Abstract
Background
Aedes (Stegomyia) albopictus (Skuse) is an indigenous species and the predominant vector of dengue fever in China. Understanding of genetic diversity and structure of the mosquito would facilitate dengue prevention and vector control. Sympatric cryptic species have been identified in the Ae. albopictus subgroup in Southeast Asia; however, little is known about the presence and distribution of cryptic species in China. This study aimed to examine the genetic diversity, evaluate potential new cryptic sibling species, and assess the prevalence of Wolbachia infections in field populations.
Methods
Aedes adult female specimens were collected from five provinces in southern and central China during 2015–2016. Morphological identification was performed under dissection microscope. The mitochondrial DNA cytochrome c oxidase subunit 1 (cox1, DNA barcoding) locus and the ribosomal DNA internal transcribed spacer region 2 (ITS2) marker were used to examine the genetic variation, evaluate cryptic sibling species, and population structure in the field populations. Screening for the presence of Wolbachia was performed using multiplex PCR.
Results
A total of 140 individual specimens with morphological characteristics similar to Ae. albopictus were sequenced for DNA barcoding. Among these, 129 specimens (92.1%) were confirmed and identified as Ae. albopictus. The remaining 11 specimens, from 2 provinces, were identified as 2 distinct sequence groups, which were confirmed by ITS2 marker sequencing, suggesting the existence of potential cryptic species of Ae. albopictus. In Ae. albopictus, we found significant genetic differentiation and population structure between populations collected from different climate zones. Medium to high frequencies of Wolbachia infections were observed in natural Ae. albopictus populations, whereas Wolbachia was infrequent or absent in cryptic species populations.
Conclusions
Our findings highlight the population differentiation by climate zone and the presence of novel, cryptic Aedes species in China. The low prevalence of Wolbachia infections in cryptic species populations could reflect either a recent invasion of Wolbachia in Ae. albopictus or different host immune responses to this symbiont in the cryptic species. The study provides useful information for vector control and host-symbiont coevolution. Further study is needed to investigate the potential for arbovirus infection and disease transmission in the emerged cryptic species.
Electronic supplementary material
The online version of this article (10.1186/s13071-018-2814-8) contains supplementary material, which is available to authorized users.
Keywords: Aedes albopictus, Sympatric cryptic species, Wolbachia endosymbiont, Mitochondrial DNA, Genetic diversity, Population structure
Background
Over the past decades, more than 11 sibling species or cryptic species have been identified and characterized in the Aedes albopictus subgroup of the Scutellaris group in the subgenus Stegomyia of Aedes [1–3]. Of these species, Ae. albopictus, originating from Asia, is the most widely distributed and has invaded on every continent except Antarctica [4, 5]. Aedes albopictus is considered a medically important species and is a major vector of several human arboviruses, including dengue, Zika, chikungunya, yellow fever and West Nile viruses [5–10]. Dengue fever has experienced a 30-fold increase in incidence worldwide over the past 50 years and shows no signs of slowing down [11]. Since the 1970s there have been several major outbreaks of dengue fever in southern China, including in Hainan, Guangxi, Fujian, Zhejiang, Yunnan and Guangdong provinces [12–14]. Due to climate change, the transmission of the dengue virus has spread gradually from southern tropical or subtropical regions to the surrounding northern and western regions, and even to the central China Henan Province with a generally warm temperate continental climate [15]. The most recent outbreak of dengue fever occurred in 2014 in Guangdong Province, with a total of 45,224 dengue fever cases and 6 deaths [12, 16, 17]. Aedes albopictus mosquitoes are regarded as the sole vector for dengue transmission in nearly all these epidemics [14, 18].
In China, Ae. albopictus is an indigenous species, closely associated with human migration, transportation, commerce and urbanization. It is the most important dengue vector species and has different susceptibilities to dengue virus in different geographical areas [19, 20]. Due to the lack of effective treatments or vaccines for dengue fever, vector control through chemical or biological measures targeting mosquitoes or their breeding sites is essential for dengue prevention. With the progressive spread of insecticide resistance, the threat of Ae. albopictus is growing, and development of efficient surveillance methods is more urgent than ever before [21, 22]. Population genetic studies of arthropod disease vectors can provide information about the transmission dynamics of specific pathogens, which aids in the design of strategies for controlling vector-borne disease epidemics [23, 24]. The recent waves of dengue outbreak in China highlight the need to improve our knowledge of Ae. albopictus population distribution and dynamics. Although scientists have studied the diversity of the cox1 gene in Ae. albopictus in several localities in China [9, 25–27], there have been no systematic studies of the genetic diversity of Ae. albopictus field populations and its cryptic species.
Cryptic species are defined as sibling species of two or more morphologically indistinguishable biological groups that are closely related and live in the same habitat [28]. Cryptic species may be medically important in vector-borne disease transmission, vector ecology and evolutionary biology. A number of new cryptic species have been identified in mosquito genera (Diptera: Culicidae), including Culex [29–31], Anopheles [32–39] and Aedes [40, 41]. In Ae. albopictus, a novel cryptic species has been reported in Vietnam [40], and the divergence between the cryptic species and Ae. albopictus was confirmed by analysis of nuclear ribosomal genes and mitochondrial genes. However, there are no reports of cryptic species of Ae. albopictus in other Asian countries, including China.
Natural infections of Wolbachia microbiota are common in Ae. albopictus, and the two Wolbachia biotypes, wAlbA and wAlbB, co-occur at a rate near 100% in many areas [42–46]. Maternally inherited Wolbachia bacteria can cause cytoplasmic incompatibility (CI) in many insect species, including Ae. albopictus mosquitoes [47, 48]. Wolbachia mediates antiviral protection of Aedes mosquitoes against a broad range of RNA viruses, including dengue, yellow fever, chikungunya and Rift Valley fever virus [49]. Understanding the distribution and prevalence of Wolbachia in Ae. albopictus and its cryptic species will provide useful information for vector control and host-symbiont coevolution.
In this study, we investigated the genetic diversity and population structure of Ae. albopictus from different climate regions in China, uncovered and molecularly identified the cryptic Aedes species and its polymorphism, and detected Wolbachia infection in the natural Aedes populations. The prevalence of Wolbachia endosymbiont was evaluated by multiplex PCR genotyping and DNA sequencing of individuals in the natural Aedes populations.
Methods
Sample collection
Adult Aedes mosquito specimens were collected from April 2015 to October 2016 using BG-sentinel traps (Bioquip Products, Inc. California, USA) or electric aspirator mosquito catches [50] at 14 collection sites in five provinces in China: Henan, Guangdong, Guangxi, Yunnan and Hainan (Fig. 1). These sites are highly diverse in environmental conditions and most of them have experienced dengue epidemics in the past. The sampling site in Henan Province, located in central China, has a temperate climate with a distinct seasonality characterized by hot, humid summers and generally cold, windy and dry winters. The sampling sites in Guangdong and Guangxi provinces have a subtropical monsoon climate with long summers and year-round abundant precipitation. The sampling sites in Hainan and Yunnan provinces are tropical areas with a wet climate. Hainan Province had dengue epidemics in the late 1970s and early 1980s, however, no dengue epidemic has been reported from there since 1990. Guangdong Province has experienced multiple major dengue and chikungunya epidemics since 1980 and dengue has remained every year since 1994. In 2013, Guangdong, Yunnan and Henan provinces had dengue outbreaks. Dengue transmission occurred in southern China from July to November and the peak season is usually September and October whereas dengue epidemic in Henan Province is limited in summer from July to September, and showed earlier peaks and shorter epidemic periods [15]. Aedes albopictus mosquitoes are the primary vector of dengue virus across China, especially in urban areas. Aedes aegypti mosquitoes are only found in a small portion of southern China, including Hainan Province and small portions of Yunnan Province and southern tip of Guangdong Province [51]. Ten mosquito specimens from each collection site were used in the study. All mosquito specimens were morphologically identified under a stereomicroscope (Nikon) using morphological keys as described by Lu et al. [52]. All mosquito samples were stored at -20 °C prior to DNA extraction.
Fig. 1.
