Abstract
Background
Euglena gracilis, a unicellular phytoflagellate within Euglenida, has attracted much attention as a potential feedstock for renewable energy production. In outdoor open-pond cultivation for biofuel production, excess direct sunlight can inhibit photosynthesis in this alga and decrease its productivity. Carotenoids play important roles in light harvesting during photosynthesis and offer photoprotection for certain non-photosynthetic and photosynthetic organisms including cyanobacteria, algae, and higher plants. Although, Euglenida contains β-carotene and xanthophylls (such as zeaxanthin, diatoxanthin, diadinoxanthin and 9′-cis neoxanthin), the pathway of carotenoid biosynthesis has not been elucidated.
Results
To clarify the carotenoid biosynthetic pathway in E. gracilis, we searched for the putative E. gracilis geranylgeranyl pyrophosphate (GGPP) synthase gene (crtE) and phytoene synthase gene (crtB) by tblastn searches from RNA-seq data and obtained their cDNAs. Complementation experiments in Escherichia coli with carotenoid biosynthetic genes of Pantoea ananatis showed that E. gracilis crtE (EgcrtE) and EgcrtB cDNAs encode GGPP synthase and phytoene synthase, respectively. Phylogenetic analyses indicated that the predicted proteins of EgcrtE and EgcrtB belong to a clade distinct from a group of GGPP synthase and phytoene synthase proteins, respectively, of algae and higher plants.
In addition, we investigated the effects of light stress on the expression of crtE and crtB in E. gracilis. Continuous illumination at 460 or 920 μmol m−2 s−1 at 25 °C decreased the E. gracilis cell concentration by 28–40 % and 13–91 %, respectively, relative to the control light intensity (55 μmol m−2 s−1). When grown under continuous light at 920 μmol m−2 s−1, the algal cells turned reddish-orange and showed a 1.3-fold increase in the crtB expression. In contrast, EgcrtE expression was not significantly affected by the light-stress treatments examined.
Conclusions
We identified genes encoding CrtE and CrtB in E. gracilis and found that their protein products catalyze the early steps of carotenoid biosynthesis. Further, we found that the response of the carotenoid biosynthetic pathway to light stress in E. gracilis is controlled, at least in part, by the level of crtB transcription. This is the first functional analysis of crtE and crtB in Euglena.
Electronic supplementary material
The online version of this article (doi:10.1186/s12870-015-0698-8) contains supplementary material, which is available to authorized users.
Keywords: Euglena gracilis, Light stress, Carotenoid biosynthesis, Geranylgeranyl pyrophosphate synthase, CrtE, Phytoene synthase, CrtB
Background
Euglena gracilis, a eukaryotic unicellular phytoflagellate within Euglenida, is a secondary plant [1] in which the chloroplasts carry chlorophylls a and b and carotenoids, similar to what is observed in green algae (Chlorophyta) and higher plants [2]. This alga has attracted much attention as a potential feedstock for renewable energy production. In outdoor open-pond cultivation for biofuel production, the productivity of this alga depends on several environmental factors such as light intensity and temperature. Excess direct sunlight can inhibit photosynthesis in this alga and decrease its productivity.
Carotenoids play important roles in photosynthesis and photoprotection of photosynthetic organisms and certain non-photosynthetic organisms. More than 750 natural carotenoids have been isolated from various organisms. Carotenoids are synthesized by phototrophs and non-phototrophs including bacteria, archaea, fungi, algae, and higher plants [3]. In photosynthetic pathways, both carotenoids and chlorophylls constitute light-harvesting pigment-protein complexes in chloroplast membranes. Carotenoids also play important roles in the stabilization of thylakoid membranes [4], in photoprotection (i.e. non-photochemical quenching, the xanthophyll cycle, and scavenging reactive oxygen species) [5], and in the synthesis of abscisic acid [6] and strigolactones [7].
Carotenoids are classified into two classes, carotenes (hydrocarbons) and xanthophylls (oxygenated derivatives of carotenes). Geranylgeranyl pyrophosphate (GGPP; C20), the precursor of carotenes, is synthesized from farnesyl pyrophosphate (C15) and isopentenyl pyrophosphate (C5) by geranylgeranyl pyrophosphate synthase (CrtE, also known as GGPPS or GGPS). Then phytoene (C40), the first carotene, is synthesized by the condensation of two molecules of GGPP by phytoene synthase (CrtB, also called Psy or Pys). Subsequently, phytoene is converted into lycopene through desaturation steps and isomerization catalyzed by phytoene desaturase (CrtP, also called Pds), ζ-carotene desaturase (CrtQ, also called Zds) and cis-carotene isomerase (CrtH, also called CrtISO) in oxygenic phototrophs. Bicyclic carotenes, α-carotene and β-carotene and their oxygenated derivatives (xanthophylls), are synthesized from lycopene [3, 8].
The distribution of carotenoid species in algae including cyanobacteria, red algae, brown algae, and green algae, has been summarized [8] and suggests that algae have several carotenoid biosynthetic pathways in common with higher plants based on similarities among carotenoid chemical structures. The genes whose products catalyze the early steps of the carotenoid biosynthetic pathways in common with higher plants have been functionally identified in several eukaryotic algae such as Pyropia umbilicalis (ggps), Chlamydomonas reinhardtii (crtB), Haematococcus pluvialis (pys), and Chlorella zofingiensis (psy and crtP) and as well as cyanobacteria such as Thermosynechococcus elongatus (crtE), Gloeobacter violaceus PCC 7421 (crtB), Synechococcus elongatus PCC 7942 (pys), and Synechocystis sp. PCC 6803 (crtQ and crtH) [8–10].
Euglenida contains β-carotene and xanthophylls such as zeaxanthin, diatoxanthin, diadinoxanthin and 9′-cis neoxanthin [8, 11–13], however, the biosynthetic pathways and the corresponding genes of carotenoid synthesis in this alga have not been elucidated. In the present study, to clarify the carotenoid biosynthetic pathway of E. gracilis within Euglenida, we searched for the orthologs of the GGPP synthase gene and phytoene synthase gene from a series of E. gracilis cDNA sequences (Yoshida et al., unpublished observations) using tblastn, and we identified E. gracilis crtE (EgcrtE) and EgcrtB encoding GGPP synthase and phytoene synthase, respectively. Phylogenetic analyses indicated that E. gracilis CrtE and CrtB belong to a clade that is distinct from groups of algae and higher plants, respectively. In addition, we investigated the effects of light stress on the expression of crtE and crtB in E. gracilis, and revealed that the carotenoid biosynthetic pathway in E. gracilis responded to excess light stress at the level of crtB transcription.
Results
Cloning of EgcrtE and EgcrtB
We performed BLAST (tblastn) searches against a series of Euglena full-length cDNA sequences (Yoshida et al., unpublished observations) using Capsicum annuum GGPS [GenBank: CAA56554] and C. annuum PSY1 [GenBank: CAA48155] as queries. We obtained cDNA sequences of the putative GGPP synthase gene (crtE) and phytoene synthase gene (crtB) in E. gracilis. The cDNA sequences that encode EgcrtE and EgcrtB from the RNA-seq data each contained a spliced-leader (SL) sequence 5′-TTTTTTTTCG-3′, a characteristic sequence transferred to the 5′ extremity of mRNAs by trans-splicing [14]. The presence of SL sequences at the 5′ ends of the cDNAs corresponding to EgcrtE and EgcrtB indicated that the obtained sequences code for the full-length cDNA. The cDNAs for putative EgcrtE and EgcrtB (Additional files 1 and 2) were isolated from E. gracilis by RT-PCR with primers designed according to the RNA-seq data. The sequences of EgcrtE and EgcrtB cDNA were submitted to the DDBJ under accession numbers LC062706 and LC062707, respectively.
