants, contains a table with doubling times for several mutant strains used in the study, and also shows deletion of NUP100 increases RLS in BY4741 cells. Fig. S2 shows several experiments comparing young and aged yeast, including those that compare Nup53 protein levels, NUP116, NUP100, NUP53, NSP1, and PGK1 transcript levels, and the relative amounts of Nsp1-mCherry to Nup100-GFP. Survival curves for nup116ΔGLFG fob1Δ, sir2Δ nup100Δ, gsp1-322 nup100Δ, and kap121-7 nup100Δ mutants are shown in Fig. S3. Fig. S4 shows mitochondrial defects are specific to nup116ΔGLFG mutants and also contains additional microscopy data characterizing nup116ΔGLFG cells. Fig. S5 shows the mitochondrial defect of nup116ΔGLFG cells is rescued by GSP1 overexpression, that GSP1 overexpression extends RLS on glycerol, and contains brightness-adjusted images of kap121 mutant cells. Table S1 lists strains used for the study. Table S2 lists plasmids used for this study. Online supplemental material is available at http://www.jcb.org/cgi/content/full/jcb.201412024/DC1.
Supplementary Material
Acknowledgments
The authors thank members of the Wente laboratory for discussions and comments on the manuscript. We also thank Christopher Browne and Andrew Link for technical assistance and equipment for polysome profiles. FRAP experiments were performed in part through the use of the Vanderbilt University Medical Center Cell Imaging Shared Resource (supported by National Institutes of Health grants CA68485, DK20593, DK58404, DK59637, and EY08126). Pamela Silver, Eric Shows, Anita Corbett, David Goldfarb, and John Aitchison kindly provided strains and/or plasmids.
This research was supported by a training position on National Cancer Institute 2T32CA9582-26 (to C.L. Lord), the National Institute on Aging under award number F32AG047737 (to C.L. Lord), and National Institutes of Health grant R37GM051219 (to S.R. Wente).
The authors declare no competing financial interests.
Footnotes
Abbreviations used in this paper:
- ANOVA
- analysis of variance
- CCFW
- calcofluor white
- cNLS
- classical NLS
- DR
- dietary restriction
- ERC
- extrachromosomal ribosomal DNA circle
- HGPS
- Hutchinson–Gilford progeria syndrome
- Kap
- karyopherin
- LS
- low salt
- NPC
- nuclear pore complex
- Nup
- nucleoporin
- RLS
- replicative life span
- SB
- sample buffer
- SC
- synthetic complete
References
- Bailer S.M., Balduf C., Katahira J., Podtelejnikov A., Rollenhagen C., Mann M., Panté N., and Hurt E.. 2000. Nup116p associates with the Nup82p-Nsp1p-Nup159p nucleoporin complex. J. Biol. Chem. 275:23540–23548 10.1074/jbc.M001963200 [DOI] [PubMed] [Google Scholar]
- Belhumeur P., Lee A., Tam R., DiPaolo T., Fortin N., and Clark M.W.. 1993. GSP1 and GSP2, genetic suppressors of the prp20-1 mutant in Saccharomyces cerevisiae: GTP-binding proteins involved in the maintenance of nuclear organization. Mol. Cell. Biol. 13:2152–2161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradatsch B., Katahira J., Kowalinski E., Bange G., Yao W., Sekimoto T., Baumgärtel V., Boese G., Bassler J., Wild K., et al. 2007. Arx1 functions as an unorthodox nuclear export receptor for the 60S preribosomal subunit. Mol. Cell. 27:767–779 10.1016/j.molcel.2007.06.034 [DOI] [PubMed] [Google Scholar]
- Breker M., Gymrek M., and Schuldiner M.. 2013. A novel single-cell screening platform reveals proteome plasticity during yeast stress responses. J. Cell Biol. 200:839–850 10.1083/jcb.201301120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaves S.R., and Blobel G.. 2001. Nuclear import of Spo12p, a protein essential for meiosis. J. Biol. Chem. 276:17712–17717 10.1074/jbc.M010760200 [DOI] [PubMed] [Google Scholar]
- Colombi P., Webster B.M., Fröhlich F., and Lusk C.P.. 2013. The transmission of nuclear pore complexes to daughter cells requires a cytoplasmic pool of Nsp1. J. Cell Biol. 203:215–232 10.1083/jcb.201305115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corral-Debrinski M., Belgareh N., Blugeon C., Claros M.G., Doye V., and Jacq C.. 1999. Overexpression of yeast karyopherin Pse1p/Kap121p stimulates the mitochondrial import of hydrophobic proteins in vivo. Mol. Microbiol. 31:1499–1511 10.1046/j.1365-2958.1999.01295.x [DOI] [PubMed] [Google Scholar]