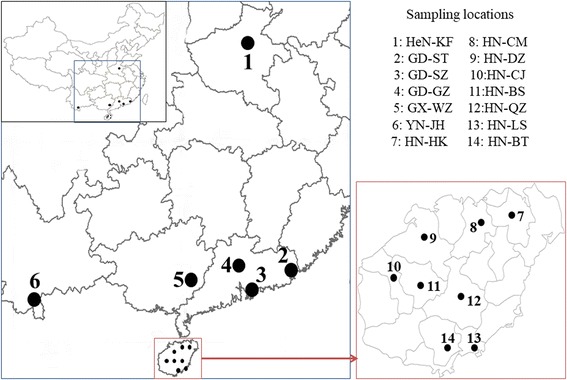
Locations of the 14 sampling sites. Site 1: Kaifeng (HeN-KF, 34°47'53" N, 114°18'05"E) in Henan Province. Sites 2–4: Shantou (GD-ST, 23°21'22"N, 116°40'40"E), Shenzhen (GD-SZ, 22°32'11"N, 113°55'32"E), and Guangzhou (GD-GZ, 23°07'54"N, 113°15'33"E) in Guangdong Province. Site 5: Wuzhou (GX-WZ, 23°53'43"N, 110°32'54"E) in Guangxi Province. Site 6: Jinghong (YN-JH, 22°00'10"N, 100°46'14"E) in Yunnan Province. Sites 7–14: Haikou (HN-HK, 20°02'47"N, 110°11'44"E), Chengmai (HN-CM, 19°44'25"N, 110°00'02"E), Danzhou (HN-DZ, 19°31'23"N, 109°34'36"E), Changjiang (HN-CJ, 19°17'60"N, 109°03'05"E), Baisha (HN-BS, 19°13'37"N, 109°26'51"E), Qiongzhong (HN-QZ, 19°02'06"N, 109°50'03"E), Lingshui (HN-LS, 18°30'27"N, 110°01'59"E) and Baoting (HN-BT, 18°38'27"N, 109°41'54"E) in Hainan Province. The map was created using the R package ‘maptools’, version: 0.9–2, URL: http://r-forge.r-project.org/projects/maptools/
PCR amplification and sequencing of mitochondrial DNA (mtDNA)
Total DNA was extracted from individual adult mosquitoes using the Insect DNA Kit (OMEGA Bio-Tek, D0926-01, Guangzhou, China) according to the manufacturer’s standard protocol. Extracted DNA was preserved at -20 °C until molecular analysis. The mitochondrial gene cytochrome c oxidase subunit 1 (cox1) was used to examine sequence polymorphism among mosquito samples. PCR was performed to amplify a 651 bp fragment of the 5' cox1 region of mtDNA using the DNA primer pairs LCOI490 (5'-GGT CAA CAA ATC ATA AAG ATA TTG G-3') and HCO2198 (5'-TAA ACT TCA GGG TGA CCA AAA AAT CA-3') [53, 54]. PCR amplification was performed in a 25 μl reaction volume with 12.5 μl GoTaq Green Master Mix (Promega, Guangzhou, China), 1 μl each of the forward and reverse primers at 10 μmol/l, 2 μl of template DNA and sufficient nuclease-free water to make 25 μl. PCR conditions were as follows: an initial denaturation at 94 °C for 1 min followed by five cycles of 94 °C for 40 s (denaturation), 45 °C for 40 s (annealing), and 72 °C for 1 min (extension); 30 cycles of 94 °C for 40 s (denaturation), 53 °C for 40 s (annealing), and 72 °C for 1 min (extension); and a final extension at 72 °C for 5 min. The amplified fragments were run on a 1% agarose gel to check integrity, stained with ethidium bromide and analyzed under UV light. PCR products were purified using a gel extraction kit (OMEGA Bio-Tek, D2500-02) and sequenced with PCR primers in both directions using the ABI 3730XL automatic sequencer (Applied Biosystems, Guangzhou, China). The sequences of cox1 unique haplotypes were deposited to the GenBank database under the accession numbers KY765450-KY765506.
PCR amplification and sequencing of ribosomal DNA (rDNA)
The internal transcribed spacer 2 (ITS2) region of ribosomal DNA was amplified from the DNA samples using the universal primers ITS2A (5'-ATC ACT CGG CTC GTG GAT CG-3') and ITS2B (5'-ATG CTT AAA TTT AGG GGG TAG TC-3'), which anneal to highly conserved sequences in the 5.8S and 28S rDNA genes flanking the entire ITS2 region [21, 55]. PCR amplification was performed in a 25 μl reaction volume with 12.5 μl GoTaq Green Master Mix (Promega, Guangzhou, China), 1 μl each of the forward and reverse primers at 10 μmol/l, 2 μl of template DNA (1~2 ng/μl), and sufficient nuclease-free water to make 25 μl. PCR conditions were as follows: an initial denaturation at 94 °C for 3 min followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min; and a final extension at 72 °C for 5 min. Purification and sequencing of the PCR products were the same as described above for the cox1 gene. The sequences of ITS2 unique haplotypes were submitted to the GenBank database under the accession numbers MF623839-MF623851.
PCR detection of Wolbachia infection in mosquitoes
The Wolbachia infection status of individual mosquitoes was determined by PCR amplification of Wolbachia ribosomal DNA using primers specific for Wolbachia 16S rDNA (WF: 5'-CAT ACC TAT TCG AAG GGA TAG-3' and WR: 5'-AGC TTC GAG TGA AAC CAA TTC-3') [56]. To further classify infected mosquitoes by Wolbachia group, we amplified the Wolbachia surface protein gene (wsp) using wAlbA primers (328F: 5'-CCA GCA GAT ACT ATT GCG-3' and 691R: 5'-AAA AAT TAA ACG CTA CTC CA-3') for A group and wAlbB primers (183F: 5'-AAG GAA CCG AAG TTC ATG-3' and 691R: 5'-AAA AAT TAA ACG CTA CTC CA-3') for B group [57]. PCR amplification was performed in a 25 μl reaction volume with 12.5 μl GoTaq Green Master Mix (Promega, Guangzhou, China), 1 μl each of the forward and reverse primers at 10 μmol/l, 2 μl of template DNA, and sufficient nuclease-free water to make 25 μl. PCR conditions were as follows: an initial denaturation at 94 °C for 3 min followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min; and a final extension at 72 °C for 5 min. PCR-amplified fragments of 408 bp, 364 bp, and 509 bp for 16S rDNA, wAlbA and wAlbB, respectively, were revealed under UV light after electrophoresis on 1% agarose gel. Negative and positive controls for the PCR assay were included in each run. To obtain the positive control, we sequenced PCR fragments from the 16S rDNA and wsp genes and confirmed that the amplified PCR product was Wolbachia by using BLAST search to compare it with existing sequences in the NCBI database.
Data analysis
The cox1 gene sequences from 140 mosquitoes were aligned using Clustal W multiple alignment in BioEdit (version 7.2.6.1) [58]. The number of segregating sites, haplotype diversity (Hd), and nucleotide diversity (π) within each population were determined using DnaSP version 5 [59]. Pairwise sequence divergences were calculated using a Kimura 2-parameter (K2P) distance model in MEGA 7.0.20 [60]. The K2P model was used to make our results comparable with most other studies on mosquito DNA barcoding. To examine population expansion, we also performed neutrality tests for each population. Deviations from selective neutrality were tested using Fu’s Fs statistic [61] and Tajima’s D [62]. To determine the genealogical relationships among haplotypes, we constructed a haplotype network using a statistical parsimony algorithm implemented in TCS version 1.21 [63]. The minimum number of mutational steps between sequences was calculated with > 95% confidence. A haplotype network shows the haplotype frequencies in each population and their relatedness, which is useful in inferring the plausible geographical origin of a population [64]. Genetic differentiation among populations was estimated using Arlequin 3.5 [65]. Analysis of molecular variance (AMOVA) was conducted to determine the distribution of genetic variation within and among populations and among groups (tropical, subtropical, and temperate zone).
To examine the evolutionary relationships between the individuals of Ae. albopictus and its cryptic species, we performed phylogenetic analysis using sequences of the mtDNA cox1 gene and the rDNA ITS2 region. Sequences from different species of the family Culicidae were selected from GenBank and used to build the phylogenetic trees. The cox1 phylogenetic tree was built using sequences of Ae. aegypti (AF390098, AY056597 and AF425846), Cx. tritaeniorhynchus (KT851544), Cx. pipiens (KT851543), Cx. tarsalis (AF425847), Ae. flavopictus (KT358463 and LC054359) and Ae. albopictus and its cryptic species (KF406577, JQ728019, KY378914, KF406649, KX495909, KX495922, KX495910, JQ728198, KY378918 and KY378935). The ITS2 phylogenetic tree was built using sequences of Ae. aegypti (GU980956, M95126 and KF471584), Ae. flavopictus (AF353532 and AF353548), Cx. pipiens (U22131 and U33044), Cx. quinquefasciatus (GU562872) and Ae. albopictus and its cryptic species (KU497617, DQ168420, KX495928, AF305554, KY382426, KX495936, KX495942 and KX495949). The predominant haplotype sequences of Ae. albopictus (cox1: KY765450, KY765468 and KY765479; ITS2: MF623839, MF623840 and MF623841) and all the haplotypes of cryptic species identified in the study, together with above GenBank sequences, were aligned using MAFFT v7.31 [66]. Phylogenetic trees were constructed based on the aligned nucleotide sequences using the neighbor-joining method via the maximum composite likelihood substitution model in MEGA 7.0.20 [60]. The statistical significance of tree branching was tested by performing 1000 bootstrap replications [67].