The first ATG downstream of the SL sequence in both EgcrtE and EgcrtB cDNA was considered the start codon of the respective mRNA. The deduced amino acid sequences of EgcrtE and EgcrtB are predicted to be 402 and 406 amino acids in length, respectively (Figs. 1 and 2, and Additional files 1 and 2). The typical signal motif for plastid-targeted proteins in E. gracilis [15] was not found in either EgCrtE or EgCrtB with the TMHMM program [16]. Furthermore, no characteristic signal motif was predicted in EgCrtE and EgCrtB with the TargetP program [17].
Fig. 1.

Alignment of the deduced E. gracilis CrtE amino acid sequence with known GGPP synthases. The accession numbers are Arabidopsis thaliana GGPPS1, [GenBank: NP_175376]; GGPPS4 [GenBank: NP_179960]; Capsicum annuum GGPS, [GenBank: CAA56554] and Thermosynechococcus elongatus BP-1 CrtE, [GenBank: NP_680811]. Sequence data for GGPS of Pyropia umbilicalis [P_umbilicalis_esContig5139] was obtained from NoriBLAST [58]. Underlined sequences indicate the first and second aspartate-rich motifs, FARM and SARM, respectively. The boxed residues comprise the chain-length determination (CLD) region. Multiple sequence alignment was conducted with Clustal W using MEGA version 6.0 [59]
Fig. 2.
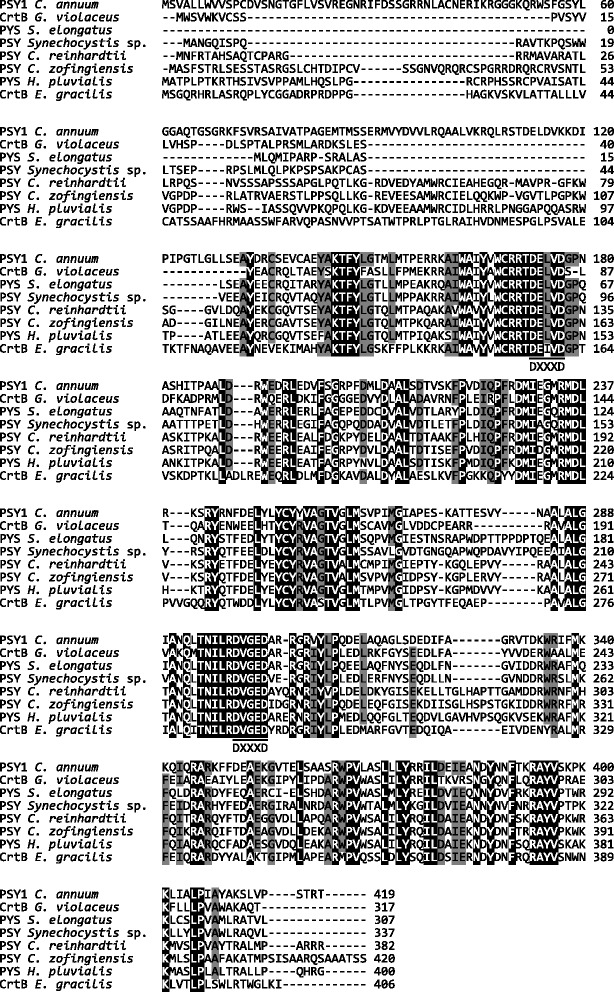

Alignment of the deduced E. gracilis CrtB amino acid sequence with known phytoene synthases. The accession numbers are Capsicum annuum PSY1, [GenBank: CAA48155]; Gloeobacter violaceus PCC 7421 CrtB [GenBank: BAC89685]; Synechococcus elongatus PCC 7942 PYS [GenBank: CAA45350]; Synechocystis sp. PCC 6803 PYS [GenBank: CAA48922]; Chlamydomonas reinhardtii PSY [GenBank: XP_001701192]; Chlorella zofingiensis PSY [GenBank: CBW37867] and Haematococcus pluvialis PYS [GenBank: AAY53806]. Underlined sequences indicate the two aspartate-rich motifs (DXXXD). Multiple sequence alignment was conducted with Clustal W using MEGA version 6.0 [59]
In the phylogenetic tree for GGPP synthases (Additional file 3), the predicted protein encoded by EgcrtE is relatively close to an algal clade including Cyanophyta and Rhodophyta. The amino acid sequence of E. gracilis CrtE is 46 and 44 % identical to GGPP synthases of T. elongatus and P. umbilicalis, respectively, and the corresponding sequence similarities are 59 and 55 %, as aligned with Needle in EMBOSS [18]. EgCrtE contains the typical aspartate-rich motifs conserved in type II GGPPS of eubacteria and plants, namely the first aspartate-rich motif (FARM: DDXXXD) in the chain-length determination (CLD) region, and the second aspartate-rich motif (SARM: DDXXD) [19, 20] (Fig. 1). In the phylogenetic tree (Additional file 4), EgCrtB is in a distinct clade apart from clades of phytoene synthases of cyanobacteria (Cyanophyta) and green algae (Chlorophyta). The deduced amino acid sequence for EgCrtB is 38, 39 and 40 % identical with phytoene synthases of H. pluvialis, C. zofingiensis, and C. reinhardtii, respectively, and the corresponding sequence similarities are 52, 53 and 56 %, as aligned with Needle in EMBOSS [18]. EgCrtB contains two aspartate-rich motifs (DXXXD) conserved among phytoene synthases [21] (Fig. 2).
Functional analysis of EgcrtE and EgcrtB
The function of isolated EgcrtE and EgcrtB cDNA was analyzed with color complementation studies in Escherichia coli carrying the carotenoid biosynthetic gene cluster of Pantoea ananatis (formerly Erwinia uredovora) [22]. E. coli transformed with pET-EgcrtE and pACCAR25ΔcrtE [22], which carries P. ananatis carotenoid biosynthetic gene cluster (crtB, crtI; phytoene desaturase, crtY; lycopene cyclase, crtZ; β-carotene hydroxylase and crtX; zeaxanthin glucosidase, but missing crtE), showed accumulation of yellow-orange pigments (Fig. 3a). In contrast, this pigmentation was not observed in E. coli carrying pACCAR25ΔcrtE and pETDuet-1 (vector control). In the same way, the function of EgcrtB was analyzed in E. coli with pACCAR25ΔcrtB [23] carrying P. ananatis crtE, crtI, crtY, crtZ and crtX, but missing crtB. E. coli co-transformed with pET-EgcrtB and pACCAR25ΔcrtB showed the yellow-orange color (Fig. 3b). These results suggested that the proteins predicted to be encoded by EgcrtE and EgcrtB have GGPP synthase and phytoene synthase activity, respectively.
Fig. 3.
