Abstract
The ability of individuals with end-stage osteoarthritis (OA) to functionally recover from total joint arthroplasty is highly inconsistent. The molecular mechanisms driving this heterogeneity have yet to be elucidated. Furthermore, OA disproportionately impacts females, suggesting a need for identifying female-specific therapeutic targets. We profiled the skeletal muscle transcriptome in females with end-stage OA (n = 20) undergoing total knee or hip arthroplasty using RNA-Seq. Single-gene differential expression (DE) analyses tested for DE genes between skeletal muscle overlaying the surgical (SX) joint and muscle from the contralateral (CTRL) leg. Network analyses were performed using Pathway-Level Information ExtractoR (PLIER) to summarize genes into latent variables (LVs), i.e., gene circuits, and link them to biological pathways. LV differences in SX versus CTRL muscle and across sources of muscle tissue (vastus medialis, vastus lateralis, or tensor fascia latae) were determined with ANOVA. Linear models tested for associations between LVs and muscle phenotype on the SX side (inflammation, function, and integrity). DE analysis revealed 360 DE genes (|Log2 fold-difference| ≥ 1, FDR ≤ 0.05) between the SX and CTRL limbs, many associated with inflammation and lipid metabolism. PLIER analyses revealed circuits associated with protein degradation and fibro-adipogenic cell gene expression. Muscle inflammation and function were linked to an LV associated with endothelial cell gene expression highlighting a potential regulatory role of endothelial cells within skeletal muscle. These findings may provide insight into potential therapeutic targets to improve OA rehabilitation before and/or following total joint replacement.
Keywords: osteoarthritis, skeletal muscle, transcriptomics
INTRODUCTION
Osteoarthritis (OA) is a degenerative, inflammatory disease of the joints affecting ∼300 million people in the hip or knee globally (1, 2). OA in the hip and knee joints accounts for a tremendous impact on both individuals and the healthcare system. Incidence is greater in females than in males (3, 4), and the physiological burden is expansive, undermining physical function and quality of life (5, 6). Much of the physical implications of OA are known, including disability and loss of mobility (7, 8), skeletal muscle functional decline (9, 10), and pain (11). Impairments in single myofiber contractile properties appear in part to explain these functional declines (9), and individuals with OA are approximately three times as likely to develop age-associated muscle atrophy (sarcopenia) compared with healthy counterparts (12). Our research team and others have shown that periarticular skeletal muscles (around the diseased joint) show heightened expression of proinflammatory genes, greater atrophy-associated signaling, higher myofiber type IIax/x proportion, and more severe tissue fibrosis (13–15). These deleterious factors together contribute to impaired integrity of the skeletal muscle near the diseased joint.
To mitigate the negative impact of OA on physical function, many interventions have been established (16–18), with exercise training being particularly effective during early disease progression (16, 18–23). Despite this, interventions cannot be expected to fully halt disease progression. Consequently, severe mobility limitations may still occur and often result in elective total joint arthroplasty (TJA) surgery. Unfortunately, rehabilitation from TJA remains inconsistently prescribed, and approximately one-third of the patients with total knee or hip arthroplasty (TKA and THA, respectively) suffer from activity limitations years following their surgery (8) and persistent reductions in strength (24). These physical limitations impair movement and threaten independence, particularly in advanced age. It is vital to determine molecular mechanisms that may predict a patient’s ability to physically recover from TKA/THA to optimize rehabilitative success through precision-based medicine approaches.
This may be best achieved at the point of surgery, based on key phenotypic signatures that we have identified based on skeletal muscle inflammatory gene expression. Although these genes play a variety of important physiological roles (25–27), we have identified overt inflammation in periarticular skeletal muscle in some patients with OA, classified as muscle inflammation susceptibility (MuIS) (15, 28, 29). MuIS+ status limits muscle regenerative capacity (28), and the protein products of transcripts Fn14 and TNF-αR may be key drivers of muscle impairment in patients with OA (15, 29). Well beyond these target genes, an array of coordinated changes in the molecular phenotype of skeletal muscle in OA is likely contributing to functional and structural impairments.
Although transcriptome-wide association studies (TWAS) have begun to establish candidate markers for genetic predisposition to OA (30, 31), transcriptomic differences in periarticular skeletal muscle have been underexplored in OA. This is surprising given the importance of skeletal muscle in a number of functions relevant to OA progression or TKA/THA recovery, notably movement and locomotion, metabolism, and immune/inflammatory function. Our recent findings suggest that perioperative features of muscle may be important consideration in recovery from TKA/THA (29). Extending these findings to the protein-coding muscle transcriptome in end-stage OA (i.e., at the time of TKA/THA) is considered a major advance toward understanding the impact of OA on the surrounding muscle and may potentially provide insight into designing interventions that overcome individual heterogeneity in response to post-TJA rehabilitation.
In the present study, we performed RNA sequencing (RNA-Seq) of the muscle overlaying the diseased joint on the surgical (SX) side and a comparator muscle on the contralateral/nonsurgical (CTRL) side in females with end-stage knee or hip OA undergoing TKA/THA. To establish precedent for potential site-specific differences in skeletal muscle surrounding the OA joint, we also compared the muscle overlaying the SX hip to muscle overlaying the SX knee across individuals. Gene expression data were analyzed in two stages: single-gene-level differential expression (DE) analysis, followed by modeling of transcriptomic expression circuits using Pathway-Level Information ExtractoR (PLIER). PLIER anchors gene expression circuits to biological pathways, aiding in the interpretation of transcriptomic data while also summarizing the biology of novel associations, or “latent variables” (LVs) of genes (32, 33). Our team has leveraged PLIER to establish differential LVs in skeletal muscle in the context of aging, Parkinson’s disease (PD), rehabilitation, and acute exercise responses in skeletal muscle (34, 35). PLIER has also been used to profile the gene expression patterns in various cell types that exist within skeletal muscle tissue homogenate (33), some of which have a high degree of relevance for OA [e.g., fibro-adipogenic progenitor cells (FAPS)] (35). Our findings provide valuable insight into skeletal muscle perturbations in end-stage OA and may be leveraged to refine therapeutic interventions to improve patients’ quality of life following TKA/THA.
METHODS
Participants
Participants with end-stage osteoarthritis (OA) were recruited from the University of Alabama at Birmingham (UAB) and the University of Arkansas for Medical Sciences for the parent clinical trial (NCT02628795, R01HD084124). Inclusion and exclusion criteria have previously been described in detail (29). All participants provided written informed consent to participate and permission for their collected biospecimens to be used for follow-on analyses. The parent clinical trial was approved by the UAB and UAMS Institutional Review Boards. Twenty female participants from the parent trial were selected for the present skeletal muscle transcriptomic profiling. Data for knee extension strength and power, thigh lean mass, fibrosis, and the targeted gene expression (qPCR) of TNF-αR and Fn14 are a subset of data from our previously published manuscript (29).
Knee Extensor Strength, Power, and Thigh Lean Mass
A subset of previously published data for knee extension maximum voluntary isometric contraction (MVIC) strength and power, as well as thigh lean mass measurements were used in these analyses (29). MVIC was assessed by fixing the participant’s leg at 60° of knee flexion then following two familiarization repetitions, three maximum efforts to extend the shank against a fixed dynamometer (Biodex System 4 Pro Shirley, NY, and Cybex Isokinetic System, Ronkonkoma, NY) were measured. Knee extension power was assessed by participants completing five full unilateral knee extension repetitions, with the concentric action being completed as explosively as possible against a resistance equal to 40% MVIC (Biodex System 4 Pro, Shirley, NY, and Cybex Isokinetic System, Ronkonkoma, NY). SX thigh lean mass (TLM) was assessed using Dual Energy X-ray Absorptiometry (UAB: DXA, Lunar Prodigy Model No. 8743, GE Lunar Corporation, Madison, WI; UAMS: Hologic Discovery model W) before arthroplasty surgery. TLM was normalized to participant’s height (cm) and analyzed using enCORE/APEX software (v. 18/13.3) according to the manufacturer’s instructions.
Biospecimen Collection
Tissue collection procedures have previously been described in detail (29). Briefly, skeletal muscle was collected in a fasting state at the time of the participant’s scheduled joint replacement surgery, shortly after the onset of anesthesia and before the surgery itself. CTRL muscle was collected from the vastus lateralis, whereas the muscle collected from the SX limb was either the vastus medialis [for individuals undergoing total knee arthroplasty (TKA)], or tensor fascia latae [for individuals undergoing total hip arthroplasty (THA)]. Following collection, muscle biopsies were cleaned of excess blood, fat, and connective tissue. Muscle tissue intended for RNA-Seq (∼20 mg) was immediately frozen in liquid nitrogen (LN2), whereas ∼50 mg was aligned for histochemical analyses in a bed of mounting media made up of tragacanth gum combined with Tissue Tek O.C.T. compound (Sakura Finetek, Torrance, CA), and frozen in LN2 cooled isopentane, as previously described (29). All biospecimens were stored at −80°C until processed for analyses.