Results
Genetic polymorphism of Ae. albopictus and its cryptic species
PCR amplification and sequencing of the mitochondrial cox1 gene resulted in a 651 bp fragment for each individual study subject, with no insertions or deletions. We compared the cox1 sequences with the existing sequences in the NCBI database by BLAST search. Of the 140 individuals, 129 sequences (92.1%) were identical or possessed > 98% similarity with Ae. albopictus (GenBank: KR068634) (Additional file 1: Table S1). The remaining 11 individuals (7.9%) were approximately 10% K2P divergent from Ae. albopictus, indicating the existence of cryptic species (namely, Aedes sp.) of Ae. albopictus in China (Additional file 2: Table S2). Of these 11 individuals, 9 were from samples collected in Wuzhou, Guangxi Province, with one each from Baisha and Baoting, Hainan Province. Since there is only one individual of Ae. albopictus in GX-WZ population and one each of Aedes sp. from HN-BS, and HN-BT populations, which is insufficient for population genetic analysis, these individuals were excluded from the analysis of the population genetic structure and genetic diversity. Thus, only 9 individuals from each of these three populations (GX-WZ, HN-BS and HN-BT) were included in the genetic polymorphism analysis (Table 1). A high level of genetic diversity (S = 33; Hd = 0.972; π = 1.327; k = 8.639) was found in the Wuzhou (GX-WZ) population (cryptic species) compared with the 13 Ae. albopictus populations. Varied levels of genetic diversity were identified among the Ae. albopictus populations (Table 1). The Haikou (HN-HK) population had the highest number of polymorphism sites (S = 18), the greatest haplotype diversity (Hd = 0.978), and the highest average number of nucleotide differences (k = 4.333), followed by the Kaifeng (HeN-KF) population (k = 3.356). The three populations from Guangdong Province (GD-SZ, GD-ST and GD-GZ) had relatively low genetic diversity compared with populations from the other provinces. Varied genetic diversity was also found in the 8 populations from Hainan Province, with nucleotide diversity (π) ranging from 0.123 in specimens from Qiongzhong to 0.666 in specimens from Haikou. Tajima’s D tests for all study populations were not statistically significant (Table 1), indicating that the populations are in genetic equilibrium, consistent with the neutral mutation hypothesis. Likewise, Fu’s Fs test was not statistically significant and rejected the population expansion/bottleneck model for all study localities, with the exception of one Yunnan population (YN-JH, Fs = -4.738, P < 0.01) and two Hainan populations (HN-HK, Fs = -4.086, P < 0.05; HN-BT, Fs = -4.034, P < 0.05) (Table 1).
Table 1.
Genetic polymorphism and neutrality tests of Aedes albopictus and its cryptic species in China
| Province | Locality | Name | Species | n | S | h | H d | π | k | Tajima’s D | Fu's Fs |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Henan | Kaifeng | HeN-KF | Ae. albopictus | 10 | 15 | 5 | 0.756 | 0.515 | 3.356 | -1.691 | 0.584 |
| Guangdong | Shantou | GD-ST | Ae. albopictus | 10 | 5 | 5 | 0.667 | 0.178 | 1.156 | -1.388 | -1.896 |
| Shenzhen | GD-SZ | Ae. albopictus | 10 | 2 | 2 | 0.2 | 0.061 | 0.4 | -1.401 | 0.586 | |
| Guangzhou | GD-GZ | Ae. albopictus | 10 | 3 | 4 | 0.644 | 0.116 | 0.756 | -1.034 | -1.466 | |
| Guangxi | Wuzhou | GX-WZ | Cryptic Aedes sp. | 9 | 33 | 8 | 0.972 | 1.327 | 8.639 | -1.559 | -1.358 |
| Ae. albopictus | 1 | – | – | – | – | – | – | – | |||
| Yunnan | Jinghong | YN-JH | Ae. albopictus | 10 | 8 | 8 | 0.933 | 0.335 | 2.178 | -0.992 | -4.738** |
| Hainan | Haikou | HN-HK | Ae. albopictus | 10 | 18 | 9 | 0.978 | 0.666 | 4.333 | -1.489 | -4.086* |
| Chengmai | HN-CM | Ae. albopictus | 10 | 5 | 5 | 0.8 | 0.205 | 1.333 | -0.985 | -1.547 | |
| Danzhou | HN-DZ | Ae. albopictus | 10 | 5 | 5 | 0.867 | 0.294 | 1.911 | 0.326 | -0.706 | |
| Changjiang | HN-CJ | Ae. albopictus | 10 | 10 | 7 | 0.867 | 0.447 | 2.911 | -0.782 | -2.134 | |
| Baisha | HN-BS | Ae. albopictus | 9 | 4 | 5 | 0.833 | 0.179 | 1.166 | -0.843 | -2.109 | |
| Cryptic Aedes sp. | 1 | – | – | – | – | – | – | – | |||
| Qiongzhong | HN-QZ | Ae. albopictus | 10 | 4 | 3 | 0.378 | 0.123 | 0.8 | -1.667 | 0.058 | |
| Lingshui | HN-LS | Ae. albopictus | 10 | 4 | 5 | 0.8 | 0.164 | 1.067 | -0.943 | -2.096 | |
| Baoting | HN-BT | Ae. albopictus | 9 | 12 | 8 | 0.972 | 0.503 | 3.277 | -1.221 | -4.034* | |
| Cryptic Aedes sp. | 1 | – | – | – | – | – | – | – |
Abbreviations: n, number of samples; S, number of segregating sites; h, number of haplotypes; Hd, haplotype diversity; π, nucleotide diversity (× 102, average number of nucleotide differences per site); k, average number of nucleotide differences
**P < 0.01
*0.01 < P < 0.05
A total of 57 haplotypes of mtDNA cox1 were detected in the 140 specimens, including 47 haplotypes derived from 129 Ae. albopictus mosquitoes and 10 haplotypes derived from cryptic species (Additional file 3: Table S3, GenBank: KY765450–KY765506). Three predominant haplotypes were identified in Ae. albopictus populations: H01 (21.5%) from Guangdong Province, H19 (22.1%) from Yunnan and Hainan provinces, and H30 (10%) from Hainan Province. Other haplotypes were either unique to a specific population or had a limited geographical distribution (Additional file 3: Table S3). To determine the relationships among the samples, we constructed a median-joining network using haplotypes based on sequence variation. Haplotypes were connected when the probability of parsimony was at least 0.95. Two networks were constructed based on all the 57 haplotypes, one based on haplotypes from Ae. albopictus (Fig. 2a-c) and the other based on haplotypes from the cryptic species (Fig. 2d). Three haplotypes could not be connected to the networks at a 95% confidence level: H28 for Ae. albopictus, and H16 and H44 for the cryptic species. The Ae. albopictus haplotypes can be classified into three clusters corresponding to three climate zones (tropical, subtropical and temperate) as shown in Fig. 2a-c. Haplotype H01 from Guangdong Province (subtropical) was connected to haplotypes from Hainan Province (tropical) through haplotype H30 by one mutation step at nucleotide position 342 (T-342-C), and to haplotypes from Henan Province (temperate) through H26 by one mutation step at nucleotide position 624 (G-624-A). Similar patterns were also observed for haplotype H19 from Yunnan and Hainan provinces (tropical zones). These results may imply a multiple origin for Ae. albopictus populations in China. In the cryptic species, haplotype H17 had at least 4 connections with other haplotypes, suggesting it as a potential ancestral haplotype (Fig. 2d).
Fig. 2.
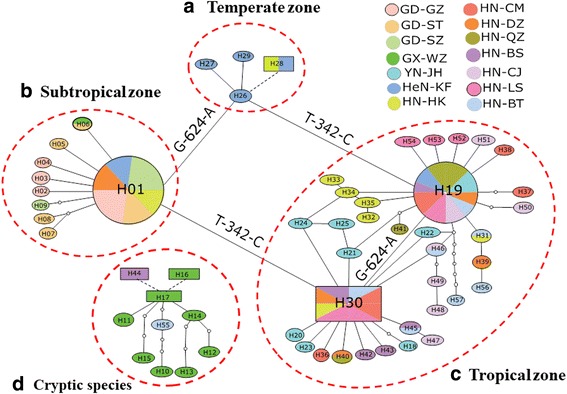
Cox1 haplotype networks showing the genealogical relationships. a-c Aedes albopictus. d Cryptic Aedes species. Each haplotype is represented by a pie chart with size proportional to its frequency in each population. A black dotted line indicates that a mutation step could not be determined between haplotypes at probability of parsimony above the 0.95 limit
A phylogenetic tree based on cox1 sequence variation indicated three clades, which were assigned to three different species (Fig. 3). The first clade (solid blue circles in Fig. 3) corresponds to Ae. albopictus from China and Vietnam, the second clade (red squares) corresponds to cryptic Aedes species previously identified in Vietnam, and the third clade (purple diamond) corresponds to cryptic Aedes species previously identified in Pakistan. The haplotype Aedes sp. CH-H44 (BS07) (purple diamond) is a different cryptic species compared to the haplotypes from Wuzhou, Guangxi Province (red squares). A similar pattern was observed in the phylogenetic tree based on ITS2 sequence variation (Fig. 4). The first clade (solid blue circles in Fig. 4) corresponds to Ae. albopictus from China and Vietnam and the second clade (red squares) corresponds to cryptic Aedes species previously identified in Vietnam. The ITS2 haplotype of Aedes sp. CH-H44 (BS07) (purple diamond) is also in different clade compared to the haplotypes from Wuzhou, Guangxi Province (red squares). The two taxa were distinguished using PCR length polymorphism at the ITS2 locus. Aedes albopictus had an amplicon size of ~580 bp, whereas the cryptic species had an amplicon size of ~415 bp, allowing for easy and accurate identification of the cryptic species through 1% agarose gel electrophoresis.
Fig. 3.