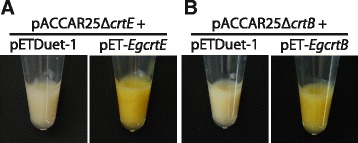
Color complementation experiments in E. coli with the P. ananatis carotenoid synthetic gene cluster. a E. coli carrying pACCAR25ΔcrtE [22] with pETDuet-1 (vector control) or pET-EgcrtE. b E. coli cells carrying pACCAR25ΔcrtB [23] with pETDuet-1 (vector control) or pET-EgcrtB. E. coli strain BL21(DE3) was used as the host. Data are representative of at least eight E. coli transformants with similar results
The ability of EgCrtE and EgCrtB to function in phytoene production was also investigated by high-performance liquid chromatography (HPLC). Phytoene was detected in E. coli harboring crtE of E. gracilis (pET-EgcrtE) and crtB of P. ananatis (pAC-PacrtB) with a retention time of 28.6 min (Fig. 4a, d). Similarly, phytoene production was also observed in E. coli carrying crtE of P. ananatis (pACCRT-E plasmid [23]) and crtB of E. gracilis (pET-EgcrtB) (Fig. 4b). In addition, E. coli transformed with pET-EgcrtEB carrying EgcrtE and EgcrtB synthesized phytoene (Fig. 4c). In contrast, phytoene was not detected in E. coli carrying either EgcrtE or EgcrtB alone (Additional file 5A and B). Furthermore, phytoene production was not observed in E. coli carrying pAC-PacrtB or pACCRT-E with pETDuet-1 (vector control) (Additional file 5C and D). Taken together, these findings indicate that the crtE and crtB cDNAs isolated from E. gracilis code for the GGPP synthase and the phytoene synthase, respectively.
Fig. 4.
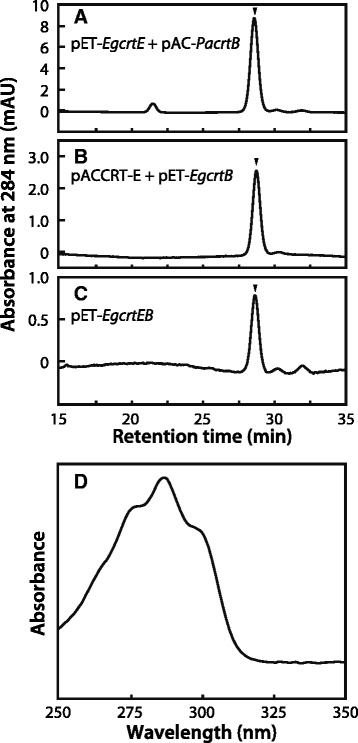
Analysis of phytoene production in E. coli by HPLC. HPLC chromatogram (284 nm) of extracts from E. coli carrying a pET-EgcrtE with pAC-PacrtB, b pACCRT-E [23] with pET-EgcrtB and c pET-EgcrtEB. d Absorbance spectrum of phytoene detected at a retention time of 28.6 min. Phytoene was extracted from E. coli transformants and analyzed with HPLC in accordance with the method of Takaichi [57]. The arrowheads in the chromatograms indicate the position of phytoene elution. Data are representative of three or four experiments with similar results
crtE and crtB expression in E. gracilis in response to light stress
Figure 5a shows a time course of E. gracilis cell concentration grown under various light intensities. When the cells were grown under continuous light at 55 μmol m−2 s−1 (control) for 7 days, the cell concentration increased from 3 × 103 cells ml−1 to 1.4 − 1.5 × 106 cells ml−1. Illumination at 27 μmol m−2 s−1 did not affect the cell concentration compared with the control throughout the cultivation period. In contrast, a significant decrease in the cell concentration was observed in the algae grown under illumination at 460 and 920 μmol m−2 s−1 (Fig. 5a). The treatment with light intensity at 460 μmol m−2 s−1 significantly (P < 0.05) decreased the cell concentration to 72, 60, and 77 % of the control after 4, 5, and 6 days of cultivation, respectively. Illumination at 920 μmol m−2 s−1 decreased the cell concentration to 87 % of the control 1 day after the cultivation, and the degree of inhibition of cell growth increased in a time-dependent manner. After 6 days of cultivation, the concentration of cells illuminated at 920 μmol m−2 s−1 was decreased to 9 % (1.5 × 105 cells ml−1) of the control. After 7 days of treatment at 460 and 920 μmol m−2 s−1, the cell concentration reached 1.4 × 106 cells ml−1 (99 % of control) and 2.4 × 105 cells ml−1 (16 % of control), respectively.
Fig. 5.
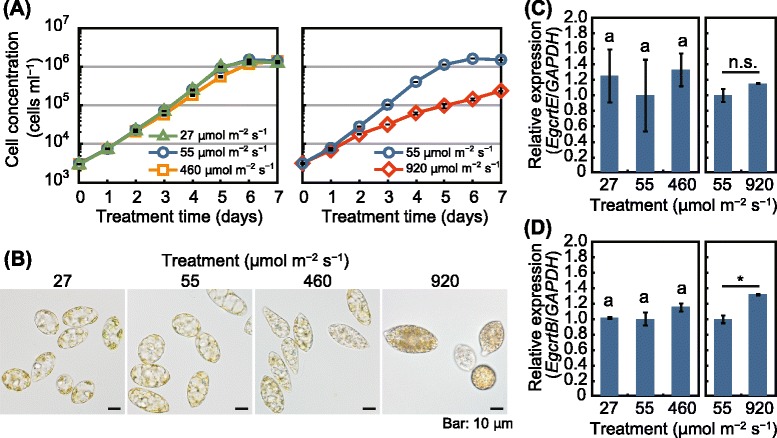
Effects of light intensity on crtE and crtB expression levels in E. gracilis. a Time-course of cell concentration of E. gracilis grown under continuous light at 27, 55, 460, and 920 μmol m−2 s−1 at 25 °C. b Cells of the alga cultured under the indicated light-stress treatments for 7 days. c and d Expression levels of EgcrtE (c) and EgcrtB (d) in the algal cells treated with the 7-day light-stress treatments. Data are the mean ± SE (n = 3). Data are representative of at least two individual experiments with similar results. Bars labeled with the same letter are not significantly different (Tukey’s multiple range test, P < 0.05). n.s., not significant; *P < 0.05, t-test
Compared with the control, no remarkable difference was observed in the appearance of the algal cells grown under continuous light at 27 μmol m−2 s−1 for 7 days (Fig. 5b). The cells subjected to the control (55 μmol m−2 s−1) and to 27 μmol m−2 s−1 contained translucent granules thought to be paramylon. The translucent granules were also observed in the cells illuminated at 460 μmol m−2 s−1 for 7 days, although grayish-colored granules (1–2 μm in diameter) also appeared in the cells (Fig. 5b). The cells illuminated at 920 μmol m−2 s−1 possessed more grayish granules than the cells illuminated at 460 μmol m−2 s−1. Furthermore, grown under illumination at 920 μmol m−2 s−1, the cells looked more reddish-orange than the control.
The expression of crtE mRNA in E. gracilis was not significantly affected by the various light intensities examined when the cells were cultured at 25 °C under continuous illumination (Fig. 5c). In contrast, the expression of crtB in the cells illuminated at 920 μmol m−2 s−1 increased 1.3-fold relative to the control (Fig. 5d). These results indicate that the response of the carotenoid biosynthetic pathway to light stress in E. gracilis is controlled, at least in part, at the level of crtB transcription.
