Abstract
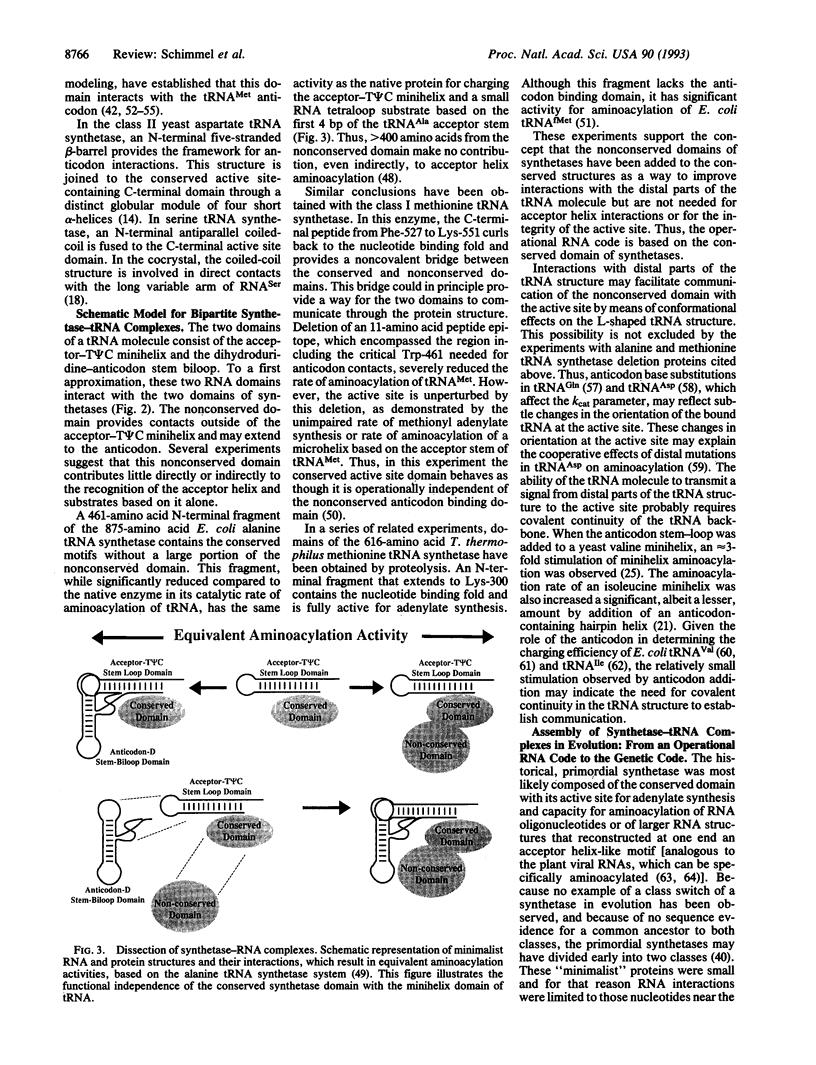
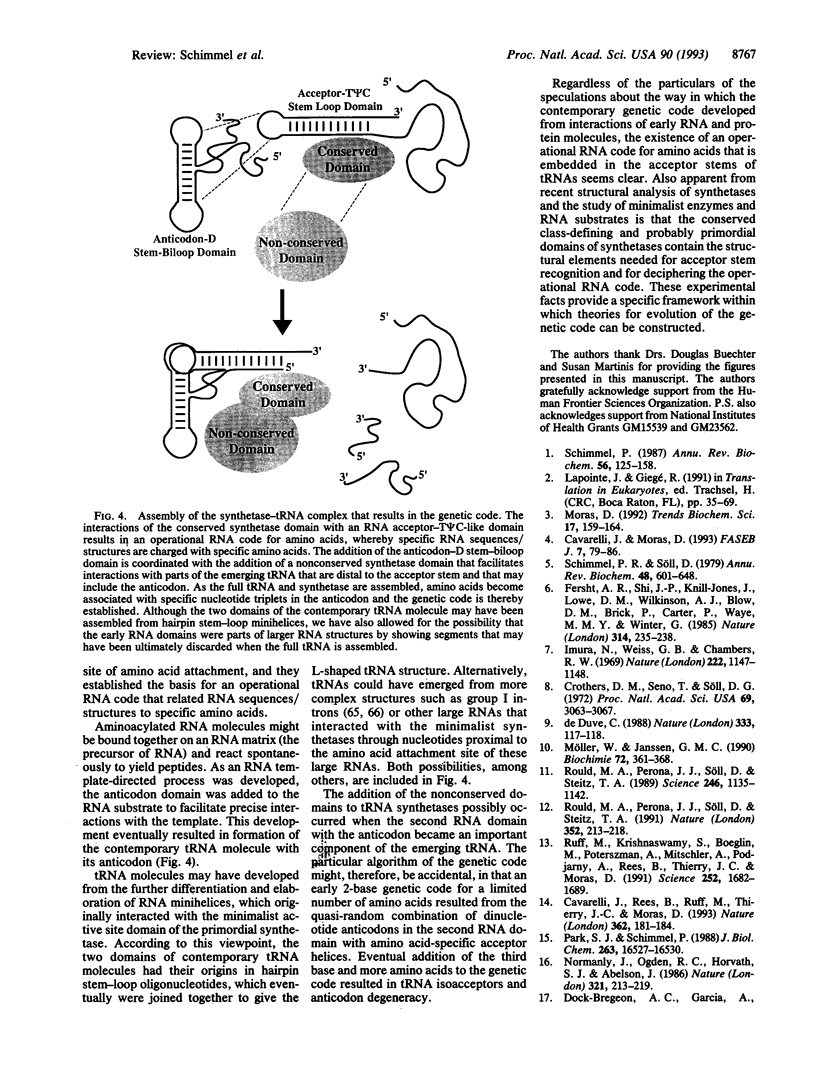
RNA helical oligonucleotides that recapitulate the acceptor stems of transfer RNAs, and that are devoid of the anticodon trinucleotides of the genetic code, are aminoacylated by aminoacyl tRNA synthetases. The specificity of aminoacylation is sequence dependent, and both specificity and efficiency are generally determined by only a few nucleotides proximal to the amino acid attachment site. This sequence/structure-dependent aminoacylation of RNA oligonucleotides constitutes an operational RNA code for amino acids. To a rough approximation, members of the two different classes of tRNA synthetases are, like tRNAs, organized into two major domains. The class-defining conserved domain containing the active site incorporates determinants for recognition of RNA mini-helix substrates. This domain may reflect the primordial synthetase, which was needed for expression of the operational RNA code. The second synthetase domain, which generally is less or not conserved, provides for interactions with the second domain of tRNA, which incorporates the anticodon. The emergence of the genetic from the operational RNA code could occur when the second domain of synthetases was added with the anticodon-containing domain of tRNAs.
