Abstract
Inflammasome is an intracellular signaling complex of the innate immune system. Activation of inflammasomes promotes the secretion of interleukin 1β (IL-1β) and IL-18 and triggers pyroptosis. Caspase-1 and -11 (or -4/5 in human) in the canonical and non-canonical inflammasome pathways, respectively, are crucial for inflammasome-mediated inflammatory responses. Here we report that gasdermin D (GSDMD) is another crucial component of inflammasomes. We discovered the presence of GSDMD protein in nigericin-induced NLRP3 inflammasomes by a quantitative mass spectrometry-based analysis. Gene deletion of GSDMD demonstrated that GSDMD is required for pyroptosis and for the secretion but not proteolytic maturation of IL-1β in both canonical and non-canonical inflammasome responses. It was known that GSDMD is a substrate of caspase-1 and we showed its cleavage at the predicted site during inflammasome activation and that this cleavage was required for pyroptosis and IL-1β secretion. Expression of the N-terminal proteolytic fragment of GSDMD can trigger cell death and N-terminal modification such as tagging with Flag sequence disrupted the function of GSDMD. We also found that pro-caspase-1 is capable of processing GSDMD and ASC is not essential for GSDMD to function. Further analyses of LPS plus nigericin- or Salmonella typhimurium-treated macrophage cell lines and primary cells showed that apoptosis became apparent in Gsdmd−/− cells, indicating a suppression of apoptosis by pyroptosis. The induction of apoptosis required NLRP3 or other inflammasome receptors and ASC, and caspase-1 may partially contribute to the activation of apoptotic caspases in Gsdmd−/− cells. These data provide new insights into the molecular mechanisms of pyroptosis and reveal an unexpected interplay between apoptosis and pyroptosis.
Keywords: GSDMD, inflammasome, pyroptosis, apoptosis, IL-1β, caspase-1, caspase-11
Introduction
Inflammasome is a large multiprotein complex that serves critical roles in host defense against microbial pathogens1. Inflammasomes control the maturation and secretion of proinflammatory cytokines interleukin (IL)-1β and IL-18 and induce an inflammatory cell death mode termed as pyroptosis2. Pyroptosis is a type of programmed cell death that features pore formation on the plasma membrane, cell swelling and plasma membrane disruption, similar to that of necrosis but not apoptosis3. IL-1β has been known for a long time as a critical pyrogen. IL-1β and IL-18 affect innate and adaptive immunities through a multitude of mechanisms that contribute greatly to host responses against infected pathogens4. Although the role of pyroptosis in host defense is still largely unknown, it was shown to restrict in vivo replication and dispersion of microbes5. The proinflammatory effect of IL-1β/IL-18 and pyroptosis could contribute to the development of autoimmune and inflammatory diseases6.
Activation of inflammasomes often requires a priming signal induced by Toll-like receptors. Different subsets of inflammasomes contain different cytosolic pattern-recognition receptors and their assembly is initiated by different stimuli7. Members of the Nod-like receptor (NLR) family and the HIN-200 family are receptors in inflammasomes to recognize a diversity of pathogen- or danger-associated molecular patterns8. The NLRP3 inflammasome is assembled in response to a broad range of microbial pathogens and medically relevant substances such as crystalline. The NAIP-NLRC4 inflammasome forms upon cytosolic detection of bacterial flagellin or the rod and needle components of bacterial type III secretion systems expressed by intracellular pathogens such as Salmonella typhimurium. The NLRP1b inflammasome recognizes anthrax lethal toxin (LeTx) from Bacillus anthracis. The AIM2 or IFI16 inflammasome is activated when cytosolic or nuclear DNA is detected or by certain intracellular bacteria. Pyrin inflammasome senses bacterial modifications of host Rho GTPases. Once assembled, inflammasomes serve as activating platforms for pro-caspase-1. The activated caspase-1 then processes the precursors of IL-1β/18 to their matured forms9.
Caspase-1 is the inflammatory caspase crucial to canonical inflammasome-mediated pyroptosis and cytokine maturation2. Caspase-11 (also known as caspase-4 or -5 in human), another inflammatory caspase, is the core component of non-canonical inflammasome. Caspase-11 directly acts as a receptor of cytosolic bacterial lipopolysaccharide (LPS) and is activated by binding to LPS10. Caspase-11 can trigger pyroptosis but its effect on IL-1β/18 maturation is indirect because this process requires NLRP3-dependent activation of caspase-111.
ASC (also known as PYCARD or TMS-1) is a pyrin and CARD domain-containing protein that contributes to the assembly of inflammasomes12. ASC bridges pro-caspase-1 and pyrin-containing receptors such as NLRP3 and AIM2 and thus is required by these receptors to recruit pro-caspase-1. However, the CARD-containing receptors NLRC4 and NLRP1 can directly interact with pro-caspase-1. During inflammasome activation, ASC molecules aggregate and form ASC specks (also known as ASC foci or pyroptosomes). The ASC speck is required for pro-caspase-1 activation. However, ASC is not essential for IL-1β/18 maturation and pyroptosis induced by NLRC4 or NLRP1b inflammasome13.
Despite intensive studies of inflammasomes, the regulating mechanisms of IL-1β/IL-18 production and pyroptosis after caspase-1 or caspase-11 activation are unknown. Interference between pyroptosis and other types of programmed cell death has not been described. Here we identified Gasdermin D (GSDMD) as a new component of inflammasomes. It is required for pyroptosis and IL-1β secretion but plays no role in IL-1β processing. We showed that GSDMD was recruited to NLRP3 inflammasome with kinetics similar to caspase-1 after LPS plus nigericin stimulation. GSDMD was cleaved by pro-caspase-1 and most likely also by caspase-1 in inflammasomes, and the proteolytic cleavage of GSDMD released its N-terminal fragment to mediate pyroptosis and IL-1β secretion. In addition, gene deletion of GSDMD unmasked inflammasome-induced apoptosis, indicating a suppression of apoptotic pathway by pyroptosis.
Results
Identification of GSDMD as a required component of NLRP3 inflammasome for pyroptosis and IL-1β production
Canonical inflammasome activation primarily occurs in macrophages and dendritic cells8. Nigericin, an antibiotic derived from Streptomyces hygroscopicus, can activate NLRP3 inflammasome in LPS-primed macrophages with features of caspase-1 activation and caspase-1-mediated pyroptosis and IL-1β secretion14. J774 macrophage cell line behaves similarly to bone marrow-derived macrophages (BMDM) in terms of inflammasome activation15. RAW264.7 macrophage cell line has no ASC gene expression but it resembled BMDM when ASC gene was ectopically introduced into it (Supplementary information, Figure S1A)16. The ASC-reconstituted RAW264.7 cell line was termed as RAW-asc. We knocked out NLRP3 in J774 cell line using clustered regularly interspaced palindromic repeat (CRISPR)-Cas9 method and reconstituted NLRP3 expression with C-terminally Flag-tagged NLRP3. Similar to wild-type J774 cells, inflammasome activation occurred in the NLRP3-Flag J774 cell line upon nigericin treatment (Supplementary information, Figure S1B).