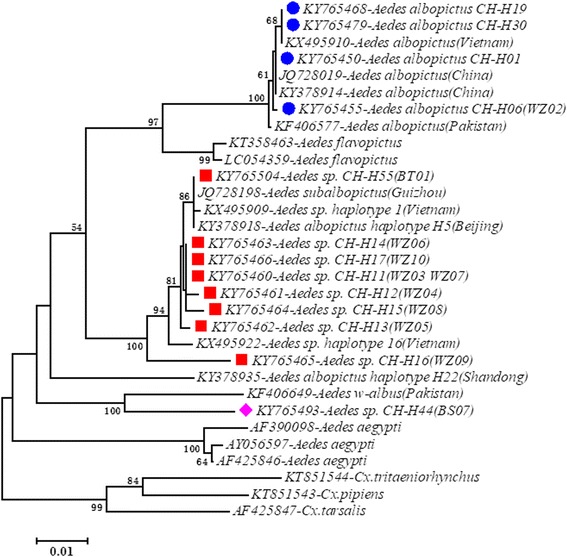

Phylogenetic analysis based on cox1 haplotype variation. Accession numbers of haplotypes marked with color symbols were identified in the current study; others were retrieved from GenBank. Haplotypes marked with a solid blue circle are associated with Ae. albopictus. Haplotypes marked with a red square or a purple diamond are associated with cryptic species of the Ae. albopictus subgroup. Neighbor-joining trees were constructed via the maximum composite likelihood substitution model using MEGA (version 7.0). Numbers at branches represent bootstrap values of 1000 replicates (values > 50 are shown). The scale-bar shows the number of nucleotide substitutions per site
Fig. 4.
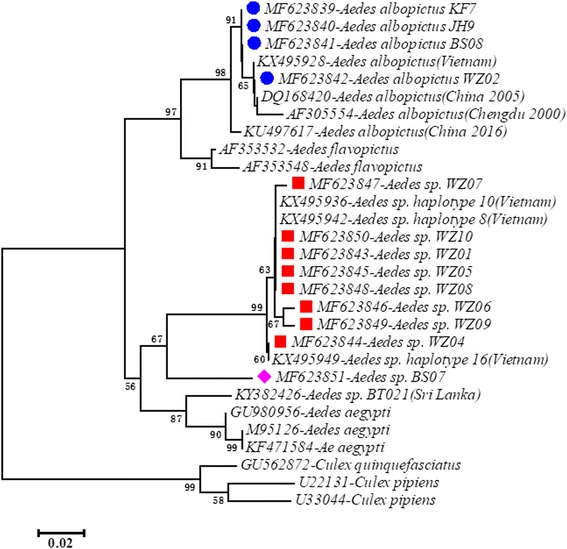

Phylogenetic analysis based on ITS2 haplotype variation. Accession numbers of haplotypes marked with color symbols were identified in the current study; others were retrieved from GenBank. Haplotypes marked with a blue circle are associated with Ae. albopictus. Haplotypes marked with a red square or a purple diamond are associated with cryptic species of the Ae. albopictus subgroup. Neighbor-joining trees were constructed via the maximum composite likelihood substitution model using MEGA (version 7.0). Numbers at branches represent bootstrap values of 1000 replicates (values > 50 are shown). The scale bar shows the number of nucleotide substitutions per site
Genetic differentiation of Ae. albopictus populations and cryptic population
Among the 13 Ae. albopictus populations, genetic differentiation was observed between populations from Henan Province (HeN-KF) and Guangdong Province (GD-ST, GD-SZ and GD-GZ) and populations from Yunnan and Hainan provinces. Thirty-four of 36 pairwise tests were significant at P < 0.05, and pairwise FST values ranged from 0.224 (between HeN-KF and HN-BT) to 0.750 (between GZ-SZ and HN-QZ), with an average of 0.411 (Table 2). No genetic differentiation was observed between populations within Hainan Province or between populations from Yunnan and Hainan provinces, indicating strong gene flow between these populations. All pairwise FST values of differentiation between cryptic population (GX-WZ) and the other 13 Ae. albopictus populations were highly significant and generally very high (FST values > 0.9) (Table 2). To further examine population structure and the extent of genetic variation between Ae. albopictus populations from tropical zones (Yunnan and Hainan provinces) and those from subtropical and temperate zones (Guangdong and Henan provinces), we conducted analysis of molecular variation (AMOVA) between the two groups of populations. Our results indicated a significant overall population structure in Ae. albopictus (FST = 0.38, P < 0.001). The majority of genetic variation (61.77%) was within populations, whereas approximately 34.21% was between the two groups, and only 4.01% of variation was among populations within groups (Table 3).
Table 2.
Pairwise genetic differentiation (FST) between Ae. albopictus populations and the cryptic population in China
| HeN-KF | GD-ST | GD-SZ | GD-GZ | YN-JH | HN-HK | HN-CM | HN-DZ | HN-CJ | HN-BS | HN-QZ | HN-LS | HN-BT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GD-ST | 0.091 | ||||||||||||
| GD-SZ | 0.106 | 0.028 | |||||||||||
| GD-GZ | 0.098 | 0.044 | 0.037 | ||||||||||
| YN-JH | 0.260* | 0.405* | 0.463* | 0.436* | |||||||||
| HN-HK | 0.118 | 0.327* | 0.360* | 0.344* | 0.059 | ||||||||
| HN-CM | 0.285* | 0.521* | 0.606* | 0.557* | 0.046 | 0.030 | |||||||
| HN-DZ | 0.139 | 0.290* | 0.358* | 0.333* | 0.054 | 0.042 | 0.057 | ||||||
| HN-CJ | 0.236* | 0.419* | 0.466* | 0.444* | 0.071 | 0.026 | 0.008 | 0.080 | |||||
| HN-BS | 0.292* | 0.488* | 0.588* | 0.539* | -0.007 | 0.082 | 0.012 | 0.035 | 0.044 | ||||
| HN-QZ | 0.347* | 0.651* | 0.750* | 0.701* | 0.182 | 0.028 | 0.048 | 0.183 | 0.081 | 0.227 | |||
| HN-LS | 0.309* | 0.573* | 0.667* | 0.620* | 0.089 | 0.029 | -0.034 | 0.092 | 0.035 | 0.070 | 0.007 | ||
| HN-BT | 0.224* | 0.348* | 0.397* | 0.377* | 0.004 | 0.055 | 0.036 | -0.003 | 0.010 | -0.011 | 0.152 | 0.067 | |
| GX-WZa | 0.905* | 0.925* | 0.931* | 0.928* | 0.917* | 0.898* | 0.923* | 0.918* | 0.911* | 0.922* | 0.927* | 0.926* | 0.905* |
aAedes sp. cryptic population
*Asterisks indicate significant values after Bonferroni correction (P < 0.05)
Table 3.
Analysis of molecular variance (AMOVA) of two groups of populations of Ae. albopictus in China
| Source of variation | df | SS | Variance components | Percentage of variation |
|---|---|---|---|---|
| Among groups | 1 | 30.34 | 0.5234 Va | 34.21 |
| Among populations within groups | 11 | 17.04 | 0.0614 Vb | 4.01 |
| Within population | 115 | 108.68 | 0.9450 Vc | 61.77 |
| Total | 127 | 156.06 | 1.5299 | – |
Abbreviations: df degrees of freedom, SS sum of squares
PCR detection of Wolbachia infection in natural mosquito populations
Wolbachia infections were detected in all 14 of the Ae. albopictus populations. Infection rates ranged from 50% (HN-DZ) to 100% (in 8 populations) with an average of ~90% (Table 4), suggesting that Wolbachia is highly prevalent in Ae. albopictus in China. In the cryptic Aedes sp. species, however, Wolbachia infection was absent (HN-BT and HN-BS) or occurred at low frequency (11%, GX-WZ) (Fig. 5a-d). Most infected individuals were infected with both the wAlbA and wAlbB strains of Wolbachia; the average superinfection rate in the 14 Ae. albopictus populations was 70%, with a range from 10% (HN-DZ) to 90% (YN-JH, HN-CJ and HN-QZ). Single-strain Wolbachia infections were found in five populations for wAlbA and eight populations for wAlbB, with low prevalence (< 20%) (Fig. 5e, f) except in HN-CM (70% wAlbA) (Table 4).
Table 4.