Full text
PDF
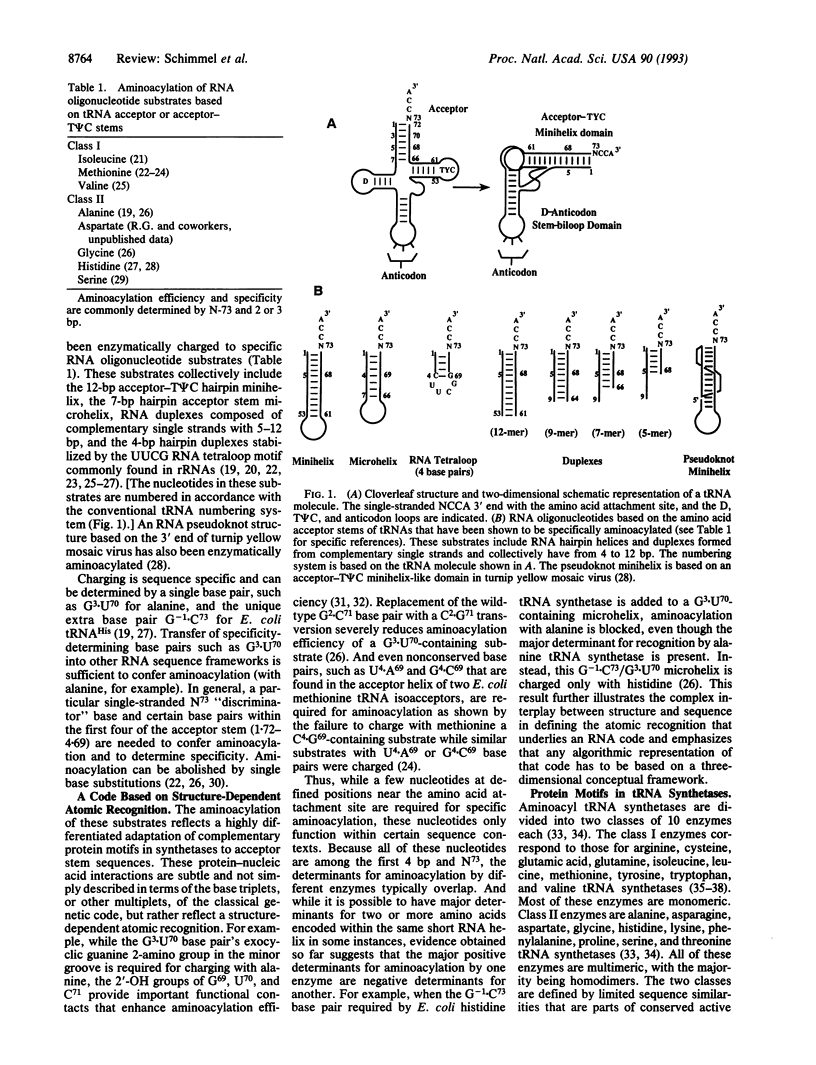




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brick P., Bhat T. N., Blow D. M. Structure of tyrosyl-tRNA synthetase refined at 2.3 A resolution. Interaction of the enzyme with the tyrosyl adenylate intermediate. J Mol Biol. 1989 Jul 5;208(1):83–98. doi: 10.1016/0022-2836(89)90090-9. [DOI] [PubMed] [Google Scholar]
- Brunie S., Zelwer C., Risler J. L. Crystallographic study at 2.5 A resolution of the interaction of methionyl-tRNA synthetase from Escherichia coli with ATP. J Mol Biol. 1990 Nov 20;216(2):411–424. doi: 10.1016/S0022-2836(05)80331-6. [DOI] [PubMed] [Google Scholar]
- Buechter D. D., Schimmel P. Dissection of a class II tRNA synthetase: determinants for minihelix recognition are tightly associated with domain for amino acid activation. Biochemistry. 1993 May 18;32(19):5267–5272. doi: 10.1021/bi00070a039. [DOI] [PubMed] [Google Scholar]
- Burbaum J. J., Schimmel P. Structural relationships and the classification of aminoacyl-tRNA synthetases. J Biol Chem. 1991 Sep 15;266(26):16965–16968. [PubMed] [Google Scholar]
- Cavarelli J., Moras D. Recognition of tRNAs by aminoacyl-tRNA synthetases. FASEB J. 1993 Jan;7(1):79–86. doi: 10.1096/fasebj.7.1.8422978. [DOI] [PubMed] [Google Scholar]
- Cavarelli J., Rees B., Ruff M., Thierry J. C., Moras D. Yeast tRNA(Asp) recognition by its cognate class II aminoacyl-tRNA synthetase. Nature. 1993 Mar 11;362(6416):181–184. doi: 10.1038/362181a0. [DOI] [PubMed] [Google Scholar]
- Crothers D. M., Seno T., Söll G. Is there a discriminator site in transfer RNA? Proc Natl Acad Sci U S A. 1972 Oct;69(10):3063–3067. doi: 10.1073/pnas.69.10.3063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cusack S., Berthet-Colominas C., Härtlein M., Nassar N., Leberman R. A second class of synthetase structure revealed by X-ray analysis of Escherichia coli seryl-tRNA synthetase at 2.5 A. Nature. 1990 Sep 20;347(6290):249–255. doi: 10.1038/347249a0. [DOI] [PubMed] [Google Scholar]
- Cusack S., Härtlein M., Leberman R. Sequence, structural and evolutionary relationships between class 2 aminoacyl-tRNA synthetases. Nucleic Acids Res. 1991 Jul 11;19(13):3489–3498. doi: 10.1093/nar/19.13.3489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dock-Bregeon A. C., Garcia A., Giegé R., Moras D. The contacts of yeast tRNA(Ser) with seryl-tRNA synthetase studied by footprinting experiments. Eur J Biochem. 1990 Mar 10;188(2):283–290. doi: 10.1111/j.1432-1033.1990.tb15401.x. [DOI] [PubMed] [Google Scholar]
- Dreher T. W., Tsai C. H., Florentz C., Giegé R. Specific valylation of turnip yellow mosaic virus RNA by wheat germ valyl-tRNA synthetase determined by three anticodon loop nucleotides. Biochemistry. 1992 Sep 29;31(38):9183–9189. doi: 10.1021/bi00153a010. [DOI] [PubMed] [Google Scholar]
- Eriani G., Delarue M., Poch O., Gangloff J., Moras D. Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature. 1990 Sep 13;347(6289):203–206. doi: 10.1038/347203a0. [DOI] [PubMed] [Google Scholar]
