Abstract
The global coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has led to unprecedented social and economic consequences. The risk of morbidity and mortality due to COVID-19 increases dramatically in the presence of coexisting medical conditions, while the underlying mechanisms remain unclear. Furthermore, there are no approved therapies for COVID-19. This study aims to identify SARS-CoV-2 pathogenesis, disease manifestations, and COVID-19 therapies using network medicine methodologies along with clinical and multi-omics observations. We incorporate SARS-CoV-2 virus–host protein–protein interactions, transcriptomics, and proteomics into the human interactome. Network proximity measurement revealed underlying pathogenesis for broad COVID-19-associated disease manifestations. Analyses of single-cell RNA sequencing data show that co-expression of ACE2 and TMPRSS2 is elevated in absorptive enterocytes from the inflamed ileal tissues of Crohn disease patients compared to uninflamed tissues, revealing shared pathobiology between COVID-19 and inflammatory bowel disease. Integrative analyses of metabolomics and transcriptomics (bulk and single-cell) data from asthma patients indicate that COVID-19 shares an intermediate inflammatory molecular profile with asthma (including IRAK3 and ADRB2). To prioritize potential treatments, we combined network-based prediction and a propensity score (PS) matching observational study of 26,779 individuals from a COVID-19 registry. We identified that melatonin usage (odds ratio [OR] = 0.72, 95% CI 0.56–0.91) is significantly associated with a 28% reduced likelihood of a positive laboratory test result for SARS-CoV-2 confirmed by reverse transcription–polymerase chain reaction assay. Using a PS matching user active comparator design, we determined that melatonin usage was associated with a reduced likelihood of SARS-CoV-2 positive test result compared to use of angiotensin II receptor blockers (OR = 0.70, 95% CI 0.54–0.92) or angiotensin-converting enzyme inhibitors (OR = 0.69, 95% CI 0.52–0.90). Importantly, melatonin usage (OR = 0.48, 95% CI 0.31–0.75) is associated with a 52% reduced likelihood of a positive laboratory test result for SARS-CoV-2 in African Americans after adjusting for age, sex, race, smoking history, and various disease comorbidities using PS matching. In summary, this study presents an integrative network medicine platform for predicting disease manifestations associated with COVID-19 and identifying melatonin for potential prevention and treatment of COVID-19.
This study uses network medicine methodologies along with multiomic observations (including SARS-CoV-2 and human interactome, transcriptome, and proteome) to identify widespread disease manifestations and drug repurposing candidates (including melatonin) for COVID-19, validating these findings using a large-scale patient registry database.
Introduction
The ongoing global coronavirus disease 2019 (COVID-19) pandemic has led to 38 million confirmed cases and 1 million deaths worldwide as of October 14, 2020. The United States alone has recorded nearly 8 million confirmed cases, with a death toll of more than 216,000 [1]. Several retrospective studies have reported the clinical characteristics of individuals with symptomatic COVID-19, and an emerging theme has been the significantly higher risk of morbidity and mortality among individuals with 1 or more comorbid health conditions, such as hypertension, asthma, diabetes mellitus, cardiovascular or cerebrovascular disease, chronic kidney disease, and malignancy [2–7]. However, these retrospective clinical studies are limited by small sample sizes and unmeasured confounding factors, leaving the underlying patho-mechanisms largely unknown. More specifically, it is unclear whether associations of disease manifestations and COVID-19 severity are merely a reflection of poorer health in general, or a clue to shared pathobiological mechanisms.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the virus that causes COVID-19, is an enveloped virus that carries a single-stranded positive-sense RNA genome [7,8]. SARS-CoV-2 is a newly discovered member of the coronavirus (CoV) family [9]. SARS-CoV-2 enters host cells via binding of its spike protein to the angiotensin converting enzyme 2 (ACE2) receptor on the surfaces of many cell types [10]. This binding is primed by transmembrane protease serine 2 (TMPRSS2) [10] and the host cell protease furin [11] (Fig 1A). Studies have shown that ACE2 and TMPRSS2 are highly co-expressed in alveolar type II (AT2) epithelial cells in the lung [12], nasal mucosa [13], bronchial secretory cells [14], and absorptive enterocytes in the ileum [15]. Yet, much remains to be learned about how these critical human proteins involved in the infection and replication of SARS-CoV-2 are associated with various disease comorbidities and complications. Systematic identification of the host factors involved in the protein–protein interactions (PPIs) of SARS-CoV-2 and the human host will facilitate identification of drug targets and advance understanding of the complications and comorbidities resulting from COVID-19 [16–20]. Studies using transcriptomics [21], proteomics [22], and interactomics (PPIs) methods [8] have contributed to a better understanding of the SARS-CoV-2–host interactome, which has enabled the investigation of the complications and comorbidities of SARS-CoV-2 and a facilitated search for effective treatment (Fig 1B).
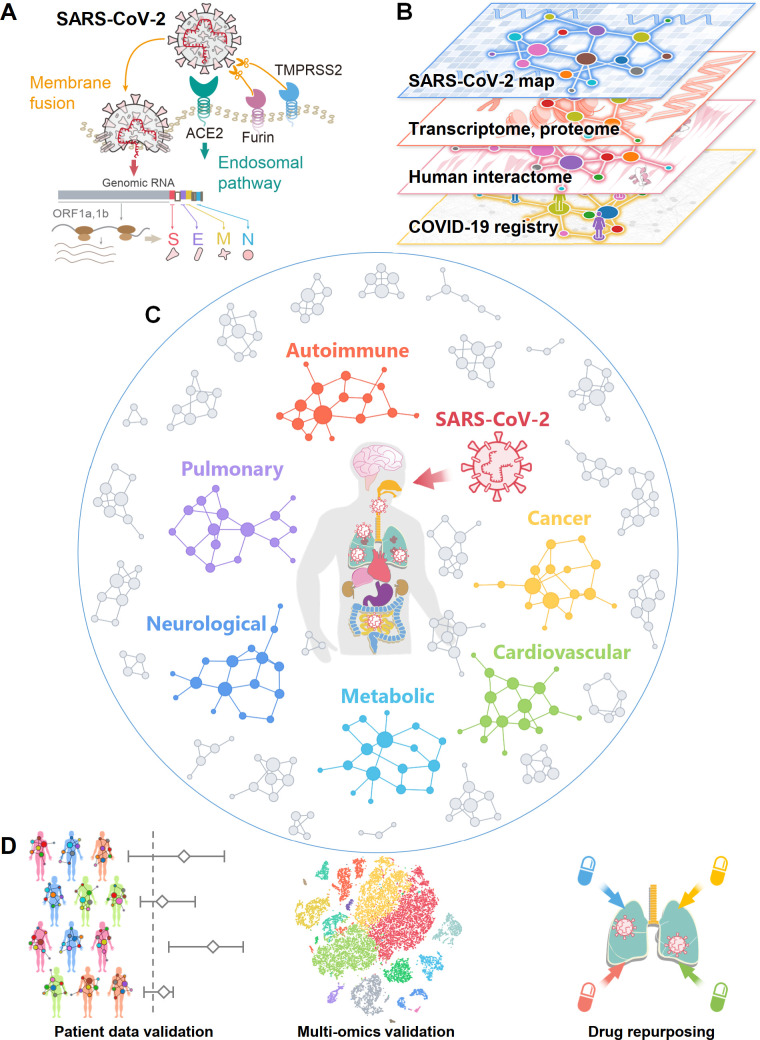
Fig 1. Overall workflow of this study.
(A) A diagram illustrating the basic pathogenesis of SARS-CoV-2. (B) A diagram illustrating how to build a global interactome map for SARS-CoV-2. We compiled the SARS-CoV-2 human target gene/protein sets from multi-omics data from the transcriptome, proteome, and human interactome, and validated network-based findings using patient data from a COVID-19 registry. (C) A diagram illustrating network-based measurement of disease manifestations associated with COVID-19. We systematically evaluated the network proximities of the SARS-CoV-2 human target genes/proteins with 64 diseases across 6 main categories: autoimmune, cancer, cardiovascular, metabolic, neurological, and pulmonary. (D) A workflow illustrating validation of network-based findings. We performed single-cell analyses to further investigate the underlying mechanisms of COVID-19 with asthma and inflammatory bowel disease. We prioritized nearly 3,000 US Food and Drug Administration–approved/investigational drugs for their potential anti-SARS-CoV-2 effects from network-based findings and validated drug–COVID-19 outcomes using an institutional review board–approved COVID-19 patient registry.
Major efforts are underway to develop safe and effective drugs to treat COVID-19: Preventive and therapeutic strategies currently being explored include vaccination, SARS-CoV-2-specific antibodies, novel nucleoside analogs such as remdesivir, and repurposed drugs [23,24]. Remdesivir, an agent originally developed for treatment of Ebola virus, was reported to shorten the time to recovery in adults who were hospitalized with COVID-19 [25]; yet, a 10-day course of remdesivir did not show a statistically significant difference in clinical status compared with standard care for patients with moderate COVID-19 [26]. Dexamethasone, an FDA-approved glucocorticoid receptor (GR) agonist, has been shown to reduce mortality by one-third in hospitalized COVID-19 patients requiring ventilation and by one-fifth in individuals requiring oxygen [27]; yet, dexamethasone did not reduce death in COVID-19 patients not receiving respiratory support [27]. Many existing drugs are currently being or have been tested in clinical trials, such as the antimalarial drug hydroxychloroquine and protease inhibitor combination lopinavir/ritonavir; results from these trials have not yet shown significant clinical benefits for COVID-19 patients [28,29]. We recently evaluated nearly 3,000 FDA-approved/investigational drugs using a network-based method and prioritized 16 drug candidates and 3 drug combinations for COVID-19 [30]. Yet, the answer to the key question of why an approved drug originally documented for other diseases might be beneficial for COVID-19 remains unclear. One possible explanation is that COVID-19 shares common disease pathobiology or functional pathways elucidated by the human PPIs [30–32]. Systematic identification of common disease pathobiological pathways shared by COVID-19 and other diseases would offer novel targets and therapies for COVID-19.
In this study, we present an integrative network medicine platform that quantifies the association of COVID-19 with other diseases across 6 categories, including autoimmune, malignant cancer, cardiovascular, metabolic, neurological, and pulmonary (Fig 1C). The rationale for these analyses rests on the notions that (1) the proteins that functionally associate with a disease (such as COVID-19) are localized in the corresponding subnetwork within the comprehensive human PPI network [31–34] and (2) proteins that are associated with a specific disease may be directly targeted by the virus or are in the close vicinity of the target host proteins. We first performed network analysis followed by single-cell RNA sequencing (RNA-Seq) data analysis to identify the underlying pathobiological relationships between COVID-19 and its associated comorbidities. Additionally, we use our network medicine findings and patient data from a large COVID-19 patient registry database to identify and prioritize existing FDA-approved drugs as potential COVID-19 drug candidates (Fig 1D).
Results
A global map of the SARS-CoV-2 virus–host interactome
We assembled 4 host gene/protein sets for SARS-CoV-2 (S1 and S2 Tables): (1) SARS2-DEG, representing the differentially expressed genes (DEGs) from the transcriptomic data of SARS-CoV-2-infected primary human bronchial epithelial cells; (2) SARS2-DEP, representing the differentially expressed proteins (DEPs) from the proteomic data of SARS-CoV-2-infected human Caco-2 cells; (3) HCoV-PPI, representing the literature-based virus–host proteins across multiple human coronaviruses (HCoVs), including SARS-CoV-1 (from the 2002–2003 pandemic) and MERS-CoV; and (4) SARS2-PPI (SARS-CoV-2-specific virus–host PPIs). Since HCoV-PPI and SARS2-PPI both involve physical virus–host PPIs, we further combined them as the fifth dataset, PanCoV-PPI.
We first performed functional enrichment analyses for the 5 different gene/protein datasets. We found that these datasets share several common pathways and ontology terms (Fig 2A; S1 Data), such as phagosome, measles, apoptosis, NF-κB signaling pathway, neutrophil-related immunity, apoptotic processes, virus transport, viral genome replication, and response to interferon, yet they differ considerably in terms of their most significantly enriched pathways (S1–S5 Figs). This is especially noticeable for SARS2-DEP and SARS2-PPI. While SARS2-DEG (S1 Fig) and HCoV-PPI (S3 Fig) show more enrichment in immune responses and viral pathways, SARS2-DEP (S2 Fig) is more related to various cellular metabolic pathways, and SARS2-PPI (S4 Fig) is more enriched in DNA replication, RNA transcription, and protein translation. These observations suggest that these different SARS-CoV-2 viral–host gene/protein sets capture complementary aspects of the biological and cellular states of the viral life cycle and host immunity. Therefore, building a global virus–host map (including interactome, transcriptome, and proteome) that incorporates data from transcriptomics, proteomics, and physical virus–host PPIs are essential for a better understanding of the pathogenesis of COVID-19. This global virus–host map for SARS-CoV-2 can offer a more complete picture of the interconnected functional pathways involved in viral pathogenesis, thereby facilitating the discovery of therapeutic targets.
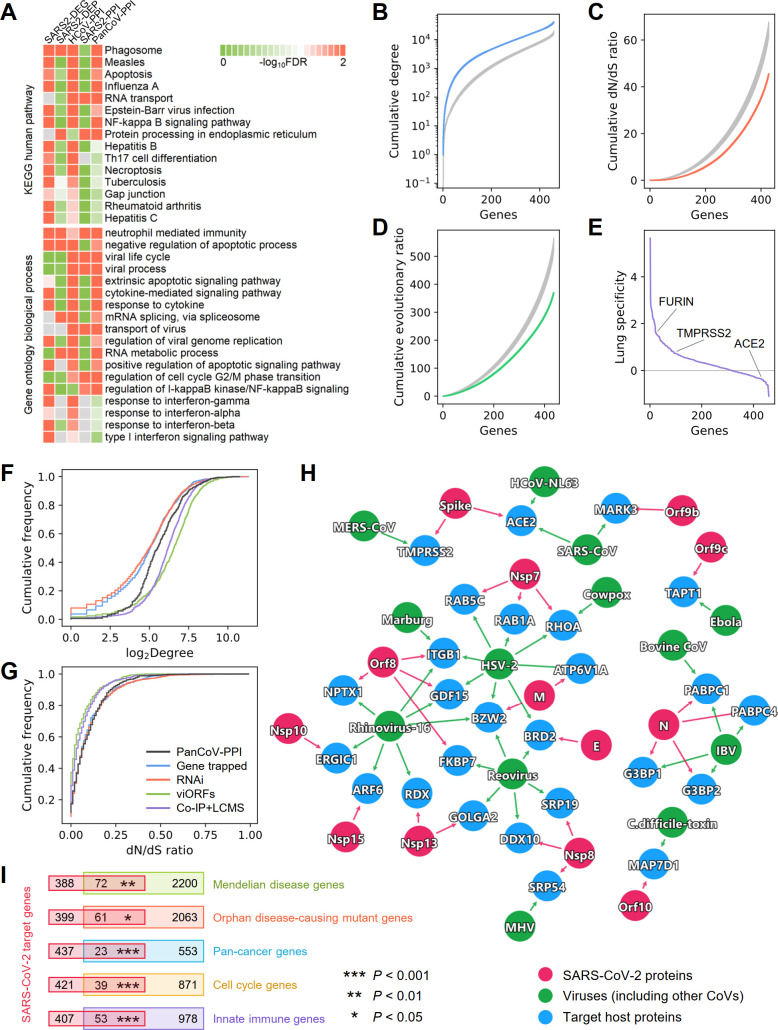
Fig 2. Network and biological characteristics of the SARS-CoV-2 interactome map.
(A) Pathway and Gene Ontology (biological process) enrichment analysis results of the SARS-CoV-2 host genes/proteins across 5 different datasets. We assembled 5 gene/protein datasets from SARS-CoV-2 host protein–protein interactions, transcriptomics, and proteomics (S2 Table). (B–D) Network and biological characteristics of the SARS-CoV-2 host genes/proteins. The proteins in PanCoV-PPI have higher node degrees (B), lower dN/dS ratios (C), and lower evolutionary ratios (D) compared to randomly selected proteins (grey, mean ± standard deviation of 100 repeats). (E) Among the 460 proteins in PanCoV-PPI, 450 (98%) are expressed in lungs, and 317 (69%) have lung-specific expression (Z > 0). (F and G) The distribution of the node degrees in the human interactome and dN/dS ratios of PanCoV-PPI and 4 published virus-related host protein sets (S3 Table). (H) The shared target human proteins (blue) of SARS-CoV-2 (red) and other viruses (green). (I) SARS-CoV-2 target proteins overlap significantly with disease-associated genes (Mendelian disease and orphan disease), cancer genes, cell cycle genes, and innate immune genes. The data underlying this figure can be found in S1 Data. Co-IP+LCMS, co-immunoprecipitation and liquid chromatography–mass spectrometry; CoV, coronavirus; dN/dS ratio, nonsynonymous to synonymous substitution rate ratio; FDR, false discovery rate; KEGG, Kyoto Encyclopedia of Genes and Genomes; RNAi, RNA interference; viORF, viral open reading frame.
Network and biological characteristics of virus–host interactome for SARS-CoV-2
In addition to identifying the functions that these viral gene/protein sets represent, we next characterized the network patterns (node degree in the human PPI network) and bioinformatics features of these SARS-CoV-2 datasets, including dN/dS ratio, evolutionary rate ratio, and lung expression specificity (Figs 2B–2G and S6). To find common as well as unique network and bioinformatic characteristics of SARS-CoV-2, we further compiled 4 additional virus–host gene/protein networks, identified by different methods, for comparison (S3 Table): (1) 900 virus–host interactions connecting 10 other viruses and 712 host genes identified by gene-trap insertional mutagenesis, (2) 2,855 known virus–host interactions connecting 2,443 host genes and 55 pathogens identified from RNA interference (RNAi), (3) 579 host proteins mediating translation of 70 innate immune-modulating viral open reading frames (viORFs), and (4) 1,292 host genes mediating influenza–host interactions identified by co-immunoprecipitation and liquid chromatography-mass spectrometry (Co-IP+LC/MS). These virus–host gene/protein networks have been well characterized [35–37] and offer high-quality datasets (S3 Table) for comparisons. We found that host proteins in PanCoV-PPI (Fig 2B) and 4 other datasets (SARS2-DEG, SARS2-DEP, HCoV-PPI, and SARS2-PPI) (S6 Fig) were more likely to be highly connected in the human PPI network. Several hub genes, such as JUN, XPO1, MOV10, NPM1, VCP, and HNRNPA1, have the highest degree (connectivity) in the PanCoV-PPI network (S2 Table). PanCoV-PPI has a comparable degree distribution with host genes/proteins identified by viORFs and Co-IP+LC/MS, although marginally higher than that identified by RNAi and gene-trap insertional mutagenesis assay (Fig 2F).
