Abstract
Background
B. cereus are of particular interest in food safety and public health because of their capacity to cause food spoilage and disease through the production of various toxins. The aim of this study was to determine the prevalence, virulence factor genes and antibiotic resistance profile of B. cereus sensu lato isolated from cattle grazing soils and dairy products in Ghana. A total of 114 samples made up of 25 soil collected from cattle grazing farm land, 30 raw milk, 28 nunu (yoghurt-like product) and 31 woagashie (West African soft cheese). Ninety-six B. cereus sensu lato isolates from 54 positive samples were screened by PCR for the presence of 8 enterotoxigenic genes (hblA, hblC, hblD, nheA, nheB, nheC, cytK and entFM), and one emetic gene (ces). Phenotypic resistance to 15 antibiotics were also determined for 96 B. cereus sensu lato isolates.
Results
About 72% (18 of 25 soil), 47% (14 of 30 raw milk), 35% (10 of 28 nunu) and 39% (12 of 31 woagashi) were positive for B. cereus sensu lato with mean counts (log10 cfu/g) of 4.2 ± 1.8, 3.3 ± 2.0, 1.8 ± 1.4 and 2.6 ± 1.8 respectively. The distribution of enterotoxigenic genes revealed that 13% (12/96 isolates) harboured all three gene encoding for haemolytic enterotoxin HBL complex genes (hblA, hblC and hblD), 25% (24/96 isolates) possessed no HBL gene, whereas 63% (60/96 isolates) possessed at least one of the three HBL genes. All three genes encoding for non-haemolytic enterotoxin (nheA, nheB and nheC) were detected in 60% (57/96) isolates, 14% (13/96) harboured only one gene, 19% (18/96) whereas 8% possessed none of the NHE genes. The detection rates of cytk, entFM, and ces genes were 75, 67 and 9% respectively. Bacillus cereus s. l. isolates were generally resistant to β-lactam antibiotics such as ampicillin (98%), oxacillin (92%), penicillin (100%), amoxicillin (100%), and cefepime (100%) but susceptible to other antibiotics tested.
Conclusions
Bacillus cereus s. l. is prevalent in soil, raw milk and dairy products in Ghana. However, loads are at levels considered to be safe for consumption. Various enterotoxin genes associated with virulence of B. cereus are widespread among the isolates.
Keywords: Bacillus cereus sensu lato, Antibiotic resistance, Virulence gene, Enterotoxin, Dairy product, Emetic toxin gene
Background
The Bacillus cereus group, also known as B. cereus sensu lato, is a species complex which shows high degree of phenotypic and genotypic similarity. The group classically consists of Gram-positive, rod-shaped, spore-forming aerobic bacteria that are widespread in natural environments. The genetic similarity within the B. cereus group has been widely studied [1–6].
Bacillus cereus s. l. has been found to have significant impact on human health, agriculture, and food processing [5]. B. cereus commonly cause spoilage in food products [7]. Additionally, it is an opportunistic pathogen which can cause two types of food poisoning in humans, characterized by either nausea and vomiting or abdominal pain and diarrhea [8, 9].
The virulence of B. cereus sensu lato is attributed to different factors. Diarrheal disease is associated with the production of enterotoxins such as hemolysin BL (HBL), non-hemolytic enterotoxin (NHE), cytotoxin K, and enterotoxin FM [10–15], whiles the virulence of emetic strains is attributed to the production of a heat stable cereulide, synthesized by a non-ribosomal peptide synthetase encoded by ces genes [16]. The emetic toxin, usually preformed in food, is not inactivated during food processing or gastrointestinal passage because it is highly resistant to heat treatments, extreme pH conditions and protease activities [17–19]. Therefore, ingestion of living B. cereus is not necessary for illness of this type to occur. On the other hand, diarrheal food poisoning is not caused by preformed toxins in food, but by viable vegetative B. cereus cells (not spores) producing enterotoxins in the small intestine, because spores do not produce enterotoxins. Additionally, spores are easily degraded under gastrointestinal conditions by the host’s digestive enzymes [20–22].
Ghanaian traditional milk products including nunu (yoghurt-like) and woagashie (cheese-like) are produced mainly by spontaneous fermentations of raw cow milk [23, 24]. These products are consumed widely throughout Ghana. However, there has been increasing public concern about the safety of consuming the Ghanaian milk products due to the crude methods of milk production, handling and processing which expose the products to possible contamination with various potential foodborne pathogens such as B. cereus. To ensure consumer safety and increase patronage of the Ghanaian traditional dairy products, attempts have been made to characterize the dominant microorganisms and to select starter cultures for the production of quality and safe products [24, 25]. The incidence of Bacillus cereus on dairy farms and in milk products has been reported elsewhere, particularly in Scandinavia, the Netherlands, Australia and Brazil [26–29]. However, there is currently no reported study on the prevalence of Bacillus cereus in dairy farms and milk products in Ghana, and their associated virulence and antibiotic resistance profile. This study therefore sought to assess the prevalence, virulence factor genes and phenotypic antibiotic resistance of Bacillus cereus sensu lato isolated from cattle grazing farms, raw milk, and traditional dairy products in Ghana.
Methods
Sampling
A total of 114 samples made up of 25 soil collected from cattle grazing farm land, 30 raw milk, 28 nunu (yoghurt-like product) and 31 woagashie (West African soft cheese) were used for the isolation of B. cereus. Raw milk, nunu and woagashie samples were purchased from retail markets in Tamale in the Northern region of Ghana. Raw unpasteurized milk and its products sampled were originally sourced from dairy farms which do not rely on supplementary feeding or routine antibiotics use. There was also no record of B. cereus infection among the cattle herds. Soil samples were collected from cattle grazing fields located within 10 mile radius in farming communities in the Northern Region of Ghana. Soil samples were taken from sites at least 500 m apart with a sterilized spatula down to a depth of about 20 cm from the ground surface into sterile stomacher bags and transferred to the laboratory for analysis. Sampling was done between January and October, 2015.
Isolation of and identification of B. cereus s. l.