Discussion
Identification of EgcrtE and EgcrtB
The GGPS of C. annuum [24] and the majority of the GGPP synthase family proteins of Arabidopsis thaliana [20] localize to plastids. Higher plants have two isoprenoid biosynthetic pathways, namely the plastidial 1-deoxy-d-xylulose 5-phosphate/2-C-methylerythritol 4-phosphate (DOXP/MEP) pathway and cytosolic acetate/mevalonate (MVA) pathway [25, 26]. Green algae (Chlorophyta) lost the MVA pathway during evolution, and thus these algae depend exclusively on the DOXP/MEP pathway [25, 26]. Higher plants and algae depend on isopentenyl pyrophosphate, which is derived from the DOXP/MEP pathway, for the biosynthesis of GGPP and subsequent synthesis of carotenoids in plastids [25]. Euglena is exceptional because it lacks the DOXP/MEP pathway and synthesizes isoprenoids via the MVA pathway [26, 27]. This is consistent with the predicted localization of EgCrtE in the cytosol based on TMHMM [16] and TargetP [17].
Phytoene synthases localize to plastids in A. thaliana, Oryza sativa, and Zea mays [21]. In the present study, however, neither TMHMM nor TargetP predicted a typical plastid transit peptide in the N-terminal region of EgCrtB, although it is difficult to exactly predict the plastid-targeted proteins of E. gracilis because the system that traffics proteins to Euglena’s plastids, which are surrounded by three membranes [1], differs from that of higher plants [28].
Most flagellate green algae have developed a light-sensitive system, the eyespot apparatus, composed of carotenoid-rich lipid globules inside the chloroplast [29]. Proteomic studies indicate that some of the β-carotene biosynthesis enzymes are localized in the eyespot apparatus of C. reinhardtii [30] and in β-carotene plastoglobuli in Dunaliella bardawil [31], suggesting that part of the β-carotene synthesis occurs in the eyespot globules. E. gracilis also possesses an eyespot apparatus (stigma) that contains carotenoids [32], although stigmata of this alga are located in the cytoplasm near the base of the major flagellum [33]. In addition, Kivic and Vesk [33] reported that the stigma of this alga is surrounded by a single membrane and has no structural similarity to the chloroplast. This suggests that EgCrtB might be transported to stigmata as well as plastids and that EgCrtB might contain an as-yet unidentified signal sequence.
Although chloroplasts in E. gracilis contain chlorophylls a and b [2], EgCrtB belongs to a distinct clade apart from groups of green algae (Chlorophyta) and higher plants (Plantae) in the phylogenetic tree (Additional files 3 and 4). This result is consistent with taxonomic relations. E. gracilis belongs to Euglenida within supergroup Excavata [34]. Euglenida is a primitive organism that has a common ancestor with Trypanosoma sp. (Kinetoplastea) [34–36]. Evolutionarily, Euglenozoa including Euglenida and Kinetoplastea is considered to have branched early from other eukaryotes carrying the symbiont, Chlorophyta [37, 38]. The phylogenetic relationships of GGPP synthase and phytoene synthase proteins among various photoautotrophs (Additional files 3 and 4) might reflect the distinctive evolutionary history of E. gracilis.
crtE and crtB expression in E. gracilis in response to light stress
Steinbrenner and Linden [39] reported that the highest growth rate of H. pluvialis is observed under continuous light at 50–150 μmol m−2 s−1, and illumination at 250 μmol m−2 s−1 reduces the cell number. Similarly, Wahidin et al. [40] showed that the cell concentration of Nannochloropsis sp. decreases under illumination at 200 μmol m−2 s−1. In our preliminary experiment, illumination at 240 μmol m−2 s−1 had no significant effect on cell concentration throughout the cultivation period compared with the control (data not shown). Illumination at an intensity of ~460 μmol m−2 s−1 is considered to be a threshold of excess light stress to E. gracilis grown under continuous light at 25 °C, and this level of illumination might begin to cause photoinhibition of photosynthesis in this alga. The cell growth delay caused by illumination at 460 μmol m−2 s−1 was slightly alleviated at the early stationary phase (6 days after the cultivation), and by the end of the cultivation, the algal cells had increased in number as much as the control (Fig. 5a). This result might be due to the shading effects of the grayish granules that accumulated in the cells (Fig. 5b).
When grown under continuous light at 920 μmol m−2 s−1, the algal cells turned reddish-orange (Fig. 5b). This result is consistent with previous studies indicating that light-stress induces the accumulation of carotenoids in certain green algae such as Dunaliella salina [41], H. pluvialis [42], and C. zofingiensis [43]. Król et al. [41] reported that excess irradiance at 2500 μmol m−2 s−1 induced a comparable accumulation of carotenoids in D. salina cells. Wang et al. [44] reported that irradiation of H. pluvialis at 350 μmol m−2 s−1 induced an increase in carotenoids, and that the astaxanthin-accumulating red cells were more resistant to very high irradiance (3000 μmol m−2 s−1) than green cells.
In higher plants, the regulation of carotenoid biosynthesis has mainly been investigated in the context of seedling de-etiolation and the accompanying burst in carotenoid biosynthesis. Lintig et al. [45] reported that the expression of the GGPP synthase gene (ggps) in Sinapsis alba seedlings remained constant during de-etiolation. This report is consistent with our data showing that EgcrtE expression remained relatively constant under the light-stress treatments examined (Fig. 5c). Flux of isoprenoids in the MEP pathway in higher plants is mainly controlled by DOXP synthase [46], DOXP reductoisomerase [47], and hydroxymethylbutenyl diphosphate reductase [48]. These three rate-determining enzymes are upregulated and control the metabolic flux to the carotenoid pathway during de-etiolation of A. thaliana [49]. Light-induction of the gene dxs encoding DOXP synthase was also reported in Phaeodactylum tricornutum (diatom) in the dark–light transition [50].
In contrast to crtE, crtB expression in E. gracilis increased by 1.3-fold in response to intense illumination (920 μmol m−2 s−1; Fig. 5d). This result is consistent with previous studies of light-regulated carotenoid biosynthetic genes. For example, expression of the phytoene synthase gene (psy) of A. thaliana is upregulated during seedling de-etiolation, resulting in an accumulation of carotenoids [48, 49, 51]. Rodríguez-Villalón et al. [49] reported that PSY is the key driver that increases carotenoid synthesis in etiolated seedlings of A. thaliana by controlling the metabolic flux to the carotenoid biosynthesis pathway. Light induction of the phytoene synthase gene has also been observed in algae. Bohne and Linden [52] reported that C. reinhardtii showed a fast upregulation of crtB with a maximum at 1–2 h after the dark-to-light transition. Steinbrenner and Linden [42] reported that continuous high-intensity light (125 μmol m−2 s−1) leads to a slight increase in pys expression followed by moderate astaxanthin accumulation in H. pluvialis. This is consistent with our finding that the carotenoid biosynthesis pathway in E. gracilis under light stress is controlled, in part, at the transcriptional level of EgcrtB downstream of the branch point for carotenoid, chlorophyll, tocopherol, plastoquinone, and gibberellin biosynthesis in isoprenoid metabolism [19].
Conclusions
We functionally identified the GGPP synthase gene (EgcrtE) and phytoene synthase gene (EgcrtB), which catalyze the early steps of the carotenoid biosynthetic pathway, in E. gracilis within supergroup Excavata. Phylogenetic analyses of GGPP synthase and phytoene synthase proteins indicated that EgCrtE and EgCrtB, respectively, belong to a clade distinct from groups of algae and higher plants, consistent with taxonomic results. In addition, we have found that the carotenoid biosynthetic pathway in E. gracilis responded to excess light stress at the level of EgcrtB expression. To the best of our knowledge, this is the first report on the functional analysis of crtE and crtB in Euglena.
