Abstract
Resistance of developed bacteria to antibiotic treatment is a very important issue, because introduction of any new antibiotic is after a little while followed by the formation of resistant bacterial isolates in the clinic. The significant increase in clinical resistance to antibiotics is a troubling situation especially in nosocomial infections, where already defenseless patients can be unsuccessful to respond to treatment, causing even greater health issue. Nosocomial infections can be identified as those happening within 2 days of hospital acceptance, 3 days of discharge or 1 month of an operation. They influence 1 out of 10 patients admitted to hospital. Annually, this outcomes in 5000 deaths only in UK with a cost to the National Health Service of a billion pounds. Despite these problems, antibiotic therapy is still the most common method used to treat bacterial infections. On the other hand, it is often mentioned that immune system plays a major role in the progress of infections. In this context, we proposed a mathematical model defining population dynamics of both the specific immune cells produced according to the properties of bacteria by host and the bacteria exposed to multiple antibiotics synchronically, presuming that resistance is gained through mutations due to exposure to antibiotic. Qualitative analysis found out infection-free equilibrium point and other equilibrium points where resistant bacteria and immune system cells exist, only resistant bacteria exists and sensitive bacteria, resistant bacteria and immune system cells exist. As a result of this analysis, our model highlights the fact that when an individual’s immune system weakens, he/she suffers more from the bacterial infections which are believed to have been confined or terminated. Also, these results was supported by numerical simulations.
Keywords: Ordinary differential equations systems, Equilibrium points, Immune system, Bacterial resistance, Antibiotics
Background
Infections have been the leading cause of most diseases in the history of mankind (Mondragón et al. 2014). Especially bacterial infections are more prevailing among these. The most common procedure known to fight bacterial infection is through antibiotic therapy applied to individuals. The expression of resistance to antimicrobial agents in this therapy is both the logical and indispensable outcome of using these agents to treat human infections (Mahmoud and Rice 1999; Ternent et al. 2014).
Resistance developed by the bacteria against antibiotics is described as the talent of bacteria to resist the effects of antibiotics generated either to eradicate or control them (Arya 2007). The release of each new class of antibiotics for treatment, shortly after, has been followed by the emergence of new strains of bacteria which are resistant to this class (Butler and Buss 2006; Clatworthy et al. 2007; Lewis 2013). In this respect, developing new treatment strategies for bacterial infections is very important (Mondragón et al. 2014).
According to its properties, antibiotics has the bacteriostatic action to stop the growth of bacteria and bactericidal action to wipe out the bacteria. However, the distinction between these properties is not explicit, as it depends on the drug concentration used and the type and the growth stage of bacteria (Zhang 2009). Hence, multiple antibiotic therapy is a more appropriate form of treatment.
In fact, the bacterial infection is a complicated process for both the infectious bacteria and the host (Carvalho et al. 2012). It is suggested that a significant role in the progress of infections is basically played by the immune system (Linares and Martinez 2005). The immune system is expressed as a system of biological structures and processes in an organism protecting the body from likely harmful substances via recognizing and responding to antigens (Alberts et al. 2002). In this context, the reactions of different hosts against the same infection can be different due to immune system’s response given by host.
In light of the above, dynamics of relations among antibiotics, immune system’s cells and bacteria are important to understand the nature of the infection.
Mathematical models are one of the significant tools used in both analyzing the spread of infectious diseases in a population of individuals (Hethcote 2000; Singer 1984), and predicting the timing and expansion of infection and possible reinfection processes in an individual (Mohtashemi and Levins 2001). While the former is usually used for planning, prevention and control scenarios, the latter can be active in therapy/intervention programs for treating the individuals exposed to the particular pathogen. In this respect, understanding and anticipating the time of occurrence and magnitude of the maximum load of the bacteria and immune system cells by mathematical modelling can be crucial in selecting effective intervention strategies (Whitman and Ashrafiuon 2006).