- Eriani G., Dirheimer G., Gangloff J. Cysteinyl-tRNA synthetase: determination of the last E. coli aminoacyl-tRNA synthetase primary structure. Nucleic Acids Res. 1991 Jan 25;19(2):265–269. doi: 10.1093/nar/19.2.265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fersht A. R. Dissection of the structure and activity of the tyrosyl-tRNA synthetase by site-directed mutagenesis. Biochemistry. 1987 Dec 15;26(25):8031–8037. doi: 10.1021/bi00399a001. [DOI] [PubMed] [Google Scholar]
- Fersht A. R., Shi J. P., Knill-Jones J., Lowe D. M., Wilkinson A. J., Blow D. M., Brick P., Carter P., Waye M. M., Winter G. Hydrogen bonding and biological specificity analysed by protein engineering. Nature. 1985 Mar 21;314(6008):235–238. doi: 10.1038/314235a0. [DOI] [PubMed] [Google Scholar]
- Francklyn C., Schimmel P. Aminoacylation of RNA minihelices with alanine. Nature. 1989 Feb 2;337(6206):478–481. doi: 10.1038/337478a0. [DOI] [PubMed] [Google Scholar]
- Francklyn C., Schimmel P. Enzymatic aminoacylation of an eight-base-pair microhelix with histidine. Proc Natl Acad Sci U S A. 1990 Nov;87(21):8655–8659. doi: 10.1073/pnas.87.21.8655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francklyn C., Shi J. P., Schimmel P. Overlapping nucleotide determinants for specific aminoacylation of RNA microhelices. Science. 1992 Feb 28;255(5048):1121–1125. doi: 10.1126/science.1546312. [DOI] [PubMed] [Google Scholar]
- Frugier M., Florentz C., Giegé R. Anticodon-independent aminoacylation of an RNA minihelix with valine. Proc Natl Acad Sci U S A. 1992 May 1;89(9):3990–3994. doi: 10.1073/pnas.89.9.3990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghosh G., Pelka H., Schulman L. H. Identification of the tRNA anticodon recognition site of Escherichia coli methionyl-tRNA synthetase. Biochemistry. 1990 Mar 6;29(9):2220–2225. doi: 10.1021/bi00461a003. [DOI] [PubMed] [Google Scholar]
- Giegé R., Puglisi J. D., Florentz C. tRNA structure and aminoacylation efficiency. Prog Nucleic Acid Res Mol Biol. 1993;45:129–206. doi: 10.1016/s0079-6603(08)60869-7. [DOI] [PubMed] [Google Scholar]
- Hou Y. M., Shiba K., Mottes C., Schimmel P. Sequence determination and modeling of structural motifs for the smallest monomeric aminoacyl-tRNA synthetase. Proc Natl Acad Sci U S A. 1991 Feb 1;88(3):976–980. doi: 10.1073/pnas.88.3.976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hountondji C., Dessen P., Blanquet S. Sequence similarities among the family of aminoacyl-tRNA synthetases. Biochimie. 1986 Sep;68(9):1071–1078. doi: 10.1016/s0300-9084(86)80181-x. [DOI] [PubMed] [Google Scholar]
- Imura N., Weiss G. B., Chambers R. W. Reconstitution of alanine acceptor activity from fragments of yeast tRNA-Ala II. Nature. 1969 Jun 21;222(5199):1147–1148. doi: 10.1038/2221147a0. [DOI] [PubMed] [Google Scholar]