Results of PCR screening for Wolbachia infection in natural Ae. albopictus populations and cryptic Aedes species in China
| Province | Locality | Name | Species | Total female | 16S rRNA n (%) | wAlbA n (%) | wAlbB n (%) | Type A and type B n (%) | Uninfected n (%) |
|---|---|---|---|---|---|---|---|---|---|
| Henan | Kaifeng | HeN-KF | Ae. albopictus | 10 | 9 (90.0) | 0 (0) | 2 (20.0) | 7 (70.0) | 1 (10.0) |
| Guangdong | Shantou | GD-ST | Ae. albopictus | 10 | 10 (100) | 0 (0) | 2 (20.0) | 8 (80.0) | 0 (0) |
| Shenzhen | GD-SZ | Ae. albopictus | 10 | 10 (100) | 1 (10.0) | 1 (10.0) | 8 (80.0) | 0 (0) | |
| Guangzhou | GD-GZ | Ae. albopictus | 10 | 10 (100) | 0 (0) | 2 (20.0) | 8 (80.0) | 0 (0) | |
| Guangxi | Wuzhou | GX-WZ | Cryptic Aedes sp. | 9 | 1 (11.1) | 0 (0) | 0 (0) | 1 (11.1) | 8 (88.9) |
| Ae. albopictus | 1 | 1 (100) | 0 (0) | 0 (0) | 1 (100) | 0 (0) | |||
| Yunnan | Jinghong | YN-JH | Ae. albopictus | 10 | 10 (100) | 0 (0) | 1 (10.0) | 9 (90.0) | 0 (0) |
| Hainan | Haikou | HN-HK | Ae. albopictus | 10 | 9 (90.0) | 0 (0) | 2 (20.0) | 7 (70.0) | 1 (10.0) |
| Chengmai | HN-CM | Ae. albopictus | 10 | 10 (100) | 7 (70.0) | 0 (0) | 3 (30.0) | 0 (0) | |
| Danzhou | HN-DZ | Ae. albopictus | 10 | 5 (50.0) | 2 (20.0) | 2 (20.0) | 1 (10.0) | 5 (50.0) | |
| Changjiang | HN-CJ | Ae. albopictus | 10 | 10 (100) | 0 (0) | 1 (10.0) | 9 (90.0) | 0 (0) | |
| Baisha | HN-BS | Ae. albopictus | 9 | 9 (100) | 1 (11.1) | 0 (0) | 8 (88.9) | 0 (0) | |
| Cryptic Aedes sp. | 1 | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | |||
| Qiongzhong | HN-QZ | Ae. albopictus | 10 | 10 (100) | 1 (10.0) | 0 (0) | 9 (90.0) | 0 (0) | |
| Lingshui | HN-LS | Ae. albopictus | 10 | 6 (60.0) | 0 (0) | 0 (0) | 6 (60.0) | 4 (40.0) | |
| Baoting | HN-BT | Ae. albopictus | 9 | 7 (77.8) | 0 (0) | 0 (0) | 7 (77.8) | 2 (22.2) | |
| Cryptic Aedes sp. | 1 | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
Fig. 5.
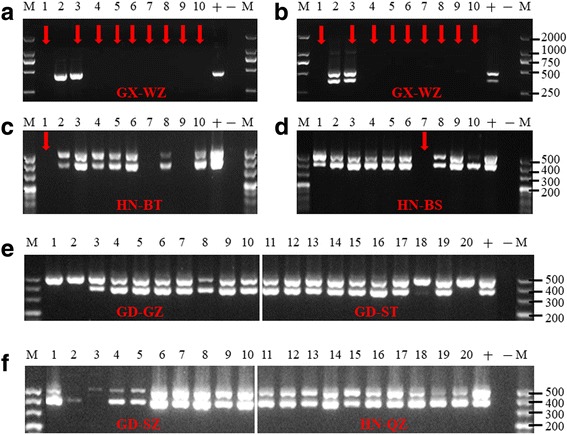
Example of banding patterns of agarose gel electrophoresis. PCR products amplified using primers for Wolbachia-specific 16S rRNA gene (a) and Wolbachia surface protein gene (wsp) (b-f) in natural populations. Lanes 1–10: PCR products of 10 individuals from specific locations; Lanes “+” and “–”: positive and negative controls, respectively; Lane M: DNA ladder. Red arrows indicate cryptic Aedes species
Discussion
Aedes albopictus, one of the cryptic species of the Ae. albopictus subgroup, is an important vector for public health. It is highly invasive and is the most widely distributed mosquito species in the world. Aedes albopictus originated at the edges of forests and bred in natural habitats, but it has adapted to being a domestic mosquito [13]. Today, the species can be found throughout tropical, subtropical and temperate zones in China. Cryptic species often occur in sympatry and are so similar they cannot be distinguished via traditional species identification using morphological keys [68, 69]. With the advent of molecular diagnosis technology and relatively inexpensive DNA sequencing technology, the discovery of cryptic species has become common in many insect groups [70]. However, little is known about the cryptic species within the Ae. albopictus subgroup in China. To the best of our knowledge, this is the first report describing the discovery of phylogenetically divergent cryptic species living in sympatry with Ae. albopictus in China. In this study, among 14 study populations collected across the tropical, subtropical and temperate climate zones of China, we found three populations in southern China in which cryptic species of Ae. albopictus coexisted in sympatry. Furthermore, multiple haplotypes of cryptic species were present in the Wuzhou population in Guangxi Province within the subtropical zone, and probably a novel cryptic species (KY765493) existed in the Baisha population in the tropical area of Hainan Province. Further investigation is needed to confirm the reproductive isolation of these species. These results may have important implications for vector control and for understanding the evolutionary processes of the species. For example, if the cryptic species is a novel disease vector with different biting or resting behaviors, current vector control interventions that target on species-specific vector behavior could lead directly to programme failure and thus these vector control strategies need to be adjusted.
The distribution of Aedes mosquito species is influenced by climatic, environmental and geographical factors, as well as by human behavior [71–73]. In this study, we found significantly higher genetic diversity in cryptic species populations than in Ae. albopictus populations. For example, the cryptic species population in Wuzhou (GX-WZ) has 33 segregation sites, with much higher nucleotide diversity and the highest average number of nucleotide differences compared with the Ae. albopictus populations. The high genetic diversity of cryptic species, as well as their coexistence with Ae. albopictus, may be explained by environmental heterogeneity in these densely forested, mountainous areas with a low level of gene flow and random genetic drift. The different population structure between tropical (Hainan and Yunnan) and subtropical (Guangdong)/temperate (Henan) populations of Ae. albopictus might be due to selective pressures exerted by specific climate, environment and human activities. The mountainous areas of Yunnan and central Hainan provinces have lower human population density and more complex tropical environments than the densely populated Guangdong (coastal) and Henan (plain) provinces. The association between genetic population structure and climate/environment has also been observed in other Ae. albopictus populations [19, 74, 75]. Gene flow between the two genetic clusters (tropical and subtropical/temperate) appeared to be restricted due to geographical isolation, which was also evidenced in the cox1 haplotype network analysis. Interestingly, both of the major tropical-zone haplotypes, H30 and H19, were just one mutation step (C-342-T) from subtropical (Guangdong Province) and temperate (Henan Province) haplotypes, and the same mutation step (G-624-A) was found between populations within tropical zones as well as between populations from subtropical and temperate zones. These results suggest that climate, geography, environment and human activity all play important roles in Ae. albopictus population structure.
The maternally transmitted endosymbiotic bacterium Wolbachia is known to have an important impact on host reproduction in many insects, including mosquitoes (Diptera: Culicidae) [47, 76, 77]. Wolbachia can inhibit human pathogens transmitted by mosquitoes, including dengue virus [78–80], yellow fever [81], filarial nematodes [82, 83], malaria parasites [84–86] and Zika virus [87]. Our results indicate significant variations in the frequency of Wolbachia infection in Ae. albopictus populations. Populations from Guangdong, Yunnan, and many sites in Hainan Province had a 100% infection rate, followed by a 90% infection rate in Henan Province in central China. Such high prevalence of Wolbachia has also been observed in other Asian countries, including Malaysia [43, 88], Thailand [89], India [90] and Sri Lanka [91]. In our study, most populations were naturally infected with two Wolbachia strains (wAlbA and wAlbB) at high frequencies, suggesting that superinfection is common in Ae. albopictus, as observed in other studies [44, 88, 89, 92]. Interestingly, a relatively low prevalence of Wolbachia was also observed in three Ae. albopictus populations (HN-DZ, HN-LS, and HN-BT) from Hainan Province, in which no more than 50% of the individuals were not detected with Wolbachia infection, indicating that there was variation in natural Wolbachia infection in these Ae. albopictus populations. Furthermore, no Wolbachia infection was detected in the Aedes cryptic species from Hainan Province, and Wolbachia infection was detected in just one individual of the cryptic species from Guangxi Province. A similar pattern was observed in the cryptic species in Vietnam [40], suggesting that the cryptic species may be resistant to Wolbachia infection or that Wolbachia are present at low cell density in the cryptic species that cannot be detected by PCR [93]. In addition, cryptic species may prevent Wolbachia introgression by reproductive isolation and maintaining ancestral levels of mitochondrial diversity. Wolbachia induced cytoplasmic incompatibility and mitochondrial selective sweep have been observed in the Ae. albopictus and other mosquito species [29, 48]. Further studies are needed to confirm the reproductive isolation between Ae. albopictus and its cryptic species.
Conclusions
Our results indicated that the genetic diversity and population structure of Ae. albopictus between tropical, subtropical and temperate zones in China appeared to be separated by a single mutation step at the mitochondrial DNA barcoding cox1 gene. Sympatric, cryptic sibling species might be common in the Ae. albopictus subgroup in China. The prevalence of high-level Wolbachia infection in most of the Ae. albopictus populations, and the absence or low prevalence of Wolbachia in the sympatric cryptic species, possibly due to Wolbachia-induced genetic hitchhiking or selective sweep that has created a barrier to gene flow among the species. Elucidating the mechanisms of the observed absence or low prevalence of Wolbachia in sympatric cryptic species may provide insight toward the development of new vector control strategies. Finally, this study will have important implications for disease vector-based control programs, Wolbachia-based disease control strategies, and host evolutionary biology. Further study is needed to investigate the potential for arbovirus infection and disease transmission in the emerged cryptic species.