Consequently, results on reproduction of sensitive and resistant bacteria to antibiotics are obtained in Austin et al. (1997), Bonten et al (2001), Esteva et al. (2011), Wiesch et al. (2011), Zhang (2009); definitions of factors responsible for resistance prevalence are studied in Austin and Anderson (1999), Linares and Martinez (2005), Opatowski et al. (2011), Rodrigues et al. (2007); bacteria behavior under different antibiotic treatments is examined in Alavez et al. (2006), Bonhoeffer et al. (1997), Bootsma et al. (2012), D’Agata et al. (2007), Sun et al. (2010); optimization results and design of control measures are investigated in Bonten et al. (2001), Haber et al. (2010), Massad et al. (2008), Sotto and Lavigne (2012); biological cost and persistence of antibiotic resistance are analyzed in Andersson et al. (2001), Andersson and Levin (1999), Antia et al. (1996), Johnson and Levin (2013), Mondragón et al. (2014); dynamics between pathogens and immune response are given in André and Gandon (2006), Carvalho et al. (2012), D’Onofrio (2005), Gilchrist and Coombs (2006), Gilchrist and Sasaki (2002), Kostova (2007), Mohtashemi and Levins (2001), Nowak and May (2000), Whitman and Ashrafiuon (2006), respectively.
In this study, we have constructed a continuous time model considering the immune response of the host and the basic mechanisms of bacterial resistance to antibiotics. Our aim is to find specific parameters determining the change in the concentrations of the immune system’s cells produced in host to fight these and the sensitive sub-populations and resistant sub-populations that has either arisen through random mutation and clonal enlargement or through cross-contamination in a special infection and under a appropriate treatment regimen.
Model formulation
When the emergence of resistant bacteria to antibiotic has modelled, there are the two basic aspects generally considered. These are within-host models and within-hospital compartmental models. Within-hospital compartmental models are generally SIR models in epidemiology, considering especially how infections will expand throughout the hospital (Ternent et al. 2014). These models are useful to develop strategies to prevent the spread of resistant individuals to antibiotic in hospital.
Current mathematical models focusing on the fact that it is within the host that resistance develops, aim to study how antibiotic treatment strategies can affect, and in addition, how the occurrence of antibiotic resistance can be prevented (Alberts et al. 2002; Hethcote 2000). In such models, the effect of immune cells generated by the host under the pressure of the bacteria are frequently either neglected or assumed to be at a constant rate. We built up the mathematical model including cell-mediated immune response. In addition, treatment regimens involving multiple antibiotic have been used in most bacterial infections due to bacterial resistant. In this sense, besides the interaction of bacterial-immune response, to investigate the effects of multiple drug therapy is biological more meaningful.
We analyzed the concentrations of the immune cells and bacteria in an individual receiving a cocktail of multi-drug treatment against bacteria via mathematical modelling. Let us denote by S(t) and R(t) the population sizes of sensitive and resistant bacteria against multiple antibiotics at time t, respectively, by B(t) population sizes of immune cells at time t, and by Ai(t) the concentration of the i-th antibiotic, for i = 1, 2, …, n at time t. The parameters used in the model are as follows.
We assume that bacteria follow a logistic growth with carrying capacity T. Let βS and (1 − c)βS the birth rate of sensitive and resistant bacteria, respectively. Specific mutations arising from resistance to chemical control frequently have an inherent fitness cost which may be resulted through reduced reproductive capacity or competitive ability (Alavez et al. 2006). We evaluate fitness cost as a reduction on the reproduction rate of the resistant strain, therefore 0 < c < 1. Likewise, we have supposed that using a logistic style term, immune cells are recruited to the region of infection at a rate k and carrying capacity of these is as ω fold of amount of present bacteria. This is biologically very significant in terms of proliferation of specific immune cells. Also, these interacts can be expressed as a generalised model of a local bacterial infection, such as a urinary tract, tuberculosis or wound infection.
During the administration of the i th antibiotic, a number of resistant bacteria to it can be showed up due to mutations of exposed sensitive bacteria to such antibiotic, we model this situation by the term where is the mutation rate of sensitive bacteria due to exposure to i th antibiotic. Sensitive and resistant bacteria have per capita death rates by response of immune cells and this rates is λ. Sensitive bacteria also die due to the action of the antibiotics, and we have assumed that this situation in model is by the term , where is the death rate of sensitive bacteria due to exposure to i-th antibiotic. Lastly, the i-th antibiotic concentration is supplied at a constant rate δi, and is taken up at a constant per capita rate μi. Under the assumptions a fore mentioned, we obtain the following system of (n + 3) ODE’s:
| 1 |
where
| 2 |
To reduce the number of parameters, the variables are changed as follows;
With the new variables, the normalized system is given as;
| 3 |
where , and . The biologically studied region is given by the set
| 4 |
The following proposition ensures that system (3) is well posed in the sense that the solutions with positive initial conditions started in Ω remain in this region for all t ≥ 0, and so, this solutions of system (3) have biological meaning.