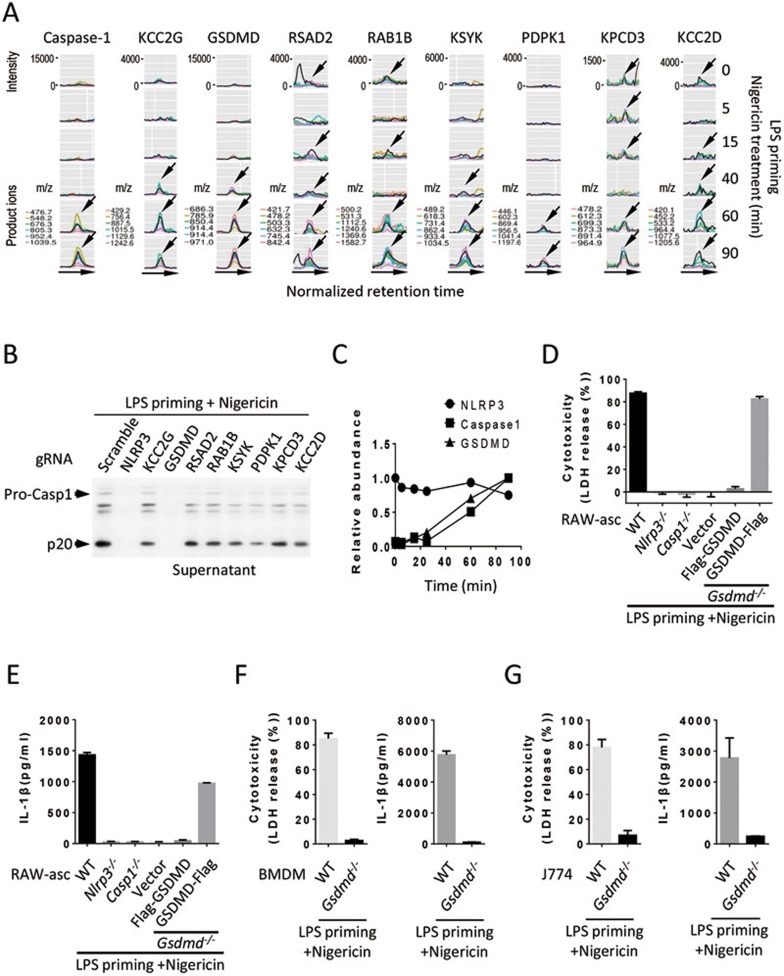
To identify new component of inflammasomes, we analyzed the assembly of NLRP3 inflammasome by high-sensitive quantitative mass spectrometry (MS). We primed NLRP3-Flag J774 cells with LPS and then treated the cells with nigericin for different periods of time. NLRP3 was immunoprecipitated with anti-Flag-M2 beads and the immune complexes were eluted with Flag peptides and subjected to quantitative MS analysis. This analysis detected more than 20 000 peptides. Our recently-developed software Group-DIA17 was used to extract the information of the amounts of different peptides detected in the NLRP3 immunocomplex and to select candidate proteins whose amount time-dependently increased in the NLRP3 immunocomplex. Nine proteins were picked by Group-DIA as the candidates that were recruited to the NLRP3 complex. We pulled out the quantitative data of these nine proteins from the data files and examined the data manually. The extracted ion chromatogram (XIC) peaks of multiple product ions of one representative peptide for each of these proteins were shown in Figure 1A. The level of peptides should correlate with XIC intensities of their product ions. Caspase-1 was among these nine proteins, which supported the effectiveness of our method (Figure 1A). Among the rest eight proteins, RSAD2, RAB1B, KPCD3 and KCC2D had some background binding to NLRP3 at time 0 and their increase in amount at different time points was not smooth. KCC2G, GSDMD, KSYK and PDPK1 were time-dependently recruited to NLRP3, but levels of these proteins except GSDMD were much lower than that of caspase-1. We used CRISPR-Cas9 to knock out each of these eight genes in RAW-asc cells and analyzed p20 caspase-1 fragment released to the cell culture medium by immunoblotting after the cells were treated with LPS plus nigericin. Similar to the knockout of Nlrp3, GSDMD deletion blocked the release of p20 caspase-1 to the culture medium while the knockouts of other seven genes had no effect (Figure 1B), suggesting that GSDMD is involved in inflammasome activation.
Figure 1.
Identification of GSDMD as a component of NLRP3 inflammasome and determination of the requirement of GSDMD in LPS plus nigericin-induced pyroptosis and IL-1β production. (A) Finding protein candidates that were recruited to NLRP3 complex by quantitative MS. NLRP3 complex was affinity-purified from NLRP3-Flag J774 cells after LPS (100 ng/ml) priming for 4 h and then nigericin (10 μM) stimulation for 0, 5, 15, 40, 60, or 90 min, digested with trypsin and subjected to high-sensitive quantitative MS analysis. MS data was analyzed with Group-DIA and peptide intensities were extracted. The results revealed nine candidate proteins whose amount time-dependently increased in NLRP3 complex. The extracted ion chromatogram (XIC) peaks of multiple product ions of one representative peptide for each of these proteins were shown. Arrows indicate the co-eluting XIC peaks of product ions of corresponding representative peptides. (B) GSDMD is involved in inflammasome activation. The genes as indicated were knocked out by CRISPR-Cas9 in RAW-asc cells. The cells were primed with LPS for 4 h and then treated with nigericin for 2 h. The release of p20 caspase-1 into the culture media was measured by immunoblotting using anti-caspase-1 antibody, which was used to assay pyroptosis. (C) Time-dependent recruitment of GSDMD and caspase-1 to NLRP3 complex. Relative abundance of NLRP3, caspase-1 and GSDMD proteins in NLRP3 immunocomplex across six time points was shown. (D) Requirement of GSDMD in pyroptosis. LDH released from RAW-asc cells with different gene deletion or gene reconstitution was measured after the cells were treated as in B. RAW-asc cells with genotype of WT, Nlrp3−/−, Caspase-1−/−(Casp1−/−), Gsdmd−/− cells reconstituted with N-terminally Flag-tagged GSDMD (Flag-GSDMD) or C-terminally Flag-tagged GSDMD (GSDMD-Flag) or a control vector were used in the experiments. (E) Requirement of GSDMD in IL-1β production. Culture supernatants of the cells described in D were analyzed by IL-1β ELISA kit. (F) Pyroptosis and IL-1β in the culture supernatants of WT and Gsdmd−/− BMDM were measured as in D, E. (G) Pyroptosis and IL-1β in the culture supernatants of WT and Gsdmd−/− J774 cells were measured as in D and E. Graphs show mean ± SD of triplicate wells and represent three independent experiments.
GSDMD is a 53 KDa gasdermin domain-containing protein with unknown biological function. We noticed that GSDMD peptides identified in our MS analyses are all located in the N-terminal half of GSDMD (Supplementary information, Figure S2A). Based on the quantitative MS data, GSDMD was time-dependently recruited to the NLRP3 complex with similar kinetics to that of caspase-1 (Figure 1C). We further analyzed GSDMD-knockout RAW-asc line (Gsdmd−/−) (Supplementary information, Figure S2B) and confirmed that Gsdmd−/− cells are resistant to nigericin-induced pyroptosis (Figure 1D), similar to what was observed in Nlrp3−/− or caspase-1−/− (Casp1−/−) RAW-asc cells (Figure 1D and Supplementary information, Figure S2B). To further validate that the resistance to LPS plus nigericin-induced pyroptosis is due to GSDMD gene deletion, we reconstituted GSDMD gene by ectopic expression of N-terminally (Flag-GSDMD) or C-terminally Flag-tagged GSDMD (GSDMD-Flag). Interestingly, the GSDMD-Flag but not the Flag-GSDMD can reconstitute GSDMD's function in pyroptosis (Figure 1D and Supplementary information, Figure S2B), indicating that the structure of N-terminal end is important for GSDMD to function.
We then analyzed the requirement of GSDMD in inflammasome-mediated IL-1β production. The cells were primed with LPS and then treated with nigericin. IL-1β in the culture medium was almost completely eliminated in Gsdmd−/− RAW-asc cells and its production was restored when GSDMD-Flag but not the Flag-GSDMD was ectopically expressed in Gsdmd−/− cells (Figure 1E). Nlrp3−/− and Casp1−/− RAW-asc cells were included as controls and showed no production of IL-1β. The requirement of GSDMD in LPS plus nigericin-induced pyroptosis and IL-1β production was also confirmed by using BMDM derived from Gsdmd−/− C57BL/6 mice and Gsdmd−/− J774 cells (Figure 1F and 1G). Collectively, our data demonstrated that GSDMD is recruited to NLRP3 inflammasome after LPS-primed macrophages are treated with nigericin, and that GSDMD is required for NLRP3 inflammasome to mediate pyroptosis and IL-1β production.
