Abstract
The human gastric pathogen Helicobacter pylori causes chronic gastritis, peptic ulcer disease, gastric carcinoma, and mucosa-associated lymphoid tissue (MALT) lymphoma. It infects over 50% of the worlds' population, however, only a small subset of infected people experience H. pylori-associated illnesses. Associations with disease-specific factors remain enigmatic years after the genome sequences were deciphered. Infection with strains of Helicobacter pylori that carry the cytotoxin-associated antigen A (cagA) gene is associated with gastric carcinoma. Recent studies revealed mechanisms through which the cagA protein triggers oncopathogenic activities. Other candidate genes such as some members of the so-called plasticity region cluster are also implicated to be associated with carcinoma of stomach. Study of the evolution of polymorphisms and sequence variation in H. pylori populations on a global basis has provided a window into the history of human population migration and co-evolution of this pathogen with its host. Possible symbiotic relationships were debated since the discovery of this pathogen. The debate has been further intensified as some studies have posed the possibility that H. pylori infection may be beneficial in some humans. This assumption is based on increased incidence of gastro-oesophageal reflux disease (GERD), Barrett's oesophagus and adenocarcinoma of the oesophagus following H. pylori eradication in some countries. The contribution of comparative genomics to our understanding of the genome organisation and diversity of H. pylori and its pathophysiological importance to human healthcare is exemplified in this review.
Introduction
Helicobacter pylori is a bacterium that colonizes the harshly acidic milieu of the human stomach. More than half of the world's population carries this infection. Infection rates vary among the developed and developing countries of the world. H. pylori infection is on a steep decline in most of the western countries mainly due to the success of combination therapies and improved personal hygiene and community sanitation to prevent re-infection. However, the situation is not improving in many of the developing countries due to failure of treatment regimes and emergence of drug resistance. The infection in some cases leads to chronic superficial gastritis, chronic active gastritis, peptic ulcer disease and gastric adenocarcinoma [1-4]. One of the most distinctive features of H. pylori is the genetic diversity between clinical isolates obtained from different patient populations. Most H. pylori isolates can be discriminated from others by DNA profiling [5-8] or sequencing of corresponding genes due to mainly a high degree of sequence divergence between orthologs (3–5%) [9,10]. Also, H. pylori has a panmictic or freely recombining population structure [11] and is naturally competent [12]. These characteristics facilitate an inter-strain recombination due mainly to horizontal exchange of alleles from other strains colonising the same niche, which is extremely common in H. pylori chromosome. However, such genetic recombinations in the H. pylori genome might not be deleterious because they occur in the plasticity zone, a special cluster of DNA rearrangements that protects the essential complement of genes by acting as a bed for foreign DNA insertion or abrogation. DNA loss and rearrangement are therefore a norm for H. pylori, and flexibility and diversity in gene content may contribute to bacterial fitness in different members of the diverse human host population. Post genomic analyses have revealed interesting attributes of H. pylori pathogenicity and novel mechanisms of causation of ulcer disease and cancer have been envisaged. Efforts to know the cause and potential benefits of the genetic diversity of this bacterium has led to some interesting discoveries relating to its co-evolution with the human host, microevolution while during infection and quasi-species development, virulence determinants and eradication strategies. Recent studies reviewed herein collectively aim at testing the speculation about whether H. pylori may be beneficial to human health in certain circumstances and whether eradication of this organism is always necessary? Epidemiological studies are needed in the context of such intriguing hypotheses. Results obtained from such studies might enable the development of a high-throughput screening system for high-risk groups within the huge population of H. pylori-infected individuals. Recent studies have shown that H. pylori infection protects against gastro-oesophageal reflux and oesophageal carcinoma. So it will be important to selectively eradicate H. pylori in people that are at the highest risk of developing gastric carcinoma. Eradication of highly pathogenic H. pylori specifically from high-risk groups would markedly reduce the worldwide incidence of ulcer and gastric cancer.
Epidemiology and Evolution
H. pylori infection is usually acquired during childhood, where transmission occurs predominantly within families [13]. A couple of recent studies demonstrated the possible co-existence of a large array of clonal lineages within H. pylori populations that are evolving in each individual separately from one another [14,10]. It is therefore probable that via this semi-vertical transmission of H. pylori strains, there are distinct sets of H. pylori genotypes colonising different human populations. With different strains evolving separately of one another and the fact that H. pylori is a genetically diverse (panmictic) organism, distinct genotypes have been found to be associated with particular geographic regions [15,16]. For example, the shuffling of variant regions within the vacA gene (a gene encoding a vacuolating cytotoxin) within a local H. pylori population has led to predominant vacA genotypes being characteristic of isolates from different geographic regions. In addition, worldwide studies encompassing H. pylori isolates from many geographic regions have demonstrated weak clonal groupings and geographic partitioning of H. pylori isolates [9,17]. If recombination only occurs between a resident H. pylori population, exchange of genetic sequences can genetically homogenise this population. As H. pylori is naturally competent and recombination occurs frequently [11], specific genotypes associated with different geographic regions occur as a result of this homogenising force.
Introduction of polymorphisms and sequence variants from one H. pylori population from a particular geographic region to another H. pylori population from another geographic region via human migration makes the association of particular genotypes with specific geographic locations more difficult. Although the introduction of new polymorphisms into a particular H. pylori population poses a problem with identifying specific genotypes within certain geographic locales, it may, however, provide information on the ancestry of the hosts in whose stomachs the strains were carried. Studies have been aimed at demonstrating the path of human migration to Latin America with conflicting results regarding whether European or Asian populations brought H. pylori to South America [16,11]. However, a recent and comprehensive study by Flaush et al. [19] demonstrated that sequence analysis of H. pylori isolates recovered from twenty-seven countries displayed geographic partitioning. Thus, polymorphisms within the H. pylori genome can serve as useful markers for studying ancient human migrations. However, a mix up of H. pylori strains between migrated and native populations can sometimes complicate analysis. Accordingly, the study of migrated populations that have remained isolated from the native populations is essential.
Genome organization, genetic diversity and microevolution
Since its successful isolation in 1983 by Warren and Marshall, H. pylori has been linked to various pathologies and a strong association with gastric carcinoma and mucosa-associated lymphoid tissue lymphoma [1,3] has been established. However, although H. pylori is definitely responsible for these diseases, only less than 10% of people colonized with H. pylori portray disease symptoms. This suggests that specific H. pylori strains may be responsible for virulence in different hosts. Many studies have shown that certain allotypes of the vacA gene and the presence a functional cagA gene are associated with an increased risk of peptic ulceration and gastric cancer, respectively [20-22]. However, these correlations vary based on the host population studied and efforts to correlate other H. pylori alleles with clinical diseases have failed.