For the isolation of Bacillus cereus s. l., 25 g of each sample was transferred into 225 ml of sterile phosphate buffered saline (PBS) in a sterile stomacher and homogenized for 2 min using BagMixer stomacher (Inter science, St Nom, France). The homogenate was serially diluted (10-fold) in sterile PBS and 0.1 mL of each dilution was inoculated onto duplicated agar plates containing B. cereus agar base (Oxoid, UK) supplemented with 100 ml/l of Egg Yolk Emulsion (Oxoid,) and 5 ml/l of Polymyxin B Selective Supplement (Oxoid). Plates were incubated at 30 °C for 24 h and observed for growth. Suspected B. cereus colonies with blue appearance (typically mannitol-negative) and lecithinase positive (zone of precipitation around colonies) were selected from each plate and sub-cultured on nutrient agar (Oxoid). Suspected colonies were further identified by phenotypic and biochemical tests [30] including cell shape and motility, hemolysis, production of catalase, oxidase, urease and lecithinase, nitrate reduction, fermentation of D-glucose, maltose, D-xylose, lactose and D-mannitol, and growth at a temperature of 10 °C. Bacillus cereus ATCC 11778 and B. cereus ATCC 14579 were used as reference strains for phenotypic tests.
PCR detection of virulence factor genes in B. cereus s. l.
Prior to DNA extraction, bacterial cultures were grown by streaking on nutrient agar and incubating at 30 °C for 24 h for preparation of template DNA for PCR screening. The bacterial genomic DNA of was extracted using the InstaGene Matrix DNA extraction kit following the instructions of the manufacturer (Bio-Rad, Hercules, CA, USA).
PCR screening was done to detect the presence of 8 enterotoxigenic genes (hblA, hblC, hblD, nheA, nheB, nheC, cytK and entFM), and one emetic gene (ces). The primer pair sequences used for the amplification of virulence factor genes of B. cereus in this study are shown in Table 1. The PCR reaction was carried out as described by Kim et al. [31]. Briefly, the PCR reaction mixture contained 25 ng of template DNA, 0.5 U dreamTaq DNA polymerase (Fermentas GmbH, St. Leon-Rot, Germany), 10 mM Tris-HCl (pH 8.3), 10 mM KCl, 0.2 mM each deoxynucleoside triphosphate (dNTPs), 2.5 mM MgCl2, and 1 μM each primer. Sterile MilliQ water was used for the preparation of the PCR mixture and for all negative control reactions. The reaction was performed in an automatic thermal cycler (Biotron, Göttingen, Germany) under the following optimized cycling program: an initial denaturation step of 3 min at 95 °C; 35 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 45 s, extension at 72 °C for 1.5 min; and a final extension at 72 °C for 5 min.
Table 1.
Sequences of PCR primers targeting various targeting various virulent factor genes in this study
| Target gene | Primer | Primer sequence (5′ – 3′) | Amplicon size (bp) | Reference |
|---|---|---|---|---|
| nheA | nheA 344 S | TACGCTAAGGAGGGGCA | 480 | [11] |
| nheA 843 A | GTTTTTATTGCTTCATCGGCT | |||
| nheB | nheB 1500 S | CTATCAGCACTTATGGCAG | 754 | [11] |
| nheB 2269 A | ACTCCTAGCGGTGTTCC | |||
| nheC | nheC 2820 S | CGGTAGTGATTGCTGGG | 564 | [11] |
| nheC 3401 A | CAGCATTCGTACTTGCCAA | |||
| hblA | HBLA1 | GTGCAGATGTTGATGCCGAT | 301 | [11] |
| HBLA2 | ATGCCACTGCGTGGACATAT | |||
| hblC | L2A | AATGGTCATCGGAACTCTAT | 731 | [11] |
| L2B | CTCGCTGTTCTGCTGTTAAT | |||
| hblD | L1A | AATCAAGAGCTGTCACGAAT | 411 | [11] |
| L1B | CACCAATTGACCATGCTAAT | |||
| cytK | CK-F-1859 | ACAGATATCGG(GT)CAAAATGC | 809 | [46] |
| CK-R-2668 | TCCAACCCAGTT(AT)(GC)CAGTTC | |||
| entFM | ENTA | ATGAAAAAAGTAATTTGCAGG | 1, 269 | [63] |
| ENTB | TTAGTATGCTTTTGTGTAACC | |||
| Ces | cesF1 | GGTGACACATTATCATATAAGGTG | 1, 271 | [16] |
| cesR2 | GTAAGCGAACCTGTCTGTAACAACA |
The amplified PCR fragments were analysed by submerged 1.5% agarose gel electrophoresis in 1x buffer (108 g Trisbase/l, 55 g boric acid/l and 40 ml of 0.5 M EDTA, pH 8.0). Following electrophoresis, gels were stained with ethidium bromide, photographed under UV illumination. A reaction mixture without DNA template served as a general control for extraneous nucleic acid contamination. Other controls including sterile MilliQ water and template DNA were used to detect false-positive and false-negative reactions. PCR amplification and electrophoresis experiments were all carried out in triplicates.
Antibiotics susceptibility testing
Resistance/susceptibility to antimicrobials by B. cereus sesu lato isolates were determined in Mueller-Hinton (MH) broth using the broth dilution method recommended by the standard criteria of the Clinical and Laboratory Standards Institute (CLSI) guide [32–35]. The 15 antibiotics including Amoxicillin, Ampicillin, Cefepime, Chloramphenicol, Ciprofloxacin, Clindamycin, Erythromycin, Gentamycin, Oxacillin, Penicillin, Quinupristin/Dalfopristin, Rifampin, Tetracycline, Trimethoprim /sulfumethoxazole and Vancomycin were each diluted in two-fold in the range of 64 to 0.015 mg/L of MH-broth. The final inoculum of B. cereus sesu lato suspension in the broth media were equivalent to approximately 1 × 103 CFU/ml. Growth was carried out at 35 ± 2 °C for 18–20 h incubation period and examined in a microplate reader (OD at 610 nm). The breakpoints against B. cereus in the CLSI guideline M45A2E (2010) and M45-P (2005) were used for all antimicrobial agents except oxacillin and quinupristin/dalfopristin for which the breakpoints for Staphylococcus spp. in the CLSI guideline M 100-S22 (2012) and S24 (2014) were used according to Luna et al. [35].
