Abstract
The TRK2 gene in Saccharomyces cerevisiae encodes a membrane protein involved in potassium transport and is expressed at extremely low levels. Dominant cis-acting mutations (TRK2D), selected by their ability to confer TRK2-dependent growth on low-potassium medium, identified an upstream repressor element (URS1-TRK2) in the TRK2 promoter. The URS1-TRK2 sequence (5'-AGCCGCACG-3') shares six nucleotides with the ubiquitous URS1 element (5'-AGCCGCCGA-3'), and the protein species binding URS1-CAR1 (URSF) is capable of binding URS1-TRK2 in vitro. Sequence analysis of 17 independent repression-defective TRK2D mutations identified three adjacent nucleotides essential for URS1-mediated repression in vivo. Our results suggest a role for context effects with regard to URS1-related sequences: several mutant alleles of the URS1 element previously reported to have little or no effect when analyzed within the context of a heterologous promoter (CYC1) [Luche, R.M., Sumrada, R. & Cooper, T.G. (1990) Mol. Cell. Biol. 10, 3884-3895] have major effects on repression in the context of their native promoters (TRK2 and CAR1). TRK2D mutations that abolish repression also reveal upstream activating sequence activity either within or adjacent to URS1. Additivity between TRK2D and sin3 delta mutations suggest that SIN3-mediated repression is independent of that mediated by URS1.
Full text
PDF

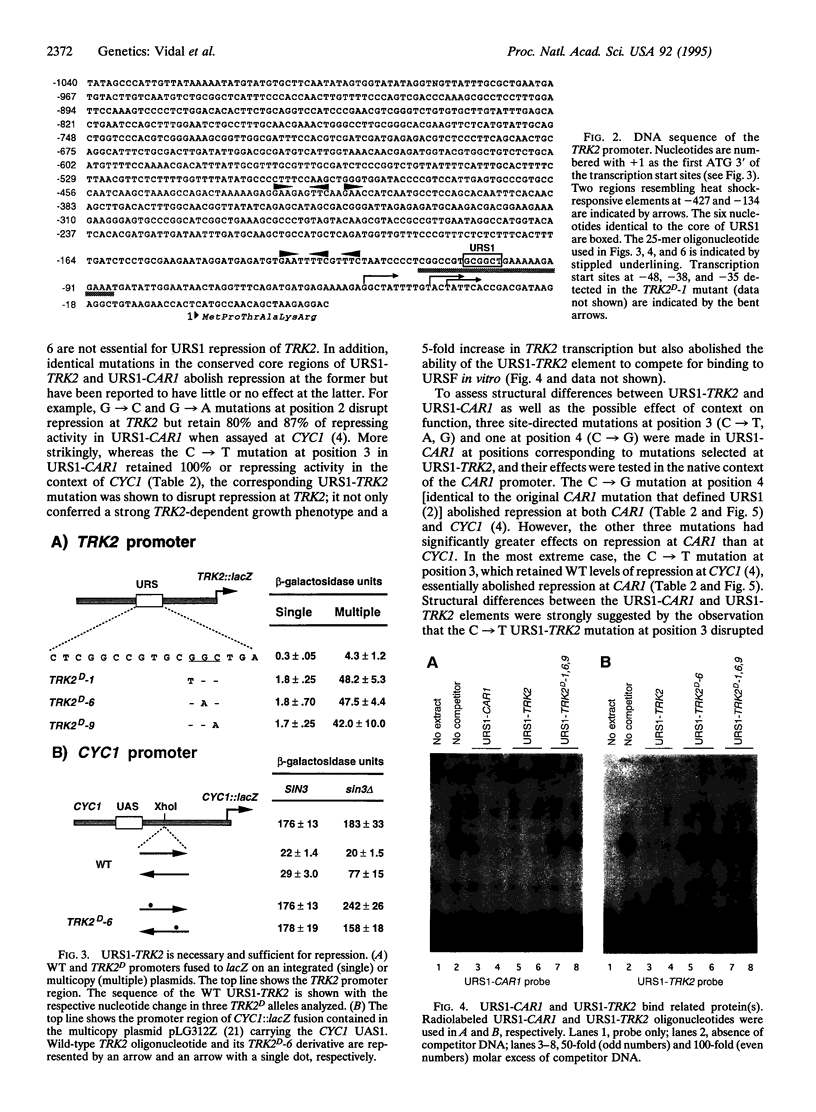


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bowdish K. S., Mitchell A. P. Bipartite structure of an early meiotic upstream activation sequence from Saccharomyces cerevisiae. Mol Cell Biol. 1993 Apr;13(4):2172–2181. doi: 10.1128/mcb.13.4.2172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buckingham L. E., Wang H. T., Elder R. T., McCarroll R. M., Slater M. R., Esposito R. E. Nucleotide sequence and promoter analysis of SPO13, a meiosis-specific gene of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1990 Dec;87(23):9406–9410. doi: 10.1073/pnas.87.23.9406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen R., Yokoi T., Holland J. P., Pepper A. E., Holland M. J. Transcription of the constitutively expressed yeast enolase gene ENO1 is mediated by positive and negative cis-acting regulatory sequences. Mol Cell Biol. 1987 Aug;7(8):2753–2761. doi: 10.1128/mcb.7.8.2753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaber R. F., Styles C. A., Fink G. R. TRK1 encodes a plasma membrane protein required for high-affinity potassium transport in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jul;8(7):2848–2859. doi: 10.1128/mcb.8.7.2848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner M. M., Revzin A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981 Jul 10;9(13):3047–3060. doi: 10.1093/nar/9.13.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guarente L. Yeast promoters and lacZ fusions designed to study expression of cloned genes in yeast. Methods Enzymol. 1983;101:181–191. doi: 10.1016/0076-6879(83)01013-7. [DOI] [PubMed] [Google Scholar]
- Herskowitz I. A regulatory hierarchy for cell specialization in yeast. Nature. 1989 Dec 14;342(6251):749–757. doi: 10.1038/342749a0. [DOI] [PubMed] [Google Scholar]
- Hudak K. A., Lopes J. M., Henry S. A. A pleiotropic phospholipid biosynthetic regulatory mutation in Saccharomyces cerevisiae is allelic to sin3 (sdi1, ume4, rpd1). Genetics. 1994 Feb;136(2):475–483. doi: 10.1093/genetics/136.2.475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ko C. H., Gaber R. F. TRK1 and TRK2 encode structurally related K+ transporters in Saccharomyces cerevisiae. Mol Cell Biol. 1991 Aug;11(8):4266–4273. doi: 10.1128/mcb.11.8.4266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luche R. M., Smart W. C., Cooper T. G. Purification of the heteromeric protein binding to the URS1 transcriptional repression site in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7412–7416. doi: 10.1073/pnas.89.16.7412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luche R. M., Smart W. C., Marion T., Tillman M., Sumrada R. A., Cooper T. G. Saccharomyces cerevisiae BUF protein binds to sequences participating in DNA replication in addition