So how do a few H. pylori strains trigger higher virulence as compared to other strains? Current approaches in functional genomics based on protein- protein interaction and microarray based transcription profiling are helping to decode this mystery. Functional genomics often uses the gene chip based expression profiling to provide a condition-dependent and time-specific genome wide profile of an organism's transcriptome [23,24]. Whereas, comparative genomics juxtaposes two or more genome sequences at the level of gene content and organization [25,26]. Both the approaches harness extensive computer algorithms and in silico modelling to summarise gene encoded (or putative) functions. H. pylori has been the first prokaryote wherein full genome sequence of two different patient isolates [J99 and 26995] were characterized and compared [27,28]. As H. pylori is a freely recombining or panmictic organism [11] the question of whether the two genome sequences would accurately represent the myriad of genetic diversity found among the strains was posed. Since the 2 sequenced strains were obtained a decade apart from the two different continents and cultured from the lesions of different gastric disorders, it has been widely assumed that their genome sequences will more likely portray the genetic diversity exhibited by clinical isolates. Comparative genomic analysis of the two completely sequenced strains revealed a significant amount of genetic variation between their genomes. For instance, the J99 genome is shorter (1,643,831 bp) than that of strain 26695 (1,667,867 bp) and has 57 less predicted ORF's [27,28]. Strains 26695 and J99 contain 110 and 52 strain specific genes, respectively [29,30], in which more than a half reside within a locus termed the plasticity cluster. A recent approach helped revised annotation and comparison of the two sequenced H. pylori genomes [31] and reclassified the coding sequences. Based on this study the total number of hypothetical proteins was reduced from 40% to 33%. A large amount of size variation was also discovered between orthologous genes mostly due to natural polymorphisms arising as a result of natural transformation and free recombination within H. pylori chromosome. Recombinational events including the presence of insertion elements, pathogenicity islands, horizontally acquired genes (restriction recombination genes), mosaics and chromosomal rearrangements were frequently annotated in subsequent bioinformatics based attempts. It has been argued that such diversity is a result of a lack of direct competition between strains, even when resident within different individuals within the same community [32]. However, a recent study has demonstrated that integration of foreign gene fragments acquired via natural transformation is often prevented by the well-developed restriction-modification systems in H. pylori genome [33]. It has been demonstrated that H. pylori has extensive, non-randomly distributed repetitive chromosomal sequences, and that recombination between identical repeats contributes to the variation within individual hosts [34]. That H. pylori is representative of prokaryotes, especially those with smaller (<2 megabases) genomes, that have similarly extensive direct repeats, suggests that recombination between such direct DNA repeats is a widely conserved mechanism to promote genome diversification [33]. In addition, although H. pylori has been termed as a panmictic organism [11,35], it is surprising that clonal lineages within H. pylori populations exist [36-39]. Recent reports demonstrated that H. pylori in some populations shows a clonal descent and suggest that a large array of H. pylori clonal lineages co-exist, which evolve in isolation from on another [14]. Moreover, in certain parts of the world H. pylori isolates have been shown to exhibit little genetic heterogeneity, based on fingerprint profiles [17,40]. Functional genomics, utilising microarray technology, has provided researchers with a powerful tool to investigate the genetic diversity of clinical isolates [41], the transcriptional profiles of isolates grown under different conditions [42], the identification of strain-specific and species-specific genes [29,30] and the diversity between strains giving rise to differing clinical illnesses [43]. One of the interesting findings using microarray based genotyping has been the discovery that H. pylori isolates undergo 'microevolution' and give rise to sub-species during prolonged colonisation of a single host [44-46]. The presence of stable sub-species within a single individual suggests an adaptation of a H. pylori population to specific host niches, facilitated by unknown advantages conferred to them by select plasticity region genes. Bjorkholm et al. [45] demonstrated that several loci differed within two genetically related isolates from the same host, one major difference being the presence of the cag pathogenicity island (cag PAI) in one isolate and not the other. As the cagA gene and cag PAI are principle virulence factors within the strains, the excision or abrogation of the cag PAI within a strain may indicate that attenuating the virulence of a strain could be a favourable adaptation.
The conundrum of strain diversity: How many more genome sequences do we need to understand this bug?
Within the bacterial populations, genome content may not be fixed, as changing selective forces favour particular phenotypes; however, organisms well adapted to particular niches may have evolved mechanisms to facilitate such plasticity. The highly diverse H. pylori is a model for studying genome plasticity in the colonization of individual hosts. For H. pylori, neither point mutation, nor intergenic recombination requiring the presence of multiple colonizing strains, is sufficient to fully explain the observed diversity.
The two H. pylori genomes sequenced to date are each from ethnic Europeans, and genomic comparisons modelled on these data are sufficient to identify novel loci from new strains, especially from understudied Asian populations. However, these genome sequences may not be fully representative of the entire diversity of the gene pool. Identification and characterization of such loci which are more abundant in the Asian gene pool may lead to newer insights into the mechanisms of H. pylori colonization, carriage and virulence in the countries of Asia which are more seriously under threat from H. pylori. Therefore, future high throughput efforts involving a large number of strains are clearly needed. Taking the Indian example for instance, according to the Ethnologue database http://www.ethnologue.com, there are about 1683 languages and dialects ('mother tongues') in this country and H. pylori diversity therefore can be assumed to coincide with this figure. So one has to roughly look at the inter-strain genomic diversity contributed by approximately 1683 different strains representing each dialect and or a community.
Nonetheless, genotypic data from each geographic area or a community is extremely vital and might constitute a missing piece of a large, biologic jigsaw puzzle.
Natural competence and transformation
Independent of the other two pathogenesis associated type IV systems, H. pylori harbours a dedicated type IV apparatus, the comB gene cluster [47] linked to the natural transformation and competence. The comB gene cluster is essential for the bacterium to take up plasmid and chromosomal DNA during natural transformation. To identify the genes essential for natural transformation competence in H. pylori, genetic approach of transposon shuttle mutagenesis has been used and the comB locus was located, consisting of orf2 and comB1-comB3 [48,49]. This cluster contains four tandemly arranged genes, ORF2, comB1, comB2 and comB3 as a single transcriptional unit. Subsequently, the components of comB cluster namely Orf2, comB1, comB2 and comB3 were renamed (according to homology with the Agrobacterium tumifaciens type IV secretory apparatus) as comB7, comB8, comB9 and comB10 respectively (Figure 1). Another ORF in HP26695, HP0017 was found to be homologous to the virB4 gene in Agrobacterium tumifaciens type IV secretory apparatus and was named as comB4 [50]. From this study it also appeared that each of the gene products of ORFs comB8 to comB10 were absolutely essential for the development of natural transformation competence. It appears that the comB transformation apparatus has evolved conservatively and is typically present in all the strains. This conservation is interestingly in agreement with the need for genomic fluidity in H. pylori where deletions and rearrangements due to natural transformation and transposition are the norm. This is therefore necessary for the pathogen to keep the gene content flexible and as diverse as possible to probably acclimatise itself to diverse host niches during the process of infection. Both these systems, the cag-PAI encoded type IV export system and the transformation-associated type IV system seem to act completely independently, since the deletion of one system from the chromosome does apparently not affect the function of the other system.
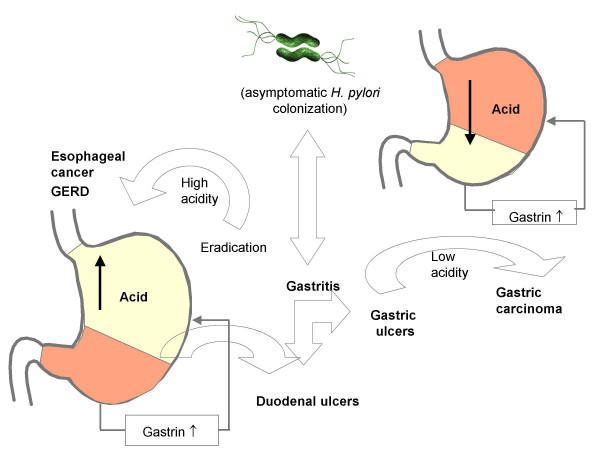
Figure 1.