Results and discussion
Prevalence and phenotypic characteristics of Bacillus cereus s. l.
The prevalence of B. cereus sensu lato in soil from cattle grazing farms, raw milk, nunu, and woagashie are shown in Table 2. Eighteen of 25 (72.0%) soil, 14 of 30 (46.6%) raw milk, 10 of 28 (35.7%) nunu and 12 of 31 (38.7%) woagashi samples were positive for B. cereus sensu lato with mean counts (log10 CFU/g) of 4.2 ± 1.8, 3.3 ± 2.0, 1.8 ± 1.4 and 2.6 ± 1.8 respectively (Table 2). All the isolates showed common phenotypic and biochemical characteristics that are consistent with the identification of Bacillus cereus sensu lato. The isolates were motile rods with peritrichous flagella, and haemolytic with β-haemolysis or lavender-green coloration under heavy growth which is an indication of proteolytic activity. Additionally, they produced catalase, lecithinase, and reduced nitrate. The isolates fermented D-glucose and maltose but were variable in their ability to ferment sucrose and lactose. None of the isolates fermented of D-xlyose and D-mannitol. Production of oxidase and urease was variable while there was no production of indole. All the isolates were able to grow at 10 °C. Thus the phenotypic and biochemical characteristics suggest that the pool of isolates selected did not include B. anthracis which is non-hemolytic, and B. cytotoxicus which has a minimum growth temperature of 20 °C [36]. Previous reports suggests that B. cereus strains isolated from dairy products adapt to environmental culture conditions [37] which might explain the ability of some B. cereus isolated from milk to ferment lactose. In analysing 334 samples of pasteurized milk, TeGiffel et al. [37] found that 40% of the samples were contaminated with B. cereus, out of which 53% of the B. cereus isolates could grow at 7 °C, and 20% fermented lactose which is an uncommon carbon source for B. cereus [37]. Similarly, 17 B. cereus s. l. isolated from milk and dairy products in the current work fermented lactose, whiles no isolate from soil could ferment lactose.
Table 2.
Prevalence of B. cereus sensu lato in soil, raw milk and milk products
| Sample type | Number of positive samples (%) | Mean counta (log CFU/g) |
|---|---|---|
| Soils (n = 25) | 18 (72.0) | 4.2 ± 1.8 |
| Raw milk (n = 30) | 14 (46.7) | 3.3 ± 2.0 |
| Nunu (n = 28) | 10 (35.7) | 1.8 ± 1.4 |
| Woagashie (n = 31) | 12 (38.7) | 2.6 ± 1.8 |
aValues are means ± standard deviations (SD)
B. cereus is widespread in the environment and shows great ecological diversity, enhancing their ability to contaminate many raw and finished food products including milk and milk products, although usually at low levels of ≤ 103 CFU/g [38–42]. In the present study, raw milk and traditional milk products (nunu and woagashie) all had B. cereus counts of <4 log CFU/g. In general, it is estimated that consumption of food containing B. cereus cells and/or spores between 105 and 108 can cause disease [9, 43]. Therefore, the load of B. cereus sensu lato in milk and milk products in the present study are within acceptable limits for consumption according to the EFSA recommended level of <105 CFU/g at the point of consumption. However, there are reported cases of both emetic and diarrhoeal diseases involving lower levels (below 105 cfu/g) of B. cereus [43]. Therefore the potential for B. cereus infections through the consumption of unpasteurized milk and milk products in Ghana cannot be underestimated. Additionally, Ghana lacks proper foodborne diseases surveillance systems to provide reliable data on the burden of foodborne illnesses involving B. cereus in milk and milk products. Thus a number of illnesses or sporadic outbreaks of B. cereus infections resulting from the consumption of unpasteurized milk and milk products may go unreported. Because B. cereus are generally resident flora of soil and frequently associated with farm environments and the fecal shedding of cattle, there is a higher risk of contamination of milk, and subsequent entry into the dairy food chain where they can cause spoilage and/or diseases. It is therefore important for dairy farmers and processors of traditional milk products to practice high level of good hygienic practices (GHP) and Good manufacturing practices (GMP), as well as implement the use of starter cultures for the production fermented dairy products.
Distribution of virulence factor genes among B. cereus s. l. isolates
PCR based detection of Virulence factors in B. cereus sensu lato targeted genes encoding enterotoxins and emetic toxin. These included genes encoding haemolytic (hblA, hblC, and hblD) and non-haemolytic (nheA, nheB, and nheC) enterotoxin complexes, cytotoxin K (cytK), enterotoxin FM (entFM), and cereulide (ces) as shown in Fig. 1. All primers used produced amplicons of the expected size from their respective target virulence genes with reproducible results in repeated experiments. The distribution of virulence genes among 96 B. cereus s. l. isolates are shown in Table 3. For HNE encoding genes, about 60% (58/96) of B. cereus s. l. isolates were found to harbour simultaneously the nheABC genes, 13% (12/96) harboured only one gene, 19% (18/96) harboured simultaneously two genes, and 8% possessed none of the NHE encoding genes. For HBL encoding genes, 38% (37/96) possessed only one gene, 24% (23/96) possessed simultaneously two genes, 13% (12/96) possessed simultaneously all three bhlACD genes, and 25% (24/96) possessed no HBL encoding gene at all. The prevalence of cytk, entFM, and the emetic gene ces among B. cereus s. l. isolates were 75, 67 and 9% respectively. The emetic gene was only detected in B. cereus s. l. isolated from milk and milk products but not from soil samples.
Fig. 1.