Additional files
Table S1. K2P divergence of cox1 sequences in Ae. albopictus from China. (XLSX 76 kb)
Table S2. K2P divergence of cox1 sequences in Aedes sp. identified in China. (XLSX 9 kb)
Table S3. cox1 haplotype distribution in the 14 mosquito populations from China. (XLSX 14 kb)
Acknowledgments
We thank Chenglong Li, Defang Huo, Bing Liang, Jiu Zheng, Chunfu Yin, Rui Hong, Yishuang Fu, Xiuxian Tan, Haiyan Chen, Mingjie Yang and Xianzhao Ke for help in collecting mosquito samples in Hainan Province. We are also grateful to Yunyan Guo, Kanglian Zheng, Zhendong Wu and Junhao Jiang for help in collecting mosquito samples from other provinces.
Funding
This work was supported by National Natural Science Foundation of China (No. 31630011), Science and Technology Plan Project of Guangdong Province (No. 2013B021800042) and Natural Science Foundation of Guangdong Province (No. 2015A030313784). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Availability of data and materials
The data supporting the findings of this article are included within the article. Nucleotide sequence data reported in this paper have been deposited to the NBCI GenBank database under the accession numbers: KY765450-KY765506 and MF623839-MF623851.
Abbreviations
- AMOVA
Analysis of molecular variance
- cox1
cytochrome c oxidase subunit 1
- GD-GZ
Guangzhou
- GD-ST
Shantou
- GD-SZ
Shenzhen
- GX-WZ
Wuzhou
- HeN-KF
Kaifeng
- HN-BS
Baisha
- HN-BT
Baoting
- HN-CJ
Changjiang
- HN-CM
Chengmai
- HN-DZ
Danzhou
- HN-HK
Haikou
- HN-LS
Lingshui
- HN-QZ
Qiongzhong
- ITS2
DNA internal transcribed spacer region 2
- K2P
Kimura 2-parameter
- mtDNA
mitochondrial DNA
- rDNA
ribosomal DNA
- wsp
Wolbachia surface protein gene
- YN-JH
Jinghong
Authors’ contributions
All authors have contributed significantly to this study. Conceived and designed the experiments: XZ, GZ and DZ. Performed the experiments: YG, ZS, LL, QW and DY. Analysed the data: YG, GZ and DZ. Wrote and revised the manuscript: YG, XZ, GZ, and DZ. All authors read and approved the final manuscript.
Ethics approval and consent to participate
No specific permits were required for the field studies. For mosquito collection on private lands or in private residential areas, oral consent was obtained from field owners in each location. No sites were protected by law and this study did not involve endangered or protected species.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Footnotes
Electronic supplementary material
The online version of this article (10.1186/s13071-018-2814-8) contains supplementary material, which is available to authorized users.
Contributor Information
Yuyan Guo, Email: guoyuyan66721@163.com.
Zhangyao Song, Email: 805786385@qq.com.
Lei Luo, Email: 724468786@qq.com.
Qingmin Wang, Email: 839579497@qq.com.
Guofa Zhou, Email: zhoug@uci.edu.
Dizi Yang, Email: 1519903906@qq.com.
Daibin Zhong, Email: dzhong@uci.edu.
Xueli Zheng, Email: zhengxueli2001@126.com.
References
- 1.Rai KS. Genetics of Aedes albopictus. J Am Mosq Control Assoc. 1986;2:429–436. [PubMed] [Google Scholar]
- 2.Huang YM. Contributions to the mosquito fauna of Southeast Asia. XIV. The subgenus Stegomyia of Aedes in Southeast Asia. I - The scutellaris group of species. Contrib. Am Entomol Inst. 1972;9:1–109. [Google Scholar]
- 3.McLain DK, Rai KS, Fraser MJ. Intraspecific and interspecific variation in the sequence and abundance of highly repeated DNA among mosquitoes of the Aedes albopictus subgroup. Heredity. 1987;58:373–381. doi: 10.1038/hdy.1987.65. [DOI] [PubMed] [Google Scholar]
- 4.Kraemer MU, Sinka ME, Duda KA, Mylne AQ, Shearer FM, Barker CM, et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. eLife. 2015;4:e08347. doi: 10.7554/eLife.08347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Enserink MA. A mosquito goes global. Science. 2008;320(5878):864–6. [DOI] [PubMed]
- 6.Enserink M. An obscure mosquito-borne disease goes global. Science. 2015;350(6264):1012–1013. doi: 10.1126/science.350.6264.1012. [DOI] [PubMed] [Google Scholar]
- 7.Amraoui F, Vazeille M, Failloux AB. French Aedes albopictus are able to transmit yellow fever virus. Euro Surveill. 2016;21(39). 10.2807/1560-7917.ES.2016.21.39.30361. [DOI] [PMC free article] [PubMed]
- 8.Ngoagouni C, Kamgang B, Nakouné E, Paupy C, Kazanji M. Invasion of Aedes albopictus (Diptera: Culicidae) into central Africa: what consequences for emerging diseases? Parasit Vectors. 2015;8:191. doi: 10.1186/s13071-015-0808-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhong D, Lo E, Hu R, Metzger ME, Cummings R, Bonizzoni M, et al. Genetic analysis of invasive Aedes albopictus populations in Los Angeles County, California and its potential public health impact. PLoS One. 2013;8:e68586. doi: 10.1371/journal.pone.0068586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Battaglia V, Gabrieli P, Brandini S, Capodiferro MR, Javier PA, Chen X-G, et al. The worldwide spread of the tiger mosquito as revealed by mitogenome haplogroup diversity. Front Genet. 2016;7:208. doi: 10.3389/fgene.2016.00208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Achee NL, Gould F, Perkins TA, Reiner RC, Morrison AC, Ritchie SA, et al. A critical assessment of vector control for dengue prevention. PLoS Negl Trop Dis. 2015;9:e0003655. doi: 10.1371/journal.pntd.0003655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xiao J-P, He J-F, Deng A-P, Lin H-L, Song T, Peng Z-Q, et al. Characterizing a large outbreak of dengue fever in Guangdong Province, China. Infect Dis Poverty. 2016;5:44. doi: 10.1186/s40249-016-0131-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wu J-Y, Lun Z-R, James AA, Chen X-G. Dengue fever in Mainland China. Am J Trop Med Hyg. 2010;83:664–671. doi: 10.4269/ajtmh.2010.09-0755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li M-T, Sun G-Q, Yakob L, Zhu H-P, Jin Z, Zhang W-Y. The driving force for 2014 dengue outbreak in Guangdong, China. PLoS One. 2016;11:e0166211. doi: 10.1371/journal.pone.0166211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lai S, Huang Z, Zhou H, Anders KL, Perkins TA, Yin W, et al. The changing epidemiology of dengue in China, 1990–2014: a descriptive analysis of 25 years of nationwide surveillance data. BMC Med. 2015;13:100. doi: 10.1186/s12916-015-0336-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhu G, Liu J, Tan Q, Shi B. Inferring the spatio-temporal patterns of dengue transmission from surveillance data in Guangzhou, China. PLoS Negl Trop Dis. 2016;10:e0004633. doi: 10.1371/journal.pntd.0004633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lin H, Liu T, Song T, Lin L, Xiao J, Lin J, et al. Community involvement in dengue outbreak control: an integrated rigorous intervention strategy. PLoS Negl Trop Dis. 2016;10:e0004919. [DOI] [PMC free article] [PubMed]
- 18.Jin LQ, Li D. A recent survey of mosquito fauna in Guangdong Province, southern China, with a review of past records [corrected] Med Vet Entomol. 2008;22:359–363. doi: 10.1111/j.1365-2915.2008.00758.x. [DOI] [PubMed] [Google Scholar]
- 19.Paupy C, Girod R, Salvan M, Rodhain F, Failloux A-B. Population structure of Aedes albopictus from La Reunion Island (Indian Ocean) with respect to susceptibility to a dengue virus. Heredity. 2001;87:273–283. doi: 10.1046/j.1365-2540.2001.00866.x. [DOI] [PubMed] [Google Scholar]
- 20.Gubler DJ, Rosen L. Variation among geographic strains of Aedes albopictus in susceptibility to infection with dengue viruses. Am J Trop Med Hyg. 1976;25:318–325. doi: 10.4269/ajtmh.1976.25.318. [DOI] [PubMed] [Google Scholar]
- 21.Manni M, Gomulski LM, Aketarawong N, Tait G, Scolari F, Somboon P, et al. Molecular markers for analyses of intraspecific genetic diversity in the Asian tiger mosquito, Aedes albopictus. Parasit Vectors. 2015;8:188. doi: 10.1186/s13071-015-0794-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li Y, Xu J, Zhong D, Zhang H, Yang W, Zhou G, et al. Evidence for multiple-insecticide resistance in urban Aedes albopictus populations in southern China. Parasit Vectors. 2018;11:4. doi: 10.1186/s13071-017-2581-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ferreira-de-Lima VH, Lima-Camara TN. Natural vertical transmission of dengue virus in Aedes aegypti and Aedes albopictus: a systematic review. Parasit Vectors. 2018;11:77. doi: 10.1186/s13071-018-2643-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Goubert C, Minard G, Vieira C, Boulesteix M. Population genetics of the Asian tiger mosquito Aedes albopictus, an invasive vector of human diseases. Heredity. 2016;117:125–134. doi: 10.1038/hdy.2016.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Guo S, Feng L, Wand J-N, Wu Y-Y, Juan H, Gong Z-Y. Genetic polymorphism analysis of cytochrome c oxidase subunit I gene in Aedes albopictus from Zhejiang Province, China. Chin J Zoonoses. 2016;32:133–6.