Methods
Biological materials
Euglena gracilis Klebs (strain Z) was cultured in 100 ml of Cramer-Myers medium [53] containing 0.1 % ethanol at an initial cell concentration of 3.0 × 103 cells ml−1 in a 300-ml conical flask. Algal cells were grown in an incubator (LH-350SP, NK system) with agitation (90 rpm), and illuminated with fluorescent lamps (FL20S EX-N-HG and FL40S EX-N-HG, NEC Lighting). To clone EgcrtE and EgcrtB, the algal cells were grown at 25 °C under continuous illumination at 55 μmol m−2 s−1 for 7 days. To analyze the expression levels of EgcrtE and EgcrtB gene in E. gracilis under light stress, algal cells were grown at 25 °C under continuous illumination at 27, 55 (control), 460, and 920 μmol m−2 s−1 for 7 days. For illumination at 460 and 920 μmol m−2 s−1, white LED lamps (LLM0175A, Stanley Electric) were used in combination with the fluorescent lamps. Cell concentration was measured daily by counting with a plankton counter (MPC-200, Matsunami Glass Ind.) under a microscope. At 7 days after the cultivation, algal cells were harvested by centrifugation (1000 × g, 2 min), and the collected cells were frozen immediately and stored at −60 °C until the RNA was isolated.
Cloning of EgcrtE and EgcrtB
Total RNA was isolated from the algal cells with RNAqueous kit (Ambion) and Plant RNA Isolation Aid (Ambion). First-strand cDNA was synthesized with SuperScript First-Strand Synthesis System for RT-PCR (Invitrogen) from total RNA treated with DNase I (Invitrogen). cDNAs containing EgcrtE and EgcrtB coding sequences were amplified by RT-PCR with PrimeSTAR GXL DNA Polymerase (Takara Bio). Primers used for RT-PCR were as follows: EgcrtE, 5′-TTTCGCTCACACGCACAATG-3′ and 5′-CCCAGCGTACAGAAAAGCTA-3′; EgcrtB, 5′-TTCGGTCGCTCCCCTTCCA-3′ and 5′-AGCAGCCGAGTATGATACGA-3′. The amplified fragments were gel-purified (Gel/PCR Extraction kit, FastGene) and sub-cloned into pMD20-T vector with Mighty TA-cloning Reagent Set for PrimeSTAR (Takara Bio) and sequenced. E. coli strain JM109 (Takara Bio) was used as a host for the plasmids and grown in LB medium [54] at 37 °C in the dark. Ampicillin (50 μg ml−1) was added to the medium as needed.
Construction of plasmids for complementation experiments
The coding sequence of EgcrtE cDNA was amplified with PrimeSTAR GXL DNA Polymerase and the primers 5′-TGAATTCCACACGCACAATGGCC-3′ and 5′-ATAAGCTTCAGTTGGTGCGGGC-3′, which contain EcoRI and HindIII restriction sites, respectively. The coding sequence of EgcrtB cDNA was amplified with primers 5′-CTTCCATATGTCCGGCCAGAG-3′ and 5′-TCTCGAGTAAGATCTTCAAGCCC-3′, which contain NdeI and XhoI restriction sites, respectively. The amplified fragments were gel-purified and sub-cloned into pMD20-T vector with Mighty TA-cloning Reagent Set for PrimeSTAR. E. coli strain JM109 was used as a host for the plasmids and grown as described above.
To construct the pET-EgcrtE, the coding sequence for EgcrtE was cloned into the EcoRI/HindIII site (multi cloning site 1, MCS1) of pETDuet-1 vector (Novagen) with Ligation Mighty Mix (Takara Bio). pET-EgcrtB plasmid was created by cloning the EgcrtB sequence into the NdeI/XhoI sites (MCS2) of pETDuet-1. pET-EgcrtEB was created by cloning EgcrtE and EgcrtB into the EcoRI/HindIII site (MCS1) and NdeI/XhoI site (MCS2) of pETDuet-1, respectively.
pAC-PacrtB was constructed as follows. The open reading frame of P. ananatis crtB was amplified from pACCAR25ΔcrtE [22] with primers 5′-GAACATATGGCAGTTGGCTCGA-3′ and 5′-ACCTCGAGCTAGAGCGGGC-3′, which contain NdeI and XhoI restriction site, respectively, and was then cloned into MCS2 of pACYCDuet-1 (Novagen). Restriction enzymes used in this study were purchased from Takara Bio. E. coli strain JM109 was used as a host for the plasmids, and grown as described above. Ampicillin (50 μg ml−1) and chloramphenicol (30 μg ml−1) were added to the medium as needed.
Complementation experiments
pACCAR25ΔcrtE, which carries the P. ananatis carotenoid synthetic gene cluster (crtB, crtI, crtY, crtZ and crtX) with the exception of crtE was introduced into E. coli strain BL21(DE3) (New England BioLabs). The transformant harboring pACCAR25ΔcrtE was made competent in accordance with the method of Inoue et al. [55] and then was transformed with pET-EgcrtE. For the functional analysis of EgcrtB, E. coli strain BL21(DE3) was transformed with both pET-EgcrtB and pACCAR25ΔcrtB [23] carrying the P. ananatis gene cluster for zeaxanthin biosynthesis (crtE, crtI, crtY, crtZ and crtX) with the exception of crtB. The transformed E. coli cells were grown in 5 ml of LB medium at 37 °C in the dark until the OD600 of the culture medium reached 0.6 − 0.8 and then were cultured at 21 °C for 2 days in the medium with 50 μM of isopropyl-β-d-thiogalactopyranoside (IPTG) [56]. Ampicillin (50 μg ml−1) and chloramphenicol (30 μg ml−1) were added to the medium as needed. The E. coli cells were harvested from the medium by centrifugation (3000 × g, 5 min).
Phytoene extraction from E. coli and HPLC analysis
For the functional analysis of EgcrtE, E. coli strain BL21(DE3) was transformed with both pET-EgcrtE and pAC-PacrtB. For the functional analysis of EgcrtB, E. coli was co-transformed with pET-EgcrtB and pACCRT-E [23], which carries P. ananatis crtE. E. coli carrying pET-EgcrtEB was also created. The transformed cells were incubated in 5 ml of LB medium at 37 °C until the OD600 of the culture medium reached 0.6 − 0.8 and were then grown in the medium with 50 μM IPTG at 21 °C for 2 days in the dark [56]. The E. coli cells were harvested by centrifugation (3000 × g, 5 min) and frozen at −60 °C until the pigments extraction. Ampicillin (50 μg ml−1) and chloramphenicol (30 μg ml−1) were added to the medium as needed.
Pigments were extracted twice from the cells with 2 ml of acetone/methanol (7:2, v/v). After centrifugation, extracts were dried with a rotary evaporator. The pigments were dissolved in a small volume of n-hexane and then loaded on a silica gel (Wakogel C-300, Wako) column. The extracts were eluted from the column with 1–2 ml of n-hexane, and the n-hexane phase was evaporated to dryness with the rotary evaporator. The residue was dissolved in a small volume of ethanol and analyzed with an HPLC system as described below. The extraction procedure was conducted under dim light just before HPLC analysis.
The HPLC system was equipped with PEGASIL ODS SP100 column (6φ × 150 mm, Senshu Scientific Co.). The mobile phase was acetonitrile/methanol/tetrahydrofuran (58:35:7, v/v/v) [57] at a flow rate of 1.0 ml min−1. Absorbance spectra (250–350 nm, 1.2-nm resolution) and retention times were recorded with SPD-M20A, Photodiode Array Detector (Shimadzu).