To determine whether GSDMD is required for the activation of different subsets of inflammasomes, we examined IL-1β production and pyroptosis in RAW-asc cells upon three different stimuli including intracellular LPS that activates non-canonical inflammasomes. GSDMD deletion blocked pyroposis and IL-1β production induced by all the stimuli tested (Supplementary information, Figure S3A-S3C). Thus, GSDMD is a common element in inflammasome pathways.
GSDMD has no effect on pro-caspase-1 auto-processing and caspase-1-mediated maturation of IL-1β
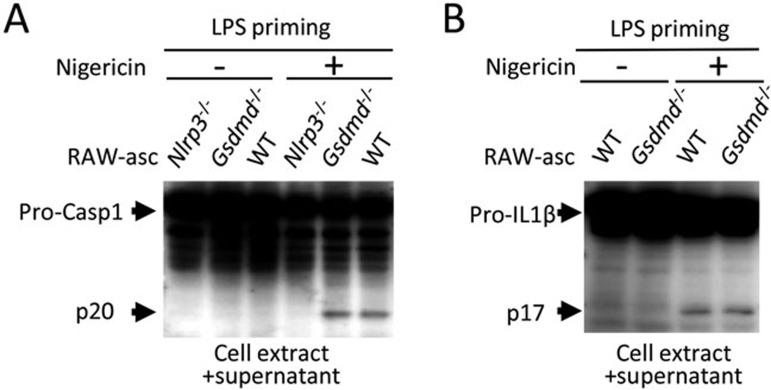
To understand how GSDMD regulates pyroptosis and IL-1β production, we analyzed pro-caspase-1 cleavage in LPS plus nigericin-treated RAW-asc cells and found that unlike Nlrp3 knockout, deletion of Gsdmd did not affect pro-caspase-1 cleavage (Figure 2A). This result indicates that the function of GSDMD in inflammasome pathway is downstream of NLRP3 and either downstream of or parallel to caspase-1 auto-proteolytic activation. Caspase-1 is known to be responsible for IL-1β proteolytic maturation. To examine whether the processing of caspase-1 in Gsdmd−/− cells leads to maturation of IL-1β, we analyzed IL-1β maturation. As expected, we detected cleavage of pro-IL-1β in LPS plus nigericin-treated Gsdmd−/− cells (Figure 2B). Since IL-1β was not detected in the culture media of Gsdmd−/− cells (Figure 1E-1G), this result indicates that GSDMD controls the release of matured IL-1β into culture medium.
Figure 2.
GSDMD is not required for proteolytic maturation of caspase-1 and IL-1β. (A) Processing of caspase-1 in Gsdmd−/− cells. Culture supernatants together with their corresponding cell extracts from WT, Nlrp3−/− and Gsdmd−/− RAW-asc cells treated as in Figure 1B were analyzed by immunoblotting with anti-caspase-1 antibody. (B) Processing of IL-1β in Gsdmd−/− cells. Culture supernatants together with their corresponding cell extracts from WT and Gsdmd−/− RAW-asc cells treated as in Figure 1B were analyzed by immunoblotting with anti-IL-1β antibody.
Cleavage of GSDMD by caspase-1 is required for pyroptosis and release of matured IL-1β
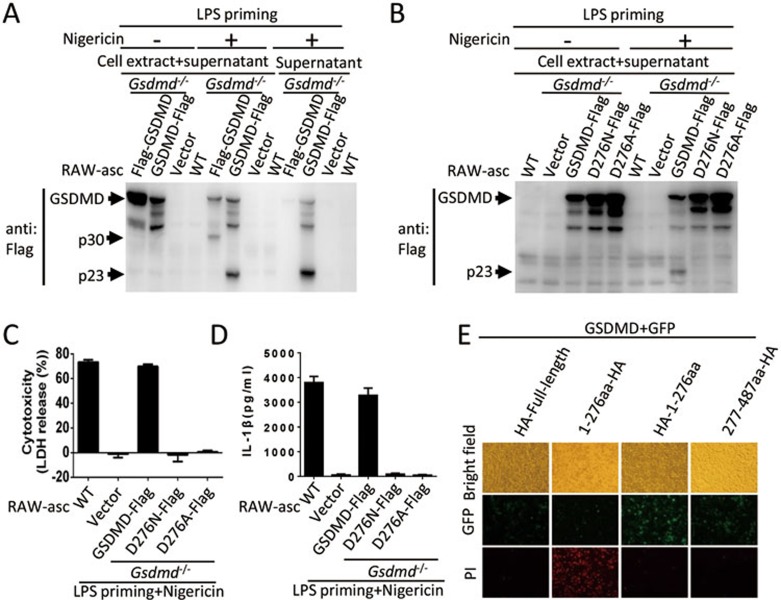
A proteomics analysis by Agard et al.18 revealed that GSDMD is a substrate of caspase-1. We analyzed GSDMD protein in Gsdmd−/− cells expressing N-Flag GSDMD or C-Flag GSDMD before and after LPS plus nigericin treatment. Both N-Flag and C-Flag GSDMD were cleaved after the treatment (Figure 3A) though no pyroptosis induction and IL-1β production occurred in the cells expressing N-Flag GSDMD (Figure 1D and 1E). Consistent with the occurrence of pyroptosis, only the proteolytic product of C-Flag GSDMD was released into the culture medium (supernatant; Figure 3A). The size of N-terminal and C-terminal fragments fits the prediction by Agard et al.18. We mutated the predicted cleavage site D276 in GSDMD to A and N and both mutations blocked GSDMD cleavage (Figure 3B). The D276 mutations abolished the function of GSDMD as reconstituted GSDMD expression with D276N or D276A mutant of GSDMD cannot restore the induction of pyroptosis and IL-1β secretion by LPS plus nigericin (Figure 3C and 3D), demonstrating that cleavage of GSDMD by caspase-1 is a critical step in inflammasome signaling.
Figure 3.
Cleavage of GSDMD at D276 is required for pyroptosis and IL-1β secretion and the N-terminal proteolytic fragment of GSDMD executes cell death. (A) Processing of GSDMD. WT RAW-asc cells and Gsdmd−/− RAW-asc cells reconstituted with Flag-GSDMD, GSDMD-Flag or a control vector were treated as in Figure 1B. Cell extracts together with their corresponding culture supernatants or supernatants alone were analyzed by immunoblotting with anti-Flag antibody. (B) D276 is the cutting site in GSDMD upon NLRP3 inflammasome activation. Gsdmd−/− RAW-asc cells reconstituted with the expression of GSDMD-Flag, D276N mutant of GSDMD (D276N-Flag) or D276A mutant of GSDMD (D276A-Flag) were treated as in Figure 1B. Cell extracts together with their corresponding culture supernatants were analyzed by immunoblotting with anti-Flag antibody. (C) Requirement of GSDMD cleavage in pyroptosis. LDH release was measured in the cells described in B. (D) Requirement of GSDMD cleavage in IL-1β secretion. The same as in C except that IL-1β in the culture media was measured by ELISA. (E) Pyroptosis induced by N-terminal domain of GSDMD. Green fluorescence protein (GFP) and one of the GSDMD fragments were co-expressed in 293T cells and images were taken 16 h after transfection. Graphs show mean ± SD of triplicate wells and represent three independent experiments.
N-terminal proteolytic fragment of GSDMD triggers pyroptosis
To explore the function of GSDMD cleavage by caspase-1 in pyroptosis, we expressed full-length, N-terminal or C-terminal fragment of GSDMD in 293T cells. The N-fragment but not the C-fragment or the full-length GSDMD caused propidium iodide (PI)-positive cell death (Figure 3E), indicating that N-terminal fragment is the executor of pyroptosis. Consistent with the data shown in Figure 1D, the N-terminal tagging also blocked the cytotoxicity of N-fragment of GSDMD (Figure 3E). It can be concluded that the N-terminal structure is essential for GSDMD to execute pyroptosis and removal of C-terminal domain is the mechanism for N-terminal domain of GSDMD to function.