Different outcomes of H. pylori infection. Some studies argue that eradication of H. pylori might trigger some of the worst forms of heart burns and increased acidity, leading ultimately to oesophageal cancer and or GERD.
Pathogenic apparatuses
The cag pathogenicity island
Molecular analysis of bacterial transport has been attempted in several bacterial pathogens. Among such transport systems, Type IV secretion systems have been described in greater detail in diverse bacteria. In H. pylori, 3 different kinds of type IV secretion apparatuses have been identified. The first such secretion system identified in H. pylori was the one comprised of 29 genes encoding the cag pathogenicity island (cag-PAI). One of the principal virulence factors of H. pylori, the cagA antigen is contained in the 40 kb cag-PAI. The tyrosine-phosphorylated cagA protein is translocated to the epithelial cells by the type IV secretion system (forming a sort of syringe like structure) [51-53]. Upon tyrosine phosphorylation, the cagA protein elicits growth factor like stimuli in epithelial cells (hummingbird phenotype) coupled with interleukin-8 induction for the recruitment of neutrophils. Mutations in several genes of the cag-PAI interfere with tyrosine phosphorylation and induction of interleukin-8 secretion [54]. In recent studies, in order to analyse which genes of the cag-PAI are essential for cagA translocation and/or interleukin 8 induction, a complete mutagenesis of the cag-PAI was performed [55]. In general, it appears that most of the cag genes are involved in assembly and arrangement of the secretory apparatus. Five of these genes namely HP0524 (virD4), HP0525 (virB11), HP0527 (virB10), HP0528 (virB9) and HP0544 (virB4/cagE) constitute the main apparatus of the type IV secretory system of H. pylori [56]. All these genes except HP0524 are associated with IL8 production [55]. However, the presence of strains eliciting IL 8 responses irrespective of intactness of the cag-PAI underlines the fact that it's 'not' the only factor linked to IL8 secretion [57].
Very recently, and for the first time, ultrastructure analysis of the surface of H. pylori 26695 has revealed a sheathed, surface organelle, coded by the cag-PAI genes, HP0527 (forms sheath around the pilus needle) and HP0532/ cagT (forms the base of the pilus) [58]. This structure, although uncommon, could be the special adaptation of H. pylori to the host niches and this might mediate biological as well as transport functions of the cag-PAI encoded proteins. Computational analyses to predict the macromolecular assemblies of such apparatuses are needed to have a more simplified understanding of the entire model of the H. pylori type IV secretion mechanism
Link with cancer: the oncogenic cagA protein
A large-scale prospective study revealed that the risk for development of gastric carcinoma was much greater in the H. pylori-infected population than in the H. pylori-uninfected population [59]. The cagA gene of H. pylori is assumed as partially responsible for eliciting signaling mechanisms that lead to the development of gastric adenocarcinoma. Based on the carriage of a functional cagA as a marker for the cag PAI, the H. pylori species is divided into cagA-positive and cagA-negative strains. The cagA-positive strains are associated with higher grades of gastric or duodenal ulceration and are more virulent than the cagA-negative strains [60]. Some epidemiological studies have demonstrated roles of cagA positive H. pylori in the development of atrophic gastritis, peptic-ulcer disease and gastric carcinoma [61,62]. The cagA gene product, cagA, is translocated to the gastric epithelial cells to undergo tyrosine phosphorylation by SRC family kinases [63]. Tyrosine phosphorylation is known to occur at the EPIYA motifs on the cagA. The cagA protein upon phosphorylation binds and activates a SHP2 phosphatase that acts as a human oncoprotein. As SHP2 transmits positive signals for cell growth and motility, deregulation of SHP2 by cagA is an important mechanism by which cagA-positive H. pylori promotes gastric carcinogenesis. Cag A is noted for its variation at the SHP2 binding site and, based on the sequence variation, it is sub- classified into two main types – East-Asian cagA and Western cagA. East-Asian cagA shows stronger SHP2 binding and greater biological activity than Western cagA. In East-Asian countries, endemic circulation of H. pylori strains that carry biologically active forms of cagA might underlie the high incidence of gastric carcinoma. One puzzling attribute of H. pylori infection is why some populations with high incidences of H. pylori infection, such as those in Japan and Korea, have high incidences of gastric carcinoma, whereas other highly infected populations, such as populations in central Africa, do not. Possible reasons could be the differences in genetic susceptibility among populations, environmental factors such as dietary habits, and strain differences of H. pylori. Among these, diversity of cagA in H. pylori strains might be involved in determination of the type and severity of disease. As discussed above, East-Asian and Western forms of cagA possess the distinctly structured tyrosine phosphorylation/ SHP2-binding sites – EPIYA-D and EPIYA-C, respectively [64]. Notably, the grades of inflammation, activity of gastritis, and atrophy are significantly higher in patients with gastritis who were infected with the East-Asian cagA-positive strain than in patients infected with the cagA-negative or Western cagA-positive strain [65]. Furthermore, the prevalence of the East-Asian cagA-positive strain is associated with the mortality rate of gastric cancer in Asia. Therefore, populations infected with East-Asian cagA positive H. pylori are at greater risk for gastric cancer than those infected with Western cagA-positive strains.
Among Western CagA species, the number of EPIYA-C sites directly correlates with levels of tyrosine phosphorylation, SHP2-binding activity and morphological transformation [64]. Furthermore, molecular epidemiological studies have shown that the number of EPIYA-C sites is associated with the severity of atrophic gastritis and gastric carcinoma in patients infected with Western CagA-positive strains of H. pylori [66].
The number-2 virulence determinant: vacuolating cytotoxin (VacA) of H. pylori
H. pylori has a single copy of the vacA gene. Screening of H. pylori chromosomal fragments permitted the identification of a 3864-base pair open reading frame (vacA) that encoded the vacuolating cytotoxin [67]. The sequence of the vacA gene includes a 33 amino acid signal sequence. With the exception of a hydrophobic region at the N terminus, the mature 90-kDa protein (amino acids 34 to 842) is mainly hydrophilic [67]. The cytotoxic activity of VacA has been shown to increase substantially under acidic conditions. VacA protein, a secreted 95 kD peptide, varies in the signal sequence (alleles s1a, s1b, s1c, s2) and/or its middle region (alleles m1, m2) between different H. pylori strains. The different combinations of s and m regions determine the production of cytotoxic activity. Strains with the genotype s1 m1 produce high levels of vacuolating cytotoxin in vitro. Strains with the genotype s2 produce an inactive toxin. Whereas, strains with the genotype m2 produce toxic activity with a different target cell specificity from those of m1 genotype. Genotypic variations in the vacA gene structure specific to a geographic locale have been recognised. While the vacA m1a allele is specific for the European strains [16], the vacA m1b genotype is typical of the Asian strains [68]. Yet another signal region genotype, s1c is also characteristic of the East Asians [69]. Among other functions, VacA selectively inhibits the invariant chain (Ii)-dependent pathway of antigen presentation mediated by the MHC class II and might induce apoptosis in epithelial cells. VacA, so far mainly regarded as a cytotoxin of the gastric epithelial cell layer, now turns out to be a potent immunomodulatory toxin, targeting the adapted immune system. Thus, in addition to the well-known vacuolating activity, VacA has been reported to induce apoptosis in epithelial cells, to affect B lymphocyte antigen presentation, to inhibit the activation and proliferation of T lymphocytes, and to modulate the T cell-mediated cytokine response.