Proposition 1
The regionΩ defined in (4) is positively invariant with respect to the system (3).
Proof
By adding the first two equations of the system (3),
| 5 |
is obtained. Considering the region Ω, we have the following inequality;
| 6 |
By the solution according to (s + r) of inequality (6), it follows that 0 ≤ s + r ≤ 1 for all t ≥ 0. Furthermore, the solutions of the last n equations of system (3) are
| 7 |
where initial conditions satisfy 0 ≤ ai(0) ≤ 1 for i = 1, 2, …, n. Lastly, let . Then the solution of third equation in system (3) is
| 8 |
where initial conditions satisfy 0 < b(0) ≤ s(0) + r(0). From (8), it is obtained that 0 ≤ b ≤ s + r ≤ 1.
Hence, the vector field of system (3) restricted to the boundary of Ω does not include a point at the exterior of it. In this context, solutions starting in remain in the region Ω for all t ≥ 0.
Qualitative analysis of system (3)
We have examined the existence and stability of equilibria of system (3).
Proposition 2
Let
| 9 |
We accept that the general expressions of the system’s equilibria show as for andj = 0, 1, 2, 3. Then, system (3) always hasE0 = (0, 0, 0, 1, 1, …, 1) (namely, the infection-free equilibrium point), E1 = (0, 1, 0, 1, 1, …, 1), E2 = (0, B, B, 1, 1, …, 1) contained inΩ. WhenA > B, in addition toE0, E1 andE2, there exists a fourth the equilibrium point,, inΩ.
Proof
In (9), it is clear that
| 10 |
The equilibria of system (3) are given by the solutions of the system of following algebraic equations;
| 11 |
From the last n equation of system (11), we have ai = 1 for i = 1, 2, …, n. Consequently, the system (11) turns into following system;
| 12 |
From (12), it is obtained that either or .
- When , we have that the equilibrium points are , and
Although the equilibrium points E0 and E1 always exist in Ω, the equilibrium point EI where signs of and are opposite, is biologically meaningless. Therefore, EI is not in Ω. -
In case of , it is obtained the equilibrium points as following;
and .
Equilibrium point E2 always exists in Ω. In addition that, the equilibrium point E3 is in Ω, when .
Taking (9) into account, we have that equilibria of system (3) in Ω are
| 13 |
Theorem 3
(Routh–Hurwitz Criteria) Given the polynomial,
where the coefficients for are real constants, define then Hurwitz matrices using the coefficients ai of the characteristic polynomial:
whereaj = 0 ifj > n. All of the roots of polynomialP(λ) are negative or have negative real parts, if and only if the determinants of all Hurwitz matrices are positive: . For polynomial of degree and 5, the Routh–Hurwitz criteria are summarized.
This criteria has given necessary and sufficient conditions for all of the roots of the characteristic polynomial (with real coefficients) to lie in the left half of the complex plane (Allen 2007).
Theorem 4
Suppose is a nonlinear first-order autonomous system with an equilibrium. Denote the Jacobian matrix ofF evaluated at as. If the characteristic equation of the Jacobian matrix,
satisfies the conditions of the Routh–Hurwitz criteria in Theorem 3, that is, the determinants of all of the Hurwitz matrices are positive, , then the equilibrium is locally asimptotically stable. If, for some, then the equilibrium is unstable (Allen2007).
The following proposition is shown conditions that equilibrium points in the Proposition 2 are locally asimptotically stability (LAS).
Proposition 4
The equilibrium points of system (3) inΩ satisfy
-
(i)
E0 andE1 are unstable points.
-
(ii)
IfA < B, thenE2 is LAS.
-
(iii)
LetB < A, thenE3 is LAS.
Proof
For the stability analysis, the functions of the right side of system (3) are determined as follows;
| 14 |
That jacobian matrix obtained from the equations in (14) is
| 15 |
For ease of examination, the τ-th eigenvalue of equilibrium point Ek has shown as λk,τ for k = 0, 1, 2, 3 and .
-
(i)
Some of the eigenvalues evaluated at the equilibrium point E0 in Ω are and λ0,2 = βR. The eigenvalue λ0,2 is positive, due to (2). According to Theorem 4, the infection-free equilibrium point E0 is unstable point for system (3).