to those mediating transcriptional repression (URS1) and activation. Mol Cell Biol. 1993 Sep;13(9):5749–5761. doi: 10.1128/mcb.13.9.5749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luche R. M., Sumrada R., Cooper T. G. A cis-acting element present in multiple genes serves as a repressor protein binding site for the yeast CAR1 gene. Mol Cell Biol. 1990 Aug;10(8):3884–3895. doi: 10.1128/mcb.10.8.3884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasmyth K., Stillman D., Kipling D. Both positive and negative regulators of HO transcription are required for mother-cell-specific mating-type switching in yeast. Cell. 1987 Feb 27;48(4):579–587. doi: 10.1016/0092-8674(87)90236-4. [DOI] [PubMed] [Google Scholar]
- Park H. O., Craig E. A. Positive and negative regulation of basal expression of a yeast HSP70 gene. Mol Cell Biol. 1989 May;9(5):2025–2033. doi: 10.1128/mcb.9.5.2025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park H. O., Craig E. A. Transcriptional regulation of a yeast HSP70 gene by heat shock factor and an upstream repression site-binding factor. Genes Dev. 1991 Jul;5(7):1299–1308. doi: 10.1101/gad.5.7.1299. [DOI] [PubMed] [Google Scholar]
- Perisic O., Xiao H., Lis J. T. Stable binding of Drosophila heat shock factor to head-to-head and tail-to-tail repeats of a conserved 5 bp recognition unit. Cell. 1989 Dec 1;59(5):797–806. doi: 10.1016/0092-8674(89)90603-x. [DOI] [PubMed] [Google Scholar]
- Slater M. R., Craig E. A. Transcriptional regulation of an hsp70 heat shock gene in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1987 May;7(5):1906–1916. doi: 10.1128/mcb.7.5.1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spevak W., Fessl F., Rytka J., Traczyk A., Skoneczny M., Ruis H. Isolation of the catalase T structural gene of Saccharomyces cerevisiae by functional complementation. Mol Cell Biol. 1983 Sep;3(9):1545–1551. doi: 10.1128/mcb.3.9.1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sternberg P. W., Stern M. J., Clark I., Herskowitz I. Activation of the yeast HO gene by release from multiple negative controls. Cell. 1987 Feb 27;48(4):567–577. doi: 10.1016/0092-8674(87)90235-2. [DOI] [PubMed] [Google Scholar]
- Strich R., Slater M. R., Esposito R. E. Identification of negative regulatory genes that govern the expression of early meiotic genes in yeast. Proc Natl Acad Sci U S A. 1989 Dec;86(24):10018–10022. doi: 10.1073/pnas.86.24.10018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Point mutation generates constitutive expression of an inducible eukaryotic gene. Proc Natl Acad Sci U S A. 1985 Feb;82(3):643–647. doi: 10.1073/pnas.82.3.643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Ubiquitous upstream repression sequences control activation of the inducible arginase gene in yeast. Proc Natl Acad Sci U S A. 1987 Jun;84(12):3997–4001. doi: 10.1073/pnas.84.12.3997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vidal M., Buckley A. M., Hilger F., Gaber R. F. Direct selection for mutants with increased K+ transport in Saccharomyces cerevisiae. Genetics. 1990 Jun;125(2):313–320. doi: 10.1093/genetics/125.2.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vidal M., Gaber R. F. RPD3 encodes a second factor required to achieve maximum positive and negative transcriptional states in Saccharomyces cerevisiae. Mol Cell Biol. 1991 Dec;11(12):6317–6327. doi: 10.1128/mcb.11.12.6317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vidal M., Strich R., Esposito R. E., Gaber R. F. RPD1 (SIN3/UME4) is required for maximal activation and repression of diverse yeast genes. Mol Cell Biol. 1991 Dec;11(12):6306–6316. doi: 10.1128/mcb.11.12.6306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winston F., Chumley F., Fink G. R. Eviction and transplacement of mutant genes in yeast. Methods Enzymol. 1983;101:211–228. doi: 10.1016/0076-6879(83)01016-2. [DOI] [PubMed] [Google Scholar]
- Wright C. F., Zitomer R. S. A positive regulatory site and a negative regulatory site control the expression of the Saccharomyces cerevisiae CYC7 gene. Mol Cell Biol. 1984 Oct;4(10):2023–2030. doi: 10.1128/mcb.4.10.2023. [DOI] [PMC free article] [PubMed] [Google Scholar]