The plasticity region cluster
The fascinating genomic landmark discovered post genomic era in both the sequenced strains is the one where 48% and 46% of the strain specific genes are located in J99 and 26695 respectively. This region is called as the 'plasticity zone' [28]. Genome sequence comparisons have revealed that nearly half of the strain specific genes fall in this zone. Recently, a new type IV secretion apparatus has been located in this plasticity zone [70]. This type IV cluster is comprised of 7 genes, homologous to the vir B operon of A. tumifaciens carried in a 16.3 kb genomic fragment called tfs3 (Figure 1). This cluster was discovered by Kersulyte et al. as a result of subtractive hybridization and chromosome walking and sequence homology. They also tested conservation of this island in clinical isolates and found that full length and partially disrupted tfs3 occur in 20% and19% of the strains respectively, from Spain, Peru, India and Japan. Although there is no correct role assigned to this cluster, it might be an unusual transposon linked to many deletion events occuring in the plasticity region that contribute to bacterial fitness in diverse host populations via exercising flexibility in gene content and gene order. The plastic nature of H. pylori and the evidence of horizontal transfer of genes from other H. pylori isolates and bacterial species could explain the ability of this organism to persist in a changing environment and why only a subset of clinical isolates exert an adverse effect on patients.
Link with cancer: are plasticity region genes involved?
The plasticity region as a whole displays certain characteristics of pathogenicity islands [71] with relatively low G+C content (35%) compared to the rest of the genome (39%). This region is about 45 kb long in J99 and 68 kb long in strain 26695. Genomic analysis revealed the region to be highly mosaic with a majority of the genes being transcribed suggesting their functional role. They also express protein level homology to various other recombinases, integrases and topoisomerases [72] accounting to natural transformation and recombination. In addition to these, many ORFs were identified as differentially expressed (JHP0927-JHP0928-JHP0931 and JHP-042-JHP0944-JHP0945-JHP0947-JHP0960). They share same chromosomal orientation and therefore they potentially represent a bacterial operon. It is interesting to study the expression or suppression of these ORFs in strains linked to different clinical conditions. Recent studies have posed a possibility to explore the presence of any new pathogenicity markers in the plasticity zone, although the functions of most of the putatively encoded proteins in this cluster are unknown. But they are thought to play a role in increasing the virulence capacity of H. pylori strains either directly or by encoding factors that could lead to variance in the clinical out come of the infection. More interestingly, it is also noted that some of the genes of the plasticity regions were co-inherited along with cagA. However their co-association with the disease status or with the severity of gastric inflammation was not established either due to small sample size or lack of clinical information [73]. Interestingly, a novel pathogenicity marker, JHP947 has been detected within the plasticity zone [74]. Many genes putatively linked to the development of gastric cancer have been assigned to the plasticity zone [72]. Researchers have looked for genetic markers in H pylori strains isolated from patients with gastric extranodal marginal zone B cell lymphoma (MZBL) of the mucosa-associated lymphoid tissue (MALT)-type and strains from age matched patients with gastritis only [75]. Two ORFs were significantly linked with gastric MZBL over gastritis strains: JHP950 (74% v 49%) and JHP1462 (26% v 3%). JHP950 proved specific for gastric MZBL when tested against a group of strains from patients with duodenal ulcer and patients with adenocarcinoma, with significant prevalence (49% and 39%, respectively), and is therefore the candidate marker for gastric MZBL. Interestingly, the candidate ORF JHP950 is located in the plasticity region of the J99 genome [75,76]. In view of such findings it can be speculated that some members of the plasticity region cluster provide selective advantage to some of the strains to adapt to changing host niches and become more and more invasive. In what way such advantage is gained? This needs to be discovered.
Do we need to eradicate H. pylori from this earth?
How long humans carried H. pylori is still controversial. However, it is accepted that this organism has colonized humans possibly for many thousands of years, and the successful persistence of H. pylori in human stomach for such a long period may be because this organism offers some advantages to the host. Unfortunately, the H. pylori infection is on steep decline in the western world. This is mainly due to the success rate of combination therapies and subsequent prevention of re-infection due to improvement in sanitation and personal hygiene. This may seem good news to many gastro-enterologists around the world, but having a H. pylori infection may be advantageous. A study has shown that H. pylori produces a cecropin-like peptide (antibacterial peptide) with high antimicrobial properties [77]. A German study revealed that children infected with H. pylori were less likely to have diarrhoea than children without an infection [78], implying that H. pylori may have beneficial properties to human hosts. Interestingly, there has been a marked decline in the instances of peptic ulcer disease and gastric cancer in the 20th century. Concurrent with this is a dramatic increase in the incidences of gastro-oesophageal reflux disease (GERD), Barrett's oesophagus and adenocarcinoma of the oesophagus in Western countries [79]. This observation led to the speculation that H. pylori may in some way be associated with these diseases and perhaps capable of preventing their onset. Studies have also shown that cagA+ H. pylori strains have a more protective effect than cagA- strains [80]. The presence of cagA+ H. pylori strains can reduce the acidity of the stomach, and it is believed that the raising of the pH by H. pylori prevents GERD, Barrett's oesophagus and adenocarcinoma of the oesophagus (Figure). Conversely, arguments have been made that, although H. pylori may prevent these reflux-associated diseases, the risks of acquiring gastric cancer via H. pylori infection far outweigh any possible benefits it may provide [81]. However, it has been stated that, if H. pylori does provide protection from GERD, the notion of restriction of anti-H. pylori treatment to only a few cases (peptic ulcer disease and MALT lymphoma) could be justified [82]. In spite of this controversy, recent reports have demonstrated a protective role for H. pylori in erosive reflux oesophagitis [83-85]. However, as safe and potent anitsecretory drugs to prevent gastro-oesophageal reflux are available [86] it seems unjustified to use a dangerous organism that has been associated with extremely dangerous outcomes such as a carcinoma.
On the other hand, eradication also is not an ultimate choice. Some ulcers recur even after successful eradication of H. pylori in non-users of non-steroidal anti-inflammatory drug (NSAID). In addition, the incidence of H. pylori-negative, non-NSAID peptic ulcer disease (PUD) (idiopathic PUD) is reported to increase with time. Moreover, it appears that H. pylori-positive ulcers are not always H. pylori-induced ulcers because there are two paradoxes of the H. pylori myth, first the existence of H. pylori-positive non-recurring ulcer and secondly, recurring ulcer after cure of H. pylori infection. To summarise, H. pylori is not the only cause of peptic ulcer disease. Therefore, it is still necessary to seriously consider the need for eradication in all cases of PUD, which may exist even after the elimination of H. pylori.