In the same way, the eigenvalues for E1 in Ω are that , λ1,2 = −βR, λ1,3 = k and λ1,i+3 = −μi for i = 1, 2, …, n. The eigenvalue λ1,3 is positive, due to (2). From Theorem 4, E1 is unstable point for system (3).
-
(ii)The jacobian matrix in (15) evaluated at the equilibrium point E2 in Ω is given below
16 The eigenvalues of matrix (16) are for i = 1, 2, …, n and the remain three eigenvalues are found from following matrix;
where matrix is the block matrix of (16). From (2), it is clear that Reλ2,i+3 < 0. Characteristic equation of (17) is17
By the solution of (18), these eigenvalues are determined. Hence, it is obtained that18
and λ2,2 and λ2,3 are gained by solving following equation,19 20 It can be observed that , due to (2) and (10). From Theorem 3 (n = 2), . According to Theorem 4, the LAS conditions for E2 are provided for the eigenvalues, and , exceptionally λ2,1in the (19). It is sufficient to examine the sign of λ2,1. By (9), (19) it can be rewrited as λ2,1 = (βS + η)(A − B). In this respect, λ2,1 is negative, when A < B.
As a result, the equilibrium point E2 is LAS for system (3), when A < B.
-
(iii)
Let B < A. Then the equilibrium point E3 is revealed in Ω. The Jacobian matrix for E3 is
where21 22 The eigenvalues obtained from (21) are λ3,i+3 = −μi for i = 1, 2, …, n and the remaining three eigenvalues, λ3,1, λ3,2 and λ3,3, are found from following matrix;
where the matrix is the block matrix of (21). By (2), it is Reλ3,i+3 < 0. Characteristic equation of matrix (23) is obtained as follows:23
where24 25 We have a1, a3 > 0 due to (2) and (22). By (25), we have that
and so,
Therefore, it is obtained that a1a2 − a3 > 0 due to (2) and (22). From Theorem 3(n = 3), Reλ3,1, Reλ3,2 and Reλ3,3 < 0. According to Theorem 4, The equilibrium point E3 in Ω is LAS for the system (3), when it exists biological, that is, B < A. Hence, proof is completed.
The LAS conditions found for equilibrium points in (13) are summarized in the Table 1.
Table 1.
Existence and stability conditions of the equilibria of system (3)
| Equilibrium points | Biological existence conditions | LAS conditions |
|---|---|---|
| Always exists | Unstable | |
| Always exists | Unstable | |
| Always exists | ||
| When it exists biological |
Where the values , and are as indicated in (9)
Proposition 6
Let us denote by Γ2 the LAS region of the equilibrium pointE2inΩ. In the same way,Γ3 is forE3. ThenΓ2 ∩ Γ3 = ∅.
Proof
It can be clearly observed in Table 1.
The parameter values and it’s references used for numerical study are given in Table 2.
Table 2.
Interpretation and considered values of the parameters
| Parameter | Description | Value | References |
|---|---|---|---|
| Growth rate of sensitive bacteria | 0.8 day−1 | Mondragón et al. (2014) | |
| Growth rate of resistant bacteria | 0.4–0.1 day−1 | Mondragón et al. (2014)-hypothesis | |
| Growth rate of immune cells | 0.6 day−1 | Pugliese and Gandolfi (2008) | |
| Rate of bacteria destroyed by immune cells | 0.3 day−1 | Pugliese and Gandolfi (2008) | |
| Rate to the amount of present bacteria of carrying capacity of immune cells | 1 | Hypothesis | |
| Carrying capacity of bacteria | 109 bacteria | Alavez et al. (2006) | |
| Mutation rate of INH | 10−6 mut×gen | Coll (2009) | |
| Mutation rate of PZA | 0 | Mondragón et al. (2014) | |
| Elimination rate of sensitive bacteria due INH | 0.0039 day−1 | Zhang (2009) | |
| Elimination rate of sensitive bacteria due PZA | 0.0001625 day−1 | Alavez et al. (2006) | |
| Daily dose of INH | 5 mg/kg/day | Coll (2009) | |
| Daily dose of ZPA | 35–20 mg/kg/day | Coll (2009) | |
| Uptake rate of INH | 0.06 day−1 | Esteva et al. (2011) | |
| Uptake rate of PZA | 0.03 day−1 | Esteva et al. (2011) |
Datas are deduced from the literature
The values obtained from this table are that (i) in the first case, , and so, is LAS. (ii) in the second case, , and so, is LAS
Among the treatment regimen recommended by WHO includes isoniazid (INH) and pyrazinamide (PZA) for some bacterial infectious (such as Mycobacterium tuberculosis) (Coll 2009)
Proposition 7
IfA < B, then the equilibrium pointE2 = (0, B, B, 1, …, 1) is globally asimptotically stable (GAS) inΩ.