Real-time quantitative PCR (qPCR) analysis of EgcrtE and EgcrtB expression
Total RNA was extracted from E. gracilis cells using RNAqueous kit and Plant RNA Isolation Aid. First-strand cDNA was synthesized from total RNA with QuantiTect Reverse Transcription kit (Qiagen) and used as the template. qPCR was conducted with Fast SYBR Green Master Mix (Applied Biosystems) on 7500 Fast Real-Time PCR System (Applied Biosystems). GAPDH [GenBank: L21903.1] was used as a reference gene for normalization of gene expression levels across samples. Primer sequences were as follows: GAPDH, 5′-GGTCTGATGACCACCATCCAT-3′ and 5′-TGAGGGTCCATCGACAGTCTT-3′; EgcrtE, 5′-GGTCTGGCGTTCCAAATCAT-3′ and 5′-TCATCCTTACCCGCTGTCTTG-3′; and EgcrtB, 5′-CGGAGTGACGGAGGATCAGA-3′ and 5′-ATCAAGGCCCGGTAATTCTCA-3′. qPCR analysis was performed in triplicate on each of three independent samples for each treatment.
Availability of supporting data
The data sets supporting the results of this article are included within the article and its additional files.
Acknowledgements
The authors are grateful to Dr. N. Misawa (Ishikawa Prefectural University, Japan), who kindly provided pACCAR25ΔcrtE, pACCAR25ΔcrtB, and pACCRT-E plasmids. We also thank to Dr. M. Takemura (Ishikawa Prefectural University, Japan) for helpful suggestions for the color complementation experiments. A part of this work was supported by grants from the JSPS KAKENHI (25450308) and MEXT-supported Program for the Strategic Research Foundation at Private Universities (S1311014) to SK and TS.
Abbreviations
- CrtB Psy, Pys
Phytoene synthase
- CrtE GGPPS, GGPS
geranylgeranyl pyrophosphate synthase
- CrtH CrtISO
cis-carotene isomerase
- CrtI CrtP
Phytoene desaturase
- CrtQ
ζ-carotene desaturase
- CrtX
Zeaxanthin glucosidase
- CrtY
Lycopene cyclase
- CrtZ
β-carotene hydroxylase
- DOXP
1-deoxy-d-xylulose 5-phosphate
- GGPP
Geranylgeranyl pyrophosphate
- HPLC
High-performance liquid chromatography
- IPTG
Isopropyl-β-d-thiogalactopyranoside
- MEP
2-C-methylerythritol 4-phosphate
- MEV
Mevalonate
Additional files
Nucleotide sequence of E. gracilis crtE and its deduced amino acid sequence. (PDF 1127 kb)
Nucleotide sequence of E. gracilis crtB and its deduced amino acid sequence. (PDF 1130 kb)
Phylogenetic relationships of the deduced EgCrtE amino acid sequence and known GGPP synthases. Numbers in parentheses are accession numbers of GGPP synthases. Sequence data for GGPS of Pyropia umbilicalis [P_umbilicalis_esContig5139] was obtained from NoriBLAST [58]. The phylogenetic tree was constructed with the neighbor-joining method using MEGA version 6.0 [59]. Bootstrap values from the percentages of 1000 replications are indicated beside each node. (PDF 972 kb)
Phylogenetic relationships of the deduced EgCrtB amino acid sequence and known phytoene synthases. Numbers in parentheses are accession numbers of phytoene synthases. The phylogenetic tree was constructed with the neighbor-joining method using MEGA version 6.0 [59]. Bootstrap values from the percentages of 1000 replications are indicated beside each node. (PDF 988 kb)
Analysis of phytoene production in E. coli by HPLC. HPLC chromatogram (284 nm) of extracts from E. coli cells carrying (A) pET-EgcrtE, (B) pET-EgcrtB, (C) pAC-PacrtB with pETDuet-1 (vector control), and (D) pACCRT-E [23] with pETDuet-1. Data are representative of three or four experiments with similar results. Phytoene was eluted at 28.6 min (Fig. 4). The peak at 21.5 min was not carotenoid. (PDF 1108 kb)
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
SK designed the experiments and conducted the algal culture, cDNA cloning, HPLC, and gene expression analyses; and drafted the manuscript. TI provided the RNA-seq data including unpublished observations. MA cooperated with SK in the molecular genetic studies including the cDNA cloning and gene expression analyses. SK and ShT performed phylogenetic analyses of GGPP synthase and phytoene synthase proteins. ShT and SeT established the analysis method of carotenoids in E. coli cells for the functional analysis of EgcrtE and EgcrtB with HPLC in cooperation with SK. TS conceived of the study, and participated in its design and coordination; and helped to draft the manuscript. All authors read and approved the final manuscript.
Authors’ information
Shota Kato Plant Molecular and Cellular Biology Laboratory, Department of Biosciences, School of Science and Engineering, Teikyo University, 1–1 Toyosatodai, Utsunomiya, Tochigi, 320–8551, Japan. Phone: +81-28-627-7111 Email: shota.kato.680@gmail.com
Contributor Information
Shota Kato, Email: shota.kato.680@gmail.com.
Shinichi Takaichi, Email: takaichi@nms.ac.jp.
Takahiro Ishikawa, Email: ishikawa@life.shimane-u.ac.jp.
Masashi Asahina, Email: asahina@nasu.bio.teikyo-u.ac.jp.
Senji Takahashi, Email: takahasi@nasu.bio.teikyo-u.ac.jp.
Tomoko Shinomura, Phone: +81-28-627-7724, Email: shinomura@nasu.bio.teikyo-u.ac.jp.
References
- 1.Gibbs SP. The chloroplasts of Euglena may have evolved from symbiotic green algae. Can J Bot. 1978;56:2883–9. doi: 10.1139/b78-345. [DOI] [Google Scholar]
- 2.Durnford DG, Deane JA, Tan S, McFadden GI, Gantt E, Green BR. A phylogenetic assessment of the eukaryotic light-harvesting antenna proteins, with implications for plastid evolution. J Mol Evol. 1999;48:59–68. doi: 10.1007/PL00006445. [DOI] [PubMed] [Google Scholar]
- 3.Sandmann G. Molecular evolution of carotenoid biosynthesis from bacteria to plants. Physiol Plant. 2002;116:431–40. doi: 10.1034/j.1399-3054.2002.1160401.x. [DOI] [Google Scholar]
- 4.Havaux M. Carotenoids as membrane stabilizers in chloroplasts. Trends Plant Sci. 1998;3:147–51. doi: 10.1016/S1360-1385(98)01200-X. [DOI] [Google Scholar]
- 5.Demmig-Adams B. Carotenoids and photoprotection in plants: a role for the xanthophyll zeaxanthin. Biochim Biophys Acta. 1990;1020:1–24. doi: 10.1016/0005-2728(90)90088-L. [DOI] [Google Scholar]
- 6.Seo M, Koshiba T. Complex regulation of ABA biosynthesis in plants. Trends Plant Sci. 2002;7:41–8. doi: 10.1016/S1360-1385(01)02187-2. [DOI] [PubMed] [Google Scholar]
- 7.Matusova R, Rani K, Verstappen FWA, Franssen MCR, Beale MH, Bouwmeester HJ. The strigolactone germination stimulants of the plant-parasitic Striga and Orobanche spp. are derived from the carotenoid pathway. Plant Physiol. 2005;139:920–34. doi: 10.1104/pp.105.061382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Takaichi S. Carotenoids in algae: distributions, biosyntheses and functions. Mar Drugs. 2011;9:1101–18. doi: 10.3390/md9061101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cordero BF, Couso I, León R, Rodríguez H, Vargas MÁ. Enhancement of carotenoids biosynthesis in Chlamydomonas reinhardtii by nuclear transformation using a phytoene synthase gene isolated from Chlorella zofingiensis. Appl Microbiol Biotechnol. 2011;91:341–51. doi: 10.1007/s00253-011-3262-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yang LE, Huang XQ, Lu QQ, Zhu JY, Lu S. Cloning and characterization of the geranylgeranyl diphosphate synthase (GGPS) responsible for carotenoid biosynthesis in Pyropia umbilicalis. J Appl Phycol. 2015. doi:10.1007/s10811-015-0593-6.