- D’Angelo M.A., Raices M., Panowski S.H., and Hetzer M.W.. 2009. Age-dependent deterioration of nuclear pore complexes causes a loss of nuclear integrity in postmitotic cells. Cell. 136:284–295 10.1016/j.cell.2008.11.037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Defossez P.A., Prusty R., Kaeberlein M., Lin S.J., Ferrigno P., Silver P.A., Keil R.L., and Guarente L.. 1999. Elimination of replication block protein Fob1 extends the life span of yeast mother cells. Mol. Cell. 3:447–455 10.1016/S1097-2765(00)80472-4 [DOI] [PubMed] [Google Scholar]
- Enenkel C., Blobel G., and Rexach M.. 1995. Identification of a yeast karyopherin heterodimer that targets import substrate to mammalian nuclear pore complexes. J. Biol. Chem. 270:16499–16502 10.1074/jbc.270.28.16499 [DOI] [PubMed] [Google Scholar]
- Giaever G., Chu A.M., Ni L., Connelly C., Riles L., Véronneau S., Dow S., Lucau-Danila A., Anderson K., André B., et al. 2002. Functional profiling of the Saccharomyces cerevisiae genome. Nature. 418:387–391 10.1038/nature00935 [DOI] [PubMed] [Google Scholar]
- Gotta M., Strahl-Bolsinger S., Renauld H., Laroche T., Kennedy B.K., Grunstein M., and Gasser S.M.. 1997. Localization of Sir2p: the nucleolus as a compartment for silent information regulators. EMBO J. 16:3243–3255 10.1093/emboj/16.11.3243 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halfmann R., Wright J.R., Alberti S., Lindquist S., and Rexach M.. 2012. Prion formation by a yeast GLFG nucleoporin. Prion. 6:391–399 10.4161/pri.20199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes A.L., and Gottschling D.E.. 2012. An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast. Nature. 492:261–265 10.1038/nature11654 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huh W.-K., Falvo J.V., Gerke L.C., Carroll A.S., Howson R.W., Weissman J.S., and O’Shea E.K.. 2003. Global analysis of protein localization in budding yeast. Nature. 425:686–691 10.1038/nature02026 [DOI] [PubMed] [Google Scholar]
- Hülsmann B.B., Labokha A.A., and Görlich D.. 2012. The permeability of reconstituted nuclear pores provides direct evidence for the selective phase model. Cell. 150:738–751 10.1016/j.cell.2012.07.019 [DOI] [PubMed] [Google Scholar]
- Hung N.-J., Lo K.-Y., Patel S.S., Helmke K., and Johnson A.W.. 2008. Arx1 is a nuclear export receptor for the 60S ribosomal subunit in yeast. Mol. Biol. Cell. 19:735–744 10.1091/mbc.E07-09-0968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Izaurralde E., Kutay U., von Kobbe C., Mattaj I.W., and Görlich D.. 1997. The asymmetric distribution of the constituents of the Ran system is essential for transport into and out of the nucleus. EMBO J. 16:6535–6547 10.1093/emboj/16.21.6535 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaeberlein M., McVey M., and Guarente L.. 1999. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13:2570–2580 10.1101/gad.13.19.2570 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelley J.B., Datta S., Snow C.J., Chatterjee M., Ni L., Spencer A., Yang C.-S., Cubeñas-Potts C., Matunis M.J., and Paschal B.M.. 2011. The defective nuclear lamina in Hutchinson-gilford progeria syndrome disrupts the nucleocytoplasmic Ran gradient and inhibits nuclear localization of Ubc9. Mol. Cell. Biol. 31:3378–3395 10.1128/MCB.05087-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keminer O., and Peters R.. 1999. Permeability of single nuclear pores. Biophys. J. 77:217–228 10.1016/S0006-3495(99)76883-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitakawa M., Graack H.-R., Grohmann L., Goldschmidt-Reisin S., Herfurth E., Wittmann-Liebold B., Nishimura T., and Isono K.. 1997. Identification and characterization of the genes for mitochondrial ribosomal proteins of Saccharomyces cerevisiae. Eur. J. Biochem. 245:449–456 10.1111/j.1432-1033.1997.t01-2-00449.x [DOI] [PubMed] [Google Scholar]
- Laun P., Pichova A., Madeo F., Fuchs J., Ellinger A., Kohlwein S., Dawes I., Fröhlich K.U., and Breitenbach M.. 2001. Aged mother cells of Saccharomyces cerevisiae show markers of oxidative stress and apoptosis. Mol. Microbiol. 39:1166–1173 10.1111/j.1365-2958.2001.02317.x [DOI] [PubMed] [Google Scholar]