GSDMD cleavage can be mediated by pro-caspase-1 and does not need ASC
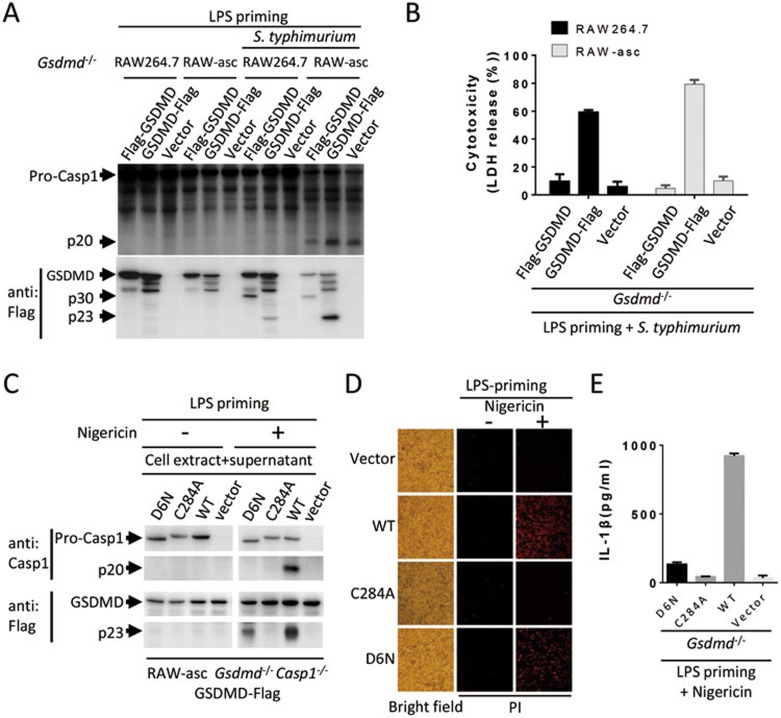
Since caspase-1 auto-cleavage was not found in S. typhimurium-induced pyroptosis in ASC-deleted cells19, we examined whether pro-caspase-1 is capable of cleaving GSDMD. RAW264.7 cell line was used as ASC-deficient cells16. N-Flag and C-Flag GSDMD were expressed in Gsdmd−/− RAW264.7 and Gsdmd−/− RAW-asc cells. Then the cells were primed with LPS, followed by S. typhimurium treatment. As anticipated, we detected pro-caspase-1 cleavage in RAW-asc but not in RAW264.7 cells (Figure 4A). Interestingly, we detected cleavage of both Flag-GSDMD and GSDMD-Flag in both RAW264.7 and RAW-asc cells (Figure 4A), indicating that the cleavage of GSDMD can occur without ASC. It also suggested that pro-caspase-1 can cleave GSDMD. Consistent with the data that N-terminal tagging destroys the function of GSDMD and that GSDMD was cleaved in both RAW264.7 and RAW-asc cells, GSDMD-Flag- but not Flag-GSDMD-reconstituted RAW264.7 and RAW-asc cells underwent pyroptosis upon LPS plus S. typhimurium treatment (Figure 4B). Thus the cleavage of GSDMD could be parallel to the activation of caspase-1, and ASC is not required for GSDMD activation.
Figure 4.
GSDMD can be cleaved by pro-caspase-1. (A) GSDMD is cleaved in ASC-deficient cells in the absence of pro-caspase-1 processing. Gsdmd−/− RAW264.7 and Gsdmd−/− RAW-asc cells reconstituted with the expression of Flag-GSDMD, GSDMD-Flag or a control vector were primed with LPS for 4 h followed by S. typhimurium (100 MOI) treatment for 2 h. The cells were analyzed by immunoblotting with anti-caspase-1 and anti-Flag antibodies. (B) LDH release was measured in the cells described in A. (C) The auto-cleave-deficient pro-caspase-1 mutant can cleave GSDMD. Gsdmd and caspase-1 double knockout RAW-asc cells (Gsdmd−/− Casp1−/−) were reconstituted with GSDMD-Flag expression first and then with WT, auto-cleavage-deficient mutant D6N, and enzymatic dead mutant C284A of pro-caspase-1, respectively. The cells were treated as in Figure 1B and analyzed by immunoblotting with anti-caspase-1 and anti-Flag antibodies. (D) Pyroptosis can be mediated by auto-cleavage-deficient pro-caspase-1. The cells described in C were stained with PI and analyzed under microscope. (E) IL-1β production cannot be mediated by auto-cleavage-deficient pro-caspase-1. IL-1β in the culture media of the cells described in C was measured by ELISA. Graphs show mean ± SD of triplicate wells and represent three independent experiments.
To confirm that pro-caspase-1 is able to cleave GSDMD, we reconstituted Gsdmd and caspase-1 double knockout RAW-asc cells (Gsdmd−/− Casp1−/−) with GSDMD-Flag expression first and then with WT, auto-cleavage-deficient mutant D6N, and enzymatic dead mutant C284A of caspase-1, respectively. The cells were primed with LPS and then stimulated with nigericin. As anticipated, caspase-1 cleavage was only observed in WT caspase-1-reconstituted cells (Figure 4C). However, GSDMD was cleaved in both WT and D6N caspase-1-reconstituted cells (Figure 4C). As a negative control, no GSDMD cleavage was observed in C284A caspase-1-reconstituted cells. We examined pyroptosis in these cells and found cell death in WT and D6N caspase-1-reconstituted cells (Figure 4D). As anticipated, D6N cannot mediate IL-1β production (Figure 4E) as D6N caspase-1 cannot process pro-IL-1β. We noticed that D6N mutant-reconstituted cells had less GSDMD cleavage and pyroptosis upon LPS plus nigericin treatment, which suggested that either matured caspase-1 participated in the cleavage of GSDMD or the D6N mutant has less activity than WT pro-caspase-1. Nonetheless, we can conclude that pro-caspase-1 is able to cleave and activate GSDMD.