Expression patterns of genes in a specific disease-related tissue play a crucial role for elucidation of disease pathogenesis and drug discovery [31,32]. Given the major impact of SARS-CoV-2 on pulmonary function and lung injury [2], we inspected the lung-specific expression of genes in PanCoV-PPI using a Z score measure (see Materials and Methods—Tissue specificity analysis) compared to expression in other tissues from the GTEx database [38]. We found that most host genes for SARS-CoV-2 have high expression in lung (Fig 2E; S2 Table) compared to other tissues; yet, ACE2 has a low expression in lung compared to other tissues. A recent study showed that ACE2 was primarily expressed in the epithelial cells in lungs, and only 3.8% of AT2 pneumocytes expressed both ACE2 and TMPRSS2, but ACE2 is significantly upregulated in smokers and 24–48 hours following SARS-CoV-2 infection [12,39]. Another study also showed that despite relatively low expression of ACE2 in the lung, ACE2 was expressed in multiple epithelial cell types along the airway [13]. Therefore, it is important to understand the cell-type-specific SARS-CoV-2 pathogenesis at the single-cell level.
To inspect the evolutionary factors underlying the SARS-CoV-2–human PPIs, we investigated the selective pressure and evolutionary rates quantified by the dN/dS ratio using human–mouse orthologous gene pairs. We found that PanCoV-PPI shows stronger purifying selection (lower dN/dS ratio [Fig 2C] and evolutionary rate ratio [Fig 2D]) compared to the same number of random genes. PanCoV-PPI is also comparable to 4 other virus–host genes/protein datasets identified by different assays (Fig 2F and 2G) in terms of node degrees and dN/dS ratios. Altogether, these observations suggest that the virus–host PPIs assembled in this study offer a high-quality interactome map for SARS-CoV-2 for identifying pathogenesis and potential treatments for COVID-19.
To inspect shared viral pathways across different viruses and SARS-CoV-2, we further performed network overlap analysis of PanCoV-PPI with 712 host genes across 10 types of other viruses identified by gene-trap insertional mutagenesis assays [35]. We found a significant overlap of the SARS-CoV-2 host proteins with non-coronaviruses (P < 0.002, Fisher’s exact test; Fig 2H). For example, BRD2, a transcriptional regulator that belongs to the Bromodomain and Extra-Terminal motif family, is connected to SARS-CoV-2 and 2 other viruses, herpes simplex virus 2 (HSV-2) and reovirus. RHOA, encoding a small GTPase protein in the Rho family of GTPases, is connected to SARS-CoV-2, cowpox, and HSV-2 as well. RHOA has been reported to be involved in multiple human diseases, including cardiovascular disease [40] and cancer [41]. These observations indicate possible disease manifestations associated with SARS-CoV-2.
SARS-CoV-2 cellular network perturbations of disease manifestations
Investigation of the relationships between human host proteins targeted by SARS-CoV-2 and disease susceptibility genes may offer crucial information for identifying COVID-19-associated disease manifestations. We thus inspected the overlap between SARS-CoV-2 host genes/proteins and the susceptibility gene sets implicated in different diseases and biological events (Fig 2I). We found that host genes/proteins targeted by SARS-CoV-2 are significantly enriched in Mendelian disease (P = 0.002, Fisher’s exact test), orphan disease (P = 0.044), and cancer (P < 0.001). Mechanistically, SARS-CoV-2 target host genes are significantly enriched in cell cycle genes (P < 0.001) and innate immune genes (P < 0.001).
Using the 5 SARS-CoV-2 host gene/protein sets, we next tried to identify potential COVID-19 comorbidities. To achieve this, we assembled the disease–protein network of COVID-19 and 6 disease categories (S4 Table). For pan-cancer analysis, the somatic driver genes were retrieved from the Cancer Gene Census Tier 1 gene table [42,43]. The putative somatic driver genes for individual cancer types were identified from The Cancer Genome Atlas projects and were downloaded from a previous study [44]. For autoimmune, pulmonary, neurological, cardiovascular, and metabolic categories, we extracted their associated genes/proteins from the Human Gene Mutation Database (HGMD) [45,46]. Each gene from HGMD has at least 1 reported mutation associated with the disease. This information on the genes, in addition to the sources, counts, and terms used in HGMD to identify the diseases, can be found in S4 Table. Using the disease–protein network together with the SARS2-PPI (SARS-CoV-2 virus–host interactome) and HCoV-PPI (HCoV–host interactome) sets, we examined the overall connectivity of the disease-associated proteins and the SARS-CoV-2 target host proteins in the human PPI network (Fig 3A; S1 File; S2 Data). To build the global network for the disease comorbidities, we extracted the PPIs from the human interactome for the virus target proteins and disease-associated proteins. Each node indicates a virus target host protein (blue) or a disease-associated protein (green). Some disease-associated proteins can be directly targeted by the viruses, as shown in orange. Edges among these protein nodes indicate PPIs. For SARS-CoV-2, the targets of its individual viral proteins are shown based on the SARS2-PPI dataset. Due to their tendency of having common disease-associated proteins, some disease categories tend to cluster closely, e.g., cancer and neurological. Diseases from other categories, such as autoimmune and pulmonary, are scattered. Most of the virus target proteins are connected with the disease-associated proteins, which suggests shared pathobiological pathways of COVID-19 and these diseases. Various cancer types formed a relatively distant module from the virus targets, compared to other disease categories.
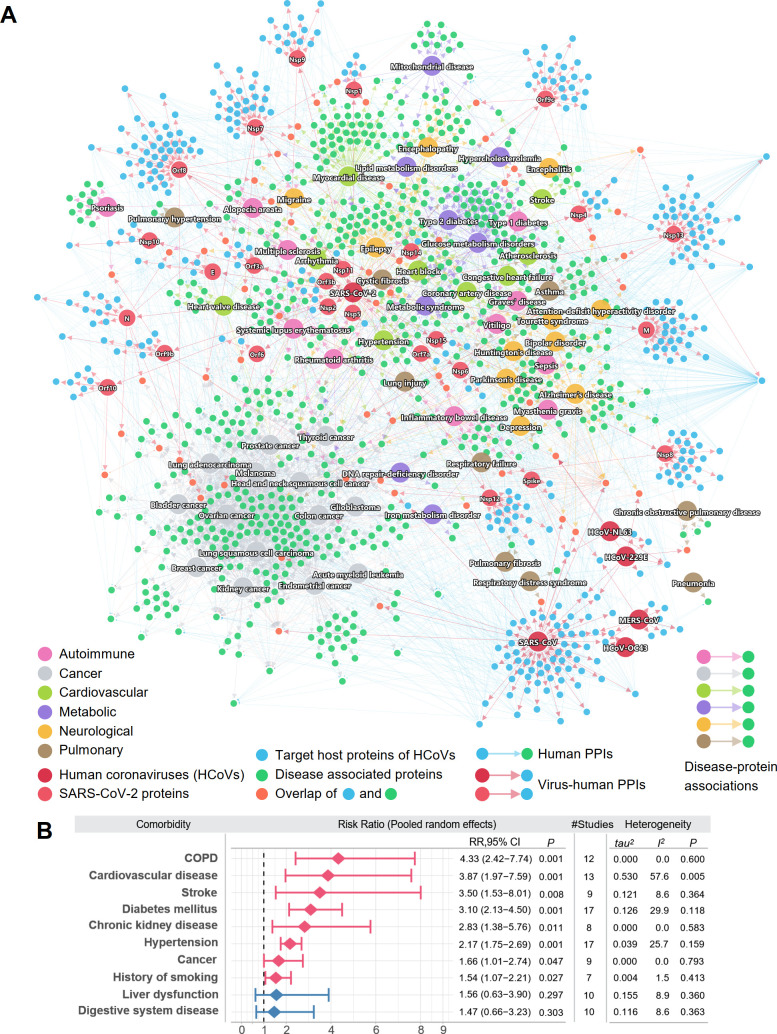
Fig 3. A global network illustrating disease manifestations associated with human coronavirus.
(A) The target human proteins of SARS-CoV-2 are connected to the disease-associated proteins. Blue links (edges) indicate physical protein–protein interactions. For SARS-CoV-2, its viral proteins are shown by light red nodes. The target human proteins (blue) of the viruses are intricately connected to the disease-associated proteins (green). Human disease nodes are colored by different disease categories: autoimmune, cancer, cardiovascular, metabolic, neurological, and pulmonary. (B) Estimation of the pooled risk ratio using random effects meta-analysis for 10 comorbidities between patients with severe versus non-severe COVID-19. The tau2 and I2 statistics were calculated to quantify the heterogeneity among studies. I2 ≤ 50% was considered as low heterogeneity among studies, 50% < I2 ≤ 75% was considered as moderate heterogeneity, and I2 > 75% was considered as high heterogeneity. The data underlying this figure can be found in S2 Data. COPD, chronic obstructive pulmonary disease; PPI, protein–protein interaction.
Shown in Fig 3A, the disease genes can interact with the SARS-CoV-2 viral proteins either directly or indirectly in the human protein–protein interactome. For example, among the 4 chronic obstructive pulmonary disease (COPD)–associated proteins shown in the network, TGFB1 is the direct target of HCoV-229E, and all 4 proteins (TGFB1, DEFB1, SNAI1, and ADAM33) interact with at least 1 SARS-CoV-2 target protein. The risk of various cardiovascular diseases was found to be increased in COVID-19 patients, including heart block, coronary artery disease, congestive heart failure, and arrhythmia, which is consistent with clinically reported myocardial injury [47] and cardiac arrest [48]. These observations reveal a common network relationship between COVID-19 and human diseases (Fig 3A). We then performed meta-analysis of 34 COVID-19 clinical studies (S5 Table) to evaluate the pooled risk ratios of 10 comorbidities among 4,973 COVID-19-positive patients (including 2,268 with mild and 731 with severe COVID-19). The random effects model was used to estimate the pooled risk ratio of disease severity. The tau2 and I2 statistics were used to evaluate the heterogeneity among studies (see Materials and Methods—Risk ratio analysis for COVID-19 patients). We found that patients with several disease comorbidities or risk factors—including COPD, cardiovascular disease, stroke, diabetes mellitus, chronic kidney disease, hypertension, cancer, and history of smoking—have significantly higher risks of severe COVID-19 (Fig 3B). The overall pooled risk ratio for patients with COPD was 4.33 (95% CI 2.42–7.74, P = 0.001) in 12 low heterogeneous clinical studies (I2 = 0.0%, P = 0.6). The COVID-19 patients with cardiovascular diseases had a risk ratio of 3.87 (95% CI 1.97–7.59, P = 0.001), and there was slightly higher heterogeneity across the 13 studies (I2 = 57.6%, P = 0.005). We next turned to quantify the network-based relationships between COVID-19 and human diseases in the human interactome model using state-of-the-art network proximity analysis [32,33,49].
Network-based measurement of COVID-19-associated disease manifestations
We systematically evaluated the network-based relationships of the 64 diseases across the 6 categories to COVID-19 (Fig 4A; S3 Data). We used the state-of-the-art network proximity measure to evaluate the connectivity and the closeness of the disease proteins and SARS-CoV-2 host proteins, taking the topology of the human interactome network into consideration. To test the significance of the proximity, Z scores and P values were calculated based on permutation tests (see Materials and Methods—Network proximity measure) and are shown in Fig 4A. We found that each disease–disease pair has a well-defined network-based footprint. If the footprint between the COVID-19 module and another disease module is significantly close (low Z score and P < 0.05), the magnitude of the proximity is indicative of their biological relationship: Closer network proximity (Fig 4A) of SARS-CoV-2 host genes/proteins with a disease module indicates higher potential of manifestation between COVID-19 and a specific disease. We first noticed that immunological, pulmonary, and neurological diseases showed significant network proximity to different SARS-CoV-2 maps more frequently than did cancer, cardiovascular, and metabolic diseases. Several diseases have significant network proximities to more than 1 SARS-CoV-2 dataset, most notably inflammatory bowel disease (IBD), attention-deficit/hyperactivity disorder, and stroke, which achieved significant P values for all 5 SARS-CoV-2 protein sets. Pulmonary diseases, including COPD, lung injury, pulmonary fibrosis, and respiratory failure, achieved 4 significant proximities. Some diseases have significant proximities to certain SARS-CoV-2 datasets, indicating associations at certain levels, e.g., asthma (transcriptomic), respiratory distress syndrome (proteomic), and hypertension (HCoV-PPI and PanCoV-PPI).
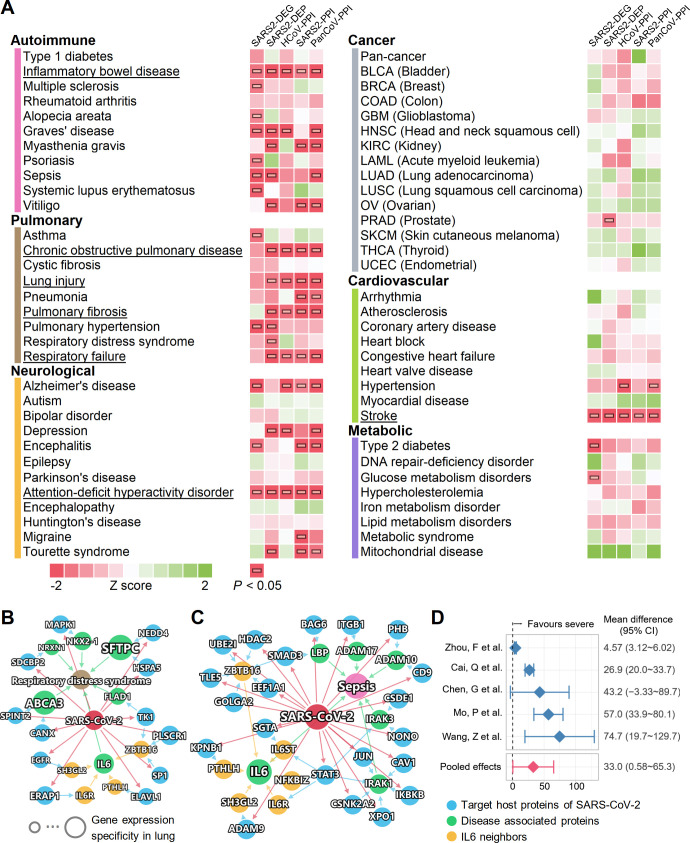
Fig 4. A landscape of disease manifestations associated with COVID-19 quantified by network proximity measure.
(A) Heatmaps showing the network proximities of COVID-19 with 64 diseases across 6 categories. The network proximities of the disease modules and the 5 SARS-CoV-2 datasets were evaluated using the “closest” network proximity measure (see Materials and Methods—Network proximity measure). The magnitude of the proximity is indicative of their biological relationship: Closer network proximity of SARS-CoV-2 host genes/proteins and a disease indicates higher potential of manifestation between COVID-19 and the disease. P < 0.05 computed by permutation test was considered significant (indicated by horizontal rectangles). Three categories, autoimmune, pulmonary, and neurological, frequently show significant proximities to COVID-19. Inflammatory bowel disease, attention-deficit/hyperactivity disorder, and stroke achieved significance with all 5 SARS-CoV-2 datasets. Underlined diseases are those highlighted in the Results. (B and C) Highlighted subnetworks between SARS-CoV-2 host genes/proteins and the disease-associated proteins of respiratory distress syndrome (B) and sepsis (C). (D) Clinical data analyses showed an association of COVID-19 severity with IL-6 expression levels in patients. Meta-analysis of the random effects model was performed using the mean difference in IL-6 (pg/ml). There was high heterogeneity among these studies (I2 = 94%, P < 0.001). The data underlying this figure can be found in S3 Data.
Network visualization can further show the connections between SARS-CoV-2 and other diseases, for example, respiratory distress syndrome (Fig 4B), sepsis (Fig 4C), and COPD (S7 Fig). Respiratory distress syndrome and sepsis are the 2 main causes of mortality in patients with severe COVID-19 [50,51]. We found that multiple SARS-CoV-2 host proteins are directly connected with the disease-associated proteins (Fig 4B). ABCA3 is a lipid transporter located in the outer membrane of lamellar bodies in AT2 cells; mutations of the ABCA3 gene can disrupt pulmonary surfactant homeostasis and lead to inherited pulmonary diseases [52]. Another membrane surface protein, the pulmonary-associated surfactant protein C encoded by SFTPC, can cause lung injury when misfolded [53]. For sepsis, we noticed several inflammatory and immune-related proteins, such as IRAK1, IRAK3, IKBKB, and STAT3, in the network, suggesting overlap of the inflammatory response activated in COVID-19 and sepsis (Fig 4C). It has been reported that an overzealous production of certain cytokines, such as IL-6, caused by dysregulation of innate immune responses to SARS-CoV-2 infection, can result in a “cytokine storm,” better known as cytokine release syndrome (CRS) [54]. The potential prognoses of acute respiratory distress syndrome and sepsis using IL-6 expression levels have also been established [55–57]. IL-6 also has a significantly increased expression level in the human bronchial epithelial cells infected with SARS-CoV-2 from the SARS2-DEG dataset [21]. In addition, it is also potentially affected by SARS-CoV-2 through multiple PPIs (Fig 4B and 4C, IL-6 neighbors), such as IL-6R, endophilin A1 (encoded by SH3GL2), and parathyroid hormone like hormone (encoded by PTHLH). Our random effects meta-analysis of 5 clinical studies of COVID-19 revealed that there was an increase of IL-6 levels in patients with severe COVID-19 compared to those with non-severe COVID-19 (Fig 4D). The mean difference was 33.0 pg/ml (95% CI 0.58–65.3; Fig 4D), with high literature heterogeneity (I2 = 94%, P < 0.001). These results indicate that IL-6 plays a critical role in COVID-19-associated respiratory distress syndrome and sepsis. Due to the importance of IL-6 in SARS-CoV-2 infection, IL-6 antagonists including tocilizumab [58] (NCT04315480) and sarilumab (NCT04327388) are being investigated in clinical trials for treatment of patients with severe COVID-19.