- 26.Wang G, Li C, Guo X, Xing D, Dong Y, Wang Z, et al. Identifying the main mosquito species in China based on DNA barcoding. PLoS One. 2012;7:e47051. doi: 10.1371/journal.pone.0047051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ruiling Z, Peien L, Xuejun W, Zhong Z. Molecular analysis and genetic diversity of Aedes albopictus (Diptera, Culicidae) from China. Mitochondrial DNA A DNA Mapp Seq Anal. 2017. 10.1080/24701394.2017.1325481. [DOI] [PubMed]
- 28.Bickford D, Lohman DJ, Sodhi NS, Ng PK, Meier R, Winker K, et al. Cryptic species as a window on diversity and conservation. Trends Ecol Evol. 2007;22(3):148–155. doi: 10.1016/j.tree.2006.11.004. [DOI] [PubMed] [Google Scholar]
- 29.Dumas E, Atyame CM, Malcolm CA, Le Goff G, Unal S, Makoundou P, et al. Molecular data reveal a cryptic species within the Culex pipiens mosquito complex. Insect Mol Biol. 2016;25:800–809. doi: 10.1111/imb.12264. [DOI] [PubMed] [Google Scholar]
- 30.Laurito M, Ayala AM, Almirón WR, Gardenal CN. Molecular identification of two Culex (Culex) species of the Neotropical region (Diptera: Culicidae). PLoS One. 2017;12:e0173052. [DOI] [PMC free article] [PubMed]
- 31.Cook S, Moureau G, Harbach RE, Mukwaya L, Goodger K, Ssenfuka F, et al. Isolation of a novel species of flavivirus and a new strain of Culex flavivirus (Flaviviridae) from a natural mosquito population in Uganda. J Gen Virol. 2009;90:2669–2678. doi: 10.1099/vir.0.014183-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Müller P, Pflüger V, Wittwer M, Ziegler D, Chandre F, Simard F, et al. Identification of cryptic Anopheles mosquito species by molecular protein profiling. PLoS One. 2013;8:e57486. doi: 10.1371/journal.pone.0057486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lehr MA, Kilpatrick CW, Wilkerson RC, Conn JE. Cryptic species in the Anopheles (Nyssorhynchus) albitarsis (Diptera: Culicidae) complex: Incongruence between random amplified polymorphic DNA-polymerase chain reaction identification and analysis of mitochondrial DNA COI gene sequences. Ann Entomol Soc Am. 2005;98:908–917. doi: 10.1603/0013-8746(2005)098[0908:CSITAN]2.0.CO;2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lobo NF, Laurent BS, Sikaala CH, Hamainza B, Chanda J, Chinula D, et al. Unexpected diversity of Anopheles species in Eastern Zambia: implications for evaluating vector behavior and interventions using molecular tools. Sci Rep. 2015;5:17952. doi: 10.1038/srep17952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Paredes-Esquivel C, Donnelly MJ, Harbach RE, Townson H. A molecular phylogeny of mosquitoes in the Anopheles barbirostris subgroup reveals cryptic species: implications for identification of disease vectors. Mol Phylogenet Evol. 2009;50:141–151. doi: 10.1016/j.ympev.2008.10.011. [DOI] [PubMed] [Google Scholar]
- 36.Roux O, Diabaté A, Simard F. Larvae of cryptic species of Anopheles gambiae respond differently to cues of predation risk. Freshw Biol. 2013;58:1178–1189. doi: 10.1111/fwb.12117. [DOI] [Google Scholar]
- 37.St. Laurent B, Cooke M, Krishnankutty SM, Asih P, Mueller JD, Kahindi S, et al. Molecular characterization reveals diverse and unknown malaria vectors in the western Kenyan highlands. Am J Trop Med Hyg. 2016;94:327–335. doi: 10.4269/ajtmh.15-0562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Conn JE. News from Africa: Novel anopheline species transmit Plasmodium in western Kenya. Am J Trop Med Hyg. 2016;94(2):251. doi: 10.4269/ajtmh.16-0020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Fouet C, Kamdem C, Gamez S, White BJ. Genomic insights into adaptive divergence and speciation among malaria vectors of the Anopheles nili group. Evol Appl. 2017;10:897–906. doi: 10.1111/eva.12492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Minard G, Tran Van V, Tran FH, Melaun C, Klimpel S, Koch LK, et al. Identification of sympatric cryptic species of Aedes albopictus subgroup in Vietnam: new perspectives in phylosymbiosis of insect vector. Parasit Vectors. 2017;10:276. doi: 10.1186/s13071-017-2202-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ashfaq M, Hebert PDN, Mirza JH, Khan AM, Zafar Y, Mirza MS. Analyzing mosquito (Diptera: Culicidae) diversity in Pakistan by DNA barcoding. PLoS One. 2014;9:e97268. doi: 10.1371/journal.pone.0097268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Raquin V, Valiente Moro C, Saucereau Y, Tran F-H, Potier P, Mavingui P. Native Wolbachia from Aedes albopictus blocks chikungunya virus infection in cellulo. PLoS One. 2015;10:e0125066. [DOI] [PMC free article] [PubMed]
- 43.Ahmad NA, Vythilingam I, Lim YAL, Zabari NZAM, Lee HL. Detection of Wolbachia in Aedes albopictus and their effects on chikungunya virus. Am J Trop Med Hyg. 2017;96:148–156. doi: 10.4269/ajtmh.16-0516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kittayapong P, Baisley KJ, Sharpe RG, Baimai V, O'Neill SL. Maternal transmission efficiency of Wolbachia superinfections in Aedes albopictus populations in Thailand. Am J Trop Med Hyg. 2002;66:103–107. doi: 10.4269/ajtmh.2002.66.103. [DOI] [PubMed] [Google Scholar]
- 45.ALd A, Magalhães T, Ayres CFJ. High prevalence and lack of diversity of Wolbachia pipientis in Aedes albopictus populations from Northeast Brazil. Mem Inst Oswaldo Cruz. 2011;106(6):773. doi: 10.1590/S0074-02762011000600021. [DOI] [PubMed] [Google Scholar]
- 46.Minard G, Tran FH, Van VT, Goubert C, Bellet C, Lambert G, et al. French invasive Asian tiger mosquito populations harbor reduced bacterial microbiota and genetic diversity compared to Vietnamese autochthonous relatives. Front Microbiol. 2015;6:970. doi: 10.3389/fmicb.2015.00970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Werren JH, Baldo L, Clark ME. Wolbachia: master manipulators of invertebrate biology. Nat Rev Micro. 2008;6:741–751. doi: 10.1038/nrmicro1969. [DOI] [PubMed] [Google Scholar]
- 48.Dobson SL, Marsland EJ, Rattanadechakul W. Mutualistic Wolbachia infection in Aedes albopictus: accelerating cytoplasmic drive. Genetics. 2002;160:1087–1094. doi: 10.1093/genetics/160.3.1087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Johnson KN. The impact of Wolbachia on virus infection in mosquitoes. Viruses. 2015;7:5705–5717. doi: 10.3390/v7112903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Duo-quan W, Lin-hua T, Zhen-cheng G, Xiang Z, Man-ni Y, Wei-kang J. Comparative evaluation of light-trap catches, electric motor mosquito catches and human biting catches of Anopheles in the Three Gorges Reservoir. PLoS One. 2012;7:e28988. [DOI] [PMC free article] [PubMed]
- 51.Wang G, Zhang H, Cao X, Zhang X, Wang G, He Z, et al. Using GARP to predict the range of Aedes aegypti in China. Southeast Asian J Trop Med Public Health. 2014;45:290–298. [PubMed] [Google Scholar]
- 52.Lu B. Fauna Sinica Insecta (Vol.8). Diptera Culicidae 1. Beijing: Science Press; 1997. [Google Scholar]
- 53.Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol. 1994;3:294–9. [PubMed]
- 54.Hebert PDN, Cywinska A, Ball SL, deWaard JR. Biological identifications through DNA barcodes. Proc Biol Sci. 2003;270(1512):313–321. doi: 10.1098/rspb.2002.2218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.De Jong L, Moreau X, Dalia J, Coustau C, Thiery A. Molecular characterization of the invasive Asian tiger mosquito, Aedes (Stegomyia) albopictus (Diptera: Culicidae) in Corsica. Acta Trop. 2009;112:266–269. doi: 10.1016/j.actatropica.2009.08.008. [DOI] [PubMed] [Google Scholar]
- 56.Werren JH, Windsor DM. Wolbachia infection frequencies in insects: evidence of a global equilibrium? Proc Biol Sci. 2000;267(1450):1277–1285. doi: 10.1098/rspb.2000.1139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhou W, Rousset F, O'Neil S. Phylogeny and PCR-based classification of Wolbachia strains using wsp gene sequences. Proc Biol Sci. 1998;265(1395):509–515. doi: 10.1098/rspb.1998.0324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41:95–98. [Google Scholar]
- 59.Librado P, Rozas J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics. 2009;25:1451–1452. doi: 10.1093/bioinformatics/btp187. [DOI] [PubMed] [Google Scholar]