Depletion of GSDMD unmasks inflammasome-induced apoptosis
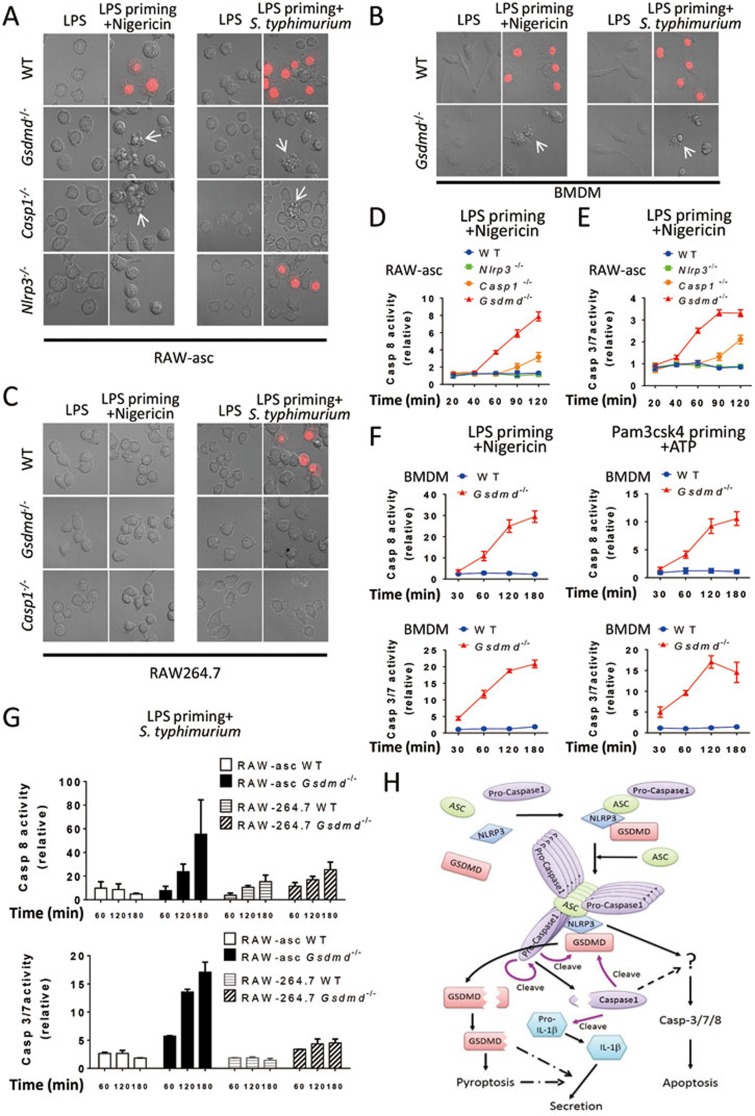
We investigated pyroptosis in WT, Nlrp3−/−, casp1−/−, Gsdmd−/− RAW-asc cells via microscopy and made a couple of interesting observations (Figure 5A). LPS plus nigericin-induced pyroptosis was completely abolished by NLRP3 deletion. The morphology of nigericin-treated Nlrp3−/− cells was almost the same as that of untreated cells (Figure 5A). Both caspase-1 deletion and GSDMD deletion blocked pyroptosis (shown by PI-positive cells), but the morphology of the cells still changed to be apoptotic-like (Figure 5A). We counted the number of apoptotic-like cells and pyroptotic cells in Gsdmd−/− (or Casp1−/−) and WT cells, respectively, after 4 h of LPS priming and 1 h of nigericin treatment. Results showed that the number of apoptotic-like cells in Gsdmd−/− cells was about 2/3 of that of pyroptotic cells in WT cells. The number of apoptotic-like cells in Casp1−/− cells was about half of that in Gsdmd−/− cells. Unlike Nlrp3−/− cells, Casp1−/− and Gsdmd−/− cells cannot efficiently re-grow after removal of the inflammasome stimulus. We used S. typhimurium to treat LPS-primed cells and obtained the same result except that S. typhimurium-induced pyroptosis does not need NLRP3 (Figure 5A), which is consistent with the published result19.
Figure 5.
GSDMD deletion unmasks inflammasome-induced apoptosis. (A) Apoptotic-like cells in GSDMD- or Caspase-1-deleted RAW-asc cells. WT, Gsdmd−/−, Casp1−/−, and Nlrp3−/− RAW-asc cells were used in the experiments. Images were taken after LPS priming for 4 h and then nigericin or S. typhimurium treatment for 2 h. PI was added to detect the loss of plasma membrane integrity. Arrows point to apoptotic-like cells. (B) Apoptotic-like cells in GSDMD-deleted BMDM. The same as in A except that WT and Gsdmd−/− BMDM were used and nigericin or S. typhimurium stimulation time was 1 h. (C) No apoptotic-like cells in GSDMD- or casppase-1-deleted RAW264.7 cells. The same as in A except that WT, Gsdmd−/− and Casp1−/− RAW264.7 cells were used in the experiments. (D) Apoptotic caspase-8 is activated in Gsdmd−/− and Casp1−/− RAW-asc cells after LPS plus nigericin stimulation. WT, Nlrp3−/−, Casp1−/− and Gsdmd−/− RAW-asc cells were primed with LPS for 4 h and then treated with nigericin for different time periods as indicated. The activities of caspase-8 were measured and shown. (E) Apoptotic caspase-3/7 is activated in Gsdmd−/− and Casp1−/− RAW-asc cells. The same as in D except that the activities of caspase-3/7 were measured and shown. (F) Apoptotic caspases are activated in inflammasome stimuli-treated Gsdmd−/− BMDM. WT and Gsdmd−/− BMDM were primed with LPS or Pam3csk4 (1 μg/ml) for 5 h and stimulated with nigericin or ATP (5 mM), respectively, for different periods of time as indicated. The activities of caspase-8 and caspase-3/7 were measured and shown. (G) Apoptotic caspases are not activated in ASC-deficient cells. WT and Gsdmd−/− RAW264.7 cells were primed with LPS for 4 h and stimulated with S. typhimurium for different time periods as indicated. WT and Gsdmd−/− RAW-asc cells were included for comparison. The activities of caspase-8 and caspase-3/7 were measured and shown. (H) Proposed model for pyroptosis and IL-1β production induced by NLRP3 inflammasome pathway. Graphs show mean ± SD of triplicate wells and represent three independent experiments.
We analyzed BMDM from WT and Gsdmd−/− mice and also observed apoptotic-like morphological change of Gsdmd−/− cells after LPS plus nigericin or LPS plus S. typhimurium treatment (Figure 5B). To assess the role of ASC, we analyzed RAW264.7 cells and did not find apoptotic-like cells after LPS plus nigericin treatment (Figure 5C). S. typhimurium treatment induced pyroptosis in RAW264.7 cells but it also cannot lead to apoptotic-like morphological changes in Gsdmd−/− or Casp1−/− cells (Figure 5C), indicating the involvement of ASC in triggering this non-pyroptotic pathway.
To determine whether apoptotic pathway was activated when pyroptosis was blocked by GSDMD deletion, we analyzed the activities of caspase-8 and caspase-3 or -7, these so-called apoptotic caspases. WT, Gsdmd−/−, Nlrp3−/−, and Casp1−/− RAW-asc cells were stimulated with LPS plus nigericin for different periods of time, and the activities of caspase-8 and caspase-3/7 were measured (Figure 5D and 5E). LPS plus nigericin treatment increased significantly the activities of caspase-8 and caspase-3/7 in Gsdmd−/− cells and slightly the activities of these caspases in Casp1−/− cells, but it had no influence in WT and Nlrp3−/− cells. BMDMs appear to be more susceptible to apoptosis than RAW-asc cells as much higher activation of the apoptotic caspases was observed in Gsdmd−/− but not WT BMDM (Figure 5F, left panel). The activation of apoptotic pathway seems to be common in Gsdmd−/− cells when the cells were treated with different subsets of inflammasome stimuli since ATP also induced the activation of caspase-8 and caspase-3/7 in Pam3CSK4-primed BMDM (Figure 5F). We also assessed the requirement of ASC in the induction of apoptosis in Gsdmd−/− cells and found that unlike RAW-asc cells, Gsdmd−/− RAW264.7 cells showed no or little S. typhimurium- induced activation of caspase-8 and caspase-3/7 (Figure 5G). Thus, Nlrp3 (or other receptors) and ASC together can activate the apoptotic pathway when GSDMD is absent.
GSDMD deletion can effectively block nigericin-induced LDH release in LPS-primed BMDM at 1 h time point but not at later time points (Supplementary information, Figure S4A). Similar phenomenon can be observed in RAW-asc cells although LDH was detected later than that in BMDM (Supplementary information, Figure S4B). The release of LDH in Casp1−/− was delayed and less in comparison with Gsdmd−/− RAW-asc and Nlrp3−/− cells had no LDH release even at much later time points (Supplementary information, Figure S4B). The LDH release in BMDM and RAW-asc cells with different gene deletions correlates very well with the levels of activities of apoptotic caspases in these different cells (Figure 5D-5F). Thus, the release of LDH at later time points could result from secondary necrosis of the apoptotic cells. Indeed, we observed PI-positive cells after apoptotic morphological change of Gsdmd−/− BMDM and RAW-asc cells (Supplementary information, Figure S4C and S4D). Collectively, our data indicated that apoptotic pathway was suppressed in wild-type cells and emerged when GSDMD was deleted. Blocking pyroptosis may not completely block cytolysis since secondary necrosis of apoptosis unmasked by it is still able to disrupt plasma membrane.