- Jahn M., Rogers M. J., Söll D. Anticodon and acceptor stem nucleotides in tRNA(Gln) are major recognition elements for E. coli glutaminyl-tRNA synthetase. Nature. 1991 Jul 18;352(6332):258–260. doi: 10.1038/352258a0. [DOI] [PubMed] [Google Scholar]
- Kim S., Schimmel P. Function independence of microhelix aminoacylation from anticodon binding in a class I tRNA synthetase. J Biol Chem. 1992 Aug 5;267(22):15563–15567. [PubMed] [Google Scholar]
- Kohda D., Yokoyama S., Miyazawa T. Functions of isolated domains of methionyl-tRNA synthetase from an extreme thermophile, Thermus thermophilus HB8. J Biol Chem. 1987 Jan 15;262(2):558–563. [PubMed] [Google Scholar]
- Ludmerer S. W., Schimmel P. Gene for yeast glutamine tRNA synthetase encodes a large amino-terminal extension and provides a strong confirmation of the signature sequence for a group of the aminoacyl-tRNA synthetases. J Biol Chem. 1987 Aug 5;262(22):10801–10806. [PubMed] [Google Scholar]
- Martinis S. A., Schimmel P. Enzymatic aminoacylation of sequence-specific RNA minihelices and hybrid duplexes with methionine. Proc Natl Acad Sci U S A. 1992 Jan 1;89(1):65–69. doi: 10.1073/pnas.89.1.65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinis S. A., Schimmel P. Microhelix aminoacylation by a class I tRNA synthetase. Non-conserved base pairs required for specificity. J Biol Chem. 1993 Mar 25;268(9):6069–6072. [PubMed] [Google Scholar]
- Meinnel T., Mechulam Y., Le Corre D., Panvert M., Blanquet S., Fayat G. Selection of suppressor methionyl-tRNA synthetases: mapping the tRNA anticodon binding site. Proc Natl Acad Sci U S A. 1991 Jan 1;88(1):291–295. doi: 10.1073/pnas.88.1.291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Westhof E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J Mol Biol. 1990 Dec 5;216(3):585–610. doi: 10.1016/0022-2836(90)90386-Z. [DOI] [PubMed] [Google Scholar]
- Moras D. Structural and functional relationships between aminoacyl-tRNA synthetases. Trends Biochem Sci. 1992 Apr;17(4):159–164. doi: 10.1016/0968-0004(92)90326-5. [DOI] [PubMed] [Google Scholar]
- Muramatsu T., Nishikawa K., Nemoto F., Kuchino Y., Nishimura S., Miyazawa T., Yokoyama S. Codon and amino-acid specificities of a transfer RNA are both converted by a single post-transcriptional modification. Nature. 1988 Nov 10;336(6195):179–181. doi: 10.1038/336179a0. [DOI] [PubMed] [Google Scholar]
- Musier-Forsyth K., Schimmel P. Aminoacylation of RNA oligonucleotides: minimalist structures and origin of specificity. FASEB J. 1993 Feb 1;7(2):282–289. doi: 10.1096/fasebj.7.2.7680012. [DOI] [PubMed] [Google Scholar]
- Musier-Forsyth K., Schimmel P. Functional contacts of a transfer RNA synthetase with 2'-hydroxyl groups in the RNA minor groove. Nature. 1992 Jun 11;357(6378):513–515. doi: 10.1038/357513a0. [DOI] [PubMed] [Google Scholar]