Histologically Assessed Skeletal Muscle Fibrosis
Skeletal muscle fibrosis was determined using wheat germ agglutinin (WGA) as previously published (29, 36). Cryosections (6-µm) were placed in a humidity chamber for 10 min at room temperature, fixed for 3 min in 1:2 acetone and methanol, washed three times for 5 min in 1× PBS, incubated in Texas Red conjugated WGA for 1 h, washed again with 1× PBS three times for 5 min, coated with mounting media, and secured with a coverslip. Images were captured at ×20 using an Olympus BX51 fluorescent microscope with an Olympus XM10 camera (Olympus) and CellSens Dimensions software (Olympus). Images were analyzed for percent positive red pixel area using Image Pro Premier software (v. 9.1).
Biospecimen Processing for Omics Pipelines
Skeletal muscle total RNA was extracted using the Qiagen miRNeasy kits (Qiagen, Valencia CA, Cat. No. 217004) as previously described (29). RNA quality was assessed using an Agilent Bioanalyzer (Santa Clara, CA) and was of sufficient quality for sequencing [RNA Integrity Number (RIN): means ± SD 7.8 ± 0.7]. RNA samples were then sent to the Translational Genomics Research Institute (TGen) for downstream RNA processing and sequencing. Both samples (i.e., RNA from CTRL and SX muscle) were prepared in the same library batch to minimize batch effects within a participant. Allocation of paired samples to a given batch was randomized. Total RNA was DNase-treated with TURBO DNA-free (Thermo Fisher, Cat. No. AM1907), then purified with RNeasy MinElute Cleanup (Qiagen, Cat. No. 74204) using a modified MinElute Cleanup protocol to capture both small and long RNA species (in step 2 of the standard MinElute protocol, 950 µL 100% EtOH was added to each sample instead of 250 µL). RNA was measured for quantity with Quant-iT Ribogreen RNA Assay (Thermo Fisher, Cat. No. R11490) and quality with High Sensitivity RNA ScreenTape and buffer (Agilent, Cat. Nos. 5067–5579 and 5067–5580). For each RNA sample, a uniquely dual-indexed, Illumina-compatible, double-stranded cDNA whole transcriptome library was synthesized from 10 ng of total RNA with Takara Bio’s SMARTer Stranded Total RNA-Seq kit v2 Pico Input Mammalian (Takara Bio, Cat. No. 634418) and SMARTer RNA Unique Dual Index Kit (Takara Bio, Cat. No. 634452). Briefly, this library preparation included RNA fragmentation, a 5-cycle Indexing PCR, ribosomal cDNA depletion, and a 12-cycle enrichment PCR. The RNA fragmentation incubation time and temperature were dictated by each sample’s RIN and %DV200 according to the library prep protocol. Each library was measured for size with Agilent’s High-Sensitivity D1000 ScreenTape and buffer (Agilent, Cat. Nos. 5067–5584 & 5067–5603). One microliter of each library was combined into a nonequimolar pool that was then measured for size via TapeStation and concentration via Roche’s KAPA SYBR FAST Universal qPCR Kit (Roche, Cat. No. KK4824), diluted to 70 pM, then loaded into an iSeq flowcell cartridge (Illumina, Cat. No. 20031371) with a 1% v/v PhiX Control v3 spike-in (Illumina, Cat. No. FC-110–3001), and sequenced at 101 × 8 × 8 × 101 cycles. Passing filter cluster counts per library were generated from this data and used to make a rebalanced pool that was subsequently measured for size and concentration, diluted to 2 nM with a 1% v/v PhiX Control v3 spike-in, denatured and further diluted, loaded into a NovaSeq 6000 flow cell cartridge (Illumina, Cat. No. 20028313) and sequenced at 101 × 9 ± 9 × 101 cycles with standard workflow and a final flow cell concentration of 400 pM. Each library generated at least 100 million paired-end reads. STAR (v2.6+) was used to align RNA to the human genome (GCHr38). Count data were generated using featureCounts, and compiled annotations were used to subset only protein-coding genes. Raw FASTQ files and the associated count tables have been deposited in GEO (GSE205431).
Targeted Gene Expression (qPCR)
Targeted skeletal muscle gene expression of two genes was assessed using TaqMan qPCR as previously described (29): TNF receptor 1 A (TNF-αR, Hs00533560_m1) and TWEAK receptor (Fn14, Hs0017993_m1) (29). The arithmetic mean of glyceraldehyde 3-phosphate dehydrogenase (GAPDH, Hs02758991_g1) and chromosome-1 open reading frame 43 (CH1ORF43, Hs01555103_T1 g1) served as internal controls. All samples were run in triplicate using the Quantstudio 3 (Applied Biosystems, StepOne software v. 2.2.2). Gene expression was determined using the arbitrary units 2−DCT method (37).
Computational Analyses
Gene counts were filtered by biotype to focus on protein-coding genes and analyzed with a modified version of our established workflow (34, 38), which is shown in Fig. 1. All samples passed sequencing QC. Count data were filtered by expression. Statistical outliers were identified and verified using principal component and median absolute deviation analyses (MAD). Samples with a MAD > 6 from principal components 1 and 2 were excluded from the analyses. If one sample in the paired analyses was deemed an outlier, the corresponding sample was also removed (e.g., SX sample also dropped if CTRL was an outlier).
Figure 1.
Methodological workflow utilized for the complementary single-gene (Limma-Voom) and network-based (PLIER) analysis pipelines. Transcripts were filtered for low expression, protein-coding biotype, then by study design. Batch effects and outliers were assessed: two surgical (SX) samples were outliers, and these samples along with corresponding contralateral (CTRL) samples were removed from further analyses. Limma-voom was used to detect single-gene differences in muscle gene expression between the SX and CTRL limbs. For PLIER, counts were converted to counts per million then z-scored. An input matrix linking genes to biological pathways from published prior knowledge was constructed resulting in 4,515 available pathways. Singular value decomposition (SVD) was performed in PLIER and generated 14 latent variables (LVs). SX vs. CTRL and TKA vs. THA differences were assessed with analysis of variance (ANOVA). For SX only, PLIER scores for each LV were assessed using linear models to determine associations between LVs and individual phenotypes of interest, including gene expression as measured by qPCR, histologically assessed fibrosis, and muscle function. PLIER, Pathway-Level Information ExtractoR; THA, total hip arthroplasty; TKA, total knee arthroplasty.
Single-gene-level DE analyses were conducted using the limma-voom pipeline (39) to distinguish differences between the CTRL and SX muscle, as well as SX muscle overlaying the knee versus the hip. Voom was used to normalize samples and establish predicted variance-based weights for each gene. To control for between-subject variability, participant ID was used as a blocking variable in the model design such that CTRL and SX samples within a participant were effectively paired. DE genes (DEGs) were deemed significant based on an absolute log2 fold-difference of ≥ 1 and a Benjamini–Hochberg-adjusted false discovery rate (FDR) ≤ 0.05. Significant genes with an absolute log2 fold difference ≥ 3 were assessed for representation within biological processes using the WEB-based Gene SeT AnaLysis Toolkit [WebGestalt (40)].
In our stage 2 parallel analysis, expression circuits were summarized as LVs using PLIER. As previously described, PLIER generated LVs using singular value decomposition (SVD) guided by a compendium of prior knowledge established from relevant literature (32, 34, 38). Briefly, fiber type-specific expression, skeletal muscle mononuclear cell type-specific expression (33), as well as canonical pathways [e.g., REACTOME, KEGG, and Gene Ontology (41–46)] were loaded as prior knowledge to inform biological annotations for protein-coding transcripts as previously described (38) (n total input transcripts = 13,221; n total input pathways = 4,515). The fiber type and muscle-derived mononuclear cell expression profiles were established from the study by Rubenstein et al. (33), wherein single-cell techniques were coupled with PLIER and bulk RNA-Seq to establish methods for cell/fiber type deconvolution within skeletal muscle homogenate. PLIER down selected input data to 10,771 common genes shared between the prior knowledge and our transcriptomic data and 1,274 pathways in which these transcripts were adequately annotated.
Based on the inherent structure of the data, PLIER generated 14 LVs. The 50 genes best associated with each LV were examined for relative expression across samples using heatmaps (32, 34). ANOVA was used to test differences between the CTRL and SX muscle as well as site of joint arthroplasty (TKA or THA). P values were corrected for multiple comparisons using Bonferroni adjustments and deemed significant at P ≤ 0.05 with trends listed as a P value between >0.05 and <0.10. Differences between the CTRL and SX limb were determined using ANOVA main effects adjusted for the number of LVs. Any significant interaction between limb (CTRL vs. SX) and site of joint arthroplasty (TKA vs. THA) was followed by post hoc testing for multiple comparisons (Tukey’s post hoc). The resulting data for LVs with significant interaction terms (adjusted P ≤ 0.05) are provided in the supplemental material (all Supplemental material is available at https://doi.org/10.6084/m9.figshare.19387571). Associations between the LVs and perioperative functional status were determined using linear models.