Discussion
Activation of pro-caspase-1 is known to be the crucial step for inflammasomes to mediate pyroptosis and IL-1β/IL-18 production. However, how caspase-1 mediates pyroptosis and IL-1β/IL-18 secretion is largely unknown. It was known that cleavage of IL-1β/IL-18 by matured caspase-1 leads to the maturation of these cytokines and their secretion, and pyroptosis is a death mode with plasma membrane disruption which releases cytosolic components including pro- and matured IL-1β/IL-18. Here we identified GSDMD as a component of inflammasomes responsible for not only the execution of pyroptosis but also the secretion of matured IL-1β (Figure 1), pointing out an intrinsic linkage between passive release of cytosolic IL-1β by pyroptosis and active secretion of matured IL-1β. Our data indicate that when macrophages are treated with an inflammasome stimulus such as nigericin, GSDMD is recruited to NLRP3 with similar kinetics to that of pro-caspase-1 (Figure 1C). The recruitment of GSDMD does not require ASC (Figure 4), but ASC-mediated speck formation could increase the number of GSDMD in a given inflammasome, therefore enhancing the activation of GSDMD. We showed that pro-caspase-1 is capable of cleaving GSDMD (Figure 4) and thus proposed that GSDMD is cleaved in inflammasomes. However, matured caspase-1 may also participate in the cleavage of GSDMD since we observed less cleavage of GSDMD in S. typhimurium-treated ASC-deficient cells, in which pro-caspase-1 cannot be processed, than that in RAW-asc cells (Figure 4A). The N-terminal fragment of GSDMD generated by caspase-1 cleavage is responsible for executing pyroptosis and promoting the secretion of matured IL-1β generated by proteolysis by matured caspase-1 (Figure 3). We also observed that deletion of GSDMD unmasked inflammasome-induced apoptosis (Figure 5 and Supplementary information, Figure S4). Our model of how GSDMD functions in inflammasomes is shown in Figure 5H.
In this study, we used quantitative MS to find new components in NLRP3 inflammasome. Our method can efficiently obtain the information of the dynamic changes of proteins in a given signaling complex (Figure 1A). The result of this study suggested that the profile of the changes and amount (intensity) of a protein are important hints for its function. For example, the kinetics of the recruitment of GSDMD and caspase-1 were similar and their amounts recruited to the NLRP3 complex were also about the same (intensities are above 10 000), and we found that they both function in inflammasomes. Although KCC2G, KSYK and PDPK1 were recruited to the NLRP3 complex, their levels were much lower than those of caspase-1 and GSDMD and they were not required for pyroptosis. We believe that our protocol of using quantitative MS can be widely applied to the identification of new components in other signaling pathways.
Our data on GSDMD are consistent with two recent online reports that uncovered the essential role of GSDMD in inflammasome signaling20,21. In one of the publications, Kayagaki et al.20 concluded that although GSDMD can be cleaved by both caspase-1 and -11 (or -4/5 in human), it is fully required for the non-canonical pathway but its involvement in canonical inflammasome signaling is only at early time points of stimulation. However, the data in the other publication by Shi et al.21 indicated that GSDMD is essential for both canonical and non-canonical inflammasome pathways. Our data suggest that GSDMD is equally important to canonical and non-canonical inflammasome signaling (Supplementary information, Figure S3). We also observed LDH release in Gsdmd−/− cells at later time points after inflammasome activation (Supplementary information, Figure S4). However, this LDH release should be at least partially due to secondary necrosis of the apoptosis induced by inflammasome stimuli in the absence of GSDMD, because there is a very good correlation between apoptotic caspase activation and LDH release (Supplementary information, Figure S4A and S4B and Figure 5D-5F).
We showed that the apoptotic pathway in Gsdmd−/− cells is initiated downstream of NLRP3 and ASC (Figure 5). The occurrence of apoptosis in LPS plus nigericin-stimulated Casp1−/− cells is most likely due to the fact that GSDMD cannot be proteolytically processed. On the other hand, the inflammasome-induced apoptosis might be partially contributed by caspase-1 because the activation of apoptotic caspases is higher in Gsdmd−/− cells than that in Casp1−/− cells (Figure 5D and 5E).
Pore formation is a feature of pyroptosis. It is unknown whether the N-terminal fragment of GSDMD is capable of forming pore on the plasma membrane, but GSDMD is likely to mediate the secretion of matured IL-1β and trigger pyroptosis through its effect on the plasma membrane. Although it remains to be experimentally tested, GSDMD might promote IL-1β secretion via the same molecular action as it induces pyroptosis. Meanwhile, the NLRP3 inflammasome could activate an apoptotic pathway when the pyroptotic pathway is blocked. Analogous to the competition between apoptosis and necroptosis22, pyroptosis also suppresses apoptosis during the induction of inflammasomes.
Materials and Methods
Plasmids, antibodies and reagents
cDNAs for mouse Nlrp3, pro-Casp1, pro-IL1β, ASC and Gsdmd were amplified from reverse-transcribed cDNAs of J774 cells. Nlrp3, pro-casp1, pro-IL1β, ASC and Gsdmd cDNAs were inserted into a modified pBOBi plasmid vector with N-terminal or C-terminal 3× Flag or 3× HA tag. Truncation or point mutation of Gsdmd was generated by standard PCR cloning strategy. All plasmids were DNA-sequenced.
Antibody for IL-1β (5129-100) was purchased from Biovision. Antibody for Caspase-1 p20 (clone 4B4) was a kind gift from VM Dixit (Genetech, USA). Antibody for NLRP3 (AG-20B-0006-C100) was purchased from Adipogen. Antibody for GSDMD was purchased from Proteintech. Mouse anti-HA (F-7), anti-GAPDH (6C5) and ASC (sc-22514-R) antibodies were purchased from Santa Cruz Biotechnology, Inc. Mouse anti-Flag M2 and anti-HA beads, and anti-β-Tubulin (T6199) antibodies were purchased from Sigma-Aldrich. Ultrapure LPS from E. coli O111:B4, PI and Cholera Toxin B subunit were purchased from Sigma-Aldrich. Nigericin was purchased from InvivoGen. zVAD was obtained from Calbiochem. Anthrax Lethal Factor and Protective Antigen were purchased from List Biological Laboratories, Inc. CytoTox 96 Non-Radioactive Cytotoxicity Assay kit was purchased from Promega.
Cell cultures
RAW 264.7, J774, and 293T cells were obtained from ATCC. All cells were grown in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and non-essential amino acids. All cells were grown at 37 °C in a 5% CO2 incubator. Transient transfection of 293T cells was performed using calcium phosphate method. Lentiviral infection was used for stable expression. Recombinant lentiviruses were packaged in 293T cells in the presence of helper plasmids (pMDLg, pRSV-REWV and pVSV-G) using a calcium phosphate precipitation method. The transfected cells were cultured for 48 h and the viruses were then collected for infection.
Generation of knockout cell lines using CRISPR-Cas9 technique
The targeting sequence in the gRNA vector was 3′-GTGTTGTCAGGATCTCGCAT-5′ for mouse Nlrp3, 3′-TCTCTAAAAAAGGGCCCC-5′ for mouse Caspase1 and 3′-TGCAACAGCTTCGGAGTCG-5′ for mouse Gsdmd. The plasmids harboring the gene gRNA sequences and Cas9 gene were transfected into 293T in the presence of lentivirus helper plasmids, and the supernatants were collected after 48 h. The viruses were first concentrated and then used to infect J774 and RAW 264.7 cells.
Immunoprecipitation and digestion
NLRP3-3× FLAG-reconstituted J774 cells were seeded at 1 × 107 cells per 15-cm dish in DMEM supplemented with 10% FBS. After 24 h, the cells were first treated with 100 ng/ml LPS for 4 h and then treated with 10 μM Nigericin for 0, 5, 15, 25, 40, 60, 90 min. Ten 15-cm dishes of cells were collected at each time point. After Nigericin stimulation, cells were immediately washed twice with PBS and harvested by scraping and centrifugation at 100× g for 10 min. The cells were washed with PBS and then lysed for 30 min on ice in HBS lysis buffer (12.5 mM HEPES, 150 mM NaCl, 1% Nonidet P-40, pH 7.5) with protease inhibitor cocktail. Cell lysates were spun down at 50 000× g for 30 min. The soluble fraction was collected, and immunoprecipitated overnight with anti-Flag M2 antibody-conjugated agarose at 4 °C. Beads containing protein complexes were washed three times with HBS lysis buffer. Proteins were then eluted twice with 0.15 mg/ml of 3× FLAG peptide with N-terminal biotin tag in HBS lysis buffer for 30 min each time, and the elution was pooled for a final volume of 300 μl. Proteins in the elution were precipitated with 20% trichloroacetic acid (TCA) and the pellet was washed twice with 1 ml cold acetone, and dried in speedvac.