Conclusion and expert opinion
In our opinion, the intricacies of the role of H. pylori in health and disease may be fully ascertained only if we analyze genetic diversity of the pathogen as juxtaposed to the host diversity and the environment (food and dietary habits). A possible working hypothesis (that we are currently nurturing) may be that among the ocean of molecular host-pathogen interactions that could potentially occur with micro-evolution of this bacterium during long term colonization, some could prove advantageous where the bacterium and the host negotiate nearly a 'symbiotic' and balanced relationship. Such a 'friendship' might have taken thousands of years to develop. If so, why has this bacterium survived for such a long time? Microbes that have long been persisted in humans may be less harmful than recently emerged microbes, such as the human immunodeficiency viruses (HIV). This suggests that the colonization may either be beneficial or of low biological cost to the host. In addition to characterization of bacterial virulence apparatuses that are for sure linked to disease outcome, host responses to such factors must also be examined hand in hand, to completely ascertain mechanisms that lead to gastroduodenal disease. For instance polymorphisms linked to the host immune apparatus, such as IL-1β, TNF-α, and IL-10, which are responsible for elevated proinflammatory potential of the strains. These polymorphisms increase the risk for atrophic gastritis and distal gastric adenocarcinoma among H. pylori-infected persons. Cancer of stomach is a highly lethal disease and establishment of H. pylori as a risk factor for this malignancy deserves an approach to identify persons at increased risk; however, infection with this organism is extremely common and most colonized persons never develop cancer. Thus, screens to identify high-risk subpopulations must use high-resolution biological markers. Fortunately, this task appears to be highly simplified due to the availability of biological tools, which were never thought in the past. Genome sequences (H. pylori, human, C. elegans), quantitative phenotypes (cagA phosphorylation, oipA frame status, vac Aallele status), and practical animal models (Mongolian gerbils) can be harnessed to decipher the molecular basis of H. pylori-associated malignancies, which should have direct clinical applications. It is important to gain more insight into the pathogenesis of H. pylori-induced gastric adenocarcinoma, not only to develop more effective diagnostics and treatment for this common cancer, but also to validate the role of chronic inflammation in the genesis of other tumours of the alimentary tract.
Acknowledgments
Acknowledgement
Authors would like to thank Prof. Seyed E. Hasnain for his guidance, encouragement and for discussions. We are very much thankful to Farhana Kauser for her help in preparation of this manuscript.
Contributor Information
Niyaz Ahmed, Email: niyaz@cdfd.org.in.
Leonardo A Sechi, Email: sechila@uniss.it.
References
- Dunn BE, Cohen H, Blaser MJ. Helicobacter pylori. Clin Microbiol Rev. 1997;10:720–741. doi: 10.1128/cmr.10.4.720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colding H, Hartzen SH, Roshanisefat H, Andersen LP, Krogfelt KA. Molecular methods for typing of Helicobacter pylori and their applications. FEMS Immunol Med Microbiol. 1999;24:193–199. doi: 10.1016/S0928-8244(99)00026-7. [DOI] [PubMed] [Google Scholar]
- Ahmad A, Govil Y, Frank BB. Gastric mucosa-associated lymphoid tissue lymphoma. Am J Gastroenterol. 2003;98:975–986. doi: 10.1016/S0002-9270(03)00142-4. [DOI] [PubMed] [Google Scholar]
- van Belkum A. High-throughput epidemiologic typing in clinical microbiology. Clin Microbiol Infect. 2003;9:86–100. doi: 10.1046/j.1469-0691.2003.00549.x. [DOI] [PubMed] [Google Scholar]
- Akopyanz N, Bukanov NO, Westblom TU, Kresovich S, Berg DE. DNA diversity among clinical isolates of Helicobacter pylori detected by PCR-based RAPD fingerprinting. Nucleic Acids Res. 1992;20:5137–5142. doi: 10.1093/nar/20.19.5137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang JT, Sheu JC, Lin JT, Wang TH, Wu MS. Direct DNA amplification and restriction pattern analysis of Helicobacter pylori in patients with duodenal ulcer and their families. J Infect Dis. 1993;168:1544–1548. doi: 10.1093/infdis/168.6.1544. [DOI] [PubMed] [Google Scholar]
- Marshall DG, Chua A, Keeling PW, Sullivan DJ, Coleman DC, Smyth CJ. Molecular analysis of Helicobacter pylori populations in antral biopsies from individual patients using randomly amplified polymorphic DNA (RAPD) fingerprinting. FEMS Immunol Med Microbiol. 1995;10:317–323. doi: 10.1016/0928-8244(94)00087-A. [DOI] [PubMed] [Google Scholar]
- Marshall DG, Coleman DC, Sullivan DJ, Xia H, O'Morain CA, Smyth CJ. Genomic DNA fingerprinting of clinical isolates of Helicobacter pylori using short oligonucleotide probes containing repetitive sequences. J Appl Bacteriol. 1996;81:509–517. doi: 10.1111/j.1365-2672.1996.tb03540.x. [DOI] [PubMed] [Google Scholar]
- Achtman M, Azuma T, Berg DE, Ito Y, Morelli G, Pan ZJ, Suerbaum S, Thompson SA, van der Ende A, van Doorn LJ. Recombination and clonal groupings within Helicobacter pylori from different geographical regions. Mol Microbiol. 1999;32:459–470. doi: 10.1046/j.1365-2958.1999.01382.x. [DOI] [PubMed] [Google Scholar]
- Owen RJ, Xerry J. Tracing clonality of Helicobacter pylori infecting family members from analysis of DNA sequences of three housekeeping genes (ureI, atpA and ahpC), deduced amino acid sequences, and pathogenicity-associated markers (cagA and vacA) J Med Microbiol. 2003;52:515–524. doi: 10.1099/jmm.0.04988-0. [DOI] [PubMed] [Google Scholar]
- Go MF. Helicobacter pylori: its role in ulcer disease and gastric cancer and how to detect the infection. Acta Gastroenterol Latinoam. 1996;26:45–49. [PubMed] [Google Scholar]
- Israel DA, Lou AS, Blaser MJ. Characteristics of Helicobacter pylori natural transformation. FEMS Microbiol Lett. 2000;186:275–280. doi: 10.1016/S0378-1097(00)00156-7. [DOI] [PubMed] [Google Scholar]
- Drumm B, Perez-Perez GI, Blaser MJ, Sherman PM. Intrafamilial clustering of Helicobacter pylori infection. N Engl J Med. 1990;322:359–363. doi: 10.1056/NEJM199002083220603. [DOI] [PubMed] [Google Scholar]
- Han SR, Zschausch HC, Meyer HG, Schneider T, Loos M, Bhakdi S, Maeurer MJ. Helicobacter pylori: clonal population structure and restricted transmission within families revealed by molecular typing. J Clin Microbiol. 2000;38:3646–3651. doi: 10.1128/jcm.38.10.3646-3651.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Doorn LJ, Figueiredo C, Megraud F, Pena S, Midolo P, Queiroz DM, Carneiro F, Vanderborght B, Pegado MD, Sanna R, De Boer W, Schneeberger PM, Correa P, Ng EK, Atherton J, Blaser MJ, Quint WG. Geographic distribution of vacA allelic types of Helicobacter pylori. Gastroenterology. 1999;116:823–830. doi: 10.1016/s0016-5085(99)70065-x. [DOI] [PubMed] [Google Scholar]