As an illustration of the shared pathobiology and inflammatory pathways of COVID-19 with various disease manifestations (Fig 4), we next turned to focus on 2 inflammation-driven diseases, asthma and IBD.
Inflammatory molecular profile shared by COVID-19 and asthma
Patients with severe COVID-19 symptoms showed a higher prevalence of dyspnea (S8A Fig; P < 0.001). To understand the associations between COVID-19 and respiratory disease (including asthma), we adopted a multimodal analysis utilizing bulk and single-cell transcriptomics data and metabolomics data under the human interactome network model. To be specific, we identified the enzymes in the network that are associated with altered metabolites in COVID-19 patients. Comparing the DEGs from asthma patients and DEGs from COVID-19 patients, we aimed to find the common genes/proteins or interacting proteins in these patient groups. Using network analyses (degree enrichment and eigenvector centrality), we can rank the importance of these genes. Last, we examined their expression in the cell types that are more susceptible to SARS-CoV-2 (i.e., cells expressing more ACE2 and TMPRSS2).
Fig 5A shows the subnetwork of the connections among SARS-CoV-2 target host proteins and asthma-associated proteins. Most of these proteins have enriched connections within the subnetwork (Fig 5B; degree enrichment), i.e., more connections in the subnetwork than in a random network of the same size in the human interactome. To evaluate the potential influence of the nodes in the network, we also computed the eigenvector centrality of these proteins. A higher eigenvector centrality suggests a higher network influence of a specific protein node. Six overlapped proteins (orange) from both groups were identified: NFKBIA, IRAK3, TNC, IL-6, ADRB2, and CD86. A recent study showed that glucose metabolism plays a key role in influenza A–regulated cytokine storm [59]. The plasma metabolome of asthma patients versus healthy controls also suggests activated inflammatory and immune pathways [60]. Therefore, in addition to PPIs, we also integrated metabolomics data generated in a previously assembled asthma cohort [61]. By matching the enzymes of the differential metabolites and the proteins in the PPI network, we found 3 key metabolites: arachidonate, L-arginine, and L-citrulline. L-arginine and L-citrulline were decreased in the sera of COVID-19 patients [62]. These metabolites were also decreased in asthma patients [61]. Arachidonate, the precursor of a variety of products that regulate inflammatory pathways [63], was found to have an increased level in the inflamed airways of asthma patients [64]. Arachidonate can be converted by 5-lipoxygenase encoded by ALOX5 to leukotriene, which is released during an asthma attack and is responsible for the bronchoconstriction [65].
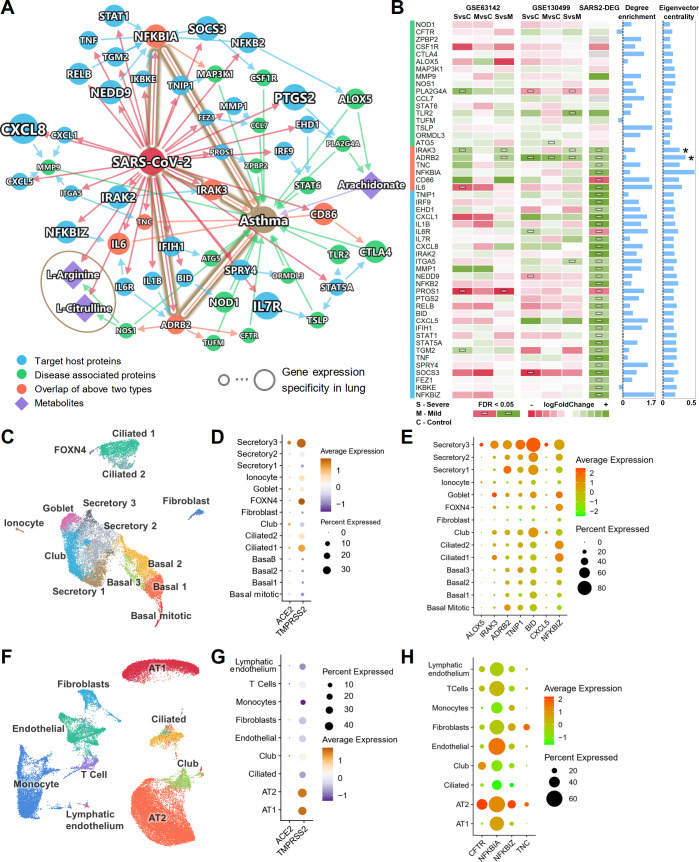
Fig 5. Shared molecular profile between asthma and COVID-19.
(A) A highlighted subnetwork between the asthma-associated genes, differential metabolites in asthma, and SARS-CoV-2 host genes, under the human interactome network model. (B) A heatmap highlighting differential gene expression analyses for the genes identified in the asthma and COVID-19 subnetwork analysis (A). We performed differential gene expression analysis using 2 existing asthma cohorts (GSE63142 and GSE130499). Blue bars show the node degree enrichment in the subnetwork (A) compared to a random network of the same size (left) and the eigenvector centrality (right). IRAK3 and ADRB2 (indicated with asterisks) are associated with asthma and are the targets of SARS-CoV-2. They had significantly elevated expression in asthma patients and SARS-CoV-2-infected human bronchial epithelial cells. (C) UMAP visualization for human bronchial epithelial cells. (D) Cell-type-specific expression levels of ACE2 and TMPRSS2 across 14 cell types in human bronchial epithelial cells. (E) Cell-type-specific expression levels of 7 highlighted inflammatory genes (A and B) show elevated expression levels in secretory 3 cells compared to other cell types. (F) UMAP visualization for lung cells. (G) Cell-type-specific expression levels of ACE2 and TMPRSS2 across 9 cell types in lung cells. (H) Cell-type-specific expression levels of 4 highlighted inflammatory genes (A and B) show elevated expression levels in alveolar type II (AT2) cells compared to other cell types. The data underlying this figure can be found in S4 Data. Single-cell data were retrieved from https://data.mendeley.com/datasets/7r2cwbw44m/1. See S1 Table for more details of the datasets. FDR, false discovery rate; MvsC, mild versus control; SvsC, severe versus control; SvsM, severe versus mild.
We further examined the gene expression data from 2 asthma cohorts from the Severe Asthma Research Program [66,67]. Utilizing 2 bulk RNA-Seq datasets (GSE63142 and GSE130499) of asthma patients compared to healthy controls, we identified that IRAK3 and ADRB2 had significantly elevated expression (false discovery rate [FDR] < 0.05) in asthma patients. Both genes also have significantly elevated expression in SARS-CoV-2-infected human bronchial epithelial cells (Fig 5B). IRAK-M, encoded by IRAK3, regulates the toll-like receptor/interleukin-1 receptor pathway and NF-κB pathway, and IRAK3 was identified as an asthma susceptibility gene [68]. ADRB2 encodes the beta2-adrenergic receptor. The polymorphisms of ADRB2 (p.Arg16Gly and p.Gln27Glu) increase the risk of asthma occurrence, and p.Gln27Glu is associated with asthma severity [69]. IRAK3 and ADRB2 also have high eigenvector centrality scores (top 5 and top 2 among these genes, respectively; Fig 5B, eigenvector centrality). Altogether, altered IRAK3 and ADRB2 expression may explain relationships between COVID-19 and asthma, though these findings require experimental and clinical validation in patients with these disorders.
To understand the expressions of the proteins in the asthma–COVID-19 network across different cell types, especially cells that express ACE2, we analyzed the single-cell RNA-Seq data from bronchial epithelium (Fig 5C) and lung (Fig 5F) [14]. Consistent with previous studies, ACE2 and TMPRSS2 have higher expression in a subtype of the secretory cells (secretory 3 cells) compared to other bronchial epithelial cell types (Figs 5D and S8B–S8F). In the lung, ACE2 and TMPRSS2 have relatively higher expression in AT2 cells (Fig 5G and S8G–S8K). We further examined the expression of the genes in the asthma–COVID-19 network (Fig 5A) in these cell types (S9 Fig). Several genes were also more highly expressed in secretory 3 cells (Fig 5E; ALOX5, IRAK3, ADRB2, TNIP1, BID, CXCL5, and NFKBIZ) and in AT2 cells (Fig 5H; CFTR, NFKBIA, NFKBIZ, and TNC), than in other cell types. IRAK3 and ADRB2 are among the 6 overlapped genes, potentially implicating the roles of IRAK3 and ADRB2 in COVID-19-associated asthma at the single-cell level as well.
Immune pathobiology shared by COVID-19 and IBD
It has been shown, using confocal and electron microscopy, that human small intestine is an additional SARS-CoV-2 target organ [70]. Diarrhea is now well described as an occasional presenting symptom of COVID-19 [71]. Our network proximity analysis showed a significant association of COVID-19 and IBD across all 5 SARS-CoV-2 datasets (Fig 4A). In addition, patients with severe COVID-19 had higher risks of abdominal pain and diarrhea (Figs 6A and S10). To understand these associations at the cellular level, we integrated network analysis and single-cell RNA-Seq analysis using publicly available data [72]. As shown in Fig 6B (S5 Data), although only 1 IBD-associated protein, HEATR3, was found to be the target of the SARS-CoV-2 protein Orf7a, other IBD-associated proteins showed an enriched number of connections to the SARS-CoV-2 target proteins (Fig 6I, degree enrichment).
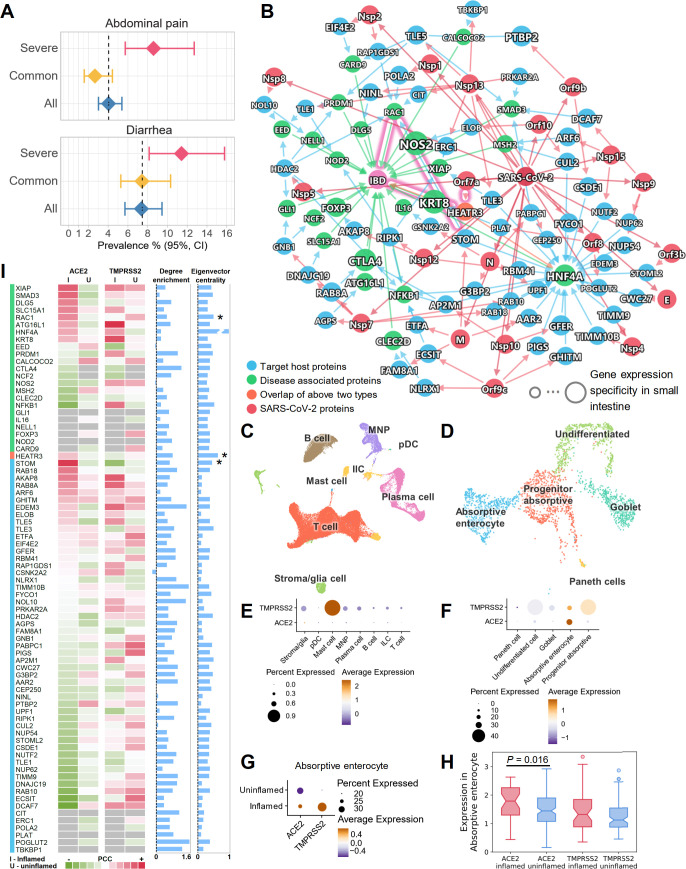
Fig 6. Inflammatory molecular profile between inflammatory bowel disease (IBD) and COVID-19.
(A) Patients with severe COVID-19 have higher risks of abdominal pain and diarrhea by meta-analysis. (B) A highlighted subnetwork between the IBD-associated genes, the SARS-CoV-2 virus proteins, and virus target proteins under the human interactome network model. (C) UMAP visualization of non-epithelial cells from the ileal tissues of patients with Crohn disease. (D) UMAP visualization of epithelial cells from the ileal tissues of patients with Crohn disease. (E) Cell-type-specific expression of ACE2 and TMPRSS2 in non-epithelial cells (C). (F) Cell-type-specific expression of ACE2 and TMPRSS2 in epithelial cells (D). (G) The co-expression of ACE2 and TMPRSS2 is elevated in absorptive enterocytes of inflamed ileal tissues compared to uninflamed tissues in patients with Crohn disease. (H) Box plot showing the expression of ACE2 and TMPRSS2 in absorptive enterocytes expressing ACE2 and TMPRSS2, respectively. (I) Co-expression analysis for the genes in the subnetwork with ACE2 and TMPRSS2. Heatmap shows the Pearson correlation coefficients (PCCs) of ACE2 and TMPRSS2 with other genes (labeled in [B]) in the absorptive enterocytes. Blue bars show the degree enrichment of the genes in the subnetwork compared to a random network of the same size (left) and the eigenvector centrality (right). Asterisks indicate that these genes may play important roles in COVID-19-associated IBD. The data underlying this figure can be found in S5 Data. Single-cell data were retrieved from the NCBI GEO database using the accession number GSE134809. See S1 Table for more details of the datasets.
Using single-cell data from the ileum (distal small bowel) in Crohn disease patients [72], we found that ACE2 and TMPRSS2 had low to undetectable expression in the non-epithelial cells (Figs 6C, 6E, and S11A–S11E). However, they showed higher expression levels in the epithelial cells, especially absorptive enterocytes (Figs 6D, 6F and S11F–S11J). We further found that both ACE2 and TMPRSS2 had elevated expression levels in inflamed cells compared to uninflamed cells in the absorptive enterocytes (Figs 6G, S11K and S11L). The Pearson correlation coefficient (PCC) of ACE2 and TMPRSS2 was also increased in the inflamed cells compared to uninflamed cells (PCC = 0.165 versus PCC = −0.006) in the absorptive enterocytes. In absorptive enterocytes expressing ACE2, the expression of ACE2 was significantly increased (Fig 6H; P = 0.016) in the inflamed ileal tissues of Crohn disease patients compared to uninflamed tissues. These observations prompted us to investigate the co-expression of the network genes in the absorptive enterocytes (Fig 6I). Several genes showed elevated co-expression with ACE2 in inflamed cells, such as XIAP, SMAD3, DLG5, SLC15A1, RAC1, STOM, RAB18, and AKAP8.
We next turned to highlight 2 potential associations between COVID-19 and IBD. First, SARS-CoV-2 protein Orf7a can directly interact with HEATR3 (top 2 eigenvector centrality; Fig 6I), whose variant was shown to be associated with increased risk of IBD by genome-wide association study [73]. Second, SARS-CoV-2 infection may impact RAC1 (top 4 eigenvector centrality; Fig 6I) signal transduction pathways. RAC proteins play important roles in many inflammatory pathways, and their dysregulation can be pathogenic. Increased RAC1 expression by single nucleotide polymorphisms promotes an inflammatory response in the colon [74]. Mercaptopurine, an effective treatment for IBD, was found to lower RAC1 expression in IBD patients [75]. Since our results show that RAC1 and ACE2 had higher co-expression in inflamed enterocytes (Fig 6I), it is highly possible that these inflamed cells are more susceptible to SARS-CoV-2 infection, and that the infection could lead to an altered RAC1 expression level through PPIs with virus target proteins STOM, HDAC2, POLA2, CIT, and RAP1GDS1 (Fig 6B). Notably STOM (top 12 eigenvector centrality) was also more co-expressed with ACE2 in inflamed cells compared to uninflamed cells (Fig 6I).
Network-based drug repurposing for COVID-19
Knowledge of the complex interplays between SARS-CoV-2 targets and human diseases indicate possibilities of drug repurposing, as the drugs that target other diseases could potentially target SARS-CoV-2 through the shared functional PPI networks [23]. In addition, drug repurposing efforts may also reveal unrecognized biological connections between originally approved indications/diseases and COVID-19. For example, the aforementioned anti-inflammatory drugs tocilizumab and sarilumab that are now being tested for COVID-19 were originally used for rheumatoid arthritis. Although not significant, our network proximity results show that rheumatoid arthritis has small network proximities (negative Z scores) across all 5 SARS-CoV-2 datasets (Fig 4A). Another drug, the thiopurine mercaptopurine, which has been used to treat IBD [76], was one of the top repurposable drugs for COVID-19 proposed in our previous work [30].
Therefore, we next performed network-based drug repurposing modeling using the existing knowledge of the drug–target network and the global map of the SARS-CoV-2 interactome built in this study. The basis for the network-based drug repurposing methodologies is the observation that for a drug with multiple targets to be effective against a disease, its target proteins should be within or in the immediate vicinity of the corresponding subnetwork of the disease in the human interactome, as we have demonstrated in multiple diseases previously [31,32]. Using our state-of-the-art network proximity framework, we measured the “closest” proximities of nearly 3,000 drugs and the 4 SARS-CoV-2 host gene/protein profiles (SARS2-DEG, SARS2-DEP, HCoV-PPI, and SARS2-PPI; S6 Table). Additionally, we performed gene set enrichment analysis (GSEA) using 5 gene/protein expression datasets: 1 SARS-CoV-2 transcriptomics dataset, 1 SARS-CoV-2 proteomics dataset, 1 MERS-CoV dataset, and 2 SARS-CoV-1 transcriptomics datasets. GSEA was used to evaluate the individual drugs for their potential to reverse the expression at the transcriptome or proteome level altered by the viruses [35].
We next prioritized the drug candidates using subject matter expertise based on a combination of factors: (1) strength of the network-based and bioinformatics-based predictions (a higher network proximity score and significant GSEA score; Fig 7; S6 Data), (2) literature-reported antiviral activities or ongoing clinical trial information, (3) availability of sufficient patient data for meaningful evaluation (exclusion of infrequently used medications) from our COVID-19 registry database, and (4) well-defined antiviral mechanisms of action (such as anti-inflammatory or immune modulators).