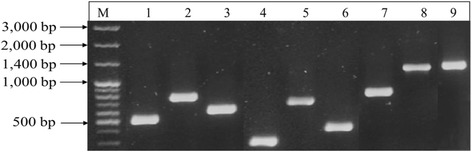
Representative PCR products detecting various virulence factor genes in B. cereus s. l. isolated from soil and various dairy products. Lane M, 100 bp molecular size DNA marker; lane 1, nheA; lane 2, nheB; lane 3, nheC; lane 4, hblA; lane 5, hblC; lane 6, hblD; lane 7, cytK; lane 8, entFM; lane 9, Ces
Table 3.
Distribution of enterotoxin and emetic toxin genes in B. cereus senso lato isolated from dairy farm and milk product
| Toxigenic genes | Number of strains (%)a positive for target gene(s) | ||||
|---|---|---|---|---|---|
| Soil (n = 30) | Raw milk (n = 24) | Nunu (n = 18) | Woagashie (n = 24) | Total (n = 96) | |
| NHE gene complexes | |||||
| nheA | 3 (10) | 1 (4) | 0 (0) | 2 (8) | 6 (6) |
| nheB | 1 (3) | 1 (4) | 0 (0) | 0 (0) | 2 (2) |
| nheC | 1 (3) | 0 (0) | 1 (6) | 2 (8) | 4 (4) |
| nheA + nheB | 2 (7) | 2 (8) | 0 (0) | 1 (4) | 5 (5) |
| nheA + nheC | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| nheB + nheC | 2 (7) | 3 (13) | 4 (22) | 4 (17) | 13 (14) |
| nheA + nheB + nheC | 19 (63) | 15 (63) | 9 (50) | 15 (63) | 58 (60) |
| None detected | 2 (7) | 2 (8) | 4 (22) | 0 (0) | 8 (8) |
| HBL gene complexes | |||||
| hblA | 4 (13) | 4 (17) | 1 (6) | 2 (8) | 11 (11) |
| hblC | 1 (3) | 5 (21) | 3 (17) | 1 (4) | 10 (10) |
| hblD | 5 (17) | 2 (8) | 2 (11) | 7 (29) | 16 (17) |
| hblA + hblC | 0 (0) | 3 (13) | 0 (0) | 0 (0) | 3 (3) |
| hblA + hblD | 6 (20) | 2 (8) | 1 (6) | 3 (13) | 12 (13) |
| hblC + hblD | 1 (3) | 3 (13) | 0 (0) | 4 (17) | 8 (8) |
| hblA + hblC + hblD | 5 (17) | 1 (4) | 4 (22) | 2 (8) | 12 (13) |
| None detected | 8 (27) | 4 (17) | 7 (39) | 5 (21) | 24 (25) |
| Other genes | |||||
| cytK | 9 (30) | 21 (88) | 18 (100) | 24 (100) | 72 (75) |
| entFM | 19 (63) | 17 (71) | 15 (83) | 13 (54) | 64 (67) |
| Ces | 0 (0) | 5 (21) | 1 (6) | 3 (13) | 9 (9) |
apercentages have been converted to the nearest whole numbers
Virulence factors HBL, NHE and cytotoxin K are primarily responsible for the production of B. cereus s. l. enterotoxins [12, 14, 44]. The isolated B. cereus s. l. commonly possessed cytk (75%) as the most prevalent toxin gene followed by entFM (67%). Similar results of high prevalence of nheABC and entFM gene complexes have previously been reported to be widespread among wild B. cereus isolates from various food and environmental sources [30, 45–48], as well as some reference strains [31]. Similarly, various studies have reported higher prevalence rates, usually between 40 and 70.6% of the HBL gene complex in B. cereus isolated from milk and dairy products [28, 49–51]. The prevalence rate of the emetic toxin gene ces was 9%. They were however detected only in B. cereus s. l. isolated milk and milk product but not from soil samples. Emetic toxin producing genes have previously been detected at different low rates (1.5 to 17.2%) in isolated B. cereus strains isolated from various food sources [42, 48, 52]; the different prevalence rates being attributed to the differences in food property [53–55]. Kim et al., [31], did not successfully generate amplicons for the emetic gene, ces, in both reference and commercial strains of B. cereus. These findings therefore seems to suggest that emetic toxin genes are not highly prevalent or are rare among B. cereus isolates.
Resistance of B. cereus s. l. to antibiotics
Resistance/susceptibility to different antimicrobial agents by B. cereus s. l. are shown in Table 4. Irrespective of the origin (soil, milk or milk products) of the isolates, they were generally resistant to ampicillin (98%), oxacillin (92%), penicillin (100%), amoxicillin (100%), cefepime (100%) and trimethoprim/sulfamethoxazole (80% with 20% intermediate resistant strains). They were however susceptible to other antimicrobials such as Chloramphenicol (99%), Ciprofloxacin (100%), Clindamycin (100%), Erythromycin (92%), Gentamicin (100%), Quinupristin/dalfopristin (100%), Rifampin (100%), Tetracycline (97%) and Vancomycin (100%).
Table 4.