RESULTS
Participant Characteristics
Participant characteristics are presented in Table 1. Although 20 participants (40 samples, i.e., 20 CTRL and 20 SX) were initially included in these analyses, outlier detection identified two SX samples that were removed along with the corresponding sample from the CTRL leg. All analyses were thus completed on n = 18 participants (36 samples, i.e., 18 CTRL and 18 SX; Table 1). As described in the methods, participants were all female, and 13/18 (∼70%) were undergoing total knee arthroplasty, whereas five were undergoing total hip arthroplasty. Body mass index (BMI) was in the obese classification (31.4 ± 4.2 kg/m2).
Table 1.
Participant characteristics
| Variable | n = 20 | n = 18 |
|---|---|---|
| Surgery | ||
| THA, n | 5 | 5 |
| TKA, n | 15 | 13 |
| Age, yr | 62.9 ± (9.4) | 61.8 ± (9.0) |
| Height, m | 1.6 ± (0.1) | 1.6 ± (0.1) |
| Weight, kg | 85.0 ± (14.5) | 85.3 ± (15.4) |
| BMI, kg/m2 | 31.5 ± (4.2) | 31.4 ± (4.2) |
Values are represented as means ± SD; n = 20 participants available for inclusion and n = 18 participants remaining after outlier exclusion. All CTRL muscle was collected from the vastus lateralis. All TKA SX muscle was collected from the vastus medialis. All THA SX muscle was collected from the tensor fascia latae. BMI, body mass index; CTRL, contralateral; THA, total hip arthroplasty; TKA, total knee arthroplasty; SX, surgical.
Differentially Expressed Genes in Skeletal Muscle of SX versus CTRL Limbs
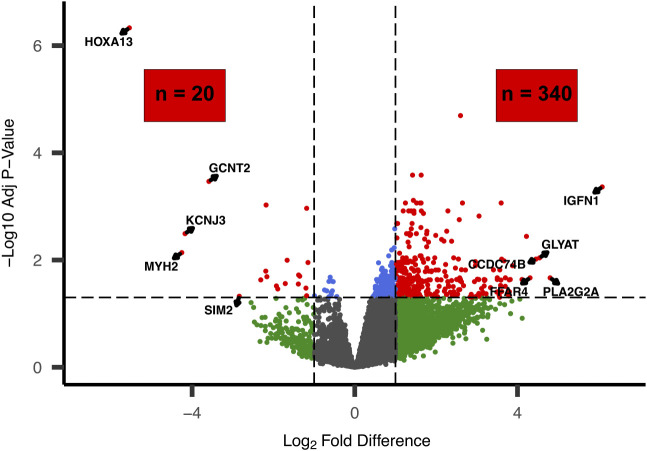
In total, 360 genes were differentially expressed between the SX and CTRL limbs (FDR ≤ 0.05, |Log2FD| ≥ 1; Fig. 2). Of these, 340 genes had significantly higher expression, whereas 20 had significantly lower expression in the muscle of the SX limb compared with the CTRL (Supplemental Table S1). Genes with an FDR ≤ 0.05 and |log2FD| ≥ 3 are shown in Table 2 (n = 49). Biological processes (via Gene Ontology) nominally associated with these 49 genes included regulation of membrane potential (GO:0042391, P = 0.0008, FDR = 0.22), ERK1 and ERK2 cascade (GO:0070371, P = 0.002, FDR = 0.26), lipid catabolic process (GO:0016042, P = 0.002, FDR = 0.26), regulation of oxidoreductase activity (GO:0051341, P = 0.002, FDR = 0.26), and humoral immune response (GO:0006959, P = 0.004, FDR = 0.38) among others. DE analysis between THA and TKA SX muscles found 141 genes with significantly higher expression and 496 genes with significantly lower expression in the vastus medialis of participants undergoing TKA compared with the muscle overlaying the hip (tensor fascia latae) in participants undergoing THA (Supplemental Table S1).
Figure 2.
Volcano plot of differentially expressed genes in skeletal muscle between SX and CTRL limbs of participants with OA. In total, 360 genes were significantly different between the SX and CTRL limbs with 340 higher and 20 lower on the SX limb, respectively. Significance was determined at a threshold of FDR ≤ 0.05 and |Log2FD| ≥ 1. Labeled genes represent the five highest log2 fold-difference (Log2FD) in the positive and negative directions. These genes are further explored in Table 2. CTRL, contralateral; FDR, false discovery rate; OA, osteoarthritis; SX, surgical.
Table 2.
Muscle genes dysregulated in the SX limb
| Gene Symbol | Description | Log2FD | FDR |
|---|---|---|---|
| Genes with higher expression in the SX compared with CTRL muscle | |||
| IGFN1 | Immunoglobulin Like and Fibronectin Type III Domain Containing 1 | 6.08 | 4.33E-04 |
| PLA2G2A | Phospholipase A2 Group IIA | 4.80 | 2.15E-02 |
| GLYAT | Glycine-N-Acyltransferase | 4.54 | 9.11E-03 |
| CCDC74B | Coiled-Coil Domain Containing 74B | 4.45 | 9.60E-03 |
| FFAR4 | Free Fatty Acid Receptor 4 | 4.31 | 2.15E-02 |
| FAM135B | Family With Sequence Similarity 135 Member B | 4.22 | 3.62E-03 |
| SCIN | Scinderin | 4.11 | 2.31E-02 |
| CIDEA | Cell Death Inducing DFFA Like Effector A | 3.88 | 1.28E-02 |
| HIST1H2BF | Histone H2B type 1-F | 3.82 | 2.31E-02 |
| CAMK1G | Calcium/Calmodulin Dependent Protein Kinase IG | 3.80 | 3.90E-02 |
| CXCL10 | C-X-C Motif Chemokine Ligand 10 | 3.80 | 2.35E-02 |
| SCN1A | Sodium Voltage-Gated Channel α Subunit 1 | 3.79 | 4.68E-02 |
| SLC26A9 | Solute Carrier Family 26 Member 9 | 3.75 | 4.26E-02 |
| CXCL1 | C-X-C Motif Chemokine Ligand 1 | 3.70 | 2.15E-02 |
| HIST1H3I | Histone gene cluster-1 H3I | 3.70 | 3.10E-02 |
| TDRD5 | Tudor Domain Containing 5 | 3.67 | 1.04E-02 |
| CNTNAP4 | Contactin Associated Protein Family Member 4 | 3.61 | 3.87E-02 |
| FAT2 | FAT Atypical Cadherin 2 | 3.61 | 9.63E-03 |
| PITX1 | Paired Like Homeodomain 1 | 3.60 | 8.63E-04 |
| FRMD1 | FERM Domain Containing 1 | 3.59 | 2.37E-02 |
| GYG2 | Glycogenin 2 | 3.58 | 2.98E-02 |
| RSPO2 | R-Spondin 2 | 3.56 | 1.53E-02 |
| PAPPA2 | Pappalysin 2 | 3.56 | 4.95E-02 |
| SHANK1 | SH3 and Multiple Ankyrin Repeat Domains 1 | 3.55 | 4.54E-02 |
| MT3 | Metallothionein 3 | 3.53 | 3.90E-02 |
| CIDEC | Cell Death Inducing DFFA Like Effector C | 3.50 | 2.15E-02 |
| SLC22A18 | Solute Carrier Family 22 Member 18 | 3.50 | 2.31E-02 |
| C4orf19 | Chromosome 4 Open Reading Frame 19 | 3.49 | 3.96E-02 |
| C6 | Complement C6 | 3.41 | 1.53E-02 |
| HOXD9 | Homeobox D9 | 3.34 | 3.28E-02 |
| B3GNT4 | UDP-GlcNAc:BetaGal Beta-1,3-N-Acetylglucosaminyltransferase 4 | 3.23 | 4.26E-02 |
| SEPT12 | Septin 12 | 3.20 | 2.35E-02 |
| KCNA3 | Potassium Voltage-Gated Channel Subfamily A Member 3 | 3.19 | 2.42E-02 |
| GALNT8 | Polypeptide N-Acetylgalactosaminyltransferase 8 | 3.18 | 2.31E-02 |
| SEC14L2 | SEC14 Like Lipid Binding 2 | 3.18 | 3.05E-02 |
| LAMC3 | Laminin Subunit Gamma 3 | 3.15 | 4.02E-02 |
| OPRD1 | Opioid Receptor Delta 1 | 3.13 | 4.37E-02 |
| SLC7A4 | Solute Carrier Family 7 Member 4 | 3.12 | 3.41E-02 |
| NPR3 | Natriuretic Peptide Receptor 3 | 3.05 | 1.51E-03 |
| KCNN4 | Potassium Calcium-Activated Channel Subfamily N Member 4 | 3.05 | 4.65E-02 |
| CD180 | CD180 Molecule | 3.05 | 4.21E-02 |
| PLIN1 | Perilipin 1 | 3.05 | 2.31E-02 |
| HS3ST3A1 | Heparan Sulfate-Glucosamine 3-Sulfotransferase 3A1 | 3.04 | 4.73E-02 |
| ADIPOQ | Adiponectin, C1Q and Collagen Domain Containing | 3.01 | 2.28E-02 |
| GFI1 | Growth Factor Independent 1 Transcriptional Repressor | 3.01 | 4.64E-02 |
| Genes with lower expression in the SX compared with CTRL muscle | |||
| HOXA13 | Homeobox A13 | −5.54 | 4.67E-07 |
| MYH2 | Myosin Heavy Chain 2 | −4.25 | 7.28E-03 |
| KCNJ3 | Potassium Inwardly Rectifying Channel Subfamily J Member 3 | −4.17 | 3.22E-03 |
| GCNT2 | Glucosaminyl (N-Acetyl) Transferase 2 (I Blood Group) | −3.59 | 3.42E-04 |
Criteria for included genes: |Log2FD| ≥ 3 and FDR ≤ 0.05. CTRL, contralateral; FD, fold-difference; FDR, false discovery rate; SX, surgical.