- Laurell E., Beck K., Krupina K., Theerthagiri G., Bodenmiller B., Horvath P., Aebersold R., Antonin W., and Kutay U.. 2011. Phosphorylation of Nup98 by multiple kinases is crucial for NPC disassembly during mitotic entry. Cell. 144:539–550 10.1016/j.cell.2011.01.012 [DOI] [PubMed] [Google Scholar]
- Lo W.-S., and Dranginis A.M.. 1998. The cell surface flocculin Flo11 is required for pseudohyphae formation and invasion by Saccharomyces cerevisiae. Mol. Biol. Cell. 9:161–171 10.1091/mbc.9.1.161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- López-Otín C., Blasco M.A., Partridge L., Serrano M., and Kroemer G.. 2013. The hallmarks of aging. Cell. 153:1194–1217 10.1016/j.cell.2013.05.039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makio T., Lapetina D.L., and Wozniak R.W.. 2013. Inheritance of yeast nuclear pore complexes requires the Nsp1p subcomplex. J. Cell Biol. 203:187–196 10.1083/jcb.201304047 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menendez-Benito V., van Deventer S.J., Jimenez-Garcia V., Roy-Luzarraga M., van Leeuwen F., and Neefjes J.. 2013. Spatiotemporal analysis of organelle and macromolecular complex inheritance. Proc. Natl. Acad. Sci. USA. 110:175–180 10.1073/pnas.1207424110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore M.S., and Blobel G.. 1993. The GTP-binding protein Ran/TC4 is required for protein import into the nucleus. Nature. 365:661–663 10.1038/365661a0 [DOI] [PubMed] [Google Scholar]
- Oki M., Noguchi E., Hayashi N., and Nishimoto T.. 1998. Nuclear protein import, but not mRNA export, is defective in all Saccharomyces cerevisiae mutants that produce temperature-sensitive forms of the Ran GTPase homologue Gsp1p. Mol. Gen. Genet. 257:624–634 10.1007/s004380050690 [DOI] [PubMed] [Google Scholar]
- Park P.U., McVey M., and Guarente L.. 2002. Separation of mother and daughter cells. Methods Enzymol. 351:468–477 10.1016/S0076-6879(02)51865-6 [DOI] [PubMed] [Google Scholar]
- Rexach M., and Blobel G.. 1995. Protein import into nuclei: association and dissociation reactions involving transport substrate, transport factors, and nucleoporins. Cell. 83:683–692 10.1016/0092-8674(95)90181-7 [DOI] [PubMed] [Google Scholar]
- Rivera-Torres J., Acín-Perez R., Cabezas-Sánchez P., Osorio F.G., Gonzalez-Gómez C., Megias D., Cámara C., López-Otín C., Enríquez J.A., Luque-García J.L., and Andrés V.. 2013. Identification of mitochondrial dysfunction in Hutchinson-Gilford progeria syndrome through use of stable isotope labeling with amino acids in cell culture. J. Proteomics. 91:466–477 10.1016/j.jprot.2013.08.008 [DOI] [PubMed] [Google Scholar]
- Rotenberg M.O., Moritz M., and Woolford J.L. Jr. 1988. Depletion of Saccharomyces cerevisiae ribosomal protein L16 causes a decrease in 60S ribosomal subunits and formation of half-mer polyribosomes. Genes Dev. 2:160–172 10.1101/gad.2.2.160 [DOI] [PubMed] [Google Scholar]
- Rout M.P., Blobel G., and Aitchison J.D.. 1997. A distinct nuclear import pathway used by ribosomal proteins. Cell. 89:715–725 10.1016/S0092-8674(00)80254-8 [DOI] [PubMed] [Google Scholar]
- Savas J.N., Toyama B.H., Xu T., Yates J.R. III, and Hetzer M.W.. 2012. Extremely long-lived nuclear pore proteins in the rat brain. Science. 335:942 10.1126/science.1217421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scheckhuber C.Q., Erjavec N., Tinazli A., Hamann A., Nyström T., and Osiewacz H.D.. 2007. Reducing mitochondrial fission results in increased life span and fitness of two fungal ageing models. Nat. Cell Biol. 9:99–105 10.1038/ncb1524 [DOI] [PubMed] [Google Scholar]
- Schleit J., Wasko B.M., and Kaeberlein M.. 2012. Yeast as a model to understand the interaction between genotype and the response to calorie restriction. FEBS Lett. 586:2868–2873 10.1016/j.febslet.2012.07.038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlenstedt G., Smirnova E., Deane R., Solsbacher J., Kutay U., Görlich D., Ponstingl H., and Bischoff F.R.. 1997. Yrb4p, a yeast ran-GTP-binding protein involved in import of ribosomal protein L25 into the nucleus. EMBO J. 16:6237–6249 10.1093/emboj/16.20.6237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seedorf M., and Silver P.A.. 1997. Importin/karyopherin protein family members required