Resistance to antimicrobials by B. cereus sensu lato isolated from dairy farms and milk products
| Antibiotic | aBreakpoints | bInterpretation n (%) | |||
|---|---|---|---|---|---|
| S (≤) | R (≥) | S | I | R | |
| Amoxicillin | 4 | 8 | 0 | 0 | 96 (100) |
| Ampicillin | 0.25 | 0.5 | 0 | 2 (0.02) | 94 (98) |
| Cefepime | 0 | 0 | 0 | 0 | 96 (100) |
| Chloramphenicol | 8 | 32 | 95 (99) | 1 (0.01) | 0 |
| Ciprofloxacin | 1 | 4 | 96 (100) | 0 | 0 |
| Clindamycin | 0.5 | 4 | 96 (100) | 0 | 0 |
| Erythromycin | 0.5 | 4 | 88 (92) | 8 (8) | 0 |
| Gentamicin | 4 | 16 | 96 (100) | 0 | 0 |
| Oxacillin | 2 | 4 | 3 (3) | 5 (5) | 88 (92) |
| Penicillin | 0.12 | 0.25 | 0 | 0 | 96 (100) |
| Quinupristin/dalfopristin | 1 | 4 | 96 (100) | 0 | 0 |
| Rifampin | 1 | 4 | 96 (100) | 0 | 0 |
| Tetracycline | 4 | 16 | 93 (97) | 3 (3) | 0 |
| Trimethoprim/sulfamethoxazole | 2 | 4 | 0 | 19 (20) | 77 (80) |
| Vancomycin | 4 | >4 | 96 (100) | 0 | 0 |
aThe breakpoints against B. cereus in the CLSI guideline M45A2E (2010) and M45-P (2005) were used for all antimicrobial agents except oxacillin and quinupristin/dalfopristin for which the breakpoints for Staphylococcus spp. in the CLSI guidelines M 100-S22 (2012) and S24 (2014) were used
b S susceptible, I intermediate, R resistant
Because B. cereus have clinical significance, determining their resistance or otherwise to antimicrobial agents is critical for treatment during outbreaks. Previous reports have shown that B. cereus is susceptible to imipenem and vancomycin, and most strains are sensitive to chloramphenicol, aminoglycosides, ciprofloxacin, erythromycin, and gentamicin [56–59]. Some strains of B. cereus are moderately sensitive to clindamycin and tetracycline [58]. In this report, B. cereus s. l. isolated from soil from cattle grazing fields and milk and dairy products were predominantly resistant to β-lactam antibiotics. The abundant production of β-lactamases by bacteria including Bacillus species is a common cause of antibiotic resistance in bacteria [39, 60]. Wild-type genomes of many bacteria, including Bacillus species have been found to possess genes encoding the production of β-Lactamase. However, these chromosomal β-lactamases do not generally provide effective antibiotic resistance in wild-type bacilli, despite evidence that the genes are not completely silenced [61, 62].
Conclusions
Bacillus cereus sensu lato is prevalent in soil from cattle grazing fields, raw milk and traditional dairy products, but load in milk and traditional dairy products are at levels considered to be safe for consumption. However, various enterotoxin genes associated with the virulence of B. cereus are widespread among isolates. Additionally, the B. cereus s. l. isolates were generally resistant to β-lactam antibiotics but susceptible to other antibiotics. There is therefore the need to observe good hygienic and manufacturing practice by milk producers and traditional dairy processors to prevent contamination and subsequent potential disease outbreak by B. cereus.
Acknowledgements
Financial assistance provided by Danida (Danish International Development Agency, Ministry of Foreign Affairs) and the Government of Ghana, through the project ‘Preserving African Food Microorganisms for Green Growth’ DFC No. 13-04KU, is sincerely acknowledged. We are also grateful to management of the Savana Agricultural Research Institute (SARI), Tamale-Ghana, for allowing us access to their molecular biology laboratory facilities.
Funding
None.
Availability of data and materials
All data generated or analyzed during this study are included in this article.
Authors’ contributions
JOK conceived the project idea, participated in laboratory experiments, analysis of data and wrote the manuscript. AW carried out most of the laboratory experiments, analyzed and interpreted the data. FA carried out part of the laboratory analysis, interpretation of the results, and reviewed the manuscript. KTD and LJ supervised the project, participated in analysis and interpretation of results and corrected the manuscript. All authors read and approved the content of the manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- CLSI
Clinical and Laboratory Standards Institute
- DNA
de-oxyribonucleic acid
- EDTA
Ethylenediaminetetraacetic acid
- EFSA
European food safety authority
- GHP
Good hygienic practices
- GMP
Good manufacturing practices
- HBL
Hemolysin BL
- MH
Muellar-Hinton
- NHE
Non-hemolytic enterotoxin
- PCR
Polymerase chain reaction
- s.l.
Sensu lato
Contributor Information
James Owusu-Kwarteng, Phone: +233 209265738, Email: jowusukwarteng@uds.edu.gh.
Alhassan Wuni, Email: alhassanwuni@yahoo.com.
Fortune Akabanda, Email: fakabanda@gmail.com.
Kwaku Tano-Debrah, Email: ktanode@ug.edu.gh.
Lene Jespersen, Email: lj@food.ku.dk.
References
- 1.Guinebretière MH, Thompson FL, Sorokin A, Normand P, Dawyndt P, Ehling‐Schulz M, Svensson B, Sanchis V, Nguyen‐The C, Heyndrickx M. Ecological diversification in the Bacillus cereus group. Environ Microbiol. 2008;10(4):851–65. doi: 10.1111/j.1462-2920.2007.01495.x. [DOI] [PubMed] [Google Scholar]