Proof
Their solutions approach ai = 1 for i = 1, 2, …, n, when last n equations of system (3) are considered separately. Replacing s = 0 and ai = 1 for i = 1, 2, …, n in system (3), we attain the asymptotically equivalent planar system (Mondragón et al. 2014) in the region Ω1 = {(r, b) ∊ R2:0 < b ≤ r ≤ 1} given by
| 26 |
According to the Dulac criterion, there exists a continuously differentiable Dulac function Φ(r, b) for a simply connected region Ω1 ⊂ R2 such that
| 27 |
Since (f(r, b), g(r, b)) is the vector field of system (26),
This result reveals that system (26) has no periodic orbits contained in the interior of Ω1 in compliance with Dulac-Bendixon criterion. In addition, the region Γ2 cited in the Proposition 6 does not include another the LAS equilibrium point. Thus, by the Poincaré–Bendixon Theorem and Dulac–Bendixon criterion, we have that equilibrium point E2 = (0, B, B, 1, …, 1) is GAS.
For status (i) obtained from the Table 2, the qualitative analysis of the system (2) has supported by numerical simulations in the following Figs. 1, 2 and 3. Here, it is shown that the equilibrium point E2 is GAS. Also, as a result of using multiple antibiotics at least for 60 days, sensitive bacteria is removed and resistant bacteria and immune cells have same equilibria value of .
Fig. 1.
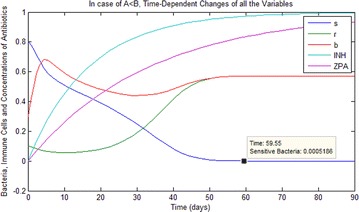
In case of (i) in the Table 2, time-dependent changes of all the variables
Fig. 2.
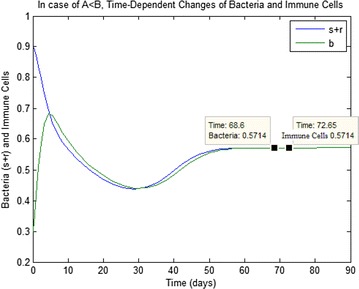
In case of (i) in the Table 2, time-dependent changes of bacteria and immune cells
Fig. 3.
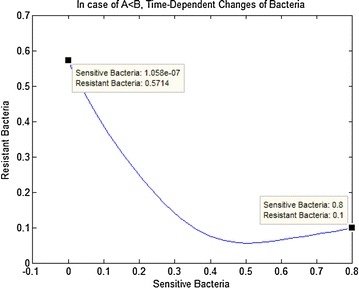
In case of (i) in the Table 2, time-dependent changes of bacteria
Proposition 8
LetB < A. Then,
| 28 |
where the values and are in (22).
Proof
Let . This shows that b is an increasing function and approach asymptotically to by increasing in Ω. From the third equation in system (3), it is obtained that except for (state of equilibrium). In this sense, we have that .
Let . In the same way, b is approaches asymptotically to by decreasing. Thus, it is obtained that . Proposition is proved.
Proposition 9
LetA > B. If and, then equilibrium point is GAS in the region
| 29 |
where.
Proof
If the last equations of system (3) are separated, then their solutions approach for . Replacing these values in the first three equations of system (3), we attain the following:
| 30 |
in the region (29). Considering (9), then the system (30) transforms to
| 31 |
where . Also, it is clear that .
In this context, the GAS of equilibrium point can be obtained by applying the LaSalle-Lyapunov Theorem. Lyapunov function is
| 32 |
where , and are arbitrary positive constants and , and are in (22). Differentiating with respect to , we get
| 33 |
Let . In this case, since and , it is obtained that
| 34 |
respectively. In addition, if , then
| 35 |
| 36 |
From (28) and (36), It is obtained that . In this respect, if and , then equilibrium point is GAS for system (31) in , and so, for system (3) in . Proposition is proved.