- Kersulyte D, Mukhopadhyay AK, Velapatino B, Su W, Pan Z, Garcia C, Hernandez V, Valdez Y, Mistry RS, Gilman RH, Yuan Y, Gao H, Alarcon T, Lopez-Brea M, Balakrish Nair G, Chowdhury A, Datta S, Shirai M, Nakazawa T, Ally R, Segal I, Wong BC, Lam SK, Olfat FO, Boren T, Engstrand L, Torres O, Schneider R, Thomas JE, Czinn S, Berg DE. Differences in genotypes of Helicobacter pylori from different human populations. J Bacteriol. 2000;182:3210–3218. doi: 10.1128/JB.182.11.3210-3218.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahmed N, Khan AA, Alvi A, Tiwari S, Jyothirmayee CS, Kauser F, Ali M, Habibullah CM. Genomic analysis of Helicobacter pylori from Andhra Pradesh, South India: Molecular evidence for three major genetic clusters. Curr Sci. 2003;85:101–108. [Google Scholar]
- Ghose C, Perez-Perez GI, Dominguez-Bello MG, Pride DT, Bravi CM, Blaser MJ. East Asian genotypes of Helicobacter pylori strains in Amerindians provide evidence for its ancient human carriage. Proc Natl Acad Sci USA. 2002;99:15107–15111. doi: 10.1073/pnas.242574599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falush D, Wirth T, Linz B, Pritchard JK, Stephens M, Kidd M, Blaser MJ, Graham DY, Vacher S, Perez-Perez GI, Yamaoka Y, Megraud F, Otto K, Reichard U, Katzowitsch E, Wang X, Achtman M, Suerbaum S. Traces of human migrations in Helicobacter pylori populations. Science. 2003;299:1582–1585. doi: 10.1126/science.1080857. [DOI] [PubMed] [Google Scholar]
- Atherton JC, Cao P, Peek , Jr, Tummuru MK, Blaser MJ, Cover TL. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. Association of specific vacA types with cytotoxin production and peptic ulceration. J Biol Chem. 1995;270:17771–17777. doi: 10.1074/jbc.270.30.17771. [DOI] [PubMed] [Google Scholar]
- Atherton JC. The clinical relevance of strain types of Helicobacter pylori. Gut. 1997;40:701–703. doi: 10.1136/gut.40.6.701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atherton JC, Peek RM, Jr, Tham KT, Cover TL, Blaser MJ. Clinical and pathological importance of heterogeneity in vacA, the vacuolating cytotoxin gene of Helicobacter pylori. Gastroenterology. 1997;112:92–99. doi: 10.1016/s0016-5085(97)70223-3. [DOI] [PubMed] [Google Scholar]
- Ferea TL, Brown PO. Observing the living genome. Curr Opin Genet Dev. 1999;9:715–722. doi: 10.1016/S0959-437X(99)00033-7. [DOI] [PubMed] [Google Scholar]
- Harrington CA, Rosenow C, Retief J. Monitoring gene expression using DNA microarrays. Curr Opin Microbiol. 2000;3:285–291. doi: 10.1016/S1369-5274(00)00091-6. [DOI] [PubMed] [Google Scholar]
- Ochman H, Jones IB. Evolutionary dynamics of full genome content in Escherichia coli. Embo J. 2000;19:6637–6643. doi: 10.1093/emboj/19.24.6637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ochman H, Moran NA. Genes lost and genes found: evolution of bacterial pathogenesis and symbiosis. Science. 2001;292:1096–1099. doi: 10.1126/science.1058543. [DOI] [PubMed] [Google Scholar]
- Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP, Gill S, Dougherty BA, Nelson K, Quackenbush J, Zhou L, Kirkness EF, Peterson S, Loftus B, Richardson D, Dodson R, Khalak HG, Glodek A, McKenney K, Fitzegerald LM, Lee N, Adams MD, Hickey EK, Berg DE, Gocayne JD, Utterback TR, Peterson JD, Kelley JM, Cotton MD, Weidman JM, Fujii C, Bowman C, Watthey L, Wallin E, Hayes WS, Borodovsky M, Karp PD, Smith HO, Fraser CM, Venter JC. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388:539–547. doi: 10.1038/41483. [DOI] [PubMed] [Google Scholar]
- Alm RA, Ling LS, Moir DT, King BL, Brown ED, Doig PC, Smith DR, Noonan B, Guild BC, deJonge BL, Carmel G, Tummino PJ, Caruso A, Uria-Nickelsen M, Mills DM, Ives C, Gibson R, Merberg D, Mills SD, Jiang Q, Taylor DE, Vovis GF, Trust TJ. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–180. doi: 10.1038/16495. [DOI] [PubMed] [Google Scholar]
- Israel DA, Salama N, Krishna U, Rieger UM, Atherton JC, Falkow S, Peek RM., Jr Helicobacter pylori genetic diversity within the gastric niche of a single human host. Proc Natl Acad Sci USA. 2001;98:14625–14630. doi: 10.1073/pnas.251551698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janssen PJ, Audit B, Ouzounis CA. Strain-specific genes of Helicobacter pylori: distribution, function and dynamics. Nucleic Acid Res. 2001;29:4395–4404. doi: 10.1093/nar/29.21.4395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boneca IG, Reuse H, de Epinat JC, Pupin M, Labigne A, Moszer I. A revised annotation and comparative analysis of Helicobacter pylori genomes. Nucleic Acids Res. 2003;31:1704–1714. doi: 10.1093/nar/gkg250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blaser MJ, Berg DE. Helicobacter pylori genetic diversity and risk of human disease. J Clin Invest. 2001;107:767–773. doi: 10.1172/JCI12672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aras RA, Small AJ, Ando T, Blaser MJ. Helicobacter pylori interstrain restriction-modification diversity prevents genome subversion by chromosomal DNA from competing strains. Nucleic Acids Res. 2002;30:5391–5397. doi: 10.1093/nar/gkf686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aras RA, Kang J, Tschumi AI, Harasaki Y, Blaser MJ. Extensive repetitive DNA facilitates prokaryotic genome plasticity. Proc Natl Acad Sci USA. 2003;100:13579–13584. doi: 10.1073/pnas.1735481100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Go MF, Kapur V, Graham DY, Musser JM. Population genetic analysis of Helicobacter pylori by multilocus enzyme electrophoresis: extensive allelic diversity and recombinational population structure. J Bacteriol. 1996;178:3934–3938. doi: 10.1128/jb.178.13.3934-3938.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell S, Fraser A, Holliss B, Schmid J, O'Toole PW. Evidence for ethnic tropism of Helicobacter pylori. Infect Immun. 1997;65:3708–3712. doi: 10.1128/iai.65.9.3708-3712.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito Y, Azuma T, Ito S, Miyaji H, Hirai M, Yamazaki Y. Analysis and typing of the vacA gene from cagA-positive strains of Helicobacter pylori isolated in Japan. J Clin Microbiol. 1997;35:1710–1714. doi: 10.1128/jcm.35.7.1710-1714.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Ende A, Pan ZJ, Bart A, van der Hulst RW, Feller M, Xiao SD, Tytgat GN, Dankert J. cagA-positive Helicobacter pylori populations in China and The Netherlands are distinct. Infect Immun. 1998;66:1822–1826. doi: 10.1128/iai.66.5.1822-1826.