- 11.Aitzetmüller K, Svec WA, Katz JJ, Strain HH. Structure and chemical identity of diadinoxanthin and the principal xanthophyll of Euglena. Chem Commun. 1968;1:32–3.
- 12.Bjørnland T. Chlorophylls and carotenoids of the marine alga Eutreptiella gymnastica. Phytochem. 1982;21:1715–9. doi: 10.1016/S0031-9422(82)85046-2. [DOI] [Google Scholar]
- 13.Takaichi S, Mimuro M. Distribution and geometric isomerism of neoxanthin in oxygenic phototrophs: 9′-cis, a sole molecular form. Plant Cell Physiol. 1998;39:968–77. doi: 10.1093/oxfordjournals.pcp.a029461. [DOI] [Google Scholar]
- 14.Tessier LH, Keller M, Chan RL, Fournier R, Weil JH, Imbault P. Short leader sequences may be transferred from small RNAs to pre-mature mRNAs by trans-splicing in Euglena. EMBO J. 1991;10:2621–5. doi: 10.1002/j.1460-2075.1991.tb07804.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Durnford DG, Gray MW. Analysis of Euglena gracilis plastid-targeted proteins reveals different classes of transit sequences. Eukaryot Cell. 2006;5:2079–91. doi: 10.1128/EC.00222-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.TMHMM Server, v. 2.0. http://www.cbs.dtu.dk/services/TMHMM/. Accessed 17 June 2015.
- 17.TargetP 1.1 Server. http://www.cbs.dtu.dk/services/TargetP/. Accessed 17 June 2015.
- 18.McWilliam H, Li W, Uludag M, Squizzato S, Park YM, Buso N, et al. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 2013;41:W597–600. doi: 10.1093/nar/gkt376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bouvier F, Rahier A, Camara B. Biogenesis, molecular regulation and function of plant isoprenoids. Prog Lipid Res. 2005;44:357–429. doi: 10.1016/j.plipres.2005.09.003. [DOI] [PubMed] [Google Scholar]
- 20.Beck G, Coman D, Herren E, Ruiz-Sola MÁ, Rodríguez-Concepción M, Gruissem W, et al. Characterization of the GGPP synthase gene family in Arabidopsis thaliana. Plant Mol Biol. 2013;82:393–416. doi: 10.1007/s11103-013-0070-z. [DOI] [PubMed] [Google Scholar]
- 21.Shumskaya M, Bradbury LMT, Monaco RR, Wurtzel ET. Plastid localization of the key carotenoid enzyme phytoene synthase is altered by isozyme, allelic variation, and activity. Plant Cell. 2012;24:3725–41. doi: 10.1105/tpc.112.104174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Misawa N, Nakagawa M, Kobayashi K, Yamano S, Izawa Y, Nakamura K, et al. Elucidation of the Erwinia uredovora carotenoid biosynthetic pathway by functional analysis of gene products expressed in Escherichia coli. J Bacteriol. 1990;172:6704–12. doi: 10.1128/jb.172.12.6704-6712.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chamovitz D, Misawa N, Sandmann G, Hirschberg J. Molecular cloning and expression in Escherichia coli of a cyanobacterial gene coding for phytoene synthase, a carotenoid biosynthesis enzyme. FEBS Lett. 1992;296:305–10. doi: 10.1016/0014-5793(92)80310-D. [DOI] [PubMed] [Google Scholar]
- 24.Kuntz M, Römer S, Suire C, Hugueney P, Weil JH, Schantz R, et al. Identification of a cDNA for the plastid-located geranylgeranyl pyrophosphate synthase from Capsicum annuum: correlative increase in enzyme activity and transcript level during fruit ripening. Plant J. 1992;2:25–34. doi: 10.1111/j.1365-313x.1992.00025.x. [DOI] [PubMed] [Google Scholar]
- 25.Lichtenthaler HK. The 1-deoxy-d-xylulose-5-phosphate pathway of isoprenoid biosynthesis in plants. Annu Rev Plant Physiol Plant Mol Biol. 1999;50:47–65. doi: 10.1146/annurev.arplant.50.1.47. [DOI] [PubMed] [Google Scholar]
- 26.Lichtenthaler HK. Biosynthesis, accumulation and emission of carotenoids, α-tocopherol, plastoquinone, and isoprene in leaves under high photosynthetic irradiance. Photosynth Res. 2007;92:163–79. doi: 10.1007/s11120-007-9204-y. [DOI] [PubMed] [Google Scholar]
- 27.Disch A, Schwender J, Müller C, Lichtenthaler HK, Rohmer M. Distribution of the mevalonate and glyceraldehyde phosphate/pyruvate pathways for isoprenoid biosynthesis in unicellular algae and the cyanobacterium Synechocystis PCC 6714. Biochem J. 1998;333:381–8. doi: 10.1042/bj3330381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sláviková S, Vacula R, Fang Z, Ehara T, Osafune T, Schwartzbach SD. Homologous and heterologous reconstitution of Golgi to chloroplast transport and protein import into the complex chloroplasts of Euglena. J Cell Sci. 2005;118:1651–61. doi: 10.1242/jcs.02277. [DOI] [PubMed] [Google Scholar]
- 29.Kreimer G. The green algal eyespot apparatus: a primordial visual system and more? Curr Genet. 2009;55:19–43. doi: 10.1007/s00294-008-0224-8. [DOI] [PubMed] [Google Scholar]
- 30.Schmidt M, Geßner G, Luff M, Heiland I, Wagner V, Kaminski M, et al. Proteomic analysis of the eyespot of Chlamydomonas reinhardtii provides novel insights into its components and tactic movements. Plant Cell. 2006;18:1908–30. doi: 10.1105/tpc.106.041749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Davidi L, Levin Y, Ben-Dor S, Pick U. Proteome analysis of cytoplasmatic and plastidic β-carotene lipid droplets in Dunaliella bardawil. Plant Physiol. 2015;167:60–79. doi: 10.1104/pp.114.248450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Heelis DV, Kernick W, Phillips GO, Davies K. Separation and identification of the carotenoid pigments of stigmata isolated from light-grown cells of Euglena gracilis strain Z. Arch Microbiol. 1979;121:207–11. doi: 10.1007/BF00425057. [DOI] [PubMed] [Google Scholar]
- 33.Kivic PA, Vesk M. Structure and function in the Euglenoid eyespot apparatus: the fine structure, and response to environmental changes. Planta. 1972;105:1–14. doi: 10.1007/BF00385158. [DOI] [PubMed] [Google Scholar]
- 34.Adl SM, Simpson AGB, Farmer MA, Andersen RA, Anderson OR, Barta JR, et al. The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J Eukaryot Microbiol. 2005;52:399–451. doi: 10.1111/j.1550-7408.2005.00053.x. [DOI] [PubMed] [Google Scholar]
- 35.Ahmadinejad N, Dagan T, Martin W. Genome history in the symbiotic hybrid Euglena gracilis. Gene. 2007;402:35–9. doi: 10.1016/j.gene.2007.07.023. [DOI] [PubMed] [Google Scholar]