- Musier-Forsyth K., Usman N., Scaringe S., Doudna J., Green R., Schimmel P. Specificity for aminoacylation of an RNA helix: an unpaired, exocyclic amino group in the minor groove. Science. 1991 Aug 16;253(5021):784–786. doi: 10.1126/science.1876835. [DOI] [PubMed] [Google Scholar]
- Möller W., Janssen G. M. Transfer RNAs for primordial amino acids contain remnants of a primitive code at position 3 to 5. Biochimie. 1990 May;72(5):361–368. doi: 10.1016/0300-9084(90)90033-d. [DOI] [PubMed] [Google Scholar]
- Nagel G. M., Doolittle R. F. Evolution and relatedness in two aminoacyl-tRNA synthetase families. Proc Natl Acad Sci U S A. 1991 Sep 15;88(18):8121–8125. doi: 10.1073/pnas.88.18.8121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Normanly J., Ogden R. C., Horvath S. J., Abelson J. Changing the identity of a transfer RNA. Nature. 1986 May 15;321(6067):213–219. doi: 10.1038/321213a0. [DOI] [PubMed] [Google Scholar]
- Pallanck L., Schulman L. H. Anticodon-dependent aminoacylation of a noncognate tRNA with isoleucine, valine, and phenylalanine in vivo. Proc Natl Acad Sci U S A. 1991 May 1;88(9):3872–3876. doi: 10.1073/pnas.88.9.3872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park S. J., Schimmel P. Evidence for interaction of an aminoacyl transfer RNA synthetase with a region important for the identity of its cognate transfer RNA. J Biol Chem. 1988 Nov 15;263(32):16527–16530. [PubMed] [Google Scholar]
- Perona J. J., Rould M. A., Steitz T. A., Risler J. L., Zelwer C., Brunie S. Structural similarities in glutaminyl- and methionyl-tRNA synthetases suggest a common overall orientation of tRNA binding. Proc Natl Acad Sci U S A. 1991 Apr 1;88(7):2903–2907. doi: 10.1073/pnas.88.7.2903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pleij C. W., Rietveld K., Bosch L. A new principle of RNA folding based on pseudoknotting. Nucleic Acids Res. 1985 Mar 11;13(5):1717–1731. doi: 10.1093/nar/13.5.1717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pütz J., Puglisi J. D., Florentz C., Giegé R. Additive, cooperative and anti-cooperative effects between identity nucleotides of a tRNA. EMBO J. 1993 Jul;12(7):2949–2957. doi: 10.1002/j.1460-2075.1993.tb05957.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Steitz T. A. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature. 1991 Jul 18;352(6332):213–218. doi: 10.1038/352213a0. [DOI] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Söll D., Steitz T. A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 A resolution. Science. 1989 Dec 1;246(4934):1135–1142. doi: 10.1126/science.2479982. [DOI] [PubMed] [Google Scholar]
- Rudinger J., Florentz C., Dreher T., Giegé R. Efficient mischarging of a viral tRNA-like structure and aminoacylation of a minihelix containing a pseudoknot: histidinylation of turnip yellow mosaic virus RNA. Nucleic Acids Res. 1992 Apr 25;20(8):1865–1870. doi: 10.1093/nar/20.8.1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruff M., Krishnaswamy S., Boeglin M., Poterszman A., Mitschler A., Podjarny A., Rees B., Thierry J. C., Moras D. Class II aminoacyl transfer RNA synthetases: crystal structure of yeast aspartyl-tRNA synthetase complexed with tRNA(Asp). Science. 1991 Jun 21;252(5013):1682–1689. doi: 10.1126/science.2047877. [DOI] [PubMed] [Google Scholar]