Gene Expression Circuits Differentially Expressed between the SX and CTRL Limbs
In a parallel analysis using PLIER as a dimension-reducing tool to summarize latent data structure, 14 LVs were generated, with 9 LVs annotating to at least one pathway from the available prior knowledge (Fig. 3). Five LVs were different at the nominal level between muscle from the SX and CTRL limbs: LV3 (electron transport chain, P = 0.04), LV5 (no annotation, P = 0.0004), LV7 (no annotation, P = 0.006), LV9 (FAP cell genes, P = 0.004), and LV11 (no annotation, P = 0.007). Of the five LVs, one survived strict Bonferroni adjustment for the number of LVs (LV5: P-adjusted = 0.006; Fig. 4, A and B), whereas trends remained for LV7 (no annotation, P-adjusted = 0.08; Supplemental Fig. 1, A and B), LV9 (FAP cell genes, P-adjusted = 0.056; Fig. 5, A and B), and LV11 (no annotation, P-adjusted = 0.10; Supplemental Fig. 1, C and D). Of the top 50 genes in LV5, eukaryotic translation elongation factor 1 alpha 1 (EEF1A1) was also significantly more highly expressed in SX than CTRL muscle at the single-gene level. Significant interactions were found between muscles collected from TKA versus THA participants and limb for LVs 2 (Type IIa Markers), 3 (Electron Transport Chain), and 13 (Endothelial Cells; P-adjusted ≤ 0.001, 0.01, and 0.03, respectively; Supplemental Fig. S2, A–F). Follow-up multiple comparisons revealed significant differences for 1) LV2 (Type IIa Markers) between CTRL and SX muscles for each respective surgery (P-adjusted ≤ 0.05) and between THA versus TKA SX muscles (P-adjusted < 0.001); 2) LV3 (Electron Transport Chain) between THA versus TKA SX muscles (P-adjusted = 0.03); and 3) LV13 (Endothelial Cells) CTRL versus SX for THA (P-adjusted ≤ 0.05), as well as THA versus TKA SX muscles (P-adjusted < 0.001).
Figure 3.
Annotated latent variables (LVs) and the top annotations associated with them. Prior knowledge inputs provided to PLIER consisted of canonical pathways (KEGG, REACTOME, MIPS), gene ontology (GO), fiber type-specific expression profiles, and vastus lateralis-derived mononuclear cell expression profiles (33, 41–46). Where available, the top pathway association was assigned as the annotation for each LV. FAP cells, fibro-adipogenic cells; FBN1+, fibrillin-1 positive; KEGG, Kyoto Encyclopedia of Genes and Genomes; MIPS, Munich Information Center for Protein Sequences; PLIER, Pathway-Level Information Extractor; PCV, postcapillary venule.
Figure 4.
A: difference between the latent variable (LV) score on the SX and CTRL limb for LV5. Individual points are indicated for each participant and circles denote total knee arthroplasty (TKA) participants and triangles denote total hip arthroplasty participants (THA). Generally, LV5 was higher in muscle on the SX limb compared with the CTRL limb (P-adjusted = 0.006). B: heatmap illustrating the normalized expression of the top 50 genes in LV5 demonstrating the difference in expression across limbs (SX and CTRL) and surgery location (TKA or THA). Significance was set as an adjusted P value ≤ 0.05, Trends were determined as an adjusted P valued ≤ 0.10. CTRL, contralateral; SX, surgical.
Figure 5.
A: difference between the latent variable (LV) score on the SX and CTRL limb for LV9 annotating to fibro-adipogenic cell expression (FAP). Individual points are indicated for each participant, and circles denote total knee arthroplasty (TKA) participants and triangles denote total hip arthroplasty participants (THA). Generally, LV9 was higher in muscle on the SX limb compared with the CTRL limb (P-adjusted = 0.056, Trend). B: heatmap illustrating the normalized expression of the top 50 genes in LV9 demonstrating the difference in expression across limbs (SX and CTRL) and surgery location (TKA or THA). Significance was determined as an adjusted P value ≤ 0.05. Trends were determined as an adjusted P valued ≤ 0.10. CTRL, contralateral; SX, surgical.
Expression Circuits Correlated With Perioperative Function on the SX Limb
Within SX limb muscle only, circuits from four LVs were associated with gene expression of Fn14 as measured by qPCR in our previous report (29): LV4 (MIPS Spliceosome, P = 0.02, adjusted R2 = 0.23), LV7 (no annotation, P = 0.04, adjusted R2 = 0.20), LV8 (GO Regulation of Transcription DNA Templated, P = 0.001, adjusted R2 = 0.45), and LV14 (Endothelial Cells, Fig. 6A, P = 0.01, adjusted R2 = 0.30). In addition, within SX only, LV4 (MIPS Spliceosome) trended toward an association with the SX qPCR gene expression of TNF-αR (P = 0.07, adjusted R2 = 0.14). Within SX only, LV14 (Endothelial Cells) trended toward association with in vivo functional phenotypes including SX knee extension MVIC and knee extension power (P = 0.08, adjusted R2 = 0.15 Fig. 6B; P = 0.06, adjusted R2 = 0.17 Fig. 6C, respectively). No SX molecular expression circuits (LVs) were correlated with SX fibrosis or TLM (P > 0.05).
Figure 6.
Associations of LV14 (annotated to endothelial cells) to markers of inflammation (A, qPCR-determined Fn14 expression, P value = 0.01, n = 18), knee extension strength (B, MVIC, P value = 0.08, n = 16), and knee extension power (C, power, P value = 0.06, n = 16). Sample sizes for strength and power are reduced because some participants were unable to complete the measurement. MVIC, maximum voluntary isometric contraction.
DISCUSSION
OA is a tremendous burden on mobility and quality of life that also negatively impacts the health of periarticular skeletal muscle. Given the greater incidence of OA among females, along with generally more limited understanding of muscle gene expression in females, we mapped the periarticular muscle transcriptome profile of females with end-stage OA to further elucidate the molecular impact of the disease and present opportunities for advancement in potential treatments. The paired design of this study allowed each individual to be their own internal control (muscle in the SX limb vs. muscle in the CTRL limb), bolstering statistical power. Dimension-reducing network analyses via PLIER enabled mapping of biologically cohesive molecular expression circuits. Furthermore, we illustrated novel and cell type-specific expression circuits that may help elucidate potential underpinnings of alterations in the skeletal muscle molecular phenotype during OA progression. These data may ultimately foster identification of therapeutic targets to improve the overall health of skeletal muscle tissue in OA. In the SX muscle, we linked a subset of the molecular circuits to inflammatory expression that we have previously described (15, 29), and strong trends toward linkages to muscle performance (knee extension MVIC and power). These results set the stage for follow-on mechanistic analyses into top genes associated with the identified circuits that may represent efficacious therapeutic targets for mitigation of OA-related muscle pathophysiology or enhanced recovery following total joint arthroplasty.
Differentially Expressed Genes between the SX and CTRL Limbs
The multifaceted nature of OA is expected to have wide-ranging effects on surrounding skeletal muscle due to the disease proper, along with functional compensations an individual makes over time to reduce pain and discomfort of the inflamed joint during locomotion and movement. In line with our previous data highlighting the role of inflammation in perioperative muscle (15, 29), the SX limb skeletal muscle had heightened expression of inflammatory genes such as C-X-C Motif Chemokine Ligand 1 (CXCL1). CXCL1 is a chemoattractant that has been found to promote immune cell infiltration in the acute stages of muscle damage (47, 48). Furthermore, CXCL1 has been suggested to contribute to IL-6 secretion in primary synovial fibroblasts collected from patients with OA, thus enhancing the inflammatory environment (49). Lipid metabolism, specifically palmitate loading, increases CXCL1 expression in C2C12 myoblasts (50). This is relevant, as lipid mediating factors such as the CIDE family members Cell Death Inducing DFFA Like Effector A (CIDEA) and Cell Death Inducing DFFA Like Effector C (CIDEC) were more highly expressed in muscle from the SX limb versus CTRL. CIDEA and CIDEC have been implicated in lipid droplet formation within skeletal muscle through antilipolytic mechanisms (51). Intramyocellular fat stores in sedentary individuals appear to promote insulin insensitivity and potentially lipid toxicity (52). Altogether, this suggests that the SX limb is in a state of heightened inflammation and dysregulated metabolism that may contribute to the decline in skeletal muscle health observed in those with OA.