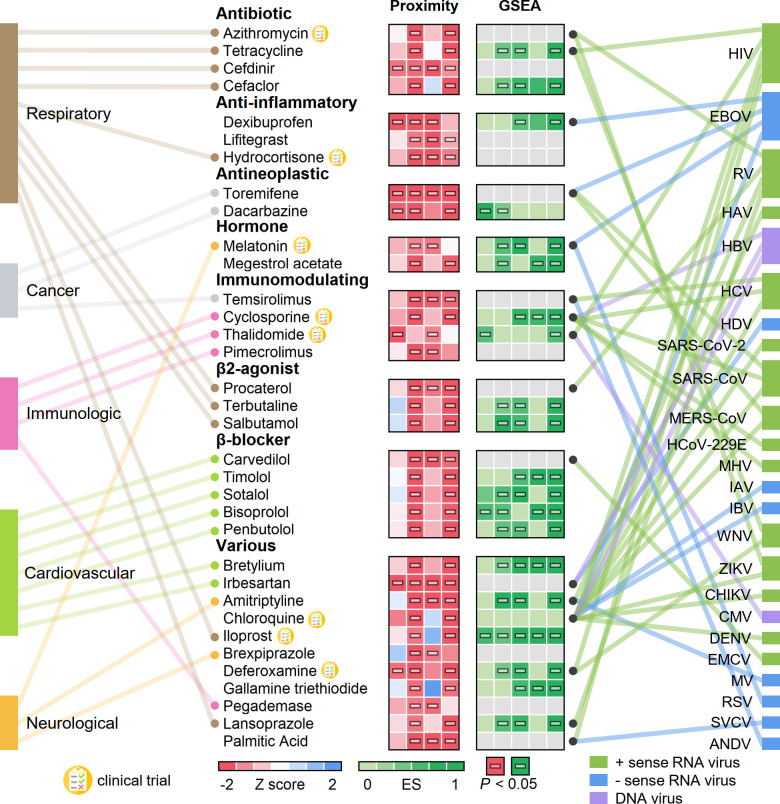
Fig 7. Network-based prediction of drug repurposing for COVID-19.
Thirty-four drugs from the top predicted list are highlighted, with the disease category they are approved for by the US Food and Drug Administration. We highlight 3 types of evidence: (1) network proximities of drugs’ targets across the 4 SARS-CoV-2 datasets (SARS2-DEG, SARS2-DEP, HCoV-PPI, and SARS2-PPI) in the human interactome, (2) gene set enrichment analysis (GSEA) scores across 5 coronavirus transcriptomics and proteomics datasets, and (3) literature-reported antivirus profiles. GSEA scores shown in grey indicate that these drugs cannot be evaluated due to the lack of data. Eight drugs that are currently being or have been tested in COVID-19 clinical trials are highlighted. Horizontal bars in boxes indicate P < 0.05. The data underlying this figure can be found in S6 Data. ANDV, Andes virus; CHIKV, chikungunya virus; CMV, cytomegalovirus; DENV, dengue virus; EBOV, Zaire ebolavirus; EMCV, encephalomyocarditis virus; ES, enrichment score; HAV, hepatitis A virus; HBV, hepatitis B virus; HCoV-229E, human coronavirus 229E; HCV, hepatitis C virus; HDV, hepatitis D virus; HIV, human immunodeficiency virus; IAV, influenza A virus; IBV, influenza B virus; MERS-CoV, Middle East respiratory syndrome coronavirus; MHV, mouse hepatitis virus; MV, measles virus; RSV, respiratory syncytial virus; RV, rhinovirus; SARS-CoV, severe acute respiratory syndrome coronavirus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SVCV, spring viremia of carp virus; WNV, West Nile virus; ZIKV, Zika virus.
In total, we computationally identified 34 drugs that were associated (Z < −1.5 and P < 0.05, permutation test) with the SARS-CoV-2 datasets (SARS2-DEG, SARS2-DEP, HCoV-PPI, and SARS2-PPI) using the above criteria. These drugs were significantly proximal to 2 or more SARS-CoV-2 host protein sets (Fig 7; S6 Data). We manually curated their reported antiviral profiles. The disease categories that these drugs have been used to treat are also shown in Fig 7. Ten drugs have been used to treat respiratory-related diseases, and the most common categories for these drugs are antibiotic and β2 agonist. The next most common disease category is cardiovascular diseases, for which 7 drugs were predicted. Among the 34 drugs, 3 drugs achieved significant network proximity with all 4 SARS-CoV-2 datasets investigated here. These drugs are (1) the antibiotic drug cefdinir, which is a cephalosporin for the treatment of bacterial infections [77]; (2) the antineoplastic drug toremifene, a selective estrogen receptor modulator that shows striking activities in blocking various viral infections at low micromolar levels, including Ebola virus [78] (50% inhibitive concentration [IC50] = approximately 1 μM), MERS-CoV [79] (50% effective concentration [EC50] = 12.9 μM), SARS-CoV-1 [80] (EC50 = 11.97 μM), and SARS-CoV-2 [81] (IC50 = 3.58 μM); and (3) the antihypertensive drug irbesartan, an angiotensin II receptor blocker (ARB) that can inhibit viral entry by inhibiting sodium/bile acid cotransporters [82].
Evidence from the COVID-19 registry data that supports the predicted drug repurposing strategies
We next evaluated drug–outcome relationships using a large-scale patient dataset from the Cleveland Clinic COVID-19 patient registry (see Materials and Methods—Patient data validation of the network-identified drugs using a COVID-19 registry). Applying subject matter expertise to the 34 repurposed drugs resulted in identifying melatonin, a physiologic hormone common to many living organisms, and carvedilol, approved for both hypertension and heart failure. A retrospective COVID-19 cohort analysis was conducted to validate the potential prevention effect of melatonin and carvedilol (Fig 8A and 8B). Among a total of 26,779 patients tested for COVID-19 in the Cleveland Clinic Health System in Ohio and Florida, 8,274 patients were diagnosed as SARS-CoV-2 positive confirmed by reverse transcription–polymerase chain reaction (RT-PCR) between March 8 and July 27, 2020 (Table 1).
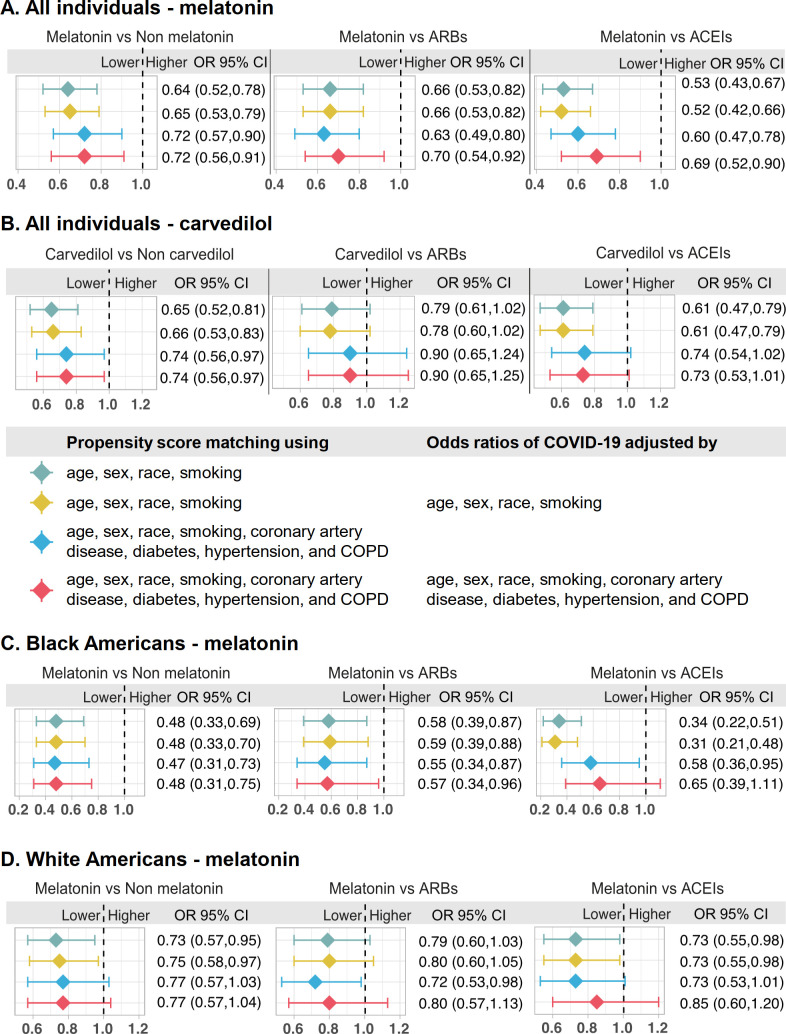
Fig 8. Patient-based validation of drug repurposing for COVID-19.
Validation for (A) melatonin and (B) carvedilol using the whole COVID-19 registry (all combined population). Validation for melatonin (C) in the black American (African American) subgroup and (D) in the white American subgroup. Patient groups were matched using propensity score matching. Four models were evaluated: (1) model 1 was matched using age, sex, race, and smoking without adjustment for the odds ratio; (2) model 2 was matched using age, sex, race, and smoking, and the odds ratio of COVID-19 was adjusted by age, sex, race, and smoking; (3) model 3 was matched using age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD without adjustment for the odds ratio; and (4) model 4 was matched using age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD, and the odds ratio of COVID-19 was adjusted by age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD. These models were adjusted for different variables using the propensity score matching approach (see Materials and Methods—Patient data validation of the network-identified drugs using a COVID-19 registry). ACEI, angiotensin-converting enzyme inhibitor; ARB, angiotensin II receptor blocker; COPD, chronic obstructive pulmonary disease; OR, odds ratio.
Table 1. Baseline characteristics of the melatonin and carvedilol use groups from the COVID-19 registry.
| Use group and characteristic | All individuals | SARS-CoV-2-positive patients | ||||
|---|---|---|---|---|---|---|
| Drug not used | Drug used | P value | Drug not used | Drug used | P value | |
| Melatonin | ||||||
| Total patients | 25,724 | 1,055 | 8,052 | 222 | ||
| Age | 48.75 ± 20.58 | 63.42 ± 19.77 | <0.001 | 49.44 ± 20.54 | 67.75 ± 18.92 | <0.001 |
| Sex (female) | 15,274 (59.4) | 549 (52.0) | <0.001 | 4,528 (56.2) | 114 (51.4) | 0.168 |
| Race other | 1,376 (5.3) | 48 (4.5) | 0.287 | 462 (5.7) | 3 (1.4) | 0.008 |
| Race black | 5,848 (22.7) | 273 (25.9) | 0.019 | 2,721 (33.8) | 70 (31.5) | 0.528 |
| Race white | 16,182 (62.9) | 695 (65.9) | 0.054 | 4,290 (53.3) | 142 (64.0) | 0.002 |
| Smoking | 3,050 (13.8) | 165 (16.1) | 0.042 | 603 (9.0) | 19 (8.8) | 1 |
| COPD & emphysema | 1,768 (10.1) | 257 (29.0) | <0.001 | 441 (10.4) | 45 (26.2) | <0.001 |
| Diabetes | 4,681 (25.4) | 417 (44.9) | <0.001 | 1,453 (31.4) | 78 (42.9) | 0.001 |
| Hypertension | 9,810 (49.7) | 764 (76.4) | <0.001 | 3,148 (59.1) | 170 (83.3) | <0.001 |
| Coronary artery disease | 2,699 (15.1) | 330 (36.3) | <0.001 | 744 (17.1) | 59 (34.7) | <0.001 |
| Asthma | 5,075 (27.5) | 265 (29.5) | 0.211 | 1,243 (26.9) | 38 (22.2) | 0.203 |
| ACEIs | 1,873 (7.3) | 208 (19.7) | <0.001 | 607 (7.5) | 48 (21.6) | <0.001 |
| ARBs | 1,371 (5.3) | 148 (14.0) | <0.001 | 447 (5.6) | 21 (9.5) | 0.019 |
| Carvedilol | ||||||
| Total patients | 25,994 | 785 | 8,070 | 204 | ||
| Age | 48.81 ± 20.67 | 66.38 ± 15.40 | <0.001 | 49.47 ± 20.65 | 68.28 ± 13.08 | <0.001 |
| Sex (female) | 15,485 (59.6) | 338 (43.1) | <0.001 | 4,550 (56.4) | 92 (45.1) | 0.002 |
| Race other | 1,385 (5.3) | 39 (5.0) | 0.717 | 460 (5.7) | 5 (2.5) | 0.066 |
| Race black | 5,841 (22.5) | 280 (35.7) | <0.001 | 2,704 (33.5) | 87 (42.6) | 0.008 |
| Race white | 16,432 (63.2) | 445 (56.7) | <0.001 | 4,328 (53.6) | 104 (51.0) | 0.497 |
| Smoking | 3,080 (13.8) | 135 (17.8) | 0.002 | 604 (9.0) | 18 (9.1) | 1 |
| COPD & emphysema | 1,862 (10.5) | 163 (26.8) | <0.001 | 447 (10.5) | 39 (27.1) | <0.001 |
| Diabetes | 4,656 (25.0) | 442 (63.4) | <0.001 | 1,422 (30.6) | 109 (66.9) | <0.001 |
| Hypertension | 9,837 (49.2) | 737 (95.7) | <0.001 | 3,124 (58.6) | 194 (97.5) | <0.001 |
| Coronary artery disease | 2,615 (14.5) | 414 (60.7) | <0.001 | 704 (16.2) | 99 (61.5) | <0.001 |
| Asthma | 5,184 (27.7) | 156 (25.1) | 0.169 | 1,243 (26.7) | 38 (26.6) | 1 |
| ACEIs | 1,866 (7.2) | 215 (27.4) | <0.001 | 597 (7.4) | 58 (28.4) | <0.001 |
| ARBs | 1,324 (5.1) | 195 (24.8) | <0.001 | 418 (5.2) | 50 (24.5) | <0.001 |
Age is shown as mean ± standard deviation. All other characteristics are shown as number of cases (percentage). Percentages were calculated using the total number of patients with known information for each variable. P values were calculated by 2-sided t test for age and Fisher’s exact test for other variables.
ACEI, angiotensin-converting enzyme inhibitor; ARB, angiotensin II receptor blocker; COPD, chronic obstructive pulmonary disease.
We found that melatonin usage was associated with a 28% reduced likelihood of a positive laboratory test result for SARS-CoV-2 (odds ratio [OR] = 0.72, 95% CI 0.56–0.91; Fig 8A) after adjusting for age, sex, race, smoking history, and various disease comorbidities (diabetes, hypertension, coronary artery disease, and COPD) using a propensity score (PS) matching method. Angiotensin-converting enzyme inhibitors (ACEIs) and ARBs are 2 common types of drugs for treatment of hypertension. A recent study showed that inpatient use of ACEI/ARB was associated with lower risk of all-cause mortality compared with ACEI/ARB non-use among hospitalized COVID-19 patients with hypertension [83]. Several recent studies also showed that there was no association of ARBs and ACEIs with the risk of SARS-CoV-2 infection [84–86]. We further performed an observational study for 3 cohorts using user active comparator design with ARBs and ACEIs used as comparators and PS adjustment for confounding factors as described in our previous study [32]. We found that melatonin usage was significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 compared to use of ARBs (OR = 0.70, 95% CI 0.54–0.92) and ACEIs (OR = 0.69, 95% CI 0.52–0.90) after adjusting for age, sex, race, smoking history, and various disease comorbidities (Fig 8A). Altogether, network-based prediction (Fig 7) and multiple observational analyses (Fig 8A) suggest that melatonin usage offers a potential prevention and treatment strategy for COVID-19; yet, randomized controlled clinical trials are urgently needed to test meaningfully the effect of melatonin for COVID-19.
We found that carvedilol use was significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 (OR = 0.74, 95% CI 0.56–0.97) after adjusting for age, sex, race, smoking history, and various disease comorbidities. Yet, carvedilol did not show a significant advantage compared to ARBs (OR = 0.90, 95% CI 0.65–1.25) or ACEIs (OR = 0.73, 95% CI 0.53–1.01).
We therefore tested whether a clinically meaningful effect of melatonin and carvedilol can be observed in different subgroups of patients. To be specific, we generated 5 different subgroups: asthma patients, hypertension patients, diabetes patients, black Americans (African Americans), and white Americans. We found that melatonin was significantly associated with a 52% reduced likelihood of a positive laboratory test result for SARS-CoV-2 in black Americans (OR = 0.48, 95% CI 0.31–0.75; Fig 8C) after adjusting for age, sex, race, smoking, and various disease comorbidities, which is stronger than the association in white Americans (OR = 0.77, 95% CI 0.57–1.04; Fig 8D). In addition, in black Americans, melatonin usage was significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 compared to ARB usage (OR = 0.57, 95% CI 0.34–0.96; Fig 8C), while there was no significant difference compared to ACEI usage (OR = 0.65, 95% CI 0.39–1.11; Fig 8C). Yet, melatonin usage was not significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 compared to use of ARBs (OR = 0.80, 95% CI 0.57–1.13; Fig 8D) and ACEIs (OR = 0.85, 95% CI 0.60–1.20; Fig 8D) in white Americans. Among the 3 comorbid disease subgroup analyses, melatonin usage was significantly associated with a reduced risk of SARS-CoV-2 positive test in diabetes patients only (OR = 0.52, 95% CI 0.36–0.75; S12B Fig); there was no significant association for asthma (OR = 0.61, 95% CI 0.36–1.06; S12A Fig) or hypertension (OR = 0.80, 95% CI 0.61–1.05; S12C Fig) patients. Carvedilol usage was not significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 among the 5 subgroups after adjusting for age, sex, race, smoking, and various disease comorbidities (S12 and S13 Figs). Thus, further observations using large-scale independent cohorts to test the meaningful effect of carvedilol in reducing risk of COVID-19 are highly needed.
Discussion
Recent studies indicated that SARS-CoV-2 infection was detected in multiple organs in addition to lungs, including heart, pharynx, liver, kidneys, brain, and intestine [70,87]. SARS-CoV-2 RNA was also found in patient stool [88]. Therefore, investigation of how SARS-CoV-2 associates with other diseases could help reveal and understand its impact on systems and organs in addition to lungs. In this study, we systematically evaluated 64 diseases across 6 categories for their potential manifestations with COVID-19. We started with assembling and characterizing 5 SARS-CoV-2 datasets representing different cellular event levels including transcriptome, proteome, and interactome. Using state-of-the-art network proximity measurement, we identified broad disease manifestations (such as autoimmune, neurological, and pulmonary; Fig 4A) associated with COVID-19. Although the number of genes associated with each disease is different (S4 Table), we did not notice any significant bias in the network proximity Z scores by different number of genes (S14 Fig). Retrospective meta-analyses using the clinical data of 4,973 patients across 34 studies confirmed our network-based findings.