- 36.Simpson AGB, Gill EE, Callahan HA, Litaker RW, Roger AJ. Early evolution within Kinetoplastids (Euglenozoa), and the late emergence of Trypanosomatids. Protist. 2004;155:407–22. doi: 10.1078/1434461042650389. [DOI] [PubMed] [Google Scholar]
- 37.Sogin ML. Early evolution and the origin of eukaryotes. Curr Opin Genet Dev. 1991;1:457–63. doi: 10.1016/S0959-437X(05)80192-3. [DOI] [PubMed] [Google Scholar]
- 38.Watanabe Y, Gray MW. Evolutionary appearance of genes encoding proteins associated with box H/ACA snoRNAs: Cbf5p in Euglena gracilis, an early diverging eukaryote, and candidate Gar1p and Nop10p homologs in archaebacteria. Nucleic Acids Res. 2000;28:2342–52. doi: 10.1093/nar/28.12.2342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Steinbrenner J, Linden H. Light induction of carotenoid biosynthesis genes in the green alga Haematococcus pluvialis: regulation by photosynthetic redox control. Plant Mol Biol. 2003;52:343–56. doi: 10.1023/A:1023948929665. [DOI] [PubMed] [Google Scholar]
- 40.Wahidin S, Idris A, Shaleh SRM. The influence of light intensity and photoperiod on the growth and lipid content of microalgae Nannochloropsis sp. Bioresour Technol. 2013;129:7–11. doi: 10.1016/j.biortech.2012.11.032. [DOI] [PubMed] [Google Scholar]
- 41.Król M, Maxwell DP, Huner NPA. Exposure of Dunaliella salina to low temperature mimics the high light-induced accumulation of carotenoids and the carotenoid binding protein (Cbr) Plant Cell Physiol. 1997;38:213–6. doi: 10.1093/oxfordjournals.pcp.a029155. [DOI] [Google Scholar]
- 42.Steinbrenner J, Linden H. Regulation of two carotenoid biosynthesis genes coding for phytoene synthase and carotenoid hydroxylase during stress-induced astaxanthin formation in the green alga Haematococcus pluvialis. Plant Physiol. 2001;125:810–7. doi: 10.1104/pp.125.2.810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Li Y, Huang J, Sandmann G, Chen F. High-light and sodium chloride stress differentially regulate the biosynthesis of astaxanthin in Chlorella zofingiensis (Chlorophyceae) J Phycol. 2009;45:635–41. doi: 10.1111/j.1529-8817.2009.00689.x. [DOI] [PubMed] [Google Scholar]
- 44.Wang B, Zarka A, Trebst A, Boussiba S. Astaxanthin accumulation in Haematococcus pluvialis (Chlorophyceae) as an active photoprotective process under high irradiance. J Phycol. 2003;39:1116–24. doi: 10.1111/j.0022-3646.2003.03-043.x. [DOI] [Google Scholar]
- 45.von Lintig J, Welsch R, Bonk M, Giuliano G, Batschauer A, Kleinig H. Light-dependent regulation of carotenoid biosynthesis occurs at the level of phytoene synthase expression and is mediated by phytochrome in Sinapsis alba and Arabidopsis thaliana seedlings. Plant J. 1997;12:625–34. doi: 10.1046/j.1365-313X.1997.00625.x. [DOI] [PubMed] [Google Scholar]
- 46.Estévez JM, Cantero A, Reindl A, Reichler S, León P. 1-Deoxy-d-xylulose-5-phosphate synthase, a limiting enzyme for plastidic isoprenoid biosynthesis in plants. J Biol Chem. 2001;276:22901–9. doi: 10.1074/jbc.M100854200. [DOI] [PubMed] [Google Scholar]
- 47.Carretero-Paulet L, Cairó A, Botella-Pavía P, Besumbes O, Campos N, Boronat A, et al. Enhanced flux through the methylerythritol 4-phosphate pathway in Arabidopsis plants overexpressing deoxyxylulose 5-phosphate reductoisomerase. Plant Mol Biol. 2006;62:683–95. doi: 10.1007/s11103-006-9051-9. [DOI] [PubMed] [Google Scholar]
- 48.Botella-Pavía P, Besumbes Ó, Phillips MA, Carretero-Paulet L, Boronat A, Rodríguez-Concepción M. Regulation of carotenoid biosynthesis in plants: evidence for a key role of hydroxymethylbutenyl diphosphate reductase in controlling the supply of plastidial isoprenoid precursors. Plant J. 2004;40:188–99. doi: 10.1111/j.1365-313X.2004.02198.x. [DOI] [PubMed] [Google Scholar]
- 49.Rodríguez-Villalón A, Gas E, Rodríguez-Concepción M. Phytoene synthase activity controls the biosynthesis of carotenoids and the supply of their metabolic precursors in dark-grown Arabidopsis seedlings. Plant J. 2009;60:424–35. doi: 10.1111/j.1365-313X.2009.03966.x. [DOI] [PubMed] [Google Scholar]
- 50.Eilers U, Bikoulis A, Breitenbach J, Büchel C, Sandmann G. Limitations in the biosynthesis of fucoxanthin as targets for genetic engineering in Phaeodactylum tricornutum. J Appl Phycol. 2015. doi:10.1007/s10811-015-0583-8.
- 51.Toledo-Ortiz G, Huq E, Rodríguez-Concepción M. Direct regulation of phytoene synthase gene expression and carotenoid biosynthesis by phytochrome-interacting factors. Proc Natl Acad Sci. 2010;107:11626–31. doi: 10.1073/pnas.0914428107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bohne F, Linden H. Regulation of carotenoid biosynthesis genes in response to light in Chlamydomonas reinhardtii. Biochim Biophys Acta. 2002;1579:26–34. doi: 10.1016/S0167-4781(02)00500-6. [DOI] [PubMed] [Google Scholar]
- 53.Cramer M, Myers J. Growth and photosynthetic characteristics of Euglena gracilis. Arch Mikrobiol. 1952;17:384–402. doi: 10.1007/BF00410835. [DOI] [Google Scholar]
- 54.Bertani G. Studies on lysogenesis I.: The mode of phage liberation by lysogenic Escherichia coli. J Bacteriol. 1951;62:293–300. doi: 10.1128/jb.62.3.293-300.1951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Inoue H, Nojima H, Okayama H. High efficiency transformation of Escherichia coli with plasmids. Gene. 1990;96:23–8. doi: 10.1016/0378-1119(90)90336-P. [DOI] [PubMed] [Google Scholar]
- 56.Takemura M, Maoka T, Misawa N. Carotenoid analysis of a liverwort Marchantia polymorpha and functional identification of its lycopene β- and ε-cyclase genes. Plant Cell Physiol. 2014;55:194–200. doi: 10.1093/pcp/pct170. [DOI] [PubMed] [Google Scholar]
- 57.Takaichi S. Characterization of carotenes in a combination of a C18 HPLC column with isocratic elution and absorption spectra with a photodiode-array detector. Photosynthesis Res. 2000;65:93–9. doi: 10.1023/A:1006445503030. [DOI] [PubMed] [Google Scholar]
- 58.NoriBLAST. http://dbdata.rutgers.edu/nori/index.php. Accessed 20 November 2015.
- 59.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]