In addition to the factors, we have previously used to describe the inflammatory burden of muscle (Fn14, TNF-αR, and IL-6R), this transcriptome-wide analysis not only revealed novel candidate biomarkers but also summarized coordinated gene expression circuits that may be partially responsible for complex processes such as chronic inflammation localized to the tissue, an ongoing focus of our research team (15, 28, 29). Future analysis is necessary to establish whether key factors are different at the protein level and/or contribute to differential activity of cellular pathways promoting inflammation within skeletal muscle.
Gene Circuits Differentially Expressed between the SX and CTRL Limbs
The use of dimension-reducing techniques to uncover coordinated expression circuits within a transcriptomics data set is a fundamental strategy to better understanding differences in molecular signatures between conditions. Leveraging PLIER to identify gene circuits different between the SX and CTRL muscles, we identified a novel expression circuit (LV5) heightened in SX limb muscle compared with the CTRL. Among the top 50 genes of LV5, eukaryotic translation elongation factor 1 alpha 1 (EEF1A1) was also significantly heightened in the SX single-gene DEG analysis. EEF1A1 has been shown to promote proapoptotic progression in cultured mouse myotubes (53) and EEF1A1 mRNA correlates positively with the rate of proteolysis in patients with multiple injuries (54). This elevation of EEF1A1, coupled with skeletal muscle atrophy implicated in the onset and progression of OA (55), suggests a potential role for EEF1A1 in promoting proteolysis in those with OA and could be an important target in future countermeasure development. Among the nominally significantly different LVs between the SX and CTRL limbs, LV9 annotated to fibro-adipogenic progenitor (FAP) cell gene expression within muscle. Among the top 50 genes in LV9 (FAP Cells), many are genes associated with collagen regulation and are more highly expressed on the SX limb. This mirrors the higher SX skeletal muscle fibrosis we previously published in the larger cohort (29). FAP cells within skeletal muscle have been implicated in development of fibrosis through paracrine signaling (56). Moreover, PLIER analysis enabled us to identify a coordinated expression circuit (LV9) indicative of the profibrosis pathology we noted in skeletal muscle surrounding a joint with OA (29). These data, although informative, should be interpreted with caution as they did not survive multiple test corrections and therefore should be validated in future work with a larger sample size.
Follow-up multiple comparisons revealed differences in LV scores between the SX and CTRL limbs and the site of surgery. These LVs annotated to type IIa markers (LV2), electron transport chain (LV3), and endothelial cells (LV13). These findings partially capture the difference in myofiber type between the periarticular muscles sampled from around the hip and the knee. Unlike the human vastus lateralis (57) and medialis muscles (roughly a 40%–60% mixture of type I and II myofibers), the muscle sampled from around the hip (tensor fascia latae) are myofiber type I dominant (58, 59). In the SX muscle of THA versus TKA, substantially lower expression of the LV2 circuit (Type IIa Markers), coupled with higher expression of the LV3 circuit (electron transport chain and Type I Myofibers), offers strong face validity for the PLIER deconvolution approach.
Finally, LV13 was annotated to endothelial cells and is also heightened in the SX THA muscle samples. Capillary endothelial cells are responsible for providing an integrated network within skeletal muscle for oxygen exchange and nutrient delivery (e.g., glucose and amino acids) (60, 61). Type I myofibers are generally in contact with more capillaries per fiber in untrained cohorts than are type II myofibers (62), and this is likely what is being reflected in the LV13 (endothelial cell-associated gene circuit). Overall, the identified circuits associated with proteolysis and fibrosis may facilitate development of targeted treatments, whereas the circuits in line with known myofiber-type differences across muscles bolster the validity of this approach.
SX Expression Circuits Correlated With Presurgical Muscle Function
In addition to identifying novel differences in gene circuits between the SX and CTRL muscles, the SX muscle transcriptional profile may provide insights into the drivers of disease severity in females at the time of surgery. This is paramount as there is a severe lack of data specific to females as a whole (63, 64), and our previous work suggests inflammatory gene expression may drive greater heterogeneity in functional performance in female participants at the time of surgery compared with their male counterparts (29). We found four molecular expression circuits associated with the expression of inflammatory receptor Fn14 as previously assessed using qPCR (29). These LVs were annotated as related to MIPS: spliceosome activity (LV4), transcription regulation (LV8), and endothelial cell-associated expression (LV14). Although LV7 was not annotated to prior knowledge, it was also statistically associated with Fn14 expression. In addition, LV4 (MIPS spliceosome expression) tended to correlate with qPCR-measured TNF-αR gene expression. These data support the notion that Fn14 and TNF-αR are important hub genes for skeletal muscle (15, 29, 65–68), and likely manifest through dysfunction of molecular networks not limited to “inflammatory” circuits.
Notably, LV14 (endothelial cells) associated circuit was linked to Fn14 qPCR expression and tended to correlate with lower levels of knee extension strength and power. Previous work has demonstrated that OA may promote myofiber-specific contractile decline (9), thus genes within the LV14 circuit may play an important role. For instance, genes related to myofiber regeneration such as hes-related family BHLH transcription factor with YRPW motif like (HEYL) and NOTCH receptor 3 (NOTCH3) (69–71) are within the top 50 genes associated with the LV14 circuit, suggesting repair-associated expression may be responsible for the observed spectrum of skeletal muscle dysfunction. In addition, LV14 is annotated to endothelial cell-associated expression and is likely depicting genes linked to muscle capillaries within the muscle homogenate. Capillary density has been implicated in age-associated anabolic resistance that may be a contributing factor to sarcopenia (72). Although the present work did not find an association of LVs with DXA thigh lean mass, this result may have differed if a higher sensitivity means of assessing muscle mass were used, such as magnetic resonance imaging or computed tomography (38, 73). Patients with OA are at higher risk for developing sarcopenia (12); thus accurate estimation of muscle mass is highly important in future work investigating its relationships to gene expression and function relationship in individuals with OA. In summary, these data identify biological signatures of interest such as skeletal muscle inflammation, FAP cell expression, and capillarization that may be key in optimizing OA therapeutic approaches moving forward, as well as novel target genes possibly responsible for driving their activity.
Limitations
Although this work reveals novel molecular expression circuits that may advance our understanding of OA-associated muscle pathophysiology and facilitate the pursuit of targeted treatments, there are limitations. First, samples size is fairly small. Despite that, the within-subject experimental design and network-based analyses strengthen statistical power. Second, comparisons across muscles varying in fiber type and in vivo function increase complexity. However, this was mitigated in multiple ways such as parsing data from SX muscles of TKA versus THA, as well as leveraging PLIER to discern differential expression specific to fiber type. PLIER defines each LV based on the main feature(s) distinguishing the variance of the data. Therefore, LVs not annotated to myofiber types should represent characteristics other than myofiber type differences across muscles (32). In addition, it should be noted that these are protein-coding transcript data only and do not depict what noncoding RNAs are being expressed or what may be occurring at the functional protein level. In future work, we acknowledge that it may be beneficial to not only use within-person controls but also plan for an age- and sex-matched healthy comparator. Finally, we caution extrapolation or direct comparison of these data with male participants and promote a larger scale analysis to capture differences associated with biological sex in the muscle of patients with OA (64).
Conclusions
We identified 360 DEGs between the SX and CTRL limbs, some of which were associated with inflammation and lipid metabolism thus presenting potential candidate markers for therapeutic intervention. Furthermore, novel gene circuits differed between the SX and CTRL limbs, pointing toward proteolysis and FAP cell expression that may be promoting the declines in muscle integrity we and others have observed. SX PLIER LVs associated with qPCR indicators of inflammation as well as markers of muscle function. LV14 (annotated to endothelial cell expression) is an identified circuit that may influence the functional capacity of skeletal muscle. Although the specific mechanisms remain unclear, it appears that skeletal muscle capillaries may be a key target area for future work to promote function improvements in the skeletal muscle of patients with OA. Future efforts are necessary to determine the impact of these periarticular muscle molecular expression circuits on OA pathology. Rehabilitation efforts following TKA/TKA may benefit from the inclusion of supervised resistance and aerobic exercise to promote muscle retention/regain and stimulate skeletal muscle capillarization.
DATA AVAILABILITY
Raw FASTQ files and the associated count tables have been deposited in GEO (GSE205431).
SUPPLEMENTAL DATA
Supplemental Figs. S1 and S2 and Supplemental Table S1: https://doi.org/10.6084/m9.figshare.19387571.
GRANTS
This work was supported by the NIH Grants R01HD084124, P2CHD086851, and T32HD071866.