We further investigated the molecular determinants of the association of COVID-19 and the comorbidities by multimodal analyses of large-scale bulk and single-cell transcriptomic profiles, metabolomic data, and the PPI networks. We identified the cell types that have the highest expression levels of ACE2 and TMPRSS2: the lung AT2 cells, secretory bronchial epithelial cells, and absorptive enterocytes in the ileum. We examined the expression of asthma- and IBD-associated genes in their relevant cell types. Combining these findings with the results of differential expression analysis, network analysis, and differential metabolites, we identified several key pathogenic pathways for asthma (including IRAK3 and ADRB2) that can be altered by the viral infection.
For asthma, our network-based findings suggest several possible shared pathobiological pathways associated with COVID-19. First, SARS-CoV-2 infection might alter the expression of several key inflammatory genes: IRAK3, which is associated with asthma [68,89,90]; ADRB2, which is an essential genetic factor for asthma [69,91]; and NFKBIA, which shows critical transcriptional responses in childhood asthma [92]. These genes show high expression in secretory 3 and AT2 cell types, suggesting a higher susceptibility to be impacted by SARS-CoV-2 infection through ACE2. SARS-CoV-2 increased the expression of IRAK3 and ADRB2, which lead to a higher risk of asthma (Fig 5B). Second, decreased levels of L-arginine and L-citrulline were found in SARS-CoV-2-infected patients [62], while it has been shown that higher levels of these metabolites are protective against asthma [61]. L-arginine can be converted to nitric oxide by the nitric oxide synthases, and it was shown that nitric oxide protects against viral infection through multiple potential mechanisms [93].
Our network proximity results show a strong connection between IBD and COVID-19 (Fig 4A). We have also shown that IBD-related pathways can potentially be affected by SARS-CoV-2 infection (Fig 6B). However, we should also note that the meta-analysis does not show a significant risk ratio of digestive system disease in COVID-19 patients (Fig 3B), while 2 specific symptoms, abdominal pain and diarrhea, showed increased risks in patients with severe COVID-19 (Fig 6A). Digestive system disease covers a broad range of gastrointestinal diseases. Future studies are needed to reveal and validate the associations of COVID-19 and individual gastrointestinal diseases.
Finally, we computationally prioritized nearly 3,000 FDA-approved/investigational drugs for their potential anti-SARS-CoV-2 effects using network proximity measurement and GSEA analysis. A list of 34 repurposable drugs with their reported antiviral profiles are highlighted, among which 8 drugs are currently in ongoing COVID-19 clinical trials (Fig 7). We further explored drug–disease outcome relationships for melatonin using a large-scale COVID-19 patient registry database. We found that among individuals who received testing for SARS-CoV-2, melatonin usage was associated with a 28% and 52% reduced likelihood of a positive laboratory test result for SARS-CoV-2 in the all combined population (Fig 8A) and black Americans (Fig 8C), respectively, after adjusting for age, sex, race, smoking, and various disease comorbidities. Using user active comparator design, we further found that melatonin usage was associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 compared to use of ARBs and ACEIs as well. Exogenous melatonin may be of benefit in older patients with COVID-19, given the aging-related reduction of endogenous melatonin and the greater vulnerability of older individuals to mortality from SARS-CoV-2 [94], the latter potentially due to declining immunity, i.e., immunosenescence [95]. Moreover, melatonin suppresses NLRP3 inflammasome activation induced by cigarette smoking and attenuates pulmonary inflammation [96], not only via reduction of NF-κB p65 and tumor necrosis factor-α (TNF-α) expression, but also via increase in anti-inflammatory cytokines such as IL-10 or IL-6, which can also have anti-inflammatory effects [97,98]. Thus, large-scale observational studies and randomized controlled trials are needed to validate the clinical benefit of melatonin for patients with COVID-19. It would be important, however, that the trials be designed with the understanding of the mechanism of the drug to be repurposed. For example, it would be obvious that drugs that decrease viral entry, e.g., part of melatonin’s action, would be beneficial in preventing infection or very early in the COVID-19 course, but may be inconsequential when utilized in severe or end-stage infection. Several randomized controlled trials are being performed to test the clinical benefits of melatonin in patients with COVID-19 (ClinicalTrials.gov NCT04409522 and NCT04353128). In addition, the Selective Estrogen Modulation and Melatonin in Early COVID-19 (SENTINEL) trial is underway to test the combination therapy [99] of melatonin with toremifene (an approved selective estrogen receptor modulator [100]) for patients with early and mild COVID-19 (ClinicalTrials.gov NCT04531748).
We acknowledge several potential limitations. First, although we integrated data from multiple sources to build the human interactome and the drug–target network, they are still incomplete. Second, this study relied on the SARS-CoV-2 target host gene/protein datasets, and their quality and literature bias may influence the performance of our network analysis. The genes in these datasets can differ significantly. DEGs identified from transcriptomics profiles can be very different from DEPs from proteomics data by multiple factors, which may influence the results of network proximity analysis as well. These datasets were from different cell types, and the COVID-19-relevant cell types may not be representative. PPIs and DEGs/proteins in different cell lines or tissues may contain false positives as well. Two recent studies suggested that chloroquine or hydroxychloroquine showed ideal antiviral activities in African green monkey kidney cells (VeroE6) but not in a model of reconstituted human airway epithelium [101] or the TMPRSS2-positive lung cell line Calu-3 [102]. These studies showed that cell lines mimicking important aspects of respiratory epithelial cells should be used when analyzing the antiviral activity of drugs targeting host cell functions. In the original SARS2-PPI dataset based on the VeroE6 cell line, a key PPI for ACE2–spike protein was absent. We posited that combining transcriptomics profiles and proteomics data derived from diverse COVID-19-relevant cell lines or tissues may provide complementary molecular information to overcome the high disease heterogeneities of COVID-19 [103]. In addition, through creating a pan-coronavirus PPI network by combining HCoV-PPI and SARS2-PPI, we aimed to identify broad-spectrum antiviral medications for SARS-CoV-2—and other emerging coronaviruses, if broadly applied—with our network medicine framework.
Third, our method can only apply to diseases with well-characterized genetic information and may not be applicable for diseases that lack such information, such as rare diseases (i.e., cerebral palsy or mental conditions). Potential literature bias of disease-associated genes and the human interactome may also influence our findings. For example, well-studied genes associated with both COVID-19 and other diseases may explain the similarity of COVID-19 with other diseases, while the understudied genes associated with both diseases may not be uncovered. Fourth, the patient data analysis is retrospective, may have selection bias, and is limited for commonly used drugs due to patient data availability. Dose and period of the medications were also missing in our current COVID-19 patient registry database. Although we performed multiple types of PS matching, residual confounding is possible despite high-dimensional covariate adjustment. Carvedilol use did not show a significant advantage compared to use of ARBs (OR = 0.90, 95% CI 0.65–1.25) or ACEIs (OR = 0.73, 95% CI 0.53–1.01) after adjusting for age, sex, race, smoking history, and multiple disease comorbidities (Fig 8B). In addition, carvedilol was not significantly associated with a reduced likelihood of a positive laboratory test result for SARS-CoV-2 among 5 subgroup analyses (S12 and S13 Figs). There are 2 possible explanations: (1) the small number of patients using carvedilol (Table 1) may yield insufficient statistical power for the current observational study or (2) carvedilol may only reduce the likelihood of a positive laboratory test result for SARS-CoV-2 for patients having existing health conditions, such as hypertension. Replication of the associations and causal inference using large-scale independent cohorts may rule out treatment effect heterogeneity and possible confounding further. Finally, although we made the intriguing observation that the use of melatonin was much less prevalent in individuals testing positive for SARS-CoV-2, we recognize that many asymptomatic or minimally symptomatic persons with the virus were not tested and, therefore, their use of melatonin was not evaluable; melatonin use in this latter group might also have been high. It should therefore be noted that all drugs we identified as therapeutic candidates in this study must be validated using experimental assays and randomized clinical trials before they can be recommended for use in patients with COVID-19.
Recent studies have suggested that COVID-19 is a systemic disease that has impacts on multiple cell types, tissues, and organs [104]. Our network methodology could potentially be improved by using tissue-specific genes for the diseases and incorporating directionality of the gene/protein biological effects. We recomputed network proximity using only genes that have a tissue specificity ≥ 0 in the associated disease: We found overall consistent results (S15 Fig) compared to lung-specific network analysis (Fig 4A). For example, IBD achieved significant proximities across all 5 SARS-CoV-2 gene/protein datasets in Fig 4A. When using genes with tissue specificity ≥ 0, IBD showed the same results in small intestine, but not in colon (S15 and S16 Figs). Type 2 diabetes showed significant network proximities across all 5 SARS-CoV-2 gene/protein datasets using pancreas-specific genes (S15 Fig), which is consistent with recent clinical observations [105]. Yet, the GTEx database used in this study was from healthy tissues, which may not represent the gene expression state in the disease condition. In addition, the disease-associated genes used this study were based on somatic or germline genetic evidence. For example, we did not observe significant association of COVID-19 with cancer. One possible explanation is that cancer is a more somatic-mutation-driven, chronic disease than COVID-19, which involves an acute immune response and inflammation-driven heterogeneous disease.
We also experimented with incorporating the directionality of the gene expression using the 2 asthma expression datasets and SARS2-DEG, since the direction of the DEGs are available in those 2 datasets. We found more significant network proximities and smaller Z scores between asthma and COVID-19 when using the up- or down-expressed genes separately (S17 Fig), suggesting that incorporating the directionalities of genes/proteins may improve the performance of network analysis. A previous study has shown that integration of the directionality of the human interactome didn’t change the results of network proximity measurement [49]. Owing to the lack of a systematic human interactome with well-documented directionalities, and without comprehensive information about whether a viral protein activates or inhibits a host protein, we didn’t test the influence of the directionality of proteins and the human interactomes on our findings in a systematic way. Therefore, future work is needed to explore well-documented directionalities in human interactome network analysis that integrates precise perturbation effects of disease-causing variants and viral proteins.
In conclusion, our study provides a powerful, integrative network medicine strategy for advancing understanding of COVID-19-associated comorbidities and facilitating the identification of drug candidates for COVID-19. This approach also promises to address the translational gap between genomic studies and clinical outcomes, which poses a significant problem when rapid development of effective therapeutic interventions is critical during a pandemic. From a translational perspective, if broadly applied, the network medicine tools applied here could prove helpful in developing effective treatment strategies for other complex human diseases as well, including other emerging infectious diseases.
Materials and methods
A list of the sources of all the datasets used in this study can be found in S1 Table.
Building the datasets of SARS-CoV-2 target host genes/proteins
We assembled 4 SARS-CoV-2 datasets of target host genes/proteins: (1) 246 DEGs in human bronchial epithelial cells infected with SARS-CoV-2 [21] (GSE147507), denoted as SARS2-DEG; (2) 293 DEPs in human Caco-2 cells infected with SARS-CoV-2 [22], denoted as SARS2-DEP; (3) 134 strong literature-evidence-based pan-human coronavirus target host proteins from our recent study [30] with 15 newly curated proteins, denoted as HCoV-PPI; and (4) 332 proteins involved in PPIs with 26 SARS-CoV-2 viral proteins identified by affinity purification–mass spectrometry (AP-MS) [8], denoted as SARS2-PPI. Finally, due to the interactome nature of HCoV-PPI and SARS2-PPI, we combined these datasets as the fifth SARS-CoV-2 dataset, which has 460 proteins and is denoted as PanCoV-PPI. Details of these datasets can be found in S2 Table.
SARS2-DEG
In the original study, the primary human bronchial epithelial cells were infected with SARS-CoV-2 for 24 hours. The transcriptome profiles of infected (3 replicates) and uninfected cells (3 replicates) were characterized, and the fold change (FC) and FDR for each gene were calculated by DESeq2 and provided in the original study. We applied a cutoff of |log2FC| > 0.5 and FDR < 0.05 to identify the DEGs.
SARS2-DEP
As described in the previous study [22], human Caco-2 cells were infected with SARS-CoV-2 for up to 24 hours. Proteomics assays of the infected and uninfected cells were measured at 24 hours in triplicates. We used the results at 24 hours, as the original study showed most DEPs at 24 hours. The P values were computed using 2-sided unpaired Student t tests with equal variance assumed in this study. We converted the P value to FDR using the “fdrcorrection” function in the Python package statsmodels v0.11.1 and used a cutoff of FDR < 0.05 to identify the DEPs.
Collection of 4 additional virus–host gene/protein networks
To characterize the SARS-CoV-2 datasets, we downloaded 4 virus–host gene/protein networks from previous studies for comparison: (1) 900 virus–host interactions identified by gene-trap insertional mutagenesis connecting 10 other viruses and 712 host genes [35]; (2) 2,855 virus–host interactions identified from RNAi connecting 2,443 host genes and 55 pathogens [35]; (3) 579 host proteins mediating translation of 70 innate immune-modulating viORFs [36]; and (4) 1,292 host genes identified by Co-IP+LC/MS that mediate influenza–host interactions [37]. All details for these 4 virus–host gene/protein networks are provided in S3 Table.
Building the disease gene profiles
We compiled the disease-associated gene sets from various sources. All databases were accessed on March 26, 2020.
Cancer
We defined a driver gene as a gene that had significantly enriched driver mutations based on whole-genome or whole-exome sequencing data or reported experimental data from the Cancer Gene Census [42,43] or the original publications from The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/). The pan-cancer driver genes were retrieved from the Cancer Gene Census [42,43]. Driver genes for individual cancer types were from a previous study [44].
Mendelian disease genes (MDGs)
A set of 2,272 MDGs were retrieved from the Online Mendelian Inheritance in Man (OMIM) database [106].
Orphan disease-causing mutant genes (ODMGs)
A set of 2,124 ODMGs were retrieved from a previous study [107].
Cell cycle genes
A set of 910 human cell cycle genes were downloaded from a previous study in which they were identified by a genome-wide RNAi screening [108].
Innate immune genes
A set of 1,031 human innate immune genes were collected from InnateDB [109].
Genes associated with autoimmune, pulmonary, neurological, cardiovascular, and metabolic diseases
The disease-associated genes/proteins were extracted from HGMD [45]. HGMD is a well-documented database, and we downloaded the whole database for data analysis and extraction using well-documented disease ontology terms [46]. We defined a disease-associated gene as a gene that has at least 1 disease-associated mutation in original publications provided in HGMD. The details, including the sources, number of genes, mutations associated with the disease, and terms used to identify diseases in HGMD, are provided in S4 Table.
Functional enrichment analysis
We performed Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) biological process enrichment analyses to reveal the biological relevance and functional pathways of the 5 SARS-CoV-2 datasets. All functional enrichment analyses were performed using Enrichr [110]. An overview of the virus infection-related pathways and ontology terms shared by 1 or more datasets was generated by searching for significant pathways or terms (FDR < 0.05). The enrichment analysis results for the 5 SARS-CoV-2 gene/protein sets can be found in S1–S5 Figs.
Selective pressure and evolutionary rate characterization
We calculated the dN/dS ratio [111] and the evolutionary rate ratio [112] as described in our previous study [113]. A dN/dS ratio below, equal to, or above 1 suggests purifying selection, neutral evolution, or positive Darwinian selection, respectively [114]. The evolutionary rate ratio was computed using the criterion that a ratio > 1 indicates a fast rate and a ratio < 1 indicates a slow rate [112]. The dN/dS and evolutionary rate ratios of the genes in the 5 SARS-CoV-2 datasets and 4 additional virus gene/protein sets can be found in S2 and S3 Tables.
Tissue specificity analysis
The RNA-Seq data (transcripts per million [TPM]) of 33 tissues from the GTEx V8 release (accessed on March 31, 2020; https://www.gtexportal.org/home/) were downloaded. Genes with count per million (CPM) ≥ 0.5 in over 90% of samples in a tissue were considered tissue-expressed genes, and otherwise tissue-unexpressed. To quantify the expression specificity of gene i in tissue t, we calculated the mean expression Ei and the standard deviation σi of a gene’s expression across all considered tissues. The significance of gene expression specificity in a tissue is defined as
| (1) |
Risk ratio analysis for COVID-19 patients
PubMed, Embase, and medRxiv databases were searched for publications as of April 25, 2020 (S18 Fig). The search was limited to articles in English describing the demographic and clinical features of SARS-CoV-2 cases. We used the search term (“SARS-COV-2” OR “COVID-19” OR “nCoV 19” OR “2019 novel coronavirus” OR “coronavirus disease 2019”) AND (“clinical characteristics” OR “clinical outcome” OR “comorbidities”). Only research articles were included; reviews, case reports, comments, editorials, and expert opinions were excluded. Three criteria were used to select studies from a total of 1,054 initial hits: (1) studies that had ≥20 COVID-19 patients; (2) studies that grouped the outcomes by degree of severity of COVID-19 (e.g., severe versus non-severe) according to the American Thoracic Society guidelines for community-acquired pneumonia; and (3) studies that were from different institutions. Two criteria were used for exclusion: (1) studies that focused on specific populations (e.g., only death cases, pregnant women, children, or family clusters) and (2) basic molecular biology research. Finally, 34 studies meeting these criteria were used for further analyses.