- Schimmel P. R., Söll D. Aminoacyl-tRNA synthetases: general features and recognition of transfer RNAs. Annu Rev Biochem. 1979;48:601–648. doi: 10.1146/annurev.bi.48.070179.003125. [DOI] [PubMed] [Google Scholar]
- Schimmel P. Aminoacyl tRNA synthetases: general scheme of structure-function relationships in the polypeptides and recognition of transfer RNAs. Annu Rev Biochem. 1987;56:125–158. doi: 10.1146/annurev.bi.56.070187.001013. [DOI] [PubMed] [Google Scholar]
- Schulman L. H., Pelka H. Anticodon switching changes the identity of methionine and valine transfer RNAs. Science. 1988 Nov 4;242(4879):765–768. doi: 10.1126/science.3055296. [DOI] [PubMed] [Google Scholar]
- Shi J. P., Francklyn C., Hill K., Schimmel P. A nucleotide that enhances the charging of RNA minihelix sequence variants with alanine. Biochemistry. 1990 Apr 17;29(15):3621–3626. doi: 10.1021/bi00467a005. [DOI] [PubMed] [Google Scholar]
- Shi J. P., Martinis S. A., Schimmel P. RNA tetraloops as minimalist substrates for aminoacylation. Biochemistry. 1992 Jun 2;31(21):4931–4936. doi: 10.1021/bi00136a002. [DOI] [PubMed] [Google Scholar]
- Shiba K., Schimmel P. Functional assembly of a randomly cleaved protein. Proc Natl Acad Sci U S A. 1992 Mar 1;89(5):1880–1884. doi: 10.1073/pnas.89.5.1880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sixma T. K., Pronk S. E., Kalk K. H., Wartna E. S., van Zanten B. A., Witholt B., Hol W. G. Crystal structure of a cholera toxin-related heat-labile enterotoxin from E. coli. Nature. 1991 May 30;351(6325):371–377. doi: 10.1038/351371a0. [DOI] [PubMed] [Google Scholar]
- Starzyk R. M., Webster T. A., Schimmel P. Evidence for dispensable sequences inserted into a nucleotide fold. Science. 1987 Sep 25;237(4822):1614–1618. doi: 10.1126/science.3306924. [DOI] [PubMed] [Google Scholar]
- Thiebe R., Harbers K., Zachau H. G. Aminoacylation of fragment combinations from yeast tRNA phe . Eur J Biochem. 1972 Mar 15;26(1):144–152. doi: 10.1111/j.1432-1033.1972.tb01750.x. [DOI] [PubMed] [Google Scholar]
- Valenzuela D., Schulman L. H. Identification of peptide sequences at the tRNA binding site of Escherichia coli methionyl-tRNA synthetase. Biochemistry. 1986 Aug 12;25(16):4555–4561. doi: 10.1021/bi00364a015. [DOI] [PubMed] [Google Scholar]
- Webster T., Tsai H., Kula M., Mackie G. A., Schimmel P. Specific sequence homology and three-dimensional structure of an aminoacyl transfer RNA synthetase. Science. 1984 Dec 14;226(4680):1315–1317. doi: 10.1126/science.6390679. [DOI] [PubMed] [Google Scholar]
- de Duve C. Transfer RNAs: the second genetic code. Nature. 1988 May 12;333(6169):117–118. doi: 10.1038/333117a0. [DOI] [PubMed] [Google Scholar]