- 2.Helgason E, Økstad OA, Caugant DA, Johansen HA, Fouet A, Mock M, Hegna I, Kolstø A-B. Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis - one species on the basis of genetic evidence. Appl Environ Microbiol. 2000;66(6):2627–30. doi: 10.1128/AEM.66.6.2627-2630.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Liu Y, Lai Q, Göker M, Meier-Kolthoff JP, Wang M, Sun Y, Wang L, Shao Z. Genomic insights into the taxonomic status of the Bacillus cereus group. Sci Rep. 2015;5:14082. doi: 10.1038/srep14082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Priest FG, Barker M, Baillie LW, Holmes EC, Maiden MC. Population structure and evolution of the Bacillus cereus group. J Bacteriol. 2004;186(23):7959–70. doi: 10.1128/JB.186.23.7959-7970.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rasko DA, Altherr MR, Han CS, Ravel J. Genomics of the Bacillus cereus group of organisms. FEMS Microbiol Rev. 2005;29(2):303–29. doi: 10.1016/j.femsre.2004.12.005. [DOI] [PubMed] [Google Scholar]
- 6.Van der Auwera GA, Feldgarden M, Kolter R, Mahillon J. Whole-genome sequences of 94 environmental isolates of Bacillus cereus sensu lato. Genome Announc. 2013;1:5. doi: 10.1128/genomeA.00380-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Arslan S, Eyi A, Küçüksarı R. Toxigenic genes, spoilage potential, and antimicrobial resistance of Bacillus cereus group strains from ice cream. Anaerobe. 2014;25:42–6. doi: 10.1016/j.anaerobe.2013.11.006. [DOI] [PubMed] [Google Scholar]
- 8.Fricker M, Messelhäußer U, Busch U, Scherer S, Ehling-Schulz M. Diagnostic real-time PCR assays for the detection of emetic Bacillus cereus strains in foods and recent food-borne outbreaks. Appl Environ Microbiol. 2007;73(6):1892–8. doi: 10.1128/AEM.02219-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Granum PE, Lund T. Bacillus cereus and its food poisoning toxins. FEMS Microbiol Lett. 1997;157(2):223–8. doi: 10.1111/j.1574-6968.1997.tb12776.x. [DOI] [PubMed] [Google Scholar]
- 10.Beecher DJ, Wong AC. Tripartite haemolysin BL: isolation and characterization of two distinct homologous sets of components from a single Bacillus cereus isolate. Microbiology. 2000;146(6):1371–80. doi: 10.1099/00221287-146-6-1371. [DOI] [PubMed] [Google Scholar]
- 11.Fagerlund A, Ween O, Lund T, Hardy SP, Granum PE. Genetic and functional analysis of the cytK family of genes in Bacillus cereus. Microbiology. 2004;150(8):2689–97. doi: 10.1099/mic.0.26975-0. [DOI] [PubMed] [Google Scholar]
- 12.Granum PE, O’sullivan K, Lund T. The sequence of the non-haemolytic enterotoxin operon from Bacillus cereus. FEMS Microbiol Lett. 1999;177(2):225–9. doi: 10.1111/j.1574-6968.1999.tb13736.x. [DOI] [PubMed] [Google Scholar]
- 13.Hansen BM, Hendriksen NB. Detection of enterotoxic Bacillus cereus and Bacillus thuringiensis strains by PCR analysis. Appl Environ Microbiol. 2001;67(1):185–9. doi: 10.1128/AEM.67.1.185-189.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lund T, De Buyser ML, Granum PE. A new cytotoxin from Bacillus cereus that may cause necrotic enteritis. Mol Microbiol. 2000;38(2):254–61. doi: 10.1046/j.1365-2958.2000.02147.x. [DOI] [PubMed] [Google Scholar]
- 15.Sergeev N, Distler M, Vargas M, Chizhikov V, Herold KE, Rasooly A. Microarray analysis of Bacillus cereus group virulence factors. J Microbiol Methods. 2006;65(3):488–502. doi: 10.1016/j.mimet.2005.09.013. [DOI] [PubMed] [Google Scholar]
- 16.Ehling-Schulz M, Vukov N, Schulz A, Shaheen R, Andersson M, Märtlbauer E, Scherer S. Identification and partial characterization of the nonribosomal peptide synthetase gene responsible for cereulide production in emetic Bacillus cereus. Appl Environ Microbiol. 2005;71(1):105–13. doi: 10.1128/AEM.71.1.105-113.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Agata N, Ohta M, Yokoyama K. Production of Bacillus cereus emetic toxin (cereulide) in various foods. Int J Food Microbiol. 2002;73(1):23–7. doi: 10.1016/S0168-1605(01)00692-4. [DOI] [PubMed] [Google Scholar]
- 18.Rajkovic A, Uyttendaele M, Vermeulen A, Andjelkovic M, Fitz‐James I, Denon Q, Verhé R, Debevere J. Heat resistance of Bacillus cereus emetic toxin, cereulide. Lett Appl Microbiol. 2008;46(5):536–41. doi: 10.1111/j.1472-765X.2008.02350.x. [DOI] [PubMed] [Google Scholar]
- 19.Shinagawa K, Konuma H, Sekita H, Sugii S. Emesis of rhesus monkeys induced by intragastric administration with the HEp-2 vacuolation factor (cereulide) produced by Bacillus cereus. FEMS Microbiol Lett. 1995;130(1):87–90. doi: 10.1016/0378-1097(95)00188-B. [DOI] [PubMed] [Google Scholar]
- 20.Ceuppens S, Rajkovic A, Hamelink S, Van de Wiele T, Boon N, Uyttendaele M. Enterotoxin production by Bacillus cereus under gastrointestinal conditions and their immunological detection by commercially available kits. Foodborne Pathog Dis. 2012;9(12):1130–6. doi: 10.1089/fpd.2012.1230. [DOI] [PubMed] [Google Scholar]
- 21.Granum PE, Brynestad S, Kramer JM. Analysis of enterotoxin production by Bacillus cereus from dairy products, food poisoning incidents and non-gastrointestinal infections. Int J Food Microbiol. 1993;17(4):269–79. doi: 10.1016/0168-1605(93)90197-O. [DOI] [PubMed] [Google Scholar]
- 22.Wijnands L, Dufrenne J, Zwietering M, Van Leusden F. Spores from mesophilic Bacillus cereus strains germinate better and grow faster in simulated gastro-intestinal conditions than spores from psychrotrophic strains. Int J Food Microbiol. 2006;112(2):120–8. doi: 10.1016/j.ijfoodmicro.2006.06.015. [DOI] [PubMed] [Google Scholar]