We performed random effects meta-analysis to estimate the pooled risk ratio with 95% CI of 10 comorbidities for patients with severe versus non-severe COVID-19. The Mantel–Haenszel method was used to estimate the pooled effects of results [115]. The DerSimonian–Laird method was used to estimate the variance among studies [116]. Continuous data such as IL-6 levels were transformed to mean and standard deviation first using Wan’s approach based on sample size, median, and interquartile range [117]. Next, we used the inverse variance method to estimate the pooled mean difference and estimated the variance among studies using the DerSimonian–Laird method. We estimated the pooled prevalence of 3 COVID-19 symptoms (abdominal pain, diarrhea, and dyspnea) and 1 comorbidity (COPD) in 3 COVID-19 patient groups (severe, non-severe, and all). A random intercept logistic regression model was used to estimate pooled prevalence, and a maximum-likelihood estimator was used to quantify the heterogeneity of studies [118]. The tau2 and I2 statistics were calculated for the heterogeneity among studies. We considered I2 ≤ 50% as low heterogeneity among studies, 50% < I2 ≤ 75% as moderate heterogeneity, and I2 > 75% as high heterogeneity. All meta-analyses were conducted using the meta and dmetar packages in the R v3.6.3 platform.
Building the human protein–protein interactome
A total of 18 bioinformatics and systems biology databases were assembled to build a comprehensive list of human PPIs with 5 types of experimental evidence: (1) protein complexes data identified by a robust AP-MS methodology collected from BioPlex V2.016 [119]; (2) binary PPIs tested by high-throughput yeast-two-hybrid (Y2H) systems from 2 publicly available high-quality Y2H datasets [120,121] and 1 in-house dataset [32]; (3) kinase–substrate interactions identified by literature-derived low-throughput or high-throughput experiments from Kinome NetworkX [122], Human Protein Resource Database (HPRD) [123], PhosphoNetworks [124], PhosphoSitePlus [125], DbPTM 3.0 [126], and Phospho.ELM [127]; (4) signaling networks identified by literature-derived low-throughput experiments from SignaLink 2.0 [128]; and (5) literature-curated PPIs identified by AP-MS, Y2H, literature-derived low-throughput experiments, or protein 3D structures from BioGRID [129], PINA [130], INstruct [131], MINT [132], IntAct [133], and InnateDB [109]. Inferred PPIs based on gene expression data, evolutionary analysis, and metabolic associations were excluded. Genes were mapped to their Entrez ID based on the NCBI database [134]. The official gene symbols were based on GeneCards (https://www.genecards.org/). The final human protein–protein interactome used in this study included 351,444 unique PPIs (edges or links) connecting 17,706 proteins (nodes). Detailed descriptions for building the human protein–protein interactome are provided in our previous studies [31–33,135]. An overview of the human protein–protein interactome can be found in S19 Fig.
Network proximity measure
We used the “closest” network proximity measure throughout this study. For 2 gene/protein sets A and B, their closest distance dAB was calculated as
| (2) |
where d(a,b) is the shortest distance of a and b in the human interactome. To evaluate the significance, we performed a permutation test using randomly selected proteins from the whole interactome that were representative of the 2 protein sets being evaluated in terms of their degree distributions. We then calculated the Z score as
| (3) |
where and σr were the mean and standard deviation of the permutation test. All network proximity permutation tests in this study were repeated 1,000 times.
Network-based comorbidity analysis
To reveal potential COVID-19 comorbidities, we computed the network proximity of the disease-associated proteins for each disease and the 5 SARS-CoV-2 datasets. SARS-CoV-2 target proteins with a non-negative tissue specificity in lung were used in the computation. The degree enrichment for protein i in a subnetwork was calculated as
| (4) |
where di is the degree of i in the subnetwork, n is number of nodes in the subnetwork, Di is the degree in the complete human protein interactome, and N is the total number of nodes in the interactome. The log10 ei value is reported.
We also computed the eigenvector centrality [136] of the nodes to evaluate their influence in the network topology while also considering the importance of their neighbors. A high eigenvector centrality value suggests that the node is connected to many other nodes with high eigenvector centrality scores as well. The computation was performed using Gephi 0.9.2 (https://gephi.org/).
Bulk and single-cell RNA-Seq data analysis
A list of the datasets used in this study can be found in S1 Table.
Bulk RNA-Seq datasets for asthma patients were retrieved from the NCBI GEO database (https://www.ncbi.nlm.nih.gov/geo/) using the accession numbers GSE63142 [66] and GSE130499 [67]. Differential expression of 3 comparisons—severe versus control, mild versus control, and severe versus mild—were performed using the GEO2R function (https://www.ncbi.nlm.nih.gov/geo/geo2r/) [137]. In GSE63142 [66], bronchial epithelial cells of 27 control samples, 72 mild asthma samples, and 56 severe asthma samples were obtained by bronchoscopy with endobronchial epithelial brushing. In GSE130499 [67], bronchial epithelial cells of 38 control samples, 72 mild asthma samples, and 44 severe asthma samples were available by bronchoscopy with endobronchial epithelial brushing as well. The differential expression analysis was performed by defining the groups in GEO2R first, then by selecting the 2 groups to compare. Genes with |log2FC| > 0.5 and FDR < 0.05 were considered significantly differentially expressed.
Single-cell data of normal lung and primary human bronchial epithelial cells were downloaded from https://data.mendeley.com/datasets/7r2cwbw44m/1 [14]. These datasets contain 39,778 lung cells and 17,451 bronchial epithelial cells with cell type annotated. GSE134809 [72] was downloaded from the NCBI GEO database. This dataset contains 67,050 inflamed and uninflamed cells from the ileal samples of 8 patients with Crohn disease. Qualifying cells based on the criteria from the original paper were used for the single-cell analysis. We used the cell type gene markers from a previous study [72] (CD3D, CD2, CD7, TNFRSF17, MZB1, BANK1, CD79B, CD22, MS4A1, HLA-DRB1, HLA-DQA1, LYZ, IL3RA, IRF7, GZMB, LILRA4, CLEC4C, TPSAB1, CMA1, KIT, PLVAP, VWF, LYVE1, CCL21, COL3A1, COL1A1, ACTA2, GPM6B, S100B) for the non-epithelial cells. We used markers from Zhang et al. [15] (DEFA5, REG3A, DEFA6, SOX4, CDCA7, KIAA0101, TOP2A, MKI67, HMGB2, STMN1, SPINK4, ITLN1, REG4, CLCA1, FCGBP, HMGA1, EIF3F, ETHE1, ADH1C, C1QBP, RBP2, APOB, APOC3, APOA1, APOA4) for the epithelial cells. The expression of these markers in the cells can be found in S20 and S21 Figs. All single-cell data analyses and visualizations were performed with the R package Seurat v3.1.4 [138]. “NormalizeData” was used to normalize the data. “FindIntegrationAnchors” and “IntegrateData” functions were used to integrate cells from different samples. UMAP was used as the dimension reduction method for visualization.
Building the metabolite–enzyme network
We built a comprehensive metabolite–enzyme network by assembling data from 4 commonly used metabolism databases: KEGG [139], Recon3D [140], the Human Metabolic Atlas (HMA) [141], and the Human Metabolome Database (HMDB) [142]. The metabolite–enzyme network contains 60,822 records of 6,725 reactions among 3,808 metabolites and 3,446 genes. Four types of enzyme functions were included in the network: biosynthesis, degradation, transformation, and transportation.
Building the drug–target network
To evaluate whether a drug is closely associated with SARS-CoV-2 target proteins in the human interactome, we gathered the drug–target interaction information from several databases: DrugBank database (v4.3) [143], Therapeutic Target Database (TTD) [144], PharmGKB database, ChEMBL (v20) [145], BindingDB [146], and the IUPHAR/BPS Guide to Pharmacology [147]. We included the interactions that have binding affinities Ki, Kd, IC50, or EC50 ≤ 10 μM and a unique UniProt accession number with “reviewed” status. The details for building the experimentally validated drug–target network can be found in our recent studies [31–33].
Network-based drug repurposing
We computed the closest network proximity as described before for 2,938 FDA-approved or investigational drugs and the 5 SARS-CoV-2 datasets. For prioritization, we ranked the drugs by their distance to the datasets (D < 2, network distance using the closest measure) and Z score (Z < −1.5) from the network proximity analysis. The antiviral profiles of the highlighted drugs were manually curated. COVID-19-related clinical trials were retrieved on August 28, 2020.
Gene set enrichment analysis (GSEA)
The GSEA was conducted as described in our recent work [30] as an additional source of evidence for drug repurposing. Briefly, for each drug and coronavirus target gene set, we computed an enrichment score (ES) to indicate whether the drug can reverse the effect of SARS-CoV-2 at the transcriptome or proteome level. Gene expression profiles for the drugs were retrieved from the Connectivity Map (CMAP) database [148]. Five gene sets were evaluated: (1) the DEGs in human bronchial epithelial cells infected with SARS-CoV-2 [21] (GSE147507); (2) the DEPs in human Caco-2 cells infected with SARS-CoV-2 [22]; (3 and 4) 2 transcriptome datasets of SARS-CoV-1-infected samples from patient’s peripheral blood [149] (GSE1739) and Calu-3 cells [150] (GSE33267), respectively; and (5) the DEGs in MERS-CoV-infected Calu-3 cells [151] (GSE122876).
The ES was calculated for up- and down-regulated genes separately first. The overall ES was calculated as
| (5) |
where ESup and ESdown were calculated using aup/down and bup/down as
| (6) |
| (7) |
j = 1,2,⋯,s were the genes in the gene sets sorted in ascending order by their rank in the drug profiles. V(j) indicates the rank of gene j, where 1≤V(j)≤r, with r being the number of genes from the drug profile. Then, ESup/down was set to aup/down if aup/down>bup/down, and was set to −bup/down if bup/down>aup/down. A permutation test was performed to evaluate the significance. Drugs were prioritized and selected if ES > 0 and P < 0.05.
Patient data validation of the network-identified drugs using a COVID-19 registry
We used institutional-review-board-approved COVID-19 registry data, including 26,779 individuals (8,274 SARS-CoV-2 positive) tested during March 8 to July 27, 2020, from the Cleveland Clinic Health System in Ohio and Florida. The pooled nasopharyngeal and oropharyngeal swab specimens were tested. SARS-CoV-2 positivity was confirmed by reverse transcription–polymerase chain reaction assay in the Cleveland Clinic Robert J. Tomsich Pathology and Laboratory Medicine Institute. All SARS-CoV-2 testing was authorized by the FDA under an Emergency Use Authorization and complied with the guidelines established by the Centers for Disease Control and Prevention. Data include COVID-19 test results, baseline demographic information, medications, and all recorded disease conditions. We conducted a series of retrospective case–control studies with a new user active comparator design to test the drug–outcome relationships for COVID-19. Patients were actively taking the evaluated drugs (carvedilol and melatonin) at the time of testing. Data were extracted from electronic health records (EPIC Systems) and were manually checked by a study team trained on uniform sources for the study variables. We collected and managed all patient data using REDCap electronic data capture tools. The exposures of drugs (including carvedilol and melatonin) were used as recorded in the medication list in the electronic medical records at the time of testing for SARS-CoV-2. A positive laboratory test result for SARS-CoV-2 was used as the primary outcome. PS was used to match patients to reduce various confounding factors. Four models, from less to more stringent in terms of patient matching and OR adjustment, were performed: (1) model 1 was matched using age, sex, race, and smoking without adjustment for the OR; (2) model 2 was matched using age, sex, race, and smoking, and the OR of COVID-19 was adjusted by age, sex, race, and smoking; (3) model 3 was matched using age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD without adjustment for the OR; and (4) model 4 was matched using age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD, and the OR of COVID-19 was adjusted by age, sex, race, smoking, coronary artery disease, diabetes, hypertension, and COPD. All analyses were conducted using the matchit package in the R v3.6.3 platform.
Statistical analysis and network visualization
Statistical tests were performed with the Python package SciPy v1.3.0 (https://www.scipy.org/). P < 0.05 was considered statistically significant throughout this study. Networks were visualized using Gephi 0.9.2 (https://gephi.org/).
Ethics statement
Procedures follow institutional guidelines for research on the COVID-19 registry database and were approved by the Cleveland Clinic Foundation Institutional Review Board.
Supporting information
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
In total, 246 differentially expressed genes (DEGs) in human bronchial epithelial cells infected with SARS-CoV-2 were obtained from the NCBI GEO database with the accession number GSE147507, denoted as SARS2-DEG. The data underlying this figure can be found in S7 Data.
(PDF)
In total, 293 differentially expressed proteins (DEPs) in human Caco-2 cells infected with SARS-CoV-2 were obtained from Bojkova et al. [22], denoted as SARS2-DEP. The data underlying this figure can be found in S7 Data.
(PDF)
This dataset contains 134 strong literature-evidence-based pan-human coronavirus target host proteins from Zhou et al. [30] with 15 newly curated proteins, denoted as HCoV-PPI. The data underlying this figure can be found in S7 Data.
(PDF)
This dataset contains 332 proteins involved in protein–protein interactions with 26 SARS-CoV-2 viral proteins identified by affinity purification–mass spectrometry from Gordon et al. [8], denoted as SARS2-PPI. The data underlying this figure can be found in S7 Data.
(PDF)
Due to the interactome nature of HCoV-PPI and SARS2-PPI, we combined these datasets as the fifth SARS-CoV-2 dataset, which has 460 proteins and is denoted as PanCoV-PPI. The data underlying this figure can be found in S7 Data.
(PDF)
Node degree (blue), dN/dS ratio (orange), evolutionary ratio (green), and lung expression specificity (purple) are shown for each dataset. Grey areas indicate mean ± standard deviation of 100 repeats using randomly selected genes. The data underlying this figure can be found in S8 Data.
(PDF)
(A) The risk of chronic obstructive pulmonary disease (COPD) is increased in severe COVID-19 patients. (B) Subnetwork shows the proteins potentially involved in the interaction between COPD and COVID-19. The data underlying this figure can be found in S9 Data.
(PDF)
(A) The risk of dyspnea is increased in patients with severe COVID-19. (B) UMAP visualization for human bronchial epithelial cells. (C and D) Expression levels of ACE2 across 14 cell types. (E and F) Expression levels of TMPRSS2 across 14 cell types. (G) UMAP visualization for lung cells. (H and I) Expression levels of ACE2 across 9 cell types. (J and K) Expression levels of TMPRSS2 across 9 cell types. The single-cell data with cell type annotation were retrieved from Lukassen et al. [14], which contains 39,778 lung cells and 17,451 bronchial epithelial cells.
(PDF)
The expression levels of the genes from the asthma–COVID-19 subnetwork in bronchial epithelial cells (A) and lung cells (B) are shown.
(PDF)
Abdominal pain (A) and diarrhea (B) have increased risks in patients with severe COVID-19.
(PDF)
(A) UMAP visualization of non-epithelial cells from the ileal tissues of patients with Crohn disease. (B and D) The expression of ACE2 in the non-epithelial cells in (A). (C and E) The expression of TMPRSS2 in the non-epithelial cells in (A). (F) UMAP visualization of epithelial cells from the ileal tissues of patients with Crohn disease. (G and I) The expression of ACE2 in the epithelial cells in (F). (H and J) The expression of TMPRSS2 in the epithelial cells in (F). (K and L) The expression levels of ACE2 and TMPRSS2 in inflamed versus uninflamed ileal absorptive enterocytes in Crohn disease patients. The single-cell data were retrieved from Martin et al. [72], which contains 67,050 inflamed and uninflamed cells from the ileal samples of 8 patients with Crohn disease.
(PDF)
The disease subgroups are (A) asthma, (B) diabetes, and (C) hypertension. Four models were evaluated. These models were matched and adjusted using different variables, as shown in the table. The variable that was used to extract each patient subgroup was not used for propensity score matching or odds ratio adjustment. ACEI, angiotensin-converting enzyme inhibitor; ARB, angiotensin II receptor blocker.
(PDF)
In black Americans, carvedilol use showed a lowered risk of a positive SARS-CoV-2 test when propensity score was matched with basic variables (age, sex, and smoking).
(PDF)
Each dot represents a disease (Z score versus number of genes). No significant bias was observed for the number of genes. The maximum R2 is 0.1468 from the SARS2-PPI dataset. The data underlying this figure can be found in S10 Data.
(PDF)
The disease-associated genes were filtered by their tissue specificity. Tissues considered are shown after the disease names. Only genes with positive specificity were retained for the network analysis. After filtering, diseases with fewer than 5 genes were removed from the evaluation. The data underlying this figure can be found in S11 Data.
(PDF)
Node sizes show their tissue specificity in colon.
(PDF)
The up- and down-expressed genes in the 2 asthma datasets (GSE63142 and GSE130499, severe versus control) were computed against the up- and down-expressed genes from the SARS2-DEG dataset. Overall, the results show more significant network proximities and smaller Z scores than when the direction is not considered, as in Fig 4.
(PDF)
We searched PubMed, Embase, and medRxiv databases for publications as of April 25, 2020, using the search term (“SARS-COV-2” OR “COVID-19” OR “nCoV 19” OR “2019 novel coronavirus” OR “coronavirus disease 2019”) AND (“clinical characteristics” OR “clinical outcome” OR “comorbidities”). Only research articles were included. Several criteria were used to filter the initial 1,054 articles to a final sample of 34 studies for meta-analyses.
(PDF)
Cytoscape 3.7.1 was used for the visualization and for generating the statistics. Clustering coefficient (ranges from 0 to 1) measures the extent to which the nodes in the network tend to cluster together. Network centralization (ranges from 0 to 1) measures the extent to which the topology resembles a star. Network density (ranges from 0 to 1) shows how densely the nodes are connected in the network. Network heterogeneity shows the tendency of the network to contain hub nodes.
(PDF)
(PDF)
(PDF)
(ZIP)
(PDF)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
Acknowledgments
We thank all helpful discussions and critical comments regarding this manuscript from the COVID-19 Research Intervention Advisory Committee members at the Cleveland Clinic.