for mRNA export from the nucleus. Proc. Natl. Acad. Sci. USA. 94:8590–8595 10.1073/pnas.94.16.8590 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shulga N., Roberts P., Gu Z., Spitz L., Tabb M.M., Nomura M., and Goldfarb D.S.. 1996. In vivo nuclear transport kinetics in Saccharomyces cerevisiae: a role for heat shock protein 70 during targeting and translocation. J. Cell Biol. 135:329–339 10.1083/jcb.135.2.329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shulga N., Mosammaparast N., Wozniak R., and Goldfarb D.S.. 2000. Yeast nucleoporins involved in passive nuclear envelope permeability. J. Cell Biol. 149:1027–1038 10.1083/jcb.149.5.1027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sinclair D.A., and Guarente L.. 1997. Extrachromosomal rDNA circles—a cause of aging in yeast. Cell. 91:1033–1042 10.1016/S0092-8674(00)80493-6 [DOI] [PubMed] [Google Scholar]
- Snow C.J., Dar A., Dutta A., Kehlenbach R.H., and Paschal B.M.. 2013. Defective nuclear import of Tpr in Progeria reflects the Ran sensitivity of large cargo transport. J. Cell Biol. 201:541–557 10.1083/jcb.201212117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steffen K.K., MacKay V.L., Kerr E.O., Tsuchiya M., Hu D., Fox L.A., Dang N., Johnston E.D., Oakes J.A., Tchao B.N., et al. 2008. Yeast life span extension by depletion of 60s ribosomal subunits is mediated by Gcn4. Cell. 133:292–302 10.1016/j.cell.2008.02.037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steffen K.K., Kennedy B.K., and Kaeberlein M.. 2009. Measuring replicative life span in the budding yeast. J. Vis. Exp. (28):e1209 10.3791/1209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinkraus K.A., Kaeberlein M., and Kennedy B.K.. 2008. Replicative aging in yeast: the means to the end. Annu. Rev. Cell Dev. Biol. 24:29–54 10.1146/annurev.cellbio.23.090506.123509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stelter P., Kunze R., Fischer J., and Hurt E.. 2011. Probing the nucleoporin FG repeat network defines structural and functional features of the nuclear pore complex. J. Cell Biol. 195:183–192 10.1083/jcb.201105042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strawn L.A., Shen T., Shulga N., Goldfarb D.S., and Wente S.R.. 2004. Minimal nuclear pore complexes define FG repeat domains essential for transport. Nat. Cell Biol. 6:197–206 10.1038/ncb1097 [DOI] [PubMed] [Google Scholar]
- Terry L.J., and Wente S.R.. 2007. Nuclear mRNA export requires specific FG nucleoporins for translocation through the nuclear pore complex. J. Cell Biol. 178:1121–1132 10.1083/jcb.200704174 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Timney B.L., Tetenbaum-Novatt J., Agate D.S., Williams R., Zhang W., Chait B.T., and Rout M.P.. 2006. Simple kinetic relationships and nonspecific competition govern nuclear import rates in vivo. J. Cell Biol. 175:579–593 10.1083/jcb.200608141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tkach J.M., Yimit A., Lee A.Y., Riffle M., Costanzo M., Jaschob D., Hendry J.A., Ou J., Moffat J., Boone C., et al. 2012. Dissecting DNA damage response pathways by analysing protein localization and abundance changes during DNA replication stress. Nat. Cell Biol. 14:966–976 10.1038/ncb2549 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toyama B.H., Savas J.N., Park S.K., Harris M.S., Ingolia N.T., Yates J.R. III, and Hetzer M.W.. 2013. Identification of long-lived proteins reveals exceptional stability of essential cellular structures. Cell. 154:971–982 10.1016/j.cell.2013.07.037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tran E.J., Zhou Y., Corbett A.H., and Wente S.R.. 2007. The DEAD-box protein Dbp5 controls mRNA export by triggering specific RNA:protein remodeling events. Mol. Cell. 28:850–859 10.1016/j.molcel.2007.09.019 [DOI] [PubMed] [Google Scholar]
- Wente S.R., and Rout M.P.. 2010. The nuclear pore complex and nuclear transport. Cold Spring Harb. Perspect. Biol. 2:a000562 10.1101/cshperspect.a000562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wente S.R., Rout M.P., and Blobel G.. 1992. A new family of yeast nuclear pore complex proteins. J. Cell Biol. 119:705–723 10.1083/jcb.119.4.705 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.