For status (ii) obtained from the Table 2, the qualitative analysis of the system (3) has supported by numerical simulations in Fig. 4. In this case, it is found that is GAS. Moreover, sensitive and resistant bacteria to multiple antibiotics and immune system cells have positive equilibrium values as in .
Fig. 4.
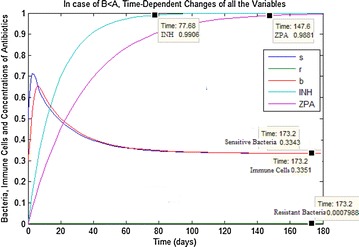
In case of (ii) in the Table 2, time-dependent changes of all the variables
Conclusions
In this paper, we formulated a mathematical model of bacterial resistance to immune system response and multiple antibiotics simultaneously, considering specific changes in bacterial DNA sequence as the only mechanism of bacterial resistance acquisition in order to evaluate the effectiveness of antibiotic treatments with respect to the mechanism above.
The parameter is interpreted as the number of bacteria produced by the fraction of sensitive bacteria that survive to the effects due to antibiotics and immune cells. Similarly, the parameter represents the bacteria produced by resistant bacteria. Also, the parameter is expresses as effect of immune cells on the bacteria .
Our model is quite appropriate when compared to the complexity of biological phenomenon and it predicts in terms of the parameters and when the bacterial progression is either for resistant bacteria and immune cellsas shown in Figs. 1, 2 and 3 or for sensitive and resistant bacteria and immune cells as shown in Fig. 4.
The model suggests that if sensitive bacteria can infect but do not produce sufficient progeny (in case of ) then they can be removed and resistant bacteria continue to survive in balance with the immune cells in the host. When sensitive bacteria persist, the model predicts the scenario that the immune response of host and antibiotics is not enough to eliminate them (in case of ), and therefore both types of bacteria continue to survive and coexist in balance with the immune cells in the host.
According to the results of this analysis, the infection never disappears. Also, the infection is continued by resistant bacteria, when the appropriate antibiotics are used, otherwise sensitive and resistant bacteria. The magnitude of infection depends on the effect of immune system in the first case and multiple antibiotics and immune system in the second case.
These results in our model highlight the fact that those whose immunity response against infections have diminished, suffer from the same bacterial infections more. Furthermore, this model shows that some of the bacterial infections believed to be limited or destroyed, make an individual whose immune system deteriorated suffer more.
Additionally, the results obtained from numerical studies in terms of bacterial infection reveal the affinity substantially with reference to the clinical treatment.
For future work we are planing consider other mechanisms such as the loss of resistance in the resistant bacteria and gaining resistance by conjugation of sensitive and resistant bacteria in order to get more accurate results.
Authors’ contributions
Both authors have contributed to the manuscript equally. Both authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Contributor Information
Bahatdin Daşbaşı, Email: dasbasibahatdin@gmail.com.
İlhan Öztürk, Email: ozturki@erciyes.edu.tr.