- 23.Akabanda F, Owusu-Kwarteng J, Glover R, Tano-Debrah K. Microbiological characteristics of Ghanaian traditional fermented milk product, Nunu. Nat Sci. 2010;8(9):178–87. [Google Scholar]
- 24.Akabanda F, Owusu-Kwarteng J, Tano-Debrah K, Glover RL, Nielsen DS, Jespersen L. Taxonomic and molecular characterization of lactic acid bacteria and yeasts in nunu, a Ghanaian fermented milk product. Food Microbiol. 2013;34(2):277–83. doi: 10.1016/j.fm.2012.09.025. [DOI] [PubMed] [Google Scholar]
- 25.Akabanda F, Owusu-Kwarteng J, Tano-Debrah K, Parkouda C, Jespersen L. The use of lactic acid bacteria starter culture in the production of Nunu, a spontaneously fermented milk product in Ghana. Int J Food Sci. 2014;2014:721067. doi: 10.1155/2014/721067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Christiansson A, Bertilsson J, Svensson B. Bacillus cereus spores in raw milk: factors affecting the contamination of milk during the grazing period. J Dairy Sci. 1999;82(2):305–14. doi: 10.3168/jds.S0022-0302(99)75237-9. [DOI] [PubMed] [Google Scholar]
- 27.Dréan P, McAuley CM, Moore SC, Fegan N, Fox EM. Characterization of the spore-forming Bacillus cereus sensu lato group and Clostridium perfringens bacteria isolated from the Australian dairy farm environment. BMC Microbiol. 2015;15(1):1. doi: 10.1186/s12866-015-0377-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Reis AL, Montanhini M, Bittencourt JV, Destro MT, Bersot LS. Gene detection and toxin production evaluation of hemolysin BL of Bacillus cereus isolated from milk and dairy products marketed in Brazil. Braz J Microbiol. 2013;44(4):1195–8. doi: 10.1590/S1517-83822013000400024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Vissers M, Te Giffel M, Driehuis F, De Jong P, Lankveld J. Minimizing the level of Bacillus cereus spores in farm tank milk. J Dairy Sci. 2007;90(7):3286–93. doi: 10.3168/jds.2006-873. [DOI] [PubMed] [Google Scholar]
- 30.Kim J-B, Kim J-M, Kim S-Y, Kim J-H, Park Y-B, Choi N-J, Oh D-H. Comparison of enterotoxin production and phenotypic characteristics between emetic and enterotoxic Bacillus cereus. J Food Prot. 2010;73(7):1219–24. doi: 10.4315/0362-028X-73.7.1219. [DOI] [PubMed] [Google Scholar]
- 31.Kim M-J, Han J-K, Park J-S, Lee J-S, Lee S-H, Cho J-I, Kim K-S. Various enterotoxin and other virulence factor genes widespread among Bacillus cereus and Bacillus thuringiensis strains. J Microb Biotechnol. 2015;25(6):872–9. doi: 10.4014/jmb.1502.02003. [DOI] [PubMed] [Google Scholar]
- 32.CLSI: Clinical and Laboratory Standards Institute (CLSI) Methods for antimicrobial dilution and disk susceptibility testing of infrequently isolated or fastidious bacteria; approved guideline M45A2E. 2. Wayne: CLSI; 2010. [Google Scholar]
- 33.CLSI: Clinical and Laboratory Standards Institute (CLSI) Performance standards for antimicrobial susceptibility testing; twenty-second informational supplement M100-S22. Wayne: CLSI; 2012. [Google Scholar]
- 34.CLSI: Clinical and Laboratory Standards Institute (CLSI) Performance standards for antimicrobial susceptibility testing; twenty-fourth informational supplement M100-S24. Wayne: CLSI; 2014. [Google Scholar]
- 35.Luna VA, King DS, Gulledge J, Cannons AC, Amuso PT, Cattani J. Susceptibility of Bacillus anthracis, Bacillus cereus, Bacillus mycoides, Bacillus pseudomycoides and Bacillus thuringiensis to 24 antimicrobials using Sensititre® automated microbroth dilution and Etest® agar gradient diffusion methods. J Antimicrob Chemother. 2007;60(3):555–67. doi: 10.1093/jac/dkm213. [DOI] [PubMed] [Google Scholar]
- 36.Guinebretière M-H, Velge P, Couvert O, Carlin F, Debuyser M-L. Ability of Bacillus cereus group strains to cause food poisoning varies according to phylogenetic affiliation (groups I to VII) rather than species affiliation. J Clin Microbiol. 2010;48(9):3388–91. doi: 10.1128/JCM.00921-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Te Giffel M, Beumer R, Granum P, Rombouts F. Isolation and characterisation of Bacillus cereus from pasteurised milk in household refrigerators in the Netherlands. Int J Food Microbiol. 1997;34(3):307–18. doi: 10.1016/S0168-1605(96)01204-4. [DOI] [PubMed] [Google Scholar]
- 38.Choma C, Guinebretiere M, Carlin F, Schmitt P, Velge P, Granum P, Nguyen‐The C. Prevalence, characterization and growth of Bacillus cereus in commercial cooked chilled foods containing vegetables. J Appl Microbiol. 2000;88(4):617–25. doi: 10.1046/j.1365-2672.2000.00998.x. [DOI] [PubMed] [Google Scholar]
- 39.Park Y-B, Kim J-B, Shin S-W, Kim J-C, Cho S-H, Lee B-K, Ahn J, Kim J-M, Oh D-H. Prevalence, genetic diversity, and antibiotic susceptibility of Bacillus cereus strains isolated from rice and cereals collected in Korea. J Food Prot. 2009;72(3):612–7. doi: 10.4315/0362-028X-72.3.612. [DOI] [PubMed] [Google Scholar]
- 40.Rahmati T, Labbe R. Levels and toxigenicity of Bacillus cereus and Clostridium perfringens from retail seafood. J Food Prot. 2008;71(6):1178–85. doi: 10.4315/0362-028X-71.6.1178. [DOI] [PubMed] [Google Scholar]
- 41.Samapundo S, Heyndrickx M, Xhaferi R, Devlieghere F. Incidence, diversity and toxin gene characteristics of Bacillus cereus group strains isolated from food products marketed in Belgium. Int J Food Microbiol. 2011;150(1):34–41. doi: 10.1016/j.ijfoodmicro.2011.07.013. [DOI] [PubMed] [Google Scholar]