- 60.Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–1874. doi: 10.1093/molbev/msw054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Fu YX. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics. 1997;147:915–925. doi: 10.1093/genetics/147.2.915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tajima F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics. 1989;123(3):585–595. doi: 10.1093/genetics/123.3.585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Clement M, Posada D, Crandall KA. TCS: a computer program to estimate gene genealogies. Mol Ecol. 2000;9:1657–1659. doi: 10.1046/j.1365-294x.2000.01020.x. [DOI] [PubMed] [Google Scholar]
- 64.Futami K, Valderrama A, Baldi M, Minakawa N, Marin Rodriguez R, Chaves LF. New and common haplotypes shape genetic diversity in Asian tiger mosquito populations from Costa Rica and Panama. J Econ Entomol. 2015;108:761–768. doi: 10.1093/jee/tou028. [DOI] [PubMed] [Google Scholar]
- 65.Excoffier L, Lischer HEL. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010;10:564–567. doi: 10.1111/j.1755-0998.2010.02847.x. [DOI] [PubMed] [Google Scholar]
- 66.Katoh K, Standley DM. A simple method to control over-alignment in the MAFFT multiple sequence alignment program. Bioinformatics. 2016;32:1933–1942. doi: 10.1093/bioinformatics/btw108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Letunic I, Bork P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016;44(W1):W242–W245. doi: 10.1093/nar/gkw290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Lukhtanov VA, Dantchenko AV, Vishnevskaya MS, Saifitdinova AF. Detecting cryptic species in sympatry and allopatry: analysis of hidden diversity in Polyommatus (Agrodiaetus) butterflies (Lepidoptera: Lycaenidae) Biol J Linn Soc Lond. 2015;116:468–485. doi: 10.1111/bij.12596. [DOI] [Google Scholar]
- 69.Muangmai N, Ammon UV, Zuccarello GC. Cryptic species in sympatry: nonrandom small-scale distribution patterns in Bostrychia intricata (Ceramiales, Rhodophyta) Phycologia. 2016;55:424–430. doi: 10.2216/16-5.1. [DOI] [Google Scholar]
- 70.Karanovic T, Djurakic M, Eberhard SM. Cryptic species or inadequate taxonomy? Implementation of 2Dtaxa geometric morphometrics based on integumental organs as landmarks for delimitation and description of copepod taxa. Syst Biol. 2016;65:304–327. doi: 10.1093/sysbio/syv088. [DOI] [PubMed] [Google Scholar]
- 71.Tabachnick WJ. Challenges in predicting climate and environmental effects on vector-borne disease episystems in a changing world. J Exp Biol. 2010;213:946–954. doi: 10.1242/jeb.037564. [DOI] [PubMed] [Google Scholar]
- 72.Gould EA, Higgs S. Impact of climate change and other factors on emerging arbovirus diseases. Trans R Soc Trop Med Hyg. 2009;103:109–121. doi: 10.1016/j.trstmh.2008.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Barrera R, Amador M, MacKay AJ. Population dynamics of Aedes aegypti and dengue as influenced by weather and human behavior in San Juan, Puerto Rico. PLoS Negl Trop Dis. 2011;5:e1378. doi: 10.1371/journal.pntd.0001378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Delatte H, Toty C, Boyer S, Bouetard A, Bastien F, Fontenille D. Evidence of habitat structuring Aedes albopictus populations in Réunion Island. PLoS Negl Trop Dis. 2013;7:e2111. doi: 10.1371/journal.pntd.0002111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Waldock J, Chandra NL, Lelieveld J, Proestos Y, Michael E, Christophides G, et al. The role of environmental variables on Aedes albopictus biology and chikungunya epidemiology. Pathog Glob Health. 2013;107:224–241. doi: 10.1179/2047773213Y.0000000100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Xi Z, Dean JL, Khoo C, Dobson SL. Generation of a novel Wolbachia infection in Aedes albopictus (Asian tiger mosquito) via embryonic microinjection. Insect Biochem Mol Biol. 2005;35:903–910. doi: 10.1016/j.ibmb.2005.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Xi Z, Khoo CC, Dobson SL. Wolbachia establishment and invasion in an Aedes aegypti laboratory population. Science. 2005;310(5746):326–328. doi: 10.1126/science.1117607. [DOI] [PubMed] [Google Scholar]
- 78.Bian G, Xu Y, Lu P, Xie Y, Xi Z. The endosymbiotic bacterium Wolbachia induces resistance to dengue virus in Aedes aegypti. PLoS Pathog. 2010;6:e1000833. doi: 10.1371/journal.ppat.1000833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Moreira LA, Iturbe-Ormaetxe I, Jeffery JA, Lu G, Pyke AT, Hedges LM, et al. A Wolbachia symbiont in Aedes aegypti limits infection with dengue, chikungunya, and Plasmodium. Cell. 2009;139:1268–1278. doi: 10.1016/j.cell.2009.11.042. [DOI] [PubMed] [Google Scholar]
- 80.Walker T, Johnson PH, Moreira LA, Iturbe-Ormaetxe I, Frentiu FD, McMeniman CJ, et al. The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature. 2011;476(7361):450–453. doi: 10.1038/nature10355. [DOI] [PubMed] [Google Scholar]
- 81.van den Hurk AF, Hall-Mendelin S, Pyke AT, Frentiu FD, McElroy K, Day A, et al. Impact of Wolbachia on infection with chikungunya and yellow fever viruses in the mosquito vector Aedes aegypti. PLoS Negl Trop Dis. 2012;6:e1892. doi: 10.1371/journal.pntd.0001892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Townson H. Wolbachia as a potential tool for suppressing filarial transmission. Ann Trop Med Parasitol. 2002;96(Suppl. 2):S117–S127. doi: 10.1179/000349802125002464. [DOI] [PubMed] [Google Scholar]
- 83.Kambris Z, Cook PE, Phuc HK, Sinkins SP. Immune activation by life-shortening Wolbachia and reduced filarial competence in mosquitoes. Science. 2009;326(5949):134–136. doi: 10.1126/science.1177531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Hughes GL, Koga R, Xue P, Fukatsu T, Rasgon JL. Wolbachia infections are virulent and inhibit the human malaria parasite Plasmodium falciparum in Anopheles gambiae. PLoS Pathog. 2011;7:e1002043. [DOI] [PMC free article] [PubMed]
- 85.Bian G, Joshi D, Dong Y, Lu P, Zhou G, Pan X, et al. Wolbachia invades Anopheles stephensi populations and induces refractoriness to Plasmodium infection. Science. 2013;340(6133):748–751. doi: 10.1126/science.1236192. [DOI] [PubMed] [Google Scholar]
- 86.Murdock CC, Blanford S, Hughes GL, Rasgon JL, Thomas MB. Temperature alters Plasmodium blocking by Wolbachia. Sci Rep. 2014;4:3932. doi: 10.1038/srep03932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Caragata EP, Dutra HL, Moreira LA. Inhibition of Zika virus by Wolbachia in Aedes aegypti. Microb Cell. 2016;3:293–295. doi: 10.15698/mic2016.07.513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Afizah AN, Roziah A, Nazni WA, Lee HL. Detection of Wolbachia from field collected Aedes albopictus Skuse in Malaysia. Indian J Med Res. 2015;142:205–210. doi: 10.4103/0971-5916.164259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Kitrayapong P, Baimai V, O’Neill SL. Field prevalence of Wolbachia in the mosquito vector Aedes albopictus. Am J Trop Med Hyg. 2002;66:108–111. doi: 10.4269/ajtmh.2002.66.108. [DOI] [PubMed] [Google Scholar]
- 90.Sivan A, Shriram AN, Bhattacharya D, Vijayachari P. Wolbachia endobacterium in wild population of Aedes albopictus (Skuse) (Diptera: Culicidae) and phylogeny from Andaman and Nicobar Islands, India. J Vector Borne Dis. 2014;51:235–238. [PubMed] [Google Scholar]
- 91.Nugapola NWNP, De Silva WAPP, Karunaratne SHPP. Distribution and phylogeny of Wolbachia strains in wild mosquito populations in Sri Lanka. Parasit Vectors. 2017;10:230. doi: 10.1186/s13071-017-2174-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Armbruster P, Damsky WE, Jr, Giordano R, Birungi J, Munstermann LE, Conn JE. Infection of New- and Old-World Aedes albopictus (Diptera: Culicidae) by the intracellular parasite Wolbachia: implications for host mitochondrial DNA evolution. J Med Entomol. 2003;40:356–360. doi: 10.1603/0022-2585-40.3.356. [DOI] [PubMed] [Google Scholar]
- 93.Baldini F, Segata N, Pompon J, Marcenac P, Robert Shaw W, Dabiré RK, et al. Evidence of natural Wolbachia infections in field populations of Anopheles gambiae. Nat Commun. 2014;5:3985. doi: 10.1038/ncomms4985. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. K2P divergence of cox1 sequences in Ae. albopictus from China. (XLSX 76 kb)
Table S2. K2P divergence of cox1 sequences in Aedes sp. identified in China. (XLSX 9 kb)
Table S3. cox1 haplotype distribution in the 14 mosquito populations from China. (XLSX 14 kb)
Data Availability Statement
The data supporting the findings of this article are included within the article. Nucleotide sequence data reported in this paper have been deposited to the NBCI GenBank database under the accession numbers: KY765450-KY765506 and MF623839-MF623851.