DISCLOSURES
J.A.S. has received consultant fees from Schipher, Crealta/Horizon, Medisys, Fidia, PK Med, Two laboratories Inc., Adept Field Solutions, Clinical Care options, Clearview healthcare partners, Putnam associates, Focus forward, Navigant consulting, Spherix, MedIQ, Jupiter Life Science, UBM LLC, Trio Health, Medscape, WebMD, and Practice Point communications; and the National Institutes of Health and the American College of Rheumatology. J.A.S. has received institutional research support from Zimmer Biomet Holdings. J.A.S. received food and beverage payments from Intuitive Surgical Inc./Philips Electronics North America. J.A.S. owns stock options in TPT Global Tech, Vaxart pharmaceuticals, Atyu biopharma, Adaptimmune Therapeutics, GeoVax Labs, Pieris Pharmaceuticals, Enzolytics Inc., Seres Therapeutics, Tonix Pharmaceuticals Holding Corp., and Charlotte’s Web Holdings, Inc. J.A.S. previously owned stock options in Amarin, Viking and Moderna pharmaceuticals. J.A.S. is on the speaker’s bureau of Simply Speaking. J.A.S. is a member of the executive of Outcomes Measures in Rheumatology (OMERACT), an organization that develops outcome measures in rheumatology and receives arms-length funding from 8 companies. J.A.S. serves on the FDA Arthritis Advisory Committee. J.A.S. is the chair of the Veterans Affairs Rheumatology Field Advisory Committee. J.A.S. is the editor and the Director of the University of Alabama at Birmingham (UAB) Cochrane Musculoskeletal Group Satellite Center on Network Meta-analysis. J.A.S. previously served as a member of the following committees: member, the American College of Rheumatology’s (ACR) Annual Meeting Planning Committee (AMPC) and Quality of Care Committees, the Chair of the ACR Meet-the-Professor, Workshop and Study Group Subcommittee and the co-chair of the ACR Criteria and Response Criteria subcommittee. None of the other authors has any conflicts of interest, financial or otherwise, to disclose.
AUTHOR CONTRIBUTIONS
D.J.D., K.M.L., I.A., J.A., R.R., A.A.F., S.L.B.Jr., and M.M.B. conceived and designed research; D.J.D., K.M.L., R.S., H.J.S., E.G., J.A.S., A.B., J.A., R.R., E.H., K.V.K.-J., S.E.S., C.L.B., A.A.F., and S.L.B.Jr. performed experiments; D.J.D. and K.M.L. analyzed data; D.J.D., K.M.L., Z.A.G., and M.M.B. interpreted results of experiments; D.J.D. and K.M.L. prepared figures; D.J.D. drafted manuscript; D.J.D., K.M.L., Z.A.G., S.M.O., J.S.M., M.E.L., R.S., I.A., H.J.S., E.G., J.A.S., A.B., J.A., R.R., E.H., K.V.K.-J., S.E.S., C.L.B., A.A.F., S.L.B.Jr., and M.M.B. edited and revised manuscript; D.J.D., K.M.L., Z.A.G., S.M.O., J.S.M., M.E.L., R.S., I.A., H.J.S., E.G., J.A.S., A.B., J.A., R.R., E.H., K.V.K.-J., S.E.S., C.L.B., A.A.F., S.L.B.Jr., and M.M.B. approved final version of manuscript.
ACKNOWLEDGMENTS
The authors thank all of the study participants as well as exercise testing (S. C. Tuggle and B. Peoples) and laboratory (A. Dunning, C. Kelley, D. Wiggins, and S. Yang) staff involved in data collection.
REFERENCES
- 1. Mobasheri A, Saarakkala S, Finnilä M, Karsdal MA, Bay-Jensen A-C, van Spil WE. Recent advances in understanding the phenotypes of osteoarthritis. F1000Res 8: 2091, 2019. doi: 10.12688/f1000research.20575.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Safiri S, Kolahi A-A, Smith E, Hill C, Bettampadi D, Mansournia MA, Hoy D, Ashrafi-Asgarabad A, Sepidarkish M, Almasi-Hashiani A, Collins G, Kaufman J, Qorbani M, Moradi-Lakeh M, Woolf AD, Guillemin F, March L, Cross M. Global, regional and national burden of osteoarthritis 1990-2017: a systematic analysis of the Global Burden of Disease Study 2017. Ann Rheum Dis 79: 819–828, 2020. doi: 10.1136/annrheumdis-2019-216515. [DOI] [PubMed] [Google Scholar]
- 3. Tschon M, Contartese D, Pagani S, Borsari V, Fini M. Gender and sex are key determinants in osteoarthritis not only confounding variables. a systematic review of clinical data. J Clin Med 10: 3178, 2021. doi: 10.3390/jcm10143178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Anderson AS, Loeser RF. Why is osteoarthritis an age-related disease? Best Pract Res Clin Rheumatol 24: 15–26, 2010. doi: 10.1016/j.berh.2009.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Mobasheri A, Batt M. An update on the pathophysiology of osteoarthritis. Ann Phys Rehabil Med 59: 333–339, 2016. doi: 10.1016/j.rehab.2016.07.004. [DOI] [PubMed] [Google Scholar]
- 6. Kawano MM, Araújo ILA, Castro MC, Matos MA. Assessment of quality of life in patients with knee osteoarthritis. Acta Ortop Bras 23: 307–310, 2015. doi: 10.1590/1413-785220152306150596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. McDonough CM, Jette AM. The contribution of osteoarthritis to functional limitations and disability. Clin Geriatr Med 26: 387–399, 2010. doi: 10.1016/j.cger.2010.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Singh JA, Lewallen DG. Predictors of activity limitation and dependence on walking aids following primary total hip arthroplasty. J Am Geriatr Soc 58: 2387–2393, 2010. doi: 10.1111/j.1532-5415.2010.03182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Callahan DM, Tourville TW, Slauterbeck JR, Ades PA, Stevens-Lapsley J, Beynnon BD, Toth MJ. Reduced rate of knee extensor torque development in older adults with knee osteoarthritis is associated with intrinsic muscle contractile deficits. Exp Gerontol 72: 16–21, 2015. doi: 10.1016/j.exger.2015.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Suetta C, Aagaard P, Magnusson SP, Andersen LL, Sipilä S, Rosted A, Jakobsen AK, Duus B, Kjaer M. Muscle size, neuromuscular activation, and rapid force characteristics in elderly men and women: effects of unilateral long-term disuse due to hip-osteoarthritis. J Appl Physiol (1985) 102: 942–948, 2007. doi: 10.1152/japplphysiol.00067.2006. [DOI] [PubMed] [Google Scholar]
- 11. Loeser RF, Goldring SR, Scanzello CR, Goldring MB. Osteoarthritis: a disease of the joint as an organ. Arthritis Rheum 64: 1697–1707, 2012. doi: 10.1002/art.34453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Petermann-Rocha F, Chen M, Gray SR, Ho FK, Pell JP, Celis-Morales C. Factors associated with sarcopenia: a cross-sectional analysis using UK Biobank. Maturitas 133: 60–67, 2020. doi: 10.1016/j.maturitas.2020.01.004. [DOI] [PubMed] [Google Scholar]
- 13. Callahan DM, Tourville TW, Miller MS, Hackett SB, Sharma H, Cruickshank NC, Slauterbeck JR, Savage PD, Ades PA, Maughan DW, Beynnon BD, Toth MJ. Chronic disuse and skeletal muscle structure in older adults: sex-specific differences and relationships to contractile function. Am J Physiol Cell Physiol 308: C932–C943, 2015. doi: 10.1152/ajpcell.00014.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Noehren B, Kosmac K, Walton RG, Murach KA, Lyles MF, Loeser RF, Peterson CA, Messier SP. Alterations in quadriceps muscle cellular and molecular properties in adults with moderate knee osteoarthritis. Osteoarthritis Cartilage 26: 1359–1368, 2018. doi: 10.1016/j.joca.2018.05.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Bamman MM, Ferrando AA, Evans RP, Stec MJ, Kelly NA, Gruenwald JM, Corrick KL, Trump JR, Singh JA. Muscle inflammation susceptibility: a prognostic index of recovery potential after hip arthroplasty? Am J Physiol Endocrinol Physiol 308: E670–E679, 2015. doi: 10.1152/ajpendo.00576.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kolasinski SL, Neogi T, Hochberg MC, Oatis C, Guyatt G, Block J, Callahan L, Copenhaver C, Dodge C, Felson D, Gellar K, Harvey WF, Hawker G, Herzig E, Kwoh CK, Nelson AE, Samuels J, Scanzello C, White D, Wise B, Altman RD, DiRenzo D, Fontanarosa J, Giradi G, Ishimori M, Misra D, Shah AA, Shmagel AK, Thoma LM, Turgunbaev M, Turner AS, Reston J. 2019 American College of Rheumatology/Arthritis Foundation Guideline for the management of osteoarthritis of the hand, hip, and knee. Arthritis Rheumatol 72: 220–233, 2020. doi: 10.1002/art.41142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Jette DU, Hunter SJ, Burkett L, Langham B, Logerstedt DS, Piuzzi NS, Poirier NM, Radach LJL, Ritter JE, Scalzitti DA, Stevens-Lapsley JE, Tompkins J, Zeni J Jr;. American Physical Therapy Association. Physical therapist management of total knee arthroplasty. Phys Ther 100: 1603–1631, 2020. doi: 10.1093/ptj/pzaa099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Bamman MM, Wick TM, Carmona‐Moran CA, Bridges SL. Exercise medicine for osteoarthritis: research strategies to maximize effectiveness. Arthritis Care Res (Hoboken) 68: 288–291, 2016. doi: 10.1002/acr.22680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Bannuru RR, Osani MC, Vaysbrot EE, Arden NK, Bennell K, Bierma-Zeinstra SMA, Kraus VB, Lohmander LS, Abbott JH, Bhandari M, Blanco FJ, Espinosa R, Haugen IK, Lin J, Mandl LA, Moilanen E, Nakamura N, Snyder-Mackler L, Trojian T, Underwood M, McAlindon TE. OARSI guidelines for the non-surgical management of knee, hip, and polyarticular osteoarthritis. Osteoarthritis Cartilage 27: 1578–1589, 2019. doi: 10.1016/j.joca.2019.06.011. [DOI] [PubMed] [Google Scholar]