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Doorn LJ, Figueiredo C, Sanna R, Blaser MJ, Quint WG. Distinct variants of Helicobacter pylori cagA are associated with vacA subtypes. J Clin Microbiol. 1999;37:2306–2311. doi: 10.1128/jcm.37.7.2306-2311.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carroll IM, Ahmed N, Beesley SM, Khan AA, Ghousunnissa S, O'Morain CA, Smyth CJ. Fine- structure molecular typing of Irish Helicobacter pylori isolates and their genetic relatedness to strains from four different continents. J Clin Microbiol. 2003;41:5755–5759. doi: 10.1128/JCM.41.12.5755-5759.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salama N, Guillemin K, McDaniel TK, Sherlock G, Tompkins L, Falkow S. A whole-genome microarray reveals genetic diversity among Helicobacter pylori strains. Proc Natl Acad Sci. 2000;97:14668–14673. doi: 10.1073/pnas.97.26.14668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ang S, Lee CZ, Peck K, Sindici M, Matrubutham U, Gleeson MA, Wang JT. Acid-induced gene expression in Helicobacter pylori: study in genomic scale by microarray. Infect Immun. 2001;69:1679–1686. doi: 10.1128/IAI.69.3.1679-1686.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maeda S, Otsuka M, Hirata Y, Mitsuno Y, Yoshida H, Shiratori Y, Masuho Y, Muramatsu M, Seki N, Omata M. cDNA microarray analysis of Helicobacter pylori-mediated alteration of gene expression in gastric cancer cells. Biochem Biophys Res Commun. 2001;284:443–449. doi: 10.1006/bbrc.2001.5006. [DOI] [PubMed] [Google Scholar]
- Covacci A, Rappuoli R. Helicobacter pylori: molecular evolution of a bacterial quasi-species. Curr Opin Microbiol. 1998;1:96–102. doi: 10.1016/S1369-5274(98)80148-3. [DOI] [PubMed] [Google Scholar]
- Bjorkholm B, Lundin A, Sillen A, Guillemin K, Salama N, Rubio C, Gordon JI, Falk P, Engstrand L. Comparison of genetic divergence and fitness between two subclones of Helicobacter pylori. Infect Immun. 2001;69:7832–7838. doi: 10.1128/IAI.69.12.7832-7838.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Israel DA, Salama N, Arnold CN, Moss SF, Ando T, Wirth HP, Tham KT, Camorlinga M, Blaser MJ, Falkow S, Peek RM., Jr Helicobacter pylori strain-specific differences in genetic content, identified by microarray, influence host inflammatory responses. J Clin Invest. 2001;107:611–620. doi: 10.1172/JCI11450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofreuter D, Odenbreit S, Haas R. Natural transformation competence in Helicobacter pylori is mediated by the basic components of a type IV secretion system. Mol Microbiol. 2001;41:379–391. doi: 10.1046/j.1365-2958.2001.02502.x. [DOI] [PubMed] [Google Scholar]
- Odenbreit S, Till M, Haas R. Optimized BlaM-transposon shuttle mutagenesis of Helicobacter pylori allows the identification of novel genetic loci involved in bacterial virulence. Mol Microbiol. 1996;20:361–373. doi: 10.1111/j.1365-2958.1996.tb02623.x. [DOI] [PubMed] [Google Scholar]
- Hofreuter D, Odenbreit S, Henke G, Haas R. Natural competence for DNA transformation in Helicobacter pylori: identification and genetic characterization of the comB locus. Mol Microbiol. 1998;28:1027–1038. doi: 10.1046/j.1365-2958.1998.00879.x. [DOI] [PubMed] [Google Scholar]
- Hofreuter D, Karnholz A, Haas R. Topology and membrane interaction of Helicobacter pylori ComB proteins involved in natural transformation competence. Int J Med Microbiol. 2003;293:153–165. doi: 10.1078/1438-4221-00258. [DOI] [PubMed] [Google Scholar]
- Asahi M, Azuma T, Ito S, Ito Y, Suto H, Nagai Y, Tsubokawa M, Tohyama Y, Maeda S, Omata M, Suzuki T, Sasakawa C. Helicobacter pylori CagA protein can be tyrosine phosphorylated in gastric epithelial cells. J Exp Med. 2000;191:593–602. doi: 10.1084/jem.191.4.593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Odenbreit S, Püls J, Sedlmaier B, Gerland E, Fischer W, Haas R. Translocation of Helicobacter pylori CagA into gastric epithelial cells by type IV secretion. Science. 2000;287:1497–1500. doi: 10.1126/science.287.5457.1497. [DOI] [PubMed] [Google Scholar]
- Stein M, Rappuoli R, Covacci A. Tyrosine phosphorylation of the Helicobacter pylori CagA antigen after cag-driven host cell translocation. Proc Natl Acad Sci. 2000;97:1263–1268. doi: 10.1073/pnas.97.3.1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Censini S, Lange C, Xiang ZY, Crabtree JE, Ghiara P, Borodovsky M. Cag, a pathogenicity island of Helicobacter pylori, encodes type I-specific and disease-associated virulence factors. Proc Natl Acad Sci USA. 1996;93:14648–14653. doi: 10.1073/pnas.93.25.14648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fischer W, Püls J, Buhrdorf R, Gebert B, Odenbreit S, Haas R. Systematic mutagenesis of the Helicobacter pylori cag pathogenicity island: essential genes for CagA translocation in host cells and induction of interleukin-8. Mol Microbiol. 2001;42:1337–1348. doi: 10.1046/j.1365-2958.2001.02714.x. [DOI] [PubMed] [Google Scholar]
- Buhrdorf R, Forster C, Haas R, Fischer W. Topological analysis of a putative virB8 homologue essential for the cag type IV secretion system in Helicobacter pylori. Int J Med Microbiol. 2003;293:213–217. doi: 10.1078/1438-4221-00260. [DOI] [PubMed] [Google Scholar]
- Audibert C, Burucoa C, Janvier B, Fauchere JL. Implication of the structure of the Helicobacter pylori cag pathogenicity island in induction of interleukin-8 secretion. Infect Immun. 2001;69:1625–1629. doi: 10.1128/IAI.69.3.1625-1629.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rohde M, Puls J, Buhrdorf R, Fischer W, Haas R. A novel sheathed surface organelle of the Helicobacter pylori Cag type IV secretion system. Mol Microbiol. 2003;49:219–234. doi: 10.1046/j.1365-2958.2003.03549.x. [DOI] [PubMed] [Google Scholar]
- Uemura N, Okamoto S, Yamamoto S, Matsumura N, Yamaguchi S, Yamakido M, Taniyama K, Sasaki N, Schlemper RJ. Helicobacter pylori infection and the development of gastric cancer. N Engl J Med. 2001;345:784–789. doi: 10.1056/NEJMoa001999. [DOI] [PubMed] [Google Scholar]
- Kuipers EJ, Perez-Perez GI, Meuwissen SG, Blaser MJ. Helicobacter pylori and atrophic gastritis: importance of the cagA status. J Natl Cancer Inst. 1995;87:1777–1780. doi: 10.1093/jnci/87.23.1777. [DOI] [PubMed] [Google Scholar]
- Blaser MJ, Perez-Perez GI, Kleanthous H, Cover TL, Peek RM, Chyou PH, Stemmermann GN, Nomura A. Infection with Helicobacter pylori strains possessing cagA is associated with an increased risk of developing adenocarcinoma of the stomach. Cancer Res. 1995;55:2111–2115. [PubMed] [Google Scholar]
- Parsonnet J, Friedman GD, Orentreich N, Vogelman H. Risk for gastric cancer in people with CagA positive or CagA negative Helicobacter pylori infection. Gut. 1997;40:297–301. doi: 10.1136/gut.40.3.297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatakeyama M. Oncogenic mechanisms of Helicobacter pylori cagA protein. Nat Rev Cancer. 2004;4:688–694. doi: 10.1038/nrc1433. [DOI] [PubMed] [Google Scholar]