TCA-precipitated proteins were dissolved in 50 μl of 8 M urea in 50 mM NH4HCO3, and 10 mM TCEP (tris(2-carboxyethyl)phosphine) and 40 mM chloroacetamide was added for 30 min at 37 °C. Next, 8 M urea was diluted to 1.6 M urea with NH4HCO3 and trypsin (Washtington) was added at 50:1 ratio of protein to trypsin. Digestion lasted for 16-18 h at 37 °C. The reactions were incubated with 40 μl of Streptavidin-sepharose beads (bed volume) for removal of FLAG peptides. After adding 1% formic acid, the peptides were purified with C18 STAGETips. After desaltion, peptides were eluted with 10% acetonitrile/1% formic acid and dried.
MS and data analysis
Peptides were dissolved in 0.1% formic acid and analyzed by MS in SWATH (the sequential window acquisition of all theoretical mass spectra) mode23. MS analysis was performed on a TripleTOF 5600 (AB Sciex) MS coupled to NanoLC Ultra 2D Plus (Eksigent) HPLC system as previously described24. Peptides first bound to a 5 mm × 500 μm trap column packed with Zorbax C18 5 μm 200 Å resin using 0.1% formic acid/2% acetonitrile in H2O at 10 μl/min for 5 min, and then separated on a about 30 cm × 75 μm in-house pulled emitter-integrated column packed with Magic C18 AQ 3-μm 200-Å resin. The 240-min linear gradient is from 2%-35% buffer B (buffer A: 0.1% formic acid, 5% DMSO in H2O; buffer B: 0.1% formic acid, 5% DMSO in acetonitrile), and the whole run is 256 min.
For SWATH-MS, the MS was operated such that a 250-ms survey scan (TOF-MS) which was collected in 350-1 500 m/z was performed followed by 100 33-ms MS2 experiments which were collected in 100-1 800 m/z, resulting in an about 3.6 s cycle time. These MS2 experiments used the variable isolation window widths to cover the precursor mass range of 400-1 200 m/z. The 100 variable isolation windows are “399.5-409.9,408.9-418.9,417.9-427.4,426.4-436,435-443.6,442.6-450.8,449.8-458,457-464.8,463.8-471.1,470.1-476.9,475.9-482.8,481.8-488.6,487.6-494,493-499,498-504.4,503.4-509.3,508.3-514.3,513.3-519.2,518.2-524.2,523.2-529.1,528.1-534.1,533.1-539,538-543.5,542.5-548.5,547.5-553,552-558,557-562.5,561.5-567,566-571.5,570.5-576,575-580.5,579.5-585,584-589.5,588.5-594,593-598,597-602.5,601.5-607,606-611.1,610.1-615.6,614.6-620.1,619.1-624.6,623.6-628.6,627.6-633.1,632.1-637.6,636.6-642.1,641.1-646.6,645.6-651.1,650.1-655.6,654.6-660.1,659.1-665.1,664.1-669.6,668.6-674.5,673.5-679,678-684,683-688.5,687.5-693.4,692.4-698.4,697.4-703.3,702.3-708.7,707.7-713.7,712.7-719.1,718.1-724.5,723.5-729.9,728.9-735.3,734.3-740.7,739.7-746.5,745.5-751.9,750.9-757.8,756.8-763.6,762.6-769.5,768.5-775.3,774.3-781.2,780.2-787,786-793.3,792.3-800.1,799.1-806.4,805.4-813.1,812.1-820.3,819.3-827.5,826.5-835.2,834.2-843.3,842.3-851.4,850.4-859.9,858.9-868.9,867.9-878.4,877.4-888.3,887.3-899.1,898.1-910.3,909.3-922.9,921.9-936,935-949.5,948.5-963.4,962.4-978.7,977.7-994.9,993.9-1015.6,1014.6-1042.2,1041.2-1070.1,1069.1-1100.7,1099.7-1140.7,1139.7-1196.5”. Ions were fragmented for MS2 experiment in the collision cell using a collision energy according to the equation of a doubly charged peptide, ramped ±15 V from the calculated collision energy.
The wiff files from SWATH-MS were converted to profile mzXML files using proteoWizard MSConvert v. 3.0.4472. mzXML files were analyzed by Group-DIA. Briefly, Group-DIA first aligned the retention time of six runs using MS1 signal, and then extracted precursor-product ion pairs by combing the MS1 and MS2 information of six runs. The pseudo-spectra were subjected to database search using X!tandem (Version 2013.06.15.1, Native and Kscore) which was integrated into Trans-Proteomics Pepline software (Version 4.7.1)25 against the full non-redundant, canonical mouse genome as annotated by UniprotKB/Swiss-Prot (downloaded in September 2014) appendant with common contaminants and reversed sequence decoys (33 864 sequences include decoys). “DECOY” label was added before the name of proteins. The database search parameters were set as follows: carbamidomethylation (C) was set as fixed modification; methionine oxidation was set as variable modification, semi-tryptic peptides and peptides with up to two missed cleavages were allowed, and mass tolerance of the precursor ion and product ion were set at 50 p.p.m. and 0.05 Da, respectively. After database searching, the pepXML files were validated with PeptideProphet26 with “-OAdPE-PPM-p0.05-l7-dDECOY” option, and subsequently combined and rescored using iProphet27 with “DECOY=DECOY” option. The file generated by iProphet was filtered at 1% peptide-spectrum match level. For each peptide precursor, at most six product ions were selected for extraction of quantitation information in SWATH files. For quantitation results, peptides with swath_score ≥ 0.99 were kept for further analysis. The sum of six product ion intensities represents peptide intensities, and the sum of three most intense peptide intensities represents protein intensities.
The protein intensity values were transformed with Log2, and normalized with the values at 0 min. The proteins with normalized values (≥–2 and ≤2) were considered unchanged. XICs of the peptide with normalized values (≥2) were manually inspected for correct peak assignment.
Inflammasome assays
The NLRP3 inflammasome was activated by 10 μM Nigericin treatment for 2 h after 100 ng/ml LPS priming for 4 h. The NLRP1b inflammasome was activated by treatment of 2 μg/ml LeTx for 2 h after 100 ng/ml LPS-priming for 4 h. The NLRC4 inflammasome was activated by 100 MOI S. typhimurium for 2 h after 100 ng/ml LPS priming for 4 h. The non-canonical inflammasome was activated by 100 ng/ml LPS priming for 4 h and then transfected with 20 μg/ml Ctb.
Microscopy imaging of cell death
To examine cell death morphology, cells were treated as indicated in 12-well plates or 35-mm glass bottom dishes for image capture. PI (5 ng/ml) was added to the medium for monitoring cell membrane integrity. Static bright field images of pyroptotic cells were captured using an Olympus IX71 or a Zeiss LSM 780 at room temperature. The pictures were processed using ImageJ or the ZEN 2012 Image program.
Cytotoxicity assay and IL-1β ELISA
A CytoTox 96 Non-Radioactive Cytotoxicity assay was used to measure cell death. To measure IL-1β release, RAW-ASC cells were primed with 100 ng/ml LPS for 4 h and then treated with 10 μM Nigericin for 2 h. The release of mature IL-1 was determined by using the IL-1β ELISA kit (Neobioscience Technology Company).