- 20. Gohir SA, Eek F, Kelly A, Abhishek A, Valdes AM. Effectiveness of internet-based exercises aimed at treating knee osteoarthritis. JAMA Netw Open 4: e210012, 2021. [Erratum in JAMA Netw Open 4: e216209, 2021]. doi: 10.1001/jamanetworkopen.2021.0012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. de Almeida AC, Aily JB, Pedroso MG, Gonçalves GH, de Carvalho Felinto J, Ferrari RJ, Pastre CM, Mattiello SM. A periodized training attenuates thigh intermuscular fat and improves muscle quality in patients with knee osteoarthritis: results from a randomized controlled trial. Clin Rheumatol 39: 1265–1275, 2020. doi: 10.1007/s10067-019-04892-9. [DOI] [PubMed] [Google Scholar]
- 22. Liao C-D, Liao Y-H, Liou T-H, Hsieh C-Y, Kuo Y-C, Chen H-C. Effects of protein-rich nutritional composition supplementation on sarcopenia indices and physical activity during resistance exercise training in older women with knee osteoarthritis. Nutrients 13: 2487, 2021. doi: 10.3390/nu13082487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Liao C-D, Chen H-C, Kuo Y-C, Tsauo J-Y, Huang S-W, Liou T-H. Effects of muscle strength training on muscle mass gain and hypertrophy in older adults with osteoarthritis: a systematic review and meta-analysis. Arthritis Care Res (Hoboken) 72: 1703–1718, 2020. doi: 10.1002/acr.24097. [DOI] [PubMed] [Google Scholar]
- 24. Berghmans DDP, Lenssen AF, Emans PJ, de Bie RA. Functions, disabilities and perceived health in the first year after total knee arthroplasty; a prospective cohort study. BMC Musculoskelet Disord 19: 250, 2018. doi: 10.1186/s12891-018-2159-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Tidball JG. Inflammatory processes in muscle injury and repair. Am J Physiol Regul Integr Comp Physiol 288: R345–R353, 2005. doi: 10.1152/ajpregu.00454.2004. [DOI] [PubMed] [Google Scholar]
- 26. Bhatnagar S, Kumar A. The TWEAK-Fn14 system: breaking the silence of cytokine-induced skeletal muscle wasting. Curr Mol Med 12: 3–13, 2012. doi: 10.2174/156652412798376107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Chazaud B. Inflammation during skeletal muscle regeneration and tissue remodeling: application to exercise-induced muscle damage management. Immunol Cell Biol 94: 140–145, 2016. doi: 10.1038/icb.2015.97. [DOI] [PubMed] [Google Scholar]
- 28. Merritt EK, Stec MJ, Thalacker-Mercer A, Windham ST, Cross JM, Shelley DP, Tuggle SC, Kosek DJ, Kim J, Bamman MM. Heightened muscle inflammation susceptibility may impair regenerative capacity in aging humans. J Appl Physiol (1985) 115: 937–948, 2013. doi: 10.1152/japplphysiol.00019.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Drummer DJ, McAdam JS, Seay R, Aban I, Lavin KM, Wiggins D, Touliatos G, Yang S, Kelley C, Tuggle SC, Peoples B, Siegel H, Ghanem E, Singh J, Schutzler S, Barnes CL, Ferrando AA, Bridges SL Jr, Bamman MM. Perioperative assessment of muscle inflammation susceptibility in patients with end-stage osteoarthritis. J Appl Physiol (1985) 132: 984–994, 2022. doi: 10.1152/japplphysiol.00428.2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Qi X, Yu F, Wen Y, Li P, Cheng B, Ma M, Cheng S, Zhang L, Liang C, Liu L, Zhang F. Integration of transcriptome-wide association study and messenger RNA expression profile to identify genes associated with osteoarthritis. Bone Joint Res 9: 130–138, 2020. doi: 10.1302/2046-3758.93.BJR-2019-0137.R1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Xu J, Zeng Y, Si H, Liu Y, Li M, Zeng J, Shen B. Integrating transcriptome-wide association study and mRNA expression profile identified candidate genes related to hand osteoarthritis. Arthritis Res Ther 23: 81, 2021. doi: 10.1186/s13075-021-02458-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Mao W, Zaslavsky E, Hartmann BM, Sealfon SC, Chikina M. Pathway-level information extractor (PLIER) for gene expression data. Nat Methods 16: 607–610, 2019. doi: 10.1038/s41592-019-0456-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Rubenstein AB, Smith GR, Raue U, Begue G, Minchev K, Ruf-Zamojski F, Nair VD, Wang X, Zhou L, Zaslavsky E, Trappe TA, Trappe S, Sealfon SC. Single-cell transcriptional profiles in human skeletal muscle. Sci Rep 10: 229, 2020. doi: 10.1038/s41598-019-57110-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Lavin KM, Ge Y, Sealfon SC, Nair VD, Wilk K, McAdam JS, Windham ST, Kumar PL, McDonald M-LN, Bamman MM. Rehabilitative impact of exercise training on human skeletal muscle transcriptional programs in Parkinson’s disease. Front Physiol 11: 653, 2020. doi: 10.3389/fphys.2020.00653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Rubenstein AB, Hinkley JM, Nair VD, Nudelman G, Standley RA, Yi F, Yu G, Trappe TA, Bamman MM, Trappe SW, Sparks LM, Goodpaster BH, Vega RB, Sealfon SC, Zaslavsky E, Coen PM. Skeletal muscle transcriptome response to a bout of endurance exercise in physically active and sedentary older adults. Am J Physiol Endocrinol Physiol 322: E260–E277, 2022. doi: 10.1152/ajpendo.00378.2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Yarar-Fisher C, Bickel CS, Kelly NA, Stec MJ, Windham ST, McLain AB, Oster RA, Bamman MM. Heightened TWEAK-NF-κB signaling and inflammation-associated fibrosis in paralyzed muscles of men with chronic spinal cord injury. Am J Physiol Endocrinol Physiol 310: E754–E761, 2016. doi: 10.1152/ajpendo.00240.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Lavin KM, Perkins RK, Jemiolo B, Raue U, Trappe SW, Trappe TA. Effects of aging and lifelong aerobic exercise on basal and exercise-induced inflammation. J Appl Physiol (1985) 128: 87–99, 2020. doi: 10.1152/japplphysiol.00495.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Lavin KM, Bell MB, McAdam JS, Peck BD, Walton RG, Windham ST, Tuggle SC, Long DE, Kern PA, Peterson CA, Bamman MM. Muscle transcriptional networks linked to resistance exercise training hypertrophic response heterogeneity. Physiol Genomics 53: 206–221, 2021. doi: 10.1152/physiolgenomics.00154.2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Law CW, Alhamdoosh M, Su S, Dong X, Tian L, Smyth GK, Ritchie ME. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Res 5: ISCB Comm J-1408, 2018. doi: 10.12688/f1000research.9005.3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res 47: W199–W205, 2019. doi: 10.1093/nar/gkz401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Gillespie M, Jassal B, Stephan R, Milacic M, Rothfels K, Senff-Ribeiro A, Griss J, Sevilla C, Matthews L, Gong C, Deng C, Varusai T, Ragueneau E, Haider Y, May B, Shamovsky V, Weiser J, Brunson T, Sanati N, Beckman L, Shao X, Fabregat A, Sidiropoulos K, Murillo J, Viteri G, Cook J, Shorser S, Bader G, Demir E, Sander C, Haw R, Wu G, Stein L, Hermjakob H, D'Eustachio P. The reactome pathway knowledgebase 2022. Nucleic Acids Res 50: D687–D692, 2022. doi: 10.1093/nar/gkab1028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28: 27–30, 2000. doi: 10.1093/nar/28.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Kanehisa M. Toward understanding the origin and evolution of cellular organisms. Protein Sci 28: 1947–1951, 2019. doi: 10.1002/pro.3715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res 49: D545–D551, 2021. doi: 10.1093/nar/gkaa970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. Nat Genet 25: 25–29, 2000. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gene Ontology Consortium. The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res 49: D325–D334, 2021. doi: 10.1093/nar/gkaa1113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Tidball JG. Regulation of muscle growth and regeneration by the immune system. Nat Rev Immunol 17: 165–178, 2017. doi: 10.1038/nri.2016.150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Howard EE, Pasiakos SM, Blesso CN, Fussell MA, Rodriguez NR. Divergent roles of inflammation in skeletal muscle recovery from injury. Front Physiol 11: 87, 2020. doi: 10.3389/fphys.2020.00087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Hou S-M, Chen P-C, Lin C-M, Fang M-L, Chi M-C, Liu J-F. CXCL1 contributes to IL-6 expression in osteoarthritis and rheumatoid arthritis synovial fibroblasts by CXCR2, c-Raf, MAPK, and AP-1 pathway. Arthritis Res Ther 22: 251, 2020. doi: 10.1186/s13075-020-02331-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Masuda S, Tanaka M, Inoue T, Ohue-Kitano R, Yamakage H, Muranaka K, Kusakabe T, Shimatsu A, Hasegawa K, Satoh-Asahara N. Chemokine (C-X-C motif) ligand 1 is a myokine induced by palmitate and is required for myogenesis in mouse satellite cells. Acta Physiol (Oxf) 222: e12975, 2018. doi: 10.1111/apha.12975. [DOI] [PubMed] [Google Scholar]
- 51. Meex RCR, Schrauwen P, Hesselink MKC. Modulation of myocellular fat stores: lipid droplet dynamics in health and disease. Am J Physiol Regul Integr Comp Physiol 297: R913–R924, 2009. doi: 10.1152/ajpregu.91053.2008. [DOI] [PubMed] [Google Scholar]
- 52. Li X, Li Z, Zhao M, Nie Y, Liu P, Zhu Y, Zhang X. Skeletal muscle lipid droplets and the athlete’s paradox. Cells 8: 249, 2019. doi: 10.3390/cells8030249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Ruest L-B, Marcotte R, Wang E. Peptide elongation factor eEF1A-2/S1 expression in cultured differentiated myotubes and its protective effect against caspase-3-mediated apoptosis. J Biol Chem 277: 5418–5425, 2002. doi: 10.1074/jbc.M110685200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Bosutti A, Scaggiante B, Grassi G, Guarnieri G, Biolo G. Overexpression of the elongation factor 1A1 relates to muscle proteolysis and proapoptotic p66(ShcA) gene transcription in hypercatabolic trauma patients. Metabolism 56: 1629–1634, 2007. doi: 10.1016/j.metabol.2007.07.003. [DOI] [PubMed] [Google Scholar]
- 55. Shorter E, Sannicandro AJ, Poulet B, Goljanek-Whysall K. Skeletal muscle wasting and its relationship with osteoarthritis: a mini-review of mechanisms and current interventions. Curr Rheumatol Rep 21: 40, 2019. doi: 10.1007/s11926-019-0839-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Bensalah M, Muraine L, Boulinguiez A, Giordani L, Albert V, Ythier V, Dhiab J, Oliver A, Hanique V, Gidaro T, Perié S, Lacau St-Guily J, Corneau A, Butler-Browne G, Bigot A, Mouly V, Negroni E, Trollet C. A negative feedback loop between fibroadipogenic progenitors and muscle fibres involving endothelin promotes human muscle fibrosis. J Cachexia Sarcopenia Muscle 13: 1771–1784, 2022. doi: 10.1002/jcsm.12974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Stec MJ, Thalacker-Mercer A, Mayhew DL, Kelly NA, Tuggle SC, Merritt EK, Brown CJ, Windham ST, Dell'Italia LJ, Bickel CS, Roberts BM, Vaughn KM, Isakova-Donahue I, Many GM, Bamman MM. Randomized, four-arm, dose-response clinical trial to optimize resistance exercise training for older adults with age-related muscle atrophy. Exp Gerontol 99: 98–109, 2017. doi: 10.1016/j.exger.2017.09.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Willemse H, Theodoratos A, Smith PN, Dulhunty AF. Unexpected dependence of RyR1 splice variant expression in human lower limb muscles on fiber-type composition. Pflugers Arch 468: 269–278, 2016. doi: 10.1007/s00424-015-1738-9. [DOI] [PubMed] [Google Scholar]
- 59. Pumberger M, VON Roth P, Preininger B, Mueller M, Perka C, Winkler T. Microstructural changes of tensor fasciae latae and gluteus medius muscles following total hip arthroplasty: a prospective trial. Acta Chir Orthop Traumatol Cech 84: 97–100, 2017. [PubMed] [Google Scholar]
- 60. Poole DC, Copp SW, Ferguson SK, Musch TI. Skeletal muscle capillary function: contemporary observations and novel hypotheses. Exp Physiol 98: 1645–1658, 2013. doi: 10.1113/expphysiol.2013.073874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Hendrickse P, Degens H. The role of the microcirculation in muscle function and plasticity. J Muscle Res Cell Motil 40: 127–140, 2019. doi: 10.1007/s10974-019-09520-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Kelly NA, Hammond KG, Stec MJ, Bickel CS, Windham ST, Tuggle SC, Bamman MM. Quantification and characterization of grouped type I myofibers in human aging. Muscle Nerve 57: E52–E59, 2018. doi: 10.1002/mus.25711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Costello JT, Bieuzen F, Bleakley CM. Where are all the female participants in sports and exercise medicine research? Eur J Sport Sci 14: 847–851, 2014. doi: 10.1080/17461391.2014.911354. [DOI] [PubMed] [Google Scholar]
- 64. O’Bryan SM, Connor KR, Drummer DJ, Lavin KM, Bamman MM. Considerations for sex-cognizant research in exercise biology and medicine. Front Sports Act Living 4: 903992, 2022. doi: 10.3389/fspor.2022.903992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Kumar A, Bhatnagar S, Paul PK. TWEAK and TRAF6 regulate skeletal muscle atrophy. Curr Opin Clin Nutr Metab Care 15: 233–239, 2012. doi: 10.1097/MCO.0b013e328351c3fc. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Bhatnagar S, Mittal A, Gupta SK, Kumar A. TWEAK causes myotube atrophy through coordinated activation of ubiquitin-proteasome system, autophagy, and caspases. J Cell Physiol 227: 1042–1051, 2012. doi: 10.1002/jcp.22821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Mittal A, Bhatnagar S, Kumar A, Lach-Trifilieff E, Wauters S, Li H, Makonchuk DY, Glass DJ, Kumar A. The TWEAK-Fn14 system is a critical regulator of denervation-induced skeletal muscle atrophy in mice. J Cell Biol 188: 833–849, 2010. doi: 10.1083/jcb.200909117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Hayden MS, Ghosh S. Regulation of NF-κB by TNF family cytokines. Semin Immunol 26: 253–266, 2014. doi: 10.1016/j.smim.2014.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Gerrard JC, Hay JP, Adams RN, Williams JC, Huot JR, Weathers KM, Marino JS, Arthur ST. Current thoughts of Notch’s role in myoblast regulation and muscle-associated disease. Int J Environ Res Public Health 18: 12558, 2021. doi: 10.3390/ijerph182312558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Fukuda S, Kaneshige A, Kaji T, Noguchi Y-T, Takemoto Y, Zhang L, Tsujikawa K, Kokubo H, Uezumi A, Maehara K, Harada A, Ohkawa Y, Fukada S-I. Sustained expression of HeyL is critical for the proliferation of muscle stem cells in overloaded muscle. eLife 8: e48284, 2019. doi: 10.7554/eLife.48284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Carey KA, Farnfield MM, Tarquinio SD, Cameron-Smith D. Impaired expression of Notch signaling genes in aged human skeletal muscle. J Gerontol A Biol Sci Med Sci 62: 9–17, 2007. doi: 10.1093/gerona/62.1.9. [DOI] [PubMed] [Google Scholar]
- 72. Banks NF, Rogers EM, Church DD, Ferrando AA, Jenkins NDM. The contributory role of vascular health in age-related anabolic resistance. J Cachexia Sarcopenia Muscle 13: 114–127, 2022. doi: 10.1002/jcsm.12898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Haun CT, Vann CG, Roberts BM, Vigotsky AD, Schoenfeld BJ, Roberts MD. A critical evaluation of the biological construct skeletal muscle hypertrophy: size matters but so does the measurement. Front Physiol 10: 247, 2019. doi: 10.3389/fphys.2019.00247. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figs. S1 and S2 and Supplemental Table S1: https://doi.org/10.6084/m9.figshare.19387571.
Data Availability Statement
Raw FASTQ files and the associated count tables have been deposited in GEO (GSE205431).