References
- Alavez J, Avenda R, Esteva L, et al. Within-host population dynamics of antibiotic-resistant M. tuberculosis. Math Med Biol. 2006;24:35–56. doi: 10.1093/imammb/dql026. [DOI] [PubMed] [Google Scholar]
- Alberts B, Johnson A, Lewis J et al (2002) Molecular biology of the cell. In: The adaptive immune system, 4th edn. Garland Science, New York. ISBN 10: 0-8153-3218-1
- Allen LJS (2007) An introduction to mathematical biology. ISBN 10: 0-13-035216-0
- Andersson DI, Levin BR. The biological cost of antibiotic resistance. Curr Opin Microbiol. 1999;2:489–493. doi: 10.1016/S1369-5274(99)00005-3. [DOI] [PubMed] [Google Scholar]
- Andersson DI, Björkholm B, Sjölund M, et al. Mutation frequency and biological cost of antibiotic resistance in Helicobacterpylori. Proc Natl Acad Sci. 2001;98(25):14607–14612. doi: 10.1073/pnas.241517298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- André JB, Gandon S. Vaccination, within-host dynamics, and virulence evolution. Evolution. 2006;60(1):13–23. doi: 10.1111/j.0014-3820.2006.tb01077.x. [DOI] [PubMed] [Google Scholar]
- Antia R, Koella JC, Perrot V. Models of within-host dynamics of persistent mycobacterial infections. Proc R Soc Lond B. 1996;263:257–263. doi: 10.1098/rspb.1996.0040. [DOI] [PubMed] [Google Scholar]
- Arya DP. Aminoglycoside antibiotics: from chemical biology to drug discovery. New Jersey: Wiley; 2007. [Google Scholar]
- Austin D, Anderson R. Studies of antibiotic resistance within the patient, hospitals and the community using simple mathematical models. Philos Trans R Soc Lond B. 1999;354(1384):721–738. doi: 10.1098/rstb.1999.0425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Austin D, Kakehashi M, Anderson RM. The transmission dynamics of antibiotic-resistant bacteria: the relationship between resistance in commensal organisms and antibiotic consumption. Proc R Soc Lond B. 1997;264(1388):1629–1638. doi: 10.1098/rspb.1997.0227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonhoeffer S, Lipsitch M, Levin BR. Evaluating treatment protocols to prevent antibiotic resistance. Proc Natl Acad Sci USA. 1997;94(22):12106–12111. doi: 10.1073/pnas.94.22.12106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonten M, Austin J, Lipsitch M. Understanding the spread of antibiotic resistant pathogens in hospitals, mathematical models as tools for control. Clin Infect Dis. 2001;33(10):1739–1746. doi: 10.1086/323761. [DOI] [PubMed] [Google Scholar]
- Bootsma MCJ, Van Der Horrst MA, Guryeva T, et al. Modeling non-inherited antibiotic resistance. Bull Math Biol. 2012;74:1691–1705. doi: 10.1007/s11538-012-9731-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butler MS, Buss AD. Natural products—the future scaffolds for novel antibiotics? Biochem Pharmacol. 2006;71(7):919–929. doi: 10.1016/j.bcp.2005.10.012. [DOI] [PubMed] [Google Scholar]
- Carvalho RV, Kleijn J, Meijer A, et al. Modeling innate immuneresponse to early mycobacterium infection. Comput Math Methods Med. 2012;790482:1–12. doi: 10.1155/2012/790482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clatworthy AE, Pierson EP, Hung DT. Targeting virulence: a new paradigm for antimicrobial therapy. Nat Chem Biol. 2007;3:541–548. doi: 10.1038/nchembio.2007.24. [DOI] [PubMed] [Google Scholar]
- Coll P. Fármacos con actividad frente a Mycobacterium tuberculosis. Enferm Infecc Microbiol Clin. 2009;27(8):474–480. doi: 10.1016/j.eimc.2009.06.010. [DOI] [PubMed] [Google Scholar]
- D’Agata E, Magal P, Olivier D, et al. Modeling antibiotic resistance in hospitals: the impact of minimizing treatment duration. J Theor Biol. 2007;249:487–499. doi: 10.1016/j.jtbi.2007.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Onofrio A. A general framework for modeling tumor–immune system competition and immunotherapy: mathematical analysis and biomedical inferences. Phys D. 2005;208:220–235. doi: 10.1016/j.physd.2005.06.032. [DOI] [Google Scholar]
- Esteva L, Mondragon EI, Leyton JR. Un modelo matemático sobre bacterias sensibles y resistentes a antibióticos. Mat Ensen͂ Univ. 2011;19(2):55–73. [Google Scholar]
- Gilchrist M, Coombs D. Evolution of virulence: interdependence, constraints and selection using nested models. Theor Popul Biol. 2006;69:145–153. doi: 10.1016/j.tpb.2005.07.002. [DOI] [PubMed] [Google Scholar]