Abbreviations
- ACEI
angiotensin-converting enzyme inhibitor
- AP-MS
affinity purification–mass spectrometry
- ARB
angiotensin II receptor blocker
- AT2
alveolar type II
- Co-IP+LC/MS
co-immunoprecipitation and liquid chromatography–mass spectrometry
- COPD
chronic obstructive pulmonary disease
- COVID-19
coronavirus disease 2019
- DEG
differentially expressed gene
- DEP
differentially expressed protein
- dN/dS ratio
nonsynonymous to synonymous substitution rate ratio
- ES
enrichment score
- FDA
US Food and Drug Administration
- FDR
false discovery rate
- GSEA
gene set enrichment analysis
- HCoV
human coronavirus
- HGMD
Human Gene Mutation Database
- IBD
inflammatory bowel disease
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- OR
odds ratio
- PPI
protein–protein interaction
- PS
propensity score
- RNA-Seq
RNA sequencing
- RNAi
RNA interference
- SARS-CoV-2
severe acute respiratory syndrome coronavirus 2
- viORF
viral open reading frame
Data Availability
All data sets used in this study and their sources for downloading can be found in S1 Table. The bulk and single-cell RNA-Seq data used in this study were downloaded from the NCBI GEO database with accession numbers GSE63142, GSE130499, and GSE134809. The lung and human bronchial epithelial single-cell data were downloaded from https://data.mendeley.com/datasets/7r2cwbw44m/1. Source code, the human protein-protein interactome, and drug-target network can be downloaded from https://github.com/ChengF-Lab/COVID-19_Map. All other relevant data are within the paper and its Supporting Information files.
Funding Statement
This work was supported by the National Institute of Aging (R01AG066707 and 3R01AG066707-01S1) and the National Heart, Lung, and Blood Institute (R00HL138272) to F.C. This work has been also supported in part by the VeloSano Pilot Program (Cleveland Clinic Taussig Cancer Institute) to F.C. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis. 2020;20(5):533–4. 10.1016/S1473-3099(20)30120-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bhatraju PK, Ghassemieh BJ, Nichols M, Kim R, Jerome KR, Nalla AK, et al. Covid-19 in critically Ill patients in the Seattle Region—case series. N Engl J Med. 2020;382(21):2012–22. 10.1056/NEJMoa2004500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, et al. Clinical characteristics of Coronavirus Disease 2019 in China. N Engl J Med. 2020;382(18):1708–20. 10.1056/NEJMoa2002032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, et al. Clinical characteristics of 138 hospitalized patients with 2019 novel Coronavirus-infected pneumonia in Wuhan, China. JAMA. 2020;323(11):1061–9. 10.1001/jama.2020.1585 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yang X, Yu Y, Xu J, Shu H, Xia J, Liu H, et al. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir Med. 2020;8(5):475–81. 10.1016/S2213-2600(20)30079-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395(10229):1054–62. 10.1016/S0140-6736(20)30566-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–13. 10.1016/S0140-6736(20)30211-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020;583(7816):459–68. 10.1038/s41586-020-2286-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382(8):727–33. 10.1056/NEJMoa2001017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, Erichsen S, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181(2):271–80.e8. 10.1016/j.cell.2020.02.052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hoffmann M, Kleine-Weber H, Pohlmann S. A multibasic cleavage site in the Spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol Cell. 2020;78(4):779–84.e5. 10.1016/j.molcel.2020.04.022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ziegler CGK, Allon SJ, Nyquist SK, Mbano IM, Miao VN, Tzouanas CN, et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell. 2020;181(5):1016–35.e19. 10.1016/j.cell.2020.04.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sungnak W, Huang N, Becavin C, Berg M, Queen R, Litvinukova M, et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat Med. 2020;26(5):681–7. 10.1038/s41591-020-0868-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lukassen S, Chua RL, Trefzer T, Kahn NC, Schneider MA, Muley T, et al. SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J. 2020;39(10):e105114 10.15252/embj.20105114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhang H, Kang Z, Gong H, Xu D, Wang J, Li Z, et al. Digestive system is a potential route of COVID-19: an analysis of single-cell coexpression pattern of key proteins in viral entry process. Gut. 2020;69(6):1010–8. 10.1136/gutjnl-2020-320953 [DOI] [Google Scholar]
- 16.Yang S, Fu C, Lian X, Dong X, Zhang Z. Understanding human-virus protein-protein interactions using a human protein complex-based analysis framework. mSystems. 2019;4(2):e00303–18. 10.1128/mSystems.00303-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Liu C, Ma Y, Zhao J, Nussinov R, Zhang Y, Cheng F, et al. Computational network biology: data, models, and applications. Phys Rep. 2020;846:1–66. 10.1016/j.physrep.2019.12.004 [DOI] [Google Scholar]
- 18.Zumla A, Chan JF, Azhar EI, Hui DS, Yuen KY. Coronaviruses—drug discovery and therapeutic options. Nat Rev Drug Discov. 2016;15(5):327–47. 10.1038/nrd.2015.37 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pfefferle S, Schopf J, Kogl M, Friedel CC, Muller MA, Carbajo-Lozoya J, et al. The SARS-coronavirus-host interactome: identification of cyclophilins as target for pan-coronavirus inhibitors. PLoS Pathog. 2011;7(10):e1002331 10.1371/journal.ppat.1002331 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fung TS, Liu DX. Human coronavirus: host-pathogen interaction. Annu Rev Microbiol. 2019;73:529–57. 10.1146/annurev-micro-020518-115759 [DOI] [PubMed] [Google Scholar]
- 21.Blanco-Melo D, Nilsson-Payant BE, Liu WC, Uhl S, Hoagland D, Moller R, et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell. 2020;181(5):1036–45.e9. 10.1016/j.cell.2020.04.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bojkova D, Klann K, Koch B, Widera M, Krause D, Ciesek S, et al. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature. 2020;583(7816):469–72. 10.1038/s41586-020-2332-7 [DOI] [PubMed] [Google Scholar]
- 23.Zhou Y, Wang F, Tang J, Nussinov R, Cheng F. Artificial intelligence in COVID-19 drug repurposing. Lancet Digit Health. 2020. September 18 10.1016/S2589-7500(20)30192-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Krammer F. SARS-CoV-2 vaccines in development. Nature. 2020;586(7830):516–27. 10.1038/s41586-020-2798-3 [DOI] [PubMed] [Google Scholar]
- 25.Beigel JH, Tomashek KM, Dodd LE, Mehta AK, Zingman BS, Kalil AC, et al. Remdesivir for the treatment of Covid-19—Final report. N Engl J Med. 2020. October 8 10.1056/NEJMoa2007764 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Spinner CD, Gottlieb RL, Criner GJ, Arribas Lopez JR, Cattelan AM, Soriano Viladomiu A, et al. Effect of remdesivir vs standard care on clinical status at 11 days in patients with moderate COVID-19: a randomized clinical trial. JAMA. 2020;324(11):1048–57. 10.1001/jama.2020.16349 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Horby P, Lim WS, Emberson JR, Mafham M, Bell JL, Linsell L, et al. Dexamethasone in hospitalized patients with COVID-19—preliminary report. N Engl J Med. 2020. July 17 10.1056/NEJMoa2021436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, et al. A trial of lopinavir-ritonavir in adults hospitalized with severe COVID-19. N Engl J Med. 2020;382(19):1787–99. 10.1056/NEJMoa2001282 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yan L, Zhang H, Goncalves J, Xiao Y, Wang M, Guo Y, et al. An interpretable mortality prediction model for COVID-19 patients. Nat Mach Intell. 2020;2:283–8. 10.1038/s42256-020-0180-7 [DOI] [Google Scholar]
- 30.Zhou Y, Hou Y, Shen J, Huang Y, Martin W, Cheng F. Network-based drug repurposing for novel coronavirus 2019-nCoV/SARS-CoV-2. Cell Discov. 2020;6:14 10.1038/s41421-020-0153-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cheng F, Lu W, Liu C, Fang J, Hou Y, Handy DE, et al. A genome-wide positioning systems network algorithm for in silico drug repurposing. Nat Commun. 2019;10(1):3476 10.1038/s41467-019-10744-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cheng F, Desai RJ, Handy DE, Wang R, Schneeweiss S, Barabasi AL, et al. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat Commun. 2018;9(1):2691 10.1038/s41467-018-05116-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Cheng F, Kovacs IA, Barabasi AL. Network-based prediction of drug combinations. Nat Commun. 2019;10(1):1197 10.1038/s41467-019-09186-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gysi DM, Do Valle I, Zitnik M, Ameli A, Gan X, Varol O, et al. Network medicine framework for identifying drug repurposing opportunities for COVID-19. arXiv: 2004.07229v2. 2020. August 9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cheng F, Murray JL, Zhao J, Sheng J, Zhao Z, Rubin DH. Systems biology-based investigation of cellular antiviral drug targets identified by gene-trap insertional mutagenesis. PLoS Comput Biol. 2016;12(9):e1005074 10.1371/journal.pcbi.1005074 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Pichlmair A, Kandasamy K, Alvisi G, Mulhern O, Sacco R, Habjan M, et al. Viral immune modulators perturb the human molecular network by common and unique strategies. Nature. 2012;487(7408):486–90. 10.1038/nature11289 [DOI] [PubMed] [Google Scholar]
- 37.Watanabe T, Kawakami E, Shoemaker JE, Lopes TJ, Matsuoka Y, Tomita Y, et al. Influenza virus-host interactome screen as a platform for antiviral drug development. Cell Host Microbe. 2014;16(6):795–805. 10.1016/j.chom.2014.11.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Consortium GTEx. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45(6):580–5. 10.1038/ng.2653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li G, He X, Zhang L, Ran Q, Wang J, Xiong A, et al. Assessing ACE2 expression patterns in lung tissues in the pathogenesis of COVID-19. J Autoimmun. 2020;112:102463 10.1016/j.jaut.2020.102463 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shimokawa H, Sunamura S, Satoh K. RhoA/Rho-kinase in the cardiovascular system. Circ Res. 2016;118(2):352–66. 10.1161/CIRCRESAHA.115.306532 [DOI] [PubMed] [Google Scholar]
- 41.Zaoui K, Boudhraa Z, Khalife P, Carmona E, Provencher D, Mes-Masson AM. Ran promotes membrane targeting and stabilization of RhoA to orchestrate ovarian cancer cell invasion. Nat Commun. 2019;10(1):2666 10.1038/s41467-019-10570-w [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D, et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 2011;39(Database issue):D945–50. 10.1093/nar/gkq929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sondka Z, Bamford S, Cole CG, Ward SA, Dunham I, Forbes SA. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat Rev Cancer. 2018;18(11):696–705. 10.1038/s41568-018-0060-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Liu C, Zhao J, Lu W, Dai Y, Hockings J, Zhou Y, et al. Individualized genetic network analysis reveals new therapeutic vulnerabilities in 6,700 cancer genomes. PLoS Comput Biol. 2020;16(2):e1007701 10.1371/journal.pcbi.1007701 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Stenson PD, Ball EV, Mort M, Phillips AD, Shiel JA, Thomas NS, et al. Human Gene Mutation Database (HGMD): 2003 update. Hum Mutat. 2003;21(6):577–81. 10.1002/humu.10212 [DOI] [PubMed] [Google Scholar]
- 46.Bello SM, Shimoyama M, Mitraka E, Laulederkind SJF, Smith CL, Eppig JT, et al. Disease Ontology: improving and unifying disease annotations across species. Dis Model Mech. 2018;11(3):dmm032839 10.1242/dmm.032839 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Clerkin KJ, Fried JA, Raikhelkar J, Sayer G, Griffin JM, Masoumi A, et al. COVID-19 and cardiovascular disease. Circulation. 2020;141(20):1648–55. 10.1161/CIRCULATIONAHA.120.046941 [DOI] [PubMed] [Google Scholar]
- 48.Baldi E, Sechi GM, Mare C, Canevari F, Brancaglione A, Primi R, et al. Out-of-hospital cardiac arrest during the COVID-19 outbreak in Italy. N Engl J Med. 2020;383(5):496–8. 10.1056/NEJMc2010418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Menche J, Sharma A, Kitsak M, Ghiassian SD, Vidal M, Loscalzo J, et al. Disease networks. Uncovering disease-disease relationships through the incomplete interactome. Science. 2015;347(6224):1257601 10.1126/science.1257601 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Fan E, Beitler JR, Brochard L, Calfee CS, Ferguson ND, Slutsky AS, et al. COVID-19-associated acute respiratory distress syndrome: is a different approach to management warranted? Lancet Respir Med. 2020;8(8):816–21. 10.1016/S2213-2600(20)30304-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Prescott HC, Girard TD. Recovery from severe COVID-19: leveraging the lessons of survival from sepsis. JAMA. 2020;324(8):739–40. 10.1001/jama.2020.14103 [DOI] [PubMed] [Google Scholar]
- 52.Beers MF, Mulugeta S. The biology of the ABCA3 lipid transporter in lung health and disease. Cell Tissue Res. 2017;367(3):481–93. 10.1007/s00441-016-2554-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Nogee LM. Abnormal expression of surfactant protein C and lung disease. Am J Respir Cell Mol Biol. 2002;26(6):641–4. 10.1165/ajrcmb.26.6.f241 [DOI] [PubMed] [Google Scholar]
- 54.Tay MZ, Poh CM, Renia L, MacAry PA, Ng LFP. The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol. 2020;20(6):363–74. 10.1038/s41577-020-0311-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Swaroopa D, Bhaskar K, Mahathi T, Katkam S, Raju YS, Chandra N, et al. Association of serum interleukin-6, interleukin-8, and acute physiology and chronic health evaluation II score with clinical outcome in patients with acute respiratory distress syndrome. Indian J Crit Care Med. 2016;20(9):518–25. 10.4103/0972-5229.190369 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hou T, Huang D, Zeng R, Ye Z, Zhang Y. Accuracy of serum interleukin (IL)-6 in sepsis diagnosis: a systematic review and meta-analysis. Int J Clin Exp Med. 2015;8(9):15238–45. [PMC free article] [PubMed] [Google Scholar]
- 57.Song J, Park DW, Moon S, Cho HJ, Park JH, Seok H, et al. Diagnostic and prognostic value of interleukin-6, pentraxin 3, and procalcitonin levels among sepsis and septic shock patients: a prospective controlled study according to the Sepsis-3 definitions. BMC Infect Dis. 2019;19(1):968 10.1186/s12879-019-4618-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Xu X, Han M, Li T, Sun W, Wang D, Fu B, et al. Effective treatment of severe COVID-19 patients with tocilizumab. Proc Natl Acad Sci U S A. 2020;117(20):10970–5. 10.1073/pnas.2005615117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wang Q, Fang P, He R, Li M, Yu H, Zhou L, et al. O-GlcNAc transferase promotes influenza A virus-induced cytokine storm by targeting interferon regulatory factor-5. Sci Adv. 2020;6(16):eaaz7086 10.1126/sciadv.aaz7086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Comhair SA, McDunn J, Bennett C, Fettig J, Erzurum SC, Kalhan SC. Metabolomic endotype of asthma. J Immunol. 2015;195(2):643–50. 10.4049/jimmunol.1500736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Holguin F, Grasemann H, Sharma S, Winnica D, Wasil K, Smith V, et al. L-Citrulline increases nitric oxide and improves control in obese asthmatics. JCI Insight. 2019;4(24):e131733 10.1172/jci.insight.131733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Shen B, Yi X, Sun Y, Bi X, Du J, Zhang C, et al. Proteomic and metabolomic characterization of COVID-19 patient sera. Cell. 2020;182(1):59–72.e15. 10.1016/j.cell.2020.05.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Samuelsson B. Arachidonic acid metabolism: role in inflammation. Z Rheumatol. 1991;50(Suppl 1):3–6. [PubMed] [Google Scholar]
- 64.Calabrese C, Triggiani M, Marone G, Mazzarella G. Arachidonic acid metabolism in inflammatory cells of patients with bronchial asthma. Allergy. 2000;55(Suppl 61):27–30. 10.1034/j.1398-9995.2000.00504.x [DOI] [PubMed] [Google Scholar]
- 65.Berger A. What are leukotrienes and how do they work in asthma? BMJ. 1999;319(7202):90 10.1136/bmj.319.7202.90 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Modena BD, Tedrow JR, Milosevic J, Bleecker ER, Meyers DA, Wu W, et al. Gene expression in relation to exhaled nitric oxide identifies novel asthma phenotypes with unique biomolecular pathways. Am J Respir Crit Care Med. 2014;190(12):1363–72. 10.1164/rccm.201406-1099OC [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Weathington N, O’Brien ME, Radder J, Whisenant TC, Bleecker ER, Busse WW, et al. BAL cell gene expression in severe asthma reveals mechanisms of severe disease and influences of medications. Am J Respir Crit Care Med. 2019;200(7):837–56. 10.1164/rccm.201811-2221OC [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Balaci L, Spada MC, Olla N, Sole G, Loddo L, Anedda F, et al. IRAK-M is involved in the pathogenesis of early-onset persistent asthma. Am J Hum Genet. 2007;80(6):1103–14. 10.1086/518259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.de Paiva AC, Marson FA, Ribeiro JD, Bertuzzo CS. Asthma: Gln27Glu and Arg16Gly polymorphisms of the beta2-adrenergic receptor gene as risk factors. Allergy Asthma Clin Immunol. 2014;10(1):8 10.1186/1710-1492-10-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Lamers MM, Beumer J, van der Vaart J, Knoops K, Puschhof J, Breugem TI, et al. SARS-CoV-2 productively infects human gut enterocytes. Science. 2020;369(6499):50–4. 10.1126/science.abc1669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.D’Amico F, Baumgart DC, Danese S, Peyrin-Biroulet L. Diarrhea during COVID-19 infection: pathogenesis, epidemiology, prevention, and management. Clin Gastroenterol Hepatol. 2020;18(8):1663–72. 10.1016/j.cgh.2020.04.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Martin JC, Chang C, Boschetti G, Ungaro R, Giri M, Grout JA, et al. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell. 2019;178(6):1493–508.e20. 10.1016/j.cell.2019.08.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zhang W, Hui KY, Gusev A, Warner N, Ng SM, Ferguson J, et al. Extended haplotype association study in Crohn’s disease identifies a novel, Ashkenazi Jewish-specific missense mutation in the NF-kappaB pathway gene, HEATR3. Genes Immun. 2013;14(5):310–6. 10.1038/gene.2013.19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Muise AM, Walters T, Xu W, Shen-Tu G, Guo CH, Fattouh R, et al. Single nucleotide polymorphisms that increase expression of the guanosine triphosphatase RAC1 are associated with ulcerative colitis. Gastroenterology. 2011;141(2):633–41. 10.1053/j.gastro.2011.04.057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Seinen ML, van Nieuw Amerongen GP, de Boer NK, Mulder CJ, van Bezu J, van Bodegraven AA. Rac1 as a potential pharmacodynamic biomarker for thiopurine therapy in inflammatory bowel disease. Ther Drug Monit. 2016;38(5):621–7. 10.1097/FTD.0000000000000326 [DOI] [PubMed] [Google Scholar]