- 42.Wijnands L, Dufrenne J, Rombouts F, Van Leusden F. Prevalence of potentially pathogenic Bacillus cereus in food commodities in the Netherlands. J Food Prot. 2006;69(11):2587–94. doi: 10.4315/0362-028X-69.11.2587. [DOI] [PubMed] [Google Scholar]
- 43.EFSA Biohazrd Panel (EFSA panel on biological hazards) Scientific opinion on the risk for public health related to the precence of Bacillus cereus and other Bacillus spp. including Bacillus thuringiensis in foodstuffs. EFSA J. 2016;14(7):4524, 93. [Google Scholar]
- 44.Hendriksen NB, Hansen BM, Johansen JE. Occurrence and pathogenic potential of Bacillus cereus group bacteria in a sandy loam. Antonie Van Leeuwenhoek. 2006;89(2):239–49. doi: 10.1007/s10482-005-9025-y. [DOI] [PubMed] [Google Scholar]
- 45.Chaves JQ, Cavados CFG, Vivoni AM. Molecular and toxigenic characterization of Bacillus cereus and Bacillus thuringiensis strains isolated from commercial ground roasted coffee. J Food Prot. 2012;75(3):518–22. doi: 10.4315/0362-028X.JFP-11-325. [DOI] [PubMed] [Google Scholar]
- 46.Guinebretière M-H, Broussolle V. Enterotoxigenic profiles of food-poisoning and food-borne Bacillus cereus strains. J Clin Microbiol. 2002;40(8):3053–6. doi: 10.1128/JCM.40.8.3053-3056.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Yang I, Shih DY-C, Huang T-P, Huang Y-P, Wang J-Y, Pan T-M. Establishment of a novel multiplex PCR assay and detection of toxigenic strains of the species in the Bacillus cereus group. J Food Prot. 2005;68(10):2123–30. doi: 10.4315/0362-028X-68.10.2123. [DOI] [PubMed] [Google Scholar]
- 48.Yim JH, Kim KY, Chon JW, Kim DH, Kim HS, Choi DS, Choi IS, Seo KH. Incidence, antibiotic susceptibility, and toxin profiles of Bacillus cereus sensu lato isolated from Korean fermented soybean products. J Food Sci. 2015;80(6):M1266–70. doi: 10.1111/1750-3841.12872. [DOI] [PubMed] [Google Scholar]
- 49.Aragon-Alegro LC, Palcich G, Lopes GV, Ribeiro VB, Landgraf M, Destro MT. Enterotoxigenic and genetic profiles of Bacillus cereus strains of food origin in Brazil. J Food Prot. 2008;71(10):2115–8. doi: 10.4315/0362-028X-71.10.2115. [DOI] [PubMed] [Google Scholar]
- 50.Chitov T, Dispan R, Kasinrerk W. Incidence and diarrhegenic potential of Bacillus cereus in pasteurized milk and cereal products in Thailand. J Food Saf. 2008;28(4):467–81. doi: 10.1111/j.1745-4565.2008.00125.x. [DOI] [Google Scholar]
- 51.Rather MA, Aulakh RS, Gill JPS, Rao TS, Hassan MN. Direct detection of Bacillus cereus and its enterotoxigenic genes in meat and meat products by polymerase Chain reaction. J Adv Vet Res. 2011;1(3):99–104. [Google Scholar]
- 52.Hoton FM, Fornelos N, N’guessan E, Hu X, Swiecicka I, Dierick K, Jääskeläinen E, Salkinoja‐Salonen M, Mahillon J. Family portrait of Bacillus cereus and Bacillus weihenstephanensis cereulide‐producing strains. Environ Microbiol Rep. 2009;1(3):177–83. doi: 10.1111/j.1758-2229.2009.00028.x. [DOI] [PubMed] [Google Scholar]
- 53.Finlay W, Logan N, Sutherland A. Bacillus cereus emetic toxin production in cooked rice. Food Microbiol. 2002;19(5):431–9. doi: 10.1006/fmic.2002.0505. [DOI] [Google Scholar]
- 54.Naranjo M, Denayer S, Botteldoorn N, Delbrassinne L, Veys J, Waegenaere J, Sirtaine N, Driesen RB, Sipido KR, Mahillon J. Sudden death of a young adult associated with Bacillus cereus food poisoning. J Clin Microbiol. 2011;49(12):4379–81. doi: 10.1128/JCM.05129-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zuberovic MA, Tröger R, Granelli K, Hellenäs K-E. Quantitative analysis of cereulide toxin from Bacillus cereus in rice and pasta using synthetic cereulide standard and 13C6-cereulide standard - a short validation study. Toxins. 2014;6(12):3326–35. doi: 10.3390/toxins6123326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.David DB, Kirkby GR, Noble BA. Bacillus cereus endophthalmitis. Br J Ophthalmol. 1994;78(7):577. doi: 10.1136/bjo.78.7.577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Logan NA, Rodrigez-Diaz M. Bacillus spp. and related genera. Principles Pract Clin Bacteriol. 2006;2:139–58. doi: 10.1002/9780470017968.ch9. [DOI] [Google Scholar]
- 58.Murray PR, Barron EJ, Jorgensen JH, Landry ML, Pfaller MA, editors. Manual of clinical microbiology. 9. Washington: ASM Press; 2007. [Google Scholar]
- 59.Rosovitz M, Voskuil M, Chambliss G. Bacillus. Systematic bacteriology. London: Arnold Press; 1998. pp. 709–20. [Google Scholar]
- 60.Majiduddin FK, Materon IC, Palzkill TG. Molecular analysis of beta-lactamase structure and function. Int J Med Microbiol. 2002;292(2):127–37. doi: 10.1078/1438-4221-00198. [DOI] [PubMed] [Google Scholar]
- 61.Chen Y, Succi J, Tenover FC, Koehler TM. β-Lactamase genes of the penicillin-susceptible Bacillus anthracis Sterne strain. J Bacteriol. 2003;185(3):823–30. doi: 10.1128/JB.185.3.823-830.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Ozer JH, Lowery DL, Saz AK. Derepression of β-Lactamase (Penicillinase) in Bacillus cereus by Peptidoglycans. J Bacteriol. 1970;102(1):52–63. doi: 10.1128/jb.102.1.52-63.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Asano S-I, Nukumizu Y, Bando H, Iizuka T, Yamamoto T. Cloning of novel enterotoxin genes from Bacillus cereus and Bacillus thuringiensis. Appl Environ Microbiol. 1997;63:1054–7. doi: 10.1128/aem.63.3.1054-1057.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data generated or analyzed during this study are included in this article.