Measurement of Caspase-3, -7 and -8 activity
Caspases-3/7 and -8 activities were determined by using a Caspase-Glo 3/7 or 8 assay kit (Promega) according to the manufacturer's instructions. In brief, 1.0 × 105 cells were seeded in 96-well plate with white wall (Nunc). After treatment, equal volume of Caspase-Glo 3/7 or 8 reagent was added to the cell culture medium, which had been equilibrated to room temperature for 1 h, cells were shaked for 5 min and incubated at room temperature for 30 min. Luminescent recording was performed with POLAR star Omega (BMG Labtech).
Knockout mice and isolation of mouse BMDM cells
Mice were housed in a specific pathogen-free facility. All experiments were conducted in compliance with the regulations of Xiamen University. The Gsdmd-knockout mice were generated by co-microinjection of in vitro-translated Cas9 mRNA and gRNA into the C57BL/6 zygotes. New-born mouse tail DNA samples were amplified with PrimeStar (Takara, Dalian, China) and the PCR products were sequenced for the screening of heterozygous or knockout descendants. The gRNA sequence used to generate the knockout mice is 3′-CACGACTCCGAAGCTGTTGC-5′.
Mouse BMDMs were differentiated in vitro from isolated bone marrow cells. Bone marrow cells collected from mouse femurs and tibias were incubated for 7 days in DMEM containing 20% heat-inactivated FBS, penicillin, streptomycin, and 30% L929 conditional medium.
Acknowledgments
We thank Prof Xingxu Huang in Shanghai University for pST1374-NLS-flag-linker-Cas9 and pUC57-sgRNA plasmids. This work was supported by the National Basic Research Program of China (973 Program; 2015CB553800), the National Natural Science Foundation of China (91429301, 31420103910, 31330047, and 31221065), the National Scientific and Technological Major Project (2013ZX10002-002), the Hi-Tech Research and Development Program of China (863 program; 2012AA02A201), the 111 Project (B12001), the Science and Technology Foundation of Xiamen (3502Z20130027), the National Science Foundation of China for Fostering Talents in Basic Research (J1310027) and the Open Research Fund of State Key Laboratory of Cellular Stress Biology, Xiamen University.
Footnotes
(Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary Material
Pyroptosis, caspase-1 activation and IL-1β secretion in RAW264.7, J774 and their derived cell lines.
Identification of GSDMD by MS and characterization of RAW-asc derived cells.
GSDMD is required for canonical and non-canonical inflammasome-mediated pyroptosis and IL-1β secretion.
Secondary necrosis of apoptotic cells in inflammasome stimuli-treated Gsdmd−/− cells.
References
- 1Martinon F, Mayor A, Tschopp J. The inflammasomes: guardians of the body. Annu Rev Immunol 2009; 27:229–265. [DOI] [PubMed] [Google Scholar]
- 2Lamkanfi M, Dixit VM. Inflammasomes and their roles in health and disease. Annu Rev Cell Dev Biol 2012; 28:137–161. [DOI] [PubMed] [Google Scholar]
- 3Fink SL, Cookson BT. Apoptosis, pyroptosis, and necrosis: mechanistic description of dead and dying eukaryotic cells. Infect Immun 2005; 73:1907–1916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4Netea MG, Simon A, van de Veerdonk F, et al. IL-1beta processing in host defense: beyond the inflammasomes. PLoS Pathog 2010; 6:e1000661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5Miao EA, Leaf IA, Treuting PM, et al. Caspase-1-induced pyroptosis is an innate immune effector mechanism against intracellular bacteria. Nat Immunol 2010; 11:1136–1142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6Lamkanfi M, Dixit VM. Mechanisms and functions of inflammasomes. Cell 2014; 157:1013–1022. [DOI] [PubMed] [Google Scholar]
- 7Davis BK, Wen H, Ting JP. The inflammasome NLRs in immunity, inflammation, and associated diseases. Annu Rev Immunol 2011; 29:707–735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8Schroder K, Tschopp J. The inflammasomes. Cell 2010; 140:821–832. [DOI] [PubMed] [Google Scholar]
- 9Dinarello CA. Immunological and inflammatory functions of the interleukin-1 family. Annu Rev Immunol 2009; 27:519–550. [DOI] [PubMed] [Google Scholar]
- 10Shi J, Zhao Y, Wang Y, et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 2014; 514:187–192. [DOI] [PubMed] [Google Scholar]
- 11Kayagaki N, Warming S, Lamkanfi M, et al. Non-canonical inflammasome activation targets caspase-11. Nature 2011; 479:117–121. [DOI] [PubMed] [Google Scholar]
- 12Masumoto J, Taniguchi S, Ayukawa K, et al. ASC, a novel 22-kDa protein, aggregates during apoptosis of human promyelocytic leukemia HL-60 cells. J Biol Chem 1999; 274:33835–33838. [DOI] [PubMed] [Google Scholar]
- 13Van Opdenbosch N, Gurung P, Vande Walle L, et al. Activation of the NLRP1b inflammasome independently of ASC-mediated caspase-1 autoproteolysis and speck formation. Nat Commun 2014; 5:3209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14Mariathasan S, Weiss DS, Newton K, et al. Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 2006; 440:228–232. [DOI] [PubMed] [Google Scholar]
- 15Hu Y, Mao K, Zeng Y, et al. Tripartite-motif protein 30 negatively regulates NLRP3 inflammasome activation by modulating reactive oxygen species production. J Immunol 2010; 185:7699–7705. [DOI] [PubMed] [Google Scholar]
- 16Pelegrin P, Barroso-Gutierrez C, Surprenant A. P2X7 receptor differentially couples to distinct release pathways for IL-1beta in mouse macrophage. J Immunol 2008; 180:7147–7157. [DOI] [PubMed] [Google Scholar]
- 17Li Y, Zhong CQ, Xu X, et al. Group-DIA: analyzing multiple data-independent acquisition mass spectrometry data files. Nat Methods 2015 Oct 5. doi:10.1038/nmeth.3593 [DOI] [PubMed]
- 18Agard NJ, Maltby D, Wells JA. Inflammatory stimuli regulate caspase substrate profiles. Mol Cell Proteomics 2010; 9:880–893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19Broz P, von Moltke J, Jones JW, et al. Differential requirement for Caspase-1 autoproteolysis in pathogen-induced cell death and cytokine processing. Cell Host Microbe 2010; 8:471–483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20Kayagaki N, Stowe IB, Lee BL, et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signaling. Nature 2015; 526:666–671. [DOI] [PubMed] [Google Scholar]
- 21Shi J, Zhao Y, Wang K, et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 2015; 526:660–665. [DOI] [PubMed] [Google Scholar]
- 22Han J, Zhong CQ, Zhang DW. Programmed necrosis: backup to and competitor with apoptosis in the immune system. Nat Immunol 2011; 12:1143–1149. [DOI] [PubMed] [Google Scholar]
- 23Gillet LC, Navarro P, Tate S, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics 2012; 11:O111 016717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24Wu X, Tian L, Li J, et al. Investigation of receptor interacting protein (RIP3)-dependent protein phosphorylation by quantitative phosphoproteomics. Mol Cell Proteomics 2012; 11:1640–1651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25Deutsch EW, Mendoza L, Shteynberg D, et al. A guided tour of the Trans-Proteomic Pipeline. Proteomics 2010; 10:1150–1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26Keller A, Nesvizhskii AI, Kolker E, et al. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 2002; 74:5383–5392. [DOI] [PubMed] [Google Scholar]
- 27Shteynberg D, Deutsch EW, Lam H, et al. iProphet: multi-level integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates. Mol Cell Proteomics 2011; 10:M111 007690. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Pyroptosis, caspase-1 activation and IL-1β secretion in RAW264.7, J774 and their derived cell lines.
Identification of GSDMD by MS and characterization of RAW-asc derived cells.
GSDMD is required for canonical and non-canonical inflammasome-mediated pyroptosis and IL-1β secretion.
Secondary necrosis of apoptotic cells in inflammasome stimuli-treated Gsdmd−/− cells.