- 76.Korelitz BI. Expert opinion: experience with 6-mercaptopurine in the treatment of inflammatory bowel disease. World J Gastroenterol. 2013;19(20):2979–84. 10.3748/wjg.v19.i20.2979 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Perry CM, Scott LJ. Cefdinir: a review of its use in the management of mild-to-moderate bacterial infections. Drugs. 2004;64(13):1433–64. 10.2165/00003495-200464130-00004 [DOI] [PubMed] [Google Scholar]
- 78.Johansen LM, Brannan JM, Delos SE, Shoemaker CJ, Stossel A, Lear C, et al. FDA-approved selective estrogen receptor modulators inhibit Ebola virus infection. Sci Transl Med. 2013;5(190):190ra79 10.1126/scitranslmed.3005471 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Cong Y, Hart BJ, Gross R, Zhou H, Frieman M, Bollinger L, et al. MERS-CoV pathogenesis and antiviral efficacy of licensed drugs in human monocyte-derived antigen-presenting cells. PLoS ONE. 2018;13(3):e0194868 10.1371/journal.pone.0194868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Dyall J, Coleman CM, Hart BJ, Venkataraman T, Holbrook MR, Kindrachuk J, et al. Repurposing of clinically developed drugs for treatment of Middle East respiratory syndrome coronavirus infection. Antimicrob Agents Chemother. 2014;58(8):4885–93. 10.1128/AAC.03036-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Jeon S, Ko M, Lee J, Choi I, Byun SY, Park S, et al. Identification of antiviral drug candidates against SARS-CoV-2 from FDA-approved drugs. Antimicrob Agents Chemother. 2020;64(7):e00819–20. 10.1128/AAC.00819-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Wang XJ, Hu W, Zhang TY, Mao YY, Liu NN, Wang SQ. Irbesartan, an FDA approved drug for hypertension and diabetic nephropathy, is a potent inhibitor for hepatitis B virus entry by disturbing Na(+)-dependent taurocholate cotransporting polypeptide activity. Antiviral Res. 2015;120:140–6. 10.1016/j.antiviral.2015.06.007 [DOI] [PubMed] [Google Scholar]
- 83.Zhang P, Zhu L, Cai J, Lei F, Qin JJ, Xie J, et al. Association of inpatient use of angiotensin-converting enzyme inhibitors and angiotensin II receptor blockers with mortality among patients with hypertension hospitalized with COVID-19. Circ Res. 2020;126(12):1671–81. 10.1161/CIRCRESAHA.120.317134 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Vaduganathan M, Vardeny O, Michel T, McMurray JJV, Pfeffer MA, Solomon SD. Renin-angiotensin-aldosterone system inhibitors in patients with COVID-19. N Engl J Med. 2020;382(17):1653–9. 10.1056/NEJMsr2005760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Jarcho JA, Ingelfinger JR, Hamel MB, D’Agostino RB Sr, Harrington DP. Inhibitors of the renin-angiotensin-aldosterone system and COVID-19. N Engl J Med. 2020;382(25):2462–4. 10.1056/NEJMe2012924 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Mehta N, Kalra A, Nowacki AS, Anjewierden S, Han Z, Bhat P, et al. Association of use of angiotensin-converting enzyme inhibitors and angiotensin II receptor blockers with testing positive for Coronavirus Disease 2019 (COVID-19). JAMA Cardiol. 2020;5(9):1020–6. 10.1001/jamacardio.2020.1855 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Puelles VG, Lutgehetmann M, Lindenmeyer MT, Sperhake JP, Wong MN, Allweiss L, et al. Multiorgan and renal tropism of SARS-CoV-2. N Engl J Med. 2020;383(6):590–2. 10.1056/NEJMc2011400 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Xiao F, Tang M, Zheng X, Liu Y, Li X, Shan H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. 2020;158(6):1831–3.e3. 10.1053/j.gastro.2020.02.055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Nakashima K, Hirota T, Obara K, Shimizu M, Jodo A, Kameda M, et al. An association study of asthma and related phenotypes with polymorphisms in negative regulator molecules of the TLR signaling pathway. J Hum Genet. 2006;51(4):284–91. 10.1007/s10038-005-0358-1 [DOI] [PubMed] [Google Scholar]
- 90.Pino-Yanes M, Sanchez-Machin I, Cumplido J, Figueroa J, Torres-Galvan MJ, Gonzalez R, et al. IL-1 receptor-associated kinase 3 gene (IRAK3) variants associate with asthma in a replication study in the Spanish population. J Allergy Clin Immunol. 2012;129(2):573–5, 575.e1–10. 10.1016/j.jaci.2011.10.001 [DOI] [PubMed] [Google Scholar]
- 91.Reihsaus E, Innis M, MacIntyre N, Liggett SB. Mutations in the gene encoding for the beta 2-adrenergic receptor in normal and asthmatic subjects. Am J Respir Cell Mol Biol. 1993;8(3):334–9. 10.1165/ajrcmb/8.3.334 [DOI] [PubMed] [Google Scholar]
- 92.Ali S, Hirschfeld AF, Mayer ML, Fortuno ES 3rd, Corbett N, Kaplan M, et al. Functional genetic variation in NFKBIA and susceptibility to childhood asthma, bronchiolitis, and bronchopulmonary dysplasia. J Immunol. 2013;190(8):3949–58. 10.4049/jimmunol.1201015 [DOI] [PubMed] [Google Scholar]
- 93.Akaike T, Maeda H. Nitric oxide and virus infection. Immunology. 2000;101(3):300–8. 10.1046/j.1365-2567.2000.00142.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Verity R, Okell LC, Dorigatti I, Winskill P, Whittaker C, Imai N, et al. Estimates of the severity of coronavirus disease 2019: a model-based analysis. Lancet Infect Dis. 2020;20(6):669–77. 10.1016/S1473-3099(20)30243-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Aw D, Silva AB, Palmer DB. Immunosenescence: emerging challenges for an ageing population. Immunology. 2007;120(4):435–46. 10.1111/j.1365-2567.2007.02555.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Wang X, Bian Y, Zhang R, Liu X, Ni L, Ma B, et al. Melatonin alleviates cigarette smoke-induced endothelial cell pyroptosis through inhibiting ROS/NLRP3 axis. Biochem Biophys Res Commun. 2019;519(2):402–8. 10.1016/j.bbrc.2019.09.005 [DOI] [PubMed] [Google Scholar]
- 97.Shang Y, Xu SP, Wu Y, Jiang YX, Wu ZY, Yuan SY, et al. Melatonin reduces acute lung injury in endotoxemic rats. Chin Med J (Engl). 2009;122(12):1388–93. [PubMed] [Google Scholar]
- 98.Zhang J, Wang L, Xie W, Hu S, Zhou H, Zhu P, et al. Melatonin attenuates ER stress and mitochondrial damage in septic cardiomyopathy: a new mechanism involving BAP31 upregulation and MAPK-ERK pathway. J Cell Physiol. 2020;235(3):2847–56. 10.1002/jcp.29190 [DOI] [PubMed] [Google Scholar]
- 99.Cheng F, Rao S, Mehra R. COVID-19 treatment: combining anti-inflammatory and antiviral therapeutics using a network-based approach. Cleve Clin J Med. 2020. June 30 10.3949/ccjm.87a.ccc037 [DOI] [PubMed] [Google Scholar]
- 100.Martin WR, Cheng F. Repurposing of FDA-approved toremifene to treat COVID-19 by blocking the Spike glycoprotein and NSP14 of SARS-CoV-2. J Proteome Res. 2020. September 27 10.1021/acs.jproteome.0c00397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Maisonnasse P, Guedj J, Contreras V, Behillil S, Solas C, Marlin R, et al. Hydroxychloroquine use against SARS-CoV-2 infection in non-human primates. Nature. 2020;585(7826):584–7. 10.1038/s41586-020-2558-4 [DOI] [PubMed] [Google Scholar]
- 102.Hoffmann M, Mosbauer K, Hofmann-Winkler H, Kaul A, Kleine-Weber H, Kruger N, et al. Chloroquine does not inhibit infection of human lung cells with SARS-CoV-2. Nature. 2020;585(7826):588–90. 10.1038/s41586-020-2575-3 [DOI] [PubMed] [Google Scholar]
- 103.Behzad S, Aghaghazvini L, Radmard AR, Gholamrezanezhad A. Extrapulmonary manifestations of COVID-19: radiologic and clinical overview. Clin Imaging. 2020;66:35–41. 10.1016/j.clinimag.2020.05.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Gupta A, Madhavan MV, Sehgal K, Nair N, Mahajan S, Sehrawat TS, et al. Extrapulmonary manifestations of COVID-19. Nat Med. 2020;26(7):1017–32. 10.1038/s41591-020-0968-3 [DOI] [PubMed] [Google Scholar]
- 105.Rubino F, Amiel SA, Zimmet P, Alberti G, Bornstein S, Eckel RH, et al. New-onset diabetes in COVID-19. N Engl J Med. 2020;383(8):789–90. 10.1056/NEJMc2018688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2005;33(Database issue):D514–7. 10.1093/nar/gki033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Zhang M, Zhu C, Jacomy A, Lu LJ, Jegga AG. The orphan disease networks. Am J Hum Genet. 2011;88(6):755–66. 10.1016/j.ajhg.2011.05.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Kittler R, Pelletier L, Heninger AK, Slabicki M, Theis M, Miroslaw L, et al. Genome-scale RNAi profiling of cell division in human tissue culture cells. Nat Cell Biol. 2007;9(12):1401–12. 10.1038/ncb1659 [DOI] [PubMed] [Google Scholar]
- 109.Breuer K, Foroushani AK, Laird MR, Chen C, Sribnaia A, Lo R, et al. InnateDB: systems biology of innate immunity and beyond—recent updates and continuing curation. Nucleic Acids Res. 2013;41(Database issue):D1228–33. 10.1093/nar/gks1147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44(W1):W90–7. 10.1093/nar/gkw377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Hirsh AE, Fraser HB, Wall DP. Adjusting for selection on synonymous sites in estimates of evolutionary distance. Mol Biol Evol. 2005;22(1):174–7. 10.1093/molbev/msh265 [DOI] [PubMed] [Google Scholar]
- 112.Bezginov A, Clark GW, Charlebois RL, Dar VU, Tillier ER. Coevolution reveals a network of human proteins originating with multicellularity. Mol Biol Evol. 2013;30(2):332–46. 10.1093/molbev/mss218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Cheng F, Jia P, Wang Q, Lin CC, Li WH, Zhao Z. Studying tumorigenesis through network evolution and somatic mutational perturbations in the cancer interactome. Mol Biol Evol. 2014;31(8):2156–69. 10.1093/molbev/msu167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Yang Z, Bielawski JP. Statistical methods for detecting molecular adaptation. Trends Ecol Evol. 2000;15(12):496–503. 10.1016/s0169-5347(00)01994-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959;22(4):719–48. 10.1093/jnci/22.4.719 [DOI] [PubMed] [Google Scholar]
- 116.DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7(3):177–88. 10.1016/0197-2456(86)90046-2 [DOI] [PubMed] [Google Scholar]
- 117.Wan X, Wang W, Liu J, Tong T. Estimating the sample mean and standard deviation from the sample size, median, range and/or interquartile range. BMC Med Res Methodol. 2014;14(1):135 10.1186/1471-2288-14-135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Stijnen T, Hamza TH, Ozdemir P. Random effects meta-analysis of event outcome in the framework of the generalized linear mixed model with applications in sparse data. Stat Med. 2010;29(29):3046–67. 10.1002/sim.4040 [DOI] [PubMed] [Google Scholar]
- 119.Huttlin EL, Ting L, Bruckner RJ, Gebreab F, Gygi MP, Szpyt J, et al. The BioPlex Network: a systematic exploration of the human interactome. Cell. 2015;162(2):425–40. 10.1016/j.cell.2015.06.043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Rolland T, Tasan M, Charloteaux B, Pevzner SJ, Zhong Q, Sahni N, et al. A proteome-scale map of the human interactome network. Cell. 2014;159(5):1212–26. 10.1016/j.cell.2014.10.050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature. 2005;437(7062):1173–8. 10.1038/nature04209 [DOI] [PubMed] [Google Scholar]
- 122.Cheng F, Jia P, Wang Q, Zhao Z. Quantitative network mapping of the human kinome interactome reveals new clues for rational kinase inhibitor discovery and individualized cancer therapy. Oncotarget. 2014;5(11):3697–710. 10.18632/oncotarget.1984 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Keshava Prasad TS, Goel R, Kandasamy K, Keerthikumar S, Kumar S, Mathivanan S, et al. Human Protein Reference Database—2009 update. Nucleic Acids Res. 2009;37(Database issue):D767–72. 10.1093/nar/gkn892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Hu J, Rho HS, Newman RH, Zhang J, Zhu H, Qian J. PhosphoNetworks: a database for human phosphorylation networks. Bioinformatics. 2014;30(1):141–2. 10.1093/bioinformatics/btt627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Hornbeck PV, Zhang B, Murray B, Kornhauser JM, Latham V, Skrzypek E. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 2015;43(Database issue):D512–20. 10.1093/nar/gku1267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Lu CT, Huang KY, Su MG, Lee TY, Bretana NA, Chang WC, et al. DbPTM 3.0: an informative resource for investigating substrate site specificity and functional association of protein post-translational modifications. Nucleic Acids Res. 2013;41(Database issue):D295–305. 10.1093/nar/gks1229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Dinkel H, Chica C, Via A, Gould CM, Jensen LJ, Gibson TJ, et al. Phospho.ELM: a database of phosphorylation sites—update 2011. Nucleic Acids Res. 2011;39(Database issue):D261–7. 10.1093/nar/gkq1104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Csabai L, Olbei M, Budd A, Korcsmaros T, Fazekas D. SignaLink: multilayered regulatory networks. Methods Mol Biol. 2018;1819:53–73. 10.1007/978-1-4939-8618-7_3 [DOI] [PubMed] [Google Scholar]
- 129.Oughtred R, Stark C, Breitkreutz BJ, Rust J, Boucher L, Chang C, et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019;47(D1):D529–41. 10.1093/nar/gky1079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Cowley MJ, Pinese M, Kassahn KS, Waddell N, Pearson JV, Grimmond SM, et al. PINA v2.0: mining interactome modules. Nucleic Acids Res. 2012;40(Database issue):D862–5. 10.1093/nar/gkr967 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Meyer MJ, Das J, Wang X, Yu H. INstruct: a database of high-quality 3D structurally resolved protein interactome networks. Bioinformatics. 2013;29(12):1577–9. 10.1093/bioinformatics/btt181 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Licata L, Briganti L, Peluso D, Perfetto L, Iannuccelli M, Galeota E, et al. MINT, the molecular interaction database: 2012 update. Nucleic Acids Res. 2012;40(Database issue):D857–61. 10.1093/nar/gkr930 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Orchard S, Ammari M, Aranda B, Breuza L, Briganti L, Broackes-Carter F, et al. The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 2014;42(Database issue):D358–63. 10.1093/nar/gkt1115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2016;44(D1):D7–19. 10.1093/nar/gkv1290 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Smith IN, Thacker S, Seyfi M, Cheng F, Eng C. Conformational dynamics and allosteric regulation landscapes of germline PTEN mutations associated with autism compared to those associated with cancer. Am J Hum Genet. 2019;104(5):861–78. 10.1016/j.ajhg.2019.03.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Golbeck J. Analyzing the social web. Boston: Morgan Kaufmann; 2013. [Google Scholar]
- 137.Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 2013;41(Database issue):D991–5. 10.1093/nar/gks1193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36(5):411–20. 10.1038/nbt.4096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44(D1):D457–62. 10.1093/nar/gkv1070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Brunk E, Sahoo S, Zielinski DC, Altunkaya A, Drager A, Mih N, et al. Recon3D enables a three-dimensional view of gene variation in human metabolism. Nat Biotechnol. 2018;36(3):272–81. 10.1038/nbt.4072 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Pornputtapong N, Nookaew I, Nielsen J. Human metabolic atlas: an online resource for human metabolism. Database (Oxford). 2015;2015:bav068 10.1093/database/bav068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Wishart DS, Feunang YD, Marcu A, Guo AC, Liang K, Vazquez-Fresno R, et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 2018;46(D1):D608–17. 10.1093/nar/gkx1089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 2014;42(Database issue):D1091–7. 10.1093/nar/gkt1068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Yang H, Qin C, Li YH, Tao L, Zhou J, Yu CY, et al. Therapeutic target database update 2016: enriched resource for bench to clinical drug target and targeted pathway information. Nucleic Acids Res. 2016;44(D1):D1069–74. 10.1093/nar/gkv1230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012;40(Database issue):D1100–7. 10.1093/nar/gkr777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK. BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 2007;35(Database issue):D198–201. 10.1093/nar/gkl999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Pawson AJ, Sharman JL, Benson HE, Faccenda E, Alexander SP, Buneman OP, et al. The IUPHAR/BPS Guide to PHARMACOLOGY: an expert-driven knowledgebase of drug targets and their ligands. Nucleic Acids Res. 2014;42(Database issue):D1098–106. 10.1093/nar/gkt1143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 2006;313(5795):1929–35. 10.1126/science.1132939 [DOI] [PubMed] [Google Scholar]
- 149.Reghunathan R, Jayapal M, Hsu LY, Chng HH, Tai D, Leung BP, et al. Expression profile of immune response genes in patients with severe acute respiratory syndrome. BMC Immunol. 2005;6(1):2 10.1186/1471-2172-6-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Josset L, Menachery VD, Gralinski LE, Agnihothram S, Sova P, Carter VS, et al. Cell host response to infection with novel human coronavirus EMC predicts potential antivirals and important differences with SARS coronavirus. mBio. 2013;4(3):e00165–13. 10.1128/mBio.00165-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Yuan S, Chu H, Chan JF, Ye ZW, Wen L, Yan B, et al. SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat Commun. 2019;10(1):120 10.1038/s41467-018-08015-x [DOI] [PMC free article] [PubMed] [Google Scholar]