- Gilchrist M, Sasaki A. Modeling host–parasite coevolution: a nested approach based on mechanistic models. J Theor Biol. 2002;218:289–308. doi: 10.1006/jtbi.2002.3076. [DOI] [PubMed] [Google Scholar]
- Haber M, Levin BR, Kramarz P. Antibiotic control of antibiotic resistance in hospitals: a simulation study. BMC Infect Dis. 2010 doi: 10.1186/1471-2334-10-254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hethcote HW. The mathematics of infectious diseases. SIAM Rev. 2000;42:599–653. doi: 10.1137/S0036144500371907. [DOI] [Google Scholar]
- Johnson PJT, Levin BR. Pharmacodynamics, population dynamics and the evolution of persistence in Staphyloccus aureus. PLoS Genet. 2013;9(1):1–13. doi: 10.1371/journal.pgen.1003123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kostova T. Persistence of viral infections on the population level explained by an immunoepidemiological model. Math Biosci. 2007;206(2):309–319. doi: 10.1016/j.mbs.2005.08.003. [DOI] [PubMed] [Google Scholar]
- Lewis K. Platforms for antibiotic discovery. Nat Rev Drug Discov. 2013;12:371–387. doi: 10.1038/nrd3975. [DOI] [PubMed] [Google Scholar]
- Linares J, Martinez J. Resistencia a los antimicrobianos y virulencia bacteriana. Enferm Infecc Microbiol Clin. 2005;23(2):86–93. doi: 10.1157/13071612. [DOI] [PubMed] [Google Scholar]
- Mahmoud AG, Rice LB. Antifungal agents: mode of action, mechanisms of resistance, and correlation of these mechanisms with bacterial resistance, and correlation. Clin Microbiol Rev. 1999;12(4):501–517. doi: 10.1128/cmr.12.4.501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Massad E, Nascimiento M, Bezerra F. An optimization model for antibiotic use. Appl Math Comput. 2008;201:161–167. [Google Scholar]
- Mohtashemi M, Levins R. Transient dynamics and early diagnosis in infectious disease. J Math Biol. 2001;43:446–470. doi: 10.1007/s002850100103. [DOI] [PubMed] [Google Scholar]
- Mondragón EI, Mosquera S, Cerón M, et al. Mathematical modeling on bacterial resistance to multiple antibiotics caused by spontaneous mutations. BioSystems. 2014;117:60–67. doi: 10.1016/j.biosystems.2014.01.005. [DOI] [PubMed] [Google Scholar]
- Nowak MA, May R. Virus dynamics: mathematical principles of immunology and virology. Oxford: Oxford University Press; 2000. [Google Scholar]
- Opatowski L, Guillemot D, Bolle PY, et al. Contribution of mathematical modeling to the fight against bacterial antibiotic resistance. Curr Opin Infect Dis. 2011;24(3):279–287. doi: 10.1097/QCO.0b013e3283462362. [DOI] [PubMed] [Google Scholar]
- Pugliese A, Gandolfi A. A simple model of pathogen–immune dynamics including specific and non-specific immunity. Math Biosci. 2008;214:73–80. doi: 10.1016/j.mbs.2008.04.004. [DOI] [PubMed] [Google Scholar]
- Rodrigues P, Gomes M, Rebeloc C. Drug resistance in tuberculosis—a reinfection model. Theor Popul Biol. 2007;71:196–212. doi: 10.1016/j.tpb.2006.10.004. [DOI] [PubMed] [Google Scholar]
- Singer B. Mathematical Models of infectious diseases: seeking new tools for planning and evaluating control programs. Suppl Popul Dev Rev. 1984;10:347–365. doi: 10.2307/2807968. [DOI] [Google Scholar]
- Sotto A, Lavigne JP. A mathematical model to guide antibiotic treatment strategies. BMC Med. 2012 doi: 10.1186/1741-7015-10-90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun HR, Lu X, Ruan S. Qualitative analysis of models with different treatment protocols to prevent antibiotic resistance. Math Biosci. 2010;227:56–67. doi: 10.1016/j.mbs.2010.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ternent L, Dyson RJ, Krachler A-M, et al. Bacterial fitness shapes the population dynamics of antibiotic resistant and susceptible bacteria in a model. J Theor Biol. 2014;372:1–11. doi: 10.1016/j.jtbi.2015.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitman A, Ashrafiuon H. Asymptotic theory of an infectious disease model. J Math Biol. 2006;53(2):287–304. doi: 10.1007/s00285-006-0009-y. [DOI] [PubMed] [Google Scholar]
- Wiesch PA, Kouyos R, Engelstadter J, et al. Population biological principles of drug- resistance evolution in infectious diseases. Lancet Infect Dis. 2011;11(3):236–247. doi: 10.1016/S1473-3099(10)70264-4. [DOI] [PubMed] [Google Scholar]
- Zhang Y. Mechanisms of drug resistance in Mycobacterium tuberculosis. Int J Tuberc Lung Dis. 2009;13(11):1320–1330. [PubMed] [Google Scholar]