- Higashi H, Tsutsumi R, Fujita A, Yamazaki S, Asaka M, Azuma T, Hatakeyama M. Biological activity of the Helicobacter pylori virulence factor CagA is determined by variation in the tyrosine phosphorylation sites. Proc Natl Acad Sci USA. 2002;99:14428–14433. doi: 10.1073/pnas.222375399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azuma T, Yamazaki S, Yamakawa A, Ohtani M, Muramatsu A, Suto H, Ito Y, Dojo M, Yamazaki Y, Kuriyama M, Keida Y, Higashi H, Hatakeyama M. Association between diversity in the Src homology 2 domain-containing tyrosine phosphatase binding site of Helicobacter pylori CagA protein and gastric atrophy and cancer. J Infect Dis. 2004;189:820–827. doi: 10.1086/381782. [DOI] [PubMed] [Google Scholar]
- Yamaoka Y, El-Zimaity HM, Gutierrez O, Figura N, Kim JG, Kodama T, Kashima K, Graham DY. Relationship between the cagA 3' repeat region of Helicobacter pylori, gastric histology, and susceptibility to low pH. Gastroenterol. 1999;117:342–349. doi: 10.1053/gast.1999.0029900342. [DOI] [PubMed] [Google Scholar]
- Cover TL, Tummuru MKM, Cao P, Thompson SA, Blaser MJ. Divergence of genetic sequences for the vacuolating cytotoxin among Helicobacter pylori strains. J Biol Chem. 1994;269:10566–10573. [PubMed] [Google Scholar]
- Pan ZJ, Berg DE, van der Hulst RWM, Su W-W, Raudonikiene A, Xia SD. High prevalence of vacuolating cytotoxin production and unusual distribution of vacA alleles in Helicobacter pylori from China. J Infect Dis. 1998;178:220–226. doi: 10.1086/515601. [DOI] [PubMed] [Google Scholar]
- van Doorn LJ, Figueiredo C, Sanna R, Pena S, Midolo P, Ng EK, Atherton JC, Blaser MJ, Quint WG. Expanding allelic diversity of Helicobacter pylori vacA. J Clin Microbiol. 1998;36:2597–2603. doi: 10.1128/jcm.36.9.2597-2603.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kersulyte D, Velapatino B, Mukhopadhyay AK, Cahuayme L, Bussalleu A, Combe J, Gilman RH, Berg DE. Cluster of type IV secretion genes in Helicobacter pylori 's plasticity zone. J Bacteriol. 2003;185:3764–3772. doi: 10.1128/JB.185.13.3764-3772.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doig P, de Jonge BL, Alm RA, Brown ED, Uria-Nickelsen M, Noonan B, Mills SD, Tummino P, Carmel G, Guild BC, Moir DT, Vovis GF, Trust TJ. Helicobacter pylori physiology predicted from genomic comparison of two strains. Microbiol Mol Biol Rev. 1999;63:675–707. doi: 10.1128/mmbr.63.3.675-707.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Occhialini A, Marais A, Alm R, Garcia F, Sierra R, Megraud F. Links of open reading frames of plasticity region of strain J99 in Helicobacter pylori strains isolated from gastric carcinoma and gastritis patients in Costa Rica. Infect Immun. 2000;68:6240–6249. doi: 10.1128/IAI.68.11.6240-6249.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salama N, Guillemin K, McDaniel TK, Sherlock G, Tompkins L, Falkow S. A whole-genome microarray reveals genetic diversity among Helicobacter pylori strains. Proc Natl Acad Sci USA. 2000;97:14668–14673. doi: 10.1073/pnas.97.26.14668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santos A, Queiroz DM, Menard A, Marais A, Rocha GA, Oliveira CA, Nogueira AM, Uzeda M, Megraud F. New pathogenicity marker found in the plasticity region of the Helicobacter pylori genome. J Clin Microbiol. 2003;41:1651–1655. doi: 10.1128/JCM.41.4.1651-1655.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehours P, Dupouy S, Bergey B, Ruskone-Foumestraux A, Delchier JC, Rad R, Richy F, Tankovic J, Zerbib F, Megraud F, Menard A. Identification of a genetic marker of Helicobacter pylori strains involved in gastric extranodal marginal zone B cell lymphoma of the MALT-type. Gut. 2004;53:931–937. doi: 10.1136/gut.2003.028811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehours P, Menard A, Dupouy S, Bergey B, Richy F, Zerbib F, Ruskone-Fourmestraux A, Delchier JC, Megraud F. Evaluation of the association of nine Helicobacter pylori virulence factors with strains involved in low-grade gastric mucosa-associated lymphoid tissue lymphoma. Infect Immun. 2004;72:880–888. doi: 10.1128/IAI.72.2.880-888.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Putsep K, Branden CI, Boman HG, Normark S. Antibacterial peptide from H. pylori. Nature. 1999;398:671–672. doi: 10.1038/19439. [DOI] [PubMed] [Google Scholar]
- Rothenbacher D, Blaser MJ, Bode G, Brenner H. Inverse relationship between gastric colonization of Helicobacter pylori and diarrheal illnesses in children: results of a population-based cross-sectional study. J Infect Dis. 2000;182:1446–1449. doi: 10.1086/315887. [DOI] [PubMed] [Google Scholar]
- Richter JE, Falk GW, Vaezi MF. Helicobacter pylori and gastroesophageal reflux disease: the bug may not be all bad. Am J Gastroenterol. 1998;93:1800–1802. doi: 10.1111/j.1572-0241.1998.00523.x. [DOI] [PubMed] [Google Scholar]
- Chow WH, Blaser MJ, Blot WJ, Gammon MD, Vaughan TL, Risch HA, Perez-Perez GI, Schoenberg JB, Stanford JL, Rotterdam H, West AB, Fraumeni JF., Jr Inverse relation between cagA+ strains of Helicobacter pylori infection and risk of esophageal and gastric cardia adenocarcinoma. Cancer Res. 1998;58:588–590. [PubMed] [Google Scholar]
- Graham DY. Helicobacter pylori, GERD, NSAIDs, and cancer: where we really stand. Am J Gastroenterol. 1999;94:1420–1421. doi: 10.1111/j.1572-0241.1999.01420.x. [DOI] [PubMed] [Google Scholar]
- Vieth M, Stolte M. possibly protective properties of Helicobacter pylori in connection with GERD. Am J Gastroenterol. 1999;94:3068–3069. doi: 10.1111/j.1572-0241.1999.03068.x. [DOI] [PubMed] [Google Scholar]
- Koike T, Ohara S, Sekine H, Iijima K, Abe Y, Kato K, Toyota T, Shimosegawa T. Helicobacter pylori infection prevents erosive reflux oesophagitis by decreasing gastric acid secretion. Gut. 2001;49:330–334. doi: 10.1136/gut.49.3.330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warburton-Timms VJ, Charlett A, Valori RM, Uff JS, Shepherd NA, Barr H, McNulty CA. The significance of cagA (+) Helicobacter pylori in reflux oesophagitis. Gut. 2001;49:341–346. doi: 10.1136/gut.49.3.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaji Y, Mitsushima T, Ikuma H, Okamoto M, Yoshida H, Kawabe T. Inverse background of Helicobacter pylori antibody and pepsinogen in reflux oesophagitis compared with gastric cancer: analysis of 5732 Japanese subjects. Gut. 2001;49:335–340. doi: 10.1136/gut.49.3.335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham DY. The changing epidemiology of GERD: geography and Helicobacter pylori. Am J Gastroenterol. 2003;98:1462–1470. doi: 10.1016/S0002-9270(03)00362-9. [DOI] [PubMed] [